Abstract
Mitochondrial dysfunction manifests as different neurological diseases, but the mechanisms underlying the clinical variability remain poorly understood. To clarify whether different brain cells have differential sensitivity to mitochondrial dysfunction, we induced mitochondrial DNA (mtDNA) depletion in either neurons or astrocytes of mice, by inactivating Twinkle (TwKO), the replicative mtDNA helicase. Here we show that astrocytes, the most abundant cerebral cell type, are chronically activated upon mtDNA loss, leading to early-onset spongiotic degeneration of brain parenchyma, microgliosis and secondary neurodegeneration. Neuronal mtDNA loss does not, however, cause symptoms until 8 months of age. Findings in astrocyte-TwKO mimic neuropathology of Alpers syndrome, infantile-onset mitochondrial spongiotic encephalopathy caused by mtDNA maintenance defects. Our evidence indicates that (1) astrocytes are dependent on mtDNA integrity; (2) mitochondrial metabolism contributes to their activation; (3) chronic astrocyte activation has devastating consequences, underlying spongiotic encephalopathy; and that (4) astrocytes are a potential target for interventions.
Similar content being viewed by others
Introduction
Primary mitochondrial dysfunction is the most common cause of inherited metabolic disease1. The central nervous system manifestations of mitochondrial diseases are typically progressive, with manifestations ranging from infantile catastrophic epileptic encephalopathy to ataxia-epilepsy syndrome, leukoencephalopathy, neurodegeneration or parkinsonism1,2. Among the most severe disorders are Leigh syndrome, a subacute necrotizing encephalopathy, and Alpers–Huttenlocher syndrome with spongiotic changes, laminar cortical necrosis and liver failure3,4,5. These devastating, lethal disorders were both characterized originally by their typical neuropathological signs, which are the key determinant of the clinical diagnosis even today6,7.
Alpers–Huttenlocher syndrome is caused by recessive mutations in mitochondrial DNA (mtDNA) maintenance proteins: the nuclear-encoded DNA polymerase gamma or replicative helicase Twinkle8,9. These proteins, together with the mitochondrial transcription factor A (TFAM) form the minimal mtDNA replisome, and are essential for faithful mtDNA replication10: their total inactivation in mice is embryonic lethal at ~E8.511,12,13. Knocking out TFAM or expressing mitochondrial-targeted endonuclease in forebrain or dopaminergic neurons of mice resulted in progressive loss of mtDNA, eventual death of the targeted neurons and shortened lifespan of the animals14,15,16. The direct consequence of mtDNA loss is deficiency of the 13 mtDNA-encoded subunits that together with 76 nuclear-encoded subunits build up the multisubunit enzyme complexes of oxidative phosphorylation (OXPHOS) in which the Complexes I–IV (CI-CIV) form the respiratory chain (RC) and Complex V is the ATP synthase. The late manifestation of neurological symptoms in neuronal TFAM-knockout mice and the mitochondrial endonuclease over-expressor mice indicates that neurons can sustain relatively prolonged periods of mtDNA depletion and severe RC deficiency, questioning the role of neurons as the primary culprit in mtDNA maintenance disorders.
Astrocytes are the most abundant cell type in the brain, providing trophic and metabolic support for neurons, pruning synapses and synthesizing neurotransmitters17. They are metabolically dynamic, and considered to be glycolytic, i.e. to have capacity to perform non-oxygen-dependent ATP-synthesis and survive OXPHOS deficiency in vivo18,19. The contribution of astrocytes in mitochondrial disorders has been considered to be secondary, forming a glial scar at the sites of neuronal damage. However, no evidence exists of the importance of mtDNA maintenance in astrocytes in vivo.
Here we investigate the consequences of inactivation of Twinkle, the mtDNA replicative helicase, in astrocytes or neurons in vivo. We show that mtDNA maintenance is critical for both cell types, but in a different manner. Progressive loss of Twinkle and mtDNA in neurons causes late-onset neurodegeneration and rapid progressive late-stage neurological deterioration. Twinkle-KO in astrocytes, however, leads to wide-spread activation of the cell type, severe early-onset progressive neurological disease, accompanied by spongiotic encephalopathy and inflammation. The results propose that primary astrocyte dysfunction underlies mitochondrial spongiotic encephalopathies, such as Alpers syndrome, caused by mtDNA maintenance defects.
Results
Generation of constitutive and conditional Twinkle-KO mice
To clarify whether mtDNA maintenance is similarly essential for neurons and glia, we designed cell-type-specific conditional knockout mice for the Twnk gene encoding mitochondrial DNA helicase Twinkle (TwKO) (Fig. 1a). We first generated early ubiquitous inactivation of Twinkle, using PGK-promoter, driving cre-recombinase expression from the zygote stage20. These homozygous full-TwKO embryos showed drastically reduced mtDNA content already at E7.5 leading to lethality at E8.5, as previously described12 (Fig. 1b). Then we targeted Twinkle knockout (TwKO) to postnatal forebrain neurons (TwKOneuro; CamKII-cre) or astrocytes (TwKOastro; mouseGFAP73.12-cre) (Fig. 1a). From the existing transgenic cre-lines targeting astrocytes, we chose the mGFAP73.12 promoter since (1) it starts to express postnatally, allowing brain to develop normally; (2) embryonic neuronal stem cells remain unaffected; (3) the profile of expression is well characterized and specific for astrocytes21,22,23, except for some expression in a population of adult neuronal stem cells generating a small number of neurons during adult life. Furthermore, cre-activation affects Twinkle protein and mtDNA amounts with a delay, the latter having a half-life in brain of several weeks24. Reporter mice expressing GFP upon recombination, crossed with mGfap73.12, indeed did not demonstrate GFP-expression at P2, and in adult mice showed intensive expression in cortex, hippocampus and cerebellum (Supplementary Fig. 1), and low expression in midbrain, resembling the pattern of expression previously reported for this promoter. The TwKOneuro and TwKOastro lines were viable, and the cre transgene segregated to Mendelian ratio (TwKOneuro 42/78 and TwKOastro 24/41). The successful TwKO was confirmed by Twinkle cDNA analysis (Fig. 1c).
Knockout of mtDNA maintenance in astrocytes causes early-onset neurodegenerative disease. a Cre-promoters and timing of activation (PGK = phosphoglycerate kinase, GFAP = glial fibrillary-acidic-protein, CaMKII = calcium/calmodulin-dependent protein kinase-II). Targeting construct: Boxes = exons, triangles = LoxP sites. b MtDNA amount in constitutive Twinkle knockout (TwKO), embryonal day 7.5 (Ctr = control, Het = heterozygote; TwKO = homozygous). c Size of Twinkle-cDNA in TwKO lines; cerebral cortex. d TwKO mouse phenotypes at the age of 6 months. TwKOastro mice: small, paraparetic posture. TwKOneuro mice are control-like. e Body weight progression: red = TwKOastro, blue = TwKOneuro, black = Control. **p < 0.01 for the first time-point; subsequent time-points p < 0.05. f Automated grip strength test for forelimbs (6 months of age). g Beam walk test for motor coordination (large d = large diameter, small d = small diameter beam). h Kaplan–Meier survival curves. Red = TwKOastro, blue = TwKOneuro, black = Control. i Brains of TwKOastro, TwKOneuro, and Control mice are similar. Scale bar is 0.5 cm. The data are presented as mean and error bars indicate standard deviation. Statistical significance: *p < 0.05, **p < 0.01
TwKO in astrocytes causes early-onset neurological disease
Postnatal inactivation of Twinkle in neurons versus astrocytes led to grossly different phenotypes. TwKOastro mice were significantly smaller in size than their control littermates already at the age of 3 months, and showed kyphosis and progressive weight loss (Fig. 1d, e), which was not due to reduced food intake (Supplementary Fig. 2a). At 5–6 months of age, the mice showed an abnormal symmetric hopping gait with compromised motor performance and weakness of hind limbs, with low pelvic posture, progressing to paraparesis (Fig. 1f, g). The animals had a limited lifespan of 7–8 months of age (Fig. 1h). The TwKOneuro mice showed no difference in body weight from littermates until ~8 months of age (Fig. 1e). They developed moderate kyphosis, but showed no motor defects and appeared wild-type-like, until rapid deterioration of general condition and weight loss a few days prior to death at the age of 7–9 months (Fig. 1d–h). The overall gross morphology and size of the TwKOastro and TwKOneuro brains was unremarkable (Fig. 1i). No specific pathology was found in spinal cord (Supplementary Fig. 3).
TwKOneuro, although asymptomatic at the age of 3–4 months, did demonstrate mtDNA depletion already at this age (Fig. 2a). TwKOastro manifested with body weight loss and brain pathology starting from the age of 2–3 months, but showed significant mtDNA depletion only prior to death at 8 months (Fig. 2a). To assess consequences of the two genetic manipulations, we chose to analyze TwKOastro mainly at 5–6 months of age, when the mice were already in advanced neurological disease, and neuronal-KO at 7–8 months, as only then they showed morphological findings and manifested with symptoms, and consequences to other cell types and brain pathology could be studied.
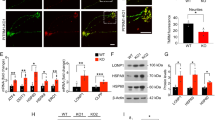
Mitochondrial consequences of Twinkle inactivation. a MtDNA amount in different ages in TwKO mice. b Western blot of OXPHOS complexes CI–CV and quantification of signals normalized to total protein. c NAD+, NADH, NADP+, NADPH amounts and their ratios in cortex lysates. The data are presented as mean and error bars indicate standard deviation. Statistical significance: *p < 0.05, **p < 0.01, ***p < 0.001
At terminal stage of TwKOneuro and TwKOastro, mtDNA amount showed reduction to 25 and 50% of controls’ mean, respectively (Fig. 2a). Consequently, in TwKOastro, mtDNA-encoded RC enzyme complex amounts (CI, III, IV) were reduced in the brain tissue, with also OXPHOS enzyme activities, measured by high-resolution respirometry (Fig. 2b; Supplementary Fig. 2b). Fully nuclear-encoded CII showed normal protein amount. TwKOneuro demonstrated a combined RC protein deficiency, with CI and CIV affected the most (~5% and 40% of the controls’ mean, respectively; CI contains seven mtDNA-encoded subunits; CIII one, CIV three and CV one; Fig. 2b). The severe CI deficiency in TwKOneuro reflects the major contribution of neuronal mtDNA to total CI amount in mouse brain. In brain of TwKOneuro mice also the overall redox status was altered with significantly reduced NAD/NADH ratio, contributed by CI deficiency, as the complex is a major acceptor of reducing equivalents from NADH. NAD is the precursor of both NADPH and NADP, and their levels were significantly decreased (Fig. 2c). As NAD and NADP form regulate major biosynthetic reactions in all cells, the effects of mtDNA depletion extend beyond RC, to the whole cellular metabolism.
Loss of mtDNA maintenance activates astrocytes
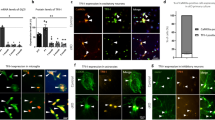
Embryonic lethality of the full TwKO predicted progressive death of the targeted neurons or astrocytes. Unexpectedly, however, mtDNA depletion in astrocytes caused a remarkable increase of GFAP-positive cells with reactive astrocyte morphology in the targeted brain regions (Fig. 3a, b). Such morphology is typical for disease-related reactive gliosis. Activation was progressive, starting at 2–3 months of age (absent at the age of 1 month, Supplementary Fig. 4b), peaking in the cortex at terminal stage. The TwKOneuro showed only modestly increased cortical astrogliosis at the terminal stage, with increased number of GFAP-positive cells of stellar morphology in cortex (Fig. 3a, b). Part of the population of reactive astrocytes in TwKOastro also co-expressed nestin, another marker of reactive gliosis. Nestin-positive cells are not found in the cortex of controls or TwKOneuro (Fig. 3c). The amount of GFAP+Nestin+ double-positive cells was progressive (<30% at the age of 3–4 months, and > 50% at 5–6 months). We then addressed the question, whether the remarkable number of activated astrocytes was due to their proliferation. In all three genotypes we found rare KI67-positive cells, marking cell division, but not co-localizing in the nuclei of the GFAP-positive cells (Supplementary Fig. 4c). Furthermore, most of the astrocytes in the mouse cortex are Aldhl1-positive, and in TwKOastro they showed slightly swollen morphology, but their number was not changed (Fig. 3d). These data imply that the gliosis is due to astrocytes activated on site, but not their proliferation.
MtDNA loss in astrocytes causes massive astrocyte activation. a, b Astrocytes (GFAP, red) in somatosensory cortex (a) and in hippocampus, CA1 area (b) of 7–8-month-old mice and 3–4 months as shown in inset of (a). In b, boxed area is enlarged on right. c Astrocytes (GFAP, red) immuno-co-stained with reactive astrocyte marker (Nestin, green). Insets enlarge the boxed area. Arrows show the same cells. d Astrocytic marker Aldh1l1 immunohistochemistry, counterstaining hematoxylin. TwKOastro show intensive vacuolization. Brain graphs show sampling site. Fluorescence figures are counterstained with DAPI (blue). Bar graphs on right hand side: corresponding quantifications of GFAP+ cells (cortex and hippocampus); GFAP+Nestin+/GFAP+, and Aldh1l1-positive cells in somatosensory cortex. The data are presented as mean and error bars indicate standard deviation. Statistical significance: *p < 0.05, **p < 0.01, ***p < 0.001. Scale bars: 20 μm (a), 50 μm (b–d)
To examine whether the activated astrocytes were RC deficient, we tested in situ the mitochondrial RC enzyme activities and mitochondrial morphology. Consistent with mtDNA depletion in astrocytes of TwKOastro mice, we found a large number of cells with astrocyte-like stellar morphology showing typical signs of RC deficiency: succinate dehydrogenase-positive (SDH; CII, nuclear-encoded) and cytochrome c oxidase-negative (COX; CIV, partially encoded by mtDNA)25 (Fig. 4a, Supplementary Fig. 5a). Also in TwKOneuro we found COX-negative cells of neuronal morphology at 5–6 months of age, consistent with neuronal mtDNA depletion (Fig. 4a, Supplementary Fig. 5a). The activated astrocytes in TwKOastro harbored accumulations of mitochondria, a sign of mitochondrial dysfunction, both perinuclearly and in cell processes (Fig. 4b, Supplementary Fig. 5b). GFAP-positive cells of controls or the reactive astrocytes of TwKOneuro mice showed no such accumulations (Fig. 4b, Supplementary Fig. 5b). These findings indicate that the targeted cell populations developed RC deficiency as a consequence of lack of mtDNA-encoded RC subunits. Furthermore, the mitochondrial accumulation in astrocytic processes, similar that has been previously found for example in Purkinje dendrites of Twinkle-mutant Deletor mice25, suggests dysfunctional trafficking or impaired turnover of the organelle. Electron microscopic examination of TwKOastro cortex revealed clusters of giant vacuolated mitochondria in cells with astrocytic morphology (Fig. 4c). In TwKOneuro, the neuronal mitochondria appeared swollen, but showed no vacuolization (Fig. 4c). The findings strongly suggest that (1) astrocyte functions are dependent on mitochondrial metabolism; (2) astrocyte activation is a cell-autonomous consequence of Twinkle inactivation and consequent mtDNA loss; (3) astrocyte activation caused by primary mtDNA depletion is different from secondary activation after neuronal mtDNA depletion, further pointing the importance of mtDNA maintenance for correct astrocyte functioning.
OXPHOS activity and mitochondrial structure in Twinkle KOs. a In situ histochemical activity analysis of mitochondrial respiratory chain enzymes in cortex. Brown: Complex IV, contains mtDNA-encoded subunits. Blue: fully nuclear-encoded Complex II. Arrows: respiratory chain deficient cells with astrocyte (TwKOastro) or neuronal (TwKOneuro) morphology. Inset in TwKOneuro: hippocampus, COX-deficient cells pointed with arrows. b Mitochondrial Complex II (SDHA, green) immuno-co-stained with astrocytes (GFAP, red), arrows point the same cells. TwKOastro astrocytes show punctate mitochondrial accumulations. c Ultrastructure of mitochondria (arrows and purple pseudocolor). Green = nuclei, yellow = neuronal projections. Brain graph shows sampling site. Scale bars: 50 μm (a, b), 5 μm (c)
MtDNA loss in astrocytes causes secondary neurodegeneration
We then asked whether chronic astrocyte activation in TwKOastro had secondary consequences for neurons. Twinkle inactivation in astrocytes resulted in significantly disorganized neuronal dendrite arborization in the hippocampal stratum radiatum, but the somata of pyramidal neurons appeared intact (Fig. 5a). To visualize the neuronal morphology in detail, we used Golgi staining of neurons. Indeed, dendrite complexity was dramatically decreased in both hippocampus and cortex of TwKOastro. The few and short dendrites carried dense varicosities (Fig. 5b), potentially as a consequence of deficient synaptic pruning. TwKOastro secondarily challenged viability of cortical neurons close to terminal stage (Fig. 5c, d), but the death was not occurring through caspase-3-dependent manner. However, occasional caspase-positive neurons/glia were present in the neuropil of TwKOastro mice (Fig. 5d). In the hippocampal CA1 region and cortex of TwKOneuro, Golgi-staining demonstrated short dendrites, decreased dendrite complexity and reduced neuronal body size, accompanied by abundant cell death, significantly more than in TwKOastro (Fig. 5b–d). These results indicate that the early-onset phenotype of TwKOastro mice is not explained by cell death. Instead, the chronic activation of the astrocytes challenges their functions in neuronal maintenance, leading to progressive changes in neuron structure and late-stage secondary neurodegeneration.
MtDNA loss in astrocytes or neurons challenges neuronal dendrite maintenance and causes neurodegeneration. a Immunofluorescent co-staining (overlay) of neuronal nuclei (NeuN, green), dendrites (microtubular MAP2, red) and nucleus (DAPI, blue) in hippocampus of 7–8 months old mice. b Neuronal fine morphology with Golgi staining in 7–8 months old mice. Arrows point gross morphological changes with dendritic degeneration in both KOs and varicosities in TwKOastro. c Dying neurons (arrows), Fluorojade-C (FJ-C) staining. Quantification below. d Apoptotic cells (arrows); immunohistochemical staining for cleaved caspase-3 and counterstaining with hematoxylin (parallel sections to FJ-C (c)). Quantification in somatosensory cortex and CA1 region of hippocampus below. Inset: enlarged caspase-3-positive cell. The data are presented as mean and error bars indicate standard deviation. Statistical significance: *p < 0.05, **p < 0.01, ***p < 0.001. Scale bars: 20 μm (a, b), 50 μm (c, d)
TwKOastro affects neuron subtypes and promotes inflammation
We then asked what types of cortical neurons were differentially affected by TwKO. The overall cellularity in the cortex was relatively well preserved in all mice (Fig. 6a). However, TwKOastro mice showed a nearly complete loss of GABA-ergic calbindin-positive interneurons in the somatosensory cortical layers I-II at an advanced disease stage (Fig. 6a, b). In TwKOneuro cortex, the calbindin-positive neurons remained relatively well preserved. As the cre-expression is prominent in these regions in both KO models, the TwKOneuro data suggest relative resistance of these cortical neurons to mtDNA loss, but high sensitivity of these cells to dysfunction of their astrocytic support.
MtDNA loss in astrocytes depletes Calbindin-positive interneurons in cortex, upper neuronal layers. Immunofluorescnece staining of mid-sagittal sections of 7–8 months old mice, brain graph shows sampling site. a Neuronal nuclei (NeuN, green), Calbindin-positive neurons (red) and nucleus (DAPI, blue). b Quantification of NeuN-and Calbindin-positive neurons in somatosensory cortex, cortical layers I and II are boxed. Scale bars: 20 μm. The data are presented as mean and error bars indicate standard deviation. Statistical significance: ***p < 0.001
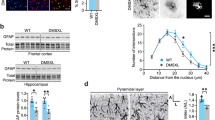
Neuronal damage typically promotes an inflammatory response in the form of microglial activation. IBA1-positive microglia in both KO models adopted a rounded morphology, typical for activated microglia, whereas the control mice microglia had quiescent ramified morphology with elongated, branched processes (Fig. 7a). The microglial activation in TwKOastro appeared earlier than in TwKOneuro, despite the moderate number of apoptotic cells in TwKOastro compared to TwKOneuro, suggesting a link between astrocyte and microglial activation. Starting from the age of 5–6 months, the TwKOastro mice had wide-spread chronic microglial and astrocyte activation accompanied by prominent myelin disorganization, while the number of Olig2-positive oligodendrocytes and total MBP content were unchanged (Fig. 7b, d). The axonal structure in the white matter of corpus callosum did not show signs of disruption (Fig. 7c). Myelin abnormalities were not present in TwKOneuro. These results indicate that chronic astrocyte activation and dysfunction promote inflammation and myelin changes.
Inactivation of Twinkle in astrocytes leads to microglial activation and demyelination. Immunofluorescent staining of mid-sagittal sections from 6 months old mice, brain graph shows sampling site. a Immunofluorescent staining of microglia (IBA1, red) and nucleus (DAPI, blue) in cortex. b Myelin content (MBP, green), oligodendrocytes (Olig2, red), and nuclei (DAPI, blue). c Pan-axonal staining of long axons in corpus callosum. d Quantification of Olig2-positive cells and length of processes of IBA1-positive cells in cortex; western blot of MBP, and its quantification. Insets: enlarged boxed area. Scale bars: 20 μm (a), 50 μm (b, c). The data are presented as mean and error bars indicate standard deviation. Statistical significance: *p < 0.05, ***p < 0.001
TwKO in astrocytes results in spongiotic encephalopathy
The most remarkable pathological consequence of astrocyte-specific TwKO was development of spongiotic vacuolization of the brain (Fig. 8a). The vacuoles appeared in a progressive manner, starting from cortex at 2 months of age (not present at 1 month of age, Supplementary Fig. 4a). At the terminal stage, we found a severe spongiotic encephalopathy with profound vacuolization of the cortex and to a lesser extent of hippocampus, cerebellum, and olfactory bulb (Fig. 8b). The vacuoles are often formed close to intact astrocytic nuclei, and activated astrocytes enwrap the vacuoles (Fig. 8c, d). Ultrastructural analysis revealed clusters of round and oval shaped vacuoles of varying sizes, ranging from 1 to 15 μm (Fig. 8e). We also found vacuolization of myelinated axons at terminal stage, indicating neuronal degeneration (Fig. 8f). Intriguingly, these changes of TwKOastro resemble closely the spongiotic pathology in infantile mitochondrial diseases, especially Leigh syndrome and Alpers–Huttenlocher syndrome, the latter caused by defects of mtDNA maintenance3,6,9,26.
Inactivation of Twinkle in astrocytes leads to brain vacuolization and spongiotic encephalopathy. a Cortical morphology with hematoxylin/eosin staining from terminal stage of TwKOastro and TwKOneuro mice. Representative images of the brain mid-sagittal sections. Extensive brain vacuolization in TwKOastro indicated with arrows. b Spongiotic encephalopathy of TwKOastro has progressive nature. Brain graphs show the affected regions with severity of pathology demonstrated with intensity of the color shading. c Vacuoles (blue arrows) and astrocyte nuclei (black arrows) in hematoxylin/eosin staining. d Immunohistochemistry of GFAP in TwKOastro and vacuoles enwrapped by an astrocyte (blue arrow). e Representative electron micrograph of vacuoles. Upper arrow indicates a vacuolated mitochondrion, lower arrow indicates group of vacuoles in the brain of TwKOastro. f Representative electron micrograph of vacuolated myelinated axon. Brain graph shows sampling site. Scale bars: 100 μm (a, b); 10 μm (c), 5 μm (d). g Time-course development of brain pathology in TwKOastro and TwKOneuro mice
Figure 8g summarizes the consequences of Twinkle inactivation in the two mouse models and the progression of different changes.
Discussion
Glial cells constitute at least half of the cells in human neocortex27. They are a metabolically active and dynamic cell type, but considered to be mainly glycolytic, compared to high oxidative reliance of neurons. We report here that mtDNA maintenance is of absolute importance for astrocyte function, as Twinkle loss and progressive mtDNA depletion in these cells causes an early-onset, severe neurological disease, whereas a similar defect in neurons results in late-manifesting rapidly progressive disease course. The manifestations of TwKOastro are among the most severe reported in any model targeting astrocytes in vivo28,29. Our findings are not consequences of a general decline of mitochondrial respiratory functions, because inactivation of COX assembly factor, COX10, does not lead to severe brain pathology19. Therefore, we claim that mtDNA helicase and replisome activity or mtDNA amount contribute to signaling that modulates astrocyte activity.
Surprisingly, mtDNA loss in TwKOastro induced a wide-spread chronic activation of astrocytes, followed by severe secondary changes in neuronal morphology and progressive spongiotic encephalopathy, with relative sparing of neurons. The findings closely resemble those in children with mitochondrial encephalopathies, Alpers–Huttenlocher and Leigh syndromes. The neuropathologic characteristics of these disorders include spongiotic encephalopathy with astrocyte activation—often described as glial scar—relative sparing of neurons and cortical laminar necrosis3,6,30,31. Such pathology is also typical for adult-onset mitochondrial spinocerebellar ataxia-neuropathy-epilepsy syndromes caused by mtDNA maintenance defects32,33. We propose that chronic activation and dysfunction of astrocytes, as a consequence of mtDNA maintenance defect, is a primary underlying cause of spongiotic mitochondrial encephalopathies of children.
What type of dysfunction could underlie the spongiosis? Normal astrocytes promote CNS homeostatic functions by pruning synapses, maintaining axons, providing metabolites and trophic factors for neurons, and phagocytosing myelin debris34. However, during a CNS insult, activation of astrocytes may both promote and hinder recovery: microglial activation was reported to induce neurotoxic transformation of astrocytes, to “A1” type astrocytes. These cells lose important homeostatic functions, which leads to poor maintenance of synapses, neurodegeneration, and myelin changes34,35,36. The neurotoxic factor remains still unknown. We find activated astrocytes enwrapping pathological vacuoles, and neuronal morphology and myelin showing major secondary changes, indicative of possible toxicity. As increasing microgliosis in TwKOastro follows astrocyte activation, it remains possible that the interaction of the two cell types contributes to pathology, with devastating secondary consequences to CNS.
Our evidence suggests that defective mtDNA maintenance in astrocytes drives their chronic activation, which promotes pathology of mitochondrial encephalopathy in mice. The findings offer an alternative mechanistic explanation for pathogenesis of the most common mitochondrial encephalopathies in infants. Furthermore, the results emphasize the potential importance of astrocyte dysfunction in all primary and secondary disorders with mtDNA maintenance defects, including dopaminergic neurons in Parkinson’s disease37. We propose that supporting of astrocytic mitochondrial function should be considered as an attractive target for intervention, to combat neurodegeneration.
Methods
Generation of knockout mice and phenotype testing
All the animal experiments were conducted according to relevant national and international guidelines, and approved by the Finnish Committee of Experimental Animal Research. The generation and validation of the loxP-mice is described elsewhere38. Cre-expressing mouse lines: [Tg(Pgk1-cre)1Lni, Camk2a-cre, GFAP(73.12)-cre and mT/mG; The Jackson Laboratory, Stock No: 2178050, 005359, 012886, 007576 ME, USA] were crossed to lox-P mice. Cre-mediated excision of the floxed fragment produced a frame-shift mutation leading to a premature stop codon. For staging, the day of vaginal plug was counted as embryonic day 0.5 (E0.5). For PGK-cre-mice, embryos were collected at E7.5, at the time of which all genotypes were present. The astro/neuro-TwKO offspring were born in Mendelian ratios. The mice were regularly followed up by inspection for activity, and weighted. Beam walk test, conducted as previously published39. Briefly, the mice were trained to navigate the 1 m poles of two different diameters (1 and 1.5 cm) for two constitutive days and recorded on the third day. We also used the grip strength tester (BioSEB, USA) with each mouse recoded three times with 5 min intervals.
Tissue collection, expression analysis
The animals were sacrificed with CO2 and decapitated. One brain hemisphere was snap-frozen in liquid nitrogen and stored at −80 °C, and the other was sliced and immersion fixed in 4% PFA for immunohistological analysis. Cortex DNA and RNA isolation was performed using NucleoSpin TriPrep kit (Macherey-Nagel). The deletion caused by cre-excision was analyzed from Twinkle cDNA by PCR with the following primers sites 5′-CGTTTTGAGGACCTGCCTCT-3′, 5′-TTGGACACCTGCAGATACCG-3′. Primers used for Twinkle quantification by quantitative PCR were following: Twinkle 5′-ACGAGCAGCTCTCCTCTGA-3′, 5′-TGCTGTCTGCAGTTCCTTGT-3′ normalized to β-2 microglobulin level (primers: 5′-CATGGCTCGCTCGGTGACC-3′, 5′-AATGTGAGGCGGGTGGAACTG-3′).
Histology, immunohistochemistry, and immunofluorescence
The animals were terminally anesthetized with Mebunat (Orion, Finland) and perfused intracardially with 4% paraformaldehyde in PBS or sacrificed with CO2. Brains were removed, post-fixed with 4% paraformaldehyde in PBS followed by embedding in paraffin. All histological and immunoassays were carried out on 7–10 μm sagittal sections. Immunohistochemical and immunofluorescence stainings were described previously40 and performed on a minimum of n = 3 of each genotype. COX/SDH activity analysis is described elsewhere38. Images were acquired with either Zeiss AxioImager epifluorescent microscope or Zeiss LSM 780 confocal microscope. Primary antibodies were rabbit Calbindin (Abcam), GFAP (Millipore), ALDH1L1 (Abcam), cleaved Caspase 3 (Cell Signaling Technology), IBA1 (Wako), Olig2 (Merck Millipore), KI67 (Merck Millipore); mouse Nestin (Merck Millipore), SDHA (CII) (Abcam), NeuN (Chemion), GAD67 (Merck Millipore); Neurofilament Marker pan axonal cocktail (BioLegend); chicken MAP2 (Thermo Scientific); rat MBP (Nordic BioSite). Golgi staining was processed using the FD Rapid GolgiStain Kit (FD Neuro Technologies). Fluoro-Jade C was from Millipore. Morphometrical analysis of coronal 80-μm-thick sections was described in ref. 41. For cell counting, images from mid-sagittal sections were taken from three to six mice per genotype.
Electron microscopy
Ultrastructural analysis was performed as described elsewhere42. Briefly, tissue samples were fixed in phosphate buffer with 2.5% glutaraldehyde, post-fixed with 1% osmium tetroxide, dehydrated through ascending concentrations of ethanol and embedded in Epon-resin. Ultrathin sections of 60 nm were cut with Leica ultramicrotome and post stained with uranylacetate and lead citrate. Sections were imaged on a Phillips CM12 electron microscope operated at 80 kV.
Western blot
Protein from frozen tissue was extracted in 150 mM NaCl 50 mM Tris buffer with 1% Triton-X, pH 7.6. Proteins were separated by SDS-PAGE using 4–20% gradient gels and transferred to PVDF membranes. Blots were blocked with 5% milk in Tris-buffered saline with Tween 20 (TBST) buffer for 1 h and antibodies were incubated at +4 °C overnight. Bio-Rad TGS stain-free technology was used for quantification of the total protein signal. Images were acquired with ChemiDoc Imager and quantified with ImageLab software. Original blots are shown in Supplementary Fig. 6.
NAD+, NADH, NADP+, and NADPH measurement
Pyridine nucleotide concentrations in tissues were measured using cycling assay similar to previous publication43.
High-resolution respirometry
Oxygen consumption rates were measured using a high-resolution oxygraph (OROBOROS instruments using a substrate-uncoupler-inhibitor protocol as described elsewhere44). Specific oxygen consumption rates are expressed as pmol/(s*mg). Briefly, half of 6 months old cerebellum was homogenized with 6–8 strokes in a 2 ml potter-Elvehjem in respiration buffer (0.5 mM EGTA, 3 mM MgCl2, 60 mM Lactobionic acid, 20 mM Taurine, 10 mM KH2PO4, 20 mM HEPES, 110 mM d-sucrose, 1% fat-free BSA). Oxygen consumption rates were performed in the presence of pyruvate-glutamate-malate and activities determined with +ADP (CI), +succinate (CI + CII), maximal uncoupled respiration by FCCP titration, and CIV by ascorbate + TMPD.
MtDNA copy number analysis
Analysis of mtDNA amount was done by quantitative PCR, as described elsewhere45. Briefly, level of mtDNA (primers: 5′-AGGAGCCTGTTCTATAATCGATAAA-3′, 5′-GATGGCGGTATATAGGCCGAA-3′) was normalized to a nuclear-encoded single-copy gene RBM15 (primers: 5′-GGACAGTTTTCTTGGGCAAC-3′, 5′-AGTTTGGCCCTGTGAGACAT -3′).
Statistical methods
For statistical significance analysis, we used unpaired Student’s t test or one-way ANOVA followed by Bonferroni tests (P ≤ 0.05).
Data availability
Authors can confirm that all relevant data are included in the paper and/or its Supplementary Information files.
References
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012).
Chinnery P. F. in Gene Reviews (eds Pagon, R. A. A. M. et al.) Ch. Mitochondrial Disorders Overview (University of Washington, Seattle, USA, 2014).
Lake, N. J., Bird, M. J., Isohanni, P. & Paetau, A. Leigh syndrome: neuropathology and pathogenesis. J. Neuropathol. Exp. Neurol. 74, 482–492 (2015).
Suomalainen, A. & Isohanni, P. Mitochondrial DNA depletion syndromes--many genes, common mechanisms. Neuromuscul. Disord. 20, 429–437 (2010).
Jellinger, K. & Seitelberger, F. Spongy glio-neuronal dystrophy in infancy and childhood. Acta Neuropathol. 16, 125–140 (1970).
Leigh, D. Subacute necrotizing encephalomyelopathy in an infant. J. Neurol. Neurosurg. Psychiatry 14, 216–221 (1951).
Alpers, B. J. Diffuse progressive degeneration of the gray matter of the cerebrum. Arch. Neurol. Psychiatry 25, 469–505 (1931).
Naviaux, R. K. et al. Mitochondrial DNA polymerase gamma deficiency and mtDNA depletion in a child with Alpers’ syndrome. Ann. Neurol. 45, 54–58 (1999).
Hakonen, A. H. et al. Recessive Twinkle mutations in early onset encephalopathy with mtDNA depletion. Brain 130, 3032–3040 (2007).
Korhonen, J. A., Pham, X. H., Pellegrini, M. & Falkenberg, M. Reconstitution of a minimal mtDNA replisome in vitro. EMBO J. 23, 2423–2429 (2004).
Larsson, N. G. et al. Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat. Genet. 18, 231–236 (1998).
Milenkovic, D. et al. TWINKLE is an essential mitochondrial helicase required for synthesis of nascent D-loop strands and complete mtDNA replication. Hum. Mol. Genet. 22, 1983–1993 (2013).
Hance, N., Ekstrand, M. I. & Trifunovic, A. Mitochondrial DNA polymerase gamma is essential for mammalian embryogenesis. Hum. Mol. Genet. 14, 1775–1783 (2005).
Sorensen, L. et al. Late-onset corticohippocampal neurodepletion attributable to catastrophic failure of oxidative phosphorylation in MILON mice. J. Neurosci. 21, 8082–8090 (2001).
Ekstrand, M. I. et al. Progressive parkinsonism in mice with respiratory-chain-deficient dopamine neurons. Proc. Natl Acad. Sci. USA 104, 1325–1330 (2007).
Pickrell, A. M., Fukui, H. & Moraes, C. T. The role of cytochrome c oxidase deficiency in ROS and amyloid plaque formation. J. Bioenerg. Biomembr. 41, 453–456 (2009).
Sofroniew, M. V. & Vinters, H. V. Astrocytes: biology and pathology. Acta Neuropathol. 119, 7–35 (2010).
Pauwels, P. J., Opperdoes, F. R. & Trouet, A. Effects of antimycin, glucose deprivation, and serum on cultures of neurons, astrocytes, and neuroblastoma cells. J. Neurochem. 44, 143–148 (1985).
Supplie, L. M. et al. Respiration-deficient astrocytes survive as glycolytic cells in vivo. J. Neurosci. 37, 4231–4242 (2017).
Lallemand, Y., Luria, V., Haffner-Krausz, R. & Lonai, P. Maternally expressed PGK-Cre transgene as a tool for early and uniform activation of the Cre site-specific recombinase. Transgenic Res. 7, 105–112 (1998).
Garcia, A. D., Doan, N. B., Imura, T., Bush, T. G. & Sofroniew, M. V. GFAP-expressing progenitors are the principal source of constitutive neurogenesis in adult mouse forebrain. Nat. Neurosci. 7, 1233–1241 (2004).
Herrmann, J. E. et al. STAT3 is a critical regulator of astrogliosis and scar formation after spinal cord injury. J. Neurosci. 28, 7231–7243 (2008).
Tao, J. et al. Deletion of astroglial Dicer causes non-cell-autonomous neuronal dysfunction and degeneration. J. Neurosci. 31, 8306–8319 (2011).
Gross, N. J., Getz, G. S. & Rabinowitz, M. Apparent turnover of mitochondrial deoxyribonucleic acid and mitochondrial phospholipids in the tissues of the rat. J. Biol. Chem. 244, 1552–1562 (1969).
Tyynismaa, H. et al. Mutant mitochondrial helicase Twinkle causes multiple mtDNA deletions and a late-onset mitochondrial disease in mice. Proc. Natl Acad. Sci. USA 102, 17687–17692 (2005).
Naviaux, R. K. & Nguyen, K. V. POLG mutations associated with Alpers’ syndrome and mitochondrial DNA depletion. Ann. Neurol. 55, 706–712 (2004).
Azevedo, F. A. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Colucci-Guyon, E. et al. Mice lacking vimentin develop and reproduce without an obvious phenotype. Cell 79, 679–694 (1994).
Pekny, M. et al. Mice lacking glial fibrillary acidic protein display astrocytes devoid of intermediate filaments but develop and reproduce normally. EMBO J. 14, 1590–1598 (1995).
Harding, B. N. et al. Progressive neuronal degeneration of childhood with liver disease (Alpers’ disease) presenting in young adults. J. Neurol. Neurosurg. Psychiatry 58, 320–325 (1995).
Sparaco, M., Bonilla, E., DiMauro, S. & Powers, J. M. Neuropathology of mitochondrial encephalomyopathies due to mitochondrial DNA defects. J. Neuropathol. Exp. Neurol. 52, 1–10 (1993).
Hakonen, A. H. et al. Mitochondrial DNA polymerase W748S mutation: a common cause of autosomal recessive ataxia with ancient European origin. Am. J. Hum. Genet. 77, 430–441 (2005).
Lax, N. Z. et al. Cerebellar ataxia in patients with mitochondrial DNA disease: a molecular clinicopathological study. J. Neuropathol. Exp. Neurol. 71, 148–161 (2012).
Liddelow, S. & Barres, B. SnapShot: astrocytes in health and disease. Cell 162, 1170–1170.e1171 (2015).
Liddelow, S. A. & Barres, B. A. Reactive astrocytes: production, function, and therapeutic potential. Immunity 46, 957–967 (2017).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Bender, A. et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat. Genet. 38, 515–517 (2006).
Nikkanen, J. et al. Mitochondrial DNA replication defects disturb cellular dNTP pools and remodel one-carbon metabolism. Cell Metab. 23, 635–648 (2016).
Luong, T. N., Carlisle, H. J., Southwell, A & Patterson, P. H. Assessment of motor balance and coordination in mice using the balance beam. J. Vis. Exp. 49, 2376 (2011).
Chilov, D. et al. Phosphorylated Î2-catenin localizes to centrosomes of neuronal progenitors and is required for cell polarity and neurogenesis in developing midbrain. Dev. Biol. 357, 259–268 (2011).
Vashchinkina, E., Panhelainen, A., Aitta-Aho, T. & Korpi, E. R. GABAA receptor drugs and neuronal plasticity in reward and aversion: focus on the ventral tegmental area. Front. Pharmacol. 5, 256 (2014).
Khan, N. A. et al. Effective treatment of mitochondrial myopathy by nicotinamide riboside, a vitamin B3. EMBO Mol. Med. 6, 721–731 (2014).
Matsumura, H. & Miyachi, S. Cycling assay for nicotinamide adenine dinucleotides. Methods Enzymol. 69, 465–470 (1980).
Jackson, C. B. et al. Mutations in lead to autosomal recessive encephalomyopathy and isolated mitochondrial complex II deficiency. Am. J. Med. Genet. 51, 170–175 (2014).
Ahola-Erkkilä, S. et al. Ketogenic diet slows down mitochondrial myopathy progression in mice. Hum. Mol. Gen. 19, 1974–1984 (2010).
Acknowledgements
We thank Markus Innilä, Anu Harju, Babette Hollmann, and Tuula Manninen for experimental support; the Electron Microscopy Unit of the Institute of Biotechnology, Helsinki University, for expertise in ultrastructural analysis; Dr. Fang Zhao, Advanced Microscopy Unit, Medicum, Helsinki University for consultation and data discussion. Helena Schmidt for artwork, Biomedicum Imaging Unit facility imaging expertise and Helsinki University Animal Center for professional care of animals.
We wish to acknowledge the following funding sources: Academy of Finland, Sigrid Jusélius Foundation (A.S., E.d.M.), University of Helsinki, European Research Council Advanced Grants (A.S.), Swiss National Foundation postdoc grant and the Novartis Foundation for Medical-Biological Research (C.B.J.), and Helsinki Doctoral Program for Biomedicine (O.I.).
Author information
Authors and Affiliations
Contributions
O.I., D.C., and A.S. contributed to the study design; O.I., D.C., I.P., E.d.M., C.B.J., G.C., M.T., and L.E. performed experiments, A.P. provided expertise for brain pathology analysis; O.I., D.C., and A.S. wrote the manuscript. All authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ignatenko, O., Chilov, D., Paetau, I. et al. Loss of mtDNA activates astrocytes and leads to spongiotic encephalopathy. Nat Commun 9, 70 (2018). https://doi.org/10.1038/s41467-017-01859-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-017-01859-9
This article is cited by
-
Loss of fatty acid degradation by astrocytic mitochondria triggers neuroinflammation and neurodegeneration
Nature Metabolism (2023)
-
The Relationships Among Metal Homeostasis, Mitochondria, and Locus Coeruleus in Psychiatric and Neurodegenerative Disorders: Potential Pathogenetic Mechanism and Therapeutic Implications
Cellular and Molecular Neurobiology (2023)
-
Inhibition of mitochondrial function by metformin increases glucose uptake, glycolysis and GDF-15 release from intestinal cells
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.