Abstract
We encountered a patient with mitochondrial trifunctional protein deficiency in whom the corresponding mutations were not identified by a DNA panel for newborn screening for targeted diseases. After diagnosis confirmation by an enzyme assay and immunoblotting using the autopsied liver, the re-evaluation of the panel data indicated a heterozygous deletion of exons 6–9 that was later confirmed at the genomic level. cDNA analysis also identified exonization of the 5′ region of intron 9 caused by a deep intronic mutation, c.811 + 82A>G.
Similar content being viewed by others
Introduction
Mitochondrial trifunctional protein (TFP) deficiency is a rare, autosomal recessive disorder. The main manifestations of TFP are cardiomyopathy, hypoketotic hypoglycemia, metabolic acidosis, sudden infant death, metabolic encephalopathy, liver dysfunction, peripheral neuropathy, exercise-induced myoglobinuria, and rhabdomyolysis1,2. This disease is classified into three subtypes based on the severity and onset age: the most severe form is the lethal phenotype with neonatal onset, the intermediate form is the hepatic phenotype with infant onset, and the mild form is the myopathic phenotype with late-adolescent onset3. TFP is a multienzyme complex consisting of four α and four β subunits encoded by the genes HADHA (MIM 600890) and HADHB (MIM 143450), respectively4,5. This complex exhibits three distinct enzyme activities, functioning as a long-chain enoyl-CoA hydratase (LCEH), a long-chain 3-hydroxyacyl-CoA dehydrogenase (LCHAD), and a long-chain 3-ketoacyl-CoA thiolase (LCKT). The α subunit harbors the LCHAD and LCEH functions, and the β subunit exhibits the LCKT function. TFP binds to the inner mitochondrial membrane and plays a significant role in the last three steps of the β-oxidation cycle of long-chain acyl-CoAs5,6. The incidence of TFP deficiency has been estimated to be 1 per 100,000 births in Europe7. To date, 14 TFP-deficient patients have been reported in Japan8. Seventy-two mutations have been identified in HADHA, and sixty-seven mutations have been found in HADHB (HGMD-professional-release-2019.3). We referred to the HGMD-Professional-release-2019.3 database and ClinVar_20191202 to determine whether the mutations identified in this patient were novel.
The patient was a first child born after 37 weeks and 6 days of gestation by cesarean section to nonconsanguineous parents. His birth weight was 2740 g. Newborn screening showed elevated C16-OH acylcarnitine (1.55 nmol/ml; cutoff < 0.1). Subsequent serum acylcarnitine analysis showed the C16-OH level to be 0.221 nmol/ml, with C18:1-OH at 0.219 nmol/ml. Urinary organic acid analysis was not specific. The patient was tentatively diagnosed with TFP deficiency (the patient’s clinical details are described in Supplementary Information 1). Informed consent for molecular analysis obtained from the parents, and the Ethics Committee of Gifu University Hospital approved the study (approval 29–210). Gene panel analysis using next-generation sequencing (NGS) with the MiSeq Sequencing System (Illumina, San Diego, CA, USA) was performed at the Kazusa DNA Research Institute. Genomic DNA was extracted from the patient’s peripheral blood leukocytes. We designed panel consisting of 60 genes was to detect HADHB mutations and metabolic disorders (Supplementary Table 1). This panel was designed to capture the designated protein-coding regions and 10 bp of flanking intronic sequences9. However, the gene panel analysis did not initially identify any mutations in HADHA and HADHB or in other genes related to fatty acid beta-oxidation. The patient was carefully managed but died of cardiomyopathy at 3 years 9 months of age after recurrent episodes of rhabdomyolysis.
We confirmed the diagnosis of TFP deficiency based on the lack of TFP enzyme activity and the absence of both HADHA and HADHB proteins in the liver at autopsy (Supplementary Fig. 1). Western blotting was performed using rabbit polyclonal antibodies raised against purified MTP protein as the primary antibody (provided by Dr. T. Hashimoto), as previously described10. 3-ketoacyl-CoA thiolase activity, measured as described previously, was almost null in the liver of the patient (1.85 µmol/min/mg protein, control sample 93.9 µmol/min/mg protein)11.
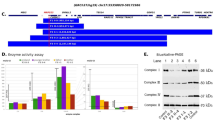
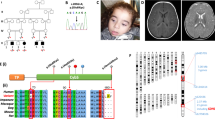
Therefore, we re-examined the gene panel data and found that the normalized read depths for exons 6–9 were lower than those of the other exons in HADHB (Fig. 1a). Polymerase chain reaction (PCR) using a forward primer for intron 5 (5′-TTCTGGACCTGGTATCAGTC-3′) and a reverse primer for intron 9 (5′-CTCTATGGAACCACAAGCCTT-3′) successfully amplified a truncated genomic fragment (781 bp) in the patient and his father (Fig. 1b). Therefore, the deletion allele of the patient was inherited from his father. Direct sequencing of this fragment revealed the breakpoint (Chr2: 26,272,937–26,279,402 in GRCh38.p13) for the deletion of exons 6–9 (Fig. 1c).
a Representation of the next-generation sequencing data coverage depth using the Integrative Genomics Viewer. A suspected heterozygous deletion of HADHB exons 6–9 was detected. b Deletion allele-specific PCR of the HADHB gene was performed. The deletion allele could only be amplified by the indicated primers flanking the deletion in the patient and his father. The wild-type allele was too large to be amplified by these primers under the applied PCR conditions; hence, no amplification products were obtained for the patient’s mother. c Sequencing analysis of the 781-bp fragment spanning the exons 6–9 deletion indicated breakpoints upstream of exon 6 (c.255–713) and downstream of exon 9 (c.811 + 86) in the HADHB gene in both the patient and his father.
To identify the patient’s other mutation inherited from his mother, cDNA analysis was performed. RNA was isolated from the patient’s autopsied heart and a control fibroblast sample (Kurabo, Osaka, Japan) with an Isogen kit (Nippon Gene, Tokyo, Japan). cDNA synthesis was performed using the HADHB-specific antisense primers Ex16rv1 (5′-GGATAAGCTTCCACTATCATAGC-3′) and Ex16rv2 (5′-GGCAAGGCTTAAGTGCAAAC-3′), an oligo (dT) primer (Thermo Fisher Scientific, Waltham, MA, USA), and a random hexamer primer (Thermo Fisher Scientific). When cDNA amplification was carried out with a forward primer for exon 8 (5′-ATGCTTGATCTCAATAAGGCC-3′) and a reverse primer for exon 12 (5′-GATCTTTTGGATCCTGAGACAC-3′), a longer cDNA fragment than that of the control was detected (Fig. 2a). Direct sequencing of the larger PCR fragment from the patient showed an insertion (81 bp) between exons 9 and 10. The insertion was derived from intron 9 and introduced a frameshift and premature termination within the insertion (p.Pro270Profs*14).
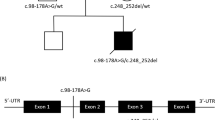
a Schematic representation of the strategy used to detect aberrant HADHB transcripts by PCR using a forward primer for exon 8 and a reverse primer for exon 12. The intronic exonized region is shown in the black box (insertion of 81 bp in intron 9). The c.811 + 82A>G variant creates a new cryptic 5′ splice site. Amplified transcripts from the patient were characterized by predominance of the 81-bp sequence of HADHB, resulting in a 583-bp amplicon. Transcripts from the control fibroblasts showed only the predicted wild-type amplicon size of 502 bp. b The patient and his mother carried the c.811 + 82A>G HADHB mutation. The other allele of the patient contained the exons 6–9 deletion, whereas the mother was heterozygous for c.811 + 82A>G and did not carry the deletion. The father was hemizygous for wild-type c.811 + 82A because his other allele contained the exons 6–9 deletion.
To shed light on the cause of the exonization of this intronic sequence, we performed direct sequencing of intron 9 and detected a HADHB NM_000183.3:c.811 + 82A>G substitution at the genomic DNA level (Fig. 2b). His mother also carried this HADHB c.811 + 82A>G mutation in a heterozygous manner. The exonization can be explained by the activation of a cryptic splice site 81 bp downstream of the authentic 5′ splice site of intron 9. This intronic mutation was found in dbSNP (https://www.ncbi.nlm.nih.gov/snp/) in rs757303269 and identified as an intron_variant (minor allele frequency from TOPMED is G = 0.000016/2).
In silico analyses were then performed on the splice sites. MaxEntScan::score5ss (http://hollywood.mit.edu/burgelab/maxent/Xmaxentscan_scoreseq.html) provided a much higher score for the activated cryptic 5′ splice site than for the authentic 5′ splice site of intron 9. The Senapathy & Shapiro (S&S) matrix12 and Human Splicing Finder (http://www.umd.be/HSF/HSF.shtml) provided similar scores for the two sites (Supplementary Table 2).
Even though the aberrantly spliced mRNA with a premature termination within the 81-bp insertion should theoretically be subjected to nonsense-mediated mRNA decay, cDNA analysis showed that the aberrantly spliced mRNA was a major transcript in the patient (Fig. 2a). Indeed, the in silico data indicated the predominant use of the activated cryptic 5′ splice site within the patient’s transcripts. Another deep intronic mutation (g.33627A>G) in HADHB resulting in intronic exonization was previously found in a Japanese patient13.
In conclusion, we report a case of TFP deficiency in a 3-year-old Japanese boy with a new pathogenic HADHB intronic mutation resulting in an atypical splice site and a large deletion. Thus, cDNA analysis can provide clues for revealing deep intronic mutations that are difficult to identify by exome or gene panel analysis.
HGV Database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2817. https://doi.org/10.6084/m9.figshare.hgv.2820.
Data availability
The data and materials described in this report are available upon request. Supplementary Information is available online through the Human Genome Variation journal.
References
Rhead, W. J. Inborn errors of fatty acid oxidation in man. Clin. Biochem. 24, 319–329 (1991).
Coates, P. M. & Tanaka, K. Molecular basis of mitochondrial fatty acid oxidation defects. J. Lipid Res. 33, 1099–1110 (1992).
Spiekerkoetter, U., Sun, B., Khuchua, Z., Bennett, M. J. & Strauss, A. W. Molecular and phenotypic heterogeneity in mitochondrial trifunctional protein deficiency due to beta-subunit mutations. Hum. Mutat. 21, 598–607 (2003).
Kamijo, T. et al. Mitochondrial trifunctional protein deficiency. Catalytic heterogeneity of the mutant enzyme in two patients. J. Clin. Invest. 93, 1740–1747 (1994).
Sims, H. F. et al. The molecular basis of pediatric long chain 3-hydroxyacyl-CoA dehydrogenase deficiency associated with maternal acute fatty liver of pregnancy. Proc. Natl Acad. Sci. USA 92, 841–845 (1995).
Carpenter, K., Pollitt, R. J. & Middleton, B. Human liver long-chain 3-hydroxyacyl-coenzyme A dehydrogenase is a multifunctional membrane-bound beta-oxidation enzyme of mitochondria. Biochem. Biophys. Res. Commun. 183, 443–448 (1992).
Sander, J. et al. Neonatal screening for defects of the mitochondrial trifunctional protein. Mol. Genet. Metab. 85, 108–114 (2005).
Bo, R. et al. Clinical and molecular investigation of 14 Japanese patients with complete TFP deficiency: a comparison with Caucasian cases. J. Hum. Genet. 62, 809–814 (2017).
Fujiki, R. et al. Assessing the accuracy of variant detection in cost-effective gene panel testing by next-generation sequencing. J. Mol. Diagn. 20, 572–582 (2018).
Towbin, H., Staehelin, T. & Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl Acad. Sci. USA 76, 4350–4357 (1979).
Uchida, Y., Izai, K., Orii, T. & Hashimoto, T. Novel fatty acid beta-oxidation enzymes in rat liver mitochondria. II. Purification and properties of enoyl-coenzyme A (CoA) hydratase/3-hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase trifunctional protein. J. Biol. Chem. 267, 1034–1041 (1992).
Shapiro, M. B. & Senapathy, P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 15, 7155–7174 (1987).
Purevsuren, J. et al. Study of deep intronic sequence exonization in a Japanese neonate with a mitochondrial trifunctional protein deficiency. Mol. Genet. Metab. 95, 46–51 (2008).
Acknowledgements
We thank the patient and family who participated in our study. We also thank Ms. Midori Furui for technical assistance. This research was supported by AMED under Grant Number JP17ek0109276 and by Health and Labor Sciences Research Grants (H29-nanchitou(nan)-ippan-051) for research on rare and intractable diseases. We thank Natasha Beeton-Kempen, Ph.D., and Rebecca Porter, Ph.D., from the Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nakama, M., Sasai, H., Kubota, M. et al. Novel HADHB mutations in a patient with mitochondrial trifunctional protein deficiency. Hum Genome Var 7, 10 (2020). https://doi.org/10.1038/s41439-020-0097-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-020-0097-z