Abstract
Heterozygous pathogenic variants in the KAT6B gene, which encodes lysine acetyltransferase 6B, have been identified in patients with congenital rare disorders, including genitopatellar syndrome and Say-Barber-Biesecker-Young-Simpson syndrome. Herein, we report another Japanese patient with a KAT6B-related disorder and a novel de novo heterozygous variant in exon 18 of KAT6B [c.3925dup, p.(Glu1309fs*33)], providing further evidence that truncating variants in exon 17 and in the proximal region of exon 18 are associated with genitopatellar syndrome-like phenotypes.
Similar content being viewed by others
Genitopatellar syndrome (GPS) is a rare autosomal dominant disorder characterized by patellar hypoplasia/agenesis, urogenital anomalies, congenital flexion contractures of the large joints, microcephaly, agenesis of corpus callosum, and hydronephrosis (OMIN #606170). Heterozygous de novo pathogenic variants in the lysine acetyltransferase 6B (KAT6B) gene have been reported as a cause of GPS1. Heterozygous truncating pathogenic variants of KAT6B are also associated with Say-Barber-Biesecker-Young-Simpson syndrome (SBBYSS) (OMIM#603736), which is characterized by long thumbs, great toes, an immobile mask-like face, blepharophimosis/ptosis, and lacrimal duct anomalies. Although GPS and SBBYSS were originally considered independent clinical entities, they are often indistinguishable and share some common phenotypes, such as delayed development, intellectual disability, congenital heart defects, thyroid dysfunction, and/or genital anomalies. Therefore, they are collectively called KAT6B-related disorders1,2. Herein, we report an additional Japanese patient with a KAT6B-related disorder and a novel heterozygous frameshift variant of KAT6B.
This male Japanese patient was born at 37 weeks of gestation as the first child of non-consanguineous phenotypically normal parents. During pregnancy, fetal echography showed fetal growth restriction, enlargement of the bilateral cerebral ventricles, and bilateral hydronephrosis. At birth, his length was 44.5 cm (−1.2 standard deviation [SD]), his weight was 2.12 kg (−1.6 SD), and his head circumference was 28.5 cm (−2.8 SD). He had distinct facial features, with a depressed nasal bridge, a bulbous nose, micrognathia, and low-set ears. He also had flexion contractures of the hips and knees, overlapping of the left toes, exostoses on the right foot, scrotal hypoplasia, and cryptorchidism (Fig. 1a). Cardiac echography showed patent ductus arteriosus. Brain magnetic resonance imaging (MRI) revealed agenesis of the corpus callosum (Fig. 1b). A skeletal survey showed bilateral radioulnar synostosis (Fig. 1c). At 4 months of age, the patient was diagnosed with bilateral patellae were revealed by knee MRI. Primary hypothyroidism, which was not detected by neonatal mass screening, was identified at 55 days of age (serum thyroid-stimulating hormone, 37.2 μIU/ml [normal range: 0.6–5.9]; free thyroxine, 0.88 ng/dl [1.37–1.51]; free triiodothyronine, 3.09 pg/ml [4.3–5.1]. Thyroid ultrasonography showed tolerably located thyroid glands of normal size. Levothyroxine sodium treatment promptly normalized thyroid function. The patient’s karyotype was normal, and urine steroid profile analysis did not show a pattern characteristic of cytochrome P450 oxidoreductase deficiency. At the final examination at 10 months of age, his height was 67.3 cm (−2.2 SD), and his weight was 7030 g (−2.0 SD). He had hypotonia, and his motor and mental development was severely delayed (DQ 55).
This study was approved by the Institutional Review Board of Nagasaki University Graduate School of Biomedical Sciences. Since clinical assessment alone did not lead to a conclusive diagnosis, we sought to identify disease-causing pathogenic variants with a trio whole-exome sequencing (WES) strategy using a SureSelect Human All Exon V5 (Agilent Technologies, Santa Clara, CA, USA) on a HiSeq 2500 platform (Illumina, San Diego, CA, USA). Written informed consent was obtained from the parents. DNA was obtained from peripheral blood samples from the patient and his parents. The reads in FASTQ format were aligned to the human reference genome using NovoAlign version 3.0 (http://www.novocraft.com/). Trio-based genomic variation information was detected by the Genome Analysis Toolkit version 3.4–463. Subsequently, de novo, homozygous, and X-linked variations were extracted and annotated by ANNOVAR software4; in this process, variants with an allele frequency >0.5% in the Exome Aggregation Consortium (http://exac.broadinstitute.org/), NHLBI GO Exome Sequencing Project (http://evs.gs.washington.edu/EVS/), Human Genetic Variation Database5 (http://www.hgvd.genome.med.kyoto-u.ac.jp), or 3.5 KJPN database of Tohoku Medical Megabank6 (https://jmorp.megabank.tohoku.ac.jp/201902/) were excluded. Heterozygous variations with the same annotation in GENCODE v19 were also extracted to detect compound heterozygous variants. Each variant was confirmed via Sanger sequencing using a BigDye terminator and 3130xl genetic analyzer (Applied Biosystems, Carlsbad, CA, USA).
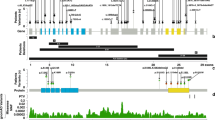
Through these investigations, we identified several candidate variants. Of these, a de novo heterozygous variant in KAT6B (c.3925dup, NM_012330.3) was proposed as the best candidate based on WES data and the Online Mendelian Inheritance in Man database of known diseases (www.omim.org). The 1-bp duplication in exon 18 of KAT6B was predicted to cause a frameshift at codon 1309 of KAT6B, resulting in termination at codon 1342 of the last exon (exon 18) [p.(Glu1309Glyfs*33), NP_036462]; this prediction indicates that the frameshift pathogenic variant can escape nonsense-mediated mRNA decay (NMD) and likely produces a truncated protein lacking the distal section of the acidic domain and the entire C-terminal transcription activation domain (Fig. 2)7.
The black and white boxes on genomic DNA (gDNA) denote the coding regions of exons 16–18 and the untranslated region, respectively. The variants leading to GPS and SBBYSS phenotypes are shown in bold and underline, respectively. a Variants with mixed or overlapping phenotypes. b Variant in the present patient.
To date, at least 62 pathogenic variants have been reported in patients with KAT6B-related disorders, which have a broad clinical spectrum, including the phenotypes of GPS and SBBYSS1,2,8. In this regard, several findings are noteworthy regarding genotype–phenotype correlations. GPS-specific phenotypes, such as patellar hypoplasia/agenesis, congenital flexion contractures of the large joints, microcephaly, hydronephrosis, and agenesis of the corpus callosum, have been identified in patients with truncating pathogenic variants between the distal region of exon 17 and the proximal region of exon 18 (Fig. 2). On the other hand, most of the truncating variants in the distal region of exon 18 have been identified in patients with SBBYSS-specific phenotypes, such as long thumbs, great toes, blepharophimosis/ptosis, and lower-extremity joint stiffness. In addition, more proximal truncating variants, which are predicted to undergo NMD, have been associated with milder phenotypes due to haploinsufficiency. These data and data on the present case indicate that truncated KAT6B proteins that lack the C-terminal transactivation domain but retain the proximal region of the acidic domain escape NMD and are strongly associated with GPS phenotypes.
The clinical features of the present patient highlight two interesting findings. First, the patient exhibited bilateral radioulnar synostosis, which has been reported in only one previous patient with GPS, and this previous patient had an unknown KAT6B genotype9. The present case is the first report of a genetically confirmed KAT6B-related disorder with bilateral radioulnar synostosis. Second, our patient presented with primary hypothyroidism, which was fortuitously diagnosed at 55 days of age. Thyroid abnormalities are often observed in patients with KAT6B-related disorders10. However, the severity and onset of the phenotype seem to be variable11,12,13,14,15. Indeed, the present patient had normal TSH levels at the time of neonatal mass screening. Hypothyroidism was diagnosed at 14 years of age in another patient with a KAT6B-related disorder16. Although the pathological mechanisms underlying the thyroid abnormalities remain to be determined, thyroid function should be frequently examined in patients with KAT6B-related disorders.
In conclusion, our study provides further evidence that heterozygous truncating pathogenic variants in the distal region of exon 17 and proximal region of exon 18 in KAT6B cause GPS phenotypes. Further studies are needed to determine the clinical spectrum of KAT6B-related disorders and the pathogenesis of KAT6B variants.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2795.
References
Campeau, P. M. et al. The KAR6B-related disorders Genitopatellar syndrome and Ohdo/SBBYS syndrome have distinct clinical features reflecting distinct molecular mechanisms. Hum. Mutat. 33, 1520–1525 (2012).
Vlckova, M. et al. A patient showing features of both SBBYSS and GPS supports the concept of a KAT6B-related disease spectrum, with mutations in mid-exon 18 possibly leading to combined phenotypes. Eur. J. Med. Genet. 58, 550–555 (2015).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Higasa, K. et al. Human genetic variation database, a reference database of genetic variations in the Japanese population. J. Hum. Genet. 61, 547–553 (2016).
Yamaguchi-Kabata, Y. et al. iJGVD: an integrative Japanese genome variation database based on whole-genome sequencing. Hum. Genome Var. 26, 15050 (2015).
Holbrook, J. A., Neu-Yilik, G., Hentze, M. W. & Kulozik, A. E. Nonsense-mediated decay approaches the clinic. Nat. Genet. 36, 801–808 (2004).
Brea-Fernández, A., Dacruz, D., Eirís, J., Barros, F. & Carracedo, Á. Novel truncating variants expand the phenotypic spectrum of KAT6B-related disorders. Am. J. Med. Genet. A 179A, 290–294 (2019).
Abdul-Rahman, O. A. et al. Genitopatellar syndrome: expanding the phenotype and excluding mutation in LMX1B and TBX4. Am. J. Med. Genet. A 140, 1567–1572 (2006).
Gannon, T. et al. Further delineation of the KAT6B molecular and phenotypic spectrum. Eur. J. Hum. Genet. 23, 1165–1170 (2015).
Yilmaz, R. et al. A recurrent synonymous KAT6B mutation causes Say-Barber-Biesecker/Young-Simpson syndrome by inducing aberrant splicing. Am. J. Med. Genet. A 167A, 3006–3010 (2015).
Penttinen, M., Koillinen, H., Niinikoski, H., Mäkitie, O. & Hietala, M. Genitopatellar syndrome in an adolescent female with severe osteoporosis and endocrine abnormalities. Am. J. Med. Genet. A 149A, 451–455 (2009).
Young, I. D. & Simpson, K. Unknown syndrome: abnormal facies, congenital heart defects, hypothyroidism, and severe retardation. J. Med. Genet. 24, 715–716 (1987).
Marangi, G. et al. A novel truncating variant within exon 7 of KAT6B associated with features of both Say-Barber-Bieseker-Young-Simpson syndrome and genitopatellar syndrome: Further evidence of a continuum in the clinical spectrum of KAT6B-related disorders. Am. J. Med. Genet. A 176, 455–459 (2018).
Szakszon, K. et al. De novo mutations of the gene encoding the histone acetyltransferase KAT6B in two patients with Say-Barber/Biesecker/Young-Simpson syndrome. Am. J. Med. Genet. A 161A, 884–888 (2013).
Penttinen, M., Koillinen, H., Niinikoski, H., Mäkitie, O. & Hietala, M. Genitopatellar syndrome in an adolescent female with severe osteoporosis and endocrine abnormalities. Am. J. Med. Genet. A 149A, 451–455 (2009).
Acknowledgements
We thank the patient and family for participating in this study. This work was supported by a grant for the Initiative on Rare and Undiagnosed Diseases in Pediatrics (no. 18gk0110012h0101) from the Japan Agency for Medical Research and Development (AMED), Tokyo, Japan. This work was supported by a grant for the Initiative on Rare and Undiagnosed Diseases in Pediatrics (no. 18gk0110012h0101) from the Japan Agency for Medical Research and Development (AMED), Tokyo, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hamaguchi, Y., Aoki, M., Watanabe, S. et al. KAT6B-related disorder in a patient with a novel frameshift variant (c.3925dup). Hum Genome Var 6, 54 (2019). https://doi.org/10.1038/s41439-019-0085-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-019-0085-3