Abstract
Temporal and spatial-specific regulation of pluripotency networks is largely dependent on the precise modifications of core transcription factors. Misregulation of glutamylation is implicated in severe physiological abnormalities. However, how glutamylation regulates cell reprogramming and pluripotency networks remains elusive. Here we show that cytosolic carboxypeptidases 1 (CCP1) or CCP6 deficiency substantially promotes induced pluripotent cell (iPSC) induction and pluripotency of embryonic stem cells (ESCs). Klf4 polyglutamylation at Glu381 by tubulin tyrosine ligase-like 4 (TTLL4) and TTLL1 during cell reprogramming impedes its lysine 48-linked ubiquitination and sustains Klf4 stability. Klf4-E381A knockin mice display impaired blastocyst development and embryonic lethality. Deletion of TTLL4 or TTLL1 abrogates cell reprogramming and early embryogenesis. Thus, Klf4 polyglutamylation plays a critical role in the regulation of cell reprogramming and pluripotency maintenance.
Similar content being viewed by others
Introduction
Reprogramming resets differentiated somatic cells to a pluripotent state, which can be achieved by nuclear transfer, cell fusion, and transduction of transcription factors1. Somatic cells can be reprogrammed to induced pluripotent cells (iPSCs) by expressing pluripotency factors Oct4, Sox2, Klf4, and c-Myc (termed OSKM)2,3. The generation of iPSCs can be derived from patient tissues and has great potential for regenerative medicine and cell replacement therapies4,5. Several hurdles, including low frequency of iPSC induction and genomic instability of iPSCs, need to be solved prior to development of a safe iPSC technology. However, the molecular mechanisms underlying reprogramming still remain ill-defined. The temporal and spatial-specific regulation of pluripotency networks largely depends on precise modifications and interaction controls of the core transcriptional factors6,7,8,9. These reprogramming factors are highly modified post-transcriptionally at the levels of mRNA stability, translation and protein activity7,10.
Protein post-translational modifications (PTMs) such as phosphorylation, acetylation, glycosylation, and ubiquitination play important roles in the regulation of activities of target proteins by changing their chemical or structural properties11,12. In-depth quantitative and dynamic proteomic studies reveal that PTMs occur on core transcription factors during the process of pluripotency maintenance and reprogramming7. Transcriptional and DNA-binding activities of Oct4 and Sox2 are regulated by phosphorylation, which exert considerable effect on pluripotency maintenance and iPSC generation7,13. Acetylation of Sox2 is critical for pluripotency control by regulating its nuclear export and protein stability14. O-GlcNacylation directly regulates transcriptional activities of Oct4 and Sox2 in maintaining pluripotency and cell reprogramming9,15. Moreover, ubiquitination of Klf4 and Oct4 modulates their half-life and subsequent protein stability16,17. It has been reported that B cells treated with C/EBPα can be efficiently reprogrammed into iPSCs by OSKM induction through enhancing chromatin accessibility and Klf4 stability18. Therefore, PTMs of reprogramming factors play critical roles in determining the cell fate decision of stem cells.
Glutamylation, a unique PTM, adds glutamate side chains onto the γ-carboxyl groups of glutamic acid residues in the primary sequence of target proteins19,20,21. Polyglutamylation of tubulins, well-known targets of glutamylation, regulates the interaction between microtubules (MTs) and their partners19, modulating MT-related processes such as ciliary motility, neurite outgrowth and neurodegeneration21,22,23,24. Glutamylation is catalyzed by polyglutamylases, also called as tubulin tyrosine ligase-like (TTLL) enzymes25,26. Glutamylation is a reversible modification that can be hydrolyzed by a family of cytosolic carboxypeptidases (CCPs)23. Misregulation of glutamylation causes several physiological abnormalities. CCP1 deficiency causes hyperglutamylation of tubulins, resulting in Purkinje cell degeneration23,27,28. We recently demonstrated that CCP6 deficiency induces hyperglutamylation of Mad2, leading to underdevelopment of megakaryocytes and abnormal thrombocytosis29. In addition, mutations of TTLL polyglutamylases are also implicated in severe disorders such as retinal dystrophy30 and pancreatic oncogenesis31. However, how glutamylation regulates cell reprogramming and pluripotency maintenance remains elusive.
The Kruppel-like factor (Klf4), together with other three Yamanaka pluripotency factors, is able to reprogram adult fibroblasts into iPSCs2,3,32, which initiates somatic gene suppression in an early phase and pluripotency gene activation in a late phase during reprogramming33,34. Klf4 is highly expressed in mouse embryonic stem cells (ESCs) and rapidly downregulated in the early stage of differentiation35. Klf4 can be phosphorylated by ERKs, whose phosphorylation recruits βTrCPs (components of a ubiquitin E3 ligase) to ubiquitinate Klf4 for its degradation16. However, the precise modification crosstalk of Klf4 in reprogramming has not clearly defined yet.
In this study, we show that CCP1 or CCP6 deficiency substantially promotes iPSC generation. Klf4 is polyglutamylated by TTLL4 or TTLL1 that impedes its lysine (K) 48-linked ubiquitination to maintain Klf4 stability. Deletion of TTLL4 or TTLL1 impairs cell reprogramming and early embryogenesis.
Results
CCP6 or CCP1 deficiency promotes somatic cell reprogramming
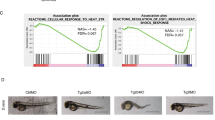
We previously demonstrated that Ccp6 (official gene name Agbl4, referred to here as Ccp6)-deficient mice exhibit underdevelopment of megakaryocytes and abnormal thrombocytosis29. We next wanted to further explore whether glutamylation modifications were involved in the regulation of cell reprogramming. We noticed that CCP6 deficiency caused higher litter size at birth (Supplementary Figure 1a), whereas Ccp6-deficient mice displayed similar sperm and ovulation numbers compared with littermate control mice, suggesting CCP6 could be implicated in the modulation of cellular reprogramming. To determine whether glutamylation is involved in somatic cell reprogramming, we first generated Ccp1−/− (official gene name Agtpbp1, referred to here as Ccp1), or Ccp6−/− mouse embryonic fibroblasts (MEFs) (Fig. 1a), and transduced Yamanaka factors OSKM for iPSC induction assays. We noticed that Ccp1−/− and Ccp6−/− MEFs produced more iPSC colonies compared with WT MEFs (Fig. 1b). These iPSCs generated from Ccp1−/− or Ccp6−/− MEFs expressed elevated pluripotent stem cell surface marker SSEA-1 and increased pluripotent genes without partial differentiation (Fig. 1c, and Supplementary Figure 1b, c). These reprogrammed iPSCs expressed similar levels of pluripotent genes compared to ESCs, validating the reprogramming system was efficient (Supplementary Figure 1b). Moreover, the iPSCs derived from Ccp1−/− or Ccp6−/− MEFs were capable of forming teratomas containing cells of all three germ layers and displayed faster growth rates (Fig. 1d). Importantly, Ccp1 and Ccp6 double knockout (DKO) MEFs showed higher reprogramming efficiency (Fig. 1b), as well as pluripotent gene expression than Ccp1−/− or Ccp6−/− MEFs alone (Supplementary Figure 1b). In addition, restoration of CCP1 or CCP6 into Ccp1−/− or Ccp6−/− MEFs still went back to a low frequency of iPSC generation (Fig. 1b and Supplementary Figure 1b). Of note, deficiency of CCP1 or CCP6 in feeder-free iPSCs really exhibited elevated pluripotent gene expression, whereas deficiency of CCP1 or CCP6 in MEFs alone did not affect pluripotent gene expression (Supplementary Figure 1d). Overall, these results indicate that deficiency of CCP1 or CCP6 promotes somatic cell reprogramming.
CCP6 or CCP1 deficiency promotes cell reprogramming. a Ccp1- or Ccp6- deficient MEFs were lyzed for immunoblotting. β-actin was used as a loading control. b WT or CCPs-deficient MEFs were infected by OSKM factors containing retrovirus and cultured in ESC media for 3 weeks. Alkaline phosphatase (AP)-positive colony numbers per 104 cells were calculated and shown as means ± S.D. **, P < 0.01. Scale bar, 50 μm. n = 5. c SSEA-1 was assayed by flow cytometry in iPSCs derived from Ccp1−/− or Ccp6−/− MEFs. d iPSCs from Ccp1−/− or Ccp6−/− MEFs were injected subcutaneously into NOD/SCID mice. One month later, teratomas were collected for H&E staining. Teratomas volumes were calculated and shown as means ± S.D. (left panel). Red arrow denotes teratomas in left panels. Black arrow, endoderm; red arrow, mesoderm; red arrowhead, ectoderm. Scale bar, 200 μm. Total 6 teratomas from 6 mice were analyzed pre condition (n = 6). The teratomas were from 2 iPSC cell lines. e 4F2A MEFs were treated with doxycycline (dox) (2 μg/ml), together with CoCl2 (10 μM) or phenanthroline (Phen, 1 μM) in ESC media for iPSC formation as in b. Reprogramming efficiency was assayed by Nanog staining after dox removal. Scale bar, 50 μm. Nanog-positive colony numbers per 104 cells were calculated and shown as means ± S.D.**P < 0.01. n = 5. f CCP6 deficiency promoted blastocyst development. Embryos were isolated at E1.5 stage and cultured in KSOM media supplied with 1 mg/ml BSA at 37 °C for 48 h. Distribution of embryonic stage at E3.5 was counted as mean ± S.D. (right panel), **P < 0.01. Scale bar, 100 μm. 105 Ccp6+/+ embryos and 122 Ccp6−/− embryos were observed. n = 4. g Embryos were isolated at E1.5 stage and cultured in KSOM media with CoCl2 (10 μM) or phenanthroline (1 μM). Distribution of embryonic stage at E3.5 was counted as means ± S.D (right panel), **P < 0.01. Scale bar, 100 μm. 131 mock-treated embryos, 118 CoCl2-treated embryos and 132 phen-treated embryos were observed. n = 4. Student’s t test was used as statistical analysis. oe overexpression, ns no significance
To further determine the physiological role of CCP1 and CCP6 in the process of reprogramming, we silenced CCP1 and CCP6 expression in MEFs with transfection of OSKM, and found CCP1 and CCP6 depletion enhanced alkaline phosphatase (AP)-positive iPSC colony formation and pluripotent gene expression (Supplementary Figure 1e-g). By contrast, overexpression of CCP1 and CCP6 impaired iPSC colony formation as well as downregulated pluripotent gene expression (Supplementary Figure 1e-g). Of note, depletion and overexpression of CCP1 and CCP6 in MEFs did not affect growth rates of MEFs (Supplementary Figure 1h). We also treated MEFs with CCP family protein agonist CoCl236 and inhibitor phenanthroline23 after OSKM induction. Consistently, the agonist CoCl2 abrogated iPSC formation, whereas the inhibitor phenanthroline remarkably enhanced iPSC generation (Fig. 1e and Supplementary Figure 1i). These data further confirm that loss of CCP1 or CCP6 virtually enhances cell reprogramming.
Fertilization initiates cellular reprogramming in zygote and subsequent blastocyst development, which also requires the establishment of pluripotency37,38. Since homozygous Ccp1-deficient mice are male sterile27, we then assessed the effect of Ccp6 deficiency on blastocyst development. We isolated 2-cell-stage embryos from Ccp6+/+ and Ccp6−/− pregnant mice and cultured the embryos ex vivo to allow following development. CCP6 deficiency substantially promoted blastocyst development at embryo day (E) 3.5 compared to WT mice (Fig. 1f). In parallel, expression levels of pluripotent genes were dramatically increased in Ccp6−/− embryos (Supplementary Figure 1j-k). To further determine the physiological relevance of glutamylation, we isolated 2-cell-stage embryos from C57BL6/J mice and cultured them ex vivo with CoCl2 and phenanthroline treatment. We observed that the agonist CoCl2 treatment overtly inhibited embryonic development, while the inhibitor phenanthroline treatment remarkably enhanced blastocyst development (Fig. 1g). Taken together, glutamylation regulations are implicated in the early embryonic development.
Glutamylation is required for pluripotency maintenance
We further tested whether CCP1 or CCP6 regulates self-renewal and pluripotency maintenance in ES cells. We silenced CCP1 or CCP6 in mouse ESC D3 cells, and noticed that CCP1- or CCP6-depleted D3 cells exhibited increased colony numbers with strong AP-positive staining and upregulated pluripotent gene expression compared with control shRNA (shCtrl)-treated cells (Fig. 2a and Supplementary Figure 2a). Double knockdown of CCP1 with CCP6 induced much more colony numbers than that of CCP1 or CCP6 knockdown alone. By contrast, ectopic expression of CCP1 or CCP6 substantially declined AP-positive colony numbers and downregulated pluripotent gene expression (Fig. 2b and Supplementary Figure 2b). In parallel, CCP1 or CCP6 overexpressing ES R1 cells displayed similar observations (Supplementary Figure 2c). Of note, we noticed that depletion of CCP1 or CCP6 promoted ESC proliferation (Supplementary Figure 2d), whereas overexpression of CCP1 or CCP6 suppressed ESC proliferation (Supplementary Figure 2d). We then measured the glutamylation dynamic changes during the process of early embryonic development. GT335 antibody specifically recognizes the branching point of glutamate side chains and detects all glutamylated forms of target proteins39. We observed that GT335 staining signals were elevated during early embryonic development and GT335 staining signals mainly resided in the nucleus of 2-cell and 4–8-cell-stage embryos (Fig. 2c). TTLL members add glutamylation modifications and CCP members remove it of their substrates19,23. We observed that polyglutamylases TTLL1 and TTLL4 were highly expressed undergoing early embryonic development (Supplementary Figure 2e). However, both CCP1 and CCP6 were downregulated in 8-cell-stage embryos (Supplementary Figure 2f).
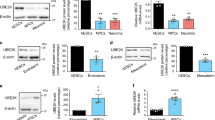
Glutamylation is required for the maintenance of mouse and human pluripotency. a Mouse ESC D3 cells were transfected with scrambled shRNA (shCtrl), shCCP1, shCCP6 or CCP1 and CCP6 overexpression (oe) plasmids as indicated and cultured in mouse ESC media. After 5 days, pluripotency was analyzed by AP staining. Colony numbers for undifferentiated, mixed or differentiated clones were calculated as means ± S.D. **P < 0.01. Scale bar, 100 μm. n = 4. b D3 cells were transfected with the indicated plasmids and assessed AP staining as in a. **P < 0.01. Scale bar, 50 μm.n = 6. c Embryos at the indicated stages were isolated and stained with GT335 antibody and PI, and visualized by confocal microscopy. Scale bar, 20 μm. Percentages of cells with GT335 signal localized in the nucleus were counted as means ± S.D. **P < 0.01. For 1-cell-stage embryos, n = 48. For 2-cell-stage embryos, n = 56. For 4- to 8-cell stage embryos, n = 62. d Human ESC H9 cells were infected with lentivirus expressing the indicated shRNAs and cultured in human ESC media for 3 weeks. Pluripotency was analyzed by AP staining. Scale bar, 100 μm. AP-positive colony numbers per well were calculated as means ± S.D. **P < 0.01. n = 5. e Depletion of CCP1 or CCP6 in human iPSCs increases AP+ colony formation. Human iPSCs were infected with lentivirus expressing the indicated shRNAs and cultured in hiPSC media for 3 weeks, followed by AP staining. Scale bar, 50 μm. AP-positive colony numbers per well were calculated as means ± S.D. **P < 0.01. n = 5. f CCP1 or CCP6 depletion in human iPSCs promotes teratomas formation. Human iPSCs infected with the indicated lentivirus were injected subcutaneously into NOD/SCID mice. Six weeks later, teratomas were collected for H&E staining. Red arrow denotes teratomas in upper panels. Black arrow, endoderm; red arrow, mesoderm; red arrowhead, ectoderm. Scale bar, 200 μm. Total 6 teratomas from 6 mice were analyzed pre condition (n = 6). The teratomas were from 2 iPSC cell lines. Student’s t test was used as statistical analysis. ns no significance
We also performed transcriptome profile assays for CCP6-depleted and shCtrl-treated ESCs. We noticed that CCP6 knockdown in ESCs caused upregulation of pluripotency transcriptional network (Supplementary Figure 2g). In addition, we analyzed RNAseq data set GSE4535240 for OSKM-induced reprogramming. We found that Ccp1 was downregulated and Ttll4 was upregulated over doxycycline-induced OSKM expression (Supplementary Figure 2h). In addition, from RNAseq data set GSE5239641, Ccp6 was downregulated during early reprogramming induction (Supplementary Figure 2h). These data suggest that glutamylation is surely involved in the regulation of cell reprogramming.
Intriguingly, similar observations were achieved in CCP1 or CCP6-silenced human ESC H9 cells and human iPSCs derived from OSKM-transduced urothelial cells (Fig. 2d,e and Supplementary Figure 2i-j). Similarly, polyglutamylases TTLL1 and TTLL4 were also highly expressed during human urothelial cell reprogramming (Supplementary Figure 2k). CCP1 and CCP6 were downregulated in human iPSCs (Supplementary Figure 2l). Moreover, CCP1- or CCP6-depleted human iPSCs were able to form teratomas with three germ layer differentiation potential and grew faster compared with shCtrl-treated iPSCs (Fig. 2f). Whereas CCP1 or CCP6 overexpressing human iPSCs impaired AP-positive colony generation and lost teratoma formation (Fig. 2f). These data suggest that glutamylation also regulates ESC and iPSC pluripotency that is evolutionally conserved in human reprogramming.
Klf4 undergoes polyglutamylation in cell reprogramming
We then measured the glutamylation dynamic changes during the process of somatic cell reprogramming. We observed that GT335 staining signals were elevated over OSKM transduction (Fig. 3a,b). Furthermore, GT335 staining signals mainly resided in the nucleus of MEFs (Fig. 3a). We observed that polyglutamylases TTLL1 and TTLL4 were highly expressed during OSKM-induced reprogramming of MEFs (Supplementary Figure 3a), whereas CCP1 and CCP6 were almost undetectable in mouse iPSCs (Supplementary Figure 3b). These observations were consistent with those of early-stage embryos (Fig. 2c, Supplementary Figure 2e-f). The specificity of GT335 antibody we used was verified (Supplementary Figure 3c). Thus, we proposed that TTLL1 and TTLL4 may mediate substrate glutamylation that is required for iPSC induction.
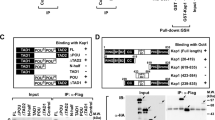
Klf4 undergoes polyglutamylation in cell reprogramming and early embryogenesis. a MEFs were transduced with STEMCCA lentivirus for the indicated days and immunostained with anti-tubulin and GT335 antibodies. Nuclei were counterstained by DAPI. Scale bar, 10 μm. Percentages of cells with GT335 signal localized in nuclei were counted as means ± S.D. **P < 0.01. More than 100 typical cells were observed and calculated. n = 134 for D0, n = 122 for D1, n = 151 for D2. b MEFs lysates induced by OSKM for the indicated days were analyzed by immunoblotting with GT335 antibody. c Recombinant CCP6-wt and inactive CCP6 mutant (CCP6-mut) were immobilized to Affi-gel10 resin. Lysates of OSKM-induced MEFs were added for affinity chromatography. The eluted fractions were visualized by SDS-PAGE followed by silver staining. The differential band around 60 kD appeared in CCP6-mut gel was cut for mass spectrometry and identified as Klf4. d MEFs Lysates induced by OSKM for the indicated days were immunoprecipitated with anti-Klf4 antibody and analyzed by immunoblotting with GT335. e Schematic representation of four putative glutamate-rich glutamate-rich regions in Klf4 (left panel). GST-Klf4wt and indicated mutant proteins were incubated with Flag-TTLL4. GST-Klf4 proteins were pulled down by Glutathione Sepharose 4B beads, followed by immunoblotting. f CCP1- and CCP6-deficient MEFs were transduced with OSKM and stained with anti-Klf4 antibody. Nuclei were stained by DAPI. Scale bar, 10 μm. Percentages of cells with Klf4/GT335 colocalization in nuclei were counted as means ± S.D. **P < 0.01. 102 Ccp1+/+ MEF cells, 112 Ccp1−/− MEF cells, 104 Ccp6+/+ MEF cells and 117 Ccp6−/− MEF cells were observed. n = 4. g CCP6-deficient embryos were isolated and immunostained with anti-Klf4 and GT335 antibodies. Percentages of cells with Klf4/GT335 colocalization in nuclei were counted as means ± S.D. **P < 0.01.110 Ccp6+/+ embryos and 116 Ccp6−/− embryos were observed. Scale bar, 20 μm. Student’s t test was used as statistical analysis. The data represent four independent experiments. MW molecular weight marker
We next wanted to identify candidate substrates for cell reprogramming. We noticed that Ccp6−/− MEF lysates with OSKM transduction displayed an around 60 kDa strong differential band compared to those of Ccp6+/+ MEFs (Supplementary Figure 3d). We then generated an enzymatically inactive mutant of CCP6 (CCP6mut) via H230S and E233Q mutations29, and performed affinity chromatography with CCP6mut-immobilized Affi-gel10 resin. Interestingly, the around 60 kDa band was identified to be Klf4 (Fig. 3c and Supplementary Figure 3e), a novel candidate substrate for CCP6 during cell reprogramming.
Expectedly, Klf4 glutamylation signals were increased after OSKM transduction via immunoprecipitation with immunoblotting assays (Fig. 3d and Supplementary Figure 4a). Since B3 antibody can detect a side chain with two or more glutamate residues42, we then used this antibody to determine monoglutamylation or polyglutamylation of glutamylated Klf4. We noticed that B3 antibody could detect glutamylated signals of Klf4 (Supplementary Figure 4b), indicating that glutamylated Klf4 is polyglutamylation. Glutamylation modifications are added at potential acceptor sites with conserved glutamate-rich stretches and flexible environment to be susceptible for catalysis43. Klf4 has four putative conserved glutamate-rich regions for glutamylation modifications (Supplementary Figure 4c). We then mutated the first glutamic residue in each putative region to alanine and generated four mutated proteins for in vitro glutamylation assays. We found that only Klf4 Glu381Ala mutant abolished TTLL4-mediated Klf4 polyglutamylation (Fig. 3e), indicating Glu381 is the acceptor site for polyglutamylation. Klf4 Glu381Ala mutant also abolished TTLL1-mediated Klf4 glutamylation (Supplementary Figure 4d). In addition, the polyglutamate chain of Klf4 could be removed by CCP1 or CCP6 with an in vitro deglutamylation assay (Supplementary Figure 4e).
We next monitored the colocalization of Klf4 and GT335 in the process of reprogramming with immunofluorescence staining. Colocalization of GT335 with Klf4 staining was observed in the nucleus of MEFs after OSKM induction (Supplementary Figure 4f). Consistently, Ccp1−/− or Ccp6−/− MEFs with OSKM transduction showed strong glutamylation signals (Fig. 3f and Supplementary Figure 4g). More importantly, Klf4 polyglutamylation also appeared in the nucleus of early embryos and enhanced after the 4- to 8-cell stage (Supplementary Figure 4h). As expected, hyperglutamylation of Klf4 also appeared in Ccp6-deficient embryos (Fig. 4g). Consistently, Klf4 polyglutamylation also appeared in mouse ESCs, and declined during the process of retinoic acid (RA)-induced differentiation (Supplementary Figure 4i). We also assessed whether the reprogramming process induced by non-OSKM factors affected endogenous Klf4 polyglutamylation. We transduced factors Sall4, Nanog, Esrrb, and Lin28 (termed SNEL) into Ccp6−/− MEFs for iPSC induction. In parallel, SNEL factor transduction in Ccp6−/− MEFs really enhanced hyperglutamylation of Klf4 as well, and consequently generated much more iPSC colonies compared to that of Ccp6+/+ MEFs (Supplementary Figure 4j). Taken together, the polyglutamylation modification of Klf4 plays a critical role in the modulation of cell reprogramming and pluripotency.
Klf4 glutamylation impedes K48-linked ubiquitination to sustain its stability. a Ccp1- and Ccp6-deficient MEFs were treated with cycloheximide (CHX) (20 μg/ml) for the indicated time and followed by immunoblotting. Percentage of residual protein levels was counted as means ± S.D. b Ccp6+/+ or Ccp6−/− MEFs were transfected by OSKM for 2 days and treated with MG132 (10 μM) for 4 h. Cells were collected and immunoprecipitated with anti-Klf4 antibody followed by detection with anti-poly-Ub antibody. c Klf4 ubiquitination is K48-linked but not K63-linked. pMX5-Klf4 together with pCDNA4-MycHis-Ubiquitin (Ub) or Ub(K48R), Ub(K63R) mutant were transfected into 293T cells for 48 h and analyzed as in b. d rGST-Klf4 was pre-incubated with lysates from 293 T cells transfected with Flag-TTLL4 at 37 °C for 2 h, followed by GST pulldown with Flag-βTrCP1 and immunoblotting with indicated antibodies. e rGST-Klf4 was pre-incubated with lysates from 293T cells transfected with Flag-TTLL4 at 37 °C for 2 h. Klf4 protein was then incubated with recombinant active ERK1 (Millipore) and 2 mM ATP at 30 °C for 30 min. His-tagged E1 (Uba1), E2 (UbcH5c), ubiquitin, Rbx, Flag-tagged βTrCP1, Skp1, Cullin1 and GST-tagged Klf4 were incubated for the in vitro ubiquitination reconstitution assay at 30 °C for 2 h. Ubiquitinated Klf4 was probed by anti-GST antibody. f TTLL4 deficiency promotes the ubiquitination of Klf4 in OSKM-induced MEFs. Ttll4+/+ or Ttll4−/− MEFs was transfected by OSKM for 2 days and assessed for ubiquitination as described in b. g Ttll4-deficient MEFs were treated with cycloheximide (CHX) (20 μg/ml) for the indicated time, followed by immunoblotting as described in a. h Ttll4-deficient 4- to 8-cell stage embryos were isolated and immunostained with anti-Klf4 and GT335 antibodies. Scale bar, 20 μm.For Ttll4+/+ embryos, n = 67. For Ttll4−/− embryos, n = 55. All data are representative of four independent experiments
Klf4 glutamylation impedes its K48-linked ubiquitination
We next wanted to explore how Klf4 glutamylation affects its activity. We noticed that Ccp1- and Ccp6-deficient MEFs impeded Klf4 degradation with OSKM transduction (Fig. 4a), whereas Klf4 was rapidly degraded in WT MEFs. In contrast, enforced overexpression of CCP1 or CCP6 in MEFs promoted Klf4 degradation (Supplementary Figure 5a). Restoration of CCP1 or CCP6 in accordance with Ccp1–/– or Ccp6–/– MEFs in turn reduced Klf4 protein levels comparable to that of Ccp1+/+ or Ccp6+/+ cells (Supplementary Figure 5a). These results indicate that deficiency of CCP1 or CCP6 sustains Klf4 protein stability.
Klf4 binds to the E3 ligase component βTrCP1 or βTrCP2 that catalyzes its ubiquitination leading to proteasome-mediated degradation16. Consistently, CCP6 deficiency remarkably declined Klf4 ubiquitination signals during iPSC induction (Fig. 4b). As expected, Klf4 ubiquitination was K48-linked but not K63-linked through K48R- and K63R-ubiquitin transfection assays (Fig. 4c). Notably, TTLL4 overexpression reduced Klf4 ubiquitination that suppressed Klf4 degradation (Fig. 4c). Furthermore, TTLL4-mediated glutamylation of Klf4 competitively inhibited its binding with the E3 ligase component βTrCP1 (Fig. 4d). Accordingly, Klf4 was ubiquitinated by the βTrCP1/Skp1/Cullin1/Rbx1 E3 ligase complex via an in vitro ubiquitination reconstitution assay (Fig. 4e). TTLL4-mediated glutamylation of Klf4 indeed repressed its ubiquitination. MLN4924 is an inhibitor that inactivates the SCF (Skp1, Cullins, and F-box proteins) complex E3 ubiquitin ligase activity44. As expected, MLN4924 treatment augmented Klf4 stability in OSKM-transduced MEFs and consequently promoted iPSC formation efficiency (Supplementary Figure 5b,c). Consistently, Klf4 depletion substantially reduced the gene expression of pluripotency factors (Supplementary Figure 5d, e). In addition, Klf4 occupied the promoter regions of these pluripotency factors (Supplementary Figure 5f).
We generated TTLL4 knockout (KO) mice via a CRISPR–Cas9 approach45. An 8-bp deletion of exon 3 led to frameshift mutation and full deletion of TTLL4 in mice (Supplementary Figure 5g, h). We found that with OSKM induction, TTLL4 KO dramatically augmented Klf4 ubiquitination and facilitated Klf4 degradation (Fig. 4f,g). In parallel, TTLL4 deficiency caused hypoglutamylation of Klf4 that resulted in its degradation in 4–8-cell-stage embryos and impaired early embryonic development (Fig. 4h). Finally, TTLL4 KO MEFs with MLN4924 treatment could restore Klf4 protein levels during OSKM transduction (Supplementary Figure 5i). The specificities of antibodies against CCP1, CCP6, TTLL1, or TTLL4 that we used were verified (Supplementary Figure 5j). Overall, TTLL4-mediated glutamylation of Klf4 impedes its K48-linked ubiquitination that sustains Klf4 stability.
A Klf4 glutamylation mutant reduces reprogramming frequency
To further confirm whether Klf4 polyglutamylation regulates reprogramming and pluripotency maintenance, we overexpressed Klf4-wt, Klf4-E381A, or Klf4-K232R (a stable form mutant46) into Klf4-silenced MEFs with OSKM transduction. We found that Klf4-E381A mutant abrogated polyglutamylation signals, iPSC formation, and pluripotent gene expression (Fig. 5a and Supplementary Figure 5k, l). By contrast, Klf4-K232R restoration achieved opposite results vs. the Klf4-E381A restoration. Consistently, Klf4-E381A restoration in mouse ESC D3 cells exhibited decreased colony numbers with AP-positive staining and downregulated pluripotent gene expression, while Klf4-K232R restoration in D3 cells displayed increased numbers of AP-positive colonies and upregulated pluripotent gene expression compared with Klf4-wt restoration (Fig. 5b and Supplementary Figure 5m). Furthermore, Klf4-E381A restoration enhanced differentiation gene expression, whereas Klf4-K232R restoration suppressed differentiation gene expression (Supplementary Figure 5n). These data indicate that Klf4 polyglutamylation is required for cell reprogramming and pluripotency maintenance.
A glutamylation-defective mutant of Klf4 reduces reprogramming efficiency. a Klf4-wt, Klf4-E381A, or Klf4-K232R were overexpressed in Klf4-silenced 4F2A MEFs and treated with dox and cultured in ESC media for 3 weeks. Scale bar, 50 μm. Reprogramming efficiency was assayed by Nanog staining after dox removal. Nanog-positive colony numbers per 104 cells were calculated and shown as means ± S.D. **P < 0.01,. n = 4. b D3 ESC cells were transfected with indicated plasmids, followed by AP staining. Scale bar, 50 μm. AP-positive colony numbers per well were calculated as means ± S.D. **P < 0.01. n = 5. c SSEA-1+ ES cells were isolated and an equal amount of cells were lyzed for nuclei extraction, followed by DNase I digestion. Total DNA was extracted and quantitated by qPCR with Nanog or Esrrb promoter-specific primers. n = 5. d Indicated plasmids along with TTLL4 were overexpressed into Klf4-silenced MEFs after OSKM transfection for 2 days, and cells were collected and immunoprecipitated with anti-Klf4 antibody followed by detection with anti-poly-Ub antibody. e Indicated proteins were incubated with labeled and unlabeled probes against Nanog promoter, followed by EMSA assay. Student’s t test was used as statistical analysis. ns, no significance
Nanog and Esrrb are downstream target genes of Klf4 during the iPSC reprogramming47,48. DNase I digestion assay showed that CCP6 deficiency augmented chromatin accessibility to DNase I digestion at the promoters of Nanog and Esrrb (Fig. 5c), indicating an open status of chromatin region of these genes. Rescue of Klf4-E381A into Klf4- silenced MEFs reduced DNase I accessibility of these two gene promoters (Fig. 5c). By contrast, restoration of Klf4-K232R into Klf4-silenced MEFs increased DNase I accessibility of these two promoters. These results suggest that Klf4 stabilization by polyglutamylation initiates transcriptional activation of its downstream target genes.
Actually, Klf4-wt glutamylation was catalyzed by TTLL4, which remarkably reduced its ubiquitination modification (Fig. 5d). By contrast, Klf4-E381A overexpression abrogated the polyglutamylation of Klf4 and augmented its ubiquitination signals (Fig. 5d). Expectedly, Klf4-K232R overexpression declined its ubiquitination modification (Fig. 5d). Similar results were achieved by treatment with the counterparts of TTLL1 overexpression (Supplementary Figure 5o). We next wanted to exclude the possibility that mutants of Klf4 could affect its target gene expression by changing their DNA-binding capacities. Binding capacity of Nanog promoter with Klf4-wt, Klf4-E381A, and Klf4-K232R was measured by an electrical mobility shift assay (EMSA). We observed that Klf4-E381A and Klf4-K232R mutants showed comparable binding affinities to Klf4-wt protein (Fig. 5e). Unlabeled Nanog probes could competitively bind to the Nanog promoter. These data suggest that Klf4 mutation per se has no direct activation effect on Nanog gene. Therefore, polyglutamylation-mediated Klf4 stability directly regulates the transcriptional activation of its downstream target genes.
To further verify the role of Klf4 polyglutamylation in the reprogramming and pluripotency regulation in vivo, we generated Klf4-E381A mutant knockin (Klf4E381A KI) mice by CRISPR/Cas9 editing technology. As expected, MEFs isolated from Klf4E381A KI mice really abrogated iPSC formation by in vitro OSKM transduction (Fig. 6a). Of note, Klf4E381A KI mice impaired blastocyst development at E3.5 (Fig. 6b). In addition, Klf4 with cell fate markers (Nanog for epiblast, Gata6 for primitive endoderm, and Cdx2 for trophectoderm) in early embryos from Klf4E381A KI mice was substantially reduced and consequently underwent apoptosis (Fig. 6c,d). Finally, Klf4E381A homozygous pups were embryonically lethal (Fig. 6e). Functions of Klf2 and Klf5 are redundant with Klf4 in ES cells35 and iPSCs49. Of interest, we found that Klf4-E381A was dimerized with Klf5 in ESCs, while Klf4-wt protein did not (Supplementary Figure 5p). We propose that the dimerization of Klf4-E381A with Klf5 could inactivate the Klf5 function to cause preimplantation lethality in Klf4E381A KI mice. It has been reported that a glutamylation site can shift to the next available glutamate residue when the main modification site is mutated in tubulin50. However, Klf4 E381A mutant knockin abrogated the Klf4 function in vivo, excluding the possibility of modification site shift for Klf4 glutamylation. Taken together, Klf4 polyglutamylation at Glu381 is required for cell reprogramming and early embryonic development.
A glutamylation-defective mutant of Klf4 impairs ESC pluripotency. a Klf4E381A knockin mice were generated by a CRISPR/Cas9 approach. MEFs were isolated and induced for iPS formation as in Fig. 5a. Scale bar, 100 μm. AP-positive colony numbers per 104 cells were calculated and shown as means ± S.D. **P < 0.01, n = 4. b Fertilized eggs at E3.5 embryonic stage were isolated from pregnant mice after heterozygotes crossing. Embryo genotype was confirmed by both GT335 immunostaining and PCR of genomic DNA. Scale bar, 100 μm. Distribution of embryonic stage at E3.5 was counted as means ± S.D, **P < 0.01. For WT embryos, n = 101. For Klf4E381A embryos, n = 87. c Indicated preimplantation embryos from WT and Klf4E381A knockin mice were stained with cell fate markers (Nanog for epiblast (EPI), Gata6 for primitive endoderm (PrE), and Cdx2 for trophectoderm (TE)) together with Klf4. n = 34 for WT embryos and n = 28 for Klf4E381A embryos (anti-Nanog staining), n = 41 for WT embryos and n = 25 for Klf4E381A embryos (anti-Gata6 staining), and n = 31 for WT embryos and n = 29 for Klf4E381A embryos (anti-Cdx2 staining). d Indicated preimplantation embryos were stained with cell fate markers as well as cleaved caspase 3 (cl-Casp3). The percentage of positive cell in each germ layer was counted as means ± S.D, **P < 0.01. Scale bar, 20 μm. n = 40 for WT embryos and n = 31 for Klf4E381A embryos (anti-Nanog staining), n = 37 for WT embryos and n = 30 for Klf4E381A embryos (anti-Gata6 staining), and n = 37 for WT embryos and n = 25 for Klf4E381A embryos (anti-Cdx2 staining). e Klf4E381A pups were genotyped after heterozygotes crossing. Student’s t test was used as statistical analysis
TTLL1 or TTLL4 deletion impairs embryonic development
To further explore the physiological role of TTLL1 and TTLL4 in the process of reprogramming, TTLL1 and TTLL4 were silenced in MEFs for iPSC formation assays. Depletion of TTLL1 or TTLL4 dramatically reduced iPSC colony numbers as well as expression levels of pluripotent genes (Fig. 7a and Supplementary Figure 6a, b). Double knockdown of TTLL1 and TTLL4 displayed a synergistic inhibitory effect. Overexpression of TTLL1 or TTLL4 in accordance with silenced MEFs restored iPSC colonies comparable to shCtrl-treated MEFs (Fig. 7a). TTLL1 or TTLL4 depletion in OSKM-transduced MEFs substantially declined Klf4 glutamylation signals (Supplementary Figure 6c). By contrast, overexpression of TTLL1 or TTLL4 in accordance with silenced MEFs rescued Klf4 glutamylation signals (Supplementary Figure 6c), which in turn promoted iPSC colony formation efficiency (Supplementary Figure 6d, e). Consequently, TTLL1 or TTLL4 depletion promoted Klf4 degradation (Fig. 7b). In contrast, overexpression of TTLL1 and TTLL4 sustained Klf4 stability. Finally, Ttll4-deficient MEFs impaired iPSC formation (Supplementary Figure 6f). These data indicate that TTLL1- or TTLL4-mediated Klf4 glutamylation is required for somatic cell reprogramming.
Deletion of TTLL1 or TTLL4 impairs iPSC induction and ESC pluripotency. a Depletion of TTLL1 or TTLL4 impairs iPSC formation. TTLL1 or TTLL4 was silenced in 4F2A MEFs as well as rescued with TTLL1 or TTLL4 into their silenced MEFs, followed by dox treatment for iPSC formation and stained with anti-Nanog antibody as in Fig. 1e. Scale bar, 50 μm. Nanog-positive colony numbers per 104 cells were calculated and shown as means ± S.D. **P < 0.01. n = 5. b Immunoblotting of Klf4 in above-treated MEFs as in a. Fold changes of relative expression of indicated proteins compared with β-actin were caculated as means ± S.D. The data represent four independent experiments. c TTLL1 or TTLL4 depletion enhances the pluripotency of mouse ESCs. D3 cells were transfected with scrambled shRNA (shCtrl), shTTLL1, shTTLL4, or TTLL1 and TTLL4 overexpression (oe) plasmids and cultured in mouse ESC media. After 5 days, pluripotency was analyzed by AP staining. Colony numbers for undifferentiated, mixed, or differentiated clones were calculated as means ± S.D. n = 5. Scale bar, 100 μm. d D3 cells were transfected with indicated plasmids. mRNA levels of the indicated genes were analyzed by real-time qPCR. Relative gene expression fold changes were counted as means ± S.D. **P < 0.01. n = 5. Primer pairs are shown in Supplementary Table 1. e Indicated plasmids were transfected into mouse D3 cells as in c. Colony numbers for undifferentiated, mixed, or differentiated clones were calculated as means ± S.D. n = 5. Scale bar, 100 μm. f mRNAs levels of the indicated genes were analyzed by real-time qPCR as in d. Relative gene expression fold changes were counted as means ± S.D. **P < 0.01. n = 5. g TTLL1 or TTLL4 depletion in human ESC H9 cells was confirmed by real-time qPCR. n = 5. h TTLL1 or TTLL4 depletion in human ESC H9 cells increases AP+ colony formation. Human H9 cells were infected with lentivirus expressing the indicated shRNAs and cultured in human ES media for 3 weeks. ESC pluripotency was analyzed by AP staining. AP+ colony numbers were calculated as means ± S.D. n = 4. Scale bar, 100 μm. Student’s t test was used as statistical analysis. oe overexpression
We next analyzed the role of TTLL1 and TTLL4 in ESC pluripotency maintenance. We silenced TTLL1 or TTLL4 expression in mouse ESC D3 cells. As expected, TTLL1- or TTLL4-depleted ESCs exhibited decreased numbers of colonies with AP-positive staining and downregulated pluripotent gene expression compared with shCtrl-treated cells (Fig. 7c,d). Double knockdown of TTLL1 with TTLL4 synergetically lowered AP-positive colony numbers and pluripotent gene expression in ESCs. Additionally, restoration of TTLL1 or TTLL4 into its corresponding TTLL1- and TTLL4-silenced cells rescued ESC pluripotency and pluripotent gene expression (Fig. 7c,d). By contrast, enforced overexpression of TTLL1 or TTLL4 increased ESC self-renewal as well as upregulated pluripotent gene expression (Fig. 7e,f). Similar results were obtained in TTLL1- or TTLL4-depleted human ESC H9 cells (Fig. 7g,h).
To further test the role of TTLL1 and TTLL4 in early embryogenesis, we also generated TTLL1 knockout (KO) mice via a CRISPR–Cas9 approach45 (Supplementary Figure 7a, b). We isolated fertilized eggs from WT, Ttll1–/–, and Ttll4–/– pregnant mice to assess ex vivo development of embryos. Both TTLL1 and TTLL4 were deleted in embryos by immunofluorescence staining (Fig. 8a). TTLL1 and TTLL4 deficiency suppressed blastocyst development (Fig. 8b), leading to decreased blastocoel areas, disordered lineage marker expression, and increased apoptotic cells of early embryos (Supplementary Figure 7c-f). Finally, both Ttll4-deficient pups and Ttll1-deficient pups were born in extremely lower numbers than those that would be predicted by Mendelian inheritance after heterozygotes crossing (Fig. 8c and Supplementary Figure 7g). Altogether, TTLL1 and TTLL4 play a critical role in the regulation of iPSC formation and early embryonic development.
Deletion of TTLL1 and TTLL4 impairs embryonic development. a TTLL1 and TTLL4 deficiency in blastomere was confirmed by immunofluorescence staining. Scale bar, 20 μm. b Fertilized eggs were isolated from WT, Ttll1−/−, and Ttll4−/− pregnant mice and assessed as in Fig. 1f. Distribution of embryonic stage at E3.5 was counted as means ± S.D (right panel), **P < 0.01. Scale bar, 100 μm. For WT embryos, n = 115. For Ttll1−/− embryos, n = 101. For Ttll4−/− embryos, n = 108. c Ttll4-deficient pups were genotyped after heterozygotes crossing. Student’s t test was used as statistical analysis. Data are representative of four independent experiments. d A working model represents Klf4 polyglutamylation in the regulation of cell reprogramming and early embryogenesis. During the process of reprogramming and pluripotency maintenance, the core pluripotency factor Klf4 is polyglutamylated by TTLL4 or TTLL1 that impedes its K48-linked ubiquitination to maintain Klf4 stability. Deletion of TTLL4 or TTLL1 impairs cell reprogramming and early embryogenesis. Klf4 glutamylation is hydrolyzed by CCP1 or CCP6. CCP1 or CCP6 deficiency substantially promotes iPSC generation and sustains ESC pluripotency. Glutamylation of Klf4 is required for activation of its downstream pluripotency genes leading to somatic reprogramming, which maintains ESC pluripotency as well as drives embryogenesis
Discussion
Transcription factors-mediated reprogramming was sufficient to reset terminally differentiated cells into induced pluripotent stem cells (iPSCs)2,32,51. The generation of iPSCs avoids ethical issues by using embryonic stem cells. Additionally, iPSCs can be established from patient’s tissues, which makes them more reliable and beneficial for treatment of human-related diseases52. In this study, we show that CCP1 or CCP6 deficiency substantially promotes iPSC generation and sustains ESC pluripotency. During the process of reprogramming, the core pluripotency factor Klf4 is polyglutamylated by TTLL4 or TTLL1 that impedes its K48-linked ubiquitination to maintain Klf4 stability (Fig. 8d). TTLL1- or TTLL4-mediated polyglutamylation of Klf4 is required for activation of its downstream pluripotency genes leading to somatic reprogramming, which maintains ESC pluripotency as well as drives embryogenesis. Deletion of TTLL4 or TTLL1 impairs cell reprogramming and early embryogenesis. Therefore, Klf4 polyglutamylation plays a critical role in the regulation of cell reprogramming and pluripotency maintenance.
Protein levels of Oct4 and Sox2 are precisely modulated in order to sustain self-renewal and pluripotency of ESCs53,54. A moderate increase of Sox2 causes differentiation of ESCs primarily into neural ectodermal cells, mesodermal, and trophectoderm-like cells53, while decreased expression of Sox2 induces differentiation of ESCs into trophectoderm-like cells55. Similarly, Klf4 activity is also tightly regulated by PTMs. For example, phosphorylation of Klf4 by ERKs at Ser123 leads to inhibition of its transcriptional activity16. In addition, Klf4 also undergoes sumoylation that enhances Klf4 transactivation activity56. However, it is unknown whether core transcription factors undergo glutamylation in cell reprogramming and ESC pluripotency. The well-known substrates of polyglutamylation are tubulins and nucleosome assembly proteins43,57,58. A recent study delineates a structural microtubule recognition basis by catalysis with TTLL719. TTLLs have different expression patterns in diverse tissues and their functions are not entirely redundant59. We recently reported that TTLL4 and TTLL6 are most highly expressed in megakaryocytes29, both of which catalyze polyglutamylation of Mad2 to modulate megakaryocyte maturation. In this study, we demonstrate that TTLL1 and TTLL4 are constitutively elevated in ESCs and early embryos, and dramatically upregulated in iPSCs during cell reprogramming. Both TTLL1 and TTLL4 can catalyze polyglutamylation of Klf4, which impedes its K48-linked ubiquitination to maintain Klf4 stability. We showed that both Ttll4-deficient pups and Ttll1-deficient pups were born in extremely lower numbers than those that would be predicted by Mendelian inheritance after heterozygotes crossing. These observations suggest that TTLL1 and TTLL4 might synergistically modify the same glutamate on Klf4 to regulate early embryonic development. However, the different biochemical activities of TTLL family enzymes in the regulation of cell reprogramming still need to be further investigated.
Protein glutamylation is a reversible modification, whose deglutamylation is hydrolyzed by a family of cytosolic carboxypeptidases (CCPs)23,60. CCP family members harbor enzymatic specificities to carry out deglutamylation. Of these CCP members, CCP1, CCP4, and CCP6 remove the shortening of penultimate polyglutamate chains of α-tubulin, while CCP5 specifically hydrolyzes the branching site glutamate23. We previously showed that Mad2 polyglutamylation can be hydrolyzed by CCP6 but not CCP123. We provide the example that CCP1 and CCP6 harbor nonredundant roles in physiological conditions. Consequently, deficiency of CCP1 or CCP6 exhibits different abnormalities27,29. Herein, we show that both CCP1 and CCP6 are highly expressed in differentiated cells and remarkably declined in ESCs and iPSCs. Both CCP1 and CCP6 can hydrolyze the glutamylation of Klf4 during cell reprogramming and ESC maintenance. CCP1 and CCP6 could synergistically shorten longer glutamate chains of glutamyated Klf4, while CCP5 may be needed to deglutamylate Klf4 at a branching glutamate site. Alternatively, CCP1 and CCP6 could also remove a branched glutamate of glutamyated Klf4, whose biochemical activities may be different from tubulins23.
Glutamylation is highly conserved in all metazoans and protists, performing the assembly and function of cilia and flagella58. For example, TTLL7, the most abundantly expressed in the mammalian nervous system, is conserved from acorn worm to primates, where it modulates neurite outgrowth and localization of dendritic MAPs. In addition, CCP and TTLL members exert their functions by enzymatic activities against their respective substrates, and their enzymatic activities are also modulated by post-translational modifications and other regulatory factors. Here, we show that Klf4 polyglutamylation mediated by TTLL1 and TTLL4 enhances mouse and human reprogramming and pluripotency. In parallel, the CCP inhibitor phenanthroline dramatically promotes somatic cell reprogramming and embryonic development. Thereby, we strongly believe that it is necessary to develop specific inhibitors or agonists for each individual polyglutamylase and cytosolic carboxypeptidase. Manipulating polyglutamylation profiles by using these compounds, we may potentially improve iPSC efficiency for future clinical applications. In sum, Klf4 glutamylation plays a critical role in the regulation of cell reprogramming and pluripotency maintenance. Our findings provide mechanistic insights into how glutamylation modulates cell fate determination.
Methods
Antibodies and reagents
Antibodies against Myc-tag (sc-40), TTLL1 (sc-86929), CCP6 (sc-138893), SSEA-1(sc-21702), and GST-tag (sc-138) were purchased from Santa Cruz Biotechnology (Santa Cruz, USA). Antibodies against Flag-tag (M2, F3165), β-actin (A-5316), His-tag (H1029), glutamylated tubulin (B3), and GFP-tag (G-1544) were from Sigma-Aldrich (St. Louis, USA). GT335 (anti-glutamylation) antibody was from AdipoGen (San Diego, USA). The antibody against TTLL4 (PAB22002) was from Abnova (Walnut, USA). Antibodies against ubiquitin (K48 linked) (ab140601) and Gata6 (ab22600) were purchased from Abcam (Cambridge, USA). Anti-Klf4 (11880-1-AP) antibody, anti-CCP1(14067-1-AP) antibody, and antibody against Nanog (14295-1-AP) were purchased from ProteinTech (Chicago, USA). The antibody against tubulin (AT819) was from Beyotime (Shanghai, China). The antibody against CDX2 (CDX2-88) was from BioGenex (Fremont, USA). The antibody against cleaved caspase 3 (#9661) was from Cell Signaling Technology (Beverly, USA). Antibodies against Myc-tag, GST-tag, Flag-tag, β-actin, GFP-tag, and His-tag were used in 1:5000 dilutions for western blotting. Antibodies against TTLL1, CCP6, TTLL4, Klf4, ubiquitin (K48 linked), glutamylated tubulin (B3), CCP1, and GT335 were used in 1:500 dilutions for western blotting. Antibodies against SSEA1 and cleaved Caspase3 were used in 1:500 dilutions for flow cytometric staining. Antibodies against ubiquitin (K48 linked) (ab140601) and Gata6, tubulin, CDX2, β-actin, Nanog, and cleaved caspase 3 were used in 1:500 dilutions for immunofluorescence staining. Alexa-405, Alexa-488, Alexa-594, and Alexa-649-labeled secondary antibodies were from Thermo Fisher Scientific (Waltham, USA). Chicago Sky Blue 6B, DAPI (4,6-diamino-2-phenyl indole), cycloheximide (CHX), MG132, phenanthroline, and NEDD8-activating enzyme inhibitor (MLN4924) were purchased from Sigma-Aldrich. DNaseI was purchased from Roche Molecular Biochemicals (Basel, Switzerland). CoCl2 was from Sinopharm Chemical Reagent Co.,Ltd (Beijing, China).
Animals
Ccp1–/– mice (Pcd mice, BALB/cByJ-Agtpbp1pcd-3J-J) were obtained from Jackson Lab. Ccp6–/– mice (FVB background) were generated by a transposon piggyBac insertional mutagenesis program (PBMICE). The strategy of breeding was to cross between heterozygotes to generate homozygote knockout mice and littermate control mice as described29. Mouse experiments were approved by the Institutional Animal Care and Use Committees at the Institute of Biophysics, Chinese Academy of Sciences. All the mice that we used were from C57BL/6 background, female, and 8–12-weeks old. We were not blinded to the group and did not use randomization in our animal studies. Littermates with the same age and gender for each group were used. We excluded the mice that were 5 g thinner than other littermates before any treatment or analysis. All mice were bred in SPF animal facility. The euthanasia method was cervical dislocation when needed.
Cell culture
Human 293 T cells were cultured with DMEM supplemented with 10% FBS and 100 U/ml penicillin and 100 mg/ml streptomycin. Retrovirus- or lentivirus-infecting MEFs were produced in 293 T cells using the standard protocols. Transfection was performed using lipofectin (Invitrogen). A total of 293 T cells were obtained from ATCC and authenticated by PCR. Mycoplasma contamination was tested by PCR. Mouse ESC D3 cells were from the stem cell core facility of Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China). D3 or R1 ESCs were cultured in DMEM supplemented with 15% FBS (Invitrogen, USA), 2 mM L-glutamine, 0.1 mM 2-mercaptoethanol, 1 mM nucleosides, 0.1 mM nonessential amino acids, and 103 units/ml mouse leukemia inhibitory factor (LIF) and were grown on B6 mouse MEFs treated with mitomycin C. Human ESC H1 cells were obtained from Wicell Research Institute (Madison, WI). Human ESCs and iPSCs were grown on CF-1 mouse MEFs treated with mitomycin C as feeder cells. Culture medium contained DMEM/F12 medium, 20% knockout serum replacement (KSR) (Invitrogen), and 2 mM L-glutamine, 0.1 mM nonessential amino acids, 0.1 mM 2-mercaptoethanol, and 4 ng/ml hFGF2 (Sigma-Aldrich, USA). Concentrations of 10 μM CoCl2 or 1 μM phenanthroline were used during iPSC induction. A concentration of 1 μM MLN4924 was used for iPSC induction44. Embryonic stem cell use was approved by the Institutional Medical Research Ethics Committee at the Institute of Biophysics, Chinese Academy of Sciences.
Plasmid construction and protein expression
Mouse CCP6, ubiquitin (Ub), Ub(K48R), and Ub(K63R) were cloned into pCDNA4-MycHis expression vector. CCP6-H230S/E233Q mutant (CCP6mut) and CCP6wt were also subcloned into H-MBP-3C vector and purified using amylose resin (New England BioLabs, Ipswich, USA) according to the manufacturer’s instruction. CCP1, TTLL4, Cullin1, and Skp1 were cloned into p3×flag-CMV-9 expression vector. pMX5-Oct4, pMX5-Sox2, pMX5-Klf4, and pMX5-cMyc plasmids were from Dr. Yang Xu’s lab (University of California, San Diego, La Jolla, USA). pDEST-Flag-PHF13-βTrCP1 plasmid was from Dr. Degui Chen’s lab (Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai, China). STEMCCA plasmid was from EMDMillipore. Mouse wild-type Klf4 (Klf4wt), Klf4E46A, Klf4E97A, Klf4K232R, Klf4E329A, and Klf4E384A mutants were generated and cloned into pGEX-6P-1 vector (GE Healthcare), expressed in E. coli, and purified using Glutathione Sepharose 4B beads. Uba1, UbcH5c, Rbx, and ubiquitin were subcloned into pET-28a vectors. CCP1, CCP6, TTLL4, TTLL1, Klf4wt, Klf4E384A mutant, and Klf4K232R mutant were also subcloned into pMY-IRES-GFP vector (Cell Biolabs) for enforced expression. RNA interference was designed according to BLOCK-IT RNAi Designer system instructions (Invitrogen). shRNA oligos encoding target mouse sequences against mKlf4 (5′-GACATCGCCGGTTTATATTGA-3′), mCCP1 (5′-GGAGAACACGAAAGATCTTCA-3′), mCCP6 (5′-CCGAGTCTGGTTCAACTTTAC-3′), mTTLL1(5′-GGATGAGCGTGCAAACCATTC-3′), mTTLL4 (5′-CCAGCCAGCCTATTTCCTTTG-3′), or their accordingly scrambled sequences were cloned into MSCV-LTRmiR30-PIG vector (LMP, Openbiosystems). shRNA oligos encoding target human sequences against hCCP1 (5′-GCATAGAAACATGCTCATTCG-3′), hCCP6 (5′-GCTGAGGACTTCTCCTATTCC-3′), or their accordingly scrambled sequences were cloned into pSUPER vector (Oligoengine, Seattle, USA).
iPSC generation
Mouse embryonic fibroblasts (MEFs) were obtained at E14.5 of Ccp1+/+, Ccp1–/–, Ccp6+/+, Ccp6–/–, Ttll4+/+, or Ttll4–/– embryos and supplied with Dulbecco’s modified Eagle’s medium (DMEM) (Invitrogen) with 15% FBS, 100 μg/ml streptomycin, and 100 U/ml penicillin. MEFs were infected with pMX5-Oct4, pMX5-Sox2, pMX5-Klf4, and pMX5-cMyc-packaged retrovirus mixture and then plated on mitomycin-C-treated feeder cells and cultured in ESC media containing DMEM supplemented with 15% FBS (Invitrogen, USA), 2 mM L-glutamine, 0.1 mM 2-mercaptoethanol, 1 mM nucleosides, 0.1 mM nonessential amino acids, and 103 units/ml mouse leukemia inhibitory factor (LIF). For higher 4-factor infection, MEFs were infected with STEMCCA lentivirus for reprogramming induction. Reprogramming experiments were also performed using MEFs obtained from E14.5 embryos of 4F2A mice (Jax stock No. 011004). The 4 Yamanaka factor expression was induced by doxycycline treatment. After 3 weeks, reprogramming efficiency was assayed by Nanog staining after dox removal.
Embryo collection and culture
A total of 5 IU of pregnant mare serum gonadotropin (PMSG) was intraperitoneally injected for superovulation and 5 IU of human chorionic gonadotropin (hCG) was injected 48 h later. Embryos were flushed with M2 medium (Millipore) from oviducts or uteri at the indicated stages and cultured ex vivo in KSOM medium (Millipore) supplied with 1 mg/ml BSA at 37 ℃ with 5% CO2. Concentrations of 10 μM CoCl2 or 1 μM phenanthroline were used for embryo culture. For each group, more than 100 typical embryos were observed.
Generation of knockout and Klf4 E381A knockin mice
The genome loci of Ttll1 and Ttll4 genes were edited by a CRISPR–Cas9 approach45. Briefly, vector pST1374 expressing Cas9 and pUC57kan-T7 expressing sgRNA for Ttll1 or Ttll4 gene was constructed as below45. Mixtures of Cas9 mRNA (100 ng/μl) and sgRNA (50 ng/μl) were microinjected into the cytoplasm of C57BL/6 fertilized eggs. The injected zygotes were cultured until blastocyst stage and transferred into the uterus of pseudopregnant ICR females. Frameshift mutations were identified by PCR screening and DNA sequencing. The genome loci of Ttll1 and Ttll4 genes were edited by a CRISPR–Cas9 approach45. For generation of Klf4E381A knockin mice, the genome locus of Klf4 gene was knocked in with Klf4E381A mutation via a CRISPR–Cas9 approach. A mixture of Cas9 mRNA, sgRNA, and Klf4-E381A templates was microinjected into the cytoplasm of C57BL/6 fertilized eggs and transferred into the uterus of pseudopregnant ICR females. Klf4E381A mutations were identified by PCR screening and DNA sequencing. sgRNA sequences are as follows: Ttll1 (5′-CGATCTGGTCGTCTGACAGCCGG-3′); Ttll4 (5′-TTTGCCTCACGTTGGTGCGGCGG-3′); and Klf4 E381A (5′-ACCGGGTTCCTGCCTGCCAGAGG-3′). Klf4 template sequence (5′-TAGGTTTTGCCACAGCCTGCATAGTCACAAGTGTGGGTGGCTGTTCTTTTCCGGGGCCACGACCTTCTTCCCCTCTTTGGCTTGGGCTCAGCTGGCAGGCA-3′).
In vivo assay of teratomas
Human iPSCs were generated by transducing OSKM factors into human urothelial cells. Urine samples were recruited from three healthy donors with informed consent based on approval by the Institutional Ethical Committees at the Institute of Biophysics, Chinese Academy of Sciences. Ccp1–/– or Ccp6–/– mouse iPS cells or human iPS cells infected with shCCP1, shCCP6, oeCCP1, and oeCCP6 lentiviruses were collected, washed twice with PBS, and then subcutaneously injected into the bilateral inguens of male NOD/SCID mice (2 × 106 cells per injection). After 4 weeks for mouse iPSCs and 6 weeks for human iPSCs, mice were killed. Tumors were weighed, fixed in 4% paraformaldehyde, and sectioned for staining with hematoxylin and eosin (H&E)61. The maximal sizes of tumors permitted were 2500 mm3 according to the ethical approval by the Institutional Animal Care and Use Committees at the Institute of Biophysics, Chinese Academy of Sciences.
Immunofluorescence assay
MEFs were fixed with 4% paraformaldehyde (PFA, Sigma-Aldrich) for 20 min after retrovirus infection. Embryos were collected and treated with acidic tyrode buffer for 1 min to remove zona pellucida followed by 4% PFA fixation. Cells were permeabilized with 1% Triton-X 100 in PBS for 20 min, and blocked with 10% donkey serum. Cells were then incubated with indicated primary antibodies at 4 ℃ overnight followed by staining with Alexa405-, Alexa488-, Alexa594-, and Alexa649-conjugated secondary antibodies. Nuclei were stained with DAPI or PI. Images were obtained with Olympus FV1000 laser-scanning confocal microscopy (Olympus, Japan). The software ImageJ was used for colocalization analysis61.
Mass spectrometry
MBP-CCP6-wt and MBP-CCP6-mut23,29 proteins were expressed in E. coli and purified using the amylose resin (New England BioLabs, Ipswich, USA) according to the manufacturer’s instruction. Proteins were immobilized with Affi-gel10 resin to go through OSKM-transduced MEF lysates for affinity chromatography. Eluted fractions were visualized by SDS-PAGE followed by silver staining. Differential bands in SDS-PAGE gels were trypsinized for mass spectrometry with LTQ Orbitrap XL (Thermo Finnigan)29.
Immunoprecipitation assay
MEFs were infected with the indicated retrovirus. Cells were lyzed with ice-cold RIPA buffer (50 mM Tris-HCl [pH 7.4], 150 mM NaCl, 0.5% sodium desoxycholate, 0.1% SDS, 5 mM EDTA, 2 mM PMSF, 20 mg/ml aprotinin, 20 mg/ml leupeptin, 10 mg/ml pepstatin A, 150 mM benzamidine, and 1% Nonidet P-40) for 1 h. Lysates were incubated with the indicated antibodies followed by immunoprecipitation with protein A/G agarose beads and immunoblotting62. For in vivo ubiquitination assay, cells were pretreated with MG132 (20 μM) for 4 h and scraped into 100-μl lysis buffer (10 mM Tris-HCl, pH 7.4, 0.5 M NaCl, and 2% SDS). Lysates were heated at 95 °C for 10 min and diluted with 10 mM Tris-HCl to reduce SDS concentration to 0.2%. Anti-Klf4 antibody was used to precipitate immunocomplexes, followed by immunoblotting with anti-Ubiquitin antibody. Uncropped scans of results were shown as Supplementary Figure 8.
In vitro glutamylation assay
A total of 293 T cells were transfected with expression plasmids of TTLL4, CCP1, and CCP6 for 48 h. Cells were collected and lyzed with PBS–0.2% NP40. Supernatants were incubated with recombinant GST-Klf4 protein at 37 ℃ for 2 h. GST-Klf4 was pulled down with Glutathione Sepharose 4B beads, followed by immunoblotting29. Uncropped scans of results were shown as Supplementary Figure 8.
In vitro ubiquitination reconstitution assay
Uba1, UbcH5c, Rbx, and ubiquitin were cloned into pET-28a vectors. Klf4 was cloned into pGEX-6p-1 vector. Plasmids were then transformed into E. coli strain BL21 (DE3). Protein expression was induced with 1 mM isopropyl-β-D-1-thiogalactopyranoside (IPTG) at 16 °C for 24 h. Proteins were purified by Ni-NTA resin column (Novagen) or GST-sepharose column. Cullin1 and Skp1 were cloned into p3×flag-CMV-9 expression vector. βTrCP1 was cloned into pDEST-Flag-PHF13 expression vector. Plasmids were transfected into 293 T cells. Cells were lyzed with RIPA buffer and Flag-tagged proteins were immunoprecipitated with anti-Flag antibody and eluted by Flag-peptide (Sigma-Aldrich). Klf4 was pretreated with TTLL4 prior to in vitro polyglutamylation assay. Before in vitro ubiquitination assay, Klf4 protein was also incubated with recombinant active ERK1 (Millipore) and 2 mM ATP at 30 °C for 30 min. In vitro ubiquitination reconstitution assay was performed by mixing E1 (Uba1), E2 (UbcH5c), E3 (βTrCP1, Cullin1, Skp1, and Rbx), ubiquitin, and Klf4 in the ubiquitination buffer (50 mM Tris-HCl, 5 mM MgCl2, 2 mM dithiothreitol, and 2 mM ATP, pH 7.4) at 30 °C for 2 h63. The uncropped scan of the result was shown as Supplementary Figure 8.
Gene expression assay by real-time qPCR
iPSCs and ESCs were sorted by flow cytometry after SSEA-1 staining. A total of 5 × 105 cells were lyzed in each sample for the analysis below. Total RNAs were extracted with the RNA miniprep kit (LCsciences, Houston, USA) according to the manufacturer’s manual. M-MLV reverse transcriptase (Promega, Madison, USA) was used to synthesize cDNA. Quantitative PCR analysis and data collection were performed on the ABI 7300 qPCR system using the primer pairs listed in Supplementary Table 1. Quantitation was normalized to an endogenous β-actin gene64.
Microarray analysis
Mouse ES cell line D3 cells were transfected with the shCCP6 or shCtrl plasmid and cultured in mouse ESC media for 3 days. RNAs from mouse shCCP6 ESCs and shCtrl ESCs were prepared for Affimatrix microarray analysis according to the manufacturer’s instruction. Briefly, total RNA was extracted by Trizol reagent. Double-strand cDNA was synthesized by an Invitrogen SuperScript cDNA synthesis kit and labeled according to the Affimatrix protocol. Microarrays were hybridized, washed and scanned according to the manufacturer’s instruction. Differentially expressed genes were identified as fold change cutoff >2.0, FDR<0.05.
DNase I accessibility assay
SSEA-1+ iPSCs were sorted by flow cytometry. Nuclei were isolated from 1 × 105 iPSC cells using the nuclei isolating kit (Sigma-Aldrich) according to the manufacturer’s protocol. Nuclei were resuspended in 200 μl of DNase I digestion buffer (1 mM EDTA, 0.1 mM EGTA, 5% sucrose, 1 mM MgCl2, and 0.5 mM CaCl2). Equal aliquots of 100 μl of nuclei were treated with the indicated units of DNase I (Sigma, USA) and incubated at 37 ℃ for 5 min. The reactions were stopped by 2 × DNase I stop buffer (20 mM Tris, pH 8.0, 4 mM EDTA, and 2 mM EGTA). DNA was extracted and analyzed by qPCR. The primer pairs aimed at the promoter region for indicated genes are listed below. mNanog promoter: Forward, 5′-ATCCACCTGCCTCTGCCGCCTAA-3′, Reverse, 5′- GCATTGGTGTTTTGCCTGCATGG-3′; mEsrrb promoter: Forward, 5′-CCAACCAGAAGTGGGTCTTGTTCCT-3′, Reverse, 5′-TGTGGAAGGATCCTGGGACACAGAT-3′64.
Electrophoretic mobility shift assay (EMSA)
Biotin-labeled double-strand probes were generated by annealing complementary single-stranded oligonucleotides. The probe sequence used for detection was from Nanog promoter region: 5′- GCCGCCTGGGTGCCTGGGAGAATAGGGGGTGGGTAGGGTAGGAGG
CTTG-3′. Recombinant GST-Klf4 and mutants were expressed by E.coli and purified as described. LightShift Chemiluminescent EMSA Kit (Thermo Scientific) was used for shift assay according to the manufacturers’ instructions. Unlabeled (1-fold or 100-fold of labeled probe) probe was used for competitive reaction61. The uncropped scan of the result was shown as Supplementary Figure 8.
Statistical analysis
Unless otherwise stated, data are presented as means ± S.D. Student’s t test was used as statistical analysis by using Microsoft Excel as described.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its supplementary information files or from the corresponding author upon reasonable request. Microarray data that support the findings of this study are available in GEO datasets with the accession code GSE106809.
References
Yamanaka, S. & Blau, H. M. Nuclear reprogramming to a pluripotent state by three approaches. Nature 465, 704–712 (2010).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Takebe, T. et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 499, 481–484 (2013).
Hallett, P. J. et al. Successful function of autologous iPSC-derived dopamine neurons following transplantation in a non-human primate model of Parkinson’s disease. Cell Stem Cell 16, 269–274 (2015).
Apostolou, E. & Hochedlinger, K. Chromatin dynamics during cellular reprogramming. Nature 502, 462–471 (2013).
Hansson, J. & Krijgsveld, J. Proteomic analysis of cell fate decision. Curr. Opin. Genet. Dev. 23, 540–547 (2013).
Christophorou, M. A. et al. Citrullination regulates pluripotency and histone H1 binding to chromatin. Nature 507, 104–108 (2014).
Jang, H. et al. O-GlcNAc regulates pluripotency and reprogramming by directly acting on core components of the pluripotency network. Cell Stem Cell 11, 62–74 (2012).
Ramakrishna, S., Kim, K. S. & Baek, K. H. Posttranslational modifications of defined embryonic reprogramming transcription factors. Cell. Reprogram. 16, 108–120 (2014).
Luca, V. C. et al. Structural biology. Structural basis for Notch1 engagement of Delta-like 4. Science 347, 847–853 (2015).
Fang, L. et al. A methylation-phosphorylation switch determines Sox2 stability and function in ESC maintenance or differentiation. Mol. Cell. 55, 537–551 (2014).
Jeong, C. H. et al. Phosphorylation of Sox2 cooperates in reprogramming to pluripotent stem cells. Stem Cells 28, 2141–2150 (2010).
Baltus, G. A. et al. Acetylation of sox2 induces its nuclear export in embryonic stem cells. Stem Cells 27, 2175–2184 (2009).
Myers, S. A. et al. SOX2 O-GlcNAcylation alters its protein-protein interactions and genomic occupancy to modulate gene expression in pluripotent cells. Elife 7, e10647 (2016).
Kim, M. O. et al. ERK1 and ERK2 regulate embryonic stem cell self-renewal through phosphorylation of Klf4. Nat. Struct. Mol. Biol. 19, 283–290 (2012).
Xu, H. et al. WWP2 promotes degradation of transcription factor OCT4 in human embryonic stem cells. Cell. Res. 19, 561–573 (2009).
Di Stefano, B. et al. C/EBPalpha creates elite cells for iPSC reprogramming by upregulating Klf4 and increasing the levels of Lsd1 and Brd4. Nat. Cell. Biol. 18, 371–381 (2016).
Garnham, C. P. et al. Multivalent microtubule recognition by tubulin tyrosine ligase-like family glutamylases. Cell 161, 1112–1123 (2015).
Edde, B. et al. Posttranslational glutamylation of alpha-tubulin. Science 247, 83–85 (1990).
Valenstein, M. L. & Roll-Mecak, A. Graded control of microtubule severing by tubulin glutamylation. Cell 164, 911–921 (2016).
Kubo, T., Yanagisawa, H. A., Yagi, T., Hirono, M. & Kamiya, R. Tubulin polyglutamylation regulates axonemal motility by modulating activities of inner-arm dyneins. Curr. Biol. 20, 441–445 (2010).
Rogowski, K. et al. A family of protein-deglutamylating enzymes associated with neurodegeneration. Cell 143, 564–578 (2010).
Lyons, P. J., Sapio, M. R. & Fricker, L. D. Zebrafish cytosolic carboxypeptidases 1 and 5 are essential for embryonic development. J. Biol. Chem. 288, 30454–30462 (2013).
van Dijk, J. et al. A targeted multienzyme mechanism for selective microtubule polyglutamylation. Mol. Cell. 26, 437–448 (2007).
Janke, C. et al. Tubulin polyglutamylase enzymes are members of the TTL domain protein family. Science 308, 1758–1762 (2005).
Fernandez-Gonzalez, A. et al. Purkinje cell degeneration (pcd) phenotypes caused by mutations in the axotomy-induced gene, Nna1. Science 295, 1904–1906 (2002).
Wang, T., Parris, J., Li, L. & Morgan, J. I. The carboxypeptidase-like substrate-binding site in Nna1 is essential for the rescue of the Purkinje cell degeneration (pcd) phenotype. Mol. Cell. Neurosci. 33, 200–213 (2006).
Ye, B. et al. Cytosolic carboxypeptidase CCP6 is required for megakaryopoiesis by modulating Mad2 polyglutamylation. J. Exp. Med. 211, 2439–2454 (2014).
Sergouniotis, P. I. et al. Biallelic variants in TTLL5, encoding a tubulin glutamylase, cause retinal dystrophy. Am. J. Hum. Genet. 94, 760–769 (2014).
Kashiwaya, K. et al. Involvement of the tubulin tyrosine ligase-like family member 4 polyglutamylase in PELP1 polyglutamylation and chromatin remodeling in pancreatic cancer cells. Cancer Res. 70, 4024–4033 (2010).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Polo, J. M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012).
Liu, L. et al. Transcriptional pause release is a rate-limiting step for somatic cell reprogramming. Cell Stem Cell 15, 574–588 (2014).
Jiang, J. et al. A core Klf circuitry regulates self-renewal of embryonic stem cells. Nat. Cell. Biol. 10, 353–360 (2008).
Xia, P. et al. Glutamylation of the DNA sensor cGAS regulates its binding and synthase activity in antiviral immunity. Nat. Immunol. 17, 369–378 (2016).
Boskovic, A. et al. Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev. 28, 1042–1047 (2014).
Wang, L. et al. INO80 facilitates pluripotency gene activation in embryonic stem cell self-renewal, reprogramming, and blastocyst development. Cell Stem Cell 14, 575–591 (2014).
Wolff, A. et al. Distribution of glutamylated alpha and beta-tubulin in mouse tissues using a specific monoclonal antibody, GT335. Eur. J. Cell. Biol. 59, 425–432 (1992).
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502, 65–70 (2013).
Di Stefano, B. et al. C/EBPalpha poises B cells for rapid reprogramming into induced pluripotent stem cells. Nature 506, 235–239 (2014).
Rudiger, A. H., Rudiger, M., Wehland, J. & Weber, K. Monoclonal antibody ID5: epitope characterization and minimal requirements for the recognition of polyglutamylated alpha- and beta-tubulin. Eur. J. Cell. Biol. 78, 15–20 (1999).
van Dijk, J. et al. Polyglutamylation is a post-translational modification with a broad range of substrates. J. Biol. Chem. 283, 3915–3922 (2008).
Luo, Z., Pan, Y., Jeong, L. S., Liu, J. & Jia, L. Inactivation of the Cullin (CUL)-RING E3 ligase by the NEDD8-activating enzyme inhibitor MLN4924 triggers protective autophagy in cancer cells. Autophagy 8, 1677–1679 (2012).
Shen, B. et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nat. Methods 11, 399–402 (2014).
Lim, K. H., Kim, S. R., Ramakrishna, S. & Baek, K. H. Critical lysine residues of Klf4 required for protein stabilization and degradation. Biochem. Biophys. Res. Commun. 443, 1206–1210 (2014).
Chan, K. K. et al. KLF4 and PBX1 directly regulate NANOG expression in human embryonic stem cells. Stem Cells 27, 2114–2125 (2009).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Nakagawa, M. et al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat. Biotechnol. 26, 101–106 (2008).
Xia, L. et al. Polyglycylation of tubulin is essential and affects cell motility and division in Tetrahymena thermophila. J. Cell. Biol. 149, 1097–1106 (2000).
Park, I. H., Lerou, P. H., Zhao, R., Huo, H. & Daley, G. Q. Generation of human-induced pluripotent stem cells. Nat. Protoc. 3, 1180–1186 (2008).
Inoue, H., Nagata, N., Kurokawa, H. & Yamanaka, S. iPS cells: a game changer for future medicine. Embo. J. 33, 409–417 (2014).
Kopp, J. L., Ormsbee, B. D., Desler, M. & Rizzino, A. Small increases in the level of Sox2 trigger the differentiation of mouse embryonic stem cells. Stem Cells 26, 903–911 (2008).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell. Biol. 9, 625–635 (2007).
Du, J. X., McConnell, B. B. & Yang, V. W. A small ubiquitin-related modifier-interacting motif functions as the transcriptional activation domain of Kruppel-like factor 4. J. Biol. Chem. 285, 28298–28308 (2010).
Regnard, C. et al. Polyglutamylation of nucleosome assembly proteins. J. Biol. Chem. 275, 15969–15976 (2000).
Garnham, C. P. & Roll-Mecak, A. The chemical complexity of cellular microtubules: tubulin post-translational modification enzymes and their roles in tuning microtubule functions. Cytoskeleton 69, 442–463 (2012).
Bosch Grau, M. et al. Tubulin glycylases and glutamylases have distinct functions in stabilization and motility of ependymal cilia. J. Cell. Biol. 202, 441–451 (2013).
Rodriguez de la Vega Otazo, M., Lorenzo, J., Tort, O., Aviles, F. X. & Bautista, J. M. Functional segregation and emerging role of cilia-related cytosolic carboxypeptidases (CCPs). FASEB J. 27, 424–431 (2013).
Wang, S. et al. Transient activation of autophagy via Sox2-mediated suppression of mTOR is an important early step in reprogramming to pluripotency. Cell Stem Cell 13, 617–625 (2013).
Xia, P. et al. Sox2 functions as a sequence-specific DNA sensor in neutrophils to initiate innate immunity against microbial infection. Nat. Immunol. 16, 366–375 (2015).
Ye, B. et al. Pcid2 inactivates developmental genes in human and mouse embryonic stem cells to sustain their pluripotency by modulation of EID1 stability. Stem Cells 32, 623–635 (2014).
Wang, Y. et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 16, 413–425 (2015).
Acknowledgements
We thank Dr. Xiaohui Wu (University of Fudan) for providing Ccp6–/– mice. We thank Dr. Degui Chen (Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences) for kindly providing βTrCP1 plasmid. We thank Drs. Yang Xu and Tongbiao Zhao (University of California, San Diego) for providing OSKM plasmids. We thank Jing Li (Cnkingbio Company Ltd, Beijing, China) for technical support. We thank Peng Xue, Xiang Ding, Xiang Shi, Yan Teng, Chunli Jiang, Junying Jia, Liangming Yao, Jing Cheng, Yihui Xu, Xudong Zhao, and Xiaofei Guo for technical support. This work was supported by the National Natural Science Foundation of China (31530093, 91640203, 91419308, 31429001, 31470864, 31670886, 81601361, 81572433, 81772646, and 31601189), and the Strategic Priority Research Programs of the Chinese Academy of Sciences (XDB19030203).
Author information
Authors and Affiliations
Contributions
B.Y., B.L., and L.Ha designed and performed the experiments; X.Z. generated TTLL4–/– mice; B.Y. analyzed data and wrote the paper; S.W., P.X., Y.D., X.Q., Y.W., X.Y., C.L., J.H., P.Z., and L.He analyzed data; L.Y., S.M., G.H., and B.L. performed some experiments; Y.T. initiated the study, crossed mice, and analyzed data; and Z.F. initiated the study, organized, designed, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ye, B., Liu, B., Hao, L. et al. Klf4 glutamylation is required for cell reprogramming and early embryonic development in mice. Nat Commun 9, 1261 (2018). https://doi.org/10.1038/s41467-018-03008-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-03008-2
This article is cited by
-
GFAT1-linked TAB1 glutamylation sustains p38 MAPK activation and promotes lung cancer cell survival under glucose starvation
Cell Discovery (2022)
-
KLF17 promotes human naive pluripotency through repressing MAPK3 and ZIC2
Science China Life Sciences (2022)
-
CCN1 gene polymorphisms associated with congenital heart disease susceptibility in Northwest Chinese population from different high-altitude areas
Environmental Science and Pollution Research (2021)
-
Structural basis for polyglutamate chain initiation and elongation by TTLL family enzymes
Nature Structural & Molecular Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.