Abstract
In photosystem II, light-induced water oxidation occurs at the Mn4CaO5 cluster. Here we demonstrate proton releases, dioxygen formation, and substrate water incorporation in response to Mn4CaO5 oxidation in the protein environment, using a quantum mechanical/molecular mechanical approach and molecular dynamics simulations. In S2, H2O at the W1 site forms a low-barrier H-bond with D1-Asp61. In the S2-to-S3 transition, oxidation of OW1H– to OW1•–, concerted proton transfer from OW1H– to D1-Asp61, and binding of a water molecule Wn-W1 at OW1•– are observed. In S4, W n -W1 facilitates oxo-oxyl radical coupling between OW1•– and corner μ-oxo O4. Deprotonation via D1-Asp61 leads to formation of OW1=O4. As OW1=O4 moves away from Mn, H2O at W539 is incorporated into the vacant O4 site of the O2-evolved Mn4CaO4 cluster, forming a μ-oxo bridge (Mn3–OW539–Mn4) in an exergonic process. Simultaneously, Wn-W1 is incorporated as W1, recovering the Mn4CaO5 cluster.
Similar content being viewed by others
Introduction
Dioxygen can be produced by removing four electrons and four protons from two substrate water molecules: 2H2O → O2 + 4 H+ + 4e–. In a protein-pigment complex photosystem II (PSII), the water-oxidation reaction proceeds at the oxygen-evolving complex via the S n -state transitions (n represents the total number of oxidation steps)1,2,3. The oxygen-evolving complex is linked with electron transfer pathways, proton transfer pathways, and substrate water intake pathways. Excitation of P680, which is composed of four chlorophyll a molecules, leads to electron transfer to the second plastoquinone, QB, via pheophytin a and the first plastoquinone, QA. The resulting P680•+ state is stabilized by electron transfer from the Mn4CaO5 cluster via redox active D1-Tyr161 (TyrZ) (Fig. 1).
Protons released from substrate water molecules are transferred via proton transfer pathways (e.g., O4-water chain4), which proceeds from the Mn4CaO5 cluster toward the protein bulk surface (Fig. 1). The releases of protons are observed with a typical stoichiometry of 1:0:1:2 for the S0 → S1 → S2 → S3 → S0 transitions, respectively5. The O4-water chain is a chain of water molecules, which forms an H-bond with the O4 site of the Mn4CaO5 cluster via a water molecule W5394. Since the rate constant is pH-independent in the S0-to-S1 transition, the absence of ionizable groups in the O4-water chain fits with the suggestion that the O4-water chain is the relevant proton transfer pathway5,6. Although release of a proton is not observed in the S1-to-S2 transition, it seems likely that migration of a proton or internal proton transfer can occur (e.g., ref. 7). On the other hand, the rate constant for the S2-to-S3 transition is pH-dependent, which suggests that ionizable groups are involved in the proton-transfer pathway5,8. Ionizable groups that are not ligands near the Mn4CaO5 cluster are D1-Asp61 and CP43-Arg3579. In the S3-to-S0 transition, O–O bond formation and release of O=O occur. The rate constant of 1–2 ms involved in the S3-to-S0 transition is pH-independent with a kinetic isotope effect (kD/kH) of 1.46, and may correspond to O2 release and incorporation of a substrate water molecule6,10. kD/kH = 1.4 is also typical for alteration in the H-bond pattern along water chains before and after proton transfer (pre-PT and post-PT H-bond patterns, respectively) in the Grotthuss mechanism11. Once proton transfer occurs, the pre-PT pattern transforms to the post-PT pattern4.
The slow-exchanging and fast-exchanging water molecules are candidate substrates in the Mn4CaO5 cluster12. In the S0-to-S1 transition, the exchange rate of the slow-exchanging water molecule decreases 600 times12 and deprotonation of the slow-exchanging water molecule may occur, indicating that it may be a μ-OH bridge of the Mn4CaO5 cluster in S013. In the S1-to-S2 transition, the exchange rate of the slow-exchanging water molecule increases 100 times12. The exchange rate of the fast-exchanging water molecule decreases three times in the S2-to-S3 transition14. The fast-exchanging water molecule may be a terminal ligand to Mn[2] and possibly form an H-bond with D1-Asp6113,15. The corresponding water molecule is W1 in the crystal structure analyzed at 1.9 Å resolution9. These water molecules approach the catalytic site from the bulk region via water channels. The D1-Glu65/D2-Glu312 channel is a candidate substrate water intake channel. The D1-Glu65/D2-Glu312 channel provides water molecules near TyrZ and the Mn4CaO5 cluster. It can also deliver water molecules to the W539 and W538 sites in the O4-water chain via D1-Asp6116.
Recently, the two-flash illuminated PSII structure analyzed at 2.35 Å resolution was reported, demonstrating several characteristic O sites in the Mn4CaO6 moiety17 (e.g., W538, W539, O4, and O6). W5389 (i.e., W66517) was unambiguously identified in both the 1.9 Å9 and radiation-damage-free18 PSII crystal structures4, whereas it was absent in the two-flash illuminated PSII structure. W538 forms an H-bond with W539. The averaged O4…OW539 (i.e., O4…OW56717) distance of 2.32 Å identified in the two-flash illuminated PSII structure is exceptionally short in comparison with the typical O–O distances of ~2.8 Å for standard (asymmetric) H-bonds in H2O. The two-flash illuminated PSII structure shows the presence of O6 near O5. The O6…O5 distance is 1.5 Å, which is close to the distance for peroxide (O–O)2–. This suggests that formation of O=O may occur at this site, which fits the reaction model proposed by Siegbahn et al.19. On the other hand, it is unclear whether the reaction model could also explain how the other characteristic O sites (e.g., W538, W539, and O4) are involved in the mechanism of water oxidation. It should be noted that the B-factor values of these O atoms are at the same level in the two-flash illuminated PSII structure.
Here we analyzed alterations in the Mn4CaO5 structure and H-bond/water network in response to Mn4CaO5 oxidation in the PSII protein environment, using a quantum mechanical/molecular mechanical (QM/MM) approach and molecular dynamics (MD) simulations based on the PSII crystal structure. In S2, the proton of H2O at W1 significantly migrates toward D1-Asp61 due to oxidation of Mn4, forming a low-barrier H-bond (HOW1–…H+…–OOCD1-Asp61). In the S2-to-S3 transition, oxidation of OW1H– to OW1•– preferentially occurs with respect to oxidation of Mn1(III) to Mn1(IV), because D1-Asp61 accepts the proton from OW1H– and facilitates oxidation to OW1•– via proton-coupled electron transfer (i.e., decreasing the redox potential of W1 for oxidation). In S3, a water molecule binds at OW1•–. Incorporation of water molecules (including W2 and W3) into the open-cubane O5 site are not observed. Since the nearest μ-oxo to OW1•– is O4, oxo-oxyl radical coupling occurs between O4 and OW1•– in S4, resulting in (OW1–O4)2– formation. As OW1=O4 moves away from the Mn3/Mn4 moiety, H2O at W539 is incorporated into the vacant O4 site near CP43-Arg357, forming a μ-oxo bridge with Mn3–OW539 and Mn4–OW539 in a concerted exergonic process. Incorporation of W539 into the O4 site requires reorientation of W539, which induces alterations in the H-bond pattern from the post-PT pattern to pre-PT pattern along the O4-water chain. A water molecule, which binds at OW1•– in S3, is also incorporated into the vacant W1 site (i.e., new W1), recovering the Mn4CaO5 cluster.
Results
Formation of a low-barrier H-bond between W1 and D1-Asp61
We used the deprotonated open-cubane S2 structure, where (Mn1,Mn2,Mn3,Mn4) = Mn(III,IV,IV,IV), as it seems to be energetically more stable than the closed-cubane S2 structure20,21,22,23. The pKa of H-bond donor and acceptor moieties in H-bonds can be analyzed from the potential-energy profiles of the H-bonds24,25,26. In H-bonds, a proton is more likely to populate the moiety with the higher pKa value between the two moieties26. The energy difference between the H-bond donor and acceptor moieties corresponds to the pKa difference and this feature also holds true for H-bonds in protein environments27. QM/MM calculations showed that a short H-bond between H2O at W1 and D1-Asp61 (OW1…OD1-Asp61 = 2.45 Å) was formed in the open-cubane S2 (Fig. 2a). The potential-energy profile suggested that the OW1…OD1-Asp61 H-bond is a low-barrier H-bond (LBHB, where the pKa difference for the donor and acceptor moieties is nearly zero25,27) (Fig. 2b), and that the proton is significantly migrated toward D1-Asp61 and delocalized over the two moieties already in S2 (Fig. 2a). This may correspond to the significant changes in the H-bond properties between D1-Asp61 and a water molecule in the S1-to-S2 transition observed in Fourier transform infrared (FTIR) spectroscopy7. It should also be noted that the spin configurations (e.g., high, low, ferromagnetic, and antiferromagnetic) did not essentially affect the resulting optimized Mn4CaO5 geometry and energies in the present case (Table 1). Potential-energy profiles also remained unaffected since proton transfer does not involve oxidation of Mn ions (Supplementary Figure 1).
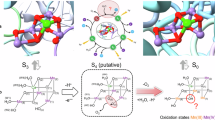
The open-cubane S2-to-S3 transition. a QM/MM geometries. Mn, Ca, O, and H atoms are represented by purple, orange, red, and black balls, respectively. Dotted lines indicate distances (excluding H atoms). b The energy profiles of the H-bonds between W1 (H2O/OH–) and D1-Asp61 in S2 and W1(OH•/O•–) and D1-Asp61 in S3. The black curve corresponds to release of the proton from H2O at W1 to D1-Asp61 in S2, whereas the red curve corresponds to release of the proton from OH• at W1 to D1-Asp61 in S3. The spin configurations (e.g., high, low, ferromagnetic, and antiferromagnetic) did not essentially affect the resulting optimized Mn4CaO5 geometries and potential-energy profiles (Table 1 and Supplementary Figure 1)
Absence of Mn oxidation in the S2-to-S3 transition
Unexpectedly, oxidation of S2 to S3 did not involve oxidation of Mn; however, oxidation of OW1H– to OW1•– and concerted proton transfer from OW1H– to D1-Asp61 did occur, resulting in Mn(III,IV,IV,IV)…OW1•–…HOOCD1-Asp61 in the open-cubane conformation (Fig. 2a). Accordingly, the first proton of W1 (from H2O/OH–), which was delocalized over the low-barrier OW1…H+…OD1-Asp61 bond in S2, seems likely to have been released toward the bulk surface via D1-Asp61 (along the D1-Glu65/D2-Glu312 water channel) in the S2-to-S3 transition16,28, since the potential-energy profile indicated that even the second proton from W1 (from OH•/O•–) can be released to D1-Asp61 (Fig. 2). MD simulations indicated that D1-Asp61 was able to accept a proton by rotating the protonated O site (Supplementary Figure 2), as reported in QM/MM-MD simulations by Narzi et al.29. Since pKa(OH–/ O2–) ( = 24) > pKa(OH•/ O•–) (=1230), the concerted oxidation and proton release of OW1H– to OW1•– occurs without stabilizing OW1H• on Mn4(IV).
In the S2-to-S3 transition, it remains unclear whether a Mn-centered oxidation (e.g., ref. 31) or a ligand-centered oxidation (e.g., ref. 32) occurs (discussed in refs. 5,13,33,34). Some theoretical studies favor the Mn-centered oxidation model over the ligand-centered oxidation model (e.g., refs. 35,36,37,38,39). Recent electrochemical analysis using the α-MnO2 electrode shows that addition of carboxylic acid (benzoic acid) induced proton-coupled electron transfer, resulting in a decrease in the redox potential required for water oxidation and an increase in evolved O2 at lower potentials40. Remarkably, FTIR spectra suggest that benzoic acids do not directly ligate to the Mn ion but form an H-bond with a ligand water molecule of the Mn ion, as identified in the Mn4…W1…D1-Asp61 moiety in PSII. The presence of benzoic acid as the proton acceptor facilitates proton-coupled electron transfer and can avoid accumulation of the positive charge in response to oxidation of Mn, leading to a decrease in the redox potential required for oxidation of water40. This may hold true for the present case, where proton-coupled electron transfer from W1 to D1-Asp61 occurs in response to the S2-to-S3 transition, which can specifically decrease the redox potential of W1 for one-electron oxidation.
The absence of the Mn-centered oxidation in the S2-to-S3 transition has been reported in spectroscopic studies by Messinger et al.; Mn was not oxidized, but a ligand substrate molecule was oxidized in the S2-to-S3 transition and an O radical was present in S3 (Mn(III)Mn(IV)3)32, which is consistent with the present results. It should be noted that although TyrZ and the H-bond network were also considered quantum-chemically (i.e., in the QM region), we did not observe formation of TyrZ•.
According to the interpretation of FTIR difference spectroscopy, a water molecule was incorporated into the Mn4CaO5 moiety in the S2 to S3 transition41. In S3, MD simulations showed that a water molecule existed near W1 (W n -W1) and could form an H-bond with OW1•– (Fig. 3). The corresponding water molecule is absent in the PSII crystal structures9,18. The binding of W n -W1 seems to be made more pronounced by the negatively charged OW1•– in S3 (Supplementary Table 1). Near W n -W1 there exists D1-Ser169, which has been proposed to provide access to water molecules for the substrate-binding site42. QM/MM calculations also showed that the H-bond between W1 and W n -W1 is remarkably short: 2.42 Å in S3 (i.e., O•– at W1; Fig. 2a) compared with 2.72 Å in S2 (i.e., H2O at W1; Fig. 2a). MD simulations suggested that water molecules at the W n -W1 binding moiety are exchangeable with bulk water via the D1-Glu65/D2-Glu312 water channel16, indicating that the D1-Glu65/D2-Glu312 water channel can serve as the intake channel for W n -W1.
Water distribution near the Mn4CaO5 cluster. a Water distribution in S3 after an equilibrating MD run at 45−50 ns (250 snapshots). The position of W n -W1 is indicated by the dotted circle. b The corresponding view in the QM/MM-optimized geometry in S3. Dotted black lines indicate interactions. c Water distribution in S1 after an equilibrating MD run at 45−50 ns (250 snapshots). W n -W1 is absent in S1 (dotted circle)
We did not observe any incorporation of water molecules (including W2 and W3) into the open-cubane O5 site, nor did we observe formation of OW2•– in QM/MM calculations, because of the absence of the proton acceptor groups (e.g., D1-Asp61). Below, we focus on the Mn4CaO5 cluster based on the obtained open-cubane S3 with OW1•–.
Oxidation of S3 and formation of (OW1–O4)2–
Oxidation of S3 resulted in Mn(IV,IV,IV,IV)…OW1•–…HOOCAsp61 (we denote the oxidized S3 states as “S4”, Fig. 4a). The presence of OW1•– in S4 suggests that W1 may be involved in O–O single-bond formation via oxo-oxyl radical coupling. The nearest μ-oxo to OW1•– is O4. Since we observed neither formation of OW2•– nor a reactive Mn(V)=O species, we focused on OW1•– and O4.
The S4 and pre-S0 states. a QM/MM geometries. Mn, Ca, O, and H atoms are represented by purple, orange, red, and black balls, respectively. Dotted lines indicate distances (excluding H atoms). Release of (OW1–O4)•– from the Mn3/Mn4 moiety (i.e., the Mn3…O4 distance was increased) resulted in formation of OW1=O4. See supplementary Figure 4 for the energy profile. b The energy profiles of the (OW1–O4)2− formation (the first and second panels in a) in the presence (blue curve) and absence (red curve) of W n -W1. The initial lower barriers correspond to reorientation of the carboxylate group of D1-Asp61 with respect to OW1•–/(OW1–O4)2–
The potential-energy profile suggests that as oxo-oxyl radical coupling (e.g., ref. 43) proceeds between O4 and OW1•–, formation of (OW1–O4)2– (confirmed by OW1–O4 = 1.44 Å and the spin density (2S) = 0 for both OW1 and O4) and concerted electron transfer from the (OW1–O4) moiety to Mn3 occur, resulting in Mn(IV,IV,III,IV)…(OW1–O4)2–…HOOCAsp61 (Fig. 4a). The energy barrier for the (OW1–O4)2– formation was 13.9 kcal/mol (Fig. 4b), which is essentially consistent with the estimated values (e.g., 12.1 kcal/mol44). In all S-state transitions, including the (OW1–O4)2– formation process, Mn2(IV) is not involved in oxidation/reduction, which could explain the similar energy barriers in the antiferromagnetically (↑↓↑↑) (Fig. 4) and ferromagnetically (↑↑↑↑) (Supplementary Figure 3) coupled forms. The energy barrier was 18.6 kcal/mol in the absence of W n -W1 (Fig. 4b and Supplementary Figure 3). Because W n -W1 lowers the energy barrier, it may serve as a catalyst for (OW1–O4)2– formation in S4. As formation of (OW1–O4)2– proceeded, the H atom of W n -W1, which was initially oriented toward OW1•–, became oriented away from (OW1–O4)2–. Simultaneously, the W n -W1…Mn4 distance decreased from 3.60 to 3.24 Å, owing to displacement of W1 toward O4, allowing W n -W1 to approach Mn4 (Fig. 4).
Formation of OW1=O4
When D1-Asp61 was deprotonated in the presence of (OW1–O4)2–, QM/MM calculations resulted in formation of (OW1–O4)•– (e.g., ref. 45, confirmed by OW1–O4 = 1.29 Å and the spin density (2S) = 0.8 for OW1 and 0.4 for O4) and concerted electron transfer to Mn1 (Fig. 4).
Furthermore, the following events occurred concertedly: (i) W n -W1 approached Mn4, (ii) (OW1–O4)•– moved away from Mn3/Mn4, (iii) W539 approached Mn3, and (iv) electron transfer from (OW1–O4)•– to Mn4 occurred, resulting in formation of OW1=O4 (confirmed by OW1–O4 = 1.22 Å and the spin density (2S) = 1.0 for OW1 and 0.9 for O4; Fig. 4) in an exergonic process (Supplementary Figure 4). The resulting oxidation state was Mn(III,IV,III,III), returning to an Mn oxidation state identical to that of S0[4].
After O2 evolution, the O2-evolved Mn4Ca“O4” cluster formed, which resembled the O4-lacking Mn4CaO4 cluster reported in the 3.5-Å PSII crystal structure46 and the O4-lacking synthetic Mn4CaO4 cluster reported by Zhang et al.47. The evolved O2 molecule is present in the hydrophobic space between Mn4 and D1-Ile60 (next to D1-Asp61), which has been proposed to serve as an O2-exiting pathway48. Indeed, O2 evolution was significantly lowered when D1-Ile60 was mutated to bulky phenylalanine49.
Release of O2 and incorporation of W539 into the O4 site
Intriguingly, when evolved O2 was absent near Mn3/Mn4, QM/MM calculations resulted in exergonic incorporation of H2O at W539 (in the O4-water chain4, Fig. 1) into the vacant O4 site of the Mn4CaO4 cluster and formation of a μ-oxo bridge with Mn3–OW539 (2.43 Å) and Mn4–OW539 (2.19 Å) (pre-S0 state, Fig. 4). While W539 was incorporated into the Mn4CaO4 cluster, W n -W1 simultaneously approached Mn4 and was incorporated into the Mn4CaO5 cluster as the new W1 ligand (W n -W1…Mn4 = 2.08 Å). The resulting QM/MM geometry (Fig. 4) was quite similar to that of the original PSII crystal structure, demonstrating the ability to recover the initial state in the Kok cycle.
D1-Asp61, the only acidic residue that is not a ligand near the Mn4CaO5 cluster (i.e., a second sphere ligand), accepted an H-bond from W539, delocalizing the negative charge over [D1-Asp61…W539]– and inducing OW539δ–. Thus, D1-Asp61 provides a driving force for the incorporation (i.e., it facilitates interactions between OW539δ– and Mn3(III)/Mn4(III)) (Fig. 5). Indeed, it has been reported that O2 release was dramatically decelerated in the D1-D61N mutant50,51,52. As long as O2 was present in the Mn3/Mn4 moiety, H2O at W539 did not incorporate into the O4-binding site (Supplementary Figure 4), i.e., the presence of O2 at the Mn3/Mn4 moiety sterically inhibited incorporation of H2O at W539 into the O4 site. Therefore, the exergonic incorporation of W539, which was facilitated by D1-Asp61, could contribute to expulsion of O2 (Supplementary Figure 4).
Difference in the orientations and interactions of the incorporating H2O molecule. a The O4 site. Anionic D1-Asp61 attracts the Hδ+ and orients the Oδ– of the incorporating H2O toward cationic Mn3(III), Mn4(III), and CP43-Arg357, resulting in attraction. b The O5 site. The absence of an anionic H-bond acceptor and the nature of the incorporating H2O as a ligand of Mn4 force the Hδ+ of the incorporating H2O to face cationic Mn1(III), Mn3(III), and Mn4(III), resulting in repulsion. The absence of D1-Asp61 near O5 would be energetically disadvantageous for incorporating H2O into the O5 site. O4 is located at the “corner” of the Mn3–O4 and O4–Mn4 bonds. Due to the presence of D1-Asp61, OW539δ– can always be oriented toward the vacant O4 site, which makes incorporation energetically favorable (Supplementary Figure 4a). On the other hand, if W2 is incorporated into the center of the Mn1–(O5)–Mn4 bond, HW2δ+ is oriented toward cationic Mn1(III) and Mn4(III), causing repulsion (Supplementary Figure 4b). Indeed, QM/MM calculations show that neither W2 nor W3 was incorporated into the vacant O5 site of the O5-depleted Mn4CaO4 cluster. Depletion of O5 did not even affect the H-bond network near O5, including the short low-barrier H-bond between TyrZ and D1-His190 (~2.5 Å)
W539 forms an H-bond with D1-Ser169 (OW539–OD1-Ser169 = 2.73 Å), which has been proposed to provide access to water molecules for the substrate-binding site42. Incorporation of W539 into the O4 site was also observed in QM/MM calculations using the original PSII crystal structure and depleting the O4 atom (Fig. 6a); the obtained geometry was consistent with the O4-incorporated geometry obtained through the S2, S3, and S4 states (Fig. 4), which indicates the robustness of the present reaction scheme.
QM/MM geometries in the O4-depleted and O5-depleted PSII proteins. a Formation of the Mn3–OW539 and Mn4–OW539 bonds in response to incorporation of H2O at W539 into the O4 site of the O4-depleted Mn4CaO4 cluster. b Shorting the Mn3–OW539 and Mn4–OW539 bonds in response to incorporation of H2O into the vacant W539 site. c QM/MM geometries in the O5-depleted PSII. (Mn1, Mn2, Mn3, Mn4) = (III, IV, III, III)
As H2O at W539 was incorporated into the Mn4CaO4 cluster, the W539 site became vacant. When the W539 site was refilled by a water molecule, both Mn3–OW539 and Mn4–OW539 were further shortened to ~2.2 Å (Fig. 6b). It seems likely that incorporation of a water molecule via D1-Asp61 into the W539 site16 led to fixation of O4 as a component of the Mn4CaO5 cluster.
Alteration of the H-bond pattern along the O4-water chain
QM/MM calculations indicated that O4 may exist as OH– in S0, and release of the proton occurs in the S0-to-S1 transition along the O4-water chain (Fig. 1), with transforming of the pre-PT to a post-PT H-bond pattern4. This implies that the pre-PT pattern must be recovered from the post-PT pattern before the next turnover.
As W539 approached Mn3 during O2 evolution, the H-bond pattern along the O4-water chain (e.g., W538) transformed from the post-PT pattern to the pre-PT pattern, since W539 (i.e., new O4) needs to reorient to form a μ-oxo bridge (Mn3–OW539–Mn4) (Fig. 7). Given that release of the first proton from H2O at O4 occurs via D1-Asp61 in the pre-S0-to-S0 transition (i.e., the O4-water chain is not used), the O4-water chain remains in the pre-PT pattern, ready for release of the proton in the S0-to-S1 transition.
The energy profiles when W539 approaches Mn3(III). As W539 approaches Mn3(III) (i.e., the Mn3…OW539 distance was decreased), concertedly (1) W n -W1 approaches Mn4 (ligating to Mn4), (2) the H-bond pattern of the O4-water chain transforms from the post-PT to pre-PT pattern, and (3) O2 releases from the Mn3 moiety. The total spin density 2S is shown for OW1=O4
The rate-limiting step in O=O formation has been reported to exhibit a kD/kH of 1.4 6, which is consistent with the kD/kH value for transformation between pre- and post-PT patterns11. This may correspond to H-bond transformation along the O4-water chain (Fig. 7), since the corresponding water chain or H-bond network is absent at the O5/O6 moiety9,17,18.
Characteristic O sites in the two-flash structure
The two-flash illuminated PSII structure shows the extremely short averaged O4…OW539 distance of 2.32 Å (2.47 and 2.17 Å)17. The present QM/MM calculations reproduced the short O4–OW539 distance (2.43 Å) when H2O at W539 was incorporated into the O4 site and the vacant W539 site was refilled by H2O (pre-S0 state, Fig. 6b). The absence of W538 (i.e., disordered W538) in the two-flash illuminated PSII structure17 can be explained by the present reaction scheme, as O4 is consumed and thus incorporation of W539 and displacement of W538 occur (Fig. 6). The vacant W539-binding site (Fig. 6a) can be refilled by H2O, as MD simulations indicated that bulk water could enter the W539 and W538 sites via D1-Asp6116. Even in the Mn-depleted PSII crystal structure, both W539 and W538 exist and are well-ordered (B-factor values are 33.9 and 43.4, respectively)53, highlighting the originally large binding affinity due to D1-Asp61. This, in turn, suggests that the absence of well-ordered W538 in the two-flash illuminated PSII structure17 is exceptional (Table 2), and can be understood only when reorganization of W539 and W538 occurs.
Discussion
Based on the findings reported here, we are able to propose the following reaction scheme (Fig. 8). In S2, the H atom of H2O at W1 is significantly migrated toward D1-Asp61 in the open-cubane Mn(III,IV,IV,IV) (Fig. 2). The S2-to-S3 transition involves oxidation of OW1H–, resulting in formation of OW1•– in Mn(III,IV,IV,IV), as suggested by Messinger et al.32. In S3, the binding of W1 at Mn4(IV) is pronounced due to formation of OW1•–. The binding of W n -W1 at OW1•– is also pronounced (Fig. 3). S4 is initially Mn(IV,IV,IV,IV) with OW1•–. Oxo-oxyl radical coupling between O4 and OW1•– leads to formation of (OW1–O4)2– and reduction to Mn(IV,IV,III,IV) (Fig. 4); W n -W1 decreases the energy barrier for the (OW1–O4)2– formation, serving as a catalyst. As OW1=O4 moves away from the Mn3/Mn4 moiety toward D1-Ile60 in the proposed O2-exiting pathway48, H2O at W539 is incorporated into the vacant O4 site, forming a μ-oxo bridge with Mn3–OW539 and Mn4–OW539 in a concerted exergonic process. W n -W1 is also incorporated into the Mn4CaO5 cluster as the new W1 ligand. The two Mn3–OW539 and Mn4–OW539 bonds are further shortened when the vacant W539 site is refilled by a water molecule, which approaches via D1-Asp6116, recovering the original Mn4CaO5 structure. Incorporation of W539 into the O4 site also seems to lead to transformation of the H-bond pattern along the O4-water chain to the pre-PT pattern (Fig. 7), in readiness for proton transfer in the S0-to-S1 transition4.
Mechanism of O2 formation and recovery of Mn4CaO5. S0 is the lowest oxidation state. In the S0-to-S1 transition, a low-barrier H-bond between OH– at O4 and W539 (as indicated in the crystal structure9) releases the proton via the O4-water chain4. In the O4-water chain, the pre-PT pattern transforms into the post PT pattern as a result of proton transfer. In the S1-to-S2 transition, oxidation of Mn4(III) to Mn4(IV) occurs, resulting in a decrease in pKa of W1 and formation of a low-barrier H-bond between W1 and D1-Asp61. In the S2-to-S3 transition, D1-Asp61 decreases the redox potential of W1 and facilitates proton-coupled oxidation of OW1H– to OW1•–. OW1•– attracts W n -W1. In S4, oxo-oxyl radical coupling occurs between OW1•– and corner μ-oxo O4, leading to formation of (OW1–O4)2–, reduction of Mn, incorporation of W n -W1 as new W1, and approach of W539. In the S4-to-pre-S0 transition, as OW1=O4 moves away from the Mn4CaO5 moiety, water incorporation occurs near two second sphere ligands, D1-Asp61 and CP43-Arg357: D1-Asp61 accepts an H-bond from W539 and incorporation of W539 into the vacant O4 site near CP43-Arg357 leads to reorientation of W539 and formation of a μ-oxo bridge with Mn3–OW539 and Mn4–OW539. Reorientation of W539 (i.e., new O4) at the O4 site propagates along the O4-water chain, transforming the post-PT pattern to the pre-PT pattern. The vacant W539 site can be filled by a water molecule from the D1-Glu65/D2-Glu312 channel via D1-Asp6116. Pre-S0, where H2O is present at O4, can be stabilized by release of a proton from the newly incorporated O4 site and proceeds to S0
The proton releasing sites are identified as W1 in the S2-to-S3 and the S3-to-S0 transitions and O4 in the pre-S0-to-S0 and S0-to-S1 4 transitions. Based on these observations, W1 and O4 are deprotonatable substrate water molecules on the current turnover, and W n -W1 and W539 serve as “holding sites7” and become W1 and O4, respectively, on the next turnover. Then, the D1-Glu65/D2-Glu312 channel serves as a water intake channel for both substrate water molecules16. The location of O4/W539 at the dead end of the narrow region of the D1-Glu65/D2-Glu312 channel16 may also make the exchange rate slow.
If the fast-exchanging and slow-exchanging water molecules represent two substrate water molecules, they might be W1 and O4, respectively. Then, the decrease in the exchange rate for the fast-exchanging water molecule in the S2-to-S3 transition12 might be explained by the pronounced binding of W1 at Mn4 (OW1•– in S3) (Supplementary Figure 5). However, as suggested by Noguchi and coworkers based on FTIR difference spectroscopy, it seems also possible that the exchanging water molecule is not necessarily a substrate water molecules on the current turnover (W1 and O4 in the present reaction scheme) but rather a substrate water molecule on the next turnover (W n -W1 as next W1 and W539 as next O4)41. In this case, the increase in the exchange rate of the slow-exchanging water molecule (i.e., W539) in the S1-to-S2 transition12 is due to the relaxed H-bond network of the O4-water chain (including O4 and W539) in the S1-to-S2 transition (i.e., after proton transfer) with respect to the strongly coupled H-bond network in the S0-to-S1 transition4. The decrease in the exchange rate for the fast-exchanging water molecule (i.e., W n -W1) in the S2-to-S3 transition12 can be explained by the pronounced binding of W n -W1 at OW1•–.
W n -W1, a substrate on the next turnover, decreases the energy barrier for (OW1–O4)2– formation in S4, serving as a catalyst. W539, another substrate on the next turnover, is the direct proton acceptor for the current turnover substrate H2O at O4. The remarkable cooperativity of the next substrate water molecules to catalyze the current substrate water molecule (Fig. 4b), as well as the self-recovery of the Mn4CaO5 cluster from the O2-evolved Mn4CaO4 cluster in a concerted exergonic process (Fig. 4a), suggest that W1/ W n -W1 and O4/W539 are substrate water molecules.
Methods
Coordinates and atomic partial charges
The atomic coordinates of PSII were taken from the X-ray structure of PSII monomer unit “A” of the PSII complexes from Thermosynechococcus vulcanus at a resolution of 1.9 Å (PDB code, 3ARC)9. During optimization of hydrogen atom positions with CHARMM54, the positions of all heavy atoms were fixed and all titratable groups (e.g., acidic and basic groups) were ionized. In QM/MM calculations, additional counter ions were added to neutralize the entire system. Atomic partial charges of the amino acids and cofactors were obtained from the CHARMM2255 parameter set and our previous studies on PSII4, respectively. D1-His337 was considered to be protonated56.
QM/MM calculations
The unrestricted density functional theory (DFT) method was employed with the B3LYP functional and LACVP* basis sets, using the QSite57 program. In the QM region, all the atomic coordinates were fully relaxed. In the MM region, the positions of H atoms were optimized using the OPLS2005 force field, while the positions of heavy atoms were fixed. The cluster was considered to be in the S2, S3, S4, and S0 states with antiferromagnetically coupled Mn ions; the resulting Mn oxidation states (Mn1, Mn2, Mn3, Mn4) and the total spin S were as follows: (III, IV, IV, IV) and S = 7/2 (↑↓↑↑) in S2; (III, IV, IV, IV) + OW1•– and S = 8/2 (↑↓↑↑ + ↑) in S3; (IV, IV, IV, IV) + OW1•– and S = 7/2 (↑↓↑↑ + ↑), (IV, IV, III, IV) + (OW1–O4)2– and S = 7/2 (↑↓↑↑), and (III, IV, III, III) + OW1=O4 and (↑↓↑↑ + ↓) in S4; (III, IV, III, III) and S = 9/2 (↑↓↑↑) in pre-S0 (Table 1, Supplementary Figures 1 and 3 for other spin configurations). The Mn4CaO5 geometry in S n was obtained as follows: first we prepared the QM/MM-optimized S n geometry with ferromagnetically coupled Mn ions, using the QM/MM-optimized S n –1 geometry. The resulting S n geometry was then optimized as antiferromagnetically coupled Mn ions. The initial-guess wave functions were obtained by using the ligand-field theory58 implemented in the QSite program. See Supplementary Data 1 for the atomic coordinates.
To analyze S2, S3 (Fig. 2), and S4 (Fig. 4), the QM region (small QM) was defined as the Mn4CaO5 cluster (including the ligand side-chains of D1-Asp170, D1-Glu189, D1-His332, D1-Glu333, D1-Asp342, and CP43-Glu354, and the backbone of D1-Ala344); the ligand water molecules of W1-W4; the O4-water chain (W539, W538, and W393); the W n -W1-binding site (W n -W1, W426, and W614); the Cl-1-binding site (Cl-1, W442, W446, and side-chains of D1-Asn181 and D2-Lys317); and the second sphere ligands (side-chains of D1-Asp61 and CP43-Arg357). Other groups were approximated by the MM force field. To mainly analyze the O4-depleted and O5-depleted PSII structures in S0 (Fig. 6), the QM region (large QM) was defined as the Mn4CaO5 cluster (including the ligand side-chains of D1-Asp170, D1-Glu189, D1-His332, D1-Glu333, D1-Asp342, and CP43-Glu354, and the backbone of D1-Ala344); the ligand water molecules of W1-W4; the O4-water chain (W539); the Cl-1-binding site (Cl-1, W442, W446, and side-chains of D1-Asn181 and D2-Lys317); the second sphere ligands (side-chains of D1-Asp61 and CP43-Arg357); the H-bond network of TyrZ (side-chains of D1-Tyr161, D1-His190, and D1-Asn298), and the diamond-shaped cluster of water molecules59 (W5, W6, and W7).
To obtain the potential-energy profiles of H-bonds (Fig. 2b and Supplementary Fig. 1), the QM/MM optimized geometry was used as the initial geometry. The H atom under investigation was moved from the H-bond donor atom (Odonor) toward the acceptor atom (Oacceptor) by 0.05 Å, after which the geometry was optimized by constraining the Odonor–H and H–Oacceptor distances and the energy was calculated. This procedure was repeated until the H atom reached the Oacceptor atom. To obtain the potential-energy profiles of (OW1–O4)2– formation (Fig. 4b and Supplementary Fig. 3) and O2 release (Fig. 7 and Supplementary Fig. 4), the QM/MM optimized geometry was used as the initial geometry; e.g., the OW1…O4 distance was then decreased by 0.05 Å, after which the geometry was optimized by constraining the OW1…O4 distance and the energy was calculated. This procedure was repeated until formation of (OW1–O4)2–.
MD simulations
The PSII structure was embedded in a lipid bilayer, which is composed of 546 1-palmitoyl-2-oleyl-sn-glycero-3-phosphocholine (POPC), and soaked in 78889 flexible water molecules (SPC-Fw)60, using the CHARMM-GUI program61. After geometry optimization with position restraints on heavy atoms, the system was heated from 0.001 K to 300 K during 5.0 ps. The position restraints on heavy atoms were gradually released over 16.5 ns. An equilibrating MD run was conducted for 45 ns using the MD engine AMBER 1462 with the SHAKE algorithm for hydrogen constraint63. The production run was conducted with an MD time step of 0.5 fs without hydrogen constraint, using AMBER 14. To control temperature and pressure, the Berendsen thermostat and barostat were employed 65. See ref. 16 for force field parameters.
Data availability
All the data supporting the findings of this study are available within the article and its Supplementary Information files or from the corresponding author upon reasonable request.
References
Dau, H., Zaharieva, I. & Haumann, M. Recent developments in research on water oxidation by photosystem II. Curr. Opin. Chem. Biol. 16, 3–10 (2012).
Cox, N. & Messinger, J. Reflections on substrate water and dioxygen formation. Biochim. Biophys. Acta 1827, 1020–1030 (2013).
Shen, J. R. The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu. Rev. Plant. Biol. 66, 23–48 (2015).
Saito, K., Rutherford, A. W. & Ishikita, H. Energetics of proton release on the first oxidation step in the water-oxidizing enzyme. Nat. Commun. 6, 8488 (2015).
Dau, H. & Haumann, M. The manganese complex of photosystem II in its reaction cycle–Basic framework and possible realization at the atomic level. Coord. Chem. Rev. 252, 273–295 (2008).
Haumann, M., Bögershausen, O., Cherepanov, D., Ahlbrink, R. & Junge, W. Photosynthetic oxygen evolution: H/D isotope effects and the coupling between electron and proton transfer during the redox reactions at the oxidizing side of Photosystem II. Photosynth. Res. 193, 193–208 (1997).
Debus, R. J. Evidence from FTIR difference spectroscopy that D1-Asp61 influences the water reactions of the oxygen-evolving Mn4CaO5 cluster of photosystem II. Biochemistry 53, 2941–2955 (2014).
Renger, G. & Hanssum, B. Studies on the reaction coordinates of the water oxidase in PS II membrane fragments from spinach. FEBS Lett. 299, 28–32 (1992).
Umena, Y., Kawakami, K., Shen, J.-R. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011).
Noguchi, T. Fourier transform infrared difference and time-resolved infrared detection of the electron and proton transfer dynamics in photosynthetic water oxidation. Biochim. Biophys. Acta 1847, 35–45 (2015).
Agmon, N. The grotthuss mechanism. Chem. Phys. Lett. 244, 456–462 (1995).
Hillier, W. & Wydrzynski, T. The affinities for the two substrate water binding sites in the O2 evolving complex of photosystem II vary independently during S-state turnover. Biochemistry 39, 4399–4405 (2000).
Messinger, J. Evaluation of different mechanistic proposals for water oxidation in photosynthesis on the basis of Mn4OxCa structures for the catalytic site and spectroscopic data. Phys. Chem. Chem. Phys. 6, 4764–4771 (2004).
Hillier, W. & Wydrzynski, T. Substrate water interactions within the Photosystem II oxygen evolving complex. Phys. Chem. Chem. Phys. 6, 4882–4889 (2004).
Singh, S., Debus, R. J., Wydrzynski, T. & Hillier, W. Investigation of substrate water interactions at the high-affinity Mn site in the photosystem II oxygen-evolving complex. Philos. Trans. R. Soc. Lond. B 363, 1229–1235 (2008).
Sakashita, N., Watanabe, H. C., Ikeda, T., Saito, K. & Ishikita, H. Origins of water molecules in the photosystem II crystal structure. Biochemistry 56, 3049–3057 (2017).
Suga, M. et al. Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543, 131–135 (2017).
Suga, M. et al. Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 517, 99–103 (2015).
Siegbahn, P. E. Structures and energetics for O2 formation in photosystem II. Acc. Chem. Res. 42, 1871–1880 (2009).
Isobe, H. et al. Theoretical illumination of water-inserted structures of the CaMn4O5 cluster in the S2 and S3 states of oxygen-evolving complex of photosystem II: full geometry optimizations by B3LYP hybrid density functional. Dalton Trans. 41, 13727–13740 (2012).
Saito, K. & Ishikita, H. Influence of the Ca2+ ion on the Mn4Ca conformation and the H-bond network arrangement in Photosystem II. Biochim. Biophys. Acta 1837, 159–166 (2014).
Yang, J., Hatakeyama, M., Ogata, K., Nakamura, S. & Li, C. Theoretical study on the role of Ca2+ at the S2 state in photosystem II. J. Phys. Chem. B 118, 14215–14222 (2014).
Amin, M., Pokhrel, R., Brudvig, G. W., Badawi, A. & Obayya, S. S. Effect of chloride depletion on the magnetic properties and the redox leveling of the oxygen-evolving complex in photosystem II. J. Phys. Chem. B 120, 4243–4248 (2016).
Perrin, C. L. & Nielson, J. B. “Strong” hydrogen bonds in chemistry and biology. Annu. Rev. Phys. Chem. 48, 511–544 (1997).
Schutz, C. N. & Warshel, A. The low barrier hydrogen bond (LBHB) proposal revisited: the case of the Asp… His pair in serine proteases. Proteins 55, 711–723 (2004).
Ishikita, H. & Saito, K. Proton transfer reactions and hydrogen-bond networks in protein environments. J. R. Soc. Interface 11, 20130518 (2014).
Ikeda, T., Saito, K., Hasegawa, R. & Ishikita, H. The existence of an isolated hydronium ion in the interior of proteins. Angew. Chem. Int. Ed. 56, 9151–9154 (2017).
Saito, K., Rutherford, A. W. & Ishikita, H. Mechanism of tyrosine D oxidation in Photosystem II. Proc. Natl Acad. Sci. USA 110, 7690–7695 (2013).
Narzi, D., Bovi, D. & Guidoni, L. Pathway for Mn-cluster oxidation by tyrosine-Z in the S2 state of photosystem II. Proc. Natl Acad. Sci. USA 111, 8723–8728 (2014).
Rabani, J. & Matheson, M. S. Pulse radiolytic determination of pK for hydroxyl ionic dissociation in water. J. Am. Chem. Soc. 86, 3175–3176 (1964).
Cox, N. et al. Photosynthesis. Electronic structure of the oxygen-evolving complex in photosystem II prior to O-O bond formation. Science 345, 804–808 (2014).
Messinger, J. et al. Absence of Mn-centered oxidation in the S2 → S3 transition: implications for the mechanism of photosynthetic water oxidation. J. Am. Chem. Soc. 123, 7804–7820 (2001).
Renger, G. Mechanism of light induced water splitting in Photosystem II of oxygen evolving photosynthetic organisms. Biochim. Biophys. Acta 1817, 1164–1176 (2012).
Yano, J. & Yachandra, V. Mn4Ca cluster in photosynthesis: where and how water is oxidized to dioxygen. Chem. Rev. 114, 4175–4205 (2014).
Siegbahn, P. E. Substrate water exchange for the oxygen evolving complex in PSII in the S1, S2, and S3 states. J. Am. Chem. Soc. 135, 9442–9449 (2013).
Isobe, H., Shoji, M., Shen, J.-R. & Yamaguchi, K. Strong coupling between the hydrogen bonding environment and redox chemistry during the S2 to S3 transition in the oxygen-evolving complex of photosystem II. J. Phys. Chem. B 119, 13922–13933 (2015).
Askerka, M., Wang, J., Vinyard, D. J., Brudvig, G. W. & Batista, V. S. S3 state of the O2-evolving complex of photosystem II: insights from QM/MM, EXAFS, and femtosecond X-ray diffraction. Biochemistry 55, 981–984 (2016).
Retegan, M. et al. A five-coordinate Mn(IV) intermediate in biological water oxidation: spectroscopic signature and a pivot mechanism for water binding. Chem. Sci. 7, 72–84 (2016).
Ugur, I., Rutherford, A. W. & Kaila, V. R. I. Redox-coupled substrate water reorganization in the active site of Photosystem II—The role of calcium in substrate water delivery. Biochim. Biophys. Acta 1857, 740–748 (2016).
Hayashi, T., Yamaguchi, A., Hashimoto, K. & Nakamura, R. Stability of organic compounds on the oxygen-evolving center of photosystem II and manganese oxide water oxidation catalysts. Chem. Commun. 52, 13760–13763 (2016).
Suzuki, H., Sugiura, M. & Noguchi, T. Monitoring water reactions during the S-state cycle of the photosynthetic water-oxidizing center: detection of the DOD bending vibrations by means of Fourier transform infrared spectroscopy. Biochemistry 47, 11024–11030 (2008).
Vassiliev, S., Zaraiskaya, T. & Bruce, D. Exploring the energetics of water permeation in photosystem II by multiple steered molecular dynamics simulations. Biochim. Biophys. Acta 1817, 1671–1678 (2012).
Siegbahn, P. E. M. Nucleophilic water attack is not a possible mechanism for O–O bond formation in photosystem II. Proc. Natl Acad. Sci. USA 114, 4966–4968 (2017).
Shoji, M. et al. Large-scale QM/MM calculations of the CaMn4O5 cluster in the S3 state of the oxygen evolving complex of photosystem II. Comparison between water-inserted and no water-inserted structures. Faraday Discuss. 198, 83–106 (2017).
Li, X. & Siegbahn, P. E. M. Alternative mechanisms for O2 release and O-O bond formation in the oxygen evolving complex of photosystem II. Phys. Chem. Chem. Phys. 17, 12168–12174 (2015).
Ferreira, K. N., Iverson, T. M., Maghlaoui, K., Barber, J. & Iwata, S. Architecture of the photosynthetic oxygen-evolving center. Science 303, 1831–1838 (2004).
Zhang, C. et al. Inorganic chemistry. A synthetic Mn4Ca-cluster mimicking the oxygen-evolving center of photosynthesis. Science 348, 690–693 (2015).
Gabdulkhakov, A. G., Kljashtorny, V. G. & Dontsova, M. V. Analysis of molecular oxygen exit pathways in cyanobacterial photosystem II: molecular dynamics studies. Crystallogr. Rep. 60, 884–888 (2015).
Dilbeck, P. L., Bao, H., Neveu, C. L. & Burnap, R. L. Perturbing the water cavity surrounding the manganese cluster by mutating the residue D1-valine 185 has a strong effect on the water oxidation mechanism of photosystem II. Biochemistry 52, 6824–6833 (2013).
Hundelt, M., Hays, A.-M. A., Debus, R. J. & Junge, W. Oxygenic photosystem II: the mutation D1-D61N in Synechocystis sp. PCC 6803 retards S-state transitions without affecting electron transfer from YZ to P680 +. Biochemistry 37, 14450–14456 (1998).
Dilbeck, P. L. et al. The D1-D61N mutation in Synechocystis sp. PCC 6803 allows the observation of pH-sensitive intermediates in the formation and release of O2 from photosystem II. Biochemistry 51, 1079–1091 (2012).
Bao, H. & Burnap, R. L. Structural rearrangements preceding dioxygen formation by the water oxidation complex of photosystem II. Proc. Natl Acad. Sci. USA 112, E6139–E6147 (2015).
Zhang, M. et al. Structural insights into the light-driven auto-assembly process of the water-oxidizing Mn4CaO5-cluster in photosystem II. eLife 6, e26933 (2017).
Brooks, B. R. et al. CHARMM: a program for macromolecular energy minimization and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983).
MacKerell, A. D. Jr et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Nakamura, S. & Noguchi, T. Infrared determination of the protonation state of a key histidine residue in the photosynthetic water oxidizing center. J. Am. Chem. Soc. 139, 9364–9375 (2017).
QSite v.5.8 (Schrödinger, LLC, New York, 2012).
Vacek, G., Perry, J. K. & Langlois, J. M. Advanced initial-guess algorithm for self-consistent-field calculations on organometallic systems. Chem. Phys. Lett. 310, 189–194 (1999).
Saito, K., Shen, J.-R., Ishida, T. & Ishikita, H. Short hydrogen-bond between redox-active tyrosine YZ and D1-His190 in the photosystem II crystal structure. Biochemistry 50, 9836–9844 (2011).
Wu, Y. J., Tepper, H. L. & Voth, G. A. Flexible simple point-charge water model with improved liquid-state properties. J. Chem. Phys. 124, 024503 (2006).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: A web-bsed graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Case, D. A. et al. AMBER 14, University of California, San Francisco (2014).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular-dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Acknowledgements
This research was supported by the JST CREST (JPMJCR1656), JSPS KAKENHI (JP26800224 to K.S., JP16H06560 to K.S. and H.I., and JP26105012 to H.I.), Japan Agency for Medical Research and Development (AMED), Materials Integration for engineering polymers of Cross-ministerial Strategic Innovation Promotion Program (SIP), and Interdisciplinary Computational Science Program in CCS, University of Tsukuba.
Author information
Authors and Affiliations
Contributions
H.I. designed research; K.K., T.T., H.K., and K.S. performed research; K.K. and H.I. analyzed data; and K.K. and H.I. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kawashima, K., Takaoka, T., Kimura, H. et al. O2 evolution and recovery of the water-oxidizing enzyme. Nat Commun 9, 1247 (2018). https://doi.org/10.1038/s41467-018-03545-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-03545-w
This article is cited by
-
Closing Kok’s cycle of nature’s water oxidation catalysis
Nature Communications (2024)
-
Binding and functions of the two chloride ions in the oxygen-evolving center of photosystem II
Photosynthesis Research (2022)
-
Role of redox-inactive metals in controlling the redox potential of heterometallic manganese–oxido clusters
Photosynthesis Research (2021)
-
Structural dynamics in the water and proton channels of photosystem II during the S2 to S3 transition
Nature Communications (2021)
-
pKa of the ligand water molecules in the oxygen-evolving Mn4CaO5 cluster in photosystem II
Communications Chemistry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.