Abstract
Characterizing the role of different mutational effect sizes in the evolution of fitness-related traits has been a major goal in evolutionary biology for a century. Such characterization in a diversity of systems, both model and non-model, will help to understand the genetic processes underlying fitness variation. However, well-characterized genetic architectures of such traits in wild populations remain uncommon. In this study, we used haplotype-based and multi-SNP Bayesian association methods with sequencing data for 313 individuals from wild populations to test the mutational composition of known candidate regions for sea age at maturation in Atlantic salmon (Salmo salar). We detected an association at five loci out of 116 candidates previously identified in an aquaculture strain with maturation timing in wild Atlantic salmon. We found that at four of these five loci, variation explained by the locus was predominantly driven by a single SNP suggesting the genetic architecture of this trait includes multiple loci with simple, non-clustered alleles and a locus with potentially more complex alleles. This highlights the diversity of genetic architectures that can exist for fitness-related traits. Furthermore, this study provides a useful multi-SNP framework for future work using sequencing data to characterize genetic variation underlying phenotypes in wild populations.
Similar content being viewed by others
Introduction
The debate over the role of different mutational effect sizes in the evolution of fitness has been ongoing for a century (Orr 1998; Orr 1999; Fisher 1930; Rockman 2012; Remington 2015; Kimura 1983). Our ability to now sequence large numbers of individuals has greatly informed this debate. Additionally, an explosion in the number of association studies has revealed the architecture of important traits. These have revealed examples of large effect loci (Hoekstra 2006; Linnen et al. 2013; Tishkoff et al. 2007; Daborn et al. 2002), but also many cases of polygenicity (Purcell et al. 2009; Loh et al. 2015; Pritchard and Di Rienzo 2010). However, even where large effect alleles are known, these can have been generated over time via sequential mutations of smaller effect that allow a slow walk to the optimum as predicted by Orr (1998), or a single mutation of large effect. The difference between these scenarios is important for our understanding of the process of adaptation, the adaptability of organisms and their resilience to rapid environmental change (Oomen et al. 2020; Kardos and Luikart 2021; Yeaman et al. 2018).
Among genome-wide association studies published to date, many complex traits appear to be polygenic (Visscher et al. 2017; Fisher 1918; Pritchard and Di Rienzo 2010) or omnigenic (i.e. affected by a large proportion of genes through a network of core genes and many peripheral genes that modify the effects of core genes, whereby a large proportion of all genes expressed in trait-related tissues has some effect on the trait) (Boyle et al. 2017; Liu et al. 2019). Although polygenicity is widespread, an increasing number of examples of major effect loci exist, whereby one locus explains a large proportion of the phenotypic variation (Barson et al. 2015; Linnen et al. 2013). In some cases, major effect loci can contain multiple tightly linked genes, coined “supergenes”, where localized reduction in recombination is often caused by larger chromosomal rearrangements. For example, this phenomenon is known to underlie phenotypic variation observed among ruff (Philomachus pugnax) mating morphs (Lamichhaney et al. 2015; Küpper et al. 2015), Atlantic cod (Gadus morhua) (Kirubakaran et al. 2016; Sinclair-Waters et al. 2018) and rainbow trout (Oncorhynchus mykiss) migratory ecotypes (Pearse et al. 2018), and Heliconius butterfly wing-pattern morphs (Joron et al. 2011). More recent work has found that major effect loci can exist alongside a polygenic background where loci with a variety of effect sizes underlie trait variation (Sinnott-Armstrong et al. 2020; Sinclair-Waters et al. 2020). Such mixed genetic architectures may be pervasive, but currently remain undetected due to the large sample sizes required for detecting loci with smaller effects (Sinclair-Waters et al. 2020) and it is possible that additional examples are to be found with future higher-powered studies. Although studies aimed at resolving genotype-phenotype links are mounting, well-characterized genetic architectures of fitness-related traits, particularly in natural populations, are still uncommon.
While some trait-associated loci have been identified, such findings lead to other crucial questions: How have trait-locus associations arisen? Has the locus arisen through a single or multiple new mutation(s)? Or alternatively, did the locus emerge via recombination that gave rise to new combinations of existing variants? Numerous studies from the past decade have shown that major effect loci involve the cumulative effects of multiple mutations, rather than a single mutation, thus highlighting the relevance of considering the latter scenarios. For example, Bickel et al. (2011) found that ~60% of variation in female abdominal pigmentation in Drosophila melanogaster can be explained by sequence variation at the bab locus, but a GWAS (genome-wide association study) analyzing the same trait did not identify a single SNP in bab that passed the genome-wide significance threshold. Alleles consisting of multiple SNPs were associated with high proportions of the variation, whereas single SNPs had only small effects and were therefore missed in the single-SNP GWAS. Additionally, Linnen at al. (2013) and Kerdaffrec et al. (2016) also identify multiple mutations within a confined region that have cumulative effects on colour traits in deer mice and seed dormancy in Arabidopsis thaliana, respectively. In natural populations with gene flow such as in Linnen et al. (2013) and Kerdaffrec et al. (2016), this is perhaps not unexpected as theory predicts that clustered and major effect loci will evolve under such scenarios (Yeaman and Whitlock 2011; Yeaman 2013). Given these findings, examining extended sequence haplotypes containing multiple SNPs, rather than each SNP independently, is important (Remington 2015). This can be achieved by using alternative strategies that look at combined effects of variants, rather than single-SNP methods typically used in GWAS.
Here we investigate the genetic basis of Atlantic salmon (Salmo salar) sea age at maturity—the number of years spent in the marine environment before reaching maturity and returning to the natal river (freshwater) to reproduce. Atlantic salmon individuals can spend anywhere from one to five years in the marine environment before maturation occurs. Prior to this marine phase, individuals spend one to seven years in their natal river before migrating to the sea. Moreover, some individuals will reach maturity in freshwater without ever having migrated to the sea, known as mature parr (Fleming 1996; Mobley et al. 2021). This variation in maturation timing contributes substantially to the diversity of life history strategies among Atlantic salmon (Erkinaro et al. 2019). Age at maturity varies both within and among Atlantic salmon populations, with multiple maturation age classes commonly occurring within single populations (Barson et al. 2015; Jonsson et al. 1991; Hutchings and Jones 1998). Furthermore, age at maturity is an important life history trait affecting fitness traits such as survival, size at maturity and reproductive success (Stearns 2000; Mobley et al. 2021). Substantial variation in Atlantic salmon sea age at maturity is maintained due to a trade-off between mating success at spawning grounds and survival, whereby individuals that mature later are larger and have higher reproductive success on the spawning grounds, but have a lower chance of surviving until reproductive age due to a high mortality in the marine environment (Chaput 2012). In contrast individuals that mature early are smaller and have lower reproductive success, but higher survival and thus higher chance of reaching reproductive age (Fleming and Einum 2011; Mobley et al. 2020).
Variation in maturation timing in Atlantic salmon is highly heritable (Gjerde 1984; Sinclair-Waters et al. 2020; Reed et al. 2018) and consequently there is substantial interest in understanding the underlying genetic architecture. A large-effect locus on chromosome 25 explaining up to 39% of the variation in sea age at maturity was found in wild European populations (Barson et al. 2015) and domesticated salmon (Ayllon et al. 2015). The primary candidate gene underlying the association of this locus is vgll3 due to its close proximity to the associated SNP variation (Sinclair-Waters et al. 2022; Ayllon et al. 2015; Barson et al. 2015) and its known function in other species. The vgll3 gene encodes a transcription cofactor that, amongst other things, regulates adipogenesis (Halperin et al. 2013) and is associated with variation in puberty timing in humans (Day et al. 2017; Perry et al. 2014). In addition to vgll3, Sinclair-Waters et al. (2020) identified 119 other candidate genes for male maturation in a GWAS using SNP-array data and including >11,000 males from an Atlantic salmon aquaculture strain originating since the 1970s and derived of founder individuals from 41 wild Norwegian rivers (Gjedrem et al. 1991). Two particularly strong associations between maturation timing were found on chromosome 9 in close proximity to six6 and chromosome 25, vgll3. The association of six6 was also found by Barson et al. (2015) in wild Atlantic salmon, but the signal disappeared after correction for the confounding effects of population structure that can increase the false positive rate of association tests (Price et al. 2010). Barson et al. (2015), however, focused solely on single-SNP associations via GWAS without considering the possible influence of combined variant effects. Interestingly, the six6 gene is also associated with age at maturity in two Pacific salmon species (Waters et al. 2021; Willis et al. 2020), humans (Perry et al. 2014) and cattle (Cánovas et al. 2014).
Characterization of genetic architecture for fitness-related traits in a number of organisms, including both model and non-model systems, and for a variety of traits, will help gain a clearer understanding of the processes underlying fitness variation (Stinchcombe and Hoekstra 2008). However, studies using sequencing data to examine variation associated with important fitness-related traits in wild populations are limited. Fortunately, due to developments in sequencing technologies and bioinformatics, studies using this approach are likely to rise in number. We therefore aim to provide a useful and timely framework for characterizing genetic variation underlying phenotypes in wild populations in the future. Here, we focus on characterizing the mutational composition of known candidate regions for sea age at maturity in wild Atlantic salmon found in the previous large sample GWAS, Sinclair-Waters et al. (2020), by testing for combined effects of variants to determine number and location of associated variants. We integrate re-sequencing data and phenotype information for 313 individuals from 53 wild population of Atlantic salmon with alternative GWAS strategies that consider the combined effects of variants, rather than single-SNP effects. This approach can provide better resolution of the variants underlying fitness-related traits when combined effects are involved, while also effective for detecting single SNP effects.
Materials and methods
Study material
Whole genome sequencing data were obtained for 313 wild individuals collected from 53 Norwegian and Finnish populations spanning the Norwegian coast and to the Barents sea in the north (59°N–71°N) (Supplementary Table S1, S2, Fig. S1) previously reported in Bertolotti et al. (2020). The 313-individual dataset includes individuals for which the sea age at maturity phenotype has been recorded, and spans populations belonging to both the Atlantic and Barents/White sea phylogeographic groups. Additionally, sampling locations were chosen to represent populations exhibiting both within and among population variation in sea-age at maturity (Barson et al. 2015). These geographic regions were studied in Barson et al. (2015) using a single SNP approach based on SNP-array data. Whole genome sequencing data allows variants not present on the SNP-array and combined SNP effects to be tested. Scales were collected from individuals and sea age was determined by examining scale growth rings as described in Mobley et al. (2021) and ICES (2011). Individuals were categorized into three maturation categories based on the number of years spent at sea prior to their first return migration to rivers for spawning: 1 (one year spent at sea), 2 (two years spent at sea), or 3 (three or more years spent at sea). Only five individuals had spent four years and were therefore combined with three-year fish for all analyses.
SNP calling and filtering
Variant calling and the first round of filtering was done in a larger set of individuals described in Bertolotti et al. (2020). Raw Illumina reads were mapped to the Atlantic salmon genome (ICSASG_v2) (Lien et al. 2016) using bcbio-nextgen v.1.1 (Chapman et al. 2020) with the bwa-mem aligner v.0.7.17 (Heng Li 2013). Genomic variation was identified using the Genome Analysis Toolkit (GATK) v4.0.3.0., following GATK’s best practice recommendations. Picard v2.18.7 (Picard Toolkit 2019) was used to mark duplicates and GATK was used for joint calling (Depristo et al. 2011). Variants were annotated using SNPeff v. 4.3 (Cingolani et al. 2012). Variant call were further filtered with GATK’s variant filtration according to the following –filterExpression: “MQRankSum < −12.5 || ReadPosRankSum < −8.0 || QD < 2.0 || FS > 60.0 || (QD < 10.0 && AD[0:1] / (AD[0:1] + AD[0:0]) < 0.25 && ReadPosRankSum < 0.0) || MQ < 30.0”. SNPs were then filtered using SNPable procedure (Li 2009), where 100 bp kmers are mapped to reference genome (ICSASG_v2) using Burrows-Wheeler Aligner (bwa aln) (Li and Durbin 2009), and only SNPs within regions with reads that uniquely map are retained. We then removed additional SNPs with vcftools using the following criteria: –min-alleles 2, –max-alleles 2, –maf 0.0000000001, –max-missing 0.7, –remove-indels, –minGQ 10, and –minDP 4. A subset 313 individuals from wild populations was then extracted from this larger dataset using vcftools (Danecek et al. 2011). This reduced dataset was used for all subsequent analyses.
Analysis of principal components used for population structure correction and genetic differentiation
We produced a reduced SNP dataset by pruning one SNP from each SNP pair with a correlation coefficient (r2) greater than 0.2 within a 50 kb block using the –indep-pairwise 50 10 0.2 function implemented in PLINK v1.9 (Purcell et al. 2007). This yielded 403,540 SNPs to examine population structure using a principal component analysis, smartpca, implemented in the EIGENSOFT v5 software (Patterson et al. 2006). Principal components were then used to correct for the confounding effects of population structure during association testing (see below). Additionally, this reduced dataset was used to estimate FST (following Weir and Cockerham (1984)) between the Atlantic and Barents/White sea phylogeographic groups with SNPRelate (Zheng et al. 2012).
Data preparation
In this study, we focus on genomic regions containing the 116 candidate loci for age at maturity identified in Sinclair-Waters et al. (2020) using an Atlantic salmon aquaculture strain. This strain is derived from founder individuals originating from 41 wild Norwegian rivers, some of which were the same rivers sampled for this study, and all belonging to the same phylogeographic lineage as the sequenced individuals used in this study (Gjedrem et al. 1991). We extracted SNP genotype data from 500 kb regions surrounding the 116 trait-associated SNPs identified in Sinclair-Waters et al. (2020) using vcftools (Danecek et al. 2011) position filtering functions –from-bp and –to-bp, as well as allele filtering function –mac 1 to keep only polymorphic sites. Trait-associated SNPs that were within 250 kb of another trait-associated SNP were combined into a single candidate region that extends 250 kb upstream of the more upstream SNP to 250 kb downstream of the more downstream SNP.
The current Atlantic salmon genome (ICSASG_v2) contains a known assembly error within the 500 kb region surrounding the known candidate loci vgll3 (Ayllon et al. 2015). A misplaced and misoriented scaffold currently placed downstream of vgll3 belongs within a gap in the assembly just upstream of vgll3 on ssa25. For this reason, we constructed a revised assembly for this chromosome. SNP calling was performed as described above. We then retained SNPs that had met the filtering criteria. A total of 8 candidate SNPs are located within regions of the genome that were moved. To find the position of these SNPs in the revised chromosome 25 sequence, we extracted 200 bp surrounding each of these SNPs from the current genome assembly (ICSASG_v2) using the getfasta function in BEDTools (Quinlan and Hall 2010). The 200 bp sequence was then blasted to the fixed assembly to determine the new position of each SNP using Blast’s blastn function (Camacho et al. 2009). Using the new SNP positions, SNP genotypes within a 500 kb region surrounding the moved candidate SNPs were extracted from the fixed dataset using vcftools.
Association testing at candidate regions
We applied three association mapping methods to describe the genetic architecture underlying sea age at maturity at each of the candidate regions identified in Sinclair-Waters et al. (2020). First, a multi-SNP approach examining associations between phenotype and haplotypes was conducted using Bayesian linear regression implemented in hapQTLv1.00 (Xu and Guan 2014). In this approach, a hidden Markov model is used to characterize haplotype structure and ancestry (Guan 2014). Haplotype sharing at each marker is then used to quantify genetic similarity among individuals. Haplotype associations are identified by testing for an association between genetic similarity at each marker and the phenotype (Xu and Guan 2014). Each of the extracted vcf files was converted to bimbam format using PLINK 1.9 (Chang et al. 2015). The resulting bimbam files were used as input for hapQTL. Second, single SNP associations were also identified using a Bayesian linear regression method implemented in hapQTL (Guan and Stephens 2011). For all hapQTL association tests, sex and the six most significant principal components (see above) were included as covariates in the models. Each hapQTL run consisted of 2 EM runs (-e 2) with 40 steps (-w 40), 2 upper clusters (-C 2), 10 lower clusters (-c 10). Three replicate hapQTL runs were performed for each of the 116 selected regions. Based on recommendations from Jeffreys (1961), Bayes factors greater than three were considered evidence for an association of either SNPs or haplotype with sea age at maturity phenotype.
Third, a multi-SNP approach aimed to estimate the number and identity of SNPs underlying trait variation at each candidate region using Bayesian Variable Selection regression implemented in PiMASS (Guan and Stephens 2011). Due to computational restrictions, the PiMASS analysis was performed for only candidate regions that had a SNP or haplotype association with Bayes factor greater than 3. Prior to the PiMASS analysis, all missing genotypes were imputed in BIMBAM (Guan and Stephens 2011) as mean genotypes (-wmg) using default settings. Additionally, our phenotype values for sea age at maturity were adjusted to correct for confounding effects of sex and population structure by regressing the phenotype on sex and the six most significant principal components (see above) using the lm function in R. PiMASS was run with the residual phenotype values. We placed priors on the proportion of variance explained by SNP(s) (hmin = 0.001 and hmax = 0.999) and the number of SNPs in the model (pmin = log\(\frac{1}{N}\) and pmax = log\(\frac{{300}}{N}\), where N is the total number of SNPs). Each run consisted of a burn-in of 1000000 steps, followed by 2500000 steps where parameter values were recorded every 1000 steps. For each analysis, we examined the posterior inclusion probability for each SNP, the distribution of the number of included SNPs and the distribution of the proportions of variance explained per model. We also examined the path of estimated Bayes factors and parameter values (h, p, s) across all recorded iterations to check for convergence of runs.
To further assess whether more than one SNP in a candidate region was significantly associated with sea age at maturity, we regressed out the top-associated SNP from the residual phenotype values described above and reran PiMASS using the previously-used priors and settings. We then examined the posterior inclusion probability for each SNP, the distribution of the number of included SNPs, and the distribution of proportion of variance explained to determine whether there was evidence for multiple SNP associations within a given candidate region.
Results
Analysis of principal components and genetic differentiation
The first six principal components (PCs) calculated with the pruned SNP dataset explained 1.96%, 0.68%, 0.63%, 0.59%, 0.56% and 0.51% of the genetic variance, respectively (Supplementary Fig. S2). These six PCs were included in subsequent association analyses to reflect population structure among samples including structure between phylogeographic groups, structuring occurring along a north-south gradient and other finer scale structuring (Supplementary Fig. S2). We do not include any PCs explaining 0.5% or less of the genetic variance. FST between the Atlantic and Barents/White sea phylogeographic groups was 0.02.
Associations identified with hapQTL
Single-SNP and haplotype association analyses with hapQTL revealed strong (Bayes factor > 3) association signals at 5 of the 116 candidate regions (Fig. 1, Supplementary Fig. S3). The strongest association observed within each region was with a single SNP, rather than an extended haplotype, suggesting a single mutation underlies the effect of each of these regions on maturation timing. However, exceptions occurred in the ssa09:24636574-25136574 and ssa25:28389273-28889273 regions, where second association signals were found upstream of the primary association signal and were most strongly linked to an extended haplotype. For instance, strong haplotype association scores (Bayes factor > 3) spanned a 26971 bp region (ssa09:24781742-24808713) containing an uncharacterized gene (LOC106610978) and pcnx4. In the ssa25:28389273-28889273 region, a strong haplotype signal was found within edar (Fig. 1).
We find differences in the location of the top-associated SNPs found here and those identified in Sinclair-Waters et al. (2020). For regions ssa06:27541960-28218141, ssa09:10915066-11415066 and ssa25:28389273-28889273, the top-associated SNP was located further upstream than in Sinclair-Waters et al. (2020). Contrastingly, the strongest associated SNPs within the regions ssa09:24636574-25136574 and ssa21:49390687-49890687 differed only slightly (<5000 bp) between studies (Table 1).
Multi-SNP associations identified using PiMASS
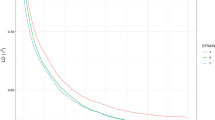
Multi-SNP association analysis with PiMASS showed that at four of five candidate regions, a single-SNP model was most commonly used to explain variation in sea age at maturity. At one candidate region, ssa09:24636574-25136574, a multi-SNP model including two SNPs was most commonly used to explain variation in sea age at maturity. Median proportion of variance explained by each candidate region ranged between 4% and 19% (Fig. 2, Table 2). Additionally, mean sea age at maturity differed substantially among genotypes at all six SNPs selected by the multi-SNP models (Supplementary Fig. S4). However, when the top-associated SNP was regressed out from the phenotype values, no SNPs were selected to explain sea age at maturity for all five candidate regions. Additionally, post-regression median proportion of variance was substantially lower—ranging between 0% and 1% (Supplementary Fig. S5, Table 2). This would suggest that sea age variation explained by each of these regions is largely explained by a single mutation. Pairwise LD among SNPs (R2) in these regions are reported in Supplementary Fig. S6. We observe no obvious trends in parameter values or Bayes factors, suggesting models converged and burn-in period was adequate (Supplementary Figs. S7 and S8).
A ssa06:27541960-28218141, B ssa09:10915066-11415066, C ssa09:24636574-25136574, D ssa21:49390687-49890687, and E ssa25:28389273-28889273. Plots display the following results for each candidate region: (i) posterior inclusion probability (PIP) indicating the probability of a SNP being included in a model explaining sea age at maturity variation, (ii) truncated distribution of the number of SNPs included in a model explaining sea age at maturity variation, and (iii) distribution of proportion of variance explained per recorded iteration (2500). Red line indicates the median proportion of variance explained.
Discussion
Despite that combined effects of multiple variants at trait-associated loci are playing an important role in controlling fitness traits across a variety of species (Linnen et al. 2013; Bickel et al. 2011; Kerdaffrec et al. 2016), our results indicate that sea age at maturation in Atlantic salmon is predominantly associated with single SNP variation at candidate regions. Using resequencing data to analyse 116 candidate loci and an analytical framework aimed at detecting multi-SNP associations, we find that single SNPs explain the variation in sea age at maturity in almost all cases. This work targeting candidate genes identified in aquaculture salmon strains suggests a mixed genetic architecture where a combination large-effect loci and smaller-effect loci also underlies age at maturity in wild Atlantic salmon populations. Two core loci, vgll3 and six6, likely play a key role in determining age at maturity and additional smaller effect loci may be important for fine-tuning the trait across heterogeneous environments.
Theoretical modelling predicts that clustering of tightly linked adaptive mutations will occur under gene flow and selection in populations inhabiting spatially and/or temporally heterogeneous environments (Yeaman and Whitlock 2011; Yeaman 2013). In Atlantic salmon, mean sea age at maturity varies among populations. Furthermore, spatially varying selection at the vgll3 locus displays homozygosity patterns consistent with selection towards local optima for sea age at maturity (Barson et al. 2015). Although these theoretical predictions seem to be a plausible scenario under which the genetic architecture of age at maturity has evolved in Atlantic salmon, our work suggests that the association at four of the five candidate regions is driven by a single mutation. We cannot rule out, however, the possibility that the examined regions have pleiotropic effects and contain SNPs controlling other adaptive traits that have weak or no correlation with maturation timing. It is also possible that we did not have sufficient power to detect additional SNPs in these regions with small effects or with rare alleles. However, previous empirical studies have found few, but complex, loci with clusters of adaptive mutations (Kerdaffrec et al. 2016; Linnen et al. 2013; Bickel et al. 2011), thus motivating our investigation of multi-SNP and haplotypic effects. Remington (2015) also highlights the importance of distinguishing between allelic effects and single mutational effects when examining the genetic architecture of adaptive variation and its evolution. Our findings, however, suggest that alternative genetic architectures are feasible. One possible explanation could relate to the multiple whole genome duplication events that have occurred in Atlantic salmon and other salmonids (Allendorf and Thorgaard 1984). The presence of multiple gene copies may impact the evolution of genetic architecture for traits such as age at maturity in Atlantic salmon. It is also possible that gene flow among Atlantic salmon populations is too restricted to neighbouring populations and/or strength of selection is insufficient for the establishment of linked mutations, as there is a rather specific balance of gene flow and selection required for clustered loci to arise (Yeaman et al. 2016). Both an extension of models predicting genetic architecture and additional empirical studies—on a wider variety organisms and traits—are needed to evaluate the generality of particular architectures and to further understand the conditions under which they evolve.
We find additional evidence that a large-effect locus on ssa25, vgll3, largely underlies age at maturity in Atlantic salmon corroborating findings from a number of association studies on Atlantic salmon maturation (Barson et al. 2015; Ayllon et al. 2015; Ayllon et al. 2019; Sinclair-Waters et al. 2020; Sinclair-Waters et al. 2022). The second strongest associated locus in this study is located in close proximity to six6 on ssa09. This locus was previously found to be associated with early maturation in male farmed Atlantic salmon (Sinclair-Waters et al. 2020), with sea age at maturity in wild Atlantic salmon prior to population structure correction (Barson et al. 2015) and two species of Pacific salmon (Sockeye salmon and Steelhead trout). Although six6 is associated with maturation in both Atlantic and several Pacific salmon species, an association between vgll3 and maturation timing has not been found in Pacific salmon species (Waters et al. 2021; Willis et al. 2020). Additionally, we found another three loci associated with sea age at maturity: pecam1, asap2aa and taar13c. The handful of loci found here suggests that wild Atlantic salmon have a mixed genetic architecture where multiple loci, with a variety of effect sizes, control maturation timing—similar to what has been found in male farmed Atlantic salmon (Sinclair-Waters et al. 2020). Knowledge of this mixed genetic architecture is highly relevant for how we predict the evolution of maturation timing in wild Atlantic salmon populations. A large body of work has shown the relevance of genetic architecture in determining evolutionary responses (Barton and Turelli 1991; Turelli 1984; Turelli and Barton 2004; Turelli and Barton 1990; Lande 1975; Bulmer 1972; Débarre et al. 2015; Fisher 1930; Yeaman 2015). Recent works highlight the relevance of the genetic architecture underlying fitness traits when predicting a population’s response to environmental changes (Kardos and Luikart 2021) and selective pressures such a fishing (Oomen et al. 2020). Future work elucidating how such mixed genetic architectures affect predicted evolution of traits, compared to that of omnigenic or polygenic architectures, will be valuable.
We find differences in locations of top-associated SNPs identified here and in Sinclair-Waters et al. (2020). This is not surprising given that we are examining sequence data that captures additional SNP variation in regions surrounding SNPs included in the SNP-array used in Sinclair-Waters et al. (2020). Furthermore, we failed to find associations between sea age at maturity and many of the candidate regions identified in Sinclair-Waters et al. (2020). For example, several candidate regions on ssa03 and ssa04 displayed particularly strong association signals in aquaculture salmon, however, no signals at these regions were found here. Additionally, only one association peak at ssa06:27541960-28218141 was found here, whereas two independent associations within this region were found in aquaculture salmon (Sinclair-Waters et al. 2020). Such differences may reflect changes in the genetic architecture of the trait evolving since the domestication of Atlantic salmon. Although, we would not expect large changes to occur given the domestication is relatively recent, just 10 to 15 generations ago (Gjerde and Gjedrem 1984). Furthermore, this study is likely under-powered to detect all previously identified loci, particularly those with smaller effect sizes or rare alleles, due to the smaller sample size of 313 individuals. Additionally, there could be differences in genetic architecture among environments (Yan et al. 2021) and/or genotype by environment interactions giving rise to distinct genetic architectures in wild populations versus aquaculture strains.
We do not find strong evidence of multi-SNP associations at candidate loci examined in this study, however, we cannot yet disregard the utility of multi-SNP association methods for further resolving the genetic architecture of Atlantic salmon maturation. First, we do not examine the entire genome due to computational restrictions, rather, we focussed on 116 previously identified candidate regions. Second, the Atlantic salmon genome is highly complex (Lien et al. 2016) and therefore errors in the assembly that may be disruptive for haplotype-based analysis could exist. As new and improved versions of the Atlantic salmon genome are published, our ability to test for haplotypic associations will improve. Furthermore, in a few cases (ssa09:10915066-11415066, ssa09:24636574-25136574, ssa25:28389273-28889273) the PiMASS analyses post-regression of the top SNP selected no SNPs for a model explaining sea age at maturity variation, however, the median proportion of variance explained across all iterations was greater than zero. This may suggest that a weak signal was present, but was being missed due to insufficient power. Although this is largely speculative, it suggests that ruling out the possibility of multi-SNP associations at these particular candidate regions may be premature. Higher-powered studies (i.e. more individuals per population) may help to resolve this in the future. Additionally, if an additional SNP is in high LD with the top-associated SNP, disentangling its effects from the top-associated SNP is challenging and its association signal could be undetectable post-regression of the top SNP. In such cases, study designs that take advantage of recombination events between highly linked SNPs are useful for characterizing genetic architectures at finer-scales (Sinclair-Waters et al. 2022).
Our analytical framework, combining both single and multi-SNP association methods, reveals that single SNP variation is sufficient for explaining the association at multiple previously identified candidate loci for Atlantic salmon maturation timing. Previous empirical and theoretical work have described trait-associated loci that have complex alleles with multiple variants, our findings therefore demonstrate the diversity of genetic architectures for fitness-related traits. Additional data, and a greater diversity of species and traits, will serve to better understand why this diversity of genetic architectures exists and how these particular genetic architectures evolve. The analytical framework used here will be a valuable resource for accomplishing this as individual-level resequencing data for wild species with phenotyped individuals becomes increasingly available.
Data availability
Genome re-sequencing data for individuals used in this study are available in the European Nucleotide Archive (ENA) or NCBI with the project accession code PRJEB38061 (Bertolotti et al. 2020).
References
Allendorf FW, Thorgaard GH (1984) Tetraploidy and the Evolution of the Salmonid Fishes. Monogr. Evol. Biol. Springer, Boston, https://doi.org/10.1007/978-1-4684-4652-4_2
Ayllon F, Kjærner-Semb E, Furmanek T, Wennevik V, Solberg MF, Dahle G, Taranger GL et al. (2015) The Vgll3 Locus Controls Age at Maturity in Wild and Domesticated Atlantic Salmon (Salmo Salar L.) Males. PLoS Genet 11(11):1–15. https://doi.org/10.1371/journal.pgen.1005628
Ayllon F, Solberg MF, Glover KA, Mohammadi F, Kjærner-semb E, Fjelldal PG, Andersson E, Hansen T, Edvardsen RB, Wargelius A (2019) The Influence of Vgll3 Genotypes on Sea Age at Maturity Is Altered in Farmed Mowi Strain Atlantic Salmon. BMC Genet 20(44):1–8. BMC Genetics
Barson NJ, Aykanat T, Hindar K, Baranski M, Bolstad GH, Fiske P, Jacq C et al. (2015) Sex-Dependent Dominance at a Single Locus Maintains Variation in Age at Maturity in Salmon. Nature 528(7582):405–8. Nature Publishing Group, a division of Macmillan Publishers Limited. All Rights Reserved. https://doi.org/10.1038/nature16062
Barton NH, Turelli M (1991) Natural and Sexual Selection on Many Loci. Genetics 127(1):229–55
Bertolotti AC, Layer RM, Gundappa MK, Gallagher MD, Pehlivanoglu E, Nome T, Robledo DM, et al. (2020) The Structural Variation Landscape in 492 Atlantic Salmon Genomes. Nat. Commun. 11(5176). https://doi.org/10.1038/s41467-020-18972-x.
Bickel RD, Kopp A, Nuzhdin SV (2011) Composite Effects of Polymorphisms near Multiple Regulatory Elements Create a Major-Effect QTL. PLoS Genet 7(1):1–8. https://doi.org/10.1371/journal.pgen.1001275
Boyle EA, Li YI, Pritchard JK (2017) An Expanded View of Complex Traits: From Polygenic to Omnigenic. Cell 169(7):1177–86. https://doi.org/10.1016/j.cell.2017.05.038. Elsevier
Bulmer MG (1972) The Genetic Variability of Polygenic Characters under Optimizing Selection, Mutation and Drift. Genet Res (Camb) 19(1):17–25
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: Architecture and Applications. BMC Bioinforma 10:1–9. https://doi.org/10.1186/1471-2105-10-421
Cánovas A, Reverter A, DeAtley KL, Ashley RL, Colgrave ML, Fortes MRS, Islas-Trejo A et al. (2014) Multi-Tissue Omics Analyses Reveal Molecular Regulatory Networks for Puberty in Composite Beef Cattle. PLoS One 9(7):1–17. https://doi.org/10.1371/journal.pone.0102551
Chang CC, Chow CC, Tellier LCAM, Vattikuti S, Purcell SM, Lee JJ (2015) Second-Generation PLINK: Rising to the Challenge of Larger and Richer Datasets. Gigascience 4(1):7. https://doi.org/10.1186/s13742-015-0047-8
Chapman B, Kirchner R, Pantano L, De Smet, M, Beltrame L, Khotiainsteva T, Naumenko S, et al (2020) Bcbio/Bcbio-Nextgen: V1.2.3, April. https://doi.org/10.5281/ZENODO.3743344.
Chaput G (2012) Overview of the Status of Atlantic Salmon (Salmo Salar) in the North Atlantic and Trends in Marine Mortality. ICES J Mar Sci 69(9):1538–48. https://doi.org/10.1093/icesjms/fss013
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A Program for Annotating and Predicting the Effects of Single Nucleotide Polymorphisms, SnpEff: SNPs in the Genome of Drosophila Melanogaster Strain W1118; Iso-2; Iso-3. Fly (Austin) 6(2):80–92. https://doi.org/10.4161/fly.19695
Daborn PJ, Yen JL, Bogwitz MR, Goff G, Le, Feil E, Jeffers S, Tijet N et al. (2002) A Single P450 Allele Associated with Insecticide Resistance in <Em>Drosophila</Em>. Sci (80-) 297(5590):2253 LP–2256. https://doi.org/10.1126/science.1074170
Danecek P, Auton A, Abecasis G, Albers C, Banks E, DePristo M (2011) The Variant Call Format and Vcftools. Bioinformatics 27(15):2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Day FR, Thompson DJ, Helgason H, Chasman DI, Finucane H, Sulem P, Ruth KS et al. (2017) Genomic Analyses Identify Hundreds of Variants Associated with Age at Menarche and Support a Role for Puberty Timing in Cancer Risk. Nat Genet 49(6):834–41. https://doi.org/10.1038/ng.3841
Débarre F, Yeaman S, Guillaume, F (2015) Evolution of Quantitative Traits under a Migration-Selection Balance: When Does Skew Matter?*. Am. Nat. 186. https://doi.org/10.1086/681717.
Depristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA et al. (2011) A Framework for Variation Discovery and Genotyping Using Next-Generation DNA Sequencing Data. Nat Genet 43(5):491–501. https://doi.org/10.1038/ng.806
Erkinaro J, Czorlich Y, Orell P, Kuusela J, Falkegård M, Länsman M, Pulkkinen H, Primmer CR, Niemelä E (2019) Life History Variation across Four Decades in a Diverse Population Complex of Atlantic Salmon in a Large Subarctic River. Can J Fish Aquat Sci 76(1):42–55. https://doi.org/10.1139/cjfas-2017-0343
Fisher R (1918) The Correlations between Relatives on the Supposition of Mendelian Inheritance. Philos Trans R Soc Edinb 52:399–433
Fisher R (1930) The Genetical Theory of Natural Selection. Clarendon, Oxford
Fleming I (1996) Reproductive Strategies of Atlantic Salmon: Ecology and Evolution. Rev Fish Biol Fish 6:379–416
Fleming IA, Einum S (2011) Reproductive Ecology: A Tale of Two Sexes. In Atl. Salmon Ecol. Wiley-Blackwell, Chichester, UK, pp 35–65.
Gjedrem T, Gjoen HM, Gjerde B (1991) Genetic Origin of Norwegian Farmed Atlantic Salmon. Aquaculture 98(1–3):41–50. https://doi.org/10.1016/0044-8486(91)90369-I
Gjerde B, Gjedrem T (1984) Estimates of Phenotypic and Genetic Parameters for Carcass Traits in Atlantic Salmon and Rainbow Trout. Aquaculture 36(1–2):97–110. https://doi.org/10.1016/0044-8486(84)90057-7
Gjerde B (1984) Response to Individual Selection for Age at Sexual Maturity in Atlantic Salmon. Aquaculture 38(3):229–40. https://doi.org/10.1016/0044-8486(84)90147-9
Guan Y (2014) Detecting Structure of Haplotypes and Local Ancestry. Genetics 196(3):625–42. https://doi.org/10.1534/genetics.113.160697
Guan Y, Stephens M (2011) Bayesian Variable Selection Regression for Genome-Wide Association Studies and Other Large-Scale Problems. Ann Appl Stat 5(3):1780–1815. https://doi.org/10.1214/11-AOAS455
Halperin DS, Pan C, Lusis AJ, Tontonoz P (2013) Vestigial-like 3 Is an Inhibitor of Adipocyte Differentiation. J Lipid Res 54(2):473–81. https://doi.org/10.1194/jlr.m032755
Jeffreys H (1961) Theory of Probability, 3rd ed. Oxford Univ. Press, Oxford
Hoekstra HE (2006) Genetics, Development and Evolution of Adaptive Pigmentation in Vertebrates. Heredity (Edinb) 97(3):222–34. https://doi.org/10.1038/sj.hdy.6800861
Hutchings JA, Jones MEB (1998) Life History Variation and Growth Rate Thresholds for Maturity in Atlantic Salmon, Salmo Salar. Can J Fish Aquat Sci 55(Suppl. 1):22–47. https://doi.org/10.1139/cjfas-55-S1-22
ICES (2011) Report of the Workshop on Age Determination of Salmon (WKADS).
Jonsson N, Hansen LP, Jonsson B (1991) Variation in Age, Size and Repeat Spawning of Adult Atlantic Salmon in Relation to River Discharge. J Anim Ecol 60(3):937–47. https://doi.org/10.2307/5423.
Joron M, Frezal L, Jones RT, Chamberlain NL, Lee SF, Haag CR, Whibley A, et al (2011) Polymorphic Supergene Controlling Butterfly Mimicry. Nature. Nature Publishing Group. https://doi.org/10.1038/nature10341.
Kardos M, Luikart G (2021) The Genetic Architecture of Fitness Drives Population Viability during Rapid Environmental Change. Am Nat. https://doi.org/10.1086/713469
Kerdaffrec E, Filiault DL, Korte A, Sasaki E, Nizhynska V, Seren Ü, Nordborg M (2016) Multiple Alleles at a Single Locus Control Seed Dormancy in Swedish Arabidopsis. Elife 5(3):1–24. https://doi.org/10.7554/eLife.22502
Kimura M (1983) The Neutral Theory of Molecular Evolution. Cambridge University Press, Cambridge, UK
Kirubakaran TG, Grove H, Kent MP, Sandve SR, Baranski M, Nome T, De Rosa MC et al. (2016) Two Adjacent Inversions Maintain Genomic Differentiation between Migratory and Stationary Ecotypes of Atlantic Cod. Mol Ecol 25:2130–43. https://doi.org/10.1111/mec.13592
Küpper C, Stocks M, Risse JE, Remedios N, Dos, Farrell LL, McRae SB, Morgan TC et al. (2015) A Supergene Determines Highly Divergent Male Reproductive Morphs in the Ruff. Nat Genet 48(1):79–83. https://doi.org/10.1038/ng.3443. Nature Publishing Group
Lamichhaney S, Fan G, Widemo F, Gunnarsson U, Thalmann DS, Hoeppner MP, Kerje S et al. (2015) Structural Genomic Changes Underlie Alternative Reproductive Strategies in the Ruff (Philomachus Pugnax). Nat Genet 48(1):84–88. https://doi.org/10.1038/ng.3430
Lande R (1975) The Maintenance of Genetic Variability by Mutation in a Polygenic Character with Linked Loci. Genet Res 26(3):221–35. https://doi.org/10.1017/S0016672300016037.
Li H (2009) SNPable Regions. http://lh3lh3.users.sourceforge.net/snpable.shtml.
Li H (2013) Aligning Sequence Reads, Clone Sequences and Assembly Contigs with BWA-MEM. ArXiv, March. https://arxiv.org/abs/1303.3997
Li H, Durbin R (2009) Fast and Accurate Short Read Alignment with Burrows-Wheeler Transform. Bioinformatics 25(14):1754–60. https://doi.org/10.1093/bioinformatics/btp324
Lien S, Koop BF, Sandve SR, Miller JR, Kent MP, Nome T, Hvidsten TR et al. (2016) The Atlantic Salmon Genome Provides Insights into Rediploidization. Nature 533(7602):200–205. Nature Publishing Group, a division of Macmillan Publishers Limited. All Rights Reserved.
Linnen CR, Poh Y-P, Peterson BK, Barrett RDH, Larson JG, Jensen JD, Hoekstra HE (2013) Adaptive Evolution of Multiple Traits through Multiple Mutations at a Single Gene. Sci (80-) 339(6125):1312–16. https://doi.org/10.1126/science.1233213
Liu X, Li YI, Pritchard JK (2019) Trans Effects on Gene Expression Can Drive Omnigenic Inheritance. Cell 177(4):1022–1034.e6. https://doi.org/10.1016/j.cell.2019.04.014.
Loh PR, Bhatia G, Gusev A, Finucane HK, Bulik-Sullivan BK, Pollack SJ, De Candia TR et al. (2015) Contrasting Genetic Architectures of Schizophrenia and Other Complex Diseases Using Fast Variance-Components Analysis. Nat Genet 47(12):1385–92. https://doi.org/10.1038/ng.3431
Mobley KB, Granroth-Wilding H, Ellmén M, Orell P, Erkinaro J, Primmer CR (2020) Time Spent in Distinct Life History Stages Has Sex-Specific Effects on Reproductive Fitness in Wild Atlantic Salmon. Mol Ecol 29(6):1173–84. https://doi.org/10.1111/mec.15390
Mobley KB, Aykanat T, Czorlich Y, House A, Kurko J, Miettinen A, Moustakas-Verho J et al. (2021) Maturation in Atlantic Salmon (Salmo Salar, Salmonidae): A Synthesis of Ecological, Genetic, and Molecular Processes. Rev Fish Biol Fish 31:523–571. https://doi.org/10.1007/s11160-021-09656-w
Oomen RA, Kuparinen A, Hutchings JA (2020) Consequences of Single-Locus and Tightly Linked Genomic Architectures for Evolutionary Responses to Environmental Change. J Hered 319–32. https://doi.org/10.1093/jhered/esaa020.
Orr HA (1998) The Population Genetics of Adaptation: The Distribution of Factors Fixed during Adaptive Evolution. Evolution (N. Y) 52(4):935–49. http://www.jstor.org/stable/2411226
Orr HA (1999) The Evolutionary Genetics of Adaptation: A Simulation Study. Genet Res 74(3):207–14. https://doi.org/10.1017/S0016672399004164
Patterson N, Price AL, Reich D (2006) Population Structure and Eigenanalysis. PLoS Genet. 2(12). https://doi.org/10.1371/journal.pgen.0020190.
Pearse DE, Barson NJ, Nome T, Gao G, Campbell MA, Abadía-Cardoso A, Anderson EC, et al (2018) Sex-Dependent Dominance Maintains Migration Supergene in Rainbow Trout. BioRxiv. 504621. https://doi.org/10.1101/504621.
Perry JRB, Day F, Elks CE, Sulem P, Thompson DJ, Ferreira T, He C et al. (2014) Parent-of-Origin-Specific Allelic Associations among 106 Genomic Loci for Age at Menarche. Nature 514(Jul):92. Nature Publishing Group, a division of Macmillan Publishers Limited. All Rights Reserved.
Picard Toolkit. (2019) Broad Institute, GitHub Repository. https://broadinstitute.github.io/picard/; Broad Institute
Price AL, Zaitlen NA, Reich D, Patterson N (2010) New Approaches to Population Stratification in Genome-Wide Association Studies. Nat Rev Genet 11:459. Nature Publishing Group, a division of Macmillan Publishers Limited. All Rights Reserved.
Pritchard JK, Di Rienzo A (2010) Adaptation - Not by Sweeps Alone. Nat Rev Genet 11(10):665–67. https://doi.org/10.1038/nrg2880
Purcell S, Neale, B, Todd-Brown, K, Thomas, L, Ferreira, M, and Bender, D. (2007) Plink: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am J Hum Genet 81. https://doi.org/10.1086/519795
Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, Sullivan PF, Ruderfer DM et al. (2009) Common Polygenic Variation Contributes to Risk of Schizophrenia and Bipolar Disorder. Nature 460(7256):748–52. https://doi.org/10.1038/nature08185
Quinlan AR, Hall IM (2010) BEDTools: A Flexible Suite of Utilities for Comparing Genomic Features. Bioinformatics 26(6):841–42. https://doi.org/10.1093/bioinformatics/btq033
Reed TE, Prodöhl PA, Bradley C, Gilbey J, McGinnity P, Primmer CR, Bacon, PJ (2018) Heritability Estimation via Molecular Pedigree Reconstruction in a Wild Fish Population Reveals Substantial Evolutionary Potential for Sea-Age at Maturity, but Not Size within Age-Classes. Can J Fish Aquat Sci., cjfas-2018-0123. https://doi.org/10.1139/cjfas-2018-0123
Remington DL (2015) Alleles versus Mutations: Understanding the Evolution of Genetic Architecture Requires a Molecular Perspective on Allelic Origins. Evolution (N. Y) 69(12):3025–38. https://doi.org/10.1111/evo.12775
Rockman MV (2012) The QTN Program and the Alleles That Matter for Evolution: All That’s Gold Does Not Glitter. Evolution (N. Y) 66(1):1–17. https://doi.org/10.1111/j.1558-5646.2011.01486.x
Sinclair-Waters M, Bradbury IR, Morris CJ, Lien S, Kent MP, Bentzen P (2018) Ancient Chromosomal Rearrangement Associated with Local Adaptation of a Post-Glacially Colonized Population of Atlantic Cod in the Northwest Atlantic. Mol Ecol 27:339–51. https://doi.org/10.1111/mec.14442
Sinclair-Waters M, Ødegård J, Korsvoll SA, Moen T, Lien S, Primmer CR, Barson NJ (2020) Beyond Large-Effect Loci: Large-Scale GWAS Reveals a Mixed Large-Effect and Polygenic Architecture for Age at Maturity of Atlantic Salmon. Genet Sel Evol 52(1):9. https://doi.org/10.1186/s12711-020-0529-8. BioMed Central
Sinclair-Waters M, Piavchenko N, Ruokolainen A, Aykanat T, Erkinaro J, Primmer CR (2022) Refining the Genomic Location of SNP Variation Affecting Atlantic Salmon Maturation Timing at a Key Large‐effect Locus. Mol Ecol 13(2):562–70. https://doi.org/10.1111/mec.16256
Sinnott-Armstrong N, Naqvi S, Rivas MA, Pritchard JK (2020) GWAS of Three Molecular Traits Highlights Core Genes and Pathways alongside a Highly Polygenic Background. BioRxiv, 2020.04.20.051631. https://doi.org/10.1101/2020.04.20.051631.
Stearns SC (2000) Life History Evolution: Successes, Limitations, and Prospects. Naturwissenschaften 87(11):476–86. https://doi.org/10.1007/s001140050763
Stinchcombe JR, Hoekstra HE (2008) Combining Population Genomics and Quantitative Genetics: Finding the Genes Underlying Ecologically Important Traits. Heredity (Edinb) 100(2):158–70. https://doi.org/10.1038/sj.hdy.6800937
Tishkoff SA, Reed FA, Ranciaro A, Voight BF, Babbitt CC, Silverman JS, Powell K et al. (2007) Convergent Adaptation of Human Lactase Persistence in Africa and Europe. Nat Genet 39(1):31–40. https://doi.org/10.1038/ng1946
Turelli M, Barton NH (2004) Polygenic Variation Maintained by Balancing Selection: Pleiotropy, Sex-Dependent Allelic Effects and G × E Interactions. Genetics 166(2):1053–79. https://doi.org/10.1534/genetics.166.2.1053
Turelli M (1984) Heritable Genetic Variation via Mutation-Selection Balance: Lerch’s Zeta Meets the Abdominal Bristle. Theor Popul Biol 25(2):138–93. https://doi.org/10.1016/0040-5809(84)90017-0. United States
Turelli M, Barton NH (1990) Dynamics of Polygenic Characters under Selection. Theor Popul Biol 38(1):1–57. https://doi.org/10.1016/0040-5809(90)90002-D
Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, Yang J (2017) 10 Years of GWAS Discovery: Biology, Function, and Translation. Am J Hum Genet 101(1):5–22. https://doi.org/10.1016/j.ajhg.2017.06.005. ElsevierCompany
Waters CD, Clemento A, Aykanat T, Garza JC, Naish KA, Narum S, Primmer CR (2021) Heterogeneous Genetic Basis of Age at Maturity in Salmonid Fishes. Mol Ecol. https://doi.org/10.1111/MEC.15822.
Weir BS, Cockerham CC (1984) Estimating F-Statistics for the Analysis of Population Structure. Evolution (N. Y) 38(6):1358–70. https://doi.org/10.2307/2408641
Willis SC, Hess JE, Fryer JK, Whiteaker JM, Brun C, Gerstenberger R, Narum, SR (2020) Steelhead (Oncorhynchus Mykiss) Lineages and Sexes Show Variable Patterns of Association of Adult Migration‐timing and Age‐at‐maturity Traits with Two Genomic Regions. Evol Appl (Jul)1–21. https://doi.org/10.1111/eva.13088.
Xu H, Guan Y (2014) Detecting Local Haplotype Sharing and Haplotype Association. Genetics 197(3):823–38. https://doi.org/10.1534/genetics.114.164814
Yan W, Wang B, Chan E, Mitchell-Olds T (2021) Genetic Architecture and Adaptation of Flowering Time among Environments. New Phytol. n/a (n/a). John Wiley & Sons, Ltd. https://doi.org/10.1111/nph.17229.
Yeaman S (2013) Genomic Rearrangements and the Evolution of Clusters of Locally Adaptive Loci. Proc Natl Acad Sci U S A 110:E1743–51. https://doi.org/10.1073/pnas.1219381110
Yeaman S (2015) Local Adaptation by Alleles of Small Effect *. Am. Nat. 186. https://doi.org/10.1086/682405.
Yeaman S, Aeschbacher S, Bürger R (2016) The Evolution of Genomic Islands by Increased Establishment Probability of Linked Alleles. Mol Ecol 25(11):2542–58. https://doi.org/10.1111/mec.13611. John Wiley & Sons, Ltd
Yeaman S, Gerstein AC, Hodgins KA, Whitlock MC (2018) Quantifying How Constraints Limit the Diversity of Viable Routes to Adaptation. PLoS Genet 14(10):1–25. https://doi.org/10.1371/journal.pgen.1007717
Yeaman S, Whitlock MC (2011) The Genetic Architecture of Adaptation under Migration-Selection Balance. Evolution (N. Y) 65(7):1897–1911. https://doi.org/10.1111/j.1558-5646.2011.01269.x
Zheng X, Levine D, Shen J, Gogarten SM, Laurie C, Weir BS (2012) A High-Performance Computing Toolset for Relatedness and Principal Component Analysis of SNP Data. Bioinformatics 28(24):3326–28. https://doi.org/10.1093/bioinformatics/bts606
Acknowledgements
We would like to acknowledge Terese Andersstuen, Dr Mariann Árnyasi and Hanne Hellerud Hansen from CIGENE for their work in organising the sequencing of samples. We thank Gunnel Østborg (NINA), Kurt Urdal (Rådgivende Biologer) and Natural Resources Institute Finland (LUKE) for their work collecting phenotype data. We also acknowledge the Aqua Genome project for providing access to data prior to public release. The Orion Computing Cluster at CIGENE-NMBU and CSC – IT Center for Science, Finland are acknowledged for computational resources. Storage resources were provided by the Norwegian National Infrastructure for Research Data (NIRD, project NS9055K). Phenotype data was provided by the Norwegian Institute for Nature Research (NINA).
Funding
Funding was provided by Academy of Finland (grant numbers 307593, 302873 and 327255), the Research Council of Norway (NFR-275310 and NFR-275862) and a Natural Sciences and Engineering Research Council of Canada postgraduate scholarship. Wild Atlantic salmon genome sequencing was funded by the Research Council of Norway (The Aqua Genome project; ref: 221734). Open Access funding provided by University of Helsinki including Helsinki University Central Hospital.
Author information
Authors and Affiliations
Contributions
CRP, NJB, MSW conceived the study. TN developed the variant calling workflow and constructed the fixed assembly of ssa25. JW developed the variant filtering criteria. MSW performed all downstream analyses with input from NJB. MPK played key role in generating whole genome sequencing data. SL led the whole genome sequencing work as part of the AquaGenome project. HS, GHB, BFL, CRP coordinated Atlantic salmon sampling and provided phenotypic information. MSW, CRP, NJB drafted the manuscript. All authors commented on and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Bastiaan Star.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sinclair-Waters, M., Nome, T., Wang, J. et al. Dissecting the loci underlying maturation timing in Atlantic salmon using haplotype and multi-SNP based association methods. Heredity 129, 356–365 (2022). https://doi.org/10.1038/s41437-022-00570-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00570-w