Abstract
The establishment of left–right (LR) asymmetry is fundamental to animal development, but the identification of a unifying mechanism establishing laterality across different phyla has remained elusive. A cilia-driven, directional fluid flow is important for symmetry breaking in numerous vertebrates, including zebrafish. Alternatively, LR asymmetry can be established independently of cilia, notably through the intrinsic chirality of the acto-myosin cytoskeleton. Here, we show that Myosin1D (Myo1D), a previously identified regulator of Drosophila LR asymmetry, is essential for the formation and function of the zebrafish LR organizer (LRO), Kupffer’s vesicle (KV). Myo1D controls the orientation of LRO cilia and interacts functionally with the planar cell polarity (PCP) pathway component VanGogh-like2 (Vangl2), to shape a productive LRO flow. Our findings identify Myo1D as an evolutionarily conserved regulator of animal LR asymmetry, and show that functional interactions between Myo1D and PCP are central to the establishment of animal LR asymmetry.
Similar content being viewed by others
Introduction
The molecular pathways governing antero-posterior and dorsoventral patterning have been extensively conserved throughout evolution. In contrast, numerous studies have revealed a striking diversity in the strategies establishing laterality both within and across phyla, making the identification of a unifying mechanism elusive1,2,3,4. Processes that have been involved in symmetry breaking range from cilia-driven fluid flows5,6,7 and localized ion flows8,9,10 to cellular rearrangements dependent on cytoskeletal polarity11,12,13,14. Body laterality can even be subject to distinct regulations in a single organism15,16. This apparent diversity raises the question whether a unifying mechanism underlying the establishment of LR asymmetry still remains to be identified?
An attractive hypothesis is that the chirality of actin filaments may provide a template for LR asymmetry1,17. Accordingly, actin-binding proteins govern molecular18 and cellular chirality19 and actin-dependent processes regulate invertebrate and vertebrate laterality1,15,20,21,22,23. Studies in Drosophila have identified the actin-based molecular motor protein Myosin1D (Myo1D, a.k.a. Myosin31DF) as an essential regulator of LR asymmetry20,21. LR asymmetry in Drosophila governs the dextral rotation of male genitalia, as well as the looping of testis, midgut and hindgut. myo1D mutants show a LR inversion of all lateralized organs, identifying Myo1D as a key regulator of dextral development24. In the adult hindgut, functional interactions of Myo1D with planar cell polarity (PCP) pathway components govern cellular chirality and promote asymmetric gut looping13. Here we study the contribution of Myo1D to vertebrate laterality, by analyzing its function in zebrafish.
The zebrafish LR organizer (LRO), Kupffer’s vesicle (KV), is a transient vesicular organ that is decorated on its inside with motile cilia whose beating creates a counter-clockwise fluid flow essential for LR asymmetry6. This directional ciliary flow has been proposed to promote symmetry breaking through the local activation of mechanosensory Ca2+ channels such as PKD2 or the lateralized transport of signaling molecules in the extracellular space8,25,26. While the mechanism through which the flow contributes to chiral morphogenesis remains debated, it is firmly established that a directional LRO flow is essential for zebrafish LR asymmetry6,25,26. The generation of a functional LRO flow requires both polarized KV cell shape remodeling14,27, as well as a PCP-dependent control of the spatial orientation of flow-generating motile cilia28.
In the present study, we provide genetic evidence that myo1D is essential for the establishment of zebrafish LR asymmetry. Myo1D controls the morphogenesis and function of the zebrafish LRO by interacting functionally with the PCP pathway component VanGogh-like2 (Vangl2). KV cilia orientation is controlled by opposing activities of Myo1D and Vangl2, the balance of which is essential for the establishment of a directional KV flow and subsequent chiral morphogenesis. Taken together, our findings identify Myo1D as a common regulator of LR asymmetry whose function is conserved in both vertebrate and invertebrate organisms.
Results
Myo1D is essential for zebrafish left–right (LR) asymmetry
We first determined whether zebrafish and Drosophila Myo1D have conserved activities by testing the ability of fish myo1D to rescue the laterality defects of fly myo1D mutants. The two orthologous proteins show 69% sequence similarity. Expressing zebrafish myo1D in Drosophila fully restored the chirality of genitalia rotation (Fig. 1a), a prominent LR marker in Drosophila20,21. These data indicate that zebrafish Myo1D is capable of mediating a conserved LR symmetry-breaking activity.
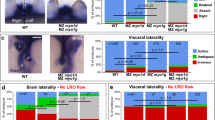
myo1D controls zebrafish left–right asymmetry. a Transgene-mediated expression of zebrafish myo1D (zmyo1D) restores the laterality of genitalia rotation in Drosophila myo1D (dmyo1D) mutant or dmyo1D RNAi flies. Lines 3 and 6 are two independent transgenic insertions. b–e Zebrafish MZ myo1D mutants present defects in leftward cardiac jogging. b–d Dorsal views of the cmlc2-expressing heart primordium at 26 hpf, anterior up. e Quantification of cardiac jogging defects in maternal (M) and maternal zygotic (MZ) myo1Dtj16b or myo1Dtj16c mutants. f–i Cardiac looping is affected in MZ myo1D mutants. f–h Frontal views of the cmlc2-expressing heart at 48 hpf, dorsal up. i Quantification of cardiac looping defects in M myo1Dtj16b and MZ myo1Dtj16b or MZ myo1Dtj16c mutants. j–m MZ myo1D mutants present defects in the leftward looping of the gut and the asymmetric development of the liver (black arrows) and pancreas (white arrows). j–l Dorsal views of foxa1-expressing visceral organs at 48 hpf, anterior up. n, o Overexpression of zebrafish myo1Cb RNA impairs cardiac jogging (n) and looping (o). MZ myo1G single mutants present laterality defects (n, o). MZ myo1D; MZ myo1G double mutants or MZ myo1D mutants injected with myo1Cb RNA present an increased frequency of defects compared to MZ myo1D mutants (n, o). Scale bars: 30 µm
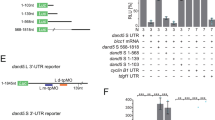
To study zebrafish myo1D function, we generated frameshift mutations that disrupt the P-loop required for Myo1D motor activity29 and delete the actin- and cargo-binding domains (Supplementary Fig. 1a). Homozygous mutants lacking zygotic myo1D function develop normally and give rise to fertile adults (Supplementary Fig. 1b). In situ hybridization reveals that, in addition to their zygotic expression, myo1D transcripts are maternally provided (Supplementary Fig. 1c–e). We therefore used homozygous mutant females to generate embryos lacking either only maternal (M) or both maternal and zygotic (MZ) myo1D activities. M and MZ myo1D mutants display LR asymmetry defects at the level of the heart (Fig. 1b–i), viscera (Fig. 1j–m), and brain (Supplementary Fig. 2a–f). Similar phenotypes were observed using two different mutant alleles (Fig. 1e, i). While LR defects of MZ myo1D mutant animals could be partially rescued through the injection of an RNA encoding wild-type (WT) Myo1D protein, no rescuing activity was observed using myo1D mutant RNA (Supplementary Fig. 2g–k). Similar defects are moreover generated when a morpholino that blocks the translation of both maternally deposited and zygotically transcribed myo1D RNA is injected in WT embryos (Supplementary Fig. 3). Altogether, our experiments identify Myo1D as a conserved regulator of laterality in both vertebrates and invertebrates.
In Drosophila, Myosin1C (Myo1C, a.k.a. Myo61F) acts as an antagonist of the essential dextral regulator Myo1D30, with myo1C overexpression phenocopying myo1D loss of function. However, Drosophila myo1C is itself dispensable for LR asymmetry30. The zebrafish genome encodes two myo1C homologues, myosin1Ca and myosin1Cb (myo1Ca & myo1Cb). Like in Drosophila, zygotic myo1Ca mutants and MZ myo1Cb mutants do not display LR defects (not shown). Conversely, Myo1Cb overexpression using mRNA microinjection causes cardiac laterality defects (Fig. 1n, o), thus mimicking myo1D loss of function, as observed in Drosophila30.
Overexpression of zebrafish myo1Cb enhances MZ myo1D mutant LR defects (Fig. 1n, o), raising the question whether additional Myo1D-like activities might contribute to the regulation of LR asymmetry. Genome analysis shows that zebrafish Myosin1G (Myo1G) is closely related to Myo1D (79% similar amino acids). To invalidate myo1G, we used two mutant alleles that introduce premature stop codons at different positions of the open reading frame (see Methods). Zygotic myo1G single mutants developed normally into fertile adults, but MZ myo1G mutant embryos displayed cardiac laterality defects (Fig. 1n, o). MZ myo1D; MZ myo1G double mutants were created through Crispr/Cas9 mutagenesis of the myo1G locus in a myo1D mutant background (see Methods). MZ myo1D; MZ myo1G double mutants presented an increased frequency of LR defects compared to myo1D single mutants, similar to the one observed upon myo1Cb overexpression (Fig. 1n, o). Our findings provide evidence that Myo1D-like agonists (Myo1D & G) and their antagonist Myo1Cb form an evolutionarily conserved pathway for the control of LR asymmetry.
Myo1D is required for LRO function
The observation that MZ myo1D mutants present laterality defects at the level of the heart, viscera, and brain (Fig. 1b–m, Supplementary Fig. 2a–f), suggests that myo1D may control the formation and/or function of the central zebrafish LRO, KV6,31. The ciliary LRO flow induces the expression of both the nodal ligand southpaw (spaw) and the transcription factor pitx2 in the left lateral plate mesodem32,33. Consistent with a potential role of Myo1D at the level of the LRO, the asymmetric expression of these two genes is disrupted in MZ myo1D mutants (Fig. 2a–f). To further test Myo1D function in the LRO, we looked at the asymmetric expression of the TGFβ antagonist charon/dand5, which is the earliest known transcriptional response to the KV flow25,34. While weak symmetric dand5 expression is initiated prior to the establishment of a functional KV, dand5 asymmetry arises as the LRO matures, peaks by the eight-somite stage and decreases again during later development (Fig. 2g–m). In accordance with a dysfunction of the LRO, dand5 asymmetry is reduced in MZ myo1D mutants (Fig. 2n, o, q). These data demonstrate that Myo1D is required prior to the first morphological manifestations of LR asymmetry for the LRO-dependent establishment of asymmetric gene expressions.
myo1D is required for left–right Organizer function. a–f MZ myo1D mutants display an impaired expression of southpaw (spaw) and pitx2 in the left lateral plate mesoderm. c, f Quantifications of spaw (c) and pitx2 (f) expression. For pitx2, the most marked difference between WT and MZ myo1D mutants is observed in the category with right-sided expression (e). g–m Temporal evolution of the asymmetry of dand5 expression between the end of gastrulation (Bud) and the 14-somite stage. m Quantifications of the relative abundance of dand5 transcripts on the left (L) and right (R) side of WT ABTÜ embryos reveal a peak of LR asymmetry at the eight-somite stage. n–q Eight-somite stage asymmetry of dand5 asymmetry is reduced in MZ myo1D mutants. No further reduction of asymmetry is observed in MZ myo1D; MZ myo1G double mutants. For values of dand5 asymmetry for individual display items, see Methods. r–t Asymmetric spaw expression is altered in MZ myo1D; MZ myo1G double mutants. The frequency of defects is comparable to MZ myo1D single mutants (c). a, b, d, e, r, s are dorsal views of 18-somite stage embryos, anterior up. g–l and n–p are dorsal views of the LRO, anterior up. Error bars in m and q indicate SEM. Scale bars: 50 µm in a, b, d, e, r, s; 20 µm in g–l and n–p
Myo1D controls the ciliary LRO flow
To further dissect the role of myo1D in zebrafish LR asymmetry, we analyzed its contribution to the formation and function of the KV. While myo1D is dispensable for the migration and coalescence of sox17-positive KV precursor cells (Supplementary Fig. 4), we found that the average KV size and the total number of cilia are reduced in MZ myo1D mutants (Fig. 3a, b, e, g–i). LRO function requires a minimal organ size and number of motile cilia25,35. We monitored KV size, total, and motile cilia number together with laterality in individual embryos. No correlation could be observed between organ size, cilia numbers, and the appearance of laterality defects (Fig. 3e, k, Supplementary Fig. 5). KV lumen inflation is promoted through the activation of the cystic fibrosis transmembrane conductance regulator (CFTR) channel36. Accordingly, treatment with IBMX and Forskolin, which activate CFTR through an elevation of cAMP levels and PKA activity, rescues KV size but not laterality defects in MZ myo1D mutants (Fig. 3c–f). While the loss of myo1D causes a modest reduction of cilia size (Fig. 3g, h, j), it has no incidence on the percentage of motile cilia (Fig. 3l). Previous studies have revealed that the majority of cilia are positioned in the anterior KV half37. This antero-posterior asymmetry is unaffected in MZ myo1D (Fig. 3m).
myo1D controls the morphogenesis of the zebrafish left–right Organizer. a–e Compared to WT (n = 104), MZ myo1D mutants (n = 118) present a reduced KV size. Treatment with IBMX and Forskolin promotes KV lumen inflation and increases organ size in WT (n = 66) and MZ myo1D mutants (n = 85). e Dot plots of KV equatorial surface area in individual embryos. KV size is similar in embryos with normal or defective laterality. f IBMX/Forskolin treatment restores KV size (d, e), but not laterality in MZ myo1D mutants. g–j Cilia number and size are reduced in MZ myo1D mutants. g, h Projection of images from confocal stacks used to quantify number and length of cilia (acetylated tubulin, magenta) in the KV (ZO-1 positive cells, green). i, j Number and average length of cilia in individual embryos are reduced in MZ myo1D (n = 117 embryos/4939 cilia) compared to WT (n = 141/6967). k–m Confocal imaging of Arl13b-GFP labeled cilia in living embryos was used to analyze the number (k), motility (l) and positioning (m) of KV cilia. k Dot plot representing motile cilia number in individual embryos. MZ myo1D (n = 42) mutants display lower motile cilia numbers compared to WT (n = 33). Motile cilia numbers are however similar in MZ myo1D mutant embryos with normal or defective laterality. Total cilia numbers for this experiment are displayed in Supplementary Fig. 5. l myo1D loss of function does not impair ciliary motility. m WT and MZ myo1D mutant animals display a similar enrichment of cilia in the anterior KV half. a–d, g, h are dorsal views of the KV, anterior up. All data collected at the eight-somites stage. Horizontal bars in e, i, j, k represent mean values. Error bars in l, m represent SEM. Scale bars: 20 µm in a–d; 10 µm in g–h
To visualize the LRO flow, we tracked fluorescent microspheres in the KV lumen. While WT controls mostly displayed circular trajectories (Fig. 4a), MZ myo1D mutants often presented aberrant flow patterns (Fig. 4b). To quantitatively assess the LRO flow, we determined its mean angular velocity as a measure of the effective circular flow strength (see Methods). MZ myo1D mutants display a decrease in angular flow velocity (Fig. 4c). Importantly, LR asymmetry defects in MZ myo1D mutants correlate with pronounced reductions in flow velocity (Fig. 4c). In contrast, treatment of control WT embryos with Ouabain to pharmacologically reduce KV size leads to an increase in flow velocity with no associated LR defects (Supplementary Fig. 6). These data show that myo1D is required for the establishment of a functional LRO flow, independently of KV size.
myo1D is required for the establishment of zebrafish left–right organizer flow. a–c KV flow is altered in MZ myo1D mutants. Temporal projections of the trajectories of fluorescent microspheres in WT (a, n = 28) and MZ myo1D mutants (b, n = 63). c MZ myo1D embryos display lower mean angular KV flow velocities than WT. MZ myo1D mutants with defective cardiac jogging show lower velocities than their sibling with normal laterality. d–g MZ myo1D (e) and vangl2 (f, n = 36) mutants present an altered LRO flow compared to WT (d) but flow pattern is restored in MZ myo1D ; vangl2+/− (g, n = 25). h MZ myo1D mutants display a reduced KV flow velocity compared to WT, as do vangl2−/− embryos (p = 0.03). Flow in MZ myo1D ; vangl2+/− is however similar to WT. Flow velocity in MZ myo1D; MZ myo1G double mutants is reduced compared to WT (n = 29, p = 0.04) but similar to MZ myo1D single mutants. Flow velocity in WT embryos injected with myo1Cb RNA is reduced compared to WT controls (n = 37, p = 0.04). Removal of one copy of vangl2 makes flow velocity again similar to WT (n = 19, p = 0.62). i, j The removal of one copy of vangl2 reduces defects in asymmetric spaw expression (i) and cardiac morphogenesis (j) in MZ myo1D (see Methods). a, b, d–g are dorsal views of eight-somites stage KVs, anterior up. Display items and quantifications in a–h are derived from the same dataset. Horizontal bars in c represent mean values. Error bars in h indicate SEM. Scale bars: 20 µm in a, b, d–g
Myo1D interacts with Vangl2 to control LR asymmetry
How does myo1D control KV flow? Embryos with no, or immotile, cilia lack a detectable flow6,25,38 and show highly penetrant LR asymmetry phenotypes (Supplementary Fig. 7). In MZ myo1D mutants, cilia are motile but the LRO flow is erratic, a phenotype reminiscent of mutants for the PCP pathway component vangl2, which displays less penetrant LR defects (Fig. 4d–f, Supplementary Fig. 7)28,39. We found that both MZ myo1D and vangl2 mutants display a reduction in overall angular velocity and a disruption of the KV flow pattern (Fig. 4h, Supplementary Fig. 8a). While myo1D and vangl2 mutants have similar flow phenotypes, the removal of one copy of vangl2 partially rescues the KV flow defects of MZ myo1D embryos (Fig. 4g, h, Supplementary Fig. 8a), suggesting that the two genes affect the LRO in opposite ways. In accordance with the restoration of a functional KV flow, the removal of one copy of vangl2 decreases the penetrance of LR asymmetry defects in myo1D-deficient animals (Fig. 4i, j).
A reduction of overall KV angular flow velocity and a disruption of circular flow geometry are also observed in MZ myo1D; MZ Myo1G double mutants or embryos injected with myo1Cb antagonist RNA (Fig. 4h, Supplementary Fig.8a–d), confirming the importance of the Myosin1 pathway for the control of LRO flow. Removal of one copy of vangl2 allows to partially restore the KV flow defects that are elicited by myoICb overexpression (Fig. 4h, Supplementary Fig. 8a, d, e), further supporting the existence of functionally relevant Myo1/PCP interactions. Our velocity measurements do not allow to explain the enhanced LR asymmetry defects of MZ myo1D; MZ Myo1G double mutants or myo1Cb-injected animals compared to MZ myo1D single mutants (Fig. 1n, o). This may be due to the fact that these measurements do not encompass the full complexity of the observed flow phenotypes. Alternatively, zebrafish Myosin1 proteins could act at different steps of the chiral morphogenesis program, in addition to their importance for LRO flow. In accordance with this hypothesis, similar defects are observed in MZ myo1D single and MZ myo1D ; MZ myo1G double mutants for the LRO-dependent expression of dand5 and spaw (Fig. 2a–c, n–t). This situation is consistent with the known role of Drosophila myo1D, which controls the morphogenesis of different organs through distinct cellular mechanisms13,21,30.
How do myo1D and vangl2 exert opposite effects on LRO morphogenesis and function? In accordance with our previous conclusion that the reduction of KV size is not the primary cause of MZ myo1D LR asymmetry defects (Fig. 3e), removal of one copy of vangl2 has no significant effect on the KV size of MZ myo1D mutant animals (Fig. 5a–c). Similarly, reducing vangl2 gene dosage does not restore the reduction of cilia length that is observed in MZ myo1D mutants (Figs. 3j and 5d–f).
KV area and cilia size are similar in MZ myo1D and MZ myo1D vangl2+/−. a–c MZ myo1D (n = 156) and MZ myo1D ; vangl2+/− (n = 60) animals have similar KV sizes. d–e Projection of images from confocal stacks used to quantify the length of cilia (acetylated tubulin, magenta) in the KV (ZO-1 positive cells, green) of MZ myo1D mutants (d), or MZ myo1D ; vangl2+/− embryos (e). f MZ myo1D (n = 117 embryos/4939 cilia) and MZ myo1D ; vangl2+/− (n = 30/1653) embryos present a similar cilia size. The MZ myo1D mutant dataset displayed in f is the same that is also displayed in Fig. 3j. a, b, d, e are dorsal views of eight-somites stage KVs, anterior up. Scale bars: 20 µm in a, b; 10 µm in d, e
PCP signaling controls the posterior positioning of the ciliary basal body and the orientation of the ciliary rotation cone to promote LRO flow11,28,40,41. Rat myo1D controls cilia orientation in multiciliated epithelia42. In this context, myo1D governs both the positioning of ciliary basal bodies (reflecting translational PCP) and the orientation of the ciliary rotation cone (rotational PCP). The inactivation of either myo1D or vangl2 causes an anterior displacement of the basal body in the zebrafish LRO (Fig. 6a–e). The observation that basal body positioning is similar in MZ myo1D and MZ myo1D ; vangl2+/− animals (Fig. 6b, d, e) suggests however that the two factors do not interact in the control of translational PCP. To determine the effect of myo1D and vangl2 on rotational PCP, we imaged cilia in live embryos and found that both genes affect cilia orientation: while vangl2 mutants present an excess of anteriorly pointing cilia in the anterior KV, the loss of myo1D increases the number of posteriorly pointing cilia in the posterior organ half (Fig. 6f–o, Supplementary Fig. 9). Importantly, the removal of one copy of vangl2 significantly alters the cilia orientation of MZ myo1D mutants, so that MZ myo1D ; vangl2+/− animals are similar to WT controls (Fig. 6f, g). These findings suggest that myo1D and vangl2 interact in the context of the control of ciliary rotational PCP, but do so in opposite ways, consistent with our genetic interaction data (Fig. 4d–j). Taken together, our data suggest that myo1D/PCP interactions are required to govern KV cilia orientation and promote the establishment of a functional directional flow in the zebrafish LRO.
myo1D interacts with vangl2 to control cilia orientation in the zebrafish left–right Organizer. a–e myo1D and vangl2 are required for basal body positioning. a–d High magnification views of KV cells (ZO-1) with cilia (acetylated tubulin) and basal bodies (γ-tubulin). e Dot plot representing average antero-posterior basal body position in individual embryos (see Methods). Compared to WT (a, n = 40 embryos/744 basal bodies), basal bodies are displaced anteriorly in MZ myo1D (b, n = 74/1521) or vangl2 (c, n = 36/526) mutants. Basal bodies are similarly affected in MZ myo1D and MZ myo1D ; vangl2+/− (d, n = 37/826). f, g MZ myo1D (n = 93 embryos/3172 cilia) and vangl2 (n = 50/1545) mutations have opposing effects on cilia orientation. Cilia orientation in MZ myo1D ; vangl2+/− (n = 35/1225) is similar to WT (n = 55/2275). h–o two-dimensional dot plots indicating the position of KV cilia pointing to the anterior (red) or posterior (green) in the KV of embryos of different genotypes. Posteriorly pointing cilia invade the posterior KV in MZ myo1D (i), while anterior pointing cilia invaded the anterior KV in vangl2−/− (n). In MZ myo1D ; vangl2+/− (k, o), cilia distribution is again more similar to WT (see Supplementary Fig. 9 for details). All data collected at eight-somites stage, anterior up in a–d, h–o. Horizontal bars in e represent mean values. Error bars in f, g indicate SEM. Scale bar: 5 µm in a–d
Discussion
The existence of an evolutionarily conserved mechanism for the establishment of animal LR asymmetry has long been subject to debate1,2,3,4. Here we identify Myo1D, an unconventional non-muscle Myosin that was initially identified for its essential function in Drosophila LR asymmetry20,21, as an evolutionarily conserved regulator of animal LR asymmetry. Our genetic analysis reveals that laterality is correctly established in zebrafish embryos that lack zygotic myo1D function, similar to zygotic rat myo1D mutants42. In contrast, laterality defects are observed in zebrafish MZ myo1D mutants (Fig. 1b–i). While a single copy of myo1D is present in flies, the zebrafish genome encodes myo1D and the closely related myo1G. MZ myo1D; MZ myo1G double mutants present an increased penetrance of LR asymmetry defects compared to MZ myo1D single mutants, revealing an essential function of myo1G in the establishment of vertebrate laterality (Fig. 1n, o). As in Drosophila, chiral morphogenesis is disrupted upon overexpression of the Myo1D-antagonist Myo1Cb (Fig. 1n, o). These findings suggest that Myo1D-like antagonists and Myo1C antagonists represent an evolutionarily conserved toolkit for the establishment of animal laterality.
MZ myo1D mutants present defects in cardiac, visceral, and brain laterality (Fig. 1b–m, Supplementary Fig. 2a–f), suggesting that myo1D function may be required at the level of the KV, which acts as a central zebrafish LRO. Accordingly, the inactivation of myo1D disrupts LRO-dependent asymmetric gene expressions at the level of the KV (dand5) and the left lateral plate mesoderm (pitx2) (Fig. 2). Through a quantitative analysis of KV architecture and LRO fluid flow in MZ myo1D mutants, we identify the failure to establish a functional LRO flow as the primary cause of MZ myo1D mutant laterality defects (Fig. 4a–c).
MZ myo1D mutants present an alteration of flow geometry and circularity that are similar to the ones resulting from the inactivation of the PCP pathway component vangl228,39 (Fig. 4d–f), suggesting that these two factors control the same biological process. Accordingly, a lowering of vangl2 gene dosage allows to partially rescue KV flow and embryonic laterality in MZ myo1D mutants (Fig. 4g–j). Studies in different vertebrate model organisms have revealed that PCP pathway activity governs the position and orientation of LRO cilia11,28,40,41. We provide evidence that, as in other species, basal bodies are located in the posterior half of zebrafish LRO cells. Inactivation of myo1D or vangl2 results in an anterior displacement of the basal bodies (Fig. 6a–e). In contrast, myo1D and vangl2 mutants present opposite defects in cilia orientation (Fig. 6f–o). The reduction in vangl2 gene dosage that partially rescues KV flow and morphological laterality defects restores the rotational orientation, but not the basal body position, of LRO cilia (Fig. 6e–g). Translational and rotational PCP of KV cilia can therefore be uncoupled, and the sole restoration of rotational PCP is sufficient to achieve a significant rescue of LRO flow.
Our observations suggest that Myo1D exerts an essential role in controlling the rotational orientation of zebrafish LRO cilia. Under WT conditions, the opposing activities of Myo1D and Vangl2 control cilia orientation to generate a productive LRO flow (Fig. 7a). The loss of either Myo1D or Vangl2 function disrupts this cellular tug of war, resulting in altered cilia orientations, chaotic LRO flow profiles, and ultimately LR asymmetry defects (Fig. 7b).
Model of Myo1D function in the zebrafish left–right organizer. a Schematic representation of a wild-type Kupffer’s vesicle (KV). Black lines indicate cell outlines and dots in magenta the basal bodies of KV cilia. Cilia pointing to the posterior are indicated in green, anteriorly pointing cilia in red. Under wild-type conditions Myo1D and Vangl2 exert opposing activities on cilia orientation, restricting thereby posteriorly pointing cilia to the anterior KV and anteriorly pointing cilia to the posterior KV. This control of cilia orientation allows the generation of a circular counter-clockwise fluid flow that promotes a proper establishment of left–right asymmetry at the level of different organs. b In myo1D mutant embryos, cilia orientation is altered so that posteriorly pointing cilia now invade the posterior KV. The KV flow loses its circular geometry and becomes chaotic, causing thereby the occurrence of subsequent morphological laterality defects at the level of the heart, viscera, and brain
The molecular mechanism through which myo1D and vangl2 interact remains to be established. A first possibility could be that myo1D functions as a general PCP pathway component. We do not favor this hypothesis. In addition to its function in the LRO, vangl2 regulates the PCP-dependent positioning and orientation of floor plate cilia28. In contrast, myo1D has no effect on floor plate cilia (Supplementary Fig. 10a, b). Similarly, MZ myo1D mutants do not present defects in vangl2/PCP-dependent convergence-extension movements (Supplementary Fig. 11). These observations suggest that myo1D and vangl2 interact specifically in the context of LR asymmetry, as observed in Drosophila13.
In the Drosophila adult hindgut, myo1D interacts differently with the global fat/dachsous and the core vangl2 PCP pathways13. Myo1D is expressed in a transient LRO where it binds the global PCP component Dachsous to trigger Vangl2-dependent chiral morphogenesis in an adjacent tissue. myo1D and vangl2 therefore interact genetically but not molecularly. In accordance with a similar scenario, Myo1D loss of function does not alter Vangl2 localization in the zebrafish LRO (Supplementary Fig. 10c, d).
A major challenge for future studies will be to determine how Myosin1 proteins interact with other components of the chiral morphogenesis program. It will notably be interesting to determine whether the residual KV flow of MZ myo1D mutant embryos is due to the activity of additional actin-dependent regulators of LR activity.
Taken together, our data show that Myo1D is essential for the formation and function of the zebrafish LRO, identifying a common regulator of LR asymmetry in both vertebrates and invertebrates. The functional interaction between zebrafish Myo1D and PCP controls the orientation of KV cilia that are required for the establishment of a directional, symmetry-breaking flow. Interestingly, recent work has shown a functional link between Myo1D and PCP pathways in Drosophila13 and in Xenopus (M. Blum, personal communication). We propose that the coupling between Myo1D and PCP may represent an evolutionarily conserved genetic framework to control animal LR asymmetry.
Methods
Fly strains and genetics
Flies were grown according to standard procedures. For inter-specific rescue experiments, the UAS::zMyo1D transgene (Zebrafish Myo1D ORF cloned into pUAST vector) was expressed either in myo1D depleted flies (w1118 ; ptc::GAL4, myo1Dk1 ; UAS::dmyo1D-RNAi21) or in a myo1Dk2 21 null mutant background (da::GAL4 ; myo1Dk2) (da::GAL4, Bloomington Drosophila Stock Center #55851). Two independent insertions of the UAS::zMyo1D transgene (lines 3 and 6) were used and showed similar results.
Rotation of the male genitalia was scored as previously described43. Briefly, male genital plates are scored according to the angle made between the dorsoventral and anus–penis axes when viewed from the posterior. Full dextral/clockwise rotation (+360°) corresponding to the WT situation; partial sinistral/counter-clockwise rotation (from –1 to –359°); full sinistral/counter-clockwise rotation (−360°).
Zebrafish strains and embryo maintenance
Embryos were raised in 0.3X Danieau medium (17.4 mM NaCl, 0.21 mM KCl, 0.12 mM MgSO4, 0.18 mM Ca(NO3)2, 1.5 mM Hepes, pH 7.6) at 28.5 °C, and staged according to standard criteria44. If necessary, 1-phenyl-2-thiourea (Sigma) was added at 30 mg/l to prevent embryonic pigmentation.
Crispr and Cas9 mutagenesis
Crispr/Cas9 mutagenesis of zebrafish myo1D (Ensembl gene ENSDARG00000036863) was performed in a WT TÜ background. gRNA design and in vitro transcription of Cas9 RNA were performed according to reported protocols45. gRNA was transcribed from a template oligo as previously described46. The sequence of the selected Crispr target site was 5′-GAGTGGAGCTGGAAAAACAGAGG-3′ (bold lettering indicates the PAM motif). The efficiency of Crispr/Cas9-induced mutagenesis was monitored at 24 hpf using a T7 endonuclease assay45 on a PCR amplicon comprising the Crispr target region (Forward primer: 5′-TCTTCACTGACACTGGTATG-3′, Reverse primer: 5′-CCATCACTGCAGCAGAAATGAGAG-3′). Adult F0 fish were outcrossed to AB WT fish and DNA extracted from F1 progeny. Mutations were identified through direct sequencing of the same PCR amplicon that was used in the T7 endonuclease assay.
To genetically invalidate zebrafish myo1G (Ensembl gene ENSDARG00000036104), we used two complementary experimental strategies. A first mutant allele (myo1Gsa17276) was generated using a TILLING-based approach by the Zebrafish Mutation Project (http://www.sanger.ac.uk/resources/zebrafish/zmp/) and obtained through the Zebrafish International Resource Center. The sa17276 point mutation causes the appearance of a premature stop codon in place of amino acid 460, deleting therefore the entire C-terminal half of the protein including the actin- and cargo-binding domains.
To generate myo1D ; myo1G double mutants, a second mutant allele (myo1Gtj18b) was created through Crispr/Cas9 mutagenesis of the myo1G locus (genomic target site 5′-GATGTCATTGAGGACTACAGGGG-3′) in a myo1D mutant background. A preassembled complex of purified Cas9 protein (NEB, 954 ng/µl) and myo1G gRNA (200 ng/µl) was injected into one cell stage embryos derived from an incross if myo1Dtj16b/tj16c trans-heterozygous parents and the cutting efficiency monitored using a T7 endonuclease assay on a myo1G PCR amplicon (Forward primer: 5′-GGAGAAGTAGTGGTGTCCGTTAAC-3′, Reverse primer: 5′-CTCACTTTTGGGCTAACAGCTC-3′). Injected F0 embryos were raised to adulthood, crossed to myo1Dtj16b homozygous fish, and DNA extracted from F1 progeny. Direct sequencing of the same PCR amplicon used in the T7 endonuclease assay lead to the identification of the myo1Gtj18b allele. The myo1Gtj18b mutation deletes a CA dinucleotide corresponding to positions 198/199 of the open reading frame and therefore causes the appearance of a premature stop codon in place of amino acid 66. Consequently, the Myo1Gtj18b mutant protein lacks all major functional domains, including the ATP-binding P-loop as well as actin- and cargo-binding regions.
Fish strains and molecular genotyping
The following lines were used for this work: Tg(-5.0sox17:EGFP)zf9947, Tg(cmlc2:RFP)48, vangl2m20949, Tg(vangl2:Vangl2-GFP)50, and dnaaf/lrrc50tm317b 38.
WT and mutant alleles of myo1D were identified through allele-specific PCRs. The allele-specific forward primers 5′-TGGAGCTGGAAAAAGGCTCGT-3′ (myo1Dtj16b) and 5′-GTGGAGCTGGAAAAAGGCTATAC-3′ (myo1Dtj16c) were used together with the generic reverse primer 5′-CCATCACTGCAGCAGAAATGAGAG-3′ to genotype for the presence of different myo1D mutant alleles. PCR amplicon size was 133 bp for myo1Dtj16b and 145 bp for myo1Dtj16c. The WT myo1D allele was amplified using the forward primer 5′-AGAGTGGAGCTGGAAAAACAGA-3′ and the reverse primer 5′-CCCATCCCTCGTGTGAAACTAAATCAC-3′ to yield a 339 bp amplicon.
The allele-specific reverse primers 5′-CCCTGCAATGAGACAGGCTTA-3′ (recognizing the myo1Gsa17276 mutation) and 5′-CCCTGCAATGAGACAGGCGTC-3′ (recognizing the corresponding WT sequence) were used together with the generic forward primer 5′-AGGAGTACCAGAGAGAGGGAATC-3′ to detect the presence of the myo1Gsa17276 mutant allele (215 bp amplicons). The allele-specific reverse primers 5′-TCTCATACAGTTCTCTTCCCCTAG-3′ (myo1Gtj18b, 115 bp amplicon) and 5′- CTCATACAGTTCTCTTCCCCTGTAG-3′ (WT, 120 bp amplicon) were used together with the generic forward primer 5′-GAGAAGAGTCGTATCTACACCTTC-3′ to detect the presence of the myo1Gtj18b mutant allele.
PCR amplifications were performed with GoTaqG2 polymerase (Promega) at 1.5 mM MgCl2 using the following cycling conditions: Initial denaturation 2 min 95 °C; 10 cycles (30 s 95 °C, 30 s 65 °C first to 55 °C last cycle, 30 s 72 °C); 20 cycles (30 s 95 °C, 30 s 55 °C, 30 s 72 °C); final extension 5 min 72 °C.
To generate M zygotic myo1D mutants (MZ myo1Dtj16b/tj16b or MZ myo1Dtj16c/tj16c), myo1D heterozygous fish were incrossed and the progeny raised to adulthood. PCR genotyping of these animals allowed identifying homozygous mutant, as well as homozygous WT fish which were then used to produce MZ myo1Dtj16b/tj16b or MZ myo1Dtj16c/tj16c mutant embryos, as well as WT myo1D+/+ control embryos. Unless mentioned otherwise, experiments were performed using the myo1Dtj16b allele. Similar procedures were used to generate MZ myo1Gsa17276 single and MZ myo1Dtj16b ; MZ myo1Gtj18b double mutants.
For the analysis of spaw expression in MZ myo1D ; vangl2+/− animals presented in Fig. 4i, myo1Dtj16b/tj16b females were crossed to myo1Dtj16b/tj16b ; vangl2+/− males. Following in situ hybridization, a total of 267 animals were first scored for spaw expression and then genotyped for the presence of the vangl2m209 allele as previously described51. PCR was performed with forwards primers specific for either the WT (5′- GTGTGTCTGCCTGTGTCTTACT-3′) or the mutant (5′-GTGTGTCTGCCTGTGTCTTACA-3′) allele together with a generic reverse primer (5′- GATAAACTCCTCCCCCAGGT-3′) to amplify 250 bp amplicons.
For the genetic interaction experiment displayed in Fig. 4j, myo1Dtj16b homozygous mutant females were crossed with myo1Dtj16b/+; vangl2m209/+ males. A total of 912 embryos derived from six independent crosses were scored and molecular genotyping performed for all embryos presenting cardiac jogging defects.
Plasmid generation
The myo1Cb ORF was amplified from a mixed stage pool of cDNAs using the primers: 5′- GATCCCATCGATTCGACGGAATCCGGGTCATGATG-3′ and 5′-AGGCTCGAGAGGCCTTGGTCACGACTCGTCCATC-3′ and cloned into the pCS2+ vector. Bold letterings indicate primer overhangs that are also present in the pCS2+ sequence and were used to recombine insert and vector using Gibson assembly (NEB). A similar strategy was used to clone the myo1D ORF into pCS2+ using the primers 5′-GATCCCATCGATTCGACAGCTTATTATGGCAGAACACG-3′ and 5′-AGGCTCGAGAGGCCTTAAATGGGATCCTGGTCCTCTAG-3′.
To test the potential residual activity of the Myo1Dtj16b mutant protein, the tj16b mutation was introduced into myo1D-pCS2+ through amplification with the primers 5′-GCATAATGTATTTACTAGCCTTTTTCCAGCTCCACTCTCCC-3′ and 5′-GGCTAGTAAATACATTATGC-3′, followed by re-ligation through Gibson assembly (NEB).
RNA and morpholino injections
myo1Cb-pCS2+, myo1D-pCS2+, and myo1Dtj16b-pCS2+ were linearized with KpnI and RNA in vitro synthesized using the mMessage mMachine SP6 transcription kit (Ambion). The resulting RNAs were injected at 50 ng/µl for myo1Cb and 25 ng/µl for myo1D and myo1Dtj16b. Arl13b-GFP-pCS2+ was linearized with ApaI and transcribed with SP6 RNA polymerase as previously described28. RNA was injected at 20 ng/µl. To inhibit myo1D function, a translation-blocking morpholino (Gene Tools) with the sequence 5′-ACTTTCGTGTTCTGCCATAATAAGC-3′ was injected at 750 µM. All reagents were injected at the one cell stage at a volume of 0.88 nl together with 0.2% phenol red.
RNA in situ hybridizations
Whole mount RNA in situ hybridizations were performed according to standard protocols52. The following probes were used in this study: cmlc253, foxa1/fkd754, lefty155, pitx233, sox1756, spaw32, dand5/charon34. To generate a myo1D probe, myo1D was amplified from myo1D-pCS2+ using the following primers 5′-CGCTAATACGACTCACTATAGGGAGAGTTCTAGAGGCTCGAGAGG-3′ and 5′-GGCAAGACCAAAGTTTTCATCCGC-3′. The in situ probe was then synthesized directly from the PCR product using the incorporated T7 promoter (bold lettering). In situ hybridizations were documented using a Leica M205FA stereomicroscope equipped with an Infinity2-5C color CCD (Lumenera).
Immunohistochemistry
Dechorionated embryos were fixed overnight at 4 °C in PEM (80 mM Sodium-Pipes, 5 mM EGTA, 1 mM MgCl2) −4% PFA −0.2% Triton X-100. After washing 2 × 5 min in PEMT (PEM −0.2% TritonX100), 10 min in PEM −50 mM NH4Cl, 2 × 5 min in PEMT and blocking in PEMT −2% BSA, embryos were incubated 2 h at room temperature with primary antibodies. Following incubation, embryos were washed during 5, 10, 15, and 20 min in PEMT, blocked in PEMT −2% BSA, and incubated again with secondary antibodies for 2 h. Embryos were again washed during 5, 10, 15, and 20 min in PEMT, mounted in PEM −0.75% LMP-Agarose and analyzed on a Zeiss LSM710 laser scanning confocal microscope. The following primary antibodies were used in this study: Mouse@ZO-1 (1:250, Invitrogen 33-9100), Mouse@Acetylated Tubulin (1:2000, Sigma T7451), and Rabbit@γ-Tubulin (1:250, Sigma T5192). The secondary antibodies Goat@MouseIgG1-Alexa568, Goat@MouseIgG2b-Alexa647, and Goat@Rabbit-Alexa488 were obtained from Invitrogen and used at a dilution of 1:500.
Pharmacological treatments
Stock solutions of IBMX (100 mM, Sigma) and Forskolin (10 mM, Sigma) were prepared in DMSO. Ouabain (1 mM, Sigma) was prepared in distilled water. Embryos were treated with 10 µM Forskolin and 40 µM IBMX, or 5 µM Ouabain at bud stage and washed in Danieau medium after imaging at the eight-somite stage.
Tracking of fluorescent beads in the LRO
Fluoresbrite Polychromatic Red Microspheres 0.5 µm (Polysciences) were manually injected into the KV lumen. The movement of fluorescent beads was imaged on a wide field Zeiss Axio Observer Z1 microscope with a 40x EC Plan-Neofluar PH/DCII NA 0.75 dry objective using an Andor Neo sCMOS camera and a DsRed filter. In this setup we obtained images with a pixel width of 0.1625 µm and recorded the KV flow for 2 min with a frame interval of 0.04 s. In addition, a single brightfield image was acquired to visualize the KV outline. Automated tracking of fluorescent beads was performed using a custom-made ImageJ script (available as Supplementary Software 1 file). The macro generates an automated tracking file that is compatible with the ImageJ MTrackJ plugin. Following automated detection, tracks were manually curated using MTrackJ to correct for any errors of the tracking algorithm and the tracking parameters exported in a .CSV datasheet for further analysis. Using this procedure, an average of 44022 tracking points were obtained per embryo. KV images were oriented precisely along the antero-posterior axis based on the position of the notochord. The center of the KV was determined as the point lying midway between the anterior and the posterior extremity of the KV on a line that extends posteriorly from the anterior pole of the KV. The angular flow velocities displayed in Fig. 4 and Supplementary Figs. 6, 8 were calculated using the center of the KV as the point of origin of a polar coordinate system.
Quantification of ciliary orientation
KV motile cilia were labeled using Arl13b-GFP RNA microinjection and imaged on a Zeiss 710 laser scanning confocal microscope. The entire KV was scanned with z-sections of 0.2 µm distance and a frame interval of 0.48 s. The 488 nm laser line was used to simultaneously record the fluorescent cilia signal as well as a transmitted light image to visualize KV morphology. Movies were then treated in ImageJ with a Mean 3D filter to visualize the ciliary rotation cone and be able to orient it from base to tip (center of the cone). Each cilium was drawn on the KV using the ImageJ arrow tool. Cilia positions and angles were extracted from the arrow ROIs using a custom-made ImageJ script (available as Supplementary Software 1 file). To compare the position and orientation of cilia in different embryos as shown in Fig. 6h–o and Supplementary Fig. 9, the relative positions of different cilia were normalized in the common space of a circular virtual KV. Each cilium was placed on the virtual KV according to its relative radial position with respect to the organ border.
Quantification of basal body positioning
Confocal Z-stacks of the entire KV were acquired with a distance of 0.6 µm between two consecutive planes. The KV was oriented along the antero-posterior axis based on the position of the notochord. Basal bodies were identified as γ-tubulin positive spots that are located at the base of an acetylated tubulin positive ciliary axoneme. The junctional marker ZO-1 was used to outline KV cells and determine the antero-posterior extent of each cell. We restrained our analysis to cells for which the entire outline could be unambiguously identified in the ZO-1 channel. As the KV is an ellipsoid structure, this applies essentially to the cells of the dorsal roof and ventral floor of the KV. Based on this criterium, we were able to determine the antero-posterior position of an average of 19.4 basal bodies per embryo. The antero-posterior location of the basal body was quantified by calculating the ratio between the distance of the basal body from the anterior-most limit of the ZO-1 signal and the full antero-posterior length of a cell inferred from the ZO-1 signal. According to this convention, a value of 0 would indicate a basal body that is located at the anterior extremity of the cell, while a value of 1 would correspond to an extreme posterior location.
dand5 and charon quantification
dand5/charon in situ hybridization pictures were analyzed with ImageJ. A bounding rectangle was fit on the dand5 expression domain and the mean gray levels of the left and right part of the rectangle measured. dand5/charon right/left enrichment was determined as the ratio between right and left mean gray levels as previously described57. For the individual display items presented in Fig. 2g–l, n–p, this approach yielded the following values of right/left asymmetry: WT Bud stage (g): 0.9824; WT three somites (h): 1.0295; WT six somites (i): 1.0712; WT eight somites (j): 1.1004; WT ten somites (k): 1.0916 ; WT 14 somites (l): 1.0450; eight-somite stage WT control (n): 1.0896; MZ myo1D (o): 1.0342; MZ myo1D; MZ myo1G (p): 1.0289.
Quantification of gastrulation stage convergence-extension movements
Bud stage embryos were imaged laterally on a Leica M205FA stereomicroscope and the angle between the anterior extremity of the hatching gland and the posterior extremity of the tail bud was quantified using the ImageJ angle tool. The reference point for the angle measurement was defined by manually fitting an oval ROI to the embryo and using the ImageJ Measure function to determine the coordinates of the centroid.
Use of research animals
Zebrafish experiments have been performed in accordance with animal welfare guidelines in the iBV zebrafish facility (authorization #A06-088-17) in the context of the authorized animal experimentation project (APAFIS#5521-201605111041958v6), approved by the animal experimentation ethical committee Ciepal Azur.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Information files or datasets (for imaging and quantitation that were generated and analyzed during the current study) are available from the corresponding author upon reasonable request.
References
Coutelis, J. B., González-Morales, N., Géminard, C. & Noselli, S. Diversity and convergence in the mechanisms establishing L/R asymmetry in metazoa. EMBO Rep. 15, 926–937 (2014).
Halpern, M. E., Hobert, O. & Wright, C. V. Left-right asymmetry: advances and enigmas. Genesis 52, 451–454 (2014).
Nakamura, T. & Hamada, H. Left-right patterning: conserved and divergent mechanisms. Development 139, 3257–3262 (2012).
Blum, M., Feistel, K., Thumberger, T. & Schweickert, A. The evolution and conservation of left-right patterning mechanisms. Development 141, 1603–1613 (2014).
Okada, Y., Takeda, S., Tanaka, Y., Izpisúa Belmonte, J. C. & Hirokawa, N. Mechanism of nodal flow: a conserved symmetry breaking event in left-right axis determination. Cell 121, 633–644 (2005).
Kramer-Zucker, A. G. et al. Cilia-driven fluid flow in the zebrafish pronephros, brain and Kupffer's vesicle is required for normal organogenesis. Development 132, 1907–1921 (2005).
Tanaka, Y., Okada, Y. & Hirokawa, N. FGF-induced vesicular release of Sonic hedgehog and retinoic acid in leftward nodal flow is critical for left-right determination. Nature 435, 172–177 (2005).
Yuan, S., Zhao, L., Brueckner, M. & Sun, Z. Intraciliary calcium oscillations initiate vertebrate left-right asymmetry. Curr. Biol. 25, 556–567 (2015).
Levin, M., Thorlin, T., Robinson, K. R., Nogi, T. & Mercola, M. Asymmetries in H+/K+-ATPase and cell membrane potentials comprise a very early step in left-right patterning. Cell 111, 77–89 (2002).
Norris, D. P. Cilia calcium and the basis of left-right asymmetry. Bmc. Biol. 10, 102 (2012).
Minegishi, K. et al. A Wnt5 activity asymmetry and intercellular signaling via PCP proteins polarize node cells for left-right symmetry breaking. Dev. Cell 40, 439–452.e434 (2017).
Gros, J., Feistel, K., Viebahn, C., Blum, M. & Tabin, C. J. Cell movements at Hensen's node establish left/right asymmetric gene expression in the chick. Science 324, 941–944 (2009).
González-Morales, N. et al. The atypical cadherin dachsous controls left-right asymmetry in Drosophila. Dev. Cell 33, 675–689 (2015).
Wang, G., Manning, M. L. & Amack, J. D. Regional cell shape changes control form and function of Kupffer's vesicle in the zebrafish embryo. Dev. Biol. 370, 52–62 (2012).
Noël, E. S. et al. A Nodal-independent and tissue-intrinsic mechanism controls heart-looping chirality. Nat. Commun. 4, 2754 (2013).
Hobert, O. Development of left/right asymmetry in the Caenorhabditis elegans nervous system: from zygote to postmitotic neuron. Genesis 52, 528–543 (2014).
Naganathan, S. R., Middelkoop, T. C., Fürthauer, S. & Grill, S. W. Actomyosin-driven left-right asymmetry: from molecular torques to chiral self organization. Curr. Opin. Cell. Biol. 38, 24–30 (2016).
Pyrpassopoulos, S., Feeser, E. A., Mazerik, J. N., Tyska, M. J. & Ostap, E. M. Membrane-bound myo1c powers asymmetric motility of actin filaments. Curr. Biol. 22, 1688–1692 (2012).
Tee, Y. H. et al. Cellular chirality arising from the self-organization of the actin cytoskeleton. Nat. Cell. Biol. 17, 445–457 (2015).
Hozumi, S. et al. An unconventional myosin in Drosophila reverses the default handedness in visceral organs. Nature 440, 798–802 (2006).
Spéder, P., Adám, G. & Noselli, S. Type ID unconventional myosin controls left-right asymmetry in Drosophila. Nature 440, 803–807 (2006).
Davison, A. et al. Formin is associated with left-right asymmetry in the pond snail and the frog. Curr. Biol. 26, 654–660 (2016).
McDowell, G. S. et al. Conserved roles for cytoskeletal components in determining laterality. Integr. Biol. 8, 267–286 (2016).
Géminard, C., González-Morales, N., Coutelis, J. B. & Noselli, S. The myosin ID pathway and left-right asymmetry in Drosophila. Genesis 52, 471–480 (2014).
Sampaio, P. et al. Left-right organizer flow dynamics: how much cilia activity reliably yields laterality? Dev. Cell. 29, 716–728 (2014).
Ferreira, R. R., Vilfan, A., Jülicher, F., Supatto, W. & Vermot, J. Physical limits of flow sensing in the left-right organizer. eL ife 6, 25078 (2017).
Compagnon, J. et al. The notochord breaks bilateral symmetry by controlling cell shapes in the zebrafish laterality organ. Dev. Cell. 31, 774–783 (2014).
Borovina, A., Superina, S., Voskas, D. & Ciruna, B. Vangl2 directs the posterior tilting and asymmetric localization of motile primary cilia. Nat. Cell Biol. 12, 407–412 (2010).
McIntosh, B. B. & Ostap, E. M. Myosin-I molecular motors at a glance. J. Cell. Sci. 129, 2689–2695 (2016).
Petzoldt, A. G. et al. DE-cadherin regulates unconventional myosin ID and myosin IC in Drosophila left-right asymmetry establishment. Development 139, 1874–1884 (2012).
Oteíza, P., Köppen, M., Concha, M. L. & Heisenberg, C. P. Origin and shaping of the laterality organ in zebrafish. Development 135, 2807–2813 (2008).
Long, S., Ahmad, N. & Rebagliati, M. The zebrafish nodal-related gene southpaw is required for visceral and diencephalic left-right asymmetry. Development 130, 2303–2316 (2003).
Faucourt, M., Houliston, E., Besnardeau, L., Kimelman, D. & Lepage, T. The pitx2 homeobox protein is required early for endoderm formation and nodal signaling. Dev. Biol. 229, 287–306 (2001).
Hashimoto, H. et al. The cerberus/Dan-family protein charon is a negative regulator of Nodal signaling during left-right patterning in zebrafish. Development 131, 1741–1753 (2004).
Gokey, J. J., Ji, Y., Tay, H. G., Litts, B. & Amack, J. D. Kupffer's vesicle size threshold for robust left-right patterning of the zebrafish embryo. Dev. Dyn. 245, 22–33 (2016).
Navis, A., Marjoram, L. & Bagnat, M. Cftr controls lumen expansion and function of Kupffer's vesicle in zebrafish. Development 140, 1703–1712 (2013).
Wang, G. et al. The Rho kinase Rock2b establishes anteroposterior asymmetry of the ciliated Kupffer's vesicle in zebrafish. Development 138, 45–54 (2011).
Sullivan-Brown, J. et al. Zebrafish mutations affecting cilia motility share similar cystic phenotypes and suggest a mechanism of cyst formation that differs from pkd2 morphants. Dev. Biol. 314, 261–275 (2008).
May-Simera, H. L. et al. Bbs8, together with the planar cell polarity protein Vangl2, is required to establish left-right asymmetry in zebrafish. Dev. Biol. 345, 215–225 (2010).
Antic, D. et al. Planar cell polarity enables posterior localization of nodal cilia and left-right axis determination during mouse and Xenopus embryogenesis. PLoS ONE 5, e8999 (2010).
Song, H. et al. Planar cell polarity breaks bilateral symmetry by controlling ciliary positioning. Nature 466, 378–382 (2010).
Hegan, P. S., Ostertag, E., Geurts, A. M. & Mooseker, M. S. Myosin Id is required for planar cell polarity in ciliated tracheal and ependymal epithelial cells. Cytoskeleton 72, 503–516 (2015).
Coutelis, J. B. et al. Drosophila left/right asymmetry establishment is controlled by the Hox gene abdominal-B. Dev. Cell. 24, 89–97 (2013).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Jao, L. E., Wente, S. R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl Acad. Sci. USA 110, 13904–13909 (2013).
Hruscha, A. et al. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 140, 4982–4987 (2013).
Mizoguchi, T., Verkade, H., Heath, J. K., Kuroiwa, A. & Kikuchi, Y. Sdf1/Cxcr4 signaling controls the dorsal migration of endodermal cells during zebrafish gastrulation. Development 135, 2521–2529 (2008).
Mably, J. D., Mohideen, M. A., Burns, C. G., Chen, J. N. & Fishman, M. C. Heart of glass regulates the concentric growth of the heart in zebrafish. Curr. Biol. 13, 2138–2147 (2003).
Solnica-Krezel, L. et al. Mutations affecting cell fates and cellular rearrangements during gastrulation in zebrafish. Development 123, 67–80 (1996).
Sittaramane, V. et al. The PCP protein Vangl2 regulates migration of hindbrain motor neurons by acting in floor plate cells, and independently of cilia function. Dev. Biol. 382, 400–412 (2013).
Gao, B. et al. Wnt signaling gradients establish planar cell polarity by inducing Vangl2 phosphorylation through Ror2. Dev. Cell. 20, 163–176 (2011).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Rottbauer, W. et al. Cardiac myosin light chain-2: a novel essential component of thick-myofilament assembly and contractility of the heart. Circ. Res. 99, 323–331 (2006).
Odenthal, J. & Nüsslein-Volhard, C. Fork head domain genes in zebrafish. Dev. Genes. Evol. 208, 245–258 (1998).
Thisse, C. & Thisse, B. Antivin, a novel and divergent member of the TGFbeta superfamily, negatively regulates mesoderm induction. Development 126, 229–240 (1999).
Alexander, J. & Stainier, D. Y. A molecular pathway leading to endoderm formation in zebrafish. Curr. Biol. 9, 1147–1157 (1999).
Superina, S., Borovina, A. & Ciruna, B. Analysis of maternal-zygotic ugdh mutants reveals divergent roles for HSPGs in vertebrate embryogenesis and provides new insight into the initiation of left-right asymmetry. Dev. Biol. 387, 154–166 (2014).
Acknowledgements
The present project was supported by the ANR project grants DRO-ASY (ANR-13-BSV2-006, S.N.) and DroZeMyo (ANR-17-CE13-0024-02, M.F., S.N.). The work in the lab of M.F. was supported by a CNRS/INSERM ATIP/Avenir 2010 grant and an HFSP Career Development Award (00036/2010). T.J. benefited from a PhD fellowship from INSERM/Région PACA/Cancéropôle PACA and the Labex SIGNALIFE (ANR-11-LABX-0028-01). We thank S. Lopes, B. Ciruna, J. Vermot, A. Chandrasekhar, and T. Lepage for the sharing of reagents and fish lines. We thank Janine Leblanc-Straceski and Martin Blum for discussions. We are particularly grateful to R. Rebillard for excellent fish care. Confocal microscopy and timelapse imaging of the KV fluid flow were performed at the iBV PRISM imaging platform with the help of Magali Mondin.
Author information
Authors and Affiliations
Contributions
The genetic analysis of myosinI function in zebrafish LR asymmetry was entirely performed by T.J. J.-B.C. performed initial morpholino knockdown experiments of zmyo1D and zmyo1G. S.P. cloned zmyo1D and G, analyzed myo1D expression and performed myo1D morpholino knockdown and myo1Cb overexpression experiments together with T.J. M.F. performed myo1D RNA rescue experiments and analyzed floor plate cilia and Vangl2-GFP localization in myo1D mutants. D.C. generated zmyo1D-transgenic Drosophila lines. C.G. performed the rescue experiments of the fly myo1D mutants with zmyo1D. S.N. and M.F. designed the study, analyzed the data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Juan, T., Géminard, C., Coutelis, JB. et al. Myosin1D is an evolutionarily conserved regulator of animal left–right asymmetry. Nat Commun 9, 1942 (2018). https://doi.org/10.1038/s41467-018-04284-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-04284-8
This article is cited by
-
Myosin1G promotes Nodal signaling to control zebrafish left-right asymmetry
Nature Communications (2024)
-
Identification of a novel MYO1D variant associated with laterality defects, congenital heart diseases, and sperm defects in humans
Frontiers of Medicine (2024)
-
Multiple pkd and piezo gene family members are required for atrioventricular valve formation
Nature Communications (2023)
-
Asymmetric activity of NetrinB controls laterality of the Drosophila brain
Nature Communications (2023)
-
Vertebrate myosin 1d regulates left–right organizer morphogenesis and laterality
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.