Abstract
Plants sense photoperiod signals to confirm the optimal flowering time. Previous studies have shown that Cryptochrome2 (CRY2) functions to promote floral transition in the long-day plant (LDP) Arabidopsis; however, the function and molecular mechanism by which CRY2 regulates floral transition in short-day plants (SDPs) is still unclear. In this study, we identified a CRY2 homologous gene, ClCRY2, from Chrysanthemum lavandulifolium, a typical SDP. The morphological changes in the C. lavandulifolium shoot apex and ClFTs expression analysis under SD conditions showed that adult C. lavandulifolium completed the developmental transition from vegetative growth to reproductive growth after eight SDs. Meanwhile, ClCRY2 mRNA exhibited an increasing trend from 0 to 8 d of SD treatment. ClCRY2 overexpression in wild-type (WT) Arabidopsis and C. lavandulifolium resulted in early flowering. The transcript levels of the CONSTANS-like (COL) genes ClCOL1, ClCOL4, and ClCOL5, and FLOWERING LOCUS T (FT) homologous gene ClFT1 were upregulated in ClCRY2 overexpression (ClCRY2-OE) C. lavandulifolium under SD conditions. The transcript levels of some circadian clock-related genes, including PSEUDO-REPONSE REGULATOR 5 (PRR5), ZEITLUPE (ZTL), FLAVIN-BINDING KELCH REPEAT F-BOX 1 (FKF1), and GIGANTEA (GI-1 and GI-2), were upregulated in ClCRY2-OE C. lavandulifolium, while the expression levels of other circadian clock-related genes, such as EARLY FLOWERING 3 (ELF3), ELF4, LATE ELONGATED HYPOCOTYL (LHY), PRR73, and REVEILLE8 (RVE8), were downregulated in ClCRY2-OE C. lavandulifolium under SD conditions. Taken together, the results suggest that ClCRY2 promotes floral transition by fine-tuning the expression of circadian clock-related gene, ClCOLs and ClFT1 in C. lavandulifolium under SD conditions.
Similar content being viewed by others
Introduction
Plants sense changes in external circumstances and integrate these signals with internal factors, such as gibberellin and age, to ensure optimal flowering time, which could guarantee the reproduction of the species1. Light length, namely photoperiod, changes regularly with different seasons. Most plants can sense photoperiod signals during seasonal changes. The plants monitor photoperiod signal changes by photoreceptors in their leaves and regulate the transcription of CONSTANS (CO) and FLOWERING LOCUS T (FT) to promote (or delay) the floral transition through different pathways2,3,4.
Plants have evolved at least five distinct families of sensory photoreceptors, including Cryptochromes (CRYs)5,6, FLAVIN-BINDING KELCH REPEAT F-BOX 1 (FKF1)/ZEITLUPE (ZTL)/LOV KELCH PROTEIN 2 (LKP2)7, Phototropins (PHOTs)8, Phytochromes (PHYs)9,10, and UV RESISTANCE LOCUS 8 (UVR8)11. Among these, PHYs, CRYs, and FKF1/ZTL/LKP2 are present as small gene families in all higher plants and they could control photoperiodic induction of flowering. CRYs are unique photoreceptors present in all major evolutionary lineages12. They are flavoproteins with similar sequences to DNA photolyases, which are light-activated DNA repair enzymes that mediate light signals to remove pyrimidine dimers from DNA to repair UV-induced DNA damage13,14. CRYs have lost DNA photolyase activity, but possess other biochemical functions15,16. For instance, they regulate floral transition and the circadian clock system in plants5,13,14. The function of CRYs in regulating photoperiodic flowering varies in different higher plants. Arabidopsis cry2 mutant exhibits a late-flowering phenotype under long day (LD) conditions rather than short day (SD) conditions17, which infers that CRY2 can specifically sense the inductive photoperiod signals to regulate the floral transition in Arabidopsis. OsCRY218 in rice, and MdCRY119 and MdCRY220 in apple act as floral stimulators, while SlCRY2 in tomato acts as an inhibitor to delay the flowering time21. PsCRY1 in pea has a slight inhibitory function on flowering, while phyacry1 in pea shows a distinct late-flowering phenotype22, indicating that PsCRY1 regulates floral transition in the presence of other photoreceptors, such as PsPHYA.
The CRY apoprotein is composed of the following two domains: (1) an N-terminal photolyase-homologous region (PHR) domain and (2) a Cryptochrome C-terminal extension domain (CCE)6. The PHR domain binds flavin adenine dinucleotide (FAD) and methenyltetrahydrofolate (MTHF)15,23,24,25. The CCE domain shares little sequence similarity among various plant species and is involved in CRY signal transduction26. Plant CRYs share a conserved DQXVP-acidic-STAESSS (DAS) motif in the CCE domain6,27. CRYs participate in the following three distinct pathways to control floral transition: (1) the CONSTITUTIVE PHOTOMORPHO-GENESIS 1 (COP1)/SUPPRESSOR of PHYA-105 (SPA1) dependent pathway, in which photoexcited CRYs promote the stability of CO protein by suppressing the activity of COP1/SPA1 complex28,29,30; (2) the CRY2-INTERACTING BHLHs (CIBs) dependent pathway28,31, in which CIBs converge signals from CRYs and ZTLs to regulate FT transcripts; and (3) the EARLY FLOWERING 3 (ELF3)/COP1-FKF1/GIGANTEA (GI) pathway, in which CRYs promote the stability of CO protein through the ELF3/COP1 and FKF1/GI complexes32. To date, studies on plant CRYs regulating photoperiodic flowering mainly focus on the CRYs-COP1/SPA and CRY2-CIB pathways. CRYs are also involved in circadian clock input and output pathways to regulate photoperiodic flowering33. However, the molecular mechanism by which CRYs regulate floral transition through circadian clock genes remains unclear.
Chrysanthemum (Chrysanthemum × morifolium Ramat.), commonly known as Flower of the Eastern, enjoys a major share of the commercial flower market. The primary objective of chrysanthemum breeding focuses on the modification of flowering time to meet the demand for marketable flowers throughout the year. CRY2 specifically regulates floral transition in Arabidopsis under inductive LD conditions17. The homologous CRY2 gene in chrysanthemum may play a crucial role in regulating the floral transition under inductive SD conditions. To date, the molecular mechanism by which CRY2 mediates SD signals to regulate the floral transition in chrysanthemum is unknown. Chrysanthemum lavandulifolium is a diploid wild species in the chrysanthemum genus. Our previous studies report that C. lavandulifolium is an obligate SD plant that is widely distributed in the northeast regions of China34, and reveal the functions of the CO and FT homologous genes in regulating the floral transition in C. lavandulifolium34,35. In this paper, we used C. lavandulifolium as a model plant for chrysanthemum cultivars to explore the role of ClCRY2 in floral transition. Our studies revealed the role of ClCRY2 in floral transition and showed the regulatory role of ClCRY2 in the circadian clock-related genes, ClCOL genes and the florigen gene ClFT1 during floral transition in C. lavandulifolium. The present study reveals a pathway whereby ClCRY2 mediates SD signals to regulate floral transition in C. lavandulifolium. ClCRY2 can also serve as a vital target for the genetic manipulation of flowering in chrysanthemum.
Materials and methods
Plant materials
C. lavandulifolium seedlings
The wild-type (WT) C. lavandulifolium lines were grown in Murashige and Skoog (MS) medium (pH 6.0 and 0.6% w/v agar) under LD conditions (16 h light/8 h dark, 50 μmol m−2 s−1), and the transgenic lines were grown in MS containing 400 mg/L carbenicillin (Car) and 7 mg/L kanamycin (Kan). When the tissue-cultured plantlets grew to four true leaves, they were transplanted to 9-cm flowerpots with vermiculite and turf (V: V = 1: 1) under LD conditions (16 h light/8 h dark, 108.4 μmol m−2 s−1). After the seedlings produced 14 leaves, they were transferred to SD conditions (12 h light/12 h dark, 108.4 μmol m−2 s−1). The light source was supplied using cool-white fluorescent lamps. The room temperature was 20 ± 1 °C with approximately 60% relative humidity.
Arabidopsis thaliana
The WT A. thaliana ecotype Columbia-0 (Col-0) and transgenic lines of the same ecotype background seeds were surface-sterilized and sowed in MS medium and MS medium containing 50 mg/L kanamycin, respectively. The seedlings were transferred into the same substrate as described above under LD conditions (16 h light/8 h dark, 108.4 μmol m−2 s−1) and SD1 conditions (8 h light/16 h dark, 108.4 μmol m−2 s−1) after 10 days. The room temperature was 20 ± 1 °C with approximately 60% relative humidity.
Anatomical observation of flower development in C. lavandulifolium
The apical buds at different developmental stages (after 0, 4, 8, 12, 16, 20, and 24 d of SD treatment) were obtained from ten individual plants and fixed in FAA (50% ethanol:formaldehyde solution:glacial acetic acid = 18: 1:1). After 24 h, we removed the apical buds from the FAA, dehydrated the buds with an ethanol series up to 100%, and then incubated the dehydrated buds with a xylene-ethanol series up to 100% xylene. The flower buds were embedded in Sigma Paraplast Plus paraffin, and the materials were sectioned into 8–10 μm sections using a rotary microtome (RM 2245; Heidelberg; LEICA; Germany). The sections were de-paraffinized and stained using safranine-fast green. Finally, the sections were detected using a light microscope (DM 2500; Heidelberg; LEICA; Germany).
Gene isolation and sequence analysis
Total RNA was extracted from leaves using a modified cetyltrimethylammonium bromide method, and first-strand cDNA was synthesized as described in a previous study36. We identified three unigenes annotated with ‘Cryptochrome2’ in a C. lavandulifolium Illumina/Solexa library37; only one unigene encoded the 5′-ends of a putative ‘Cryptochrome2’ gene. The 3′-specific primers were designed to amplify the 3′-ends of the putative ‘Cryptochrome2’gene using a 3′-rapid amplification of cDNA ends (3′-RACE) method. The full-length cDNA of the gene was amplified using PCR (specific primers are listed in Supplementary Table S1). The PCR product was cloned into the pGEM-T Easy Vector (Promega, USA) and confirmed by sequencing. The CD-search program (http://structure.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) was used to analyze the conserved protein domain sequences of ClCRY2. DNAMAN7.0 software was used to perform multiple sequence alignment. The conserved motif logo was generated by using ClustalX and WebLogo 3.4 program (http://weblogo.threeplusone.com/). MEGA version 4.0 was used to construct a phylogenetic tree.
Gene expression analysis using quantitative real-time PCR
In order to analyze the temporal expression patterns of ClCRY2, ClFT1 and ClFT2, the leaves from three individual plants at different developmental stages (after 0, 4, 8, 12, 16, 20, and 24 d of SD treatment) were harvested with the light on (ZT = 0). The roots, stems, middle leaves (the fourth true leaves from the shoot apex), petioles and shoot apices were sampled from three individual plants at the fourteenth-true-leaf stage after 8 days of SD treatment. These samples were used to analyze the tissue-specific expression patterns of the ClCRY2 in C. lavandulifolium. To investigate the effect of the SD photoperiods on ClCRY2 transcripts, the seedlings were placed under LD and SD conditions and the middle leaves were sampled at 4-h intervals over 24 h after 8 days. Three biological replicates were harvested for each sample.
The abundance of ClCRY2, ClFT1, and ClFT2 transcripts was investigated by quantitative real-time PCR (qRT-PCR) using a CFX connect Real-time PCR System (Bio-Rad, USA) based on the SYBR Premix Ex Taq (TaKaRa, Japan) as described previously36. Two reference genes were used in the assay. One was ClSAND (SAND family protein, GenBank accession number: KF752605) gene, which was used as a reference gene for analyzing ClCRY2 transcription levels in various tissues/organs and the transcripts of ClCRY2 and ClFT at different developmental stages. Another reference gene was ClMTP (Metal Tolerance Protein, GenBank accession number: KF752604), which was used to calculate the ClCRY2 transcripts in the leaves under different photoperiods38. The specific primer sequences used in the qRT-PCR assay are listed in Supplementary Table S1.
The function analysis of ClCRY2 in regulating floral transition
Full-length ClCRY2 cDNA was acquired by PCR amplification from middle leaves of C. lavandulifolium with specific primers (Supplementary Table S1). The product was cloned into the pGEM-T vector (Promega, USA). After digestion with restriction enzyme, the insert was ligated to the modified pBI121 vector39 to obtain mpBI121-ClCRY2. This construct was used to transform the Agrobacterium tumefaciens strain EHA105. Subsequently, Col-0 ecotype Arabidopsis plants were transformed with Agrobacterium using the floral-dip method. Transgenic Arabidopsis lines were selected on MS medium containing 50 mg/L Kan. Kan-resistant T1 generation Arabidopsis seedlings were transferred to soil to harvest T2 generation seeds; subsequently, homozygous T3 generation transgenic seeds were harvested for further assays. Rosette leaves from T3 generation transgenic seedlings and WT plants were harvested for qRT-PCR validation; the β-tubulin (TUB2) gene from Arabidopsis was used as a reference gene. For the analysis of flowering time, WT and T3 ClCRY2 overexpression (ClCRY2-OE) transgenic lines (>24 per line) were grown at 20 ± 1 °C under LD conditions and SD1 conditions. The flowering time was assayed by surveying the number of days from sowing to the date when the first flower opened. The number of rosette leaves of WT and T3 ClCRY2 overexpression (ClCRY2-OE) transgenic lines at bolting was recorded. Three biological replicates were assayed for the WT and transgenic lines.
Flowering-related gene expression and flowering phenotype analysis in C. lavandulifolium
Our previous study introduced a genetic transformation system suitable for C. lavandulifolium40. C. lavandulifolium hypocotyls were used as explants in the system. Each hypocotyl was cut into 2 mm sections. Subsequently, the sections were divided into two groups. Several hypocotyl sections were infected with Agrobacterium carrying ClCRY2 (OD600≈0.4) for 10 min and then laid on medium A (MS + 1.0 mg/L 2, 4-D + 0.5 mg/L 6-BA + 400 mg/L Car + 7 mg/L Kan) to obtain the calluses. The calluses were transferred to medium B (MS + 400 mg/L Car + 7 mg/L Kan) to induce the adventitious buds. Finally, single buds divided from the adventitious buds were transferred to medium B to induce roots. The other hypocotyl sections were treated as the control group. The sections were placed on MS + 1.0 mg/L 2, 4-D + 0.5 mg/L 6-BA and MS media in turn without infection to obtain the WT C. lavandulifolium.
Transgenic chrysanthemum lines were selected on MS + 7 mg/L Kan + 400 mg/L Car. Kan-resistant seedlings were transferred to substrate under LD conditions when they grew four true leaves. The seedlings were transferred to SD conditions after they produced 14 leaves. The middle leaves from three transgenic individuals were used to explore the expression levels of ClCRY2, circadian clock-related genes41, ClCOL genes35 and ClFT (ClFT1 and ClFT2) genes34 after 8 days of SD treatment. The leaves were harvested at the time point at which peak levels of circadian clock-related genes occurred based on the expression pattern of circadian clock-related genes in C. lavandulifolium41. The leaves were harvested at ZT7 (Zeitgeber time 7 h) to explore the expression levels of ClELF3, ClELF4, ClPRR73, and ClPRR37; the leaves were harvested at ZT1 to explore the expression levels of ClZTL and ClLHY; the leaves were harvested at ZT10 to explore the expression levels of ClFKF1, ClPRR5, ClGI-1, and ClGI-2; the leaves were harvested at ZT13 to explore the expression levels of ClPRR1; the leaves were harvested at ZT0 to explore the expression levels of ClCOL1, ClCOL2, and ClCRY2; the leaves were harvested at ZT2 to explore the expression levels of ClCOL4, ClCOL5, and ClFT1. The C. lavandulifolium ClMTP gene was used as an endogenous control. The primers are listed in Supplementary Table S1.
For the flowering time analysis, WT and ClCRY2-OE C. lavandulifolium (>24 per line) were grown at 20 ± 1 °C under SD conditions after they produced 14 leaves. The flowering time of WT and ClCRY2-OE chrysanthemums was determined by counting the number of days from SD treatment to the date when the first flower opened. Three biological replicates were assayed for the WT and transgenic chrysanthemums.
Results
ClCRY2 is specifically upregulated by inductive SD photoperiods
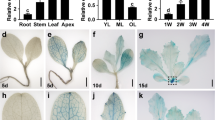
FT/TFL1 family members play crucial roles in regulating floral transition in numerous higher plants1. Morphological changes in the shoot apex and the expression patterns of FT/TFL1-related genes in C. lavandulifolium were analyzed to investigate the effects of the SD photoperiod (12 h L/ 12 h D) on floral transition. The results showed that the shoot apex changed from a conical shape to a dome shape from 0 to 8 d of SD treatment (Fig. 1a). Meanwhile, ClFT1 mRNA levels gradually accumulated, while ClFT2 mRNA levels gradually decreased from 0 to 8 d of SD treatment (Fig. 1b). Total RNA was extracted from various tissues/organs to examine ClCRY2 expression patterns, and the results showed that ClCRY2 mRNA was highest in the leaves during the C. lavandulifolium floral transition (Fig. 1c). We subsequently investigated the effects of SD on the expression patterns of ClCRY2 genes in the leaves. The results showed that ClCRY2 mRNA abundance was markedly increased with SD treatment and peaked at 8 d of SD (Fig. 1b). ClCRY2 was more highly expressed in the leaves under inductive SD (12 h L/12 h D) conditions compared with non-inductive LD (16 h L/ 8 h D) conditions (Fig. 1d).
a The shoot apex development status of C. lavandulifolium under different SD duration conditions (all bar sizes = 0.25 μm). b The expression patterns of ClFTs and ClCRY2 genes in leaves from C. lavandulifolium. c ClCRY2 expression levels in different tissues/organs during C. lavandulifolium floral transition. R roots, S stems, L the middle leaves, P petioles and Sa shoot apices. d ClCRY2 expression patterns in leaves under LD (16 h L/ 8 h D) and SD (12 h L/ 12 h D) conditions. The error bars represent the standard deviations of the data acquired from three biological replicates. The transcription levels of genes in various tissues/organs and in samples under different SD treatment days were normalized against ClSAND. The transcription levels of genes in samples under LD and SD conditions were normalized against ClMTP expression
ClCRY2 belongs to the CRY2 clade
The ClCRY2 gene was isolated from C. lavandulifolium leaves using RT-PCR and 3′-RACE. ClCRY2 contained a 1839 bp open reading frame (ORF) that encoded a 612 amino-acid residue peptide with a calculated molecular mass of 69.8 kDa and a theoretical isoelectric point of 5.78 (Accession Number KJ463737). Different CRY2 proteins are distinguished by their C-terminal sequences, which contain a DQMVP-E/D-STAESS (DAS) domain; the DAS domain contains a DQXVP motif, a short acidic motif and a STAESS motif (Fig. 2a, b). Sequence analysis showed that the STAESS motif in the C-terminal region of ClCRY2 was replaced by STAVSS (Fig. 2a). Additionally, ClCRY2 also contained three conserved motifs of CRYs, including a TGYP motif16, a WRWG motif27 and an LLDAD motif15 (Fig. 2a). CRY2 proteins are ubiquitous in monocots and dicots. Subsequence phylogenetic tree analysis revealed the evolutionary relationship between ClCRY2 and other CRY2 proteins. The results showed that all the CRY2 proteins were classified into the following two groups: (1) the dicotyledonous group (e.g., A. thaliana, Cardamine alpine, Glycine max, Medicago truncatula, and Pisum sativum) and (2) the monocotyledonous group (e.g., Oryza sativa and Triticum aestivum). Species from related families clustered together. For instance, CRY2 proteins from Glycine max, Medicago truncatula and Pisum sativum clustered into Leguminosae. ClCRY2 was related to CRY2 proteins from Cruciferae plants and was distantly related to CRY2 from rice and wheat (Fig. 2c).
a Amino-acid sequence alignment of the CRY2 protein from A. thaliana, G. max and Pisum sativum. Identical residues are highlighted by black boxes. Black lines above the sequences indicate the DAS domain located in the C-terminal region. Black rectangles and black dots represent the residues that interact with FAD and MTHF, respectively. Red lines under the sequences indicate the TGYP, WRWK and LLDAD motifs. b DAS domain sequence logos. c The phylogenetic relationship among CRY2 proteins. The phylogenetic tree is constructed with MEGA 4.0 using the neighbor-joining method. ClCRY2 is denoted by black triangles. At A. thaliana; Ca Cardamine alpine, Gm G. max, Mt Medicago truncatula, Os Oryza sativa, Ps Pisum sativum and Ta Triticum aestivum
ClCRY2 acts as a floral promoter
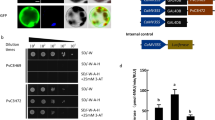
CRY2 can regulate the floral transition of higher plants18,27. To further confirm the function of ClCRY2 in the regulation of flowering, we overexpressed the ClCRY2 gene in WT Arabidopsis. Two T3 generation lines were randomly chosen to test the ClCRY2 expression levels (Fig. 3a) and to explore the role of ClCRY2. The results confirmed the presence of ClCRY2 gene in two ClCRY2-OE lines (Fig. 3a). Under LD conditions, WT plants required 33.8 days from sowing to flowering (Fig. 3b) and had similar rosette leaves with ClCRY2-OE lines (Fig. 3c). In contrast, the ClCRY2-OE showed earlier flowering, which typically required 13.9 to 17.3 days from sowing to flowering (Fig. 3b, d). Under SD1 conditions, the WT plants had more rosette leaves than ClCRY2-OE and required 44.6 days from sowing to flowering (Fig. 3b, c), while ClCRY2-OE required 24.3 to 26.2 days from sowing to flowering (Fig. 3b, e). These results indicated that ClCRY2 overexpression promoted flowering time in Arabidopsis.
a,The transcription levels of ClCRY2 in two transgenic lines. b, c The flowering time and rosette leaf number of WT Arabidopsis and T3 transgenic lines carrying 35 S::ClCRY2 under LD (16 h L/ 8 h D) and SD1 (8 h L/ 16 h D) conditions. d, e The flowering phenotypes of WT Arabidopsis and T3 transgenic lines carrying 35 S::ClCRY2 under LD (16 h L/ 8 h D) and SD1 (8 h L/ 16 h D) conditions. The error bars indicate the standard deviation of the data from three biological replicates. Different lowercase letters in columns indicate a significant difference between two columns according to F test in an analysis of variance (ANOVA) at the 5% level. The transcription levels of genes in WT and two T3 transgenic lines were normalized against TUB2
ClCRY2 acts as an upstream activator of ClFT1 by regulating the transcription of some circadian clock genes
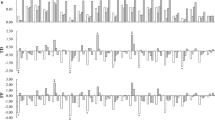
qRT-PCR was conducted in WT and transgenic chrysanthemums to confirm the expression levels of ClCRY2; the results confirmed that we obtained ClCRY2-OE chrysanthemums (Fig. 4a). The flowering phenotype analysis showed that the flowering time was significantly promoted in the two ClCRY2-OE lines compared with WT plants under SD conditions (Fig. 4b, c). WT plants required 48.8 days from SD treatment to flowering, while ClCRY2-OE lines typically required 33.2–37.8 days from SD treatment to flowering (Fig. 4c). The flowering time of the WT and transgenic lines correlated with ClCRY2 mRNA accumulation (Fig. 4a, c). In contrast, WT and two ClCRY2-OE lines did not flower under LD conditions (data not shown). Compared with WT plants, the ClCRY2-OE lines exhibited enhanced ClFT1 mRNA accumulation (Fig. 4d). These results demonstrated that ClCRY2-OE lines could rapidly flower and showed the upregulation of ClFT1 expression under SD conditions. To further explore the mechanism by which ClCRY2 mediates SD signals to promote ClFT1 transcript, we examined the expression levels of circadian clock-related genes and ClCOLs in WT plants and ClCRY2-OE plants. Compared with WT plants, the transcription level of ClELF3 and ClELF4 in input pathway was decreased in ClCRY2-OE plants (Fig. 5a). Some oscillator components, including PSEUDO-REPONSE REGULATOR homologous gene (ClPRR5), ClZTL and ClFKF1, were expressed at higher levels in ClCRY2-OE plants than in WT plants (Fig. 5c, d, g). In contrast, the transcript abundance of the other oscillator components, such as LATE ELONGATED HYPOCOTYL homologous gene (ClLHY), ClPRR73 and REVEILLE8 homologous gene (ClRVE8), decreased in transgenic plants (Fig. 5e, i, j). In the output pathway of the circadian clock, the expression levels of ClGI-1 and ClGI-2 were upregulated in transgenic plants (Fig. 5k, l). In addition, the abundance of ClCOL1, ClCOL4, and ClCOL5 transcripts was also increased in transgenic plants (Fig. 5m, o, p).
a ClCRY2 expression levels in WT and transgenic plants. b The flowering response of WT and transgenic lines under SD conditions (12 h L/ 12 h D). c The flowering time of WT and transgenic lines under SD conditions (12 h L/ 12 h D). d ClFT1 expression levels in WT and transgenic lines under SD conditions (12 h L/ 12 h D). The error bars represent the standard deviation of the data from three biological replicates. Different lowercase letters in columns indicate a significant difference between two columns according to F test in an analysis of variance (ANOVA) at the 5% level. The expression levels of genes in WT and transgenic lines were normalized against ClMTP
Note: a-p represent the expression levels of ClELF3, ClELF4, ClLHY, ClPRR1, ClPRR5, ClPRR37, ClPRR73, ClZTL, ClFKF1, ClRVE8, ClGI-1, ClGI-2, ClCOL1, ClCOL2, ClCOL4, and ClCOL5 respectively. The error bars represent the standard deviation of the data acquired from three biological replicates. The transcription levels of genes in WT and two transgenic lines were normalized against ClMTP. Different lowercase letters in columns indicate a significant difference between two columns according to F test in an analysis of variance (ANOVA) at the 5% level
Discussion
ClCRY2 is involved in the floral transition of C. lavandulifolium induced by SD photoperiods
To date, two homologous FT/TFL1 genes in C. lavandulifolium, ClFT1 and ClFT2, have been identified. It is confirmed that ClFT1 encodes a florigen protein34 and ClFT2 is a CsAFT homologue that encodes an Anti-florigenic FT/TFL1 family protein in Chrysanthemum seticuspe42 (unpublished data, the sequence of ClFT2 can be seen in Supplementary file). In this paper, the C. lavandulifolium shoot apex changed from a conical shape to a dome shape during 0–8 d of SD (12 h L/12 h D) treatment (Fig. 1a), which inferred that C. lavandulifolium completed the developmental transition from vegetative growth to reproductive growth from 0 to 8 SD days. Simultaneously, ClFT1 and ClCRY2 mRNA exhibited an increasing trend from 0 to 8 d of SD treatment; however, ClFT2 was downregulated at the same time (Fig. 1b), indicating that ClCRY2 genes may be involved in the floral transition in C. lavandulifolium by regulating ClFT1 transcription levels under SD conditions. Moreover, ClCRY2 mRNA obeyed diurnal rhythm expression patterns and more ClCRY2 transcripts were induced under inductive SD conditions compared with the non-inductive LD conditions after 8 days of treatment (Fig. 1d). The light-related cis-acting elements and circadian elements (Supplementary Table S2) in the promoter justified the SD-inducible expression patterns of ClCRY2. These results confirmed that ClCRY2 specifically recognized the inductive SD photoperiod signals to regulate the floral transition in C. lavandulifolium.
ClCRY2 enhanced the floral sensitivity to SD photoperiods in C. lavandulifolium by inducing the expression of some circadian clock-related genes and ClFT1
CRYs can regulate the floral transition by affecting the transcription of the circadian clock-related genes GI and FKF132,43. However, the mechanism by which CRYs regulate floral transition through circadian clock genes remains unclear. Conceptually, the circadian clock system comprises the following three major parts: (1) a central oscillator, (2) input pathways integrating oscillator function with light surrounding cues, and (3) output pathways that control the developmental transition44. ELF3 and ELF4 in the input pathway repress floral transition by inhibiting the expression of CO and FT45,46,47,48. LHY, an important oscillator component that encodes a MYB transcription factor49,50,51, acts as a floral inhibitor52. The PRR family contains five members, including PRR1 (TOC1), PRR3, PRR5, PRR7, and PRR9. The toc1 mutant exhibits an early-flowering phenotype under SD conditions53,54, indicating that TOC1 is a floral repressor. The ZTL family, another vital component in the oscillator system, comprises ZTL, FKF1 and LKP255, and they act as floral inducers56. RVE8 acts as a critical regulator of circadian clock and a floral repressor in Arabidopsis57. GI plays a vital role in the output pathway58, promotes flowering through the CO-FT pathway59, and also directly induces the expression of FT via a CO-independent pathway60.
The circadian clock-related genes ELF3, FKF1, and GI are involved in the CRY-COP1/ELF3-FKF1/GI pathway32,43. CRY proteins can repress the activity of the COP1/ELF3 complex and lead to GI accumulation, which results in the formation of the FKF1/GI complex. The FKF1/GI complex can promote CO accumulation by inducing the degradation of CDF proteins, which are CO repressors32,43. In the CRY2-CIBs pathway, the CRY2/CIBs complex binds to an E-box (CANNTG) in the FT promoter and induces its transcription. ZTL can stabilize the activity of CIBs and lead to flowering in this process31,56. Compared with WT C. lavandulifolium, ClCRY2-OE C. lavandulifolium exhibited increased SD sensitivity and an early flowering phenotype (Fig. 4c). ClFT1 transcription levels were upregulated in ClCRY2-OE C. lavandulifolium compared with the WT plants (Fig. 4d). The results inferred that ClCRY2 acts as a floral inducer by activating ClFT1 transcription. A previous study showed that the induction of homologous FT genes results from the integration of photoperiodic information with diurnal timing set by the plant’s circadian clock61. Therefore, we tested the changes in the expression patterns of circadian clock-related genes and ClCOLs. The results revealed that the expression levels of ClPRR5, ClZTL, ClFKF1, ClGI-1, ClGI-2, ClCOL1, ClCOL4, and ClCOL5 were also upregulated in ClCRY2-OE C. lavandulifolium compared with the WT plants (Fig. 5). Taken together, ClCRY2 promoted floral sensitivity to SD photoperiods in C. lavandulifolium by inducing the transcription of circadian clock genes, ClCOLs and ClFT1.
ClCRY2 might serve as an important gene resource for the accurate manipulation of flowering time in chrysanthemums
The flowering time of chrysanthemums can be controlled by the application of artificial SD conditions during annual production. However, this method of manipulating flowering leads to a substantial waste in manpower and material resources. The primary factor influencing adult chrysanthemum flowering time is sensitivity to SD photoperiods. Chrysanthemum cultivars, which rapidly respond to SD photoperiods, could save costs in the process of practical production. Therefore, it is vital to explain the molecular mechanism by which chrysanthemums respond to SD signals to complete the floral transition. Our results demonstrated that the ClCRY2 gene enhanced floral sensitivity to SD photoperiods in C. lavandulifolium. Therefore, it is feasible to manipulate the ClCRY2 gene to breed new chrysanthemum cultivars that can flower quickly under inductive SD conditions.
Photoreceptors may mediate light quality signal to regulate flowering in SDPs
Light quality is another parameter of ambient light signals except for photoperiod. Light quality also regulates the flowering response. For instance, blue light promotes flowering in Arabidopsis; while red light plays an opposite role62. Similar results were shown in chrysanthemum. Flowering is significantly inhibited once the night phase is interrupted by red light42,63,64,65,66,67,68. End-of-day red light inhibits FvTFL1 expression and induces flowering in SDP strawberry (Fragaria vesca)69. To date, the molecular mechanism by which light quality regulates flowering in LDP Arabidopsis has been deeply clarified28,29,30,31,32,70. However, the molecular mechanism by which light quality regulates flowering in SDPs remains unanswered. OsCRY2 promotes the flowering in rice18; blue light could hasten flowering of rice through upregulating Ehd1 expression via OsGI-dependent pathway71. Therefore, we wonder that whether OsCRY2 could mediate blue light to regulate the transcription of Ehd1 through affecting the expression of OsGI. In SDP chrysanthemum C. seticuspe, CsPHYB could mediate red light to suppress flowering by regulating the expression of CsAFT42. We wonder whether CsPHYB regulates CsAFT expression via the direct pathway or CsPHYB indirectly regulates CsAFT expression through other components.
In conclusion, we illustrated a model depicting ClCRY2-mediating SD signals to regulate floral transition in C. lavandulifolium. ClCRY2 in leaves can respond to SD signals. It upregulates some circadian clock genes (ClPRR5, ClZTL, ClFKF1, ClGI-1, and ClGI-2), downregulates other circadian clock genes (ClELF3, ClELF4, ClLHY, ClPRR73, and ClRVE8), induces the expression of ClCOLs and ClFT1, and finally leads to floral transition in C. lavandulifolium (Fig. 6). Our results indicate that ClCRY2 might serve as an important gene resource used for breeding new chrysanthemum cultivars that flower quickly under inductive SD conditions.
References
Song, Y. H., Shim, J. S., Kinmonth-Schultz, H. A. & Imaizumi, T. Photoperiodic flowering: time measurement mechanisms in leaves. Annu. Rev. Plant. Biol. 66, 441–464 (2015).
Chen, A. et al. PHYTOCHROME C plays a major role in the acceleration of wheat flowering under long-day photoperiod. Proc. Natl Acad. Sci. USA 111, 10037–10044 (2014).
Gao, H. et al. Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc. Natl Acad. Sci. USA 111, 16337–16342 (2014).
Ishikawa, R. et al. Phytochrome B regulates Heading date 1 (Hd1)-mediated expression of rice florigen Hd3a and critical day length in rice. Mol. Genet. Genom. 285, 461–470 (2011).
Cashmore, A. R. Cryptochromes: enabling plants and animals to determine circadian time. Cell 114, 537–543 (2003).
Yu, X., Liu, H., Klejnot, J. & Lin, C. The cryptochrome blue light receptors. The Arabidopsis Book 8, (e0135 (2010).
Ito, S., Song, Y. H. & Imaizumi, T. LOV domain-containing F-box proteins: light-dependent protein degradation modules in Arabidopsis. Mol. Plant 5, 573–582 (2012).
Briggs, W. R. & Huala, E. Blue-light photoreceptors in higher plants. Annu. Rev. Cell Dev. B 15, 33–62 (1999).
Quail, P. H. et al. Phytochromes: photosensory perception and signal transduction. Science 268, 675 (1995).
Smith, H. Phytochromes and light signal perception by plants-an emerging synthesis. Nature 407, 585–591 (2000).
Rizzini, L. et al. Perception of UV-B by the Arabidopsis UVR8 protein. Science 332, 103–106 (2011).
Lin, C. & Todo, T. The cryptochromes. Genome Biol. 6, 220 (2005).
Lin, C. & Shalitin, D. Cryptochrome structure and signal transduction. Annu. Rev. Plant. Biol. 54, 469–496 (2003).
Sancar, A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 103, 2203–2238 (2003).
Lin, C. et al. Association of flavin adenine dinucleotide with the Arabidopsis blue light receptor CRY1. Science 269, 968 (1995).
Malhotra, K., Kim, S. T., Batschauer, A., Dawut, L. & Sancar, A. Putative blue-light photoreceptors from Arabidopsis thaliana and Sinapis alba with a high degree of sequence homology to DNA photolyase contain the two photolyase cofactors but lack DNA repair activity. Biochem 34, 6892–6899 (1995).
Guo, H., Yang, H., Mockler, T. C. & Lin, C. Regulation of flowering time by Arabidopsis photoreceptors. Science 279, 1360–1363 (1998).
Hirose, F., Shinomura, T., Tanabata, T., Shimada, H. & Takano, M. Involvement of rice cryptochromes in de-etiolation responses and flowering. Plant Cell Physiol. 47, 915–925 (2006).
Li, Y. Y. et al. Molecular cloning of cryptochrome 1 from apple and its functional characterization in Arabidopsis. Plant Physiol. Bioch. 67, 169–177 (2013a).
Li, Y. Y. et al. Molecular cloning and functional analysis of a blue light receptor gene MdCRY2 from apple (Malus domestica). Plant Cell Rep. 32, 555–566 (2013b).
Giliberto, L. et al. Manipulation of the blue light photoreceptor cryptochrome 2 in tomato affects vegetative development, flowering time, and fruit antioxidant content. Plant Physiol. 137, 199–208 (2005).
Platten, J. D. et al. Cryptochrome 1 contributes to blue-light sensing in pea. Plant Physiol. 139, 1472–1482 (2005a).
Banerjee, R. et al. The signaling state of Arabidopsis cryptochrome 2 contains flavin semiquinone. J. Biol. Chem. 282, 14916–14922 (2007).
Bouly, J. P. et al. Cryptochrome blue light photoreceptors are activated through interconversion of flavin redox states. J. Biol. Chem. 282, 9383–9391 (2007).
Selby, C. P. & Sancar, A. A. cryptochrome/photolyase class of enzymes with single-stranded DNA-specific photolyase activity. Proc. Natl Acad. Sci. USA 103, 17696–17700 (2006).
Liscum, E., Hodgson, D. W. & Campbell, T. J. Blue light signaling through the cryptochromes and phototropins. So that’s what the blues is all about. Plant Physiol. 133, 1429–1436 (2003).
Ahmad, M. & Cashmore, A. R. HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature 366, 162 (1993).
Liu, H., Liu, B., Zhao, C., Pepper, M. & Lin, C. The action mechanisms of plant cryptochromes. Trends Plant. Sci. 16, 684–691 (2011).
Zuo, Z., Liu, H., Liu, B., Liu, X. & Lin, C. Blue light-dependent interaction of CRY2 with SPA1 regulates COP1 activity and floral initiation in Arabidopsis. Curr. Biol. 21, 841–847 (2011).
Sarid-Krebs, L. et al. Phosphorylation of CONSTANS and its COP1-dependent degradation during photoperiodic flowering of Arabidopsis. Plant J. 84, 451–463 (2015).
Liu, H. et al. Photoexcited CRY2 interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science 322, 1535–1539 (2008).
Sawa, M., Nusinow, D. A., Kay, S. A. & Imaizumi, T. FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 318, 261–265 (2007).
Lin, C. Blue light receptors and signal transduction. Plant Cell 14, S207–S225 (2002). suppl 1.
Fu, J. X. et al. Photoperiodic control of FT-like gene ClFT initiates flowering in Chrysanthemum lavandulifolium. Plant Physiol. Bioch. 74, 230–238 (2014a).
Fu, J., Yang, L. & Dai, S. Identification and characterization of the CONSTANS-like gene family in the short-day plant Chrysanthemum lavandulifolium. Mol. Genet. Genom. 290, 1039–1054 (2015).
Yang, L., Fu, J., Qi, S. & Dai, S. Molecular cloning and function analysis of ClCRY1a and ClCRY1b, two genes in Chrysanthemum lavandulifolium that play vital roles in promoting floral transition. Gene. 617, 32–43 (2017).
Wang, Y. et al. Construction and de novo characterization of a transcriptome of Chrysanthemum lavandulifolium: analysis of gene expression patterns in floral bud emergence. Plant Cell Tiss. Org. 116, 297–309 (2014).
Fu, J. X., Wang, Y., Huang, H., Zhang, C. & Dai, S. L. Reference gene selection for RT-qPCR analysis of Chrysanthemum lavandulifolium during its flowering stages. Mol. Breed. 31, 205–215 (2013).
Wang, J., Han, K. & Dai, S. Construction of expression vector and transformation of chrysanthemum with maize Lc gene. Genom. Appl. Biol. 28, 229–236 (2009).
Qi, S., Fu, J. X., Wang, Y., Yang, L. & Dai, S. Establishment of genetic trasformation system for Chrysanthemum lavandulifolium. Mol. Plant Breed. 12, 356–362 (2014).
Fu, J., Yang, L. & Dai, S. Conservation of Arabidopsis thaliana circadian clock genes in Chrysanthemum lavandulifolium. Plant Physiol. Bioch. 80, 337–347 (2014b).
Higuchi, Y. et al. The gated induction system of a systemic floral inhibitor, antiflorigen, determines obligate short-day flowering in chrysanthemums. Proc. . Natl Acad. Sci. USA 110, 17137–17142 (2013).
Fornara, F. et al. Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Dev. Cell. 17, 75–86 (2009).
Hsu, P. Y. & Harmer, S. L. Wheels within wheels: the plant circadian system. Trends Plant. Sci. 19, 240–249 (2014).
Hicks, K. A., Albertson, T. M. & Wagner, D. R. EARLY FLOWERING3 encodes a novel protein that regulates circadian clock function and flowering in Arabidopsis. Plant Cell 13, 1281–1292 (2001).
McWatters, H. G., Bastow, R. M., Hall, A. & Millar, A. J. The ELF3 zeitnehmer regulates light signalling to the circadian clock. Nature 408, 716–720 (2000).
Eriksson, M. E. & Millar, A. J. The circadian clock. A plant’s best friend in a spinning world. Plant Physiol. 132, 732–738 (2003).
McWatters, H. G. et al. ELF4 is required for oscillatory properties of the circadian clock. Plant Physiol. 144, 391–401 (2007).
Harmer, S. L. The circadian system in higher plants. Annu. Rev. Plant. Biol. 60, 357–377 (2009).
Mas, P. & Yanovsky, M. J. Time for circadian rhythms: plants get synchronized. Curr. Opin. Plant. Biol. 12, 574–579 (2009).
Imaizumi, T. Arabidopsis circadian clock and photoperiodism: time to think about location. Curr. Opin. Plant. Biol. 13, 83–89 (2010).
Mizoguchi, T. et al. LHY and CCA1 are partially redundant genes required to maintain circadian rhythms in Arabidopsis. Dev. Cell. 2, 629–641 (2002).
Kreps, J. A. & Simon, A. E. Environmental and genetic effects on circadian clock-regulated gene expression in Arabidopsis. Plant Cell 9, 297–304 (1997).
Somers, D. E., Webb, A. A., Pearson, M. & Kay, S. A. The short-period mutant, toc1-1, alters circadian clock regulation of multiple outputs throughout development in Arabidopsis thaliana. Development 125, 485–494 (1998).
Suetsugu, N. & Wada, M. Evolution of three LOV blue light receptor families in green plants and photosynthetic stramenopiles: phototropin, ZTL/FKF1/LKP2 and aureochrome. Plant Cell Physiol. 54, 8–23 (2013).
Liu, H. et al. Arabidopsis CRY2 and ZTL mediate blue-light regulation of the transcription factor CIB1 by distinct mechanisms. Proc. Natl Acad. Sci. USA 110, 17582–17587 (2013).
Farinas, B. & Mas, P. Functional implication of the MYB transcription factor RVE8/LCL5 in the circadian control of histone acetylation. Plant J. 66, 318–329 (2011).
Fowler, S. et al. GIGANTEA: a circadian clock-controlled gene that regulates photoperiodic flowering in Arabidopsis and encodes a protein with several possible membrane-spanning domains. EMBO J. 18, 4679–4688 (1999).
Mizoguchi, T. et al. Distinct roles of GIGANTEA in promoting flowering and regulating circadian rhythms in Arabidopsis. Plant Cell 17, 2255–2270 (2005).
Sawa, M. & Kay, S. A. GIGANTEA directly activates Flowering Locus T in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 108, 11698–11703 (2011).
Brambilla, V. et al. Antagonistic transcription factor complexes modulate the floral transition in Rice. Plant Cell 29, 2801–2816 (2017). tpc. 00645.
Eskins, K. Light-quality effects on Arabidopsis development. Red, blue and far-red regulation of flowering and morphology. Physiol. Plant 86, 439–444 (1992).
Shrestha, R., Gómez-Ariza, J., Brambilla, V. & Fornara, F. Molecular control of seasonal flowering in rice, arabidopsis and temperate cereals. Ann. Bot. 114, 1445–1458 (2014).
Yano, M. et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12, 2473–2483 (2000).
Hayama, R., Yokoi, S., Tamaki, S., Yano, M. & Shimamoto, K. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 422, 719 (2003).
Kojima, S. et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol. 43, 1096–1105 (2002).
Liu, B. et al. Signaling mechanisms of plant cryptochromes in Arabidopsis thaliana. J. Plant Res. 129, 137–148 (2016).
Higuchi, Y., Sumitomo, K., Oda, A., Shimizu, H. & Hisamatsu, T. Day light quality affects the night-break response in the short-day plant chrysanthemum, suggesting differential phytochrome-mediated regulation of flowering. J. Plant Physiol. 169, 1789–1796 (2012).
Rantanen, M. et al. Light quality regulates flowering in FvFT1/FvTFL1 dependent manner in the woodland strawberry Fragaria vesca. Front. Plant Sci. 5, 271 (2014).
Franklin, K. A. & Quail, P. H. Phytochrome functions in Arabidopsis development. J. Exp. Bot. 61, 11–24 (2009).
Itoh, H., Nonoue, Y., Yano, M. & Izawa, T. A pair of floral regulators sets critical day length for Hd3a florigen expression in rice. Nat. Genet. 42, 635 (2010).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31530064 and 31471907).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
National Natural Science Foundation of China (31530064) and National Natural Science Foundation of China (31471907).
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Lw., Wen, Xh., Fu, Jx. et al. ClCRY2 facilitates floral transition in Chrysanthemum lavandulifolium by affecting the transcription of circadian clock-related genes under short-day photoperiods. Hortic Res 5, 58 (2018). https://doi.org/10.1038/s41438-018-0063-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-018-0063-9
This article is cited by
-
Genome-wide identification and characterization analysis of RWP-RK family genes reveal their role in flowering time of Chrysanthemum lavandulifolium
BMC Plant Biology (2023)
-
Functional diversification and molecular mechanisms of FLOWERING LOCUS T/TERMINAL FLOWER 1 family genes in horticultural plants
Molecular Horticulture (2022)
-
Small-scale alpine topography at low latitudes and high altitudes: refuge areas of the genus Chrysanthemum and its allies
Horticulture Research (2020)