Abstract
We have previously reported the molecular signature of murine pathogenic TH17 cells that induce experimental autoimmune encephalomyelitis (EAE) in animals. Here we show that human peripheral blood IFN-γ+IL-17+ (TH1/17) and IFN-γ−IL-17+ (TH17) CD4+ T cells display distinct transcriptional profiles in high-throughput transcription analyses. Compared to TH17 cells, TH1/17 cells have gene signatures with marked similarity to mouse pathogenic TH17 cells. Assessing 15 representative signature genes in patients with multiple sclerosis, we find that TH1/17 cells have elevated expression of CXCR3 and reduced expression of IFNG, CCL3, CLL4, GZMB, and IL10 compared to healthy controls. Moreover, higher expression of IL10 in TH17 cells is found in clinically stable vs. active patients. Our results define the molecular signature of human pro-inflammatory TH17 cells, which can be used to both identify pathogenic TH17 cells and to measure the effect of treatment on TH17 cells in human autoimmune diseases.
Similar content being viewed by others
Introduction
TH17 cells are a subset of interleukin-17 (IL-17)-secreting T-helper (TH) cells implicated in the pathogenesis of multiple sclerosis (MS), rheumatoid arthritis, juvenile idiopathic arthritis (JIA), and psoriasis1,2, whose differentiation is regulated by the transcription factor RAR-related orphan nuclear receptor gamma (RORγt)3. Initially, TH17 cells were considered a uniformly pro-inflammatory population driven by IL-23 and expressed a unique pattern of pro-inflammatory cytokines different from TH1 and TH2 cells4,5,6. Subsequent studies showed the function of TH17 cells in autoimmune diseases and defense against bacterial and fungal pathogens7,8,9,10. TH17 cells can be induced to produce TH1 and TH2 cytokines11 and not all TH17 cells are pathogenic. Murine TH17 cells are pathogenic or non-pathogenic based on their ability to induce experimental autoimmune encephalomyelitis (EAE)12; pathogenic TH17 cells express higher levels of IFN-γ while non-pathogenic TH17 cells produce IL-10 with IL-1713.
As in mice, human TH17 cells can also co-produce IFN-γ or IL-10. IL-10-producing TH17 cells are induced in response to Staphylococcus aureus, whereas TH17 cells induced by Candida albicans produce IL-17 and IFN-γ. Both types of TH17 cells are enriched in a subset of human memory CD45RA–CD4+ TH cells expressing the chemokine receptors CCR6 and CCR4, while IFN-γ-secreting TH17 (TH1/17) cells may additionally express CXCR39,14. A deficiency in IL-17 or the TH17 pathway compromises host defenses against S. aureus and C. albicans, and reduces the frequency of circulating CCR6+ memory CD4+ TH cells15,16. Thus both IFN-γ and IL-10-producing TH17 cells may be protective during infection.
IFN-γ and IL-10-producing TH17 cells are considered pro-inflammatory and anti-inflammatory, respectively, and have opposite functions in autoimmunity17,18. Studies of T-cell libraries from patients with MS showed that CCR6+ myelin-reactive T cells exhibit enhanced production of IFN-γ, IL-17, and GM-CSF and reduced production of IL-10, when compared with those from healthy individuals19. In JIA, IFN-γ-secreting TH1/17 cells are highly enriched in the synovial fluid (SF) of inflamed joints20. The inflammatory environment in diseased joints can induce IFN-γ-negative TH17 cells to co-produce IFN-γ, implicating plasticity of TH17 cells21. The proportions of TH1/17-enriched CD4+CD161+ T cells in the SF of affected joints correlate with the erythrocyte sedimentation rate and serum levels of C-reactive protein, suggesting these cells function in disease pathogenicity20. These and other reports of elevated numbers of TH1/17 cells in inflamed tissues in human autoimmune diseases21,22,23 associate TH1/17 cells with human autoimmune diseases.
The complexity of TH17 function is further manifested in therapeutic studies. Although anti-IL-17 therapy benefits psoriasis, blocking the IL-17 pathway in Crohn’s disease is either ineffective or exacerbates diseases24,25,26. Similarly, in the CD45RBhi adoptive transfer mouse model of experimental colitis, a deficiency of IL-17 production or IL-17R expression in transferred CD45RBhi CD4 T cells results in accelerated disease27. The protective function of IL-17 in these studies may be due to the fact that TH17 cells that line the gut mucosa prevent invasion of the gut microbiome and promote intestinal homeostasis28. In tumors, TH17 cells are reported to have both beneficial29,30,31 and detrimental effects32 both in animal models and human disease. Hence, the function of TH17 cells in diverse immune responses is complex.
We previously reported that murine TH17 cells can be differentiated into pathogenic vs. non-pathogenic subsets, as well as characterized the molecular signature of murine pathogenic TH17 cells through global gene expression analysis12. In the present study, we compare the gene expression profiles between human IFN-γ+ and IFN-γ– TH17 subsets, and between IL-10+ and IL-10– TH17 clones. Comparative transcriptomic analyses show that human TH1/17 cells and IL-10– TH17 clones display gene signatures with marked similarities to mouse pathogenic TH17 cells. We then assess TH1/17 cells in patients with MS and find reduced expression of anti-inflammatory IL10 and elevated expression of CXCR3. When we compare clinically active vs. stable patients, we find that stable patients have higher IL10 expression in TH17 cells, whereas active patients have higher expression of STAT3 in IFN-γ–/IL-17– CD4+ T cells. Our results define the molecular signature of human pro-inflammatory TH17 cells, which can be used to both identify pathogenic TH17 cells and to measure the effect of treatment on TH17 cells.
Results
Transcriptionally distinct TH17 subsets in peripheral blood
We first performed intracellular cytokine staining of blood CD4+ T cells and identified distinct populations of IFN-γ co-producing TH17 cells, but no IL-10 co-producing TH17 cells (Fig. 1a; Supplementary Fig. 1). It is known that IFN-γ+ TH17 cells are increased in inflamed tissues in human autoimmune diseases21,22,23, and are also present in the blood of healthy individuals, whereas IL-10+ TH17 cells are barely detected14. We divided peripheral TH17 cells into IFN-γ+ (TH1/17) and IFN-γ– (TH17) subsets. We utilized a capture assay that separates live CD4+ T subsets based on differential secretion of IL-17 and/or IFN-γ to sort ex vivo TH1/17 cells and TH17 cells without in vitro polarization and with only short-term (3 h) Phorbol 12-myristate 13-acetate (PMA) plus ionomycin stimulation (Fig. 1b; Supplementary Fig. 2). Based on our global transcriptional analysis of murine TH17 cells and studies on autoimmunity from ours and other groups, we designed a nanoString nCounter CodeSet HuTH17 that detects 418 genes associated with human TH cell differentiation and activation. The HuTH17 CodeSet encompasses genes encoding transcription factors, cytokines, cell surface markers, kinases, lytic proteins, and housekeeping proteins (Supplementary Data 1). We used this CodeSet to generate high-throughput transcription profiles of isolated ex vivo TH1/17, TH17, TH1, and double negative (DN) CD4+ T cells from five healthy donors to generate high-throughput transcription profiles. We found high expression of IL17A in TH17 and TH1/17 cells and high expression of IFNG in TH1 and TH1/17 cells, whereas only minimal expression of IL17A was observed in TH1 and DN cells and minimal expression of IFNG was observed in TH17 and DN cells (Fig. 1c), thus demonstrating that we isolated pure populations of TH1/17, TH17, and TH1 cells. IL10 gene expression was detected in both TH17 and TH1/17 cells (Fig. 1d).
Transcriptionally distinct human TH17 subsets in peripheral blood. a IFN-γ and IL-10 expression in human TH17 cells. Isolated PBMCs were stimulated with PMA and ionomycin for 4 h. Production of indicated cytokines in CD4+ T cells were assessed by flow cytometry with intracellular cytokine staining assay. Dotplots shown were gated on CD4+ lymphocytes. Data are representative of two independent experiments with similar results. b Isolation of live TH1/17, TH17, TH1, and DN cells from human PBMC for nCounter analysis. CD4+ T cells isolated from the peripheral blood of healthy donors were stimulated with PMA and ionomycin for 3 h. CD3+CD4+-TH1/17 (IFN-γ+IL-17+), TH17 (IFN-γ−IL-17+), TH1 (IFN-γ+IL-17−), and DN (IFN-γ−IL-17−) cells were sorted after being stained with fluorescence-conjugated anti-CD3 and CD4 in combination with cytokine secretion detection kits (Miltenyi) (n = 5). b Isolated CD4+ T subsets were stimulated and stained as in a. c–f CD4+ T-cell subsets treated as in a were measured using the nCounter (nanoString Technologies) CodeSet HuTH17 and subsequently analyzed (hereafter abbreviated as nCounter analysis). c Differential expression analysis of mRNA levels of IL17A and IFNG. *p < 0.05, **p < 0.005, ***p < 0.0005, One-way ANOVA with Tukey’s multiple comparison test (mean ± s.d.). d Differential expression analysis of mRNA levels of IL10. Two tailed, paired Student’s t test p-value was shown (mean ± s.d.). The 326 out of the 418 measured genes in the HuTH17 CodeSet that showed unsupervised variation across the sample population were used for e hierarchical clustering of the individual samples (individual donors: A, B, C, D, and E), f hierarchical clustering of the Pearson’s linear correlations between the samples (individuals n = 5; individual donors: A, B, C, D, and E), and g principal component analysis of the samples (individuals n = 5)
For the 20 CD4+ T-cell subset samples from the five healthy individuals we analyzed, 362 of the 418 genes demonstrated unbiased variation across the population, defined as an unsupervised expression difference (difference between maximum and minimum relative gene expression values, not taking into account information about the subset classes) ≥5 across all samples, and these genes were selected for further study. Hierarchical clustering of gene expression profiles of the 20 individual samples from the CD4+ subpopulations in the context of these 362 unsupervised varying genes properly segregated the TH1/17, TH17, TH1, and DN cells into four different clusters, revealing their distinct transcriptional features (Fig. 1e). The in-group Pearson correlation values for these gene expression profiles were high for all four CD4+ T-cell subsets and ranged from 0.95 ± 0.06 for DN cells to 0.98 ± 0.02 for TH1 cells. Pearson correlation values for the gene expression profiles of TH1/17 vs. TH17 and TH1/17 vs. TH1 were 0.83 ± 0.05 and 0.78 ± 0.06, respectively, whereas the correlation coefficient for TH17 vs. TH1 was only 0.43 ± 0.06 (Fig. 1f), which were consistent with the degree of similarities observed among the cell subsets in the hierarchical clustering analysis (Fig. 1e). Principal component analysis (PCA) showed that DN cells were clearly distinct from the other three subsets, whereas TH1/17 cells lay in between TH17 and TH1 cells (Fig. 1g). These results demonstrate that human TH17 cells can be transcriptionally categorized into IFN-γ+ TH17 (TH1/17) and IFN-γ– TH17 (TH17) subsets and that TH1/17 show a close relationship to both TH17 and TH1 cells.
Human ex vivo TH17 subsets vs. murine TH17 cells
Murine TH17 cells generated with TGF-β3/IL-6 or IL-1/IL-6/IL-23 induce more severe EAE than TH17 cells generated with TGF-β1/IL-612. In comparing TGF-β3 vs. TGF-β1-induced murine TH17 cells, we defined a pathogenic transcriptional signature composed of 16 upregulated genes in TGF-β3-induced pathogenic TH17 cells including CCL3, CCL4, CLL5, CSF2, IL22, IL3, GZMB, STAT4, and TBX21 12. In humans, 10 of the 16 signature genes were upregulated in CCR6+ myelin-reactive memory CD4+ T cells in patients with MS although these cells also displayed elevated levels of IL10 19, while 12 genes were upregulated in TH1/17-enriched CCR7loCCR6+CCR4loCXCR3hi memory CD4+ T cells in healthy donors compared to TH17-enriched CCR7loCCR6+CCR4hiCXCR3lo cells33 (Supplementary Data 2). There are no known unique surface markers for TH17 cells and all of memory CD4+ T-cell populations contain not only IFN-γ+ and IFN-γ– TH17 cells but also IL-17–IFN-γ+ TH1 cells, especially for CCR7loCCR6+CCR4loCXCR3hi memory CD4+ T cells, for which the frequency of IL-17–IFN-γ+ cells may be up to four times higher than IL-17+ cells33. Among the 14 signature genes shared by murine TH17 cells and human TH17-enriched CD4+ T cells, 13 are included in the HuTH17 CodeSet (Supplementary Data 2) and we assessed their expression in TH1/17, TH17, TH1, and DN CD4+ T cells (Fig. 2a). We found that except for LRMP, the other 12 genes were detected by the CodeSet. Compared to TH1/17 cells, 10 of the 12 genes had comparable or higher messenger RNA (mRNA) levels in TH1 cells, which included CCL3, CCL4, CCL5, GZMB, ICOS, IL3, IL7R, LAG3, STAT4, and TBX21, emphasizing the importance of minimizing TH1 contamination when analyzing the gene signature of pro-inflammatory or pathogenic TH17 cells. The high purity of the TH1/17 and TH17 cells we isolated based on IL-17 and IFN-γ secretion allowed us to determine the pathogenicity-associated molecular signature of human TH17 subsets.
Gene expression comparison between human TH1/17 vs. TH17 cells and mouse pathogenic vs. non-pathogenic TH17 cells. a The expression of previously reported murine and human TH17 signature genes in purified ex vivo TH1 cells. The mRNA gene expression levels in TH1/17, TH17, TH1, and DN cells were measured as described in Fig. 1. *p < 0.05, repeated measures one-way ANOVA; **p < 0.05, pairwised group comparison with Tukey’s multiple comparison test (mean ± s.d., n = 5). b, c Gene set enrichment analysis comparing human TH1/17 vs. TH17 cells with mouse pathogenic vs. non-pathogenic TH17 cells. b Heatmap of upregulated (upper panels) and downregulated (lower panels) “leading edge” genes of comparison Scenario 1: human TH1/17 vs. TH17 cells vs. mouse TGF-β3 plus IL-6-induced TH17 cells vs. TGF-β1 plus IL-6-induced TH17 cells (Kolmogorov–Smirnov test p < 0.0001; FDR q < 0.0001 for upregulated genes; Kolmogorov–Smirnov test p = 0.0007; FDR q = 0.001 for downregulated genes). c Heatmap of upregulated (upper panels) and downregulated (lower panels) “leading edge” genes of comparison Scenario 2: human TH1/17 vs. TH17 cells vs. mouse IL-1, IL-23 plus IL-6-induced TH17 cells vs. TGF-β1 plus IL-6-induced TH17 cells (Kolmogorov–Smirnov test p < 0.0001; FDR q < 0.0001 for upregulated genes; Kolmogorov–Smirnov test p = 0.004; FDR q = 0.004 for downregulated genes). Each column represents one donor in human (n = 5) or one sample in murine (n = 4). d The robust predicted pathogenic signature (PreP-Signature) of human TH1/17 cells. Signature genes are those identified as differentially expressed between human TH1/17 and TH17 cells that are identified as enriched “leading edge” genes when assessing these human genes in the mouse profiles in b, c and that are additionally curated for robustness based on supervised absolute fold change >1.5. Two tailed, paired Student’s t test p-value < 0.05. *p < 0.05, **p < 0.05, ***p < 0.0005, n = 5
We identified 60 genes differentially expressed in TH1/17 cells vs. TH17 cells with 39 upregulated and 21 downregulated (Supplementary Data 3). To test the hypothesis that the relationship of TH1/17 to TH17 cells in humans is similar to that of mouse pathogenic vs. non-pathogenic TH17 cells, we performed gene set enrichment analysis (GSEA)34. The upregulated and downregulated genes in TH1/17 cells relative to TH17 cells were divided into two gene sets. Since both TGF-β3/IL-6 and IL-1/IL-6/IL-23 induced mouse pathogenic TH17 cells whereas TGF-β1/IL-6 induced non-pathogenic TH17 cells12, our first comparison (Scenario I) explored microarray data from TGF-β3/IL-6-induced TH17 cells (pathogenic) vs. TGF-β1/IL-6-induced TH17 cells (non-pathogenic) and the second comparison (Scenario II) explored microarray data from IL-1/IL-6/IL-23-induced TH17 cells (pathogenic) vs. TGF-β1/IL-6-induced TH17 cells (non-pathogenic). GSEA results demonstrated that genes upregulated in human TH1/17 vs. TH17 cells were enriched in mouse pathogenic TH17 cells vs. non-pathogenic TH17 cells in both scenarios with 17 “leading edge” genes in Scenario 1 and 19 genes in Scenario 2 (Kolmogorov–Smirnov test p < 0.0001; false discovery rate (FDR) q < 0.0001 for both scenarios) (upper panels of Fig. 2b, c). The “leading edge” subset of genes is defined as genes that drive the enrichment scores, thus the genes that appear in the top of the ranked list of genes at, or before, the point where the running sum reaches the maximum deviation from zero. The “leading edge” subset can be interpreted as the core subset of a gene set that accounts for the enrichment signal34. Genes upregulated in human TH17 vs. TH1/17 cells were also enriched in mouse non-pathogenic TH17 cells vs. pathogenic TH17 cells in both comparison scenarios, with seven identical “leading edge” genes (Kolmogorov–Smirnov test p = 0.0007; FDR q = 0.001 for Scenario I; Kolmogorov–Smirnov test p = 0.004; FDR q = 0.004 for Scenario II) (lower panels of Fig. 2b, c). The gene signatures obtained from both scenarios were almost identical even though the TGF-β3/IL-6-induced TH17 cells and IL-1/IL-6/IL-23-induced TH17 cells were differentiated via treating naive CD4+ T cells with different cytokines, suggesting that the integrated gene expression analysis identified common features shared by the two types of mouse pathogenic TH17 cells. Thus, we found marked similarities in differential gene expression signatures between human IFN-γ+ vs. IFN-γ– TH17 cells and mouse pathogenic vs. non-pathogenic TH17 cells indicating that human TH1/17 and TH17 cells are counterparts of murine pathogenic and non-pathogenic TH17 cells.
Among the 27 signature genes identified in both comparison scenarios, 13 had an absolute fold change >1.5 for TH1/17 vs. TH17 cells, which were selected as the robust predicted pathogenic signature (PreP-Signature) of TH1/17 cells for later analyses (Fig. 2d). The upregulated robust PreP-Signature genes with pro-inflammatory/pathogenic potential can be grouped into chemokines and cytokines (CCL3, CCL4, CCL5, CSF2, IFNG, and IL3), chemokine and cytokine receptors (CXCR3 and IL23R), cytokine responding genes (EPSTI1), effector proteins (GZMB), and signaling molecules (RGS2). The downregulated genes are the chemokine receptor (CCR6) and the transcription factor (STAT1).
We also investigated genes not shared between human and mouse pathogenic/non-pathogenic Th17 cells to identify other potentially relevant genes to test in human conditions. We identified 33 non-shared genes including 19 that were upregulated and 14 that were downregulated (Supplementary Data 4).
Human ex vivo TH17 subsets vs. TH17 clones
IL-10 has a pivotal role in regulation of both innate and adaptive immunity35 and murine TGF-β1/IL-6-induced non-pathogenic TH17 cells produce IL-1013. Human IL-10+ TH17 cells are a potential counterpart of mouse non-pathogenic TH17 cells. However, IL-10-secreting TH17 cells cannot be directly isolated from human blood for nCounter gene expression analysis because few or no TH17 cells produce IL-10 after PMA/ionomycin stimulation (Fig. 1a). This is likely due to the delayed production of IL-10 after stimulation. Nonetheless, for established human TH17 clones, IL-10+ TH17 clones can be identified by IL-10 intracellular staining 5 days after T-cell receptor activation9. Thus, we established CD4+ T cells clones from CCR6+CCR4+CXCR3– memory CD4+ T cells (Supplementary Fig. 3) that were enriched for TH17 cells14,18 and screened for clones producing IL-17 with or without co-secretion of IL-10. qRT-PCR showed that IL-10– TH17 clones expressed high levels of IFNG, low levels of IL10, and high levels of IL23R, whereas IL-10+ TH17 clones expressed minimal levels of IFNG and high levels of IL10 (Fig. 3a). To assess whether the gene expression profile of cloned TH17 cells maintained their pro or anti-inflammatory features after long-term culture, we used the HuTH17 CodeSet to analyze the gene expression profile of IL-10– and IL-10+ TH17 clones. We identified 63 genes that were differentially expressed between IL-10– vs. IL-10+ TH17 clones with 49 upregulated and 14 downregulated genes (Supplementary Data 5). To compare genes differentially expressed between human TH1/17 vs. TH17 cells (Supplementary Data 3) to those between IL-10– vs. IL-10+ TH17 clones (Supplementary Data 5), we assessed enrichment of overlaps between gene lists using the hypergeometric enrichment test. We found that genes upregulated in TH1/17 vs. TH17 cells display significant overlap with genes upregulated in IL-10– vs. IL-10+ TH17 clones (one-sided Fisher’s exact test p < 0.0001, FDR q = 0.0001) (Fig. 3b). The upregulation of CBLB, CLL5, CXCR3, IL23R, REL, TBX21, and TNFSF14 in IL-10– TH17 clones is shared by human TH1/17 cells and mouse pathogenic TH17 cells (Fig. 2b, c). To identify regulatory molecules predicted to influence the differentiation/development of TH1/17 cells and IL-10– TH17 clones, we interrogated genes differentially expressed between TH1/17 vs. TH17 cells (Supplementary Data 3) and between IL-10– vs. IL-10+ TH17 clones (Supplementary Data 5) for upstream regulator prediction analysis in ingenuity pathway analysis (IPA). TH1/17 cells and IL-10– TH17 clones displayed a similar pattern of the activation of signaling pathways involved in TH17 differentiation/development (Fig. 3c), especially for activation of IL-1β signaling, which is critical to promote co-producing IFN-γ and to inhibit IL-10 production in TH17 cells9. These results demonstrate the similarities between TH1/17 vs. TH17 cells and IL-10– vs. IL-10+ TH17 clones, and also indicate that IL-10– and IL-10+ TH17 clones maintained their pro- or anti-inflammatory characteristics after long-term culture.
Gene expression comparison between human TH1/17 vs. TH17 cells and IL-10– vs. IL-10+ TH17 clones. a Quantitative RT-PCR analysis of gene expression in human IL-10– and IL-10+ TH17 clones isolated from healthy donors (two tailed, paired Student’s t test, mean ± s.d., n = 3). For IFNG, IL17A, and IL23R, resting TH17 clones were stimulated with anti-CD3 and anti-CD28 for 4 h before RNA extraction. For IL10, resting TH17 clones were stimulated with anti-CD3 and anti-CD28 for 5 days, then cells were re-stimulated with anti-CD3 and anti-CD28 for 4 h before RNA extraction. b Hypergeometric enrichment test between human TH1/17 vs. TH17 cells and IL-10– vs. IL-10+ TH17 clones. Genes differentially expressed between human TH1/17 and TH17 cells (Supplementary Data 3) were analyzed for enrichment in those of human IL-10– vs. IL-10+ TH17 clones (Supplementary Data 5). Heatmap shows the overlapping genes (one-sided Fisher’s exact test p < 0.0001, FDR q = 0.0001). Each column represents one donor (n = 5). c Predicted upstream regulators for TH1/17 and IL-10– TH17 clone differentiation. The differentially expressed genes with corresponding fold changes and p-values from the TH1/17 vs. TH17 comparison (Supplementary Data 3) and IL-10– vs. IL-10+ TH17 clone comparison (Supplementary Data 5) were analyzed using the IPA upstream regulator analysis. Ex vivo, TH1/17 vs. TH17 comparison; Clone, IL-10– vs. IL-10+ TH17 clone comparison. d, e Gene set enrichment analysis comparing human IL-10– vs. IL-10+ TH17 clones with mouse pathogenic vs. non-pathogenic TH17 cells. d Heatmap of upregulated (upper panels) and downregulated (lower panels) “leading edge” genes of comparison Scenario 1: human IL-10– vs. IL-10+ TH17 clones vs. mouse TGF-β3 plus IL-6-induced TH17 cells vs. TGF-β1 plus IL-6-induced TH17 cells (Kolmogorov–Smirnov test p = 0.004; FDR q = 0.005 for upregulated genes; Kolmogorov–Smirnov test p = 0.004; FDR q = 0.003 for downregulated genes). e Heatmap of upregulated (upper panels) and downregulated (lower panels) ‘‘leading edge” genes of comparison Scenario 2: human IL-10– vs. IL-10+ TH17 clones vs. mouse IL-1, IL-23 plus IL-6-induced TH17 cells vs. TGF-β1 plus IL-6-induced TH17 cells (Kolmogorov–Smirnov test p = 0.005; FDR q = 0.010 for upregulated genes; Kolmogorov–Smirnov test p = 0.016; FDR q = 0.009 for downregulated genes). Each column represents one donor in human (n = 5) or one sample in murine (n = 4)
Human TH17 clones vs. murine TH17 cells
We performed GSEA to investigate the similarity between human TH17 clones and mouse TH17 cells. The upregulated and downregulated genes in IL-10– TH17 clones relative to IL-10+ TH17 clones TH17 cells were divided into two gene sets (Supplementary Data 5) to explore the enrichment of the gene sets in Scenarios I and II (defined above). GSEA showed that genes upregulated in IL-10– vs. IL-10+ TH17 clones were enriched in mouse pathogenic vs. non-pathogenic TH17 cells in both scenarios, with 17 “leading edge” genes in Scenario I (Kolmogorov–Smirnov test p = 0.004; FDR q = 0.005) and 14 “leading edge” genes in Scenario II (Kolmogorov–Smirnov test p = 0.005; FDR q = 0.010). Among the “leading edge” genes, 11 overlapped between both scenarios (upper panels of Fig. 3d, e). Genes upregulated in human IL-10+ vs. IL-10– TH17 clones were also enriched in mouse non-pathogenic vs. pathogenic TH17 cells in both scenarios, with six “leading edge” genes in Scenario I (Kolmogorov–Smirnov test p = 0.004; FDR q = 0.003) and seven “leading edge” genes in Scenario II (Kolmogorov–Smirnov test p = 0.016; FDR q = 0.009). Among the “leading edge” genes, six genes were represented in both scenarios (lower panels of Fig. 3d, e). These results demonstrate extensive similarities between human IL-10– vs. IL-10+ TH17 clones and mouse pathogenic vs. non-pathogenic TH17 cells.
TH17 PreP-signatures predict STAT3 as an upstream regulator
Integrated analysis of gene expression profiles of human ex vivo TH17 cells, human TH17 clones, and cytokine-induced mouse TH17 cells resulted in four sets of gene signatures associated with the pathogenicity of TH17 cells (Figs. 2, 3). We merged the “leading edge” genes from the above GSEA comparisons and consolidated these four gene signatures into one complete PreP-Signature for human ex vivo TH17 cells with 27 signature genes and one complete PreP-Signature for human TH17 clones with 26 signature genes. Among the signature genes upregulated in human TH1/17 cells and IL-10– TH17 clones, seven genes were shared by both types of TH17 cells, while for downregulated genes, two genes overlapped between TH1/17 cells and IL-10– TH17 clones (Fig. 4a). Since our molecular signatures derive from integrated analysis of pro-inflammatory human TH17 cells with mouse pathogenic TH17 cells in autoimmunity, they reduce the number of potential targeting genes and also help define human TH17 cells with potential pathogenicity in autoimmunity.
Predicting STAT3 as upstream transcription factor by the PreP-Signatures of TH17 cells. a Venn diagram representations of signature genes upregulated (left) and downregulated (right) in human TH1/17 cells and IL-10– TH17 clones. Complete PreP-Signatures from GSEA comparison Scenarios I and II were merged for ex vivo cells (Fig. 2b, c) and TH17 clones (Fig. 3d, e), respectively. Genes with supervised absolute fold change >1.5 in either ex vivo cells or TH17 clones were shown in italic bold letters. Ex vivo, differentially expressed “leading edge” genes from TH1/17 vs. TH17 GSEA comparisons presented in green circles; Clone, differentially expressed “leading edge” genes from IL-10– vs. IL-10+ TH17 clone GSEA comparisons presented in yellow circles. b Predicted upstream transcription factors for TH1/17 and IL-10– TH17 clone differentiation. The molecular signatures of TH1/17 cells and IL-10– TH17 clones in a were analyzed using the Enrichr ChEA2016 analysis and the predicted transcription factors with Benjamini–Hochberg adjusted p-value <0.05 were shown. TF transcription factor
We interrogated the complete PreP-Signatures of ex vivo TH1/17 cells and IL-10– TH17 clones for upstream transcription factor prediction analysis using the Enrichr ChEA2016 analysis36,37. STAT3 in CD4+ T cells was the top predicted transcription factor common for both resulting lists (Fig. 4b; Supplementary Datas 6, 7) and thus may regulate the pathogenicity of human TH17 cells in autoimmune diseases.
Investigation of PreP-Signature genes in multiple sclerosis
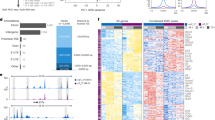
To investigate PreP-Signature genes in human pro-inflammatory TH17 cells in MS, we isolated TH1/17 and TH17 cells from untreated patients with relapse-remitting MS (RRMS) and age and sex-matched healthy controls (Table 1; Supplementary Fig. 2), assessed frequencies of TH1/17 and TH17 cells, and measured the expression of the PreP-Signature genes. TH1 and DN cells were isolated in parallel as internal controls. Of 19 patients with RRMS, sufficient RNA for quantitative PCR (qPCR) analysis was obtained from 15 TH1/17 samples, 18 TH17 samples, 19 TH1 samples, and 19 DN samples. Of 16 healthy controls, we obtained sufficient RNA for qPCR analysis from 16 TH1/17 samples, 14 TH17 samples, 16 TH1 samples, and 16 DN samples. We found no difference in frequency of TH1/17, TH17, and total TH17 cells among total CD4+ T cells in patients vs. controls (Fig. 5a). The percentage of TH1/17 in total TH17 cells was also similar (Fig. 5b). To assess purity of isolated cell subsets, we measured the expression of IL17 and IFNG by qPCR. We detected high levels of IL17 in TH1/17 and TH17 cells but not in DN cells (Supplementary Fig. 4a), and high levels of IFNG in TH1/17 cells but not in TH17 and DN cells (Supplementary Fig. 4b), demonstrating that the isolated populations were of high purity. In both groups, IL17 expression was higher in TH17 vs. TH1/17 cells (Supplementary Fig. 4a), whereas IL17 expression was similar for TH1/17 and TH17 cells (Fig. 5c). To validate the 13 robust PreP-Signature genes identified via the nCounter analysis, we measured their expression in TH1/17 and TH17 cells in healthy controls by qPCR (Fig. 5d, left). For the 11 genes upregulated in TH1/17 cells, 9 were confirmed; the 2 downregulated genes were not validated. In MS, the nine validated genes displayed the similar upregulated expression pattern (Fig. 5d, right). Thus, the transcriptional regulation of the nine validated robust PreP-Signature genes of TH1/17 cells was tightly associated with IFN-γ secretion in both controls and MS. These results indicate that studying frequency or number of IFN-γ-secreting TH17 cells may not identify important biological differences.
Frequency of TH17 subsets and differential expression of PreP-Signature genes in TH1/17 vs. TH17 cells in MS. Peripheral CD4+ T cells isolated from the PBMC of untreated RRMS patients (n = 19) and age- and sex-matched healthy controls (HC) (n = 16) were stimulated, stained, and sorted for TH1/17, TH17, TH1, and DN cells as described in Fig. 2b. RNA isolated from sorted cell subsets was subjected to low-input qPCR analysis. a Frequencies of TH1/17, TH17, and total TH17 cells in total peripheral CD4+ T cells (Welch’s t test, p-values, mean ± s.d.). b Frequency of TH1/17 in total TH17 cells (Welch’s t test, mean ± s.d.). c, d qPCR analysis of gene expression in isolated CD4+ T-cell subsets. c Comparison of IL17A expression between HC and patients in TH1/17 or TH17 cells (Welch’s t test, mean ± s.d.). d Differential expression of PreP-Signature genes between TH1/17 vs. TH17 cells in HC and MS patients. Two tailed, paired Student’s t test, *p < 0.05, **p < 0.001, ***p < 0.0001
We investigated differences between MS and control using the robust PreP signature genes we identified above. We first measured the expression of the 13 robust PreP-Signature genes in TH1/17 cells in MS and found elevated expression of CXCR3 and reduced expression of IFNG, CCL3, CLL4, and GZMB (Fig. 6a). In TH17 cells, GZMB showed reduced expression in MS (Fig. 6b). No difference was observed between MS and controls in TH1 cells (Supplementary Fig. 5). Thus, the altered expression of these five PreP-Signature genes are specific for TH1/17 and TH17 subsets, but not for TH1 cells. We then measured the expression of TBX21 and IL10, since we found downregulation of TBX21 and upregulation of IL10 in IL-10+ TH17 clones (Fig. 3a, d). We found TBX21 expression elevated in TH1/17 relative to TH17 cells in MS and controls (Fig. 6c, upper panel), with no difference in IL10 expression (Fig. 6d, upper panel). However, when we compared TH1/17 and TH17 cells between MS and controls, IL10 was reduced in TH1/17 and TH17 cells in patients (Fig. 6d, lower panel), though no difference was detected with TBX21 expression (Fig. 6c, lower panel). The predicted upstream transcription factor STAT3 was elevated in the DN cells in MS (Fig. 6e, upper panel), while the expression of its antagonistic transcription factor STAT5A 38,39,40 was reduced (Fig. 6e, lower panel).
Expression of PreP-signature genes of human pro-inflammatory TH17 cells in RRMS. RNA isolated from TH1/17, TH17, and DN cells (Fig. 5) was subjected to qPCR analysis. Comparison of PreP-Signature gene expression between HC and patients with RRMS in a TH1/17 cells and b TH17 cells. (Welch’s t test, mean ± s.d.). Comparison of c TBX21 and d IL10 expression between HC and patients with RRMS in TH1/17 and TH17 cells. e Comparison of STAT3 (upper panel) and STAT5A (lower panel) expression between HC and patients with RRMS in TH1/17, TH17, and DN cells. Welch’s t test p-values were shown (mean ± s.d.)
Thus, the altered expression of five PreP-Signature genes for TH1/17 cells in MS (CXCR3, IFNG, CCL3, CLL4, and GZMB) is TH17-specific. The low expression of IL10 in CXCR3hi TH1/17 cells in MS suggests these pro-inflammatory cells may more readily migrate to central nervous system (CNS) since CXCL10 (IP-10), a ligand for CXCR3, is increased in the inflamed CNS in MS41,42,43,44. It has been suggested that STAT3 facilitates TH17 differentiation and STAT5 facilitates Treg differentiation38,39,40, which is consistent with our observation of their expression in DN cells in MS.
We then investigated whether the PreP-Signature genes, which had altered expression in MS, were linked to disease activity in MS. Disease activity was defined as a gadolinium-enhancing lesion on magnetic resonance imaging (MRI) or disease relapse within 6 months to the time of blood sampling. All patients were untreated at the time of blood sampling. From 12 active patients, we obtained sufficient RNA for qPCR from 8 TH1/17 samples, 12 TH17 samples, and 12 DN samples. From seven stable patients, we obtained sufficient RNA for qPCR from 7 TH1/17 samples, 6 TH17 samples, and 7 DN samples. We found reduced expression of IL10 in TH17 cells (primarily in TH1/17 cells) and increased expression of STAT3 in DN cells in active patients with RRMS (Fig. 7). Given the facilitating role of STAT3 in TH17 differentiation and the anti-inflammatory function of IL-10, these results together with the altered expression of CXCR3, IFNG, CCL3, CLL4, GZMB, IL10, STAT3, and STAT5A in MS (Fig. 6) are consistent with an important role for TH17 cells in MS.
Correlation of IL10 and STAT3 expression with disease activity in MS. The mRNA levels of signature genes with altered expression in MS (Fig. 6) were compared between active and stable patients. Signature gene expression in a TH1/17 (active, n = 8; stable, n = 7), b TH17 (active, n = 12; stable, n = 6), and c DN cells (active, n = 12; stable, n = 7) (Welch’s t test, mean ± s.d.)
Discussion
We found human IL-17-secreting CD4+ helper T cells were distinct from TH1 cells, and could be transcriptionally segregated into IFN-γ+ TH17 (TH1/17) and IFN-γ– TH17 (TH17) subsets. On a transcriptional level, TH1/17 cells have features of both TH17 and TH1 cells. In addition to IFN-γ, TH1/17 cells expressed elevated levels of the pro-inflammatory molecules CCL3, CCL4, CCL5, granzyme B, IL-3, IL-22, GM-CSF, STAT1, T-bet, and IL-23R, which are pathogenic signature genes of mouse TH17 cells12. These results not only transcriptionally segregate human TH1/17 cells from other TH17 cells, but also indicate that these cells have acquired pro-inflammatory properties similar to murine pathogenic TH17 cells. This provided the foundation for the cross-species comparative transcriptomic analysis between mouse and human.
We compared the gene expression profiles of human TH1/17 cells vs. TH17 cells with mouse pathogenic vs. non-pathogenic TH17 cells via GSEA and found human TH1/17 vs. TH17 cells displayed gene expression signatures that were enriched in mouse pathogenic vs. non-pathogenic TH17 cells. We obtained similar results in comparing human IL-10– vs. IL-10+ TH17 clones with and mouse pathogenic vs. non-pathogenic TH17 cells. Through this integrated analysis, we identified gene expression features to support the hypothesis that TH1/17 cells are the pathogenic TH17 population in immune-mediated human disease.
The PreP-Signature genes derived from comparative transcriptional analysis of human and mouse studies identify genes shared between human TH1/17 vs. TH17 cells or between human IL-10– vs. IL-10+ TH17 clones and mouse pathogenic vs. non-pathogenic TH17 cells. We obtained a robust PreP-Signature for TH1/17 cells of 13 genes and assessed them in MS. We find that TH1/17 cells in patients with MS have elevated expression of CXCR3 and reduced expression of IL10. It has been shown that CXCL10 (IP-10), a ligand for CXCR3, is increased in the inflamed CNS of MS41,42,43,44. Given that IL-10 is a potent anti-inflammatory cytokine, this combination may facilitate migration of more pro-inflammatory CXCR3hiIL-10low TH1/17 cells to the inflamed CNS. However, it is unexpected to observe reduced expression of signature genes IFNG, CCL3, CLL4, and GZMB in TH1/17 cells in MS vs. healthy controls, especially the reduced expression of IFNG. Natalizumab, a humanized monoclonal antibody targeting α4 integrin used in the treatment of MS, functions by preventing immune cells from crossing the blood–brain barrier45. A study has shown that both natalizumab treated and untreated patients during relapse have lower frequencies of TH17 cells in peripheral blood compared to stable patients. Moreover, TH17 cells become almost undetectable in patients with breakthrough disease that occurs following natalizumab withdrawal46. Thus, one explanation for reduced expression of IFNG, CCL3, CLL4, and GZMB in TH1/17 cells in MS could be that migration of cells to the CNS removes CXCR3 high, active TH1/17 cells from the blood. Another possibility may be that expression of IFN-γ ensures high expression of CXCR3, which is critical for cells to migrate to inflamed CNS, and reduced IFN-γ may indicate the further enhancement of the pathogenicity of IL-10lo TH17 cells in MS. IFN-γ-stimulation is required for CXCR3 induction on T cells upon T-cell receptor stimulation47. Although counter-intuitive, the potent pro-inflammatory TH1 cytokine IFN-γ is protective during EAE induction48,49,50. It suppresses EAE induction by inhibiting generation of TH17 cells51, converting CD4+CD25– T cells to CD4+ Tregs52 and limiting myelin lipid peroxidation in CNS53. IFN-γ is dispensable for generation of pathogenic TH17 cells, however, T-bet the transcription factor for TH1 was initially considered essential due to the high resistance of Tbet−/− mice to EAE54,55. Later studies show that T-bet is essential for TH1 but not TH17-mediated EAE56,57. Studies in Tbet−/− mice have shown that reduced IFN-γ in TH17 cells does not affect their pathogenicity though the conversion of TH17 cells to TH1/17 cells as well as TH1-like IFN-γ + ex-TH17 cells is prevented56,58,59.
Interrogating the PreP-Signatures for the identification of upstream regulators and transcription factors36,37, we identified STAT3 as the top predicted upstream transcription factor from previous studies in CD4+ T cells, which suggests that STAT3 may regulate the pathogenesis of human TH17 cells in autoimmune diseases. It has been reported that STAT3 is a critical regulator for the induction of TH17 cells in humans. Humans with a genetic defect in STAT3 expression not only have reduced expression of TH17 without any impact on other T-cell subsets, but also develop a hyper-IgE syndrome with severe infections of C. albicans and S. aureus 60,61. STAT3 and STAT5 are a pair of mutual restraint transcription factors that regulate TH17 and Treg differentiation with STAT3 facilitates TH17 differentiation and STAT5 facilitates Treg differentiation38,39,40. We thus investigated the expression of STAT3 and STAT5A in MS. We found upregulation of STAT3 and downregulation of STAT5A in DN cells from patients with MS, which included naive and other memory CD4+ T cells but not TH1/17, TH17, and TH1 cells. One explanation for the different behavior of STAT3 and STAT5A between DN vs. TH1/17 and TH17 cells is that STAT3 may be critical for TH17 differentiation but not required for TH17-associated immunopathology after differentiation.
Leveraging comparative transcriptomic approaches, we integrated gene expression profiles derived from human and mouse TH17 cells to identify pathogenicity-associated signature (PreP-Signature) genes that are shared by human pro-inflammatory TH17 cells and mouse pathogenic TH17 cells, and to predict the upstream regulators or transcription factors that may be critical for the differentiation of human pro-inflammatory TH17 cells. These comparative transcriptomic analyses allowed us to identify altered gene expression associated with TH17 subsets and their differentiation in subjects with MS and to identify associations of the expression of STAT3 in DN cells and IL10 in TH17 cells that are dependent on MS disease activity. Of note, we used a nCounter codeset with a limited set of pre-selected 418 genes, which may miss other disease related, highly discriminant genes in humans. Thus, follow-up studies, such as RNA-sequencing analysis on TH17 and TH1/17 subsets isolated from MS patients, may help to identify more disease-related genes.
In summary, our study demonstrates that human TH1/17 cells and IL-10− TH17 clones display significant similarities to mouse pathogenic TH17 cells in their transcriptomic patterns. The elevated expression of pro-inflammatory cytokines and chemokines in human IFN-γ-secreting TH17 cells and the similarity in gene expression profiles between human IFN-γ-secreting TH17 cells and mouse pathogenic TH17 cells indicates a higher pro-inflammatory capacity of human TH1/17 cells and additionally provides transcriptional evidence to support the role of human TH1/17 in the pathogenesis of human autoimmune diseases. The differential molecular signature of human IFN-γ-secreting TH17 cells that we identified provides a new tool that can be utilized to assess TH17 cells both under physiologic conditions and in association with disease.
Methods
Reagents
EasySep human CD4+ T-cell enrichment kit (catalog number 19052) for CD4+ T-cell isolation was purchased from StemCell Technologies. FITC-conjugated anti-human IFN-γ (clone, B27; 1:100), Alexa 647-conjugated anti-human IL-17A (clone, N49-653; 1:20), PE-conjugated anti-human IL-10 (clone, JES3-19F; 1:660), and their corresponding isotype control antibodies for intracellular cytokine staining assay were purchased from BD Biosciences. IFN-γ cytokine secretion detection kit (APC) (catalog number 130-090-762) and IL-17 cytokine secretion detection kit (PE) (catalog number 130-094-537) were purchased from Miltenyi Biotec. nCounter CodeSet HuTH17 was custom designed and manufactured by nanoString Technologies. Fluorescence-conjugated antibodies for cell surface staining for flow cytometry were purchased from Biolegend. RNAqeous micro total RNA isolation kit (catalog number AM1931), SuperScript VILO master mix (catalog number 11755050), TaqMan preAmp master mix (catalog number 4391128), TaqMan fast universal PCR master mix (2x) (catalog number 4352042), and qPCR primers (Supplementary Data 8) were purchased from ThermoFisher Scientific.
Human subjects
Blood samples for TH17 cloning from healthy donors were obtained from the Swiss Blood Donation Center of Basel and Lugano. Informed, written consent was obtained from all donors. All uses of human material were approved by the Federal Office of Public Health (authorization no. A000197/2 to F.S.). Blood samples from MS patients and healthy controls were obtained from the Partners MS Center at Brigham and Women’s Hospital under IRB Protocol 2001P001431 and 2014P000124. Informed, written consent was obtained from all donors. MS patients were untreated for a minimum of 6 months before sampling. Disease activity was identified as a gadolinium-enhancing lesion on MRI or disease relapse within 6 months of sampling. Age- and sex-matched healthy donors did not have history of autoimmune diseases or malignancies and no acute or chronic infections. The samples from healthy donors for nCounter gene expression analysis were fresh blood samples. The MS samples and age- and sex-matched healthy control samples were frozen peripheral blood mononuclear cells (PBMCs).
Intracellular cytokine staining
For intracellular cytokine staining for PBMC or CD4+ T cells, assays were carried out with staining buffers and antibodies from BD Biosciences. Briefly, cells were seeded into a 96-well plate (up to 1 × 106 cell per well) and stimulated with PMA (100 ng/ml) and ionomycin (1 μg/ml) in the presence of GolgiStop for 4 h. After stimulation, cells were fixed with BD Cytofix fixation buffer and washed with BD Perm/Wash buffer. Cells in each well were equally divided into two wells, with one for intracellular cytokine staining and the other for isotype control staining. The following fluorophore-conjugated antibodies from BD Biosciences were used for staining analysis or as isotype controls: anti-CD4-pacific blue (clone: RPA-T4; 1:330), anti-IL-17A-Alexa647 (clone: N49-653; 1:20), anti-IFN-γ-FITC (clone: B27: 1:100), anti-IL-10-PE (clone: JES3-19F: 1:660), mouse IgG1-Alexa647 (clone: MOPC-21; 1:40), mouse IgG1-FITC (clone: MOPC-21; 1:100), and rat IgG2a-PE (clone: R35-95: 1:660). Stained cells were analyzed with a BD LSR II cytometer. Cytokine secretion in CD4+ lymphocytes was accessed with FlowJo.
Isolation of viable TH subsets from human PBMC
PBMC isolated with Ficoll-Pague PLUS (GE Healthcare) gradient centrifugation from the peripheral blood of healthy donors. Total CD4+ T cells were purified with the EasySep human CD4+ T-cell enrichment kit (StemCell Technologies). CD4+ T cells seeded in a 96-well plate (1 × 106 cells/well) were stimulated with PMA (30 ng/ml) and ionomycin (1 μg/ml) for 3–4 h (3 h for fresh blood samples and 4 h for frozen PBMC samples). Viable TH1/17, TH17, TH1, and DN cells were sorted with a FACSAria (BD Biosciences) after being stained with IFN-γ and IL-17 cytokine secretion detection kits (Miltenyi Biotec) and fluorescence-conjugated anti-CD3 and anti-CD4 antibodies following the manufacturer’s suggested protocol.
Isolation of human IL-10− and IL-10+ TH17 clones
PBMC were isolated by Ficoll-Paque PLUS (GE Healthcare) separation. CD4+ T cells were isolated from PBMC by positive selection using CD4 magnetic microbeads (Miltenyi Biotec). CCR6+CCR4+CXCR3–CD45RA–CD25–CD8–CD14–CD19–CD56– (enriched in TH17 cells) memory CD4 T cells were sorted with a FACSAria (BD Biosciences) and seeded at 0.6 cells per well in a 384 well plate. The following antibodies were used for FACS-based sorting: anti-CD45RA-Qdot655 (Life Technologies; clone: MEM-56; 1:1000); anti-CCR7-BV421 (Biolegend; clone: G043H7; 1:80); anti-CCR6-PE (BD Biosciences; clone: 11A9; 1:80) or anti-CCR6-BV605 (Biolegend; clone: G034E3; 1:60); anti-CCR4-PECy7 (BD Biosciences; clone: 1G1; 1:100); anti-CXCR3-PE-Cy5 or anti-CXCR3-APC (BD Biosciences; clone: 1C6; 1:20); anti-CD8-FITC or anti-CD8-PE-Cy5 (Beckman Coulter; clone: B9.11; 1:25); anti-CD25-FITC or anti-CD25-PE-Cy5 (Beckman Coulter; clone: B1.49.9; 1:25); anti-CD14-FITC or anti-CD14-PE-Cy5 (Beckman Coulter; clone: RMO52; 1:25); anti-CD19-FITC (BD Biosciences, clone: HIB19), or anti-CD19-PE-Cy5 (Beckman Coulter; clone: J3-119; 1:25); CD56-PE-Cy5 (Beckman Coulter; clone: N901; 1:25). CD4 T-cell clones were established in the presence of irradiated (45 Gy) allogeneic PBMC (25,000 cells per well) and phytohaemagglutinin (PHA-L) (1 μg/ml) in medium supplemented with IL-2 (500 U/ml). A small portion of cells of each CD4+ T-cell clone was used for intracellular cytokine staining assay for IL-17A (anti-IL-17A-eF660; eBioscience, catalog number 50-7179-42 clone: eBio64DEC17; 1:200), IL-10 (anti-IL-10-PE; BD Biosciences, catalog number 559330; clone: JES3-19F1; 1:20), and IFN-γ (anti-IFN-γ-FITC; BD Biosciences, catalog number 554700; clone: B27; 1:800) production. IL-10– and IL-10+ TH17 clones were selected for further analysis.
Regular quantitative real-time PCR
To measure gene expression in TH17 clones with qPCR, IL-10– and IL-10+ TH17 clones from an individual donor were pooled. They were stimulated or not with plate-bound anti-CD3 (clone TR66, 5 μg/ml) and anti-CD28 (clone CD28.2, 1 μg/ml) (αCD3/CD28) for 4 h, or stimulated with αCD3/CD28 for 5 days followed with or without a second αCD3/CD28 treatment for 4 h. Pooled TH17 clone cells were subjected to RNA isolation. Total RNA was extracted with the E.Z.N.A. Total RNA kit I (Omega Bio-tek, product number R6834). qPCR analysis was run and analyzed with the ViiA 7 Real-Time PCR System (Life Technologies). Quantitative comparison between IL-10– and IL-10+ TH17 clones was calculated using comparative ΔC T. Gene expression was normalized to the expression of β2m.
Low-input quantitative real-time PCR
To measure gene expression in TH1/17, TH17, TH1, and DN cells sorted from frozen PBMC with qPCR, total RNA was isolated and digested with DNase I with the RNAqeous micro total RNA isolation kit. cDNA was synthesized with the SuperScript VILO master mix and pre-amplified for 14 cycles with the TaqMan preAmp master mix following the manufacture’s instruction. qPCR analysis was run and analyzed with the ViiA 7 Real-Time PCR System (Life Technologies) using the TaqMan fast universal PCR master mix (2x) and qPCR primers purchased from ThermoFisher Scientific. The comparative threshold cycle method and an internal control (β2m) were used for normalization of the target genes. Relative expression was calculated as: ΔC T = C Tgene of interest – C Tβ2m; ΔΔC T = ΔC T cell subset of interest – mean of ΔC T DN of healthy control; the relative change of gene expression between the expression level of sample of interest and the mean expression level of all DN samples in healthy controls was given by this formula: (2 –ΔΔCT) × 10. All qPCR reactions were performed in duplicate.
nCounter analysis of mRNA expression
We designed a nanoString CodeSet HuTH17 that constitutes a 418-gene expression detection panel specific for human T-cell activation and differentiation (Supplementary Data 1). Cell lysates were prepared from sorted TH1/17, TH17, TH1, and DN cells with the RLT-Plus buffer of the RNeasy Plus Mini Kit (Qiagen, catalog number 74134) and gene expression levels were generated using the CodeSet HuTH17 according to the protocol provided by the manufacturer (NanoString Technologies). For gene expression in IL-10− and IL-10+ TH17 clones, total RNA was used for nCounter analysis following the manufacturer’s suggested protocol (NanoString Technologies).
Data analysis
nCounter gene expression data were normalized for code count using the geometric mean, for background using the mean, and for sample content using the geometric mean of housekeeping genes (isolated human CD4+ T-cell subsets: B2M, RPL3, and beta actin; human TH17 clones: B2M, GAPDH, and beta actin) with the R 3.2.0. NanoStringNorm package. Mouse gene expression data were downloaded from GEO (GSE39820) and normalized using RMA62 and ComBat63 in GenePattern (http://www.broadinstitute.org/cancer/software/genepattern/) as previously published12. Genes for which multiple probes were measured on the mouse microarray were collapsed to unique genes by selecting the probe with the highest average expression across all samples. Hierarchical clustering of the human samples and of the pairwise Pearson linear correlations were done based on the 362 genes that varied across the population in an unbiased manner (unsupervised expression difference (Δ = maximum expression value – minimum expression value)≥5, defined as the difference between maximum and minimum relative gene expression values across the population without considering the subset classes) using Pearson linear correlation and average linkage in GENE-E. PCA of the isolated human CD4+ T-cell subsets was done in R (prcomp) based on the same 362 genes. We selected differentially expressed genes between TH1/17 and TH17 cells using the two tailed, paired Student’s t test followed by supervised filtering for expression differences between mean TH1/17 and mean TH17 (Δ = MEANTH1/17−MEANTH17)>20 for robustness. The same approach was used to select differentially expressed genes between IL-10− and IL-10+ TH17 clones. GSEA was done in GenePattern using default settings (weighted scoring scheme, Signal2Noise metric, 1000 permutations)34,64,65 to test the enrichment of human signatures in the mouse expression profiles. Gene set overlap statistic analysis was done using the hypergeometric test in R (phyper, with N equal to the number of genes measured on the NanoString CodeSet). Storey’s q-value is used to control the FDR. Visualization of the gene expression heatmaps was done in GENE-E [http://www.broadinstitute.org/cancer/software/GENE-E/]. Il23r was not included in our microarray chip for mouse TH17 cell analysis, but we found upregulation of Il23r in murine pathogenic TH17 cells by qPCR12. Thus, we included Il23r in the murine gene expression profiles. The IPA upstream regulator analysis was performed for the differentially expressed genes (using corresponding fold changes and p-values) to identify key upstream regulatory molecules. Upstream regulators with z-scores >2 and z-scores <−2 represent activator and inhibitor mechanisms, respectively. The Enrichr ChEA2016 database and analytic tools were used to predict key upstream transcription factors36,37.
Statistical analysis
Statistical analysis was performed with Prism 7 (GraphPad Software), R statistical software (version 3.2.0), and Excel version 14.4.7. One-way analysis of variance with Tukey’s multiple comparison test was performed to compare the differential gene expression among TH subsets within HCs or MS patients when more than two cell subsets were included, while two tailed, paired Student’s t test was performed when only two cell subsets were included. Welch’s t test (unpaired t test with Welch’s correction) was performed to compare the differential gene expression between healthy controls and MS patients on a specific TH subset. Two sided p-values of <0.05 were considered statistically significant.
Code availability
Computer code used to generate results is available from the corresponding author upon request.
Data availability
The authors declare that the main data supporting the findings of this study are available within the article and its Supplementary Information files. Gene expression data that support the findings of this study have been deposited in the Gene Expression Omnibus with the accession code (GSE104024). (All other relevant data are available from the corresponding author upon reasonable request.
References
Hoe, E. et al. The contrasting roles of Th17 immunity in human health and disease. Microbiol. Immunol. 61, 49–56 (2017).
Nistala, K. et al. Interleukin-17-producing T cells are enriched in the joints of children with arthritis, but have a reciprocal relationship to regulatory T cell numbers. Arthritis Rheum. 58, 875–887 (2008).
Ivanov, I. I. et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17+T helper cells. Cell 126, 1121–1133 (2006).
Langrish, C. L. et al. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. J. Exp. Med. 201, 233–240 (2005).
Park, H. et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 6, 1133–1141 (2005).
Steinman, L. A brief history of T(H)17, the first major revision in the T(H)1/T(H)2 hypothesis of T cell-mediated tissue damage. Nat. Med. 13, 139–145 (2007).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Noack, M. & Miossec, P. Th17 and regulatory T cell balance in autoimmune and inflammatory diseases. Autoimmun. Rev. 13, 668–677 (2014).
Zielinski, C. E. et al. Pathogen-induced human TH17 cells produce IFN-gamma or IL-10 and are regulated by IL-1beta. Nature 484, 514–518 (2012).
Steinman, L. Inflammatory cytokines at the summits of pathological signal cascades in brain diseases. Sci. Signal. 6, pe3 (2013).
Cosmi, L., Santarlasci, V., Maggi, L., Liotta, F. & Annunziato, F. Th17 plasticity: pathophysiology and treatment of chronic inflammatory disorders. Curr. Opin. Pharmacol. 17, 12–16 (2014).
Lee, Y. et al. Induction and molecular signature of pathogenic TH17 cells. Nat. Immunol. 13, 991–999 (2012).
McGeachy, M. J. et al. TGF-beta and IL-6 drive the production of IL-17 and IL-10 by T cells and restrain T(H)-17 cell-mediated pathology. Nat. Immunol. 8, 1390–1397 (2007).
Acosta-Rodriguez, E. V. et al. Surface phenotype and antigenic specificity of human interleukin 17-producing T helper memory cells. Nat. Immunol. 8, 639–646 (2007).
Ma, C. S. et al. Deficiency of Th17 cells in hyper IgE syndrome due to mutations in STAT3. J. Exp. Med. 205, 1551–1557 (2008).
Puel, A. et al. Chronic mucocutaneous candidiasis in humans with inborn errors of interleukin-17 immunity. Science 332, 65–68 (2011).
Guo, B. IL-10 modulates Th17 pathogenicity during autoimmune diseases. J. Clin. Cell. Immunol. 7, 400 (2016).
Sallusto, F., Zielinski, C. E. & Lanzavecchia, A. Human Th17 subsets. Eur. J. Immunol. 42, 2215–2220 (2012).
Cao, Y. et al. Functional inflammatory profiles distinguish myelin-reactive T cells from patients with multiple sclerosis. Sci. Transl. Med. 7, 287ra274 (2015).
Cosmi, L. et al. Evidence of the transient nature of the Th17 phenotype of CD4+CD161+T cells in the synovial fluid of patients with juvenile idiopathic arthritis. Arthritis Rheum. 63, 2504–2515 (2011).
Nistala, K. et al. Th17 plasticity in human autoimmune arthritis is driven by the inflammatory environment. Proc. Natl Acad. Sci. USA 107, 14751–14756 (2010).
Kebir, H. et al. Preferential recruitment of interferon-gamma-expressing TH17 cells in multiple sclerosis. Ann. Neurol. 66, 390–402 (2009).
Reinert-Hartwall, L. et al. Th1/Th17 plasticity is a marker of advanced beta cell autoimmunity and impaired glucose tolerance in humans. J. Immunol. 194, 68–75 (2015).
Colombel, J. F., Sendid, B., Jouault, T. & Poulain, D. Secukinumab failure in Crohn’s disease: the yeast connection? Gut 62, 800–801 (2013).
Hueber, W. et al. Secukinumab, a human anti-IL-17A monoclonal antibody, for moderate to severe Crohn’s disease: unexpected results of a randomised, double-blind placebo-controlled trial. Gut 61, 1693–1700 (2012).
Fragoulis, G. E., Siebert, S. & McInnes, I. B. Therapeutic targeting of IL-17 and IL-23 cytokines in immune-mediated diseases. Annu. Rev. Med. 67, 337–353 (2016).
O’Connor, W. Jr. et al. A protective function for interleukin 17A in T cell-mediated intestinal inflammation. Nat. Immunol. 10, 603–609 (2009).
Maloy, K. J. & Kullberg, M. C. IL-23 and Th17 cytokines in intestinal homeostasis. Mucosal Immunol. 1, 339–349 (2008).
Muranski, P. et al. Tumor-specific Th17-polarized cells eradicate large established melanoma. Blood 112, 362–373 (2008).
Martin-Orozco, N. et al. T helper 17 cells promote cytotoxic T cell activation in tumor immunity. Immunity 31, 787–798 (2009).
Hus, I. et al. Th17/IL-17A might play a protective role in chronic lymphocytic leukemia immunity. PLoS ONE 8, e78091 (2013).
Lakshmi Narendra, B., Eshvendar Reddy, K., Shantikumar, S. & Ramakrishna, S. Immune system: a double-edged sword in cancer. Inflamm. Res. 62, 823–834 (2013).
Ramesh, R. et al. Pro-inflammatory human Th17 cells selectively express P-glycoprotein and are refractory to glucocorticoids. J. Exp. Med. 211, 89–104 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Ouyang, W., Rutz, S., Crellin, N. K., Valdez, P. A. & Hymowitz, S. G. Regulation and functions of the IL-10 family of cytokines in inflammation and disease. Annu. Rev. Immunol. 29, 71–109 (2011).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Laurence, A. et al. Interleukin-2 signaling via STAT5 constrains T helper 17 cell generation. Immunity 26, 371–381 (2007).
Yao, Z. et al. Nonredundant roles for Stat5a/b in directly regulating Foxp3. Blood 109, 4368–4375 (2007).
Ghoreschi, K., Laurence, A., Yang, X. P., Hirahara, K. & O’Shea, J. J. T helper 17 cell heterogeneity and pathogenicity in autoimmune disease. Trends Immunol. 32, 395–401 (2011).
Balashov, K. E., Rottman, J. B., Weiner, H. L. & Hancock, W. W. CCR5(+) and CXCR3(+) T cells are increased in multiple sclerosis and their ligands MIP-1alpha and IP-10 are expressed in demyelinating brain lesions. Proc. Natl Acad. Sci. USA 96, 6873–6878 (1999).
Sorensen, T. L. et al. Multiple sclerosis: a study of CXCL10 and CXCR3 co-localization in the inflamed central nervous system. J. Neuroimmunol. 127, 59–68 (2002).
Mahad, D. J., Howell, S. J. & Woodroofe, M. N. Expression of chemokines in the CSF and correlation with clinical disease activity in patients with multiple sclerosis. J. Neurol. Neurosurg. Psychiatry 72, 498–502 (2002).
Szczucinski, A. & Losy, J. CCL5, CXCL10 and CXCL11 chemokines in patients with active and stable relapsing-remitting multiple sclerosis. Neuroimmunomodulation 18, 67–72 (2011).
Rice, G. P., Hartung, H. P. & Calabresi, P. A. Anti-alpha4 integrin therapy for multiple sclerosis: mechanisms and rationale. Neurology 64, 1336–1342 (2005).
Haas, J. et al. Th17 cells: a prognostic marker for MS rebound after natalizumab cessation? Mult. Scler. 23, 114–118 (2017).
Nakajima, C. et al. Induction of the chemokine receptor CXCR3 on TCR-stimulated T cells: dependence on the release from persistent TCR-triggering and requirement for IFN-gamma stimulation. Eur. J. Immunol. 32, 1792–1801 (2002).
Krakowski, M. & Owens, T. Interferon-gamma confers resistance to experimental allergic encephalomyelitis. Eur. J. Immunol. 26, 1641–1646 (1996).
Ferber, I. A. et al. Mice with a disrupted IFN-gamma gene are susceptible to the induction of experimental autoimmune encephalomyelitis (EAE). J. Immunol. 156, 5–7 (1996).
Chu, C. Q., Wittmer, S. & Dalton, D. K. Failure to suppress the expansion of the activated CD4 T cell population in interferon gamma-deficient mice leads to exacerbation of experimental autoimmune encephalomyelitis. J. Exp. Med. 192, 123–128 (2000).
Spanier, J. A., Nashold, F. E., Olson, J. K. & Hayes, C. E. The Ifng gene is essential for Vdr gene expression and vitamin D(3)-mediated reduction of the pathogenic T cell burden in the central nervous system in experimental autoimmune encephalomyelitis, a multiple sclerosis model. J. Immunol. 189, 3188–3197 (2012).
Wang, Z. et al. Role of IFN-gamma in induction of Foxp3 and conversion of CD4+CD25- T cells to CD4+Tregs. J. Clin. Invest. 116, 2434–2441 (2006).
Sosa, R. A., Murphey, C., Robinson, R. R. & Forsthuber, T. G. IFN-gamma ameliorates autoimmune encephalomyelitis by limiting myelin lipid peroxidation. Proc. Natl Acad. Sci. USA 112, E5038–E5047 (2015).
Bettelli, E. et al. Loss of T-bet, but not STAT1, prevents the development of experimental autoimmune encephalomyelitis. J. Exp. Med. 200, 79–87 (2004).
Nath, N., Prasad, R., Giri, S., Singh, A. K. & Singh, I. T-bet is essential for the progression of experimental autoimmune encephalomyelitis. Immunology 118, 384–391 (2006).
O’Connor, R. A., Cambrook, H., Huettner, K. & Anderton, S. M. T-bet is essential for Th1-mediated, but not Th17-mediated, CNS autoimmune disease. Eur. J. Immunol. 43, 2818–2823 (2013).
Duhen, R. et al. Cutting edge: the pathogenicity of IFN-gamma-producing Th17 cells is independent of T-bet. J. Immunol. 190, 4478–4482 (2013).
Grifka-Walk, H. M., Lalor, S. J. & Segal, B. M. Highly polarized Th17 cells induce EAE via a T-bet independent mechanism. Eur. J. Immunol. 43, 2824–2831 (2013).
Brucklacher-Waldert, V. et al. Tbet or continued RORgammat expression is not required for Th17-associated immunopathology. J. Immunol. 196, 4893–4904 (2016).
Renner, E. D. et al. Novel signal transducer and activator of transcription 3 (STAT3) mutations, reduced T(H)17 cell numbers, and variably defective STAT3 phosphorylation in hyper-IgE syndrome. J. Allergy Clin. Immunol. 122, 181–187 (2008).
McDonald, D. R. TH17 deficiency in human disease. J. Allergy Clin. Immunol. 129, 1429–1435 (2012).
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Acknowledgements
We wish to thank Deneen Kozoriz and David Jarrossay for cell sorting, and Brian Healy for scientific discussion. This work was supported in part by National Institutes of Health P01 grant NS076410 (to H.L.W.), National Multiple Sclerosis Society research grant RG-1507-05029 (to H.L.W. & D.H.), European Research Council grant 323183 (to F.S.), and Swiss National Science Foundation grant 170213 (to F.S.). F.S. and the Institute for Research in Biomedicine are supported by the Helmut Horten Foundation.
Author information
Authors and Affiliations
Contributions
D.H. designed and supervised the study, performed experiments, analyzed data, and wrote the manuscript; S.N. performed experiments, analyzed data, and wrote the paper; M.P. and I.V.P. performed experiments; T.C., E.T., B.P., R.C., T.-H.Y., and K.K. conducted computational data analysis; N.P. supervised the comparative genomics data analyses and wrote the manuscript; D.A. performed experiments and analyzed data. Y.L. provided mouse microarray data; P.K. provided patient samples, discussions, and edited the manuscript. K.M.A., M.B., and M.G. provided patient samples; F.S. supervised the study and edited the manuscript; V.K.K. and H.L.W. designed and supervised the study, analyzed data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, D., Notarbartolo, S., Croonenborghs, T. et al. Transcriptional signature of human pro-inflammatory TH17 cells identifies reduced IL10 gene expression in multiple sclerosis. Nat Commun 8, 1600 (2017). https://doi.org/10.1038/s41467-017-01571-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-017-01571-8
This article is cited by
-
Mitochondrial and metabolic dysfunction of peripheral immune cells in multiple sclerosis
Journal of Neuroinflammation (2024)
-
Autoreactive T cells target peripheral nerves in Guillain–Barré syndrome
Nature (2024)
-
Amyloid PET scan diagnosis of Alzheimer’s disease in patients with multiple sclerosis: a scoping review study
Egyptian Journal of Radiology and Nuclear Medicine (2023)
-
miR-21 antagonism abrogates Th17 tumor promoting functions in multiple myeloma
Leukemia (2021)
-
MAIT cell alterations in adults with recent-onset and long-term type 1 diabetes
Diabetologia (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.