Abstract
Most heritable information in eukaryotic cells is encoded in the nuclear genome, with inheritance patterns following classic Mendelian segregation. Genomes residing in the cytoplasm, however, prove to be a peculiar exception to this rule. Cytoplasmic genetic elements are generally maternally inherited, although there are several exceptions where these are paternally, biparentally or doubly-uniparentally inherited. In this review, we examine the diversity and peculiarities of cytoplasmically inherited genomes, and the broad evolutionary consequences that non-Mendelian inheritance brings. We first explore the origins of vertical transmission and uniparental inheritance, before detailing the vast diversity of cytoplasmic inheritance systems across Eukaryota. We then describe the evolution of genomic organisation across lineages, how this process has been shaped by interactions with the nuclear genome and population genetics dynamics. Finally, we discuss how both nuclear and cytoplasmic genomes have evolved to co-inhabit the same host cell via one of the longest symbiotic processes, and all the opportunities for intergenomic conflict that arise due to divergence in inheritance patterns. In sum, we cannot understand the evolution of eukaryotes without understanding hereditary symbiosis.
Similar content being viewed by others
Introduction
The vast majority of genes in eukaryotes are located within chromosomal structures in the nucleus of the cell. These transmit copies of themselves to the next generation via meiosis involving strict segregation. However, eukaryotic cells also harbour smaller genomes, which reside in the cytoplasm, including: mitochondrial DNA, chloroplast DNA, and symbiont genomes. Interestingly, cytoplasmic genetic elements have been shown to have very different inheritance patterns to classic Mendelian nuclear chromosomes. The first documented evidence for this came from Carl Correns research on the four o’clock plant Mirabilis jalapa, in which he detailed the non-Mendelian inheritance of leaf colour (Correns 1909). Inheritance, in this case, was strictly maternal: a seed derived from an ovule from a non-green stem gave rise to non-green progeny, irrespective of the source of pollen. By 1952, the evidence of various forms of cytoplasmically inherited elements (CIEs) had grown, leading Joshua Lederberg to synthesise the inheritance of cellular organelles and symbionts into one framework in his treatise “Cell genetics and hereditary symbiosis” (Lederberg 1952). Furthermore, evidence for diversity in inheritance patterns (paternal or biparental) of CIEs started accumulating for a wide range of taxa (Birky 2001).
Since then, studies have demonstrated that CIEs are diverse and important - in many cases, encoding key aspects of organismal function. Cytoplasmically inherited elements vary in their level of integration with the host - in the case of organelles, the proteome is jointly encoded in nuclear and organellar DNA, in addition to integration into cellular physiology. For obligate microbial symbionts, anatomical and physiological integration are evident but generally without trafficking of host proteins into the microbe; they are commonly present in particular tissues and have host organised vertical transmission. Other microbial symbionts are less integrated, present more diffusely in the host and invade the germ line to gain vertical transmission.
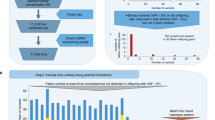
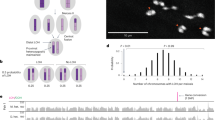
In this review, we describe the diversity of inheritance systems of CIEs, and highlight the evolutionary consequences that these inheritance systems bring to cellular, organismal and population dynamics (Fig. 1). For this, we focus on the three main groups of CIEs: mitochondrial DNA, chloroplast DNA, and symbiont genomes. We begin by outlining the origins of cytoplasmic inheritance and the evolution of uniparental inheritance, documenting the diversity of cytoplasmic inheritance systems so far observed. We discuss the diversity and patterns of genome organisation for cytoplasmic elements and examine the population genetics of CIEs, highlighting the tension between within- and between-individual spread. We summarise the evidence for the adaptive importance of cytoplasmic genes before detailing coadaptation between the cytoplasm and the nucleus, and amongst cytoplasmic components.
New mutations that arise immediately produce intra-organellar variation depicted here by differently coloured nucleoids (Mitochondria & Chloroplast panel). If mutations (differently coloured nucleoids) spread between organelles, variation between organelles is observed (Plant Cell panel). Note that mitochondria often form reticulated syncytia, rather than discrete compartments, in contrast to chloroplasts, which may facilitate recombination and therefore spread of mutations throughout the cell. Intracellular variation can give rise to intra-tissue variation, depicted here in the form of a variegated leaf (Psychotria leaf panel). Psychotria also features bacterial leaf nodules (dark green circles) that contain Burkholderia bacteria which are vertically inherited through the seed. Variation within tissues can then give rise to variation across tissues (Psychotria Plant panel). If germlines are segregated late, this can result in distinct alleles being propagated to the next generation from different parts of the plant. As a consequence, the variation that originated at the individual organelle level can finally be observed between individuals within populations (Psychotria population panel).
The evolutionary origins of vertical transmission and uniparental inheritance
The ancestors of current cytoplasmically inherited genetic material were free-living organisms (Sagan 1967), but how cytoplasmic inheritance originated and came to be limited to one sex remains an open question. For microeukaryotes (and indeed for the ancestral protoeukaryote) the presence of a microbe inside the cytoplasm would de facto produce inheritance on cell division. This would form a continuous system if replication of the microbe within the cell was occurring. Thus, the primary drive to cytoplasmic inheritance is intracellular location and replication, which could be initially host driven (symbiont capture) or symbiont driven (infection of the host, or escape from a phagolysosome).
Notably, the rate of vertical transmission for symbionts that can also transmit infectiously is evolvable: symbionts with mixed modes of transmission that are kept in continuously growing host populations (where cell division provides ample opportunity for vertical transmission) evolve a stronger tendency for heritable transmission compared to those kept in populations at carrying capacity (where the opportunity for vertical transmission is limited) (Magalon et al. 2010). Thus, there is a trajectory in which infection through the environment is lost.
When vertical transmission does evolve, there are two primary consequences. First, the population size of symbionts and mixing of strains declines, reducing within host conflicts. Second, symbiont fitness becomes a product not only of host survival but additionally host reproduction. Both of these processes drive the symbiosis towards the mutualism end of the mutualism-parasitism continuum, with current models indicating restrictions on symbiont diversity through bottlenecks and reduced mixing opportunities being most important in this transition, through quelling the conflicts associated with within-host competition (Leeks et al. 2019). Later transitions would then involve adaptation to the intracellular environment with correlated loss of capacity for free-living life and infection processes. All of these are reflected in the reductive genome evolution pattern commonly observed in heritable symbionts (Moran et al. 2008).
The evolution of inheritance for symbionts of multicellular hosts also has its origins in the association of free-living organisms, with a transition from symbiosis where the parties reform symbiosis through environmental association each generation to vertical transmission. Indeed, some symbiont clades include both symbionts acquired through the environment and heritable symbionts (e.g., (Drew et al. 2021)). Vertical transmission may arise passively through spatial structure (symbionts from a parent are more likely to infect progeny of that parent through proximity), actively through selection on the host to ensure passage of a beneficial symbiont (host driven vertical transmission) or actively through selection on the microbe to infect the next generation through the germ line. Evolution towards vertical transmission may be constrained through location (e.g., soil microbe-plant root associations have no proximity to the germ line), and made less likely in environments where partner availability is high (e.g., aquatic environments (Douglas 1998)) or where the symbiont has utility only in a restricted part of host life history (Hartmann et al. 2017).
Most cytoplasmic genetic material is inherited uniparentally - that is to say from one parent only. It is notable that uniparental inheritance is not restricted to anisogamous species - it is also commonly observed in isogamous microeukaryotes associated with mating types. This observation implies that uniparental inheritance is not a simple by-product of gamete size, but rather is an evolved state (Hurst and Hamilton 1992). Anisogamy itself is considered by some as a potential onward adaptive mechanism for imposing uniparental inheritance, in contrast to the passive view that uniparental inheritance is a by-product of anisogamy (Hurst 1990).
Evolutionary drivers of uniparental inheritance include the benefit of preventing conflicts and the damage from cytoplasmic mixing. Under a biparental inheritance scenario, we would expect a heteroplasmic state, in which multiple distinct forms of the CIE exist within the same cell, to be the norm. For mitochondria, heteroplasmy interferes with cell functioning, with empirical work demonstrating that it can cause organismal dysfunction (Nissanka and Moraes 2020). Moreover, heteroplasmy reduces the variance between cells, and if this is happening in the germ line, it then reduces the efficacy of selection (Radzvilavicius et al. 2016). Mathematical models show that selection against heteroplasmy can lead to the fixation of uniparental inheritance in an ancestrally biparental population (Christie et al. 2015; Christie and Beekman 2017). Mathematical models indicate that mitonuclear coadaptation is improved with uniparental inheritance and mitochondrial bottlenecks under a wide range of conditions (Hadjivasiliou et al. 2012).
Beyond simple maternal inheritance
Whilst the majority of anisogamous species transmit CIEs uniparentally, via the egg (Birky 2001), heteroplasmy via paternal leakage can occur when maternal inheritance is not strictly enforced (see Table 1 and Supplementary Table S1 for examples). Most species appear to exhibit some degree of leakage if sampled carefully enough (e.g., (Wagner et al. 1991; Kvist et al. 2003; Fontaine et al. 2007; Bentley et al. 2010; Nunes et al. 2013)), but the processes that contribute to variation in leakage rates are not well understood. By contrast, a variety of mechanisms that reinforce maternal inheritance by eliminating and/or silencing paternally derived elements have been documented (Sato and Sato 2013, 2017). Still, paternally derived CIEs occasionally experience positive selection, resulting in introgression and even replacement of the maternal CIE lineage by the paternal CIE lineage (Wolff et al. 2013).
Notably, there exists a biological distinction between paternal leakage versus paternal inheritance of CIEs. Paternal leakage, in which CIEs are inherited mostly maternally, but with some minor contribution from the paternal gamete, provides at least some opportunity for genetic exchange between CIEs from different lineages. By contrast, CIE lineages from separate parents are not expected to interact or recombine under systems of paternal inheritance, even if only occasional, in which all of the CIEs present in an individual host are paternally derived (e.g., Ross et al. 2016). The implications are that paternal leakage allows for breakdown of linkage disequilibrium between separate nucleotides of the CIE genome, such that beneficial mutations can be decoupled from deleterious genomic backgrounds and vice versa (Hill and Robertson 1966).
Evolutionary transitions from maternal to paternal inheritance are not especially common, but do happen (Table 1, Supplementary Table S1). Interestingly, shifts in one organelle do not always affect the other cytoplasmic elements within the cell (but see Pelargonium (Weihe et al. 2009); Sequoia (Neale et al. 1989)), indicating that the genetic machinery regulating cytoplasmic inheritance is independent across separate CIEs. For example, in Musa acuminata (banana), the mitochondria are inherited paternally and the chloroplasts are inherited maternally (Fauré et al. 1994). Some other examples of mixed uniparental inheritance include cucumbers, melons (Havey 1997) (in contrast to the rest of the Cucurbitaceae (Havey et al. 1998)), which follow the same pattern as banana, and loblolly pines, which feature maternally transmitted mitochondria and paternally transmitted chloroplasts (Neale and Sederoff 1989).
When both parents contribute CIEs to the offspring, it is probable that cells benefit from biparental mitochondrial inheritance as it provides higher individual genetic diversity, which leads to reduced susceptibility to deleterious mutation. However, this naturally leads to competition and conflict between lineages. Recent work has shown that biparental inheritance has the potential to be beneficial and remain stable for hybridising populations if the fitness cost of mitonuclear incompatibilities in hybrids is greater than a stable state of heteroplasmy (which is also detrimental to the individual, but less so) (Allison et al. 2021). In hybridised Pelargonium (geranium) cultivars, for example, mitochondrial and plastid biparental inheritance has been seen alongside chloroplast variegation (Baur 1908). In these hybrids, differing ratios of maternally and paternally derived cytoplasmic genomes were observed across different tissues (Weihe et al. 2009). Based on the phylogenetic distribution of biparental inheritance systems, it seems that the balance of reduced impact of deleterious mutations vs. increased conflict between CIE lineages favours the latter, as relatively few examples of fully biparentally inherited CIEs exist.
A particularly interesting case of biparental inheritance of mitochondria is that of doubly-uniparental inheritance (DUI) in bivalves, in which the paternal mtDNA (M-type) is passed down to male offspring and maternal mtDNA (F-type) is passed down to offspring of both sexes (Breton et al. 2007; Passamonti and Ghiselli 2009; Zouros 2013). In this scheme, two independently evolving mtDNA lineages are found in male individuals, but the sperm produced only contain the lineage they inherited paternally (Ladoukakis and Zouros 2017). The paternal mitochondria are largely confined to the gonad and the maternal mitochondria the soma, resulting in sperm that carry only the male mitochondrial line (Ghiselli et al. 2020). The separate localisation of M-type vs. F-type mtDNAs also makes it less likely that recombination between mtDNA lineages occurs (but see (Zouros 2000; Burzyński et al. 2003; Passamonti et al. 2003; Breton et al. 2006; Stewart et al. 2009)).
Heritable microbes are commonly considered as exclusively transmitted from mother to offspring only; however, this rule is not absolute. Many heritable viruses show biparental inheritance, commonly with higher fidelity through the egg than sperm (Roossinck 2010). For heritable bacteria, early studies of Volvox carteri revealed efficient paternal inheritance of what are now known to be Cand. Megaira symbionts (Lee and Kochert 1976); later work showed biparental inheritance of Rickettsia in Nephotettix planthoppers (Watanabe et al. 2014), and Sodalis glossinidia in tsetse fly hosts (De Vooght et al. 2015). Biparental inheritance allows symbionts to drive to high frequency without either conferring a benefit to their host, or exhibiting reproductive parasitism. It also creates an environment where mixed infections become common, which may lead to the evolution of increased virulence (associated with competition for transmission) and also potentiates recombination.
Genomic organisation and interactions with the nuclear genome
Genomic architectural organisation and variation in CIEs
Cytoplasmically inherited genomes are highly variable across eukaryotes in terms of both size and structure (Smith and Keeling 2015). Mitochondrial and chloroplast genomes are rarely lost entirely (but see (Hjort et al. 2010; Keeling 2010)); however, the diminutive size of mtDNAs in Plasmodium species (~6 kb) (Hikosaka et al. 2011) compared to the massive and multi-partite mitochondrial genomes of Silene conica (~11.3 Mb) (Sloan et al. 2012) and Larix siberica (~11.7 Mb) (Putintseva et al. 2020) highlight the diverse trajectories of cytoplasmic genome evolution following the original endosymbiotic event and subsequent massive transfer to the nucleus prior to the Last Eukaryotic Common Ancestor (Sloan et al. 2018).
Plastid genomes are less variable in size than mitochondrial genomes (Wu et al. 2020), but non-photosynthetic plastids have seen dramatic reductions in size compared to their photosynthetic ancestors (de Koning and Keeling 2006; Barbrook et al. 2014). While many CIEs exhibit circular genomes (e.g., most bilaterian mitochondrial genomes (Boore 1999) and most eubacterial symbionts), linear (Stampar et al. 2019; Escalante et al. 1998; Nosek et al. 2004; Shao et al. 2009), branched (Oldenburg and Bendich 1996), and multi-chromosomal arrangements (Wu et al. 2020) have arisen multiple separate times across eukaryotes. The absence of Mendelian inheritance in cytoplasmic genomes likely contributes to the tremendous variation observed there, to the extent that once multi-chromosomal genomes evolve, their inheritance is highly fragmented and inconsistent (Wu and Sloan 2019).
In parallel, heritable symbiont genomes vary greatly in genome complexity. There is a general pattern of genome reduction in symbiotic microbes, first associated with the transition from environmentally acquired to heritable, and then with the transition from facultative (not required by the host) to obligate (required by the host). Obligately required heritable symbionts have genomes that are commonly <1 Mb, and may be as small as 112 kb (Bennett and Moran 2013), in comparison to free-living microbes with genome sizes >4 Mb. Pseudogenization is rapid during the first phases of evolution, often accompanied by proliferation of mobile elements (Bennett and Moran 2013). G/C content typically reduces as the genome shrinks, with obligate symbiont genomes typically highly AT rich (Moran et al. 2008).
Interactions and molecular cross talk with the nuclear genome
It is common to observe genes in the nuclear genome which have cytoplasmic origin, and transfer of material from both mitochondria and microbial symbionts to the nucleus is ongoing (Bensasson et al. 2001; Dunning Hotopp et al. 2007). Importantly, recent nuclear derived symbiont sequences on occasion have strong phenotypic effects, potentiating retention (Leclercq et al. 2016). In deep evolutionary time, these transfers fuelled the seemingly inevitable gene transfer from the cytoplasm to the nucleus and subsequent genome streamlining that CIEs have repeatedly undergone (Timmis et al. 2004; Giannakis et al. 2021). This process has resulted in the vast majority of proteins that function in cytoplasmically inherited organelle compartments being encoded by the nucleus (Millar 2007; Meisinger et al. 2008; van Wijk and Baginsky 2011; Muthye and Lavrov 2018). The genes and gene products that are still retained in CIE genomes must therefore physically interact with nuclear-encoded gene products. To wit, four of the five multi-subunit enzymes that comprise the electron transport chain and the photosynthetic enzyme complexes of chloroplasts feature intimate interactions between subunits encoded by separately inherited and expressed genomes (Rand et al. 2004; Forsythe et al. 2019).
Much attention has been paid to the molecular nature of these cytoplasmic-nuclear interactions (Osada and Akashi 2011; van der Sluis et al. 2015; Beck et al. 2015; Adrion et al. 2016; Mossman et al. 2017; Rand and Mossman 2020; Evans et al. 2021), but relatively little is known about the stoichiometry of these interactions, except that cytoplasmic gene expression is consistently higher than expression of nuclear-encoded genes involved in the same multi-subunit complexes (Havird and Sloan 2016). Further, mitochondrial DNA depletion is associated with a number of different diseases in humans (Blokhin et al. 2008; Clay Montier et al. 2009; Monickaraj et al. 2012; Petersen et al. 2014; Pyle et al. 2016; Tin et al. 2016; Ashar et al. 2017; Liu et al. 2020), and polyploid plants exhibit elevated cytoplasmic DNA content per cell compared to diploid relatives to maintain cytonuclear stoichiometry following genome doubling (Fernandes Gyorfy et al. 2021). Even from this currently limited understanding, it is clear that complex stoichiometric relationships exist between the nuclear and cytoplasmic genomes and gene products, and perturbations to cytonuclear stoichiometry can therefore have drastic consequences for the cells that experience them.
Population and evolutionary genetics of cytoplasmic elements
Mutations and how they spread throughout the cytoplasm
Cytoplasmically inherited elements present a stark contrast to Mendelian traits as sources of variation. As Bill Birky noted, CIEs are ‘non-stringent’ genetic traits that can vary in quantity as well as sequence – in contrast to Mendelian elements limited to a copy number of two in any diploid cell (Birky 2001). Instead, CIEs are often highly multi-copy within cells (Kukat et al. 2011; Carelli et al. 2015; Schaack et al. 2020). As such, the distribution of sequence variation in CIEs is profoundly affected by the distribution of copy number variation, as mutants must first establish within the pool inside a cell, then amongst the cells within an individual, then amongst individuals in the population. Variation in copy number within the cell is also critical to the stoichiometric balance between the cytoplasmic and nuclear genomes, as they contribute to the assembly of the multi-subunit enzyme complexes that carry out bioenergetic processes like photosynthesis and respiration (Forsythe et al. 2019).
Mutational spread in cytoplasmically inherited genomes is fundamentally dependent upon the rate of occurrence of new mutations (Sung et al. 2012; Waneka et al. 2021). However, the multi-copy nature of CIEs makes it practically impossible to determine their absolute mutation rates (Schaack et al. 2020). Nevertheless, a large effort across decades has been dedicated to quantifying relative mitochondrial mutation rates and frequency spectra (Brown et al. 1979; Wolfe et al. 1987; Denver et al. 2000; Haag-Liautard et al. 2008; Howe et al. 2009; Havird and Sloan 2016; Allio et al. 2017; Konrad et al. 2017; Wu et al. 2020; Broz et al. 2021; Waneka et al. 2021), providing valuable information about the extent of heteroplasmy caused by de novo mutations (Waneka et al. 2021) and the probability of transmitting those heteroplasmies to the next generation (Konrad et al. 2017). Mutational spectra of mtDNA are also important to mutational spread (reviewed in (Katju and Bergthorsson 2019)). For example, oxidation of guanines (i.e., 8-oxo-G), especially in mitochondria, can result in elevated CG → AT transversions through mispairing with adenine (Cheng et al. 1992; Kino et al. 2017).
Despite the aforementioned difficulty in ascertaining absolute mutation rates in CIEs, it is clear that CIE mutation rates vary tremendously across taxa. For example, animal mtDNAs exhibit substantially higher mutation rates than plant mtDNAs (Wolfe et al. 1987). Indeed, animal mtDNA mutation rate varies more than two orders of magnitude across taxa (Nabholz et al. 2007). Certain plant lineages have shown episodic accelerations in cytoplasmic genome mutation rates (Sloan et al. 2014; Sloan 2015; Havird et al. 2015; Williams et al. 2019; Broz et al. 2021). There is also tremendous variation in mutation rate across different cellular genomes – animal mtDNAs exhibit higher mutation rates than animal nuclear genomes (Brown et al. 1979; Wolfe et al. 1987; Havird and Sloan 2016), but plant nuclear genomes exhibit higher mutation rates than plant cpDNA and plant mtDNAs (Wolfe et al. 1987).
Whilst mutation rate is not known for heritable microbes, it is known they vary substantially in substitution rate, with some heritable microbes evolving at a rate comparable to viruses (Gerth et al. 2021), and others, like Wolbachia, two-three orders of magnitude slower (Richardson et al. 2012). At least some of this variation can be traced back to the different mechanisms of replication and repair across taxa and compartments (Brown et al. 2005; Maréchal and Brisson 2010; Lewis et al. 2015; Gerth et al. 2021), all of which have implications for mutation rate (Longley et al. 2005; DeBalsi et al. 2017; Wu et al. 2020).
Mutational masking in cytoplasmic elements
New mutations that arise in CIEs face a dramatically different population genetic landscape compared to Mendelian elements because there are typically many competing cytoplasmic genomes present inside each cell, and because organellar cytoplasmic elements do not experience segregation, as is the case for nuclear genomes during sexual reproduction (Wilton et al. 2018). Thus, new mutations face a steep drift barrier, with their effects on host function being masked until reaching higher frequency within a cell or individual (potentially as high as 80% (King and Attardi 1989; Boulet et al. 1992; Stewart and Chinnery 2015)). As a consequence, mutations that exist at low frequencies among CIEs in a parent are likely to be lost as a result of the bottleneck that occurs between generations in multicellular organisms.
There are two sides to the mutational masking that results from harbouring many copies of cytoplasmic genomes inside cells: (1) mutations with deleterious fitness effects on the host can persist longer than they otherwise would if maintained in single-copy form within the cell (Otto 2007), and (2) mutations with beneficial fitness effects on the host can be lost at higher rates because their effects are largely invisible to selection. Recent high-resolution efforts support the existence of mutational masking, as nonsynonymous mutations are more common and exist at higher frequencies than expected (Waneka et al. 2021). Moreover, masking of the fitness effects of mutations is expected to result in a deletion bias (Lawless et al. 2020), especially under relaxed selection (Wickett et al. 2008), as CIEs with replication advantages (e.g., CIEs with smaller genomes) can rise in frequency within cells rapidly (Wallace 1989; Clark et al. 2012; Sloan and Wu 2014). This latter pattern may contribute to the observation that CIEs exhibit more streamlined genomes compared to their free-living relatives (Timmis et al. 2004; Giannakis et al. 2021).
The population of CIEs within cells, of cells within tissues, of somatic vs. germ line tissues, and of individual hosts within host populations gives rise to the expectation of multi-level selection, in which elements that have an advantage in terms of spread at a lower level of organisation do not necessarily possess the same advantage at higher levels of organisation (Fig. 1). To wit, mutations that remain at low frequencies across generations in human mtDNA can rise to high frequency in separate tissues within the same individual (Samuels et al. 2013; Rebolledo-Jaramillo et al. 2014; Li et al. 2015). Additionally, recent work in which artificial mixed infections of Buchnera were created within aphids demonstrated a ‘regular winner’, despite strong drift effects - but the winner did not necessarily confer individual level benefits (Perreau et al. 2021).
Recombination in the cytoplasm
The misconception that CIEs do not undergo recombination has been largely debunked. For example, phylogenetic and other experimental analyses of animal mitochondrial genomes consistently recover signatures of inter-molecular recombination, indicating that inheritance leakage may play a major role in CIE genome evolution (Mita et al. 1990; Kajander et al. 2000; Ladoukakis and Zouros 2001; Ladoukakis and Eyre-Walker 2004; Piganeau et al. 2004; Barr et al. 2005; Ciborowski et al. 2007; Ma and O’Farrell 2015; Leducq et al. 2017; Dahal et al. 2018). Plant plastids and plant mitochondria exhibit recombination-directed repair (Cerutti et al. 1995; Day and Madesis 2007; Maréchal and Brisson 2010; Davila et al. 2011; Gualberto and Newton 2017; Chevigny et al. 2020; Wu et al. 2020), as well as rampant structural rearrangement via repeat-mediated recombination (Palmer 1983; Ogihara et al. 1988; Palmer and Herbon 1988; Gray et al. 1999; Arrieta-Montiel and Mackenzie 2011; Cole et al. 2018; Wu and Sloan 2019; Xia et al. 2020). This latter phenomenon, termed substoichiometric shifting, makes for extensive, but heritable structural variation within individuals (Woloszynska 2009; Maréchal and Brisson 2010; Arrieta-Montiel and Mackenzie 2011; Davila et al. 2011).
Recombination has played such a large role in plant CIEs that their relatively slow rate of molecular evolution is thought to be due, at least in part, to recombination (Palmer and Herbon 1988; Chevigny et al. 2020; Wu et al. 2020), as heteroplasmies may be eliminated from intracellular populations via gene conversion. Whether the occasional and episodic accelerations in rates of cytoplasmic genome evolution in some plant lineages (Williams et al. 2019; Broz et al. 2021) is associated with altered recombinatorial activity remains an open question. Heritable microbes also show clear signatures of recombination (Baldo et al. 2006), as well as acquisition of genetic material from other bacteria (Nikoh et al. 2014), which commonly involves phage transfer (Kaur et al. 2021; Boyd et al. 2021).
The evolutionary consequences of cytoplasmic recombination are profound: recombination can act as a barrier to new mutations through gene conversion and can facilitate the rise of beneficial mutations and the elimination of deleterious mutations by separating those mutations from their genomic backgrounds (Neiman and Taylor 2009), particularly when distinct CIEs occur inside the same cell or organism.
Cytoplasmic adaptation
The seemingly asexual nature of animal cytoplasmic genomes suggests that they, like other asexual elements, will experience impaired adaptive evolution. Experimental evidence, however, has found this not to be the case, as numerous studies have reported signatures of positive selection acting within the mitochondrial genome (Mishmar et al. 2003; Ruiz-Pesini et al. 2004; Meiklejohn et al. 2007). Furthermore, clinal patterns in mtDNA genomes have been detected across several species, indicating signatures of adaptation (Camus et al. 2017; Silva et al. 2014). Early studies by Lynch and Blanchard (1998) found that mitochondrial genes had higher ratios of nonsynonymous to synonymous mutations in relation to the nuclear genome of plants, invertebrate and fungal taxa (Lynch and Blanchard 1998). Most recently, Morales et al. (2015) found evidence for positive selection on several amino acids in the mtDNA of the Australian eastern yellow robin (Eopsaltria australis) populations. The authors additionally found nuclear genome homogeneity within the robin populations sampled indicating that there were high levels of gene flow, thus the signatures of positive selection were unique to the mtDNA (Morales et al. 2015). The combined outcomes of these studies suggest that certain mtDNA protein-coding genes of natural populations might well have been shaped by positive selection.
Less evidence is available linking chloroplast genomes to adaptive processes, but this could be because of the slower rates of evolution, higher levels of complexity or the fact that dissecting the contributions of multiple organelle genomes is complicated. Nevertheless, there has been some work in domesticated species testing these questions. For example, research on rice (Oryza) has identified 14 chloroplast genes with strong signatures of positive selection, with these genes being mainly related to photosynthetic function. Interestingly, authors found that eight of these genes were independently found in sun-loving species, whereas other photosynthetic genes were selected in shade-tolerating species (Gao et al. 2019). Other studies have directly examined the effects of cytonuclear interactions across two different ecological environments. The sunflower genus Helianthus is commonly used as a model as many of its species have adapted to very distinct niches (Levin 2003). Sambatti and colleagues performed reciprocal transplant experiments between H. annus and H. petiolaris which inhabit mesic and xeric habitats respectively (Sambatti et al. 2008). In addition to examining both coevolved strains, authors used all possible backcross combinations to dissect the contribution of the cytoplasm and nuclear genome, finding that the cytoplasm was the main driver for fitness, and is therefore adapted to these two contrasting environments (Sambatti et al. 2008).
Heritable symbionts have been commonly observed to be under strong selection. Invasion of heritable symbionts into populations in real time has been observed on numerous occasions. For instance, a classic example is the wave of Wolbachia that induced cytoplasmic incompatibility (CI) which swept through Californian D. simulans populations in the 1980s (Turelli and Hoffmann 1991). Similarly, heritable Spiroplasma that provide tolerance to nematode parasitism have spread through North America, and Rickettsia spreading through whitefly populations has been observed over the last 20 years (Himler et al. 2011; Shi et al. 2021). It is also common to observe that symbionts either themselves have low diversity - or that associated mitochondria have low diversity - implying a recent history of joint selection. Finally, it is notable that heritable microbes (unlike mitochondria and chloroplasts) may segregate during host reproduction, with a fraction of progeny not inheriting them. Their maintenance thus requires some form of drive - either a benefit to host survival or reproductive parasitism. As such, it is argued they are never neutral traits, but are maintained by a selection - segregational loss balance (Jaenike 2012). In addition, they present different modularities of adaptive variation - like other CIEs they have different circulating variants in a population, but in addition, these commonly exist alongside uninfected cytotypes, which are the equivalent of a null allele.
Coadaptation
Coadaptation with nuclear encoded proteins and systems
The transfer of genetic material from cytoplasmic elements to the nucleus is thought to have created strong pressures for both nuclear and cytoplasmic genomes to cooperate with one another. Excessive amounts of conflict can have severe consequences to both host and symbiont. One of the classic demonstrations of mitonuclear coadaptation comes from studies using hybrid crosses from natural populations. The copepod species Tigriopus californicus has become one of the main wild model systems, primarily due to the high level of intraspecific divergence in mtDNA genomes. While crosses between populations give F1 offspring with normal (if not elevated) fitness compared to the parental generation, there is a drastic decrease in fitness in the F2 generations and beyond (Burton 1990). Using backcrossing approaches, they discovered that this decrease in fitness was caused by severe mitonuclear incompatibilities (Burton and Lee 1994), and the proportion of the maternal nuclear genome appears to be positively correlated with developmental rate in backcrossed individuals (Han and Barreto 2021). The nuclear-encoded mitochondrial genes (those interacting with genes encoded in the mtDNA) of the T. californicus genome have also been shown to coadapt with mtDNA, exhibiting elevated mutation-rate-corrected rates of evolution (i.e., dN/dS) compared to the rest of the nuclear-encoded DNA, matching the rapid pace of evolution in their mtDNA counterparts (Barreto et al. 2018).
Similarly, chloroplast genomes are predicted to be under strong selection to coadapt, and much work has been done to document the effects of plastid-nuclear interactions on plant fitness (Greiner et al. 2011; Postel and Touzet 2020). Nearly all of the ~75–80 proteins encoded by the chloroplast genome are involved in protein complexes which exhibit important functions that are essential to plant function, such as Rubisco and photosystems I and II. One of the best examples of coadaptation between chloroplast and nuclear genomes after intraspecific hybridisation comes from the genus Oenothera (Stubbe 1989; Greiner and Bock 2013), in which three basic haploid nuclear genomes can be paired with five different chloroplast genomes; giving a total of 30 possible chloro-nuclear combinations. Of these, only 12 produce a viable green phenotype, whereas the 18 remaining associations lead to various degrees of cytonuclear incompatibilities, from reduced phenotypic capacity to embryo lethality (Cleland 1972; Dietrich et al. 1997). Subsequent work suggests that the radiation within Oenothera is approximately 1 million years old, suggesting that incompatibilities and coadaptation mechanisms have rapidly evolved (Greiner et al. 2008). Although most chloroplast genomes evolve relatively slowly, occasional accelerations in evolutionary rate have occurred throughout angiosperms (Williams et al. 2019). In these cases, the nuclear-encoded interacting partners of chloroplast-encoded proteins exhibit corresponding increases in evolutionary rate, reflecting the co-evolutionary dynamics of plastid-nuclear interactions (Bock et al. 2014; Zhang et al. 2015; Dai et al. 2016; Weng et al. 2016; Rockenbach et al. 2016; Havird et al. 2017; Li et al. 2019; Forsythe et al. 2021).
Heritable symbionts are distinct from organelles in their interaction with the host in that symbiont proteomes are generally considered to be encoded within their genomes, rather than jointly with the nuclear genome. Thus, this route to coadaptation is less important. Nevertheless, symbionts have profound interplay with their host in terms of cell biology, organismal development, physiology and anatomy. In particular, many heritable symbionts form obligate partnerships with their host - such that neither party can live alone. For beneficial symbionts, there are systems to control symbiont number through antimicrobial production (Login et al. 2011), development of specific systems for housing and transmitting symbionts, alongside membrane systems/transporters for metabolite exchange (Feng et al. 2019), which all represent adaptations on the host to house and maintain symbiosis. On the symbiont side, there is loss of cell walls and pathways required for growth outside of the host environment, leading to reliance on host supply of nutrients. An interesting phenomenon is dependence - where the host cannot live without the symbiont for reasons other than the services supplied by the symbiont. One of the first cases of hereditary microbial symbiosis - between bacteria and plants of the family Rubiaceae - is one of these, where loss of the symbiont was observed to impede host development (Miller 1990). Further cases include the requirement of Asobara tabida for a particular strain of Wolbachia to complete oogenesis (Dedeine et al. 2001). These cases likely represent the host evolving around the presence/products of the symbiont, such that symbiont removal results in loss of function.
Coadaptation across the cytoplasm
The study of cytoplasmically inherited agents has acknowledged the diversity of genetic material in the cytoplasm, but rarely examined interactions between the parties. Interactions may be either direct (e.g., a mito-symbiont interaction), or indirect (an evolutionary response in one that impacts the other through coinheritance). Whilst little is known about the former, evidence of indirect impacts is plentiful - selection on one party feeds through to the other inherited elements. For example, the spread of Wolbachia causing CI through a population carries the linked mtDNA haplotype, and the selective sweep reduces mtDNA diversity at the population level (Turelli et al. 1992; Hurst and Jiggins 2005; Deng et al. 2021). Indeed, there are a variety of cases where symbionts are thought to have driven the movement of mtDNA across species boundaries (Turelli et al. 1992; Hurst and Jiggins 2005; Deng et al. 2021). Following spread, the presence of a symbiont at equilibrium in the population has been considered to reduce the effective population of mtDNA to that associated with the fraction which carries the symbiont, conceptually equivalent to background selection removing certain individuals from the pool of individuals out of which mutations arise and spread. More recently, it has been argued the reciprocal pattern is also likely - selective sweeps on mtDNA impact diversity (and indeed frequency and presence) of symbionts (Fenton et al. 2021). Thus, the diversity and population genetics of the cytoplasm should be taken summatively, rather than simply with regard to individual elements.
Conflicts between cytoplasmically inherited elements and their hosts
Maternal inheritance produces an association between symbiont fitness and that of their female, but not male host. This has two primary consequences - the mother’s curse (selection optimises CIEs to female phenotype fitness) and reproductive parasitism (selection optimises CIEs to maximise the production and survival of infected daughters).
Mother’s curse
The theoretical framework for the mother’s curse hypothesis was first described in the 1990s (Frank and Hurst 1996), with further theoretical support proposed a decade later. This framework is simple; the uniparental maternal inheritance of mtDNA means that males are prone to inherit mutations that are selected through the female lineage, even if these mutations are detrimental to males. Consequently, males are expected to accumulate these sexually antagonistic mutations over evolutionary time (Fig. 2A).
Differences in inheritance patterns between nuclear and cytoplasmic elements provides an arena for intergenomic conflict. A Mothers curse hypothesis: maternal inheritance of mitochondria can result in the accumulation of mutations with sexually antagonistic effects in the mtDNA genome. B Cytoplasmic sex ratio distortion in species with separate sexes. Commonly, investment into male and female offspring is equal. Maternal inheritance ties symbiont fitness to the production and survival of female hosts. This is manifested in parthenogenesis induction (left), where all progeny are daughters. Male-killing (middle), where the symbiont kills male progeny it enters, and sibling females have greater access to resources, and higher survival, as a result. Feminisation (right), where the symbiont impacts development in progeny that have a male karyotype such that they differentiate as female hosts. C Cytoplasmic incompatibility is the result of severe miscommunication between cytoplasmic and nuclear genomes, and a classic example of how cytoplasmic elements can spread through a population. When hosts carrying the symbiont (red) mate with uninfected hosts (grey), CI can result in inviable offspring in a unidrectional (left) or bidirectional (right) fashion.
The first experimental evidence for mother’s curse came from a Drosophila study that examined the effects of mtDNA genetic variation on the transcriptomic response (Innocenti et al. 2011). This study used cybrids (cytoplasmic hybrids), in which the mitochondrial genomes from five fly strains sourced from different parts of the world were coupled independently to an isogenic nuclear background, decoupling the effects of mtDNA from those of the nuclear genome. Approximately 10% of the nuclear genome was differentially expressed in males, whereas only a handful of genes were affected in females. Interestingly, these differentially expressed genes were particularly involved in the male reproductive system (testes, accessory glands, ejaculatory duct). Another clear example of mother’s curse came from humans in which a male-biased mutation in the mtDNA resulted in Leber’s hereditary optical neuropathy (Milot et al. 2017). The particular mutation was tracked over a 290-year period, by identifying via genealogical records that it was first recorded in Canada from a woman arriving from France in the 1600s. Given the large male fitness consequences conferred by the mutation, natural selection would be expected to remove this variant from the population, but authors noticed a slight increase in frequency, suggesting a female fitness benefit (Milot et al. 2017). More recently though, the scope of mother’s curse has broadened to not only include reproductive traits, but also other sex-specific life-history traits (Montooth and Dhawanjewar 2019; Nagarajan-Radha et al. 2020; Carnegie et al. 2021).
What is not currently clear is the extent to which mother’s curse impacts maternally inherited elements beyond mitochondria. Conceptually, beneficial heritable symbioses are expected to experience a parallel process in terms of adaptation - the capacity of a maternally inherited symbiont to protect a male host, for instance, derives solely from correlated selection from its impact on female hosts. However, the degree to which maternally inherited beneficial symbioses perform less well in male hosts has not been investigated.
Reproductive parasitism: Investment into female over male progeny and gametes
Both symbionts and mitochondria are known to bias the pattern of host investment into, and survival of, female hosts/gametes over male (see (Hurst and Frost 2015) for review). In hermaphroditic plants, mitochondrial variants impact the development of anthers and pollen formation in the phenotype of cytoplasmic male sterility (CMS). These variants divert resources to reproduction through ovule/seed, and in so doing, promote their own transmission (Fig. 2B).
In arthropods, symbionts variously induce parthenogenetic reproduction (thus ensuring all progeny are female and can transmit the element), feminise hosts that are otherwise ‘programmed’ to male development, or selectively kill male hosts they enter. This last phenotype appears to be spiteful, but is actually an adaptive phenotype when the death of male hosts releases resources directly (through consumption) or indirectly (through relaxed competition) to sibling females (Hurst and Majerus 1993). In ladybirds, for instance, dead male eggs (through which the symbiont cannot be transmitted) are consumed by their sisters (which carry the symbiont and can transmit them). Embryonic male-killing in dioecious species is thus conceptually equivalent to CMS in hermaphrodites, as a source of resource reallocation from male to female reproduction. Male-killing may also occur later in development, and here it is commonly associated with infectious transmission of symbionts from male hosts, with the symbiont showing mixed modes of transmission (maternal inheritance through females, infectious transmission through males (Hurst 1991).
The sex ratio/allocation distorting phenotypes described above have impacts on the individual, but also strong ecological and evolutionary consequences. Parthenogenesis inducing symbionts can spread to fixation, converting the species from sexual to asexual (Stouthamer et al. 1990). Cytoplasmic male sterility, feminisation and male-killing agents can cause strongly female-biased population sex ratios, and these may alter patterns of sexual selection (Jiggins et al. 2000; Charlat et al. 2007) and indeed the capacity of a host to effectively reproduce (Dyson and Hurst 2004). Perhaps most importantly, they engender strong selection on their hosts to restore sex allocation/sex ratio to parity. This is reflected in nuclear restorer genes against mitochondria inducing CMS (Frank 1989), and suppressor genes rescuing male function against male-killers (Hornett et al. 2006) or preventing transmission of feminizers (Rigaud and Juchault 1992). The strongly female-biased sex ratios created when cytoplasmic sex-ratio distorters are common creates intense Fisherian selection for suppression/restorer elements, such that the spread of suppression/restorer genes represents some of the strongest selective events recorded in natural populations (e.g., (Charlat et al. 2007; Hornett et al. 2014)). Further, the evolution of suppression may hide the underlying reproductive parasitism, which may only become apparent in crosses between populations (Hornett et al. 2006), hybridisation (Frank 1989), or for symbionts, transinfection to a novel host species (Sasaki et al. 2002). Indeed, the commonness with which CMS suppression evolves is reflected in the emergence of CMS in about 20% of hybridisation events in plants (Frank 1989).
There are several open questions in our understanding of sex ratio distortion. In terms of incidence, mitochondrial sex allocation distorters are commonly observed in plants, but not in animals. This may be associated with differences in coding capacity/genetics of plant vs. animal genomes (mutational constraint). Symbionts that distort sex ratio/allocation are very commonly observed in arthropods, and have been hypothesised as present in sea urchins (Carrier et al. 2021). Given heritable microbes are common, it is expected that sex ratio distortion would be present in a wider array of host than is currently recognised. In terms of impact, a key emerging question is the nature of restorer and suppression mutations. What are the host systems that are impacted in this co-evolutionary arms race? It has been widely hypothesised these may involve modifications of the sex determination system, as alterations of the signal or target of the symbiont (Hornett et al. 2014). This awaits further discovery of the mechanism of male-killing and of suppression; however, that symbionts alter splicing of key sex determination genes like doublesex supports sex determination as a focus for suppression (Sugimoto et al. 2010).
Reproductive parasitism: Cytoplasmic incompatibility
Cytoplasmic Incompatibility (CI) phenotypes describe the failure of zygote development where the male parent is infected with a symbiont and the female parent either does not have that symbiont, or carries a different strain of the symbiont. Originally described as a phenotype of the Wolbachia in arthropods (Yen and Barr 1971), this phenotype has since been associated with diverse insect symbionts, including Cardinium and Rickettsiella (Hunter et al. 2003; Rosenwald et al. 2020). The symbionts conferring CI spread as it imposes a cost solely on uninfected lineages; the positive frequency-dependent nature of the advantage means invasion either requires the symbiont to reach a threshold frequency, or has an alternate phenotype that allows initial establishment. In contrast to symbionts with sex ratio distorting phenotypes, the symbiont becomes less costly to the host when it is common, simply because symbiont-infected females are unaffected. Indeed, hosts are selected to retain the symbiont to provide immunity against CI when the symbiont is common (Fig. 2C).
Cytoplasmic Incompatibility is a very important phenotype for two reasons. First, it can be applied in the form of release of infected males to suppress target vector populations, and in the form of inoculative release of strains that combine CI and suppress viral replication, to reduce vectorial capacity. Thus, basic research on cytoplasmic symbionts (Hedges et al. 2008; Teixeira et al. 2008) has translated into applied public health protection measures (Utarini et al. 2021). Second, spread of a CI inducing symbiont in one population or species may provide a unidirectional barrier to hybridisation against another, and the spread of distinct strains may produce a bidirectional barrier (Bordenstein et al. 2001). In both cases, the symbiont spread induces reproductive isolation and thus potentiates speciation.
Conclusions
In 1919, Thomas Hunt Morgan in his Principles of Heredity wrote:
‘That there may be substances in the cytoplasm that propagate themselves and are outside the influence of the nucleus, must, of course, be at once conceded as possible despite the fact that, aside from certain plastids, all the Mendelian evidence fails to show that there are such characters’. (Morgan 1919)
By contrast, the past 100 years have created awareness of the vast amount of genetic biodiversity found inhabiting the cytoplasm. Very early on it was noted that these genetic elements did not follow Mendelian inheritance patterns, with inheritance being mostly maternal. One topic that remains a long-standing and unresolved question are why most organelle genomes transmit maternally? Since the initial observations of CIEs, many exceptions to this rule have been discovered, from paternal inheritance of mtDNA in cucumbers, to doubly-uniparental inheritance in bivalves to biparental inheritance in yeast, and symbionts combining infectious and vertical transmission. Still, a predominance for cytoplasmic elements to be inherited via the maternal lineage is evident from the phylogenetic record. The tight association of cytoplasmic genomes with the rest of the cell, plus the non-Mendelian inheritance patterns of CIEs results in fascinating co-evolutionary dynamics that manifest at multiple scales of biological organisation. Thus, the diversity, inheritance, and functional roles of CIEs across eukaryotes remain an important and open question in biology, with fundamental implications for the cells and organisms in which they reside.
References
Adrion JR, White PS, Montooth KL (2016) The roles of compensatory evolution and constraint in aminoacyl tRNA synthetase evolution. Mol Biol Evol 33:152–161
Allio R, Donega S, Galtier N, Nabholz B (2017) Large variation in the ratio of mitochondrial to nuclear mutation rate across animals: implications for genetic diversity and the use of mitochondrial DNA as a molecular marker. Mol Biol Evol 34:2762–2772
Allison TM, Radzvilavicius AL, Dowling DK (2021) Selection for biparental inheritance of mitochondria under hybridization and mitonuclear fitness interactions. Proc Biol Sci 288:20211600
Arrieta-Montiel MP, Mackenzie SA (2011) Plant mitochondrial genomes and recombination. In: Kempken F (ed.) Plant Mitochondria. Springer New York, New York, NY, p 65–82
Ashar FN, Zhang Y, Longchamps RJ, Lane J, Moes A, Grove ML et al. (2017) Association of mitochondrial DNA copy number with cardiovascular disease. JAMA Cardiol 2:1247–1255
Baldo L, Bordenstein S, Wernegreen JJ, Werren JH(2006) Widespread recombination throughout Wolbachia genomes Mol Biol Evol 23:437–449
Barbrook AC, Voolstra CR, Howe CJ (2014) The chloroplast genome of a Symbiodinium sp. clade C3 isolate. Protist 165:1–13
Barr CM, Neiman M, Taylor DR (2005) Inheritance and recombination of mitochondrial genomes in plants, fungi and animals. N. Phytol 168:39–50
Barreto FS, Watson ET, Lima TG, Willett CS, Edmands S, Li W et al(2018) Genomic signatures of mitonuclear coevolution across populations of Tigriopus californicus Nat Ecol Evolution 2:1250–1257
Baur E (1908) Das Wesen und die Erblichkeitsverhältnisse der „Varietates albomarginatae hort.“ vonPelargonium zonale. Z für Indukt Abstamm- und Vererbungslehre 1:330–351
Beck EA, Thompson AC, Sharbrough J, Brud E, Llopart A (2015) Gene flow between Drosophila yakuba and Drosophila santomea in subunit V of cytochrome c oxidase: A potential case of cytonuclear cointrogression. Evolution 69:1973–1986
Bennett GM, Moran NA (2013) Small, smaller, smallest: the origins and evolution of ancient dual symbioses in a Phloem-feeding insect. Genome Biol Evol 5:1675–1688
Bensasson D, Zhang D-X, Hartl DL, Hewitt GM (2001) Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends Ecol Evol 16:314–321
Bentley KE, Mandel JR, McCauley DE (2010) Paternal leakage and heteroplasmy of mitochondrial genomes in Silene vulgaris: evidence from experimental crosses. Genetics 185:961–968
Birky Jr CW (2001) The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu Rev Genet 35:125–148
Birky CW, Strausberg RL, Forster JL, Perlman PS (1978) Vegetative segregation of mitochondria in yeast: Estimating parameters using a random model. Mol Gen Genet 158:251–261
Blokhin A, Vyshkina T, Komoly S, Kalman B (2008) Variations in mitochondrial DNA copy numbers in MS brains. J Mol Neurosci 35:283–287
Bock DG, Andrew RL, Rieseberg LH (2014) On the adaptive value of cytoplasmic genomes in plants. Mol Ecol 23:4899–4911
Boore JL (1999) Animal mitochondrial genomes. Nucleic Acids Res 27:1767–1780
Bordenstein SR, O’Hara FP, Werren JH(2001) Wolbachia-induced incompatibility precedes other hybrid incompatibilities in Nasonia Nature 409:707–710
Boulet L, Karpati G, Shoubridge EA (1992) Distribution and threshold expression of the tRNA(Lys) mutation in skeletal muscle of patients with myoclonic epilepsy and ragged-red fibers (MERRF). Am J Hum Genet 51:1187–1200
Boyd BM, Chevignon G, Patel V, Oliver KM, Strand MR (2021) Evolutionary genomics of APSE: a tailed phage that lysogenically converts the bacterium Hamiltonella defensa into a heritable protective symbiont of aphids. Virol J 18:219
Breton S, Beaupré HD, Stewart DT, Hoeh WR, Blier PU (2007) The unusual system of doubly uniparental inheritance of mtDNA: isn’t one enough? Trends Genet 23:465–474
Breton S, Burger G, Stewart DT, Blier PU (2006) Comparative analysis of gender-associated complete mitochondrial genomes in marine mussels (Mytilus spp.). Genetics 172:1107–1119
Brown TA, Cecconi C, Tkachuk AN, Bustamante C, Clayton DA (2005) Replication of mitochondrial DNA occurs by strand displacement with alternative light-strand origins, not via a strand-coupled mechanism. Genes Dev 19:2466–2476
Brown WM, George Jr M, Wilson AC (1979) Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci USA 76:1967–1971
Broz AK, Waneka G, Wu Z, Fernandes Gyorfy M, Sloan DB (2021) Detecting de novo mitochondrial mutations in angiosperms with highly divergent evolutionary rates. Genetics 218:iyab039
Burton RS (1990) Hybrid breakdown in developmental time in the copepod Tigriopus californicus. Evolution 44:1814–1822
Burton RS, Lee BN(1994) Nuclear and mitochondrial gene genealogies and allozyme polymorphism across a major phylogeographic break in the copepod Tigriopus californicus Proc Natl Acad Sci USA 91:5197–5201
Burzyński A, Zbawicka M, Skibinski DOF, Wenne R(2003) Evidence for recombination of mtDNA in the marine mussel Mytilus trossulus from the Baltic. Mol Biol Evol 20:388–392
Camus MF, Wolff JN, Sgrò CM, Dowling DK(2017) Experimental support that natural selection has shaped the latitudinal distribution of mitochondrial haplotypes in Australian Drosophila melanogaster Mol Biol Evol 34:2600–2612
Carelli V, Maresca A, Caporali L, Trifunov S, Zanna C, Rugolo M (2015) Mitochondria: Biogenesis and mitophagy balance in segregation and clonal expansion of mitochondrial DNA mutations. Int J Biochem Cell Biol 63:21–24
Carnegie L, Reuter M, Fowler K, Lane N, Camus MF(2021) Mother’s curse is pervasive across a large mitonuclear Drosophila panel Evol Lett 5:230–239
Carrier TJ, Leigh BA, Deaker DJ, Devens HR, Wray GA, Bordenstein SR et al. (2021) Microbiome reduction and endosymbiont gain from a switch in sea urchin life history. Proc Natl Acad Sci USA 118:e2022023118
Cato SA, Richardson TE (1996) Inter- and intraspecific polymorphism at chloroplast SSR loci and the inheritance of plastids in Pinus radiata D. Don. Theor Appl Genet 93:587–592
Cerutti H, Johnson AM, Boynton JE, Gillham NW(1995) Inhibition of chloroplast DNA recombination and repair by dominant negative mutants of Escherichia coli RecA Mol Cell Biol 15:3003–3011
Charlat S, Hornett EA, Fullard JH, Davies N, Roderick GK, Wedell N et al. (2007) Extraordinary flux in sex ratio. Science 317:214
Charlat S, Reuter M, Dyson EA, Hornett EA, Duplouy A, Davies N et al. (2007) Male-killing bacteria trigger a cycle of increasing male fatigue and female promiscuity. Curr Biol 17:273–277
Cheng KC, Cahill DS, Kasai H, Nishimura S, Loeb LA (1992) 8-Hydroxyguanine, an abundant form of oxidative DNA damage, causes G-T and A-C substitutions. J Biol Chem 267:166–172
Chevigny N, Schatz-Daas D, Lotfi F, Gualberto JM (2020) DNA repair and the stability of the plant mitochondrial genome. Int J Mol Sci 21:328
Chiu W-L, Stubbe W, Sears BB(1988) Plastid inheritance in Oenothera: organelle genome modifies the extent of biparental plastid transmission Curr Genet 13:181–189
Christie JR, Beekman M (2017) Selective sweeps of mitochondrial DNA can drive the evolution of uniparental inheritance. Evolution 71:2090–2099
Christie JR, Schaerf TM, Beekman M (2015) Selection against heteroplasmy explains the evolution of uniparental inheritance of mitochondria. PLoS Genet 11:e1005112
Ciborowski KL, Consuegra S, García de Leániz C, Beaumont MA, Wang J, Jordan WC (2007) Rare and fleeting: an example of interspecific recombination in animal mitochondrial DNA. Biol Lett 3:554–557
Clark KA, Howe DK, Gafner K, Kusuma D, Ping S, Estes S et al(2012) Selfish little circles: transmission bias and evolution of large deletion-bearing mitochondrial DNA in Caenorhabditis briggsae nematodes PLoS One 7:e41433
Clay Montier LL, Deng JJ, Bai Y (2009) Number matters: control of mammalian mitochondrial DNA copy number. J Genet Genomics 36:125–131
Cleland RE (1963) The cytogenetics of Oenothera. Adv Genet 11:147–237
Cole LW, Guo W, Mower JP, Palmer JD (2018) High and variable rates of repeat-mediated mitochondrial genome rearrangement in a genus of plants. Mol Biol Evol 35:2773–2785
Correns C (1909) Zur kenntnis der rolle von kern und plasma bei der vererbung. Z Indukt Abstamm Vererbungsl 2:331–340
Dahal S, Dubey S, Raghavan SC (2018) Homologous recombination-mediated repair of DNA double-strand breaks operates in mammalian mitochondria. Cell Mol Life Sci 75:1641–1655
Dai B, Guo H, Huang C, Zhang X, Lin Z (2016) Genomic heterozygosity and hybrid breakdown in cotton (Gossypium): different traits, different effects. BMC Genet 17:58
Davila JI, Arrieta-Montiel MP, Wamboldt Y, Cao J, Hagmann J, Shedge V et al. (2011) Double-strand break repair processes drive evolution of the mitochondrial genome in Arabidopsis. BMC Biol 9:64
Day A, Madesis P (2007) DNA replication, recombination, and repair in plastids. In: Bock R (ed) Cell and Molecular Biology of Plastids. Springer Berlin Heidelberg, Berlin, Heidelberg, p 65–119
De Vooght L, Caljon G, Van Hees J, Van Den Abbeele J (2015) Paternal transmission of a secondary symbiont during mating in the viviparous tsetse fly. Mol Biol Evol 32:1977–1980
DeBalsi KL, Hoff KE, Copeland WC (2017) Role of the mitochondrial DNA replication machinery in mitochondrial DNA mutagenesis, aging and age-related diseases. Ageing Res Rev 33:89–104
Dedeine F, Vavre F, Fleury F, Loppin B, Hochberg ME, Bouletreau M (2001) Removing symbiotic Wolbachia bacteria specifically inhibits oogenesis in a parasitic wasp. Proc Natl Acad Sci USA 98:6247–6252
Deng J, Assandri G, Chauhan P, Futahashi R, Galimberti A, Hansson B et al. (2021) Wolbachia-driven selective sweep in a range expanding insect species. BMC Ecol Evol 21:181
Denver DR, Morris K, Lynch M, Vassilieva LL, Thomas WK (2000) High direct estimate of the mutation rate in the mitochondrial genome of Caenorhabditis elegans. Science 289:2342–2344
Dietrich W, Wagner WL, Raven PH (1997) Systematics of Oenothera section Oenothera subsection Oenothera (Onagraceae). Syst Bot Monogr 50:1
Douglas AE (1998) Host benefit and the evolution of specialization in symbiosis. Heredity 81:599–603
Drew GC, Budge GE, Frost CL, Neumann P, Siozios S, Yañez O et al. (2021) Transitions in symbiosis: evidence for environmental acquisition and social transmission within a clade of heritable symbionts. ISME J 15:2956–2968
Dunning Hotopp JC, Clark ME, Oliveira DCSG, Foster JM, Fischer P, Muñoz Torres MC et al. (2007) Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317:1753–1756
Dyson EA, Hurst GDD (2004) Persistence of an extreme sex-ratio bias in a natural population. Proc Natl Acad Sci USA 101:6520–6523
Escalante AA, Freeland DE, Collins WE, Lal AA (1998) The evolution of primate malaria parasites based on the gene encoding cytochrome b from the linear mitochondrial genome. Proc Natl Acad Sci USA 95:8124–8129
Evans BJ, Peter BM, Melnick DJ, Andayani N, Supriatna J, Zhu J et al. (2021) Mitonuclear interactions and introgression genomics of macaque monkeys (Macaca) highlight the influence of behaviour on genome evolution. Proc Biol Sci 288:20211756
Fauré S, Noyer JL, Carreel F, Horry JP, Bakry F, Lanaud C (1994) Maternal inheritance of chloroplast genome and paternal inheritance of mitochondrial genome in bananas (Musa acuminata). Curr Genet 25:265–269
Feng H, Edwards N, Anderson CMH, Althaus M, Duncan RP, Hsu Y-C et al. (2019) Trading amino acids at the aphid–Buchnera symbiotic interface. Proc Natl Acad Sci USA 116:16003–16011
Fenton A, Camus MF, Hurst GDD (2021) Positive selection on mitochondria may eliminate heritable microbes from arthropod populations. Proc Biol Sci 288:20211735
Fernandes Gyorfy M, Miller ER, Conover JL, Grover CE, Wendel JF, Sloan DB et al. (2021) Nuclear-cytoplasmic balance: whole genome duplications induce elevated organellar genome copy number. Plant J 108:219–230
Fontaine KM, Cooley JR, Simon C (2007) Evidence for paternal leakage in hybrid periodical cicadas (Hemiptera: Magicicada spp.). PLoS One 2:e892
Forsythe ES, Sharbrough J, Havird JC, Warren JM, Sloan DB (2019) CyMIRA: The cytonuclear molecular interactions reference for Arabidopsis. Genome Biol Evol 11:2194–2202
Forsythe ES, Williams AM, Sloan DB (2021) Genome-wide signatures of plastid-nuclear coevolution point to repeated perturbations of plastid proteostasis systems across angiosperms. Plant Cell 33:980–997
Frank SA (1989) The evolutionary dynamics of cytoplasmic male sterility. Am Nat 133:345–376
Frank SA, Hurst LD (1996) Mitochondria and male disease. Nature 383:224
Gao L-Z, Liu Y-L, Zhang D, Li W, Gao J, Liu Y et al. (2019) Evolution of Oryza chloroplast genomes promoted adaptation to diverse ecological habitats. Commun Biol 2:278
Gerth M, Martinez-Montoya H, Ramirez P, Masson F, Griffin JS, Aramayo R et al. (2021) Rapid molecular evolution of Spiroplasma symbionts of Drosophila. Microb Genom 7:000503
Ghiselli F, Milani L (2020) Linking the mitochondrial genotype to phenotype: a complex endeavour. Phil Trans Roy Soc B, 375:20190169
Giannakis K, Arrowsmith SJ, Richards L, Gasparini S (2021) Universal features shaping organelle gene retention. bioRxiv
Gray MW, Burger G, Lang BF (1999) Mitochondrial evolution. Science 283:1476–1481
Greiner S, Bock R (2013) Tuning a ménage à trois: Co-evolution and co-adaptation of nuclear and organellar genomes in plants. BioEssays 35:354–365
Greiner S, Rauwolf U, Meurer J, Herrmann RG (2011) The role of plastids in plant speciation. Mol Ecol 20:671–691
Greiner S, Wang X, Rauwolf U, Silber MV, Mayer K, Meurer J et al. (2008) The complete nucleotide sequences of the five genetically distinct plastid genomes of Oenothera, subsection Oenothera: I. Sequence evaluation and plastome evolution †. Nucleic Acids Res 36:2366–2378
Gualberto JM, Newton KJ (2017) Plant mitochondrial genomes: dynamics and mechanisms of mutation. Annu Rev Plant Biol 68:225–252
Gyllensten U, Wharton D, Josefsson A, Wilson AC (1991) Paternal inheritance of mitochondrial DNA in mice. Nature 352:255–257
Haag-Liautard C, Coffey N, Houle D, Lynch M, Charlesworth B, Keightley PD (2008) Direct estimation of the mitochondrial DNA mutation rate in Drosophila melanogaster. PLoS Biol 6:e204
Hadjivasiliou Z, Pomiankowski A, Seymour RM, Lane N (2012) Selection for mitonuclear co-adaptation could favour the evolution of two sexes. Proc Biol Sci 279:1865–1872
Han K-L, Barreto FS (2021) Pervasive mitonuclear coadaptation underlies fast development in interpopulation hybrids of a marine crustacean. Genome Biol Evol 13:evab004
Hartmann AC, Baird AH, Knowlton N, Huang D (2017) The paradox of environmental symbiont acquisition in obligate mutualisms. Curr Biol 27:3711–3716
Havey MJ (1997) Predominant paternal transmission of the mitochondrial genome in cucumber. J Hered 88:232–235
Havey MJ, McCreight JD, Rhodes B, Taurick G (1998) Differential transmission of the Cucumis organellar genomes. Theor Appl Genet 97:122–128
Havird JC, Sloan DB (2016) The roles of mutation, selection, and expression in determining relative rates of evolution in mitochondrial versus nuclear genomes. Mol Biol Evol 33:3042–3053
Havird JC, Trapp P, Miller CM, Bazos I, Sloan DB (2017) Causes and consequences of rapidly evolving mtDNA in a plant lineage. Genome Biol Evol 9:323–336
Havird JC, Whitehill NS, Snow CD, Sloan DB (2015) Conservative and compensatory evolution in oxidative phosphorylation complexes of angiosperms with highly divergent rates of mitochondrial genome evolution. Evolution 69:3069–3081
Hedges LM, Brownlie JC, O’Neill SL, Johnson KN (2008) Wolbachia and virus protection in insects. Science 322:702
Hikosaka K, Watanabe Y-I, Kobayashi F, Waki S, Kita K, Tanabe K (2011) Highly conserved gene arrangement of the mitochondrial genomes of 23 Plasmodium species. Parasitol Int 60:175–180
Hill WG, Robertson A (1966) The effect of linkage on limits to artificial selection. Genetical Res 8:269–294
Himler AG, Adachi-Hagimori T, Bergen JE, Kozuch A, Kelly SE, Tabashnik BE et al. (2011) Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science 332:254–256
Hjort K, Goldberg AV, Tsaousis AD, Hirt RP, Embley TM (2010) Diversity and reductive evolution of mitochondria among microbial eukaryotes. Philos Trans R Soc Lond B Biol Sci 365:713–727
Hornett EA, Charlat S, Duplouy AMR, Davies N, Roderick GK, Wedell N et al. (2006) Evolution of male-killer suppression in a natural population. PLoS Biol 4:e283
Hornett EA, Moran B, Reynolds LA, Charlat S, Tazzyman S, Wedell N et al. (2014) The evolution of sex ratio distorter suppression affects a 25 cM genomic region in the butterfly Hypolimnas bolina. PLoS Genet 10:e1004822
Howe DK, Baer CF, Denver DR (2009) High rate of large deletions in Caenorhabditis briggsae mitochondrial genome mutation processes. Genome Biol Evol 2:29–38
Hunter MS, Perlman SJ, Kelly SE (2003) A bacterial symbiont in the Bacteroidetes induces cytoplasmic incompatibility in the parasitoid wasp Encarsia pergandiella. Proc Biol Sci 270:2185–2190
Hurst LD (1990) Parasite diversity and the evolution of diploidy, multicellularity and anisogamy. J Theor Biol 144:429–443
Hurst LD (1991) The incidences and evolution of cytoplasmic male killers. Proc R Soc Lond Ser B: Biol Sci 244:91–99
Hurst GDD, Frost CL (2015) Reproductive parasitism: maternally inherited symbionts in a biparental world. Cold Spring Harb Perspect Biol 7:a017699
Hurst LD, Hamilton WD (1992) Cytoplasmic fusion and the nature of sexes. Proc R Soc Lond Ser B: Biol Sci 247:189–194
Hurst GDD, Jiggins FM (2005) Problems with mitochondrial DNA as a marker in population, phylogeographic and phylogenetic studies: the effects of inherited symbionts. Proc Biol Sci 272:1525–1534
Hurst GDD, Majerus MEN (1993) Why do maternally inherited microorganisms kill males? Heredity 71:81–95
Innocenti P, Morrow EH, Dowling DK (2011) Experimental evidence supports a sex-specific selective sieve in mitochondrial genome evolution. Science 332:845–848
Jaenike J (2012) Population genetics of beneficial heritable symbionts. Trends Ecol Evol 27:226–232
Jiggins FM, Hurst GD, Majerus ME (2000) Sex-ratio-distorting Wolbachia causes sex-role reversal in its butterfly host. Proc Biol Sci 267:69–73
Kajander OA, Rovio AT, Majamaa K, Poulton J, Spelbrink JN, Holt IJ et al. (2000) Human mtDNA sublimons resemble rearranged mitochondrial genomes found in pathological states. Hum Mol Genet 9:2821–2835
Katju V, Bergthorsson U (2019) Old trade, new tricks: insights into the spontaneous mutation process from the partnering of classical mutation accumulation experiments with high-throughput genomic approaches. Genome Biol Evol 11:136–165
Kaur R, Shropshire JD, Cross KL, Leigh B, Mansueto AJ, Stewart V et al. (2021) Living in the endosymbiotic world of Wolbachia: A centennial review. Cell Host Microbe 29:879–893
Keeling PJ (2010) The endosymbiotic origin, diversification and fate of plastids. Philos Trans R Soc Lond B Biol Sci 365:729–748
King MP, Attardi G (1989) Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 246:500–503
Kino K, Hirao-Suzuki M, Morikawa M, Sakaga A, Miyazawa H (2017) Generation, repair and replication of guanine oxidation products. Genes Environ 39:21
de Koning AP, Keeling PJ (2006) The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol 4:12
Konrad A, Thompson O, Waterston RH, Moerman DG, Keightley PD, Bergthorsson U et al. (2017) Mitochondrial mutation rate, spectrum and heteroplasmy in Caenorhabditis elegans spontaneous mutation accumulation lines of differing population size. Mol Biol Evol 34:1319–1334
Kukat C, Wurm CA, Spåhr H, Falkenberg M, Larsson N-G, Jakobs S (2011) Super-resolution microscopy reveals that mammalian mitochondrial nucleoids have a uniform size and frequently contain a single copy of mtDNA. Proc Natl Acad Sci USA 108:13534–13539
Kvist L, Martens J, Nazarenko AA, Orell M (2003) Paternal leakage of mitochondrial DNA in the great tit (Parus major). Mol Biol Evol 20:243–247
Ladoukakis ED, Eyre-Walker A (2004) Evolutionary genetics: Direct evidence of recombination in human mitochondrial DNA. Heredity 93:321–321
Ladoukakis ED, Zouros E (2001) Recombination in animal mitochondrial DNA: evidence from published sequences. Mol Biol Evol 18:2127–2131
Ladoukakis ED, Zouros E (2017) Evolution and inheritance of animal mitochondrial DNA: rules and exceptions. J Biol Res-Thessaloniki 24:1–7
Lawless C, Greaves L, Reeve AK, Turnbull DM, Vincent AE (2020) The rise and rise of mitochondrial DNA mutations. Open Biol 10:200061
Leclercq S, Thézé J, Chebbi MA, Giraud I, Moumen B, Ernenwein L et al. (2016) Birth of a W sex chromosome by horizontal transfer of Wolbachia bacterial symbiont genome. Proc Natl Acad Sci USA 113:15036–15041
Lederberg J (1952) Cell genetics and hereditary symbiosis. Physiol Rev 32:403–430
Leducq J-B, Henault M, Charron G, Nielly-Thibault L, Terrat Y, Fiumera HL et al. (2017) Mitochondrial recombination and introgression during speciation by hybridization. Mol Biol Evol 34:1947–1959
Lee W-SB, Kochert G (1976) Bacterial endosymbionts in Volvox carteri (Chlorophyceae)1. J Phycol 12:194–197
Leeks A, Dos Santos M, West SA (2019) Transmission, relatedness, and the evolution of cooperative symbionts. J Evol Biol 32:1036–1045
Levin DA (2003) The cytoplasmic factor in plant speciation. sbot 28:5–11
Lewis SC, Joers P, Willcox S, Griffith JD, Jacobs HT, Hyman BC (2015) A rolling circle replication mechanism produces multimeric lariats of mitochondrial DNA in Caenorhabditis elegans. PLoS Genet 11:e1004985
Li M, Schröder R, Ni S, Madea B, Stoneking M (2015) Extensive tissue-related and allele-related mtDNA heteroplasmy suggests positive selection for somatic mutations. Proc Natl Acad Sci USA 112:2491–2496
Li C, Sun X, Conover JL, Zhang Z, Wang J, Wang X et al. (2019) Cytonuclear coevolution following homoploid hybrid speciation in Aegilops tauschii. Mol Biol Evol 36:341–349
Liu X, Longchamps RJ, Wiggins K, Raffield LM, Bielak LF, Zhao W et al. (2020) Association of mitochondrial DNA copy number with cardiometabolic diseases in a large cross-sectional study of multiple ancestries. medRxiv
Login FH, Balmand S, Vallier A, Vincent-Monégat C, Vigneron A, Weiss-Gayet M et al. (2011) Antimicrobial peptides keep insect endosymbionts under control. Science 334:362–365
Longley MJ, Graziewicz MA, Bienstock RJ, Copeland WC (2005) Consequences of mutations in human DNA polymerase gamma. Gene 354:125–131
Lynch M, Blanchard JL (1998) Deleterious mutation accumulation in organelle genomes. Genetica 102–103:29–39
Ma H, O’Farrell PH (2015) Selections that isolate recombinant mitochondrial genomes in animals. Elife 4:e07247
Macey JR, Pabinger S, Barbieri CG, Buring ES, Gonzalez VL, Mulcahy DG et al. (2021) Evidence of two deeply divergent co-existing mitochondrial genomes in the Tuatara reveals an extremely complex genomic organization. Commun Biol 4:116
Magalon H, Nidelet T, Martin G, Kaltz O (2010) Host growth conditions influence experimental evolution of life history and virulence of a parasite with vertical and horizontal transmission. Evolution 64:2126–2138
Maréchal A, Brisson N (2010) Recombination and the maintenance of plant organelle genome stability. N. Phytol 186:299–317
Meiklejohn CD, Montooth KL, Rand DM (2007) Positive and negative selection on the mitochondrial genome. Trends Genet 23:259–263
Meisinger C, Sickmann A, Pfanner N (2008) The mitochondrial proteome: from inventory to function. Cell 134:22–24
Millar AH (2007) The plant mitochondrial proteome. In: Šamaj J, Thelen JJ (eds) Plant Proteomics. Springer Berlin Heidelberg, Berlin, Heidelberg, p 226–246
Miller IM (1990) Bacterial leaf nodule symbiosis. In: Callow JA (Ed.) Advances in Botanical Research. Academic Press, Vol 17 163–234
Milot E, Moreau C, Gagnon A, Cohen AA, Brais B, Labuda D (2017) Mother’s curse neutralizes natural selection against a human genetic disease over three centuries. Nat Ecol Evol 1:1400–1406
Mishmar D, Ruiz-Pesini E, Golik P, Macaulay V, Clark AG, Hosseini S et al. (2003) Natural selection shaped regional mtDNA variation in humans. Proc Natl Acad Sci USA 100:171–176
Mita S, Rizzuto R, Moraes CT, Shanske S, Arnaudo E, Fabrizi GM et al. (1990) Recombination via flanking direct repeats is a major cause of large-scale deletions of human mitochondrial DNA. Nucleic Acids Res 18:561–567
Monickaraj F, Aravind S, Gokulakrishnan K, Sathishkumar C, Prabu P, Prabu D et al. (2012) Accelerated aging as evidenced by increased telomere shortening and mitochondrial DNA depletion in patients with type 2 diabetes. Mol Cell Biochem 365:343–350
Montooth KL, Dhawanjewar AS (2019) Temperature-sensitive reproduction and the physiological and evolutionary potential for Mother’s Curse. Integr Comp Biol 59:890–899
Morales HE, Pavlova A, Joseph L, Sunnucks P (2015) Positive and purifying selection in mitochondrial genomes of a bird with mitonuclear discordance. Mol Ecol 24:2820–2837
Moran NA, McCutcheon JP, Nakabachi A (2008) Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet 42:165–190
Morgan TH (1919) The physical basis of heredity. J. B. Lippincott Co.
Mossman JA, Tross JG, Jourjine NA, Li N, Wu Z, Rand DM (2017) Mitonuclear interactions mediate transcriptional responses to hypoxia in Drosophila. Mol Biol Evol 34:447–466
Muthye V, Lavrov DV (2018) Characterization of mitochondrial proteomes of nonbilaterian animals. IUBMB Life 70:1289–1301
Nabholz B, Glémin S, Galtier N (2007) Strong variations of mitochondrial mutation rate across mammals—the longevity hypothesis. Mol Biol Evol 25:120–130
Nagarajan-Radha V, Aitkenhead I, Clancy DJ, Chown SL, Dowling DK (2020) Sex-specific effects of mitochondrial haplotype on metabolic rate in Drosophila melanogaster support predictions of the Mother’s Curse hypothesis. Philos Trans R Soc Lond B Biol Sci 375:20190178
Neale DB, Marshall KA, Sederoff RR (1989) Chloroplast and mitochondrial DNA are paternally inherited in Sequoia sempervirens D. Don Endl. Proc Natl Acad Sci USA 86:9347–9349
Neale DB, Sederoff RR (1989) Paternal inheritance of chloroplast DNA and maternal inheritance of mitochondrial DNA in loblolly pine. Theor Appl Genet 77:212–216
Neiman M, Taylor DR (2009) The causes of mutation accumulation in mitochondrial genomes. Proc Biol Sci 276:1201–1209
Nikoh N, Hosokawa T, Moriyama M, Oshima K, Hattori M, Fukatsu T (2014) Evolutionary origin of insect–Wolbachia nutritional mutualism. Proc Natl Acad Sci USA 111:10257–10262
Nissanka N, Moraes CT (2020) Mitochondrial DNA heteroplasmy in disease and targeted nuclease-based therapeutic approaches. EMBO Rep. 21:e49612
Nosek J, Novotna M, Hlavatovicova Z, Ussery DW, Fajkus J, Tomaska L (2004) Complete DNA sequence of the linear mitochondrial genome of the pathogenic yeast Candida parapsilosis. Mol Genet Genomics 272:173–180
Nunes MDS, Dolezal M, Schlötterer C (2013) Extensive paternal mtDNA leakage in natural populations of Drosophila melanogaster. Mol Ecol 22:2106–2117
Ogihara Y, Terachi T, Sasakuma T (1988) Intramolecular recombination of chloroplast genome mediated by short direct-repeat sequences in wheat species. Proc Natl Acad Sci USA 85:8573–8577
Oldenburg DJ, Bendich AJ (1996) Size and structure of replicating mitochondrial DNA in cultured tobacco cells. Plant Cell 8:447–461
Osada N, Akashi H (2011) Mitochondrial–nuclear interactions and accelerated compensatory evolution: evidence from the primate cytochrome C oxidase complex. Mol Biol Evol 29:337–346
Otto SP (2007) The evolutionary consequences of polyploidy. Cell 131:452–462
Palmer JD (1983) Chloroplast DNA exists in two orientations. Nature 301:92–93
Palmer JD, Herbon LA (1988) Plant mitochondrial DNA evolved rapidly in structure, but slowly in sequence. J Mol Evol 28:87–97
Passamonti M, Boore JL, Scali V (2003) Molecular evolution and recombination in gender-associated mitochondrial DNAs of the Manila clam Tapes philippinarum. Genetics 164:603–611
Passamonti M, Ghiselli F (2009) Doubly uniparental inheritance: two mitochondrial genomes, one precious model for organelle DNA inheritance and evolution. DNA Cell Biol 28:79–89
Perreau J, Zhang B, Maeda GP, Kirkpatrick M, Moran NA (2021) Strong within-host selection in a maternally inherited obligate symbiont: Buchnera and aphids. Proc Natl Acad Sci USA 118:e2102467118
Petersen MH, Budtz-Jørgensen E, Sørensen SA, Nielsen JE, Hjermind LE, Vinther-Jensen T et al. (2014) Reduction in mitochondrial DNA copy number in peripheral leukocytes after onset of Huntington’s disease. Mitochondrion 17:14–21
Piganeau G, Gardner M, Eyre-Walker A (2004) A broad survey of recombination in animal mitochondria. Mol Biol Evolution 21:2319–2325
Postel Z, Touzet P (2020) Cytonuclear genetic incompatibilities in plant speciation. Plants 9:487
Putintseva YA, Bondar EI, Simonov EP, Sharov VV, Oreshkova NV, Kuzmin DA et al. (2020) Siberian larch (Larix sibirica Ledeb.) mitochondrial genome assembled using both short and long nucleotide sequence reads is currently the largest known mitogenome. BMC Genomics 21:654
Pyle A, Anugrha H, Kurzawa-Akanbi M, Yarnall A, Burn D, Hudson G (2016) Reduced mitochondrial DNA copy number is a biomarker of Parkinson’s disease. Neurobiol Aging 38:216.e7–216.e10
Radzvilavicius AL, Hadjivasiliou Z, Pomiankowski A, Lane N (2016) Selection for mitochondrial quality drives evolution of the germline. PLoS Biol 14:e2000410
Rand DM, Haney RA, Fry AJ (2004) Cytonuclear coevolution: the genomics of cooperation. Trends Ecol Evol 19:645–653
Rand DM, Mossman JA (2020) Mitonuclear conflict and cooperation govern the integration of genotypes, phenotypes and environments. Philos Trans R Soc Lond B Biol Sci 375:20190188
Rebolledo-Jaramillo B, Su MS-W, Stoler N, McElhoe JA, Dickins B, Blankenberg D et al. (2014) Maternal age effect and severe germ-line bottleneck in the inheritance of human mitochondrial DNA. Proc Natl Acad Sci USA 111:15474–15479
Richardson MF, Weinert LA, Welch JJ, Linheiro RS, Magwire MM, Jiggins FM et al. (2012) Population genomics of the Wolbachia endosymbiont in Drosophila melanogaster. PLoS Genet 8:e1003129
Rigaud T, Juchault P (1992) Genetic control of the vertical transmission of a cytoplasmic sex factor in Armadillidium vulgare Latr. (Crustacea, Oniscidea). Heredity 68:47–52
Rockenbach K, Havird JC, Monroe JG, Triant DA, Taylor DR, Sloan DB (2016) Positive selection in rapidly evolving plastid-nuclear enzyme complexes. Genetics 204:1507–1522
Roossinck MJ (2010) Lifestyles of plant viruses. Philos Trans R Soc Lond B Biol Sci 365:1899–1905
Rosenwald LC, Sitvarin MI, White JA (2020) Endosymbiotic Rickettsiella causes cytoplasmic incompatibility in a spider host. Proc Biol Sci 287:20201107
Ross JA, Howe DK, Coleman-Hulbert A, Denver DR, Estes S (2016) Paternal mitochondrial transmission in intra-species Caenorhabditis briggsae hybrids. Mol Biol Evol 33:3158–3160
Ruiz-Pesini E, Mishmar D, Brandon M, Procaccio V, Wallace DC (2004) Effects of purifying and adaptive selection on regional variation in human mtDNA. Science 303:223–226
Sagan L (1967) On the origin of mitosing cells. J Theor Biol 14:255–274
Sambatti JBM, Ortiz-Barrientos D, Baack EJ, Rieseberg LH (2008) Ecological selection maintains cytonuclear incompatibilities in hybridizing sunflowers. Ecol Lett 11:1082–1091
Samuels DC, Li C, Li B, Song Z, Torstenson E, Boyd Clay H et al. (2013) Recurrent tissue-specific mtDNA mutations are common in humans. PLoS Genet 9:e1003929
Sasaki T, Kubo T, Ishikawa H (2002) Interspecific transfer of Wolbachia between two lepidopteran insects expressing cytoplasmic incompatibility: a Wolbachia variant naturally infecting Cadra cautella causes male killing in Ephestia kuehniella. Genetics 162:1313–1319
Sato M, Sato K (2013) Maternal inheritance of mitochondrial DNA by diverse mechanisms to eliminate paternal mitochondrial DNA. Biochim Biophys Acta 1833:1979–1984
Sato K, Sato M (2017) Multiple ways to prevent transmission of paternal mitochondrial DNA for maternal inheritance in animals. J Biochem 162:247–253
Schaack S, Ho EKH, Macrae F (2020) Disentangling the intertwined roles of mutation, selection and drift in the mitochondrial genome. Philos Trans R Soc Lond B Biol Sci 375:20190173
Schneider A, Stelljes C, Adams C, Kirchner S, Burkhard G, Jarzombski S et al. (2015) Low frequency paternal transmission of plastid genes in Brassicaceae. Transgenic Res 24:267–277
Shao R, Kirkness EF, Barker SC (2009) The single mitochondrial chromosome typical of animals has evolved into 18 minichromosomes in the human body louse, Pediculus humanus. Genome Res 19:904–912
Shi P-Q, Chen X-Y, Chen X-S, Lv N, Liu Y, Qiu B-L (2021) Rickettsia increases its infection and spread in whitefly populations by manipulating the defense patterns of the host plant. FEMS Microbiol Ecol 97:fiab032
Silva G, Lima FP, Martel P, Castilho R (2014) Thermal adaptation and clinal mitochondrial DNA variation of European anchovy. Proc Biol Sci 281:20141093
Skibinski DO, Gallagher C, Beynon CM (1994) Sex-limited mitochondrial DNA transmission in the marine mussel Mytilus edulis. Genetics 138:801–809
Sloan DB (2015) Using plants to elucidate the mechanisms of cytonuclear co-evolution. N. Phytol 205:1040–1046
Sloan DB, Alverson AJ, Chuckalovcak JP, Wu M, McCauley DE, Palmer JD et al. (2012) Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol 10:e1001241
Sloan DB, Triant DA, Forrester NJ, Bergner LM, Wu M, Taylor DR (2014) A recurring syndrome of accelerated plastid genome evolution in the angiosperm tribe Sileneae (Caryophyllaceae). Mol Phylogenet Evol 72:82–89
Sloan DB, Warren JM, Williams AM, Wu Z, Abdel-Ghany SE, Chicco AJ et al. (2018) Cytonuclear integration and co-evolution. Nat Rev Genet 19:635–648
Sloan DB, Wu Z (2014) History of plastid DNA insertions reveals weak deletion and at mutation biases in angiosperm mitochondrial genomes. Genome Biol Evol 6:3210–3221
van der Sluis EO, Bauerschmitt H, Becker T, Mielke T, Frauenfeld J, Berninghausen O et al. (2015) Parallel structural evolution of mitochondrial ribosomes and OXPHOS complexes. Genome Biol Evol 7:1235–1251
Smith DR, Keeling PJ (2015) Mitochondrial and plastid genome architecture: Reoccurring themes, but significant differences at the extremes. Proc Natl Acad Sci USA 112:10177–10184
Stampar SN, Broe MB, Macrander J, Reitzel AM, Daly M (2019) Linear mitochondrial genome in anthozoa (Cnidaria): A case study in Ceriantharia. Sci Rep. 9:6094
Stewart DT, Breton S, Blier PU, Hoeh WR (2009) Masculinization events and doubly uniparental inheritance of mitochondrial DNA: A model for understanding the evolutionary dynamics of gender-associated mtDNA in mussels. In Evolutionary Biology. Springer, Berlin, Heidelberg, 163–173
Stewart JB, Chinnery PF (2015) The dynamics of mitochondrial DNA heteroplasmy: implications for human health and disease. Nat Rev Genet 16:530–542
Stouthamer R, Luck RF, Hamilton WD (1990) Antibiotics cause parthenogenetic Trichogramma (Hymenoptera/Trichogrammatidae) to revert to sex. Proc Natl Acad Sci USA 87:2424–2427
Stubbe W (1989) Oenothera—An ideal system for studying the interactions of genome and plastome. Plant Mol Biol Rep. 7:245–257
Sugimoto TN, Fujii T, Kayukawa T, Sakamoto H, Ishikawa Y (2010) Expression of a doublesex homologue is altered in sexual mosaics of Ostrinia scapulalis moths infected with Wolbachia. Insect Biochem Mol Biol 40:847–854
Sung W, Ackerman MS, Miller SF, Doak TG, Lynch M (2012) Drift-barrier hypothesis and mutation-rate evolution. Proc Natl Acad Sci USA 109:18488–18492
Teixeira L, Ferreira A, Ashburner M (2008) The bacterial symbiont Wolbachia induces resistance to RNA viral infections in Drosophila melanogaster. PLoS Biol 6:e2
Timmis JN, Ayliffe MA, Huang CY, Martin W (2004) Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5:123–135
Tin A, Grams ME, Ashar FN, Lane JA, Rosenberg AZ, Grove ML et al. (2016) Association between mitochondrial DNA copy number in peripheral blood and incident CKD in the atherosclerosis risk in communities study. J Am Soc Nephrol 27:2467–2473
Turelli M, Hoffmann AA (1991) Rapid spread of an inherited incompatibility factor in California Drosophila. Nature 353:440–442
Turelli M, Hoffmann AA, McKechnie SW (1992) Dynamics of cytoplasmic incompatibility and mtDNA variation in natural Drosophila simulans populations. Genetics 132:713–723
Utarini A, Indriani C, Ahmad RA, Tantowijoyo W, Arguni E, Ansari MR et al. (2021) Efficacy of wolbachia-infected mosquito deployments for the control of dengue. N. Engl J Med 384:2177–2186
Wagner DB, Dong J, Carlson MR, Yanchuk AD (1991) Paternal leakage of mitochondrial DNA in Pinus. Theor Appl Genet 82:510–514
Wallace DC (1989) Mitochondrial DNA mutations and neuromuscular disease. Trends Genet 5:9–13
Waneka G, Svendsen JM, Havird JC, Sloan DB (2021) Mitochondrial mutations in Caenorhabditis elegans show signatures of oxidative damage and an AT-bias. Genetics 219:iyab116
Waneka G, Vasquez YM, Bennett GM, Sloan DB (2021) Mutational pressure drives differential genome conservation in two bacterial endosymbionts of sap-feeding insects. Genome Biol Evol 13:evaa254
Watanabe K, Yukuhiro F, Matsuura Y, Fukatsu T, Noda H (2014) Intrasperm vertical symbiont transmission. Proc Natl Acad Sci USA 111:7433–7437
Weihe A, Apitz J, Pohlheim F, Salinas-Hartwig A, Börner T (2009) Biparental inheritance of plastidial and mitochondrial DNA and hybrid variegation in Pelargonium. Mol Genet Genomics 282:587–593
Weng M-L, Ruhlman TA, Jansen RK (2016) Plastid–nuclear interaction and accelerated coevolution in plastid ribosomal genes in Geraniaceae. Genome Biol Evol 8:1824–1838
Wickett NJ, Fan Y, Lewis PO, Goffinet B (2008) Distribution and evolution of pseudogenes, gene losses, and a gene rearrangement in the plastid genome of the nonphotosynthetic liverwort, Aneura mirabilis (Metzgeriales, Jungermanniopsida). J Mol Evol 67:111–122
van Wijk KJ, Baginsky S (2011) Plastid proteomics in higher plants: current state and future goals. Plant Physiol 155:1578–1588
Williams AM, Friso G, van Wijk KJ, Sloan DB (2019) Extreme variation in rates of evolution in the plastid Clp protease complex. Plant J 98:243–259
Wilton PR, Zaidi A, Makova K, Nielsen R (2018) A population phylogenetic view of mitochondrial heteroplasmy. Genetics 208:1261–1274
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA 84:9054–9058
Wolff JN, Nafisinia M, Sutovsky P, Ballard JWO (2013) Paternal transmission of mitochondrial DNA as an integral part of mitochondrial inheritance in metapopulations of Drosophila simulans. Heredity 110:57–62
Woloszynska M (2009) Heteroplasmy and stoichiometric complexity of plant mitochondrial genomes—though this be madness, yet there’s method in’t. J Exp Bot 61:657–671
Wu ZQ, Liao XZ, Zhang XN, Tembrock LR, Broz A (2020) Genomic architectural variation of plant mitochondria—A review of multichromosomal structuring. J Syst Evol 60:160–168
Wu Z, Sloan DB (2019) Recombination and intraspecific polymorphism for the presence and absence of entire chromosomes in mitochondrial genomes. Heredity 122:647–659
Wu Z, Waneka G, Broz AK, King CR, Sloan DB (2020) MSH1 is required for maintenance of the low mutation rates in plant mitochondrial and plastid genomes. Proc Natl Acad Sci USA 117:16448–16455
Xia H, Zhao W, Shi Y, Wang X-R, Wang B (2020) Microhomologies are associated with tandem duplications and structural variation in plant mitochondrial genomes. Genome Biol Evol 12:1965–1974
Yen JH, Barr AR (1971) New hypothesis of the cause of cytoplasmic incompatibility in Culex pipiens L. Nature 232:657–658
Zhang J, Ruhlman TA, Sabir J, Blazier JC, Jansen RK (2015) Coordinated rates of evolution between interacting plastid and nuclear genes in Geraniaceae. Plant Cell 27:563–573
Zouros E (2000) The exceptional mitochondrial DNA system of the mussel family Mytilidae. Genes Genet Syst 75:313–318
Zouros E (2013) Biparental inheritance through uniparental transmission: the doubly uniparental inheritance (DUI) of mitochondrial DNA. Evol Biol 40:1–31
Acknowledgements
We thank three anonymous reviewers who made very helpful suggestions to improve this manuscript.
Funding
MFC was funded by a UKRI Fellowship (NE/V014307/1) and a Leverhulme Trust grant (RPG-2019-109). JS was funded by the National Science Foundation (DEB – 1753695, DEB – 1753851, IOS – 1829176, IOS – 2145811). BAL and JS were funded by the New Mexico Institute of Mining and Technology. GDDH was funded by a UKRI grant (NE/S012346/1).
Author information
Authors and Affiliations
Contributions
All authors contributed to the ideas presented and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Barbara Mable.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Camus, M.F., Alexander-Lawrie, B., Sharbrough, J. et al. Inheritance through the cytoplasm. Heredity 129, 31–43 (2022). https://doi.org/10.1038/s41437-022-00540-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00540-2
This article is cited by
-
Genesis of ectosymbiotic features based on commensalistic syntrophy
Scientific Reports (2024)
-
Egg multivesicular bodies elicit an LC3-associated phagocytosis-like pathway to degrade paternal mitochondria after fertilization
Nature Communications (2024)
-
Search for source of invasive-range populations of a haplodiploid pest species reveals Wolbachia-driven mitochondrial selective sweep in native range
Journal of Pest Science (2024)
-
Classification of Isatis indigotica Fortune and Isatis tinctoria Linnaeus via comparative analysis of chloroplast genomes
BMC Genomics (2023)
-
Oil yield prediction for sunflower hybrid selection using different machine learning algorithms
Scientific Reports (2023)