Abstract
Identifying genetic cues of functional relevance is key to understanding the drivers of evolution and increasingly important for the conservation of biodiversity. This study introduces nuclear ribosomal DNA (nrDNA) to mitochondrial DNA (mtDNA) copy number ratios as a metric with which to screen for this functional genetic variation prior to more extensive omics analyses. To illustrate the metric, quantitative PCR was used to estimate nrDNA (18S) to mtDNA (16S) copy number ratios in muscle tissue from samples of two zooplankton species: Salpa thompsoni caught near Elephant Island (Southern Ocean) and S. fusiformis sampled off Gough Island (South Atlantic). Average 18S:16S ratios in these samples were 9:1 and 3:1, respectively. nrDNA 45S arrays and mitochondrial genomes were then deep sequenced to uncover the sources of intra-individual genetic variation underlying these 18S:16S copy number differences. The deep sequencing profiles obtained were consistent with genetic changes resulting from adaptive processes, including an expansion of nrDNA and damage to mtDNA in S. thompsoni, potentially in response to the polar environment. Beyond this example from zooplankton, nrDNA:mtDNA copy number ratios offer a promising metric to help identify genetic variation of functional relevance in animals more broadly.
Similar content being viewed by others
Introduction
The use of genetic data in evolutionary and conservation biology has expanded beyond the inference of population structure, species identity, and phylogenetic relationships to incorporate inferences about function and adaptation to the environment, e.g., with the advent of ecological genomics (Ungerer et al. 2008). However, identifying genetic cues of functional relevance is not trivial because extensive omics analyses are often required—reviewed by Porcelli et al. (2015). Screening for variation at candidate loci putatively associated with function can yield valuable insights to complement subsequent and more extensive omics investigations. Nuclear ribosomal DNA (nrDNA) and mitochondrial DNA (mtDNA) are of considerable functional significance but have been under-exploited as markers within this context (Weider et al. 2005; Ballard and Pichaud 2014). In this study, I propose that nrDNA and mtDNA could be assessed for functionally relevant variation through the comparison of copy number ratios.
nrDNA and mtDNA vary in copy number between cells, tissues, individuals and species. Classic examples of the former come from studies on toad oocytes that are known to accumulate both nrDNA and mtDNA (Dawid 1966; Gall 1968). By contrast, nrDNA content is reduced during the development of copepods’ somatic tissue (Zagoskin et al. 2010). Levels of human mtDNA show more variation among tissues within individuals than between individuals (Wachsmuth et al. 2016). Nevertheless, tissue-specific inter-individual variation can be considerable. Gibbons et al. (2014) found mtDNA content to vary c.4-fold, and nrDNA more than an order of magnitude, among the human individuals included in the 1000 Genomes Project Consortium et al. (2012) dataset. Inter-individual copy number comparisons are dominated by studies involving human pathological states, such as neurodegeneration and cancer, with which differences in nrDNA and mtDNA abundance have been associated (Hallgren et al. 2014; Pyle et al. 2016; Reznik et al. 2016; Wang and Lemos 2017). Between-species comparisons of copy number have revealed substantial variation in nrDNA content and an association with genome size (Prokopowich et al. 2003; Vierna et al. 2013). There have yet to be any such multi-species comparisons of mtDNA content.
The most obvious functions of nrDNA and mtDNA relate to the structural and catalytic products these DNA regions encode, which are essential for protein synthesis and energy production (Weider et al. 2005; Ballard and Pichaud 2014 and references therein). Associations between the physiological demand for these products and copy numbers are clear in certain tissue-specific instances (e.g., toad oocytes: Dawid 1966; Gall 1968) and they have been proposed to be involved in pathological conditions (e.g., human cancers: Reznik et al. 2016; Wang and Lemos 2017). Some intriguing examples of associations between copy number and physiological demand have also emerged at the organismal level. Elser et al. (2000) hypothesized that nrDNA abundance is positively associated with growth rates, citing several examples from plants and animals (see also Weider et al. 2005). More recently, Cheng et al. (2013) and Li et al. (2016) have explored the contribution of mtDNA levels to physiological adaptation in human populations and reported associations between copy numbers and environmental parameters.
Associations between copy numbers and the physiological demand for proteins and energy are complicated by other factors including sample-specific differences in the regulation of DNA transcription and RNA translation, and the related issue of genetic and epigenetic variation affecting nrDNA and mtDNA functions. These functions have been suggested to extend beyond encoding products for protein synthesis and energy production to other roles, which in the case of yeast nrDNA, involve modulating genome integrity and adaptability (Kobayashi 2011). The nrDNA within other organisms may have similar additional functions given reported instances of redundancy in nrDNA copy numbers which exceed cellular requirements for ribosomal RNA (Weider et al. 2005). Indeed, changes in nrDNA levels that alter genome stability may have a functional role in human cancers (Wang and Lemos 2017). Copy number redundancy has also been suggested to play a role in the maintenance of mitochondrial genome integrity in mammals by provisioning for the selective degradation of damaged molecules (Alexeyev et al. 2013). Furthermore, mtDNA may have “other functions” by encoding for products involved in processes other than energy production, such as the mitochondrial-derived peptide humanin (Capt et al. 2016).
The extent to which these proposed functions of nrDNA and mtDNA ultimately drive copy number variation remains understudied. So too is the impact of other potential drivers on copy numbers, that include mechanistic cellular processes (e.g., DNA recombination and replication) and underlying evolutionary processes (e.g. drift, gene flow). Accumulating evidence from recent work on humans suggests that it is worth exploring the potential of nrDNA and mtDNA copy numbers as markers of functionally relevant genetic variation in animals more broadly. Obtaining nrDNA and mtDNA copy number data is straightforward because it can be readily estimated through quantitative PCR (qPCR) by taking advantage of archived sequences as a resource for primer design. Due to the typically high abundance of nrDNA and mtDNA in nucleic acid extractions, copy numbers can also be efficiently estimated from low coverage 1 + nth generation sequencing surveys. mtDNA levels estimated in this way are less prone to bias arising from the inadvertent qPCR amplification of nuclear-mitochondrial DNA (NUMTS). Estimating relative amounts of nrDNA and mtDNA instead of their absolute quantities offers practical advantages. Deriving copy number ratios removes the need for internal references (e.g., single copy genes), which require additional background information for qPCR primer design and deep rather than low coverage sequencing. Furthermore, the use of ratios greatly simplifies quantitative comparisons within and between studies by removing the need for data on absolute template concentrations associated with qPCR/DNA sequencing.
To explore the use of nrDNA:mtDNA copy number ratios within an evolutionary and conservation genetics setting, a comparison was made between a pair of closely related zooplankton species that might be expected to show physiological adaptations to different environments. The salp (pelagic tunicate) Salpa thompsoni was selected as the first species because it occurs within the polar waters of the Southern Ocean (Foxton 1961) and has interesting nrDNA-mtDNA properties: abundant nrDNA (Jue et al. 2016), and mtDNA duplications and heteroplasmy (Goodall-Copestake 2017). The other species selected for comparison was S. fusiformis, which is non-polar and has a near-cosmopolitan distribution (Van Soest 1975). Little is known of S. fusiformis genetics other than its nrDNA 18S sequence that confirms its close affinity to S. thompsoni (Govindarajan et al. 2011). Species distributions (Van Soest 1975) and phylogenetic relationships (Govindarajan et al. 2011) are consistent with S. thompsoni as a polar lineage of the largely tropical-temperate genus Salpa. Adaptive genetic changes in response to the polar environment might thus be expected in S. thompsoni relative to S. fusiformis, some of which may have driven differences in the nrDNA:mtDNA copy number ratio between these species. To this end, qPCR was used to determine if the ratio of 18S nrDNA to 16S mtDNA differed between samples of S. thompsoni and S. fusiformis. Traditional cloning-sequencing and deep sequencing were then used to generate 45S rRNA array (from 18S to 28S) and mitochondrial genome sequences from both salp species, high-resolution copy number data, and intra-individual single nucleotide polymorphism (SNP) and structural variation data to place the qPCR results into a broader genomic background.
Methods
Ten S. thompsoni samples (five oozooids and five blastozooids) were obtained from the archives of British Antarctic Survey cruise JR26, sampling event 184 near Elephant Island (Southern Ocean). Ten S. fusiformis (three oozooids, seven blastozooids) were sampled near Gough Island (South Atlantic) during cruise JR287 event 46. Morphological determinations followed descriptions in Foxton (1961). Body muscles were dissected out for molecular genetic analysis.
Procedures for DNA extraction, long-range PCR, amplicon visualization, cloning, Sanger sequencing-editing-assembly and annotation followed those previously described in Goodall-Copestake (2017). Novel PCR primers were designed from salp 18S (Govindarajan et al. 2011) and mtDNA (Goodall-Copestake WP, unpublished data) sequences using the Oligo Analysis Tool by Eurofins Genomics (Ebersberg, Germany). Other procedures and methodological variations were as follows.
qPCR was carried out in triplicate 10 µL reactions using a KiCqStart SYBR Green qPCR ReadyMix (Sigma-Aldrich, Staffordshire, UK), 5ng of template, and the novel 18S primers (F 5′- CAAAGATTAAGCCATGCAAGTGTAAG -3′, R 5′-TCACGCATGTATTAGCTCTAGAATTG -3′) and 16S primers (F 5′-CCTTATGCAATTGGTGTTTTACGAC-3′, R 5′-GAGATAAAATCCGGTCTAATTCTCACC-3′) on an Eco 48 machine (PCRmax Limited, Staffordshire, UK). The bundled PCRmax Eco Study software was used to determine average Cq fluorescence values from which nrDNA 18S to mtDNA 16S abundance ratios were generated.
Near complete fragments of 45S nrDNA were amplified using the primer 18S F paired with ‘11_RC’ from Machida and Knowlton (2012). Mitochondrial genomes were amplified as two overlapping fragments using the 16S qPCR primers paired with the novel trnR primers (F 5′-GAACAGGTGATTTCGGGTTACC-3′, R 5′-GGTAACCCGAAATCACCTGTTC-3′). Amplicons were single pass Sanger sequenced by Eurofins Genomics following a primer walking strategy. However, repetitive nad3-nad4-trnM-cox1 mtDNA in S. thompsoni (see Goodall-Copestake 2017) could not be sequenced using this method. Instead, the longest cloned amplicon obtained with the novel flanking primers nd2 F (5′-GGAGACTGGGTGTTGGTTACT-3′) and cx3 R (5′- TCGTGTGTGATCCCTAACTAGTC -3′) was sequenced as nested deletions following TOPO TA Cloning Kit (Life Technologies, Paisley, UK) instructions; Sbf I then Pme I were selected to make linear plasmids with blunt 5′ ends and 3′ overhangs and a 37 °C exonuclease III digestion sampled ten times every 90 s was used to generate successive 5’ end deletions (enzymes from New England BioLabs, Ipswich, USA).
TruSeq nano DNA library preparation and Illumina HiSeq 100 bp paired-end sequencing were outsourced to the University of Bristol Genomics Service (Bristol, UK). Geneious v10.2 (Biomatters, Auckland, New Zealand) was used to trim overrepresented and low-quality sequences identified with FastQC v0.11.5 (Andrews 2010). Majority (most common base call) consensus sequences for 45S nrDNA and mtDNA genomes were generated by mapping the trimmed HiSeq reads to the Sanger sequence derived templates using Geneious with a medium sensitivity and paired reads only option. MITOS (Bernt et al. 2013) was used to guide mtDNA sequence annotation.
Repeated HiSeq read mapping to the majority consensus sequences was used to estimate relative copy numbers, the tbx1 gene was applied as a single copy reference following Jue et al. (2016). A relative measure of SNP prevalence (proportion of mapped reads) was inferred by identifying sites where >10% of reads contained a base call differing from the consensus. Geneious structural variant mapper was used to identify putative deletions, small insertions (limited by read length), and inversions. Relative measures of these variants were estimated by identifying sites where >0.1% of mapped reads contained a variant junction. The lower cut-off used for structural variants compared to SNPs was based on the lower frequency of such errors in HiSeq datasets (Schirmer et al. 2016).
Results
Initial qPCR analysis revealed that the ratio of 18S nrDNA to 16S mtDNA in muscle tissue was consistently higher in the S. thompsoni samples than in S. fusiformis samples. For every copy of 16S, S. thompsoni contained on average 9 ± 2.0 copies of 18S (minimum 7, maximum 12, n = 10). By contrast, in S. fusiformis, there were on average 3 ± 0.5 copies of 18S for every copy of 16S (minimum 2, maximum 4, n = 10). Although sample sizes were small, these inter-species differences in 18S:16S copy number ratios appear to be robust to the inclusion of both oozooid and blastozooid life history stages.
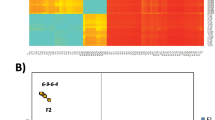
A total of 258,104,562 S. thompsoni and 279,916,278 S. fusiformis trimmed HiSeq reads were used for mapping to the long-range PCR derived Sanger sequenced templates. The resulting majority rule consensus sequences represent the first annotated 45S nrDNA and mtDNA genome accessions for any salp species and as such, will provide a valuable reference for future comparative analysis (Fig. 1a, c). Average raw pairwise differences between the two species were 0.5% for 18S-5.8S-28S, 6.9% for the ITS regions and 18.8% for mitochondrial coding DNA; the cox1 DNA barcoding difference was 15.2%. Notable structural differences involved the position of four tRNA genes, and in S. thompsoni, the presence of a near perfect sequence repeat associated with this tRNA rearrangement, and a region of tandemly repeated nad3-nad4-trnM-cox1 coding DNA (Fig. 1c).
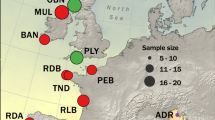
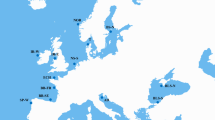
Variation (a, c) and relative abundance (b) along DNA sequences comprising salp 45S nuclear ribosomal arrays and mitochondrial genomes. Relative read depth is the proportion of total HiSeq reads mapped per nucleotide site multiplied by 10,000. Read variation includes sites where >10% of mapped reads contained a single nucleotide polymorphism (S); and sites where >0.1% mapped reads contained deletions (D), insertions (I) or inversions (V); triangles mark the mid-point of S, D, I, V feature junctions. Sampling locations and linear DNA sequence maps are provided for reference; vertical bars indicate tRNA sites, asterisks indicate duplicated DNA sequence features
The average percentage of mapped reads per site was higher for S. thompsoni nrDNA than S. fusiformis nrDNA, and conversely, proportionately fewer reads mapped to S. thompsoni mtDNA than to S. fusiformis mtDNA (Fig. 1b). Copy number ratios derived from the average read depths were 9:1 for S. thompsoni and 2:1 for S. fusiformis. Considering only those sites amplified by the qPCR primers, there were 9 and 3 copies of 18S for every copy of 16S in S. thompsoni and S. fusiformis, respectively. These results were thus in accordance with those inferred using qPCR. When normalized to the read depth of the tbx1 gene, the estimated haploid copy numbers were 484 for nrDNA and 54 for mtDNA in S. thompsoni, and 125 for nrDNA and 62 for mtDNA in S. fusiformis.
SNP and structural variant prevalence differed markedly between nrDNA and mtDNA, as well as between the salp samples (Fig. 1a, c). After correcting for sequence length, the S. thompsoni 45S HiSeq reads contained approximately one-third of the SNPs but twice as many structural variants as found in the S. fusiformis 45S reads. The S. thompsoni mtDNA HiSeq reads comprised greater numbers of both SNPs (3× more) and structural variants (33× more) than the S. fusiformis HiSeq reads.
Discussion
This study introduces nrDNA to mtDNA copy number ratios as a new metric with which to screen for functional genetic variation in evolutionary and conservation genetics studies. To investigate the potential of this metric, qPCR was used to estimate nrDNA:mtDNA copy number ratios in samples of two zooplankters, S. thompsoni and S. fusiformis, followed by deep sequencing to profile the underlying sources of genetic variation. Species-specific copy number ratios were found, as were different profiles of intra-individual nrDNA and mtDNA variation (Fig. 1). Before discussing the potential functional significance of these nrDNA and mtDNA differences, it is important to consider the life-history characteristics of S. thompsoni and S. fusiformis. In particular, the capacity of salps for opportunistic population blooms and extensive asexual reproduction, which suggests that they may be prone to reductions in effective population size (Goodall-Copestake 2017). Therefore, it is conceivable that genetic drift might have influenced, or even caused, the differences in species-specific nrDNA:mtDNA copy number ratios found in this study.
Differences in nrDNA content between S. thompsoni and S. fusiformis account for most of the difference in nrDNA:mtDNA copy number ratios between these species (Fig. 1b). The larger number of nrDNA copies found in S. thompsoni, associated with fewer SNPs but more structural variants (Fig. 1a), suggests an expansion of a subset of similar nrDNAs in S. thompsoni relative to S. fusiformis. Jue et al. (2016) also inferred an expansion of nrDNA in their analysis of the S. thompsoni genome. By incorporating gene expression results from a companion study (Batta-Lona et al. 2017), Jue et al. (2016) suggested that nrDNA content had increased in S. thompsoni to compensate for slow metabolism due to the low temperatures of its polar environment. With the added benefit of comparative data from non-polar S. fusiformis, the results obtained herein support this possibility. Salps are ectotherms and an increase in the number of actively transcribed nrDNA loci, and/or a locus advantageous in cold environments, may help S. thompsoni to achieve growth rates at temperatures ranging from −1 to 2 °C that “compare favorably” to growth rate estimates of S. fusiformis at 15 °C (Loeb and Santora 2012). At a broader functional level, the difference in nrDNA content between S. thompsoni and S. fusiformis may also reflect other environmentally induced changes, such as those associated with the regulation of genome-level integrity and adaptability (Kobayashi 2011).
mtDNA content differed less considerably than nrDNA content between S. thompsoni and S. fusiformis (Fig. 1b), and accordingly, mtDNA had less of an impact on nrDNA:mtDNA copy number ratios. Unlike nrDNA, levels of mtDNA do not appear to have increased in S. thompsoni as part of a metabolic response to low temperatures. On the contrary, mtDNA copy numbers were marginally lower in S. thompsoni than in S. fusiformis. Salpa thompsoni may thus rely on different mechanisms to ensure that energy production satisfies metabolic requirements in its polar environment. There are a number of possible mechanisms by which this may occur, prime among which are adjustments to mitochondrial membrane composition that enhance respiratory activity (Guderley 2004). However, these membrane changes are at the cost of an increase in reactive oxygen species (ROS) that are known to damage DNA (Guderley 2004; Ballard and Pichaud 2014). While speculative, the mtDNA deep sequencing results suggest that S. thompsoni could have undergone such a change in mitochondrial membrane composition as a response to the cold. The rationale for this being that the mtDNA of polar S. thompsoni contains abundant signatures of damage in the form of intra-individual SNPs and structural rearrangements, while the mtDNA of non-polar S. fusiformis does not (Fig. 1c). It is worth noting that, in addition to ROS-mediated damage, other factors might also have contributed to the signature of mtDNA damage in S. thompsoni, including genome destabilization induced by mtDNA duplications, and mutations impacting the efficacy of nuclear encoded genes involved in mtDNA replication and repair. Two other issues warrant mention with regard to copy numbers. The first is the possibility that mtDNA copy numbers in S. thompsoni might be impacted through the selective degradation of damaged mtDNA molecules (Alexeyev et al. 2013). The other is the possibility that mitochondrial ROS could influence nrDNA levels through retrograde signaling driven processes (mitochondrion to nucleus signaling; see Ballard and Pichaud 2014).
To conclude, this study reveals how nrDNA:mtDNA copy number ratios obtained from the same tissue in different individuals and species can be used as a metric of potentially functional genetic variation. In the case of S. thompsoni and S. fusiformis, differences in nrDNA:mtDNA copy number ratios and the underlying sources of sequence variation were consistent with adaptation and damage at the DNA level in S. thompsoni, potentially in response to the polar environment. This raises the intriguing possibility that S. thompsoni may be less adapted to polar conditions than its distribution might suggest. More extensive omics analyses of salps are required to ascertain if this is the case. Data from a broad range of other animal species, sampled under different conditions, are also needed to further explore the circumstances under which levels of nrDNA and mtDNA vary. These data will provide much-needed insights about the drivers of copy number variation and the utility of nrDNA:mtDNA copy number ratios as a comparative metric.
Data archiving
Sequence data have been submitted to the DNA DataBank of Japan: accession numbers LC333178-LC333181.
References
1000 Genomes Project Consortium. et al. (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65
Alexeyev M, Shokolenko I, Wilson G, LeDoux S (2013) The maintenance of mitochondrial DNA integrity—critical analysis and update. Cold Spring Harb Perspect Biol 5:a012641
Andrews S (2010). FastQC: a quality control tool for high throughput sequence data. Available at: https://www.bioinformatics.babraham.ac.uk/projects/fastqc.
Ballard JWO, Pichaud N (2014) Mitochondrial DNA: more than an evolutionary bystander. Funct Ecol 28:218–231
Batta-Lona PG, Maas AE, O’Neill RJ, Wiebe PH, Bucklin A (2017) Transcriptomic profiles of spring and summer populations of the Southern Ocean salp, Salpa thompsoni, in the Western Antarctic Peninsula region. Polar Biol 40:1261–1276
Bernt M, Donath A, Jühling F, Externbrink F, Florentz C, Fritzsch G, Pütz J, Middendorf M, Stadler PF (2013) MITOS: improved de novo Metazoan Mitochondrial genome annotation. Mol Phylogenet Evol 69:313–319
Capt C, Passamonti M, Breton S (2016) The human mitochondrial genome may code for more than 13 proteins. Mitochondrial DNA Part A 27:3098–3101
Cheng YT, Liu J, Yang LQ, Sun C, Kong QP (2013) Mitochondrial DNA content contributes to climate adaptation using Chinese populations as a model. PLoS One 8:e79536
Dawid IB (1966) Evidence for the mitochondrial origin of frog egg cytoplasmic DNA. Proc Natl Acad Sci USA 56:269–276
Elser JJ, Sterner RW, Gorokhova E, Fagan WF, Markow TA, Cotner JB et al. (2000) Biological stoichiometry from genes to ecosystems. Ecol Lett 3:540–550
Foxton P (1961) Salpa fusiformis Cuvier and related species. Discov Rep 32:1–32
Gall JG (1968) Differential synthesis of the genes for ribosomal RNA during amphibian oögenesis. Proc Natl Acad Sci USA 60:553–560
Gibbons JG, Branco AT, Yu S, Lemos B (2014) Ribosomal DNA copy number is coupled with gene expression variation and mitochondrial abundance in humans. Nat Commun 5:1–12
Goodall-Copestake WP (2017) One tunic but more than one barcode: evolutionary insights from dynamic mitochondrial DNA in Salpa thompsoni (Tunicata: Salpida). Biol J Linn Soc 120:637–648
Govindarajan AF, Bucklin A, Madin LP (2011) A molecular phylogeny of the Thaliacea. J Plankton Res 33:843–853
Guderley H (2004) Metabolic responses to low temperature in fish muscle. Biol Rev 79:409–427
Hallgren J, Pietrzak M, Rempala G, Nelson PT, Hetman M (2014) Neurodegeneration-associated instability of ribosomal DNA. Biochim Biophys Acta 1842:860–868
Jue NK, Batta-Lona PG, Trusiak S, Obergfell C, Bucklin A, O’Neill MJ et al. (2016) Rapid evolutionary rates and unique genomic signatures discovered in the first reference genome for the Southern Ocean salp, Salpa thompsoni (Urochordata, Thaliacea). Genome Biol Evol 8:3171–3186
Kobayashi T (2011) Regulation of ribosomal RNA gene copy number and its role in modulating genome integrity and evolutionary adaptability in yeast. Cell Mol Life Sci 68:1395–1403
Loeb VJ, Santora JA (2012) Population dynamics of Salpa thompsoni near the Antarctic Peninsula: growth rates and interannual variations in reproductive activity (1993–2009). Prog Oceanogr 96:93–107
Li Y, Huang W, Yu Q, Cheng YT, Kong QP (2016) Lower mitochondrial DNA content relates to high-altitude adaptation in Tibetans. Mitochondrial DNA Part A 27:753–757
Machida RJ, Knowlton N (2012) PCR primers for metazoan nuclear 18S and 28S ribosomal DNA sequences. PLoS One 7:e46180
Porcelli D, Butlin RK, Gaston KJ, Joly D, Snook RR (2015) The environmental genomics of metazoan thermal adaptation. Heredity 114:502–514
Prokopowich CD, Gregory TR, Crease TJ (2003) The correlation between rDNA copy number and genome size in eukaryotes. Genome 46:48–50
Pyle A, Anugrha H, Kurzawa-Akanbi M, Yarnall A, Burn D, Hudson G (2016) Reduced mitochondrial DNA copy number is a biomarker of Parkinson’s disease. Neurobiol Aging 38:216.e7–216.e10
Reznik E, Miller M, Senbabaoglu Y, Riaz N, Lee W, Sander C (2016) Mitochondrial DNA copy number variation across human cancers. eLife 5:e10769
Schirmer M, D’Amore R, Ijaz UZ, Hall N, Quince C (2016) Illumina error profiles: resolving fine-scale variation in metagenomic sequencing data. BMC Bioinform 17:125
Ungerer MC, Johnson LC, Herman MA (2008) Ecological genomics: understanding gene and genome function in the natural environment. Heredity 100:178–183
Van Soest RWM (1975) Zoogeography and speciation in the Salpidae (Tunicata, Thaliacea). Beaufortia 23:181–215
Vierna J, Wehner S, Höner zu Siederdissen C, Martínez-Lage A, Marz M (2013) Systematic analysis and evolution of 5S ribosomal DNA in metazoans. Heredity 111:410–421
Wachsmuth M, Hübner A, Li M, Madea B, Stoneking M (2016) Age-related and heteroplasmy-related variation in human mtDNA copy number. PLoS Genet 12:e1005939
Wang M, Lemos B (2017) Ribosomal DNA copy number amplification and loss in human cancers is linked to tumor genetic context, nucleolus activity, and proliferation. PLoS Genet 13:e1006994
Weider LJ, Elser JJ, Crease TJ, Mateos M, Cotner JB, Markow TA (2005) The functional significance of ribosomal (r)DNA variation: impacts on the evolutionary ecology of organisms. Annu Rev Ecol Evol Syst 36:219–242
Zagoskin MV, Marshak TL, Mukha DV, Grishanin AK (2010) Chromatin diminution process regulates rRNA gene copy number in freshwater copepods. Acta Nat 2:52–57
Acknowledgements
I would like to thank the RRS James Clark Ross JR26 and JR287 captains, crews and scientists for sampling and fieldwork support. Also, Sílvia Pérez-Espona, Chester Sands, and two anonymous reviewers for their valuable comments on the manuscript. Molecular work was supported by core funding from the British Antarctic Survey - Natural Environment Research Council, fieldwork by the Darwin Initiative Challenge fund grant EIDCF013 and The Pew Charitable Trusts, held by the British Antarctic Survey, the Shallow Marine Surveys Group (Falkland Islands) and the Tristan da Cunha Department of Conservation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Goodall-Copestake, W.P. nrDNA:mtDNA copy number ratios as a comparative metric for evolutionary and conservation genetics. Heredity 121, 105–111 (2018). https://doi.org/10.1038/s41437-018-0088-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-018-0088-8