Abstract
The cost of parentage assignment precludes its application in many selective breeding programmes and molecular ecology studies, and/or limits the circumstances or number of individuals to which it is applied. Pooling samples from more than one individual, and using appropriate genetic markers and algorithms to determine parental contributions to pools, is one means of reducing the cost of parentage assignment. This paper describes and validates a novel maximum likelihood (ML) parentage-assignment method, that can be used to accurately assign parentage to pooled samples of multiple individuals—previously published ML methods are applicable to samples of single individuals only—using low-density single nucleotide polymorphism (SNP) ‘quantitative’ (also referred to as ‘continuous’) genotype data. It is demonstrated with simulated data that, when applied to pools, this ‘quantitative maximum likelihood’ method assigns parentage with greater accuracy than established maximum likelihood parentage-assignment approaches, which rely on accurate discrete genotype calls; exclusion methods; and estimating parental contributions to pools by solving the weighted least squares problem. Quantitative maximum likelihood can be applied to pools generated using either a ‘pooling-for-individual-parentage-assignment’ approach, whereby each individual in a pool is tagged or traceable and from a known and mutually exclusive set of possible parents; or a ‘pooling-by-phenotype’ approach, whereby individuals of the same, or similar, phenotype/s are pooled. Although computationally intensive when applied to large pools, quantitative maximum likelihood has the potential to substantially reduce the cost of parentage assignment, even if applied to pools comprised of few individuals.
Similar content being viewed by others
Introduction
This paper describes modifications to well-established maximum likelihood (ML) parentage-assignment methods to allow parentage to be assigned to pooled samples of multiple individuals—previously published ML methods are applicable to samples of single individuals only. This novel approach uses quantitative genotype data (also referred to as continuous genotype data; Clark et al. 2019) from low-density single nucleotide polymorphism (SNP) panels (Henshall et al. 2014). Quantitative genotype data are comprised of continuous numerical genotypes, reflecting the probabilities of all possible genotype classes (i.e., ordered allele combinations or unordered allele counts), in the form of genotype probability matrices or vectors. The use of quantitative genotype data for parentage assignment negates the need to call discrete genotype classes (Kalinowski et al. 2010; Marshall et al. 1998; Meagher and Thompson 1986). This is particularly important when assigning parentage to samples comprised of multiple individuals (i.e. ‘pools’), as it negates the need to call discrete genotype classes Marshall et al. 1998; Meagher and Thompson 1986). Genotype data from pools present like polyploid data and, as for polyploids, calling discrete genotypes is prone to error due to the large number of possible genotype classes at each SNP (Clark et al. 2019; Rahman et al. 2015).
Genetic markers are used for individual parentage assignment in pedigree-based selective breeding programmes: (i) where tracking identities over the life of individuals is difficult, expensive or not possible, a circumstance common in aquaculture species (Henshall et al. 2014; Kinghorn et al. 2010; Vandeputte and Haffray 2014); (ii) where only one parent is known with certainty, such as in the case of multiple-sire joining in livestock (Henderson 1988), poly cross families in trees (Burdon and Shelbourne 1971) and aggregated full-sib families in aquaculture (Hamilton et al. 2009); or (iii) for quality-control/identity-recovery purposes (Grattapaglia et al. 2014; Hansen and Kjaer 2006). Parentage assignment is also widely used in molecular ecology, including the study of conservation biology, dispersal and recruitment patterns, quantitative genetics and sexual selection (Flanagan and Jones 2019; Jones et al. 2010). A variety of genetic markers have been adopted for parentage assignment in selective breeding programmes and molecular ecology (Flanagan and Jones 2019; Jones et al. 2010). However, SNPs are increasingly used for this purpose, due to their capacity for high-speed high-throughput screening, low mutation rate, low genotyping error rate, high prevalence in the genome, and decreasing the cost of development and implementation (Hauser et al. 2011; Liu et al. 2016). A range of assays, platforms and algorithms are available for low-density SNP genotyping—most commonly using estimates of signal intensity (or area), for two axes, X and Y, corresponding to alleles A and B, from ‘intensity-based’ assays (Henshall et al. 2014; Rahman et al. 2015; Semagn et al. 2014) or counts of allele reads from genotyping-by-sequencing (GBS) assays (Clark et al. 2019), to call discrete genotypes.
Although declining, the cost of parentage assignment per individual precludes its application in many selective breeding programmes, and/or limits the circumstances or number of individuals to which it is applied. For example, further reduction in the cost of parentage assignment per individual would potentially allow: (i) pedigree-based selective breeding in additional species; (ii) an increase in the number of families and individuals able to be generated, retained and measured in current selective breeding programmes, with associated increases in the accuracy of selection; (iii) an increase in the number of environments across which progeny tests can feasibly be conducted; and/or (iv) the genetic analysis of additional traits (Bell et al. 2017; Henshall et al. 2014; Kinghorn et al. 2010; Vandeputte and Haffray 2014). Pooling samples from more than one individual, and using appropriate genetic markers and algorithms to determine parental contributions to pools, are means of further reducing the cost of parentage assignment in selective breeding programmes and molecular ecology studies (Henshall et al. 2014; Kinghorn et al. 2010).
Two distinct approaches to pool construction and parentage assignment are addressed in this paper: (i) ‘pooling-for-individual-parentage-assignment’, whereby each individual in a pool is tagged or traceable and from a known and mutually exclusive set of possible parents; and (ii) ‘pooling-by-phenotype’, whereby individuals of the same, or similar, phenotype/s are pooled (Bell et al. 2017; Henshall et al. 2012; Kinghorn et al. 2010) into bins (i.e. classes) of, ideally, equal width to minimise heterogeneity in measurement error. Furthermore, a novel ML parentage-assignment method using quantitative genotypes that can be used to accurately assign parentage to pools is described. This approach uses the same rationale underpinning ML parentage assignment using discrete genotypes (Kalinowski et al. 2010; Marshall et al. 1998)—in that it determines the likelihood of obtaining a certain pool (or individual offspring) genotype, given a set of parental genotypes—but uses quantitative genotypes in the place of discrete genotypes. ML parentage assignment using quantitative genotypes (herein referred to as ‘quantitative ML’) is validated, using simulated data, and compared with three alternative methods—ML parentage assignment using discrete genotypes (‘discrete ML’); exclusion (Chakraborty et al. 1974); and solving the weighted least squares problem (Henshall et al. 2014; Kinghorn et al. 2010).
Materials and methods
Computation of quantitative genotypes using intensity-based assays
For intensity-based assays, a method to compute quantitative genotypes, extended from Henshall et al. (2014), is outlined in Fig. 1, with a worked example provided in Supplementary Materials 1. Methods to compute quantitative genotypes (Fig. 1)—that is, the parent genotype probability matrices (Gij) and pool unordered genotype probability vectors (gkj)—using GBS are detailed elsewhere (Clark et al. 2019).
For intensity-based SNP assays, the mean and standard deviation of allelic proportions for each genotype of each SNP (i.e., individual allelic proportion parameters—refer to Fig. 1 of Henshall et al. 2014—and pool allelic proportion parameters) are required to account for normally distributed random errors in allelic proportions in the computation of parent genotype probability matrices and pool unordered genotype probability vectors (Fig. 1). These are estimated from individual allele intensities and individual genotype calls from tissue and DNA samples of individuals (i.e., not pools). Individual samples (i.e. individual parent samples or non-parent individual samples; Fig. 1) should be sourced from the same, or a closely related, population to that for which parentage assignment is being undertaken, so as to maximise the probability that data for relevant loci and alleles are captured.
Estimation of individual allelic proportion parameters
Herein the approach and notation used to compute quantitative genotypes from intensity-based assays, accounting for normally distributed random errors in allelic proportions, is extended from Henshall et al. (2014). Accordingly, for a given individual and SNP, the individual allelic proportion (pij) takes a value between zero and one, corresponding to the polar coordinate range from 0 to \(\frac{\pi }{2}\) (Henshall et al. 2014):
where pij is the individual allelic proportion (for details refer to the Appendix of Henshall et al. 2014), and a1ij and a2ij are the individual allelic intensities for allele A and B for individual sample i and SNP j (adjusted for ‘uncertainty’ where applicable—refer to Eq. (1) of Henshall et al. 2014). Conceptually, individual allelic intensities should not be less than zero and individual allelic proportions should range between 0 and 1 but minor deviations outside these parameter boundaries can be accommodated by Eq. (1). Note that software associated with some genotyping platforms directly output a variable that is conceptually equivalent to individual (or pool) allelic proportion (e.g., ‘B-allele frequency’, refer to Bell et al. 2017).
For samples from diploid individuals, discrete genotype classes can be reliably called using proprietary or other software appropriate to a given genotyping platform (Henshall et al. 2014). Individual allelic proportion parameters, means (\(\overline x _{AAj}\), \(\overline x _{ABj}\) and \(\overline x _{BBj}\)) and standard deviations (sAAj,sABj and sBBj) by genotype, can then be estimated for each SNP from a sample of individuals. These individual allelic proportion parameters are specific to the genotyping assay, platform and method used.
Estimation of pool allelic proportion parameters
Pool allelic proportion parameters for each SNP can be approximated from individual allelic proportion parameters, as follows. First, means of allelic proportions for homozygous A (\(\overline x _{jA}\)) and homozygous B (\(\overline x _{jB}\)) genotypes may be assumed to equal \(\overline x _{jAA}\) and \(\overline x _{jBB}\), respectively, and sjA and sjB to equal sjAA and sjBB, respectively. Second, mean and standard deviation of allelic proportion for the heterozygous genotypes with equal counts of A and B alleles may be assumed to equal \(\overline x _{jAB}\) and sjAB, respectively. Finally, means and standard deviations of allelic proportion for heterozygous genotypes with unequal counts of A and B alleles may be weighted according to allele counts. For example, for a pool size of three (i.e., three individuals in a pool): \(\overline x _{jAAAAAB} \approx \frac{2}{3}\overline x _{jAA} + \frac{1}{3}\overline x _{jAB}\) and \(s_{jAAAAAB} \approx \sqrt {\frac{2}{3}s_{jAA}^2 + \frac{1}{3}s_{jAB}^2}\), \(\overline x _{jAAAABB} \approx \frac{1}{3}\overline x _{jAA} + \frac{2}{3}\overline x _{jAB}\) and \(s_{jAAAABB} \approx \sqrt {\frac{1}{3}s_{jAA}^2 + \frac{2}{3}s_{jAB}^2}\), \(\overline x _{jAABBBB} \approx \frac{2}{3}\overline x _{jAB} + \frac{1}{3}\overline x _{jBB}\) and \(s_{jAABBBB} \approx \sqrt {\frac{2}{3}s_{jAB}^2 + \frac{1}{3}s_{jBB}^2}\); and \(\overline x _{jABBBBB} \approx \frac{1}{3}\overline x _{jAB} + \frac{2}{3}\overline x _{jBB}\) and \(s_{jABBBBB} \approx \sqrt {\frac{1}{3}s_{jAB}^2 + \frac{2}{3}s_{jBB}^2}\). Note that only parameters for unordered allele combinations (i.e., ‘unordered genotypes’—genotypes corresponding to allele counts) are required. For example, the unordered genotype AAAAAB corresponds to the ordered genotypes of AAAAAB, AAAABA, AAABAA, AABAAA, ABAAAA and BAAAAA.
Alternative approaches to estimating pool allelic proportion parameters can be conceived. For example, they could be estimated directly from pools with known SNP genotypes.
Computing quantitative genotypes
For a given parent and SNP, each element of the parent genotype probability matrix (Gij) represents the probability that the true genotype is equal to the corresponding ordered genotype. Adopting the approach of Henshall et al. (2014), for each parent i and SNP j a quantitative genotype probability matrix (Gij) can be computed:
where λijAA is the height of the N(\(\overline x _{j_{AA}},s_{j_{AA}}\)) distribution at x = pij, λijAB and λijBA are the height of the N(\(\overline x _{j_{AB}},s_{j_{AB}}\)) distribution at x = pij, λijBB is the height of the N(\(\overline x _{j_{BB}},s_{j_{BB}}\)) distribution at x = pij and Σλij is the sum of λijAA, λijAB and λijBB.
Extending the method of Henshall et al. (2014), for each pool and SNP, a pool unordered genotype probability vector (gkj)—elements of which represents the probability that the true genotype is equal to the corresponding unordered genotype—can be computed by modifying Eq. (2) to accommodate the additional genotypes possible in pools:
where gkj is a vector with 2n + 1 elements, corresponding to the number of possible unordered genotypes for pool k and SNP j, n is the pool size, and ° is the Hadamard or entrywise product. The elements λkj1 to λkj2n+1 are computed as the height at the corresponding N(\(\overline x _{j_1},s_{j_1}\)) to N(\(\overline x _{j_{2n + 1}},s_{j_{2n + 1}}\)) distributions at x = pkj (where pkj is the allelic proportion for pool k, computed according to Eq. (1), replacing i with k). The elements \({\uprho}_1^{ - 1}\) to \({\uprho}_{2n + 1}^{ - 1}\) are computed as the inverse of the number of possible ordered genotypes for the unordered genotypes 1 to 2n + 1. For example in the case of two individuals per pool (n = 2): λkj1 = λkjAAAA = height of the \({\mathrm{N}}\left( {\overline x _{j_{AAAA}},s_{j_{AAAA}}} \right)\) distribution at x = pkj and ρ1 = ρAAAA = 1 (the one ordered genotype is AAAA); λkj2 = λkjAAAB = height of the \({\mathrm{N}}\left( {\overline x _{j_{AAAB}},s_{j_{AAAB}}} \right)\) distribution at x = pkj and ρ2 = ρAAAB = 4 (the four ordered genotypes are AAAB, AABA, ABAA and BAAA); λkj3 = λkjAABB = height of the \({\mathrm{N}}\left( {\overline x _{j_{AABB}},s_{j_{AABB}}} \right)\) distribution at x = pkj and ρ3 = ρAABB = 6 (the six ordered genotypes are AABB, ABAB, BAAB, ABBA, BABA, BBAA); λkj4 = λkjABBB = height of the \({\mathrm{N}}\left( {x_{j_{ABBB}},s_{j_{ABBB}}} \right)\) distribution at x = pkj and ρ4 = ρABBB = 4 (the four ordered genotypes are ABBB, BABB, BBAB, BBBA); and λkj5 = λkjBBBB = height of the \({\mathrm{N}}\left( {\overline x _{j_{BBBB}},s_{j_{BBBB}}} \right)\) distribution at x = pkj and ρ5 = τBBBB = 1 (the one ordered genotype is BBBB). Note that in the case of one individual per pool (n = 1), the elements of a pool unordered genotype probability vector are equal to the elements of the upper triangle of a corresponding quantitative genotype probability matrix for an individual offspring (i.e. gkj is equivalent to the upper triangle of Goj from Henshall et al. 2014, expressed as vector). Where data for a SNP are missing in a pool, values of gkj can be assumed to equal the expected unordered transmission vector (nnj), where nnj equals tcj, assuming tij equals fj for all 2n parents (detailed below).
ML parentage assignment using quantitative genotypes
Inputs
Required inputs to undertake quantitative ML are the parent genotype probability matrices (Gij) and the pool unordered genotype probability vectors (gkj), described above; and a list of possible parental combinations in the pool, an assumed genotype replacement error rate (\(\widehat {\varepsilon _{nj}}\)) and a critical Δ log odds (LOD) value (Fig. 2; Supplementary Material 1). The list of possible parental combinations in the pool may comprise all combinations of all possible parents but in many circumstances a shorter list can be considered, substantially reducing the computational burden. For example, in the case of pooling-for-individual-parentage-assignment, pools are constructed such that the list of possible parental combinations in a pool must include two, and only two, parents from each known and mutually exclusive set of possible parents contributing to the pool. The assumed genotype replacement error rate can theoretically range between zero and one but is generally small (e.g., <0.01). Critical Δ LOD represents a pedigree assignment acceptance threshold corresponding to a desired level of confidence in assignment (i.e., a predetermined acceptable proportion of erroneous parentage assignments)—the greater the critical Δ LOD, the lower the erroneous assignment rate in accepted assignments, but the greater the number of falsely rejected assignments (Harrison et al. 2013; Jones et al. 2010). Although computationally burdensome, critical Δ LOD is most appropriately determined by simulating populations of parents and pools (Jones et al. 2010; Kalinowski et al. 2010; Marshall et al. 1998), given an assumed genotype replacement error rate and possible parental combinations in pools.
Computation of the LOD score
Intermediate vectors required to compute the LOD score (Henshall et al. 2014; Meagher and Thompson 1986), and ultimately assign parentage, include the parent allele transmission vectors (tij), parent combination ordered transmission vectors (qcj) and parent combination unordered transition vectors (tcj; Fig. 2). For a given parent, i, and SNP, j, each element of the 2 × 1 parent allele transmission vector represents the probability that the corresponding allele (i.e., A or B) will be transmitted to its progeny. The parent allele transmission vector (tij) for parent i and SNP j can be computed as (Henshall et al. 2014):
When marker data for a SNP from a parent is missing, the corresponding parent allele transmission vector may be assumed to equal the vector of expected allele frequencies transmitted to pools for SNP j (fj). The expected allele frequencies are most simply estimated by computing the average of parental tij vectors, excluding missing data, for SNP j (Fig. 2). However, less generic, but more precise, approaches to the computation of expected allele frequencies can be conceived. For example, if expected parental contributions to pools are known, expected allele frequencies could be computed separately for each pool as the average of parental tij vectors weighted by their expected parental contribution to the pool.
For each possible combination of parents, c, and SNP, j, elements of the 1 × 2n parent combination ordered transmission vector (qcj; Fig. 2) represent the probability that the corresponding ordered genotype will be transmitted to progeny. The parent combination ordered transmission vector can be computed as the Kronecker product, ⊗, of t1j,…, t2nj. That is:
where 2n is the number of parents for possible parent combination c.
It is noteworthy that the length of qcj increases exponentially by 2n (i.e., twice the pool size). However, qcj vectors can be consolidated into parent combination unordered transition vectors (tcj) of length 2n + 1, substantially reducing the computational burden. For example, if the pool size is three (n = 3), elements corresponding to the ordered genotypes AAAAAB, AAAABA, AAABAA, AABAAA, ABAAAA, and BAAAAA in qcj can be summed with the result becoming the element in tcj corresponding to the unordered genotype AAAAAB.
In the implementation of previously described ML parentage-assignment methods, genotyping errors are accounted for by assuming they represent the replacement of true genotypes with genotypes selected at random under Hardy–Weinberg assumptions in a small proportion of SNPs (i.e., the assumed genotype replacement error rate) (Henshall et al. 2014; Jones et al. 2010; Kalinowski et al. 2010; Marshall et al. 1998). Such errors can be similarly accounted for in quantitative ML by adjusting the parent combination unordered transition vectors (tcj) and pool unordered genotype probability vectors (gkj):
where \(\widehat {\varepsilon _{nj}}\) is the assumed genotype replacement error rate for pool size n and SNP j.
For each pool with genotype probability vector \({\boldsymbol{g}}_ \ast ^{{\boldsymbol{kj}}}\), the likelihood of each possible parent combination and pool duo for each SNP can be computed as:
To compute the likelihood ratio \(\left( {\frac{{L^{\left( {ck} \right)j}}}{{L^{cj}}}} \right)\), the denominator is computed as the likelihood under the null hypothesis that the individuals in the pool are unrelated to the possible parent combination in question:
The likelihood ratio across all markers can then be computed as the sum of the natural log of the likelihood ratios, herein referred to as the LOD score:
where m is the number of SNPs.
Parentage assignment
The most likely parental combination for each pool can be identified as that with the greatest LODck. The Δ LOD for each parent in the most likely parental combination can then be computed as the difference between the LODck for the most likely parental combination and the maximum LODck of those parental combinations that were identical to the most likely combination except for the parent in question. If a parent’s Δ LOD is greater than critical Δ LOD it can then be accepted as correctly assigned.
Validation
An R package (R Core Team 2020) entitled ‘SNPpools’, available at https://github.com/mghamilton/SNPpools, was developed to implement and validate, with simulations, the quantitative ML method. Simulations were conducted for pool sizes of one (i.e., samples of individuals), two and three using the sim.parent.assign.fun function of the SNPpools package.
Generation of simulated datasets for validation and comparison
Two pedigree structures (‘Pedigree 1’ and ‘Pedigree 2’) were simulated. Pedigree 1 was constructed assuming eight dams, each crossed with eight mutually exclusive sets of ten sires (i.e. 80 sires). This pedigree structure is akin to the case of multiple-sire joining in livestock (Henderson 1988), poly cross families in trees (Burdon and Shelbourne 1971) and aggregated full-sib families in aquaculture (Hamilton et al. 2009), where the one parent is known and the other is known to be one of a finite set of possible parents. Pedigree 2 was constructed assuming 40 sires and 40 dams each crossed with one individual of the opposite sex to produce 40 full-sib families. This pedigree structure represents a mating design applicable to species where half-sib families are difficult to generate, such as shrimp (Dai et al. 2020). In both pedigree structures, all parents were assumed to be unrelated (scenarios involving half the number of possible parents/families and related parents are explored in Supplementary Materials 2 and 3, respectively).
Pedigree 1 pools were constructed to allow ‘pooling-for-individual-parentage assignment’ and for Pedigree 2 pools were constructed assuming a ‘pooling-by-phenotype’ strategy. That is, for Pedigree 1, pools were constructed ensuring that the progeny of each dam was represented no more than once in each pool. For Pedigree 2, no constraints on the ancestry of individuals placed in pools were applied—that is, individuals from families were allocated to pools at random. For each pooling strategy and pool size, parentage was assigned to the progeny of 9600 ‘unknown parents’.
Generation of simulated datasets for validation and comparison
For each simulated sample (parent or pool) and 100 SNP, individual allele intensities were generated (scenarios involving 50 SNP and 200 SNP are explored in Supplementary Materials 4 and 5, respectively). First, ‘true’ genotypes were randomly generated for parents assuming that SNPs were biallelic, SNPs were in linkage equilibrium (i.e., inherited independently) and the allele frequency for all SNPs was 0.5 (a scenario in which the allele frequency for all SNPs was 0.4 is explored in Supplementary Materials 6). Second, true genotypes for progeny were then generated from true parental genotypes assuming alleles were inherited according to Mendelian principles. Finally, pool genotypes were generated by concatenating the true genotypes of individuals in the pools (e.g., a pool of two individuals with genotypes AB and BB had a pool genotype of ABBB).
To account for lab-based genotyping errors, each SNP for each individual and pool was randomly categorised as correct (with probability 1 − εj) or erroneous (with probability εj), where εj is the true genotype replacement error rate for SNP j (Henshall et al. 2014). If categorised as erroneous, a random ‘observed genotype’, which may differ from the ‘true genotype’, was generated. For the purpose of simulation the true genotype replacement error rate was assumed to equal 0.01—a somewhat large and thus conservative value—for all SNPs and samples.
For each parent and SNP, individual allelic proportions (pij) were generated as random normal variables:
where μjq is the mean of the individual allelic proportion for the observed genotype of parent sample i, SNP j and genotype q; and σjq is the corresponding standard deviation. Assumed true inidividual allelic proportion parameters for all SNP were as follows: μAA = 0.05, μAB = 0.50, μAB = 0.05, μBB = 0.95 and σBB = 0.05 (scenarios in which standard deviations—σAA, σσAB and σBB—were assumed to equal 0.01 and 0.20 are explored in Supplementary Materials 7 and 8, respectively). Individual allele intensities were subsequently calculated as follows:
Pool allelic proportions (pkj) and pool allele intensities (a1kj and a2kj) where correspondingly computed.
For each simulated parent and pool, each SNP was randomly categorised as missing (with probability 0.1) or present (with probability 0.9). For those SNPs categorised as missing, allele intensities (i.e., a1ij, a2ij, a1kj and/or a2kj) were deleted (scenarios in which all SNP from some parents were categorised as missing are explored in Supplementary Materials 9).
Parentage assignment
For each simulated pool, quantitative ML was undertaken considering only possible parental combinations, and the assigned parents compared with the true parents of each pool. For comparison, parentage assignment was also undertaken for each pool by extending three previously described methods (Henshall et al. 2014). First, a ML parentage assignment using discrete genotypes approach was implemented. This method, ‘discrete ML’, was equivalent to quantitative ML, except that the quantitative parent genotype probability matrix (Gij) was replaced with a discrete genotype probability matrix, Dij, where \({\boldsymbol{D}}^{{\boldsymbol{ij}}} = \left[ {\begin{array}{*{20}{c}} 1 & 0 \\ 0 & 0 \end{array}} \right],\left[ {\begin{array}{*{20}{c}} 0 & {1/2} \\ {1/2} & 0 \end{array}} \right]\), or \(\left[ {\begin{array}{*{20}{c}} 0 & 0 \\ 0 & 1 \end{array}} \right]\). To compute Dij, if an element the corresponding Gij was greater than 0.98, for homozygous genotypes (i.e., diagonal elements corresponding to genotypes AA and BB), or 0.49 for heterozygous genotypes (i.e., off-diagonal elements corresponding to genotypes AB and BA), were replaced with 1 and 1/2, respectively, with all other elements equal to zero (Henshall et al. 2014). In addition, also using a threshold of 0.98, the pool unordered genotype probability vector gkj was replaced with dkj. Secondly, an exclusion method was applied. Using this method, the number of genotype mismatches, computed as the number of SNPs where \({\boldsymbol{t}}^{{\boldsymbol{cj}}}{\boldsymbol{d}}^{{\boldsymbol{kj}}^{\prime} } = 0\), for each pool and possible parental combination was calculated. The possible parental combination with the lowest proportion of mismatched SNPs (and lowest standard error of proportion, where multiple combinations with the same proportion were identified) was then identified. Finally, parental contributions to pools were estimated by solving the ‘weighted least squares problem’, as detailed in Henshall et al. (2014). The possible parental combination resulting in the minimum sum of squared difference from these estimated parental contributions was then assigned to the pool. Simple worked examples of the ‘quantitative ML’, ‘discrete ML’, ‘exclusion’ and ‘least squares’ methods of parentage assignment are provided in Supplementary Materials 1.
Results
Simulations revealed that not only can the quantitative ML method be used to assign parentage to pools, but the quantitative ML method was more accurate than discrete ML and exclusion under every scenario examined and was generally more accurate than the least squares method (Tables 1 and 2; Figs. 3 and 4; Supplementary Materials 10). This indicates that the quantitative ML method may be adopted to reduce the cost and or increase the accuracy of parentage assignment in selective breeding programmes and molecular ecology studies in many circumstances.
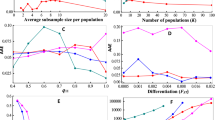
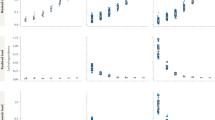
Histograms were generated using a pooling-for-individual-parentage-assignment approach—of one (a, b), two (c, d) and three (e, f) individuals. Results using quantitative maximum likelihood (a, c and e) and discrete maximum likelihood (b, d and f) parentage assignment are shown—correctly assigned individuals are in grey and incorrectly assigned individuals are represented as a black line. Where applicable, the critical Δ LOD value to achieve a 99% correct assignment rate is indicated by an arrow with the percentage of rejected assignments in parentheses.
Histograms were generated using a pooling-by-phenotype approach—of one (a, b), two (c, d) and three (e, f) individuals. Results using quantitative maximum likelihood (a, c and e) and discrete maximum likelihood (b, d and f) parentage assignment are shown—correctly assigned individuals are in grey and incorrectly assigned individuals are represented as a black line. Where applicable, the critical Δ LOD value to achieve a 99% correct assignment rate is indicated by an arrow with the percentage of rejected assignments in parentheses.
Simulations, adopting quantitative ML assigned parentage for pools of one (i.e., assignment of parentage to individuals) without error under both the pooling-for-individual-parentage assignment and pooling-phenotype scenarios (Tables 1 and 2; Figs. 3 and 4). For pools of two, correct assignment rates were very high, 0.997 and 1.000, respectively, and for pools of three, the correct assignment rate was reduced to 0.917 under pooling-for-individual-parentage assignment but remained high under the pooling-by-phenotype scenario (0.982). For pools of three, the correct assignment rate was notably poor for methods relying on discrete genotype assignments—0.175 and 0.106 for discrete ML, and 0.048 and 0.157 for exclusion. However, discrete ML and exclusion are highly dependent on the accuracy of discrete genotype calls and our results, although indicative that quantitative ML is superior to these methods, are only relevant to the approach to genotype calling adopted in simulations. Furthermore, using quantitative ML, it was necessary to reject a substantial percentage of assignments (31%; Fig. 3e) to achieve a 99% correct assignment rate (critical Δ LOD = 4.96) for pools of three using a pooling-for-individual-parentage-assignment approach. This highlights the inherent limitations of using low-density SNP panels to assign parentage to large pools.
Discussion
Quantitative ML can be used to assign parentage to pooled samples using low-density SNP data and simulations showed it to be more accurate in assigning parentage to pools than other approaches—discrete ML (Kalinowski et al. 2010; Marshall et al. 1998); exclusion (Chakraborty et al. 1974), and solving the weighted least squares problem (Henshall et al. 2014; Kinghorn et al. 2010). In addition, unlike exclusion and solving the weighted least squares problem, ML parentage assignment allows a desired level of confidence in assignment to be specified, by defining a critical Δ LOD value below which assignments are rejected (Figs. 3c, e and 4e).
Two circumstances where implementation of parentage assignment to pools is applicable were identified— pooling-for-individual-parentage-assignment and pooling-by-phenotype. In the case of pooling-for-individual-parentage assignment applied to selective breeding programmes, individuals in each pool are tagged or traceable and are from a known and mutually exclusive set of possible parents. This approach to pooling makes the construction of the additive relationship matrix for identifiable (e.g., tagged) individuals possible and is particularly suited to reconstructing full-sib pedigree from multiple-sire joinings in livestock (Henderson 1988), poly cross families in trees (Burdon and Shelbourne 1971) and aggregated full-sib families in aquaculture (Hamilton et al. 2009). However, it is also suitable for the pooling of samples from different rounds of selection or mating, different selective breeding populations (e.g., different hatcheries or seed orchards) or selective breeding programmes involving related species with common SNPs (Hamilton et al. 2019a; Hamilton et al. 2019b), where parents in each are mutually exclusive. Furthermore, it is conceivable that management of selective breeding populations could be altered to allow the adoption of pooling-for-individual-parentage-assignment. For example, in aquaculture selective breeding programmes where there are, in some circumstances, limited facilities to maintain families in separate tanks or hapas prior to tagging, families could be replicated across multiple tanks or hapas each containing multiple families in an orthogonal design so as to allow subsequent pooling-for-individual-parentage-assignment and the partitioning of common rearing environment and genetic effects in genetic analyses. In molecular ecology, pooling-for-individual-parentage assignment is applicable in circumstances where sets of mutually exclusive groups of parents can be identified (e.g., samples from different geographical areas, between which gene flow in a single generation is not possible).
In the second circumstance, pooling-by-phenotype, individuals are allocated to bins (i.e., classes) according to their phenotype, from which pools are then drawn. The primary limitation of pooling-by-phenotype is that genotypes are not assigned to tagged individuals making the re-identification of candidate parents difficult. However, pooling-by-phenotype does not preclude the adoption of a modified ‘walk-back selection’ approach (Sonesson 2005), in which individual with desirable phenotypes are tagged and individually assigned parentage. Furthermore, it can be adopted as a means of estimating additive co/variances in a cost-effective manner, to examine genotype-by-environment interaction and/or increase the accuracy of estimated breeding values for related individuals (Burdon 1977; Henderson and Quaas 1976). Estimating genetic additive co/variances using a pooling-by-phenotype approach, is most simply achieved by generating dummy identifiers for individuals in pools, allowing an additive relationship matrix to be computed using established methods (Henderson 1975; Henderson and Quaas 1976). A further drawback of pooling-by-phenotype in circumstances where multiple traits are measured is that multiple-trait bins must be generated, each containing only those individuals with phenotypes within a specified phenotypic range for each trait (Bell et al. 2017). This potentially requires the range of phenotypic values in any one pool (i.e., bin width) for any one trait to be large, resulting in less precise phenotypes.
Ability to assign parentage using ML, whether applied to pools or individuals (Jones et al. 2010), is a function of the number of possible parents (Supplementary Materials 2 and 3) and the extent to which their contributions to progeny are known; the extent to which parents are genotyped (Supplementary Materials 4), the degree of relatedness among individuals (Supplementary Materials 5); the number (Supplementary Materials 3 and 4), information content, linkage disequilibrium (LD) and neutrality of SNPs (Holman et al. 2017); and the extent of SNP expression and genotyping accuracy/error (Anderson and Garza 2006; Liu et al. 2016; Weinman et al. 2015) (Supplementary Materials 5 and 6). Furthermore, in the application of quantitative ML to pools, variation in DNA contributions among individuals in pools (e.g. due to unequal tissue contributions and/or differences in DNA amplification) reduces the accuracy of genotyping and parentage assignment (Barratt et al. 2002; Bell et al. 2017; Kinghorn et al. 2010). Variation in DNA contributions can be minimised by pooling samples after DNA extraction, rather than pooling tissue samples prior to DNA extraction. However, pooling DNA rather than tissue substantially increases DNA extraction costs and, if the benefits of parentage assignment to pools is to be fully realised, pools must be constructed in a logistically sensible and cost-effective fashion. Furthermore, before application to pools in a new population or circumstance, the quantitative ML should be validated by applying the method to pools of individuals with known pedigree using the SNPs, species, assay and platform to be applied.
Although the cost of pedigree assignment per individual can be reduced by increasing pool size it must be recognised that, in the application of quantitative ML, this increases both the percentage of individuals with Δ LOD values below the critical Δ LOD (and thus the number of rejected assignments) and the computational burden. Quantitative ML pools is computationally intensive for large pools, as the number of possible combinations of parents increases exponentially with pool size. It has been shown herein that the quantitative ML method can be practically implemented for pools of three—all simulations were conducted on a personal computer. However, access to high performance computing facilities is likely to be necessary if the method is to be applied to larger pool sizes.
Genotype data from pools present like polyploidy data. In both cases, calling of discrete genotypes is prone to error due to the large number of possible genotype classes at each SNP (Clark et al. 2019; Rahman et al. 2015), making accurate parentage assignment difficult using discrete ML and exclusion methods (Flanagan and Jones 2019; Spielmann et al. 2015; Wang and Scribner 2014). Accordingly, generalisation of the quantitative ML approach to polyploids, with differing modes of inheritance (Clark et al. 2019), would likely improve the accuracy of ML parentage assignment and merits further investigation—the use of quantitative genotypes has been adopted to increase the power of genomic prediction (de Bem Oliveira et al. 2019) and genome-wide association studies (GWAS; Grandke et al. 2016) in polyploid populations. Modification of quantitative ML and pooling to circumstances where parental genotypes are largely unknown—such as kinship analysis (Hamilton et al. 2019a; Hamilton et al. 2019b)—also warrants further development.
In conclusion, quantitative ML can be used to assign parentage to individuals and pools, to a desired level of confidence, using low-density SNP data. Moreover, parentage is assigned with greater accuracy using quantitative ML than by discrete ML (Kalinowski et al. 2010; Marshall et al. 1998); exclusion (Chakraborty et al. 1974), or solving the weighted least squares problem (Henshall et al. 2014; Kinghorn et al. 2010). The method is applicable to pools constructed using pooling-for-individual-parentage-assignment or pooling-by-phenotype approaches and has the potential to substantially reduce the cost of parentage assignment, even if applied to pools comprised of few individuals. However, before application in applied breeding programmes quantitative ML should be validated using pools of known pedigree and tissue/DNA contributions, using the SNPs, species, assay and platform to be applied; and the inherent limitations of using low-density SNP panels to assign parentage to large pools must be recognised. Generalisation of the quantitative ML approach to polyploids and kinship analysis applications warrants further investigation.
Data availability
An R package (R Core Team 2020) entitled ‘SNPpools’, available at https://github.com/mghamilton/SNPpools, was developed to implement and validate with simulations the quantitative ML method.
References
Anderson EC, Garza JC (2006) The power of single-nucleotide polymorphisms for large-scale parentage inference. Genetics 172(4):2567–2582
Barratt BJ, Payne F, Rance HE, Nutland S, Todd JA, Clayton DG (2002) Identification of the sources of error in allele frequency estimations from pooled DNA indicates an optimal experimental design. Ann Hum Genet 66(5-6):393–405
Bell AM, Henshall JM, Porto-Neto LR, Dominik S, McCulloch R, Kijas J et al. (2017) Estimating the genetic merit of sires by using pooled DNA from progeny of undetermined pedigree. Genet Sel Evol 49:ARTN 28
Burdon R, Shelbourne C (1971) Breeding populations for recurrent selection: Conflicts and possible solutions. N Z J Sci 1:174–193
Burdon RD (1977) Genetic correlation as a concept for studying genotype- environment interaction in forest tree breeding. Silvae Genet 26(5/6):168–175
Chakraborty R, Shaw M, Schull WJ (1974) Exclusion of paternity: the current state of the art. Am J Hum Genet 26(4):477
Clark LV, Lipka AE, Sacks EJ (2019) polyRAD: genotype calling with uncertainty from sequencing data in polyploids and diploids. G3 9(3):663–673
Dai P, Kong J, Liu J, Lu X, Sui J, Meng X et al. (2020) Evaluation of the utility of genomic information to improve genetic evaluation of feed efficiency traits of the Pacific white shrimp Litopenaeus vannamei. Aquaculture 527:735421
de Bem Oliveira I, Resende Jr MFR, Ferrao LFV, Amadeu RR, Endelman JB, Kirst M et al. (2019) Genomic prediction of autotetraploids; influence of relationship matrices, allele dosage, and continuous genotyping calls in phenotype prediction. G3 9(4):1189–1198
Flanagan SP, Jones AG (2019) The future of parentage analysis: From microsatellites to SNPs and beyond. Mol Ecol 28(3):544–567
Grandke F, Singh P, Heuven HC, de Haan JR, Metzler D (2016) Advantages of continuous genotype values over genotype classes for GWAS in higher polyploids: a comparative study in hexaploid chrysanthemum. BMC Genom 17:672
Grattapaglia D, Diener PSD, dos Santos GA (2014) Performance of microsatellites for parentage assignment following mass controlled pollination in a clonal seed orchard of loblolly pine (Pinus taeda L.). Tree Genet Genomes 10(6):1631–1643
Hamilton MG, Kube PD, Elliott NG, McPherson LJ, Krsinich A (2009) Development of a breeding strategy for hybrid abalone. Proc Assoc Adv Anim Breed Genet 18:350–353
Hamilton MG, Mekkawy W, Benzie JAH (2019a) Sibship assignment to the founders of a Bangladeshi Catla catla breeding population. Genet Sel Evol 51(1):17
Hamilton MG, Mekkawy W, Kilian A, Benzie JAH (2019b) Single nucleotide polymorphisms (SNPs) reveal sibship among founders of a Bangladeshi rohu (Labeo rohita) breeding population. Front Genet. 10:597
Hansen OK, Kjaer ED (2006) Paternity analysis with microsatellites in a Danish Abies nordmanniana clonal seed orchard reveals dysfunctions. Can J Res-Rev Can Rech 36(4):1054–1058
Harrison HB, Saenz-Agudelo P, Planes S, Jones GP, Berumen ML (2013) On minimizing assignment errors and the trade-off between false positives and negatives in parentage analysis. Mol Ecol 22(23):5738–5742
Hauser L, Baird M, Hilborn R, Seeb LW, Seeb JE(2011) An empirical comparison of SNPs and microsatellites for parentage and kinship assignment in a wild sockeye salmon (Oncorhynchus nerka) population Mol Ecol Resour 11(Suppl 1):150–161
Henderson CR (1975) Best linear unbiased estimation and prediction under a selection model. Biometrics 31(2):423–447
Henderson CR (1988) Use of an average numerator relationship matrix for multiple-sire joining. J Anim Sci 66(7):1614–1621
Henderson CR, Quaas RL (1976) Multiple trait evaluation using relatives’ records. J Anim Sci 43(6):1188–1197
Henshall JM, Dierens L, Sellars MJ (2014) Quantitative analysis of low-density SNP data for parentage assignment and estimation of family contributions to pooled samples. Genet Sel Evol 46:ARTN 51
Henshall JM, Hawken RJ, Dominik S, Barendse W (2012) Estimating the effect of SNP genotype on quantitative traits from pooled DNA samples. Genet Sel Evol 44:ARTN 12
Holman LE, Onoufriou A, Hillestad B, Johnston IA (2017) A workflow used to design low density SNP panels for parentage assignment and traceability in aquaculture species and its validation in Atlantic salmon. Aquaculture 476:59–64
Jones AG, Small CM, Paczolt KA, Ratterman NL (2010) A practical guide to methods of parentage analysis. Mol Ecol Resour 10(1):6–30
Kalinowski ST, Taper ML, Marshall TC (2010) Corrigendum: revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment (vol 16, pg 1099 2007). Mol Ecol 19(7):1512–1512
Kinghorn BP, Bastiaansen JWM, Ciobanu DC, van der Steen HAM (2010) Quantitative genotyping to estimate genetic contributions to pooled samples and genetic merit of the contributing entities. Acta Agr Scand a 60(1):3–12
Liu S, Palti Y, Gao G, Rexroad CE (2016) Development and validation of a SNP panel for parentage assignment in rainbow trout. Aquaculture 452:178–182
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7(5):639–655
Meagher TR, Thompson E (1986) The relationship between single parent and parent pair genetic likelihoods in genealogy reconstruction. Theor Popul Biol 29(1):87–106
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rahman A, Hellicar A, Smith D, Henshall JM (2015) Allele frequency calibration for SNP based genotyping of DNA pools: A regression based local-global error fusion method. Comput Biol Med 61:48–55
Semagn K, Babu R, Hearne S, Olsen M (2014) Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): overview of the technology and its application in crop improvement. Mol Breed 33(1):1–14
Sonesson AK (2005) A combination of walk-back and optimum contribution selection in fish: a simulation study. Genet Sel Evol 37(6):587–599
Spielmann A, Harris SA, Boshier DH, Vinson CC (2015) orchard: paternity program for autotetraploid species. Mol Ecol Resour 15(4):915–920
Vandeputte M, Haffray P (2014) Parentage assignment with genomic markers: a major advance for understanding and exploiting genetic variation ofquantitative traits in farmed aquatic animals. Front Genet 5:ARTN 432
Wang J, Scribner KT (2014) Parentage and sibship inference from markers in polyploids. Mol Ecol Resour 14(3):541–553
Weinman LR, Solomon JW, Rubenstein DR (2015) A comparison of single nucleotide polymorphism and microsatellite markers for analysis of parentage and kinship in a cooperatively breeding bird. Mol Ecol Resour 15(3):502–511
Acknowledgements
This work was supported by the CSIRO Agriculture and Food project ‘Genomics platforms to assist applied aquaculture breeding’ (AgSIP53). John Henshall shared his R scripts relating to quantitative analysis of low-density SNP data for parentage assignment and estimation of family contributions to pooled samples—code from these scripts was not used in the SNPpools package or for simulations but was used to further the author’s understanding of the methods presented in Henshall et al. (2014). Harry King, Peter Kube, James Kijas, Klara Verbyla, Sonja Dominik shared their insights into the potential application of SNP pooling in selective breeding programmes. James Kijas assisted with comments on draft versions of the manuscript. The CGIAR Research Program on Fish Agrifood Systems (FISH), led by WorldFish and supported by contributors to the CGIAR Trust Fund, financially supported completion of the manuscript subsequent to the author’s departure from CSIRO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Jinliang Wang
Supplementary information
Rights and permissions
About this article
Cite this article
Hamilton, M.G. Maximum likelihood parentage assignment using quantitative genotypes. Heredity 126, 884–895 (2021). https://doi.org/10.1038/s41437-021-00421-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-021-00421-0