Abstract
A complex conformational energy landscape determines G-protein-coupled receptor (GPCR) signalling via intracellular binding partners (IBPs), e.g., Gs and β-arrestin. Using 13C methyl methionine NMR for the β1-adrenergic receptor, we identify ligand efficacy-dependent equilibria between an inactive and pre-active state and, in complex with Gs-mimetic nanobody, between more and less active ternary complexes. Formation of a basal activity complex through ligand-free nanobody–receptor interaction reveals structural differences on the cytoplasmic receptor side compared to the full agonist-bound nanobody-coupled form, suggesting that ligand-induced variations in G-protein interaction underpin partial agonism. Significant differences in receptor dynamics are observed ranging from rigid nanobody-coupled states to extensive μs-to-ms timescale dynamics when bound to a full agonist. We suggest that the mobility of the full agonist-bound form primes the GPCR to couple to IBPs. On formation of the ternary complex, ligand efficacy determines the quality of the interaction between the rigidified receptor and an IBP and consequently the signalling level.
Similar content being viewed by others
Introduction
G-protein-coupled receptors (GPCRs) relay an extracellular stimulus across the plasma membrane, activating cellular signalling pathways via coupling to intracellular binding partners (IBP) such as heterotrimeric G-proteins and β-arrestins. GPCRs control a wide range of physiological processes and are implicated in many disease states1. Rhodopsin-like GPCRs are estimated to be targeted by ~30% of current drugs2. β-adrenergic receptors (βAR) are essential in the sympathetic nervous system; β1AR is the predominant subtype in the heart, and is targeted by drugs such as β-blockers in the context of heart failure3. βARs show constitutive (basal) activity, with β1 at a lower but nevertheless significant level, compared to β2AR4, 5.
Over 170 crystal structures have been solved of nearly 40 unique receptors6, 7 in the presence of different ligands and IBPs8,9,10, substantially developing our understanding of how these receptors function. NMR studies indicate a complex, dynamic conformational energy landscape consisting of several states in equilibrium11,12,13,14,15,16,17,18,19,20, with substantial variations for different receptors. Allosteric coupling networks are observed17 along with correlations of ligand structure and efficacy with spectral parameters12, 17.
GPCRs are known to show varying degrees of constitutive activity21, with sampling by the receptor in the absence of a ligand (apo form) of activated states observed in fluorescence22 and NMR studies18. Addition of ligands shifts this equilibrium either towards the inactive form (inverse agonists) or towards activated conformations (agonists), which can couple to multiple intracellular signalling pathways18, 23.
Crystallisation of a receptor in the fully active state requires the presence of a cytoplasmic signalling partner. A range of native binding partners and mimetics (Gs, nanobodies, β-arrestins, mini-Gs and the transducin α-subunit C-terminal peptide)10 have been used, and the resulting active structures for the μ-opioid receptor (μOR)24, adenosine A2A receptor (A2AR)25, β2AR26, 27, muscarinic acetylcholine receptor M228 and rhodopsin29 are all very similar pointing towards a unified mechanism for receptor activation. The hallmarks of an active state are a large lateral displacement of the cytoplasmic end of transmembrane helix 6 (TM6) from the helix bundle, accompanied by a characteristic rearrangement of conserved residues (Arg3.50, Tyr5.58 and Tyr7.53 (superscripts refer to Ballesteros–Weinstein numbering30)) that seem to stabilise the receptor in its active conformation10. For agonist-bound A2AR, an intermediate-active state structure is found in the absence of IBP and there is also spectroscopic evidence for an active-like intermediate19, 31. No such equivalent has been observed for agonist-bound β1AR, reflecting subtle variations in the conformational energy landscape. However, NMR studies of β1- and β2AR indicate a ligand efficacy-dependent equilibrium between inactive and active states11, 12, 17. In addition, accelerated molecular dynamics (MD) simulations may be used to assess receptor activation, e.g., following the pathway from the active (ternary) structure of β2AR to the inactive receptor, potentially providing evidence for an intermediate conformation, as well as coupling between the ligand and G-protein-binding sites32. However, currently for β1AR, there are no reported active state structures.
Evidence indicates that a range of inactive, intermediate, active-like and active states are involved in GPCR function, which likely exist in equilibrium. However, crystallographic studies are unable to address questions regarding the role of receptor dynamics accompanying the process of activation, and further questions remain related to agonist efficacy, partial agonism, constitutive activity and sampling of active states in the absence of G-protein.
Here, we use NMR to investigate the conformational diversity and dynamic nature of coupling between activated receptor states of turkey β1AR and Gs-mimetic nanobodies. We show that β1AR in an active ternary complex has differing spectral signatures when bound to diverse activating ligands that correlate ligand efficacy, providing structural insight into the mechanism of partial agonism. Structural variations on TM5 and TM6 point to differences in the receptor–IBP interaction, which we hypothesise regulates the receptor’s guanine–nucleotide exchange factor (GEF) activity. Our study also indicates extensive conformational changes that affect large parts of the receptor during ligand-independent basal activation. We show that both the ternary complexes and the nanobody-only bound forms are less dynamic, while the full agonist-bound receptor in the absence of IBP is at its most dynamic form, indicating extensive sampling of different active-like conformational states that are primed to bind a range of intracellular signalling partners. This emphasises the vital role of receptor dynamics for coupling to a variety of different signalling pathways, for biased signalling and for partial agonism.
Results
β1AR constructs and NMR signal assignment
A thermostabilised turkey β1AR33 construct, modified from the previously published β44-m23 β1AR34 was functionally expressed in baculovirus-infected insect cells (Methods). Compared to the published construct, the level of thermostabilisation was substantially reduced through reverse mutagenesis of V90M2.53, A227Y5.58 and L282A (IL3). The final β1AR construct (Supplementary Fig. 1) contained R68S1.59, E130W3.41 and F327A7.37 as the only remaining thermostabilising mutations and C116L3.27 and C358A (CT) for improved expression yields17. Selective 13C methyl methionine labelling involved supplementing methionine-deficient SF4 growth media with labelled amino acid. To reduce overlap in the NMR spectra, the number of methionine residues was reduced from 12 (including the N-terminus) to seven through removal of M44L, M48L, M179L, M281A and M338A creating β1AR-MetΔ5, with methionines M1, M902.53 (TM2), M153 (IL2), M1784.62 (TM4/EL2), M2235.54 (TM5), M2836.28 (TM6) and M2966.41 (TM6) remaining (Supplementary Fig. 1b). For some samples, L108M (EL1) and L190M (EL2) were introduced providing additional reporters in extracellular loops 1 and 2. None of these additional methionines altered the thermal stability of the receptor. A comparison with residues in other class A GPCRs is shown in Supplementary Table 1. The expressed receptor was solubilised in lauryl-maltose-neopentyl-glycol (LMNG) and purified to 95% homogeneity judged by SDS–PAGE, using one-step nickel affinity chromatography. The methyl methionine signals of [13C-methyl-Met] β1AR-MetΔ5 were assigned in 1H, 13C SOFAST heteronuclear multiple quantum coherence (HMQC) spectra using a mutagenesis approach for a range of different ligand-bound states (Supplementary Fig. 2). In the case of agonistic ligands, this assignment approach was extended to the nanobody-bound ternary complexes and to the ligand-free nanobody-bound receptor form.
Correlations between ligand efficacy and chemical shift
To examine the effect of ligand binding, 1H, 13C HMQC spectra were recorded at 308 K for β1AR-MetΔ5 in the apo form and in the presence of saturating amounts of 7-methylcyanopindolol (very weak partial agonist)35, carvedilol (weak partial agonist), cyanopindolol (weak partial agonist), salbutamol (partial agonist), isoprenaline (full agonist) and adrenaline (full agonist), respectively (Supplementary Table 2)36. Apo- and ligand-bound spectra (Supplementary Fig. 3) showed substantial chemical shift changes for M2235.54 and M2966.41 (TM5, TM6), which correlated with the ligand efficacy (Fig. 1), suggesting an equilibrium involving two states exchanging fast relative to the chemical shift difference between the two signals, known as the NMR timescale (Supplementary Note 1). M2235.54 and M2966.41, located in the cytoplasmic half of the receptor, are sensitive reporters of the receptor’s activation state. Smaller changes were observed for M2836.28 and M153 (IL2) emphasising that structural changes related to ligand binding extend as far as the cytoplasmic side of the receptor and well beyond the ligand-binding pocket (Fig. 1).
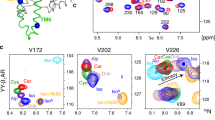
1H, 13C HMQC spectra of ligand-bound methyl 13C-Met β1AR-MetΔ5-L190M show resonance positions that correlate with ligand efficacies. Chemical shift changes for 2D 1H, 13C HMQC spectra for [13C-methyl-Met] β1AR shown for the apo form (blue) and orthosteric ligands in order of efficacy (h); 7-methylcyanopindolol (very weak partial agonist, orange), carvedilol (weak partial agonist, red), cyanopindolol (weak partial agonist, green), salbutamol (partial agonist, purple) and isoprenaline (full agonist, pink). The crystal structure of β1AR (PDB code 4BVN) is shown in b with methionine residues used in this study shown as red spheres. The various methionines are shown separately; a M90, c M153, e M223, d M283, i M296. The centres of the resonances are indicated by coloured dots. Correlations between ligand efficacy and a linear combination of the chemical shifts (\({\mathrm{a}}\delta _{1_{\mathrm{H}}} + {\mathrm{b}}\delta _{13_{\mathrm{C}}} + {\mathrm{c}}\)) are shown in f and g with M223 fit to \(- 1045.1\delta _{1_{\mathrm{H}}} - 111.0\delta _{13_{\mathrm{C}}} + 3839.8\) and M296 to \(- 634.1\delta _{1_{\mathrm{H}}} - 239.9\delta _{13_{\mathrm{C}}} + 5614.1\). Full spectra are shown in Supplementary Fig. 3
In β1AR, M902.53 is on the extracellular half of TM2 pointing towards the ligand-binding pocket. M902.53 samples a major and minor conformation in the unliganded state (M90a and M90b), indicative of at least two receptor conformations in the apo form (Fig. 1; Supplementary Fig. 3). Carvedilol-bound receptor showed two peaks for M902.53 at chemical shifts different from those in the apo form but close to the M90a state. For the partial agonists 7-methylcyanopindolol, cyanopindol and salbutamol, the major peak, which also corresponded to the major signal in the apo state, gradually shifted towards smaller ppm values. Notably, M902.53 was absent from the isoprenaline-bound spectrum, suggesting a more dynamic state in complex with a full agonist (Fig. 1; Supplementary Fig. 3). In addition, two shifts were observed for M1784.62 (TM4) in all states, consistent with sampling of at least two conformations by the EL2 region (Supplementary Fig. 3).
β1AR shows increased dynamics in the agonist-bound state
Loss of the M902.53 peak from the isoprenaline-bound β1AR spectra suggested increased signal broadening due to conformational exchange with rates similar to the NMR timescale (Supplementary Table 3). Short sample lifetimes and low experimental sensitivity limited the use of NMR spin relaxation tools, although these have been used for single-site measurements37, while interpreting signal linewidths was also unreliable (Supplementary Note 2). Instead, we used relative and normalised peak intensities as a proxy for detecting varying protein dynamics between individual receptor forms. Intensity variations were particularly pronounced for M902.53, M2235.54 and M2966.41. M2235.54 and M2966.41 are sensitive reporters of the activation state on the cytoplasmic side of β1AR and for each spectrum, their peak intensities were expressed as an intensity ratio relative to M153 (IL2) as a reference signal (Supplementary Table 4). Relative peak intensities for M2235.54 and M2966.41 were determined for two ligand series of β1AR-MetΔ5 and β1AR-MetΔ5-L190M, respectively (Supplementary Fig. 4). Smaller intensities indicated increasingly dynamic, exchange-broadened states. Normalised intensities were obtained by expressing the relative ratios as a proportion of the maximum peak intensity for a particular residue in a full series of spectra, where ‘1’ represents the least dynamic state and smaller values indicate increased conformational fluctuations on the μs-to-ms timescale (Fig. 2). While relative intensities allowed a direct comparison of M2235.54 and M2966.41 conformational dynamics, normalised intensities allowed comparison of the overall dynamics between the different receptor states and constructs. To eliminate bias from the reference signal, intensity ratios were also determined using the M190 (EL2) peak and the methyl signal of receptor-bound LMNG as the reference signal. Similar trends were obtained for all three cases (Supplementary Fig. 5a; Supplementary Table 5).
Normalised peak intensities of M223 and M296 reflect differences in dynamics that depend on the receptor state. Normalised peak intensities comparing residue M223 vs. M296 for the β1AR-MetΔ5 (a) and β1AR-MetΔ5-L190M (b) constructs. Relative intensities (Supplementary Fig. 4) were determined for each spectrum by calculating the peak intensity relative to the M153 signal. This was converted to a normalised intensity by setting the maximum intensity in each series to one, where one represents a lack of slow conformational fluctuations that lead to peak broadening and lower values represent increased µs-to-ms conformational dynamics. The different receptor states are shown as colour-coded circles: ligand bound (orange), apo (blue), nanobody bound (green) and ternary complex (red). The coloured bars along the top and the right-hand side of each graph indicate the direction of increasing µs–ms dynamics for residues M223 and M296, respectively. The red area indicates receptor states that are highly dynamic (µs–ms timescale) and blue indicates less dynamic, more rigid receptor states. Both constructs show a very similar pattern of intensities (dynamics) with full agonist-bound forms showing the greatest mobility for M223 and M296 and ternary complexes showing the greatest restriction in motion for these residues. Ligand-bound receptor and the apo form cluster around 0.5–0.6 normalised intensity units indicating some amount of motion (for a discussion of timescales see Supplementary Notes 1 and 2)
The full agonist-bound forms with isoprenaline or adrenaline showed consistently lower relative intensities for residues M2235.54 and M2966.41 than the partial agonist-bound forms (salbutamol, cyanopindolol and 7-methylcyanopindolol) or the apo state. Salbutamol-bound β1AR is more mobile than the other partial agonist-bound receptors but is still considerably more rigid than full agonist-bound receptor (Fig. 2). Very similar trends for the dynamics of the cytoplasmic side of TM5 and TM6 were also found for other mutants of β1AR-MetΔ5 (M178A, M223L, M283A and M296A) used for the residue-specific assignment of the methionine signals (Supplementary Fig. 6; Supplementary Table 4). The data consistency across a wide range of mutants and ligands supports the interpretation of signal intensities in the context of changes in receptor dynamics. However, other sources that influence R 2 relaxation, e.g., variations in dipolar interactions following side chain repacking could also affect peak heights.
At 308 K, M2235.54 and M2966.41 signals for isoprenaline-bound β1AR-MetΔ5-L190M were very weak. On lowering the temperature to 298 K, both signals increased in intensity, and a second signal for M2235.54 was resolved (Fig. 3), characteristic of intermediate exchange on the NMR timescale at 308 K, leading to extensive peak broadening; upon temperature reduction, the exchange became slow enough to resolve the exchanging states as individual signals (Supplementary Table 3). Further temperature reduction to 288 K preserved the more intense main signal compared to 308 K; however, it became difficult to determine the total number of signals as some apparent peaks were very close to the noise level. Both M2235.54 and M2966.41 13C chemical shift positions shifted towards the apo-receptor peak as the temperature was decreased (Supplementary Fig. 7; Supplementary Note 3).
Temperature dependence of chemical shifts and signal intensities in isoprenaline-bound β1AR-MetΔ5-L190M reveal a highly dynamic receptor. 2D 1H, 13C HMQC spectra (a) for the β1AR-MetΔ5-L190M construct recorded at 288 (blue), 298 (orange) and 308 K (red) showing the temperature dependence of the spectra. Resonance centres are indicated by coloured dots. All residues are seen to move with changing temperature, with substantial shifts for L190M and M1/M283, M223 and M296. Intensity changes are observed for M223 and M296 and at 298 K, M223 appears in two conformations (M223a and M223b). (Spectra were aligned using the isopropyl methyl group of unbound isoprenaline as a reference.) In b intensities, shown as signal-to-noise ratios (SNR), are plotted for M223 vs. M296 with uncorrected values in blue and corrected values accounting for the temperature and viscosity (η) changes (Supplementary Table 6) shown in orange. SNRs increase from 308 to 298 K indicating that at 308 K, M223 and M296 are in intermediate exchange leading to extensive peak broadening with a shift to slower exchange at lower temperatures, allowing separate states to be resolved as seen for M223 in a
Structural and dynamic changes in the ternary complex
To investigate conformational changes on forming the β1AR ternary complex, we recorded 1H, 13C HMQC spectra of isoprenaline-bound β1AR-MetΔ5-L190M after the addition of a stoichiometric amount of nanobody, using two nanobodies, Nb8026 and Nb6B938. In the ternary full agonist/receptor/nanobody complexes, both nanobodies are known to have nM-binding affinities to the receptor38. Substantial changes were observed upon formation of the fully active ternary complex through addition of Nb6B9 (Fig. 4). Peak positions for residues in the distant extracellular region were affected as well as in the cytoplasmic region proximal to the nanobody-binding site. The largest chemical shift changes were observed for M2235.54 and M2966.41 indicating substantial changes in TM5 and TM6 upon nanobody docking. Large displacements on the cytoplasmic side were also seen for M2836.28 (TM6) and M153 (IL2). The changes were much more substantial than for full agonist binding to the apo form (without nanobody) (Supplementary Fig. 8). Differences were also detected on the extracellular side for M1784.62 (TM4/EL2), M1905.21 (EL2) and M108 (EL1) (Fig. 4). Identical results were obtained for binding to Nb80 (Supplementary Fig. 2p).
Changes in resonance positions upon formation of the isoprenaline-bound β1AR-MetΔ5-L190M ternary complex with a nanobody. 2D 1H, 13C HMQC spectra for [13C-methyl-Met] β1AR-MetΔ5-L190M showing substantial chemical shift changes to isoprenaline-bound β1AR (blue) on addition of Nb6B9 nanobody (pink). Major changes from the isoprenaline-bound receptor to the ternary complex are indicated by arrows. The inset shows changes for M108 as observed in EL1 of β1AR-MetΔ5-L108M
Compared to ligand-only-bound receptor, normalised peak intensities for M2235.54 and M2966.41 were substantially increased in the ternary complex, indicating significantly more rigid receptor on the μs-to-ms timescale (Fig. 2). This was consistently observed for both nanobodies with β1AR-MetΔ5-L190M as well as for other β1AR-MetΔ5 mutant receptor constructs (M153A, M178A, M223L, M283A and M296A) (Supplementary Fig. 6).
Ligand efficacy affects ternary complex chemical shifts
The structural fingerprint of β1AR ternary complexes was investigated in the presence of the partial agonist cyanopindolol, agonist salbutamol and full agonist adrenaline and the spectral signatures compared with the isoprenaline-bound ternary complex, using the higher-affinity nanobody, Nb6B9. Changes were observed in both the intracellular and extracellular halves of the GPCR, with substantial shifts detected for M1784.62, and M2966.41 and smaller shifts for M153 (IL2), M2235.54 and M2836.28 (Fig. 5; Supplementary Fig. 9).
Resonance positions of ternary agonist complexes of β1AR-MetΔ5-L190M with a nanobody correlate with ligand efficacies. 2D 1H, 13C HMQC spectra for [13C-methyl-Met] β1AR-MetΔ5-L190M or [13C-methyl-Met] β1AR-MetΔ5 bound to Nb6B9 in the apo form (blue) or in a ternary complex with different orthosteric ligands; cyanopindolol (weak partial agonist, green), salbutamol (partial agonist, purple), isoprenaline (full agonist, pink) and adrenaline (full agonist, brown), in order of increasing ligand efficacy (k). The full spectrum for the apo form bound to Nb6B9 and full agonist- (isoprenaline) bound ternary complex is shown in a. Selected regions are also shown to illustrate the changes for residues M296 (b), M178 (c), M223 (f), M153 (g), M190 (h). Under each spectrum, a linear fit of ligand efficacy to a linear combination of the chemical shifts, \({\mathrm{a}}\delta _{1_{\mathrm{H}}} + {\mathrm{b}}\delta _{13_{\mathrm{C}}} + {\mathrm{c}}\), is shown. The equations determined were as follows: M178 (d), \(1510.6\delta _{1_{\mathrm{H}}} - 732.7\delta _{13_{\mathrm{C}}} + 9863.4\); M296 (e), \(- 5936.4\delta _{1_{\mathrm{H}}} + 5.93\delta _{13_{\mathrm{C}}} + 11985.7\); M223 (i), \(- 1607.4\delta _{1_{\mathrm{H}}} + 1258.4\delta _{13_{\mathrm{C}}} - 19056.1\) and M153 (j), \(- 11854.9\delta _{1_{\mathrm{H}}} + 2691.2\delta _{13_{\mathrm{C}}} - 18476.6\). Due to limited sample availability, two β1AR constructs were employed: β1AR-MetΔ5-L190M was used here with isoprenaline, cyanopindolol and in the apo form, while β1AR-MetΔ5 was used with salbutamol and adrenaline. Except for the absence of the M190 signal in the spectra of the latter, the two β1AR constructs were identical in terms of stability and binding properties and hence were used interchangeably
Notably, M2966.41 and to a lesser extent M153 (IL2), M1784.62 and M2235.54 showed a correlation between ligand efficacy and chemical shift positions (Fig. 5). As for nanobody-free receptor, M1784.62 shows two conformations, with only one showing an efficacy-dependent shift. While these measurements were done at 308 K, a repeat of the measurement for the salbutamol-bound β1AR-MetΔ5-L190M ternary complex at 293 K showed a small shift change for M2235.54 and M2966.41 away from the signal of the ternary isoprenaline towards smaller ppm (Supplementary Figs. 7, 10). Comparable observations are also made for the cyanopindolol- and isoprenaline-bound complexes. The shift changes correlating with ligand efficacy together with the observed temperature dependence suggested that M2235.54 and M2966.41 existed in equilibrium between states exchanging faster than the NMR timescale (Supplementary Table 3). For the salbutamol-bound ternary complex, the decrease in temperature caused a reduction in relative peak intensities for M2235.54 and M2966.41 consistent with fast exchange becoming comparable to the NMR timescale (Supplementary Table 6).
Ligand-free basal activity complex with a nanobody
In view of the reported basal activity for βAR5, structural evidence for an interaction between β1AR and G-protein mimetic nanobody in the absence of an agonist was sought. The nanobody–β1AR affinity was expected to be lower in the absence of agonists. Using a 15-fold nanobody excess, ligand-free receptor was observed predominantly as nanobody-bound (based on NMR signal integration: Nb6B9 99.5% bound; Nb80 96% bound). Substantial chemical shift changes were observed in the basal complex for both nanobodies relative to the apo form. Affected positions included residues at the extracellular side (M190 (EL2), M1784.62) and at the cytoplasm (M2235.54, M2966.41, M2836.28 and M153 (IL2)) (Fig. 6). Several residues showed slow exchange between the apo- and nanobody-bound receptor form (Supplementary Table 3), e.g., M2235.54 and M2966.41, allowing K d estimation using peak volumes. NMR titrations for β1AR-MetΔ5 and β1AR-MetΔ5-L190 in the apo form with Nb80 and Nb6B9 provided K d estimates of ca. 56 and 8 μM, respectively, which were several orders of magnitude weaker than in the presence of isoprenaline38.
Formation of a ligand-free basal activity complex results in large changes in resonance positions. 2D 1H, 13C HMQC spectra for [13C-methyl-Met] β1AR-MetΔ5-L190M showing substantial chemical shift changes to the apo form of β1AR (blue) on addition of Nb6B9 nanobody (orange). Major changes are indicated by arrows
For the receptor bound to Nb80 or Nb6B9, relative signal intensities for M2235.54 and M2966.41 were smaller than in the ternary agonist complexes but noticeably larger than in the apo- or ligand-bound receptor forms (Fig. 2; Supplementary Fig. 4).
Overall peak positions for the basal activity complex appeared more similar to the ternary complexes than the apo- or isoprenaline-bound receptor (Supplementary Fig. 11). Further small shifts on addition of isoprenaline resulted in a spectrum that was superimposable with the ternary complex formed by addition of a nanobody to isoprenaline-bound receptor (Supplementary Fig. 12). Isoprenaline addition affected residues including M108 (EL1), M178 (EL2) and M190 (EL2) on the extracellular side of the receptor and M2235.54 and M2966.41 on the cytoplasmic side. No changes were observed for M2836.28. Similar observations were made when adding cyanopindolol, salbutamol or adrenaline to Nb6B9-bound β1AR-MetΔ5.
Increased activation barrier for thermostabilised M90A
To further assess the influence of conformational dynamics on receptor activation, we introduced an additional thermostabilising mutation, M90A (β1AR-MetΔ5-M90A), known to increase T m for the alprenolol-bound (antagonist) β1AR34-324 by +8 °C39. M2966.41 signals for the isoprenaline- and salbutamol-bound states of the M90A mutant were slightly shifted towards the β1AR-MetΔ5 signals when bound to lower-efficacy ligands (Fig. 7a). Similar shift changes were also observed for M2235.54 with β1AR-MetΔ5-M90A bound to salbutamol. Conversely, signal positions in the corresponding apo form were almost unchanged (Supplementary Fig. 13). The ligand-only-bound form (salbutamol and isoprenaline) of M90A gave substantially higher intensity values for M2235.54 and M2966.41, indicating a more rigid and less exchange-broadened receptor following the additional thermostabilisation. The differential increase in relative intensities followed the order apo < salbutamol < isoprenaline, with little change for the apo form compared to β1AR-MetΔ5 (Fig. 7b). The relative signal intensities at 293 and 308 K showed no temperature dependence for M2966.41 and M2235.54 in the isoprenaline-bound state nor any field dependence (800 vs. 600 MHz), supporting a conformationally less dynamic, more homogeneous thermostabilised receptor (Supplementary Table 7a).
Additional thermostabilisation by M90A mutation shows β1AR-MetΔ5-M90A resonance positions and receptor dynamics characteristic of a less active receptor. 2D 1H, 13C HMQC spectra (a) for [13C-methyl-Met] β1AR-MetΔ5 (blue, green) and β1AR-MetΔ5-M90A (red, purple) in the ligand-bound form showing chemical shift changes with the orthosteric ligands isoprenaline (blue, red) and salbutamol (green, purple). Relative peak intensities (b) for ligand-bound and ternary complexes are shown for β1AR-MetΔ5 (blue) and β1AR-MetΔ5-M90A (red). The colour-coded bars at the top and the right-hand side of the peak intensity chart indicate the direction of increasing dynamics with generally less dynamic, rigid receptor states in blue, while red indicates receptor states which are increasingly dynamic on the µs-to-ms timescale. Compared to β1AR-MetΔ5, the M90A mutant shows more restricted dynamics in the ligand-only-bound forms, most notably for isoprenaline, indicating reduced conformational flexibility in this state. 2D 1H, 13C HMQC spectra (c) of the ternary isoprenaline/Nb6B9-bound β1AR-MetΔ5 (orange) and β1AR-MetΔ5-M90A (blue) show overall a similar spectral fingerprint but with distinct differences in the positions of M223 and M296 that are shifted towards the positions found in partial agonist-bound ternary complexes with the less stabilised β1AR-MetΔ5, i.e., towards (AG−)
Isoprenaline-bound M90A ternary complexes show a similar overall conformational signature to β1AR-MetΔ5 ternary complexes, with similar relative signal intensities (Fig. 7b; Supplementary Table 7b). However, the M2235.54 and M2966.41 signals of M90A were slightly shifted towards the partial agonist ternary complexes of β1AR-MetΔ5 (Fig. 7c).
Discussion
GPCRs require an IBP to populate their fully active state24,25,26,27,28,29, characterised by a large translation of the cytoplasmic end of TM6 compared to the agonist-bound state and the rearrangement of several key residues believed to stabilise the active state10, 40. GPCR activation is now understood within a complex conformational energy landscape, supported by recent NMR studies underpinning their highly dynamic nature11,12,13,14, 18,19,20. However, no active state structure of β1AR currently exists, and many questions remain surrounding the detailed mechanism of activation, e.g., how partial agonism manifests through G-protein interaction.
Using 13C methyl methionine NMR, we investigated a minimally thermostabilised turkey β1AR receptor. We studied apo- and ligand-bound states and their interaction with G-protein mimetic nanobodies (Nb80 and Nb6B9), assessing the dynamic nature of the receptor using a normalised intensity ratio for residues M2235.54 and M2966.41, which are located on the cytoplasmic side of TM5 and TM6 (Supplementary Note 2).
Ligand-bound β1AR spectra show considerable shift changes for residues near the ligand-binding pocket, e.g., M902.53 and on the cytoplasmic side of the receptor, e.g., M153 (IL2), M2235.54, M2836.28 and M2966.41 (Fig. 1; Supplementary Fig. 3). M902.53 shows multiple signals, with the major peak progressing to lower ppm values as ligand efficacy increases, comparable to M822.53 in β2AR (M90a, Fig. 1)12. However, M822.53 in β2AR12 shows a more correlated response to ligands that alter the population of the two states seen in the apo form and a third conformation observed in the presence of a full agonist12, 13. In contrast, M902.53 shows two conformations (M90a and M90c) for 7-methylcyanopindolol- and cyanopindolol-bound receptor, while apo receptor shows a different minor conformation, M90b, and carvedilol-bound receptor a second peak close to the M90a position. The salbutamol-bound form likely represents a state sampling more of the active conformation. No clear change in populations is seen with ligand efficacy. Although M822.53 is observed in formoterol-bound (full agonist) β2AR, other signals, e.g., M2155.54 (M2235.54 in β1AR) are missing, indicating increased motion in the full agonist-bound state consistent with signal loss in the isoprenaline-bound form of β1AR12. MD simulations for β2AR suggest that sterically unhindered exchange between different methyl side chain rotamer environments takes place on the ns timescale13, too fast to cause the observed multiple peaks and broadening, suggesting that the vicinity of the ligand-binding pocket in β1AR is in equilibrium between multiple slowly exchanging states with a substantial increase in intermediate exchange for isoprenaline-bound receptor.
M2235.54 and M2966.41 undergo large chemical shift changes correlating with ligand efficacy, indicating a conformational equilibrium in fast exchange (Supplementary Table 3) between the inactive state (I), unable to bind a G-protein, and the pre-active state (A), a more dynamic conformation with increased accessibility on the cytoplasmic side, populated in the full agonist-bound receptor (Fig. 1; Supplementary Note 3). The low basal activity of β1AR indicates that the apo form is almost entirely in the inactive state (I) with the cytoplasmic half of the receptor in the closed conformation. Spectral changes for M153 (IL2) and M2836.28, although smaller, are further evidence of orthosteric long-range effects that extend as far as the cytoplasmic face of the receptor.
Correlation with ligand efficacy was observed in a backbone valine 15N NMR study of tsβ1AR for Val 2265.57, although no strong correlation was observed for residues in TM6 (Val 2806.25 and Val 2986.43)17, possibly reflecting the increased sensitivity of side chain chemical shifts towards changes in conformational packing or the higher thermostabilisation of tsβ1AR which increases conformational rigidity. Notably, ligand-dependent changes in TM6 were observed in β2AR via the equivalent Met2796.41 reporter12.
Comparing normalised signal intensities for M2235.54 and M2966.41 shows that the full agonist-bound receptor signals are considerably weaker (~20% maximum intensity) than the other ligand-bound states and the apo form (~50% maximum intensity), indicating substantial broadening of the (A) state by μs-to-ms timescale exchange, with adrenaline-bound β1AR appearing even more dynamic than isoprenaline (Fig. 2; Supplementary Fig. 4). Lowering the temperature increases the intensity of these broad isoprenaline-bound receptor signals dramatically (Fig. 3a, b; Supplementary Fig. 7), suggesting a second, slower exchange process in the pre-active (A) state which is comparable to the NMR timescale, indicating sampling of multiple conformations by the (A) state. At 298 K, M2235.54 appears as two resolved signals, indicating several conformations in slow exchange (Supplementary Table 3).
Overall, our data suggest that the pre-active state (A) is highly mobile on a μs-to-ms timescale, both near the ligand-binding pocket and on the cytoplasmic side of the receptor. This indicates that the cytoplasmic side samples at least two, and possibly more active-like states (A = A′, A′′, A′′′…) in intermediate exchange on the NMR timescale at 308 K, leading to strong signal broadening. These states may couple to different IBPs, in which case the ability of the receptor to access different conformations in the pre-active state plays an important role in engaging different downstream-signalling pathways. When bound to lower-efficacy ligands or in the apo form, β1AR remains conformationally dynamic, however, less than in the full agonist-bound state. As the I ⇌ A equilibrium is shifted towards the more rigid (I) state, the receptor is conformationally less mobile as revealed by our intensity data (Fig. 2). A change in the lifetime of the (A = A′, A′′, A′′′…) states in the presence of different ligands is of course also possible.
Nanobodies, developed as a Gαs protein mimic26, were shown to interact with β1AR41, emulating the effect of Gαs-protein on the receptor and stabilising its active state. Isoprenaline-bound Nb80- or Nb6B9-β1AR-MetΔ5-L190M ternary complexes show dramatic spectral changes in the methionine region relative to isoprenaline-bound receptor (Fig. 4). The largest chemical shift changes are observed for M2235.54 and M2966.41, consistent with substantial structural rearrangements in TM5 and TM6 upon opening of the cytoplasmic half, as shown in β2AR crystal structures in complex with Nb6B9 or full G-protein26, 27. Methionine 13Cε chemical shifts report on χ 3 dihedral angles, indicating that for ligand-bound and apo states of β1AR, M2966.41 is in the trans conformation and M2235.54 shows trans/gauche exchange, shifting towards gauche on increased ligand efficacy, consistent with increased dynamics in the full agonist-bound form and the rotamer orientations in β2AR crystal structures (Supplementary Table 8). β2AR crystal structures were used due to bias in the existing β1AR structures towards antagonist-bound conformations, with limited changes in the cytoplasmic region following thermostabilisation (Supplementary Note 3; Supplementary Table 9)34, 42. On nanobody binding, M2966.41 shifts substantially towards the gauche conformation correlating with increased ligand efficacy. The gauche rotamer conformation also increases slightly for M2235.54 (Supplementary Note 3; Supplementary Fig. 14). As expected by their immediate proximity to the bound nanobody, residues M153 (IL2) and M2836.28 also show large shift changes.
Spectral changes between isoprenaline-bound receptor and the corresponding ternary complex are much more dramatic than those between apo- and isoprenaline-bound β1AR, suggesting that the (A) state corresponds to a pre-active rather than a fully active conformation (Supplementary Fig. 8). Notably, binding of a nanobody to the cytoplasmic side of the receptor transmits changes to the extracellular side of the receptor (‘outward’ effect), while binding of a full agonist in the orthosteric ligand-binding pocket results in changes as far as the cytoplasmic side of the receptor (‘inward’ effect) (Figs 1, 4; Supplementary Fig. 8). Nanobody-induced changes at the extracellular side of the ternary complex include M108 (EL1), M1784.62 (TM4/EL2) and M190 (EL2), suggesting an allosteric coupling network linking opposite faces of the receptor, where both the inward and outward effects reinforce ternary complex formation. An increase in agonist affinity frequently accompanies IBP binding and the observed conformational changes in EL1 and EL2 may be spectroscopic evidence for further contraction of the orthosteric ligand-binding environment26, 27, 41.
The structural fingerprint of Nb6B9 ternary complex formation was further investigated for other ligands (Supplementary Fig. 9). Spectra show large chemical shift changes for M2235.54 and M2966.41, characteristic of the cytoplasmic opening of the receptor and nanobody docking, as previously observed with isoprenaline. However, substantial shift differences in the ternary complexes compared to isoprenaline, which correlate with ligand efficacy (Fig. 5; Supplementary Fig. 9), indicate conformational variations among the ternary complexes relating to the specific orthosteric ligand bound. A loss in relative signal intensities suggests increased exchange broadening at lower temperature (Supplementary Table 6) and along with the observed upfield shifts at 293 K (Supplementary Figs. 7, 10; Supplementary Note 3) and the ligand efficacy correlation, indicates the presence of a two-state equilibrium for ternary complexes in fast exchange on the NMR timescale at 308 K (Supplementary Table 3).
Two mechanisms have been proposed to explain ligand-dependent efficacy: partial agonists may cause a distinct ternary complex structural state, with reduced activity compared to the full agonist43; alternatively, ligand efficacy may modulate an equilibrium between inactive and fully active ternary forms of the receptor44. Our observations suggest that ligand efficacy modulates both the equilibrium between inactive and pre-activated states, and the conformational equilibrium between a fully activated receptor ternary complex (AG+) and a less active ternary state (AG−), AG− ⇌ AG+ (Fig. 8). The two states likely differ in the ability of the cytoplasmic region to interact with IBPs, resulting in a variable response. As ligand efficacy increases, the receptor samples more of the fully activated (AG+) form, with adrenaline the most active ligand in our study. This is similar to the A2AR; however, no structural changes are observed between the pre-active state and the ternary complex possibly due to the use of a Gαs C-terminal peptide fragment or because A2AR adopts an almost fully active state in the absence of IBP19. As the study uses only one reporter (V229C6.31), structural changes distant from this site remain undetected. Ligand efficacy correlations are observed across the cytoplasmic and extracellular halves of β1AR, indicating that the AG− ⇌ AG+ equilibrium correlates with global changes in the receptor conformation (Fig. 5), which are in fast exchange on the NMR timescale (Supplementary Table 3). Relative signal intensities for M2235.54 and M2966.41 show that all ternary complexes are more rigid than the other receptor states in this study, consistent with receptor stabilisation on coupling to an IBP. The cyanopindolol-bound ternary complex remains slightly more mobile (~80% maximum intensity for M2235.54), consistent with its lower efficacy as it preferentially samples the less-active AG− state (Fig. 2; Supplementary Figs. 4, 6). In all ternary complexes, M2966.41 shows more exchange broadening than M2235.54, indicating residual TM6 motion (Supplementary Fig. 4).
Model of β1AR receptor activation showing partial agonism and basal activity. a β1AR fluctuates between conformations that vary in the accessibility of their cytoplasmic side. Ligand binding determines the equilibrium position between a closed, inactive state (I) and a less occluded, activated form (A) of the receptor and also regulates the position of a second equilibrium with the active receptor bound to Gs. Agonist binding leads to a highly dynamic (A) state of the receptor that samples multiple active-like conformations (A′, A′′, A′′′…), which vary in their ability to interact with IBPs. Coupling to Gs via these states leads to a more, AG+, and a less active form, AG−, which are in equilibrium with each other, with their relative populations influenced by the properties of the ligand bound. In the presence of a full agonist, both of these equilibria are shifted predominantly towards the (A) and in the presence of Gs, AG+ states. In contrast, when bound to inactivating ligands, or in the apo form, the receptor predominantly populates the (I) state. In the apo form, small populations of the (A) state in the presence of Gs can lead to the formation of the less active AG− state, consistent with basal activity at a low level. The basal activity form is subsequently able to bind a ligand (grey arrow), which can lead to a further shift of the active equilibrium towards the AG+ state. While our in vitro data show the formation of the ternary complexes via this route (grey arrow) (Supplementary Fig. 12), the physiological significance of this pathway is unclear and the main contribution towards ternary complex formation is likely to occur via initial binding of a ligand, followed by IBP docking (arrows in black). b The more and less active forms of the coupled receptor show conformational differences on TM5 and TM6 that modulate their ability to interact with GS. This could provide a mechanism to regulate signalling via the receptor’s GEF function affecting the rate of GDP release. In the presence of partial agonists, a shift in equilibrium in the direction of AG− results in reduced levels of receptor signalling and lower efficacy
βAR are reported to show basal activity when bound to IBPs5. For the apo state, we measured nanobody affinity in the μM range, substantially reduced from the nM affinity observed in ternary complexes38, 41. Adding an excess of nanobody demonstrated that nanobody-bound ligand-free receptor spectra have strong similarities to ternary nanobody complexes, consistent with an active state (Fig. 6; Supplementary Fig. 11).
Spectral differences between the basal form and the ternary complexes are small compared to the (unliganded) apo form, and together with the lack of substantial changes in the cytoplasmic residues M153 (IL2) and M2836.28 suggest that the main structural changes in TM6 and the cytoplasmic region occur on nanobody binding (Supplementary Fig. 11). This is endorsed by the normalised signal intensities of M2235.54 and M2966.41, which show more restricted dynamics for the basal β1AR–nanobody complexes, closer to the ternary complexes than the apo receptor (Fig. 2). It also suggests that ligand binding causes further tightening of the receptor on ternary complex formation.
Isoprenaline addition to the preformed nanobody-bound ligand-free receptor results in identical spectra to the ternary complex formed upon nanobody addition to the isoprenaline-bound receptor (Supplementary Fig. 12). Similar observations are made for the other agonists (cyanopindolol, salbutamol and adrenaline) indicating that the final conformation of the ternary complex is independent of the binding order.
Nanobody complex formation and spectral similarities between the basal state and the ternary complexes suggest that sampling of the pre-activated (A) state by the apo form of β1AR without an agonist20, 22 can allow weak coupling to G-protein with the potential for downstream signalling, providing an explanation for the basal activity. M2235.54 and M2966.41 chemical shifts in the basal state correlate with the lowest-efficacy level for nanobody-bound complexes investigated (Fig. 5), suggesting that the basal state corresponds to the minimally activated (AG−) complex in the AG− ⇌ AG+ equilibrium. Given the existence of ligand-free basal-state GPCR-G protein complexes, and the flexibility of the (A) state, receptor selectivity for intracellular signalling pathways may, in part, be influenced by which IBPs are expressed and present at the cell membrane in different cell types, potentially explaining the ability of a given ligand to activate multiple signalling pathways.
Spectra for the thermostabilising M90A mutant (β1AR-MetΔ5-M90A)39 in the apo state are hardly changed from β1AR-MetΔ5 (Supplementary Fig. 13), indicating that in the absence of a ligand, the thermostabilising mutation has little effect. The shift of M2235.54 and M2966.41 signals in ligand-bound spectra of β1AR-MetΔ5-M90A towards those from β1AR-MetΔ5 when bound to lower-efficacy ligands, both in the absence and presence of Nb6B9 (Fig. 7a, c), implies that the thermostabilising mutation reduces receptor activity. Temperature-dependent chemical shift changes of similar size and sign to β1AR-MetΔ5 are still observed for M2235.54 and M2966.41 , emphasising that the fast-exchanging equilibria, I ⇌ A and AG− ⇌ AG+, still exist, albeit shifted towards the less active state for each equilibrium (Supplementary Fig. 7).
Relative peak intensities show dramatically reduced μs-to-ms dynamics for isoprenaline- and salbutamol-bound β1AR-MetΔ5-M90A, consistent with a shift in the equilibria, revealing substantially more rigid agonist-bound receptor (Fig. 7a, b), while the dynamics of the apo form is hardly affected. The relative signal intensities do not show any temperature or field dependence, indicating reduced sampling of the (A) state via a left-shift in the I ⇌ A equilibrium (Supplementary Table 7). Based on our data, it is currently not clear if stabilisation also changes the dynamics of the (A) state, which might change the sampling of the active-like (A′, A′′, A′′′…) conformations leading to sharpening of the M2235.54 and M2966.41 signals or if the (A) state is just less populated. As the receptor retains its ability to form ternary complexes, it can be assumed that such states are being sampled. The ternary complexes show no change in dynamics compared to β1AR-MetΔ5. Overall, thermostabilisation is similar to reducing the efficacy of a particular ligand, shifting ligand-bound and ternary complex equilibria towards the less active state.
Using NMR spectroscopy, we investigated the conformational diversity and dynamic nature of β1AR activation through orthosteric ligands and Gs-mimetic nanobodies. We provide structural insight into partial agonism and basal activity, presenting our conclusions as a model (Fig. 8). The receptor populates two ligand efficacy-dependent equilibria, between an inactive (I) and pre-activated (A) form and, for the receptor bound to Gs-mimetic, between more (AG+) and less (AG−) active forms of the nanobody-bound receptor. The less active IBP-bound form is consistent with the conformation of the basal activity complex. Full agonist binding results in a highly dynamic pre-activated form that samples multiple active-like conformations, competent to bind IBPs. Two different conformations can be distinguished at lower temperature with additional broadened or low-populated states possibly remaining undetected. We suggest that the different states enable preferential interaction with particular IBPs with the relative populations of these states modulated by different ligands. Bound to a Gs-mimetic, the receptor becomes less mobile with allosteric modulation of the cytoplasmic interaction by the ligand in the orthosteric binding pocket. According to our data, orthosteric ligand efficacy is converted into a distinct conformational response on the cytoplasmic side. Differences between the (AG−) and (AG+) states occur on TM5 and TM6, indicating that these conformational variations may modulate the receptor–nanobody interaction. We speculate that when β1AR is bound to Gαs, these differences may contribute to conformational changes regulating the rate of GDP release, providing control of the GEF function of the GPCR. Through this mechanism, ligands may modulate the efficacy of downstream signalling by the receptor.
Methods
Constructs and mutagenesis
The β1AR-Met2 construct used here was derived from the truncated turkey β1AR receptor β44-m2334; through the reversal of three stabilising mutations (V90M2.53, A227Y5.58 and L282A (IL3)), and the inclusion of two further mutations (E130W3.41 and D322K7.32). To reduce spectral overlap, five methionine residues were mutated (M44L1.35, M48L1.57, M179L (EL2), M281A (IL3), M338A7.48) along with a further reversal of the stabilising mutation (K322D7.32), creating the construct Met2-Δ5. Therefore, Met2-Δ5 differs from the wild-type receptor in the truncation of the N- and C-termini, and of intracellular loop 3 (IL3), in addition to the presence of four thermostabilising point mutations, the mutation of five methionine residues as well as the presence of a mutation increasing expression yield (C116L3.27) and the removal of a palmitoylation site (C358A). For ease of purification, a C-terminal octahistidine tag was added. The sequence of Met2-Δ5 is.
MGAELLSQQWEA GLSLLLALVVLLI VAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVMGLLVVPFGA TLVVRGTWL WGSFLCELWTSL DVLCVTASIWT LCVIAIDRYLAITS PFRYQSLM TRARAKVIICTVW AISALVSFLPIMLHWWRDEDPQALKCYQDP GCCDFVTN RAYAIASSIISFYIPL LIMIFVYLRVYREAKE QIRKIDRASKR KTSRVAAMREH KALKTLGIIMGVFT LCWLPFFLVNIVNVFN RDLVPDWLFVAFNWLGY ANSAANP IIYCRSPDF RKAFKRLLAFPRKADRRLHHHH HHHH.
For assignment purposes, methionine residues were substituted using site-directed mutagenesis in a PCR reaction (primer sequences given in Supplementary Table 10) using Phusion DNA polymerase (ThermoFisher). DpnI-digested PCR product was transformed into DH5α Escherichia coli cells (New England Biolabs) along with the transfer plasmid, and the resulting plasmid DNA construct was isolated using a commercial Miniprep kit (Qiagen) ready for transfection in insect cells.
Expression and purification of β1AR
Baculovirus was generated for expression using FlashBac viral DNA (Oxford Expression Technologies). β1AR containing pBacPAK8 plasmid was diluted to 100 ng µL−1 and 1.8 µL, together with 1.8 µl of FlashBac DNA, was added to 1.8 µL of previously dissolved and NaOH-neutralised polyethylenimine (linear PEI 25000, Polysciences, 1 mg mL−1 concentration) and diluted with 360 µL of cell culture media (SF4, Bioconcept). The mixture was incubated at room temperature for 45 min to allow for DNA–polymer complexes to form. The complex mixture was added to 1 mL of mid-log phase Sf9 cells (ThermoFisher), diluted to 0.5 × 106 cells mL−1 in serum-free SF4 media and incubated at 27 °C shaking for 5 days. On day 5, a small sample was taken to confirm visual signs of infection. The resulting P0 viral stock was diluted to 4 mL with fresh media and incubated for 48 h to generate P1 viral stock. An aliquot of 100 µL of P1 stock was used to infect 50 mL of cells at a density of 1 × 106 cells mL−1, and incubated at 27 °C shaking for 48 h, generating a high-titre P2 stock for protein expression45.
For β1AR expression, Sf9 or Sf21 cells (ThermoFisher), grown in serum-free SF4 media were centrifuged (500×g, 10 min) and washed with sterile PBS, to reduce the carry-over of unlabelled methionine. The washed cells were diluted to a density of 3 × 106 cells mL−1 into methionine-deficient SF4 media at half the intended final culture volume. The culture was then infected with 4 mL L−1 of high-density viral stock, and incubated for 5 h, before supplementing the culture with 250 mg L−1 of 13C methyl methionine and diluting to a final density of 1.5 × 106 cells mL−1. The initial reduction in culture volume ensures optimal aeration in the initial phase of the viral infection. Cells were grown at 27 °C for 48 h and were harvested by centrifugation (3500×g, 15 min).
The frozen insect cell pellet was thawed with solubilisation buffer (20 mM Tris-HCl, pH 8.0, 350 mM NaCl, 3 mM imidazole, Complete Protease Inhbitor Cocktail (Roche) and 2% LMNG) and stirred for 1 h. The solubilised cells were clarified by centrifugation (175,000×g, 45 min) and the soluble fraction was loaded onto a nickel affinity column. The column was washed with equilibration buffer (20 mM Tris-HCl, pH 8.0, 350 mM NaCl, 3 mM imidazole and 0.02% LMNG) and the protein was eluted with the same buffer supplemented with 250 mM imidazole. The final sample was exchanged into 10 mM Tris-HCl, pH 8.0, 150 mM NaCl and 0.04% LMNG.
E. coli expression and purification of Nb80 and Nb6B9
Nb80- and Nb6b9-containing plasmids were transformed into BL21(DE3) (New England Biolabs) cells and grown in LB media supplemented with the relevant antibiotics. Expression cultures were inoculated with a saturated overnight culture to an OD600 density of 0.1 AU and grown to a density of 0.8 AU at 37 °C. Induction was achieved with isopropyl thiogalactopyranoside (IPTG) at a final concentration of 0.5 mM. Expression cultures were grown at 25 °C for 16 h before harvesting by centrifugation (3500×g, 20 min, 4 °C).
Frozen cell pellets were thawed in 20 mM Tris-HCl, pH 8.0, 150 mM NaCl with Complete Protease Inhibitor Cocktail (Roche) and lysed with an EmulsiFlex C5 homogeniser (Avestin). The solubilised cells were clarified by centrifugation (175,000×g, 15 min) and the soluble fraction was loaded onto a nickel affinity column. Unbound sample was removed with wash buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl), followed by a further wash of the same composition supplemented with 6 mM imidazole. Nanobodies were eluted with wash buffer containing 250 mM imidazole. A further size-exclusion chromatography step on a Superdex S200 10/300 Increase column yielded a 95% pure protein preparation in 20 mM Tris-HCl, pH 8.0, 150 mM NaCl. Typical yields of Nb80 and Nb6b9 obtained by this method were 25 and 4 mg L−1, respectively.
NMR experiments
NMR samples were prepared with 5% D2O and the ligand was added directly to the receptor in the apo form. The final ligand concentrations were 1 mM salbutamol, isoprenaline, adrenaline, 200 µM carvedilol, 140 µM or 600 µM S(−)-cyanopindolol and 100 µM 7-methylcyanopindolol. Ligand-bound populations were all > 99.9% (Supplementary Table 11). Nanobody was added in a molar excess for ternary complex formation and in 15-fold excess for basal complex formation. NMR experiments were recorded on a Bruker Avance AVIII 800 spectrometer (1H 800 MHz) equipped with a 5-mm TXI HCN/z cryoprobe or where specified on a Bruker Avance AVIII 600 spectrometer (1H 600 MHz) equipped with a 5-mm QCI HCNF/z cryoprobe.
Unless specifically mentioned experiments were acquired at 308 K using a SOFAST 1H, 13C HMQC experiment46 with gradient coherence-order selection47 and non-uniform sampling (NUS) in the 13C dimension. Gradient selection was required to reduce the intense LMNG detergent signals, while selective excitation of methyl groups enabled use of a short recycle delay of 0.5 s. Excitation used a 2.25-ms 120° PC9 pulse48 and inversion a 1.16-ms, 180° REBURP pulse49. A 60% Poisson-gap sampling schedule was used (60 complex points from a total of 100 complex points)50, with a maximum acquisition time (t max) of 25 ms and spectral width (13C) of 4000 Hz. The direct (1H) dimension was acquired with 10,000 Hz spectral width, 1024 points and t max = 51.2 ms. Spectra were recorded with 368 scans, giving an acquisition time of ~6 h. Where higher sensitivity was required, multiple 6 h experiments were recorded and added. Spectra were reconstructed using the iterative hard thresholding (IHT) compressed sensing (CS) implementation51 in the Cambridge CS package (M.J. Bostock, unpublished). Data were analysed using CCPN Analysis v252.
Data availability
The authors declare that relevant data supporting the findings of this study are available within the article and its Supplementary Information files or on reasonable request from the corresponding author. NMR shifts are available at the BioMagResBank (accession numbers 27292–27297).
References
Lagerström, M. C. & Schiöth, H. B. Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat. Rev. Drug Discov. 7, 339–357 (2008).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Taylor, M. R. G. Pharmacogenetics of the human beta-adrenergic receptors. Pharmacogenomics J. 7, 29–37 (2007).
Bond, R. A. et al. Physiological effects of inverse agonists in transgenic mice with myocardial overexpression of the β2-adrenoceptor. Nature 374, 272–276 (1995).
Engelhardt, S., Grimmer, Y., Fan, G. H. & Lohse, M. J. Constitutive activity of the human β1-adrenergic receptor in β1-receptor transgenic mice. Mol. Pharmacol. 60, 712–717 (2001).
Munk, C. et al. GPCRdb: structures overview. Available at http://gpcrdb.org/structure/statistics (accessed 18th May 2017).
Munk, C. et al. GPCRdb: the G protein-coupled receptor database-an introduction. Br. J. Pharmacol. 173, 2195–2207 (2016).
Rosenbaum, D. M., Rasmussen, S. G. F. & Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009).
Cooke, R. M., Brown, A. J. H., Marshall, F. H. & Mason, J. S. Structures of G protein-coupled receptors reveal new opportunities for drug discovery. Drug Discov. Today 20, 1355–1364 (2015).
Carpenter, B. & Tate, C. G. Active state structures of G protein-coupled receptors highlight the similarities and differences in the G protein and arrestin coupling interfaces. Curr. Opin. Struct. Biol. 45, 124–132 (2017).
Kofuku, Y. et al. Functional dynamics of deuterated β2-adrenergic receptor in lipid bilayers revealed by NMR spectroscopy. Angew. Chem. Int. Ed. 53, 13376–13379 (2014).
Kofuku, Y. et al. Efficacy of the β2-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 3, 1045 (2012).
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013).
Okude, J. et al. Identification of a conformational equilibrium that determines the efficacy and functional selectivity of the μ-opioid receptor. Angew. Chem. Int. Ed. 54, 15771–15776 (2015).
Sounier, R. et al. Propagation of conformational changes during μ-opioid receptor activation. Nature 524, 375–378 (2015).
Casiraghi, M. et al. Functional modulation of a G protein-coupled receptor conformational landscape in a lipid bilayer. J. Am. Chem. Soc. 138, 11170–11175 (2016).
Isogai, S. et al. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 530, 237–241 (2016).
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wüthrich, K. Biased signaling pathways in β2-adrenergic receptor characterised by 19F-NMR. Science 335, 1106–1110 (2012).
Ye, L., Van Eps, N., Zimmer, M., Ernst, O. P. & Prosser, R. S. Activation of the A2A adenosine G-protein-coupled receptor by conformational selection. Nature 533, 265–268 (2016).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
de Ligt, R. A. F., Kourounakis, A. P. & IJzerman, A. P. Inverse agonism at G protein-coupled receptors: (patho)physiological relevance and implications for drug discovery. Br. J. Pharmacol. 130, 1–12 (2000).
Lamichhane, R. et al. Single-molecule view of basal activity and activation mechanisms of the G protein-coupled receptor β2AR. Proc. Natl Acad. Sci. USA 112, 14254–14259 (2015).
Marinissen, M. J. & Gutkind, J. S. G-protein-coupled receptors and signaling networks: emerging paradigms. Trends Pharmacol. Sci. 22, 368–376 (2001).
Huang, W. et al. Structural insights into µ-opioid receptor activation. Nature 524, 315–321 (2015).
Carpenter, B., Nehmé, R., Warne, T., Leslie, A. G. W. & Tate, C. G. Structure of the adenosine A2A receptor bound to an engineered G protein. Nature 536, 104–107 (2016).
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature 469, 175–180 (2011).
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523, 561–567 (2015).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Dror, R. O. et al. Activation mechanism of the β2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011).
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008).
Warne, T. et al. The structural basis for agonist and partial agonist action on a β1-adrenergic receptor. Nature 469, 241–244 (2011).
Sato, T. et al. Pharmacological analysis and structure determination of 7-methylcyanopindolol-bound β1-adrenergic receptor. Mol. Pharmacol. 88, 1024–1034 (2015).
Baker, J. G., Proudman, R. G. W. & Tate, C. G. The pharmacological effects of the thermostabilising (m23) mutations and intra and extracellular (β36) deletions essential for crystallisation of the turkey β-adrenoceptor. Naunyn Schmiedebergs Arch. Pharmacol. 384, 71–91 (2011).
Kim, T. H. et al. The role of ligands on the equilibria between functional states of a G protein-coupled receptor. J. Am. Chem. Soc. 135, 9465–9474 (2013).
Ring, A. M. et al. Adrenaline-activated structure of β2-adrenoceptor stabilized by an engineered nanobody. Nature 502, 575–579 (2013).
Serrano-Vega, M. J., Magnani, F., Shibata, Y. & Tate, C. G. Conformational thermostabilization of the β1-adrenergic receptor in a detergent-resistant form. Proc. Natl Acad. Sci. USA 105, 877–882 (2008).
Venkatakrishnan, A. J. et al. Diverse activation pathways in class A GPCRs converge near the G-protein-coupling region. Nature 40, 484–487 (2016).
Miller-Gallacher, J. L. et al. The 2.1 Å resolution structure of cyanopindolol-bound β1-adrenoceptor identifies an intramembrane Na+ ion that stabilises the ligand-free receptor. PLoS ONE 9, e92727 (2014).
Lebon, G., Warne, T. & Tate, C. G. Agonist-bound structures of G protein-coupled receptors. Curr. Opin. Struct. Biol. 22, 482–490 (2012).
Kobilka, B. K. & Deupi, X. Conformational complexity of G-protein-coupled receptors. Trends Pharmacol. Sci. 28, 397–406 (2007).
Lape, R., Colquhoun, D. & Sivilotti, L. G. On the nature of partial agonism in the nicotinic receptor superfamily. Nature 454, 722–727 (2008).
Roest, S. et al. Transfection of insect cell in suspension for efficient baculovirus generation. MethodsX 3, 371–377 (2016).
Schanda, P., Kupče, Ē. & Brutscher, B. SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J. Biomol. NMR 33, 199–211 (2005).
Davis, A. L., Keeler, J., Laue, E. D. & Moskau, D. Experiments for recording pure-absorption heteronuclear correlation spectra using pulsed field gradients. J. Magn. Reson. 98, 207–216 (1992).
Kupče, Ē. & Freeman, R. Polychromatic selective pulses. J. Magn. Reson. Ser. A 102, 122–126 (1993).
Geen, H. & Freeman, R. Band-selective radiofrequency pulses. J. Magn. Reson. 93, 93–141 (1991).
Hyberts, S. G., Takeuchi, K. & Wagner, G. Poisson-gap sampling and forward maximum entropy reconstruction for enhancing the resolution and sensitivity of protein NMR data. J. Am. Chem. Soc. 132, 2145–2147 (2010).
Bostock, M. J., Holland, D. J. & Nietlispach, D. Compressed sensing reconstruction of undersampled 3D NOESY spectra: application to large membrane proteins. J. Biomol. NMR 54, 15–32 (2012).
Vranken, W. F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Acknowledgements
This work was funded in parts through a BBSRC research grant to D.N. (BB/K01983 X/1) and studentship support to A.S. (MRC Industrial CASE).
Author information
Authors and Affiliations
Contributions
A.S., M.J.B. and D.N. designed the research. A.S. performed molecular biology, protein expression and purification. Protein expression was carried out in collaboration with B.S. P.K. helped with molecular biology and construct design. T.W. and C.G.T. provided initial receptor and nanobody constructs and assisted with receptor biochemistry. A.S., M.J.B. and D.N. carried out NMR data collection, data processing, spectral assignments and analysed the data. M.J.B., A.S. and D.N. prepared the manuscript. D.N. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Solt, A.S., Bostock, M.J., Shrestha, B. et al. Insight into partial agonism by observing multiple equilibria for ligand-bound and Gs-mimetic nanobody-bound β1-adrenergic receptor. Nat Commun 8, 1795 (2017). https://doi.org/10.1038/s41467-017-02008-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-017-02008-y
This article is cited by
-
Structural and dynamic insights into the activation of the μ-opioid receptor by an allosteric modulator
Nature Communications (2024)
-
Binding kinetics drive G protein subtype selectivity at the β1-adrenergic receptor
Nature Communications (2024)
-
Structures of β1-adrenergic receptor in complex with Gs and ligands of different efficacies
Nature Communications (2022)
-
Vibrational spectroscopy analysis of ligand efficacy in human M2 muscarinic acetylcholine receptor (M2R)
Communications Biology (2021)
-
Conformational plasticity of ligand-bound and ternary GPCR complexes studied by 19F NMR of the β1-adrenergic receptor
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.