Abstract
The endosymbiotic bacterium Wolbachia suppresses the capacity for arbovirus transmission in the mosquito Aedes aegypti, and can spread spatially through wild mosquito populations following local introductions. Recent introductions in Cairns, Australia have demonstrated slower than expected spatial spread. Potential reasons for this include: (i) barriers to Ae. aegypti dispersal; (ii) higher incidence of long-range dispersal; and (iii) intergenerational loss of Wolbachia. We investigated these three potential factors using genome-wide single-nucleotide polymorphisms (SNPs) and an assay for the Wolbachia infection wMel in 161 Ae. aegypti collected from Cairns in 2015. We detected a small but significant barrier effect of Cairns highways on Ae. aegypti dispersal using distance-based redundancy analysis and patch-based simulation analysis. We detected a pair of putative full-siblings in ovitraps 1312 m apart, indicating long-distance female movement likely mediated by human transport. We also found a pair of full-siblings of different infection status, indicating intergenerational loss of Wolbachia in the field. These three factors are all expected to contribute to the slow spread of Wolbachia through Ae. aegypti populations, though from our results it is unclear whether Wolbachia loss and long-distance movement are sufficiently common to reduce the speed of spatial spread appreciably. Our findings inform the strategic deployment of Wolbachia-infected mosquitoes during releases, and show how parameter estimates from laboratory studies may differ from those estimated using field data. Our landscape genomics approach can be extended to other host/symbiont systems that are being considered for biocontrol.
Similar content being viewed by others
Introduction
The mosquito Aedes aegypti (Diptera, Culicinae) is the primary vector of arboviruses including dengue, Zika and chikungunya that impose an increasing burden on human health worldwide (Bogoch et al. 2016; Weaver and Lecuit 2015). Conventional approaches to combatting these viruses have involved the suppression of Ae. aegypti populations through source reduction or insecticide-based programmes, but these have had limited efficacy (Focks et al. 2000). One alternative strategy involves the release of Ae. aegypti carrying an infection of the virus-inhibiting, endosymbiotic bacterium Wolbachia into wild Ae. aegypti populations (McGraw and O’Neill 2013). Once they are widespread in host populations, Wolbachia are expected to diminish the disease transmission rate enough to prevent outbreaks (Ferguson et al. 2015). Although Ae. aegypti does not naturally carry Wolbachia, several Wolbachia strains have been successfully transferred into Ae. aegypti from other hosts (Walker et al. 2011; Xi et al. 2005).
The Wolbachia strain wMel, originating from D. melanogaster, has proven suitable for field deployments in Ae. aegypti given its viral blockage (Ferguson et al. 2015), moderate fitness costs (Walker et al. 2011), high intergenerational transmission fidelity (Hoffmann et al. 2014) and complete cytoplasmic incompatibility (Walker et al. 2011). Cytoplasmic incompatibility (CI) describes the phenomenon that offspring of uninfected females mated with Wolbachia-infected males are unviable, while offspring of Wolbachia-infected females are viable and will carry the infection regardless of male infection status (Hoffmann and Turelli 1997). When Wolbachia-infected males are common, CI greatly reduces the relative fitness of uninfected females, which ensures that future generations will be increasingly comprised of Wolbachia-infected individuals. However, Wolbachia also impose frequency-independent fitness costs on hosts, which for wMel in Ae. aegypti include a shorter lifespan (Walker et al. 2011) and reduced larval competitive ability (Hoffmann et al. 2014; Ross et al. 2016; Suh and Dobson 2013). The interaction between costs and benefits produces a critical frequency of Wolbachia infection (\(\widehat p\)) that must be exceeded for Wolbachia to invade the mosquito population (Hoffmann et al. 1990; Turelli 2010).
Releases of wMel-infected Ae. aegypti into two sites near Cairns, Australia, confirmed that Wolbachia can establish stably in quasi-isolated habitat patches (Hoffmann et al. 2011; Hoffmann et al. 2014). A subsequent study with releases centred within continuous mosquito habitat demonstrated the successful spatial spread of the invasion into surrounding habitat, so that the infection region grew ≈ 70–85% over the ensuing 18 months (Schmidt et al. 2017). However, the rate of spread of approximately 100–200 m per year is close to the lower bound of predicted spread speeds for this host/Wolbachia system (Turelli and Barton 2017). This suggests that various potential biological and environmental processes could be operating to restrict the speed or extent of the spread (Barton and Turelli 2011; Turelli and Barton 2017).
This study investigates three potential causes for the slow spread observed in Cairns: (i) barriers to mosquito dispersal; (ii) higher incidence of long-range dispersal; and (iii) occasional intergenerational loss of Wolbachia (designated as µ). We screened Ae. aegypti from Cairns for the wMel transinfection and genotyped individuals at genome-wide single-nucleotide polymorphisms (SNPs) using double digest restriction-site-associated DNA sequencing (ddRADseq: Peterson et al. (2012)). SNP data sets produced with ddRADseq have been used to elucidate genetic structure in Ae. aegypti within cities (Rašić et al. 2015a; Rašić et al. 2015b), and have power superior to microsatellites when inferring relationships between Ae. aegypti individuals and populations (Rašić et al. 2014).
Fine-scale dispersal of Ae. aegypti is mostly accomplished through short-range flight, which may make highways and other geographical features effective barriers to movement. In Cairns, a Mark-Release-Recapture (MRR) study with releases centred next to a 20 m-wide road recorded lower recapture rates at traps across the road (Russell et al. 2005). Similarly, patches separated by a 120 m-wide highway in Trinidad had different frequencies of mitochondrial haplotypes (Hemme et al. 2010). Observing Wolbachia invasion dynamics around highways also allows for indirect inference of highway barrier effects. Specifically, Wolbachia failed to invade a region across a highway in Gordonvale, Queensland after several years (Turelli and Barton 2017), and similar dynamics were recently observed in urban Cairns (Schmidt et al. 2017). In this study, we explicitly tested the hypothesis that the genetic structure of Ae. aegypti in Cairns is affected by highways acting as barriers to dispersal. This was tested alongside the hypotheses that genetic structure reflected a simple isolation-by-distance (IBD) pattern, and that it was affected by the recent patchy and asynchronous releases of Wolbachia-infected mosquitoes throughout the region (see Schmidt et al. 2017).
If host dispersal distances follow a leptokurtic distribution, Wolbachia spread can proceed up to ≈ 4 times slower than if the distribution of dispersal distances is Gaussian (Turelli and Barton 2017). Leptokurtic distributions are characterised by higher peaks and tails that approach zero more slowly, and when used to model dispersal distances describe a system with greater numbers of short-distance and long-distance dispersers (Clark et al. 1998). A higher frequency of long-range dispersal increases the likelihood that a Wolbachia-infected individual will move to an uninvaded region, where they will receive no frequency-dependent fitness benefits and thus contribute less to the invasion. Passive dispersal of Ae. aegypti by humans, such as when mosquitoes enter vehicles, can be over distances orders of magnitude longer than the typical 50–100 m flight range of Ae. aegypti (Harrington et al. 2005; Maciel-De-Freitas et al. 2007; Muir and Kay 1998), increasing the likelihood of ending up far from the site of invasion. Passive dispersal in Aedes is commonly observed in immatures, but movement of imagoes also occurs (Eritja et al. 2017). Aedes aegypti females lay batches of eggs over multiple gonotrophic cycles (Christophers 1960), and through multiple acts of “skip” oviposition within a gonotrophic cycle (Reiter 2007), and thus full-sibling immatures can be found at different sites. In this study, we use the distance of separation between sampled full-siblings to indicate female flight ranges or passive dispersal distances within one or two gonotrophic cycles.
Slow Wolbachia spread can also result from intergenerational loss of Wolbachia (i.e., µ > 0), which will lead to an increase in the critical infection frequency \(\widehat p\) (Turelli 2010). Loss of wMel has not been detected in laboratory-reared Ae. aegypti (Hoffmann et al. 2014; Ross et al. 2016; Walker et al. 2011). However, laboratory populations subjected to high, fluctuating temperatures similar to those of Cairns showed significant decreases in wMel density (Ross et al. 2017; Ulrich et al. 2016), which could lead to loss of the infection (Clancy and Hoffmann 1998; Ikeda et al. 2003). We looked for evidence of infection loss in Cairns Ae. aegypti by first identifying all matrilineages in which the infection was present, then looking for uninfected individuals within these matrilineages, which will have lost or failed to inherit the infection.
Identifying causes of slow spread in Cairns will help future Wolbachia deployments to achieve regional invasion faster and at lower cost. Note that we differentiate the above three potential causes of slow spatial spread from potential causes of slow establishment of the invasion, which can include larval density effects (Hancock et al. 2016).
Methods
Study site and sample collection
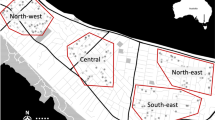
We deployed 110 ovitraps within properties of consenting householders in Cairns, Australia between 13–16 April, 2015. Traps covered a 3.3 km × 1.9 km region of central Cairns, which we partitioned into six “plots” for reference: Cairns North West (CNW), Cairns North East (CNE), Parramatta Park North (PPN), Parramatta Park South (PPS), Westcourt (WC) and Bungalow (BN) (Fig. 1). Partitions were defined by geographic location, location of highways, and the release history of wMel. Each of these had three possible assignments: the locational groupings of Cairns North [CNW, CNE], Parramatta Park [PPN, PPS] and Westcourt/Bungalow [WC, BN]; the highway groupings of southeast of Bruce Highway [PPS, BN], west of both highways [CNW, PPN, WC], or northeast of Captain Cook Highway [CNE]; and the Wolbachia release groupings of releases in 2013 [PPN, WC], releases in 2014 [CNW, BN], or no release history [CNE, PPS] (see Schmidt et al. 2017).
Sampling locations of the mosquitoes analysed with ddRADseq, set within the six sampling plots. Each sample was assigned a Wolbachia infection status, a score indicating its position relative to the two highways, and a score indicating when Wolbachia releases were carried out in the area. Plot abbreviations are: CNW (Cairns North West), CNE (Cairns North East), PPN (Parramatta Park North), PPS (Parramatta Park South), WC (Westcourt) and BN (Bungalow). (The underlying road network is derived from “Australia Oceania Continent Roads” made available by MapCruzin.com and OpenStreetMap.org under the Open Database License [https://opendatacommons.org/licenses/odbl/1.0/].)
Each ovitrap consisted of a black plastic bucket containing water and several alfalfa pellets to attract gravid female Ae. aegypti (Ritchie 2001), which oviposited on strips of red felt extending into the liquid. Traps were left for 5–7 days, then the felt strips were removed and dried. Dried strips of mosquito eggs were hatched by immersion into vessels filled with reverse osmosis water, 2–3 grains of yeast and one quarter of a tropical fish food tablet. Water, food and yeast were replaced after 3 days. Emerging virgin imagoes were transferred to freezing ethanol and stored at −20 °C until DNA extraction.
ddRADseq library preparation and SNP discovery
We extracted genomic DNA using Roche DNA Isolation Kit for Cells and Tissues (Roche, Pleasanton, CA, USA), with an additional step of RNase treatment. Seventy-four of the 110 ovitraps produced Ae. aegypti imagoes, and we selected 161 individuals for sequencing. As we expected ovitraps to contain many full-siblings from the same oviposition (Hoffmann et al. 2014), which can bias analyses of population structure (Goldberg and Waits 2010), we sequenced no more than three individuals per ovitrap.
We applied the method of Rašić et al. (2014) for ddRADseq library preparation, but selected a smaller size range (350–450 bp) of genomic fragments to accommodate more individuals per library. We generated three libraries each consisting of 60–61 individuals, which were sequenced in three Illumina HiSeq2500 lanes using 100 bp paired-end chemistry. We processed raw fastq sequences within a customised pipeline (Rašić et al. 2014), retaining reads with phred scores ≥ 13 and trimming them to 90 bp. High-quality reads were aligned to the Ae. aegypti nuclear genome assembly AaegL1 (Nene et al. 2007) using Bowtie (Langmead et al. 2009). We allowed for up to three mismatches in the alignment seed, and uniquely aligned reads were analysed using Stacks (Catchen et al. 2013), which we used to call genotypes at RAD stacks of a minimum depth of five reads.
We used the Stacks programme populations to export VCF files, which we filtered using VCFtools (Danecek et al. 2011). We kept 134 individuals with < 20% missing data, and retained loci at Hardy–Weinberg equilibrium (81% of loci) and with minor allele frequencies > 0.05 (average frequency in retained loci = 0.21). We applied thinning to ensure no SNP was within 250 kbp of another. As Aedes genome contains approximately 2.1 megabases per cM (Brown et al. 2001), 250 kbp roughly corresponds to eight SNPs per map unit, a sampling density shown to largely eradicate linkage effects in SNPs (Cho and Dupuis 2009). We retained 3784 unlinked and informative SNPs for analyses of relatedness and genetic structure.
Assessing barriers to dispersal: landscape resistance modelling
We tested the hypotheses that highways affect Ae. aegypti genetic structure in Cairns (H1), and that genetic structure was affected by recent releases of Wolbachia (H2). To test H1 and H2, we derived continuous variables we called “Highways” and “Releases” respectively, that we scored for each individual using processes outlined in Appendix A. Individuals were scored so that differences in scores between pairs of individuals would reflect the number of highways separating them and how different their release histories were.
We performed partial distance-based redundancy analyses (dbRDA: Legendre and Anderson 1999), to test H1 and H2 as explicit hypotheses, while controlling for the potentially confounding effects of IBD and latitudinal and longitudinal patterns. We sampled one individual from each matrilineage (see below) with the lowest percentage of missing data, retaining 100 individuals with 1.65–13.3% missing data. We treated Easting and Northing UTM coordinates and a binary variable indicating individual Wolbachia infection status as potentially confounding variables, which were placed inside a conditional matrix. The dependent variable was a distance matrix of Rousset’s a scores (Rousset 2000) calculated for each pair of individuals using the programme SPAGeDi (Hardy and Vekemans 2002).
All remaining model procedures were performed in the R package VEGAN (Oksanen et al. 2007). We built dbRDA models using the function capscale. For all models, we applied the effects of the conditional matrix described above, and assessed the significance of “Highways” and “Releases”. We built three models: one implementing both predictor variables and the other two implementing each variable in isolation. We assessed the marginal significance of each predictor variable with the function anova.cca, using 99,999 permutations. We then repeated the procedure with Wolbachia infection status used as a predictor variable alongside “Highways” and “Releases”. While constructing models, we calculated variance-inflation factors (VIF) to check for multicollinearity between predictor variables, using the function vif.cca. All VIFs were < 1.1, so none were rejected.
Assessing barriers to dispersal: type I error testing
Although ordination methods such as dbRDA can detect genetic structure with greater power than traditional Mantel tests (Legendre and Fortin 2010), they can also generate more Type I errors (Kierepka and Latch 2015). For instance, when genetic structure exhibits IBD, spatial dependence of barrier variables and inadequate sampling can lead to significance being observed when no barrier effect exists (Kierepka and Latch 2016). We used 100 simulations performed in CDPOP (Landguth and Cushman 2010) to evaluate whether our dbRDA analyses were at high risk of Type I error. Parameters and methodology of our CDPOP simulations are detailed in Appendix B and our CDPOP input file is supplied in Appendix C. We considered that if dbRDA variables were significant for > 5 of our simulations, Type I error was likely.
Assessing barriers to dispersal: patch-based simulation analysis
Wolbachia invasions with bistable dynamics are particularly sensitive to barrier effects (Barton and Turelli 2011). Sensitivity to barrier effects increases exponentially as \(\widehat p\) approaches 0.5, and the \(\widehat p\) ≈ 0.35 of wMel in Ae. aegypti (Turelli and Barton 2017) indicates such an invasion can be stopped by a barrier half the strength of that required to stop an invasion with \(\widehat p\) ≈ 0.25 (Barton and Turelli 2011). Likewise, higher \(\widehat p\) means that slight disruptions to dispersal can slow an advancing wave considerably (Barton and Turelli 2011; Turelli and Barton 2017). Considering this, it is likely that barriers capable of producing even slight genetic structure will have an impact.
We used a patch-based simulator derived from Nemo2 (Guillaume and Rougemont 2006) and empirical data from Schmidt et al. (2017) to investigate the restrictive effects of the two Cairns highways on Wolbachia spread. This provides a means of translating between the magnitude of genetic structure produced by highways as estimated in our dbRDAs and the magnitude of the reductions in Wolbachia frequency across highways observed following releases in Parramatta Park (Schmidt et al. 2017). Our simulations modelled the spread of wMel through Parramatta Park in the absence of barriers, and contrasted these observation with the empirical data from Schmidt et al. (2017). Parameters, methodology and results are detailed in Appendix F.
Long-distance host movement and infection loss: estimating relatedness/kinship
We used SPAGeDi (Hardy and Vekemans 2002) to calculate Loiselle’s k (Loiselle et al. 1995) among pairs of individuals, and from this estimate the specific relationship category of each pair. Estimation procedures are described in Appendix D. In summary, pairs with kinship coefficient k > 0.1875 represented putative full-siblings, and those with 0.1875 > k > 0.0938 represented putative half-siblings (Iacchei et al. 2013). These estimates were tested using maximum-likelihood simulations, involving a “standard” test, which assumed that the kinship category assigned using k was more likely than the next most likely kinship category, and a “conservative” test that assumed that the kinship category assigned using k was less likely to be correct than the next most likely kinship category. We considered the distance of separation between full-siblings to indicate movement of adult females.
Long-distance host movement and infection loss: Wolbachia infection screening
Mosquitoes were screened for Wolbachia using the protocol of Lee et al. (2012). A summary of this method and our protocols for calculating infection titres is presented in Appendix E. From our Wolbachia-infection screening and relatedness analysis, we identified wMel-infected matrilineages as being any group of full-siblings containing at least one individual infected with Wolbachia. Uninfected individuals within infected matrilineages were considered to have failed to inherit the infection maternally or to have lost it during development. We estimated the rate of infection loss per generation (µ) by comparing the number of these individuals with the total number of full-siblings from infected matrilineages.
Results
Barriers to dispersal
ANOVAs performed on partial dbRDA models showed that only the “Highways” variable was predictive of genetic structure (Table 1, P < 0.05), and was significant irrespective of the inclusion of “Releases”. Based on partial Eta squared (ƞp2: Tabachnick and Fidell (1996)), “Highways” explained 1.7% of the variance in each model. A biplot of both “Highways” and “Releases” (Fig. S1) showed that “Highways” contributed to more variance on the first axis (1.30% of variation), while “Releases” contributed to more variance on the second axis (1.19% of variation), but no obvious clustering in scores was observed for either variable. These results suggest that highways exert a significant influence on genetic structure of Cairns Ae. aegypti, whereas patchy Wolbachia release histories do not.
Following this observation, we used the individual-based simulations conducted in CDPOP (see Appendix B) to investigate potential Type I error in this observation of genetic structure. As significance in our dbRDAs was assessed at 95% confidence, we considered that if more than five of the 100 simulated samples showed significance for a variable then observations regarding that variable would be at elevated risk of Type I error. However, only two of the samples showed significant structuring by “Highways” at the P < 0.05 level, which did not change when “Releases” was included. As this is below our threshold of five significant simulations, we conclude that the structuring effect of “Highways” observed in our dbRDAs is unlikely to be a Type I error. Sensitivity analyses (see Appendix A) demonstrated robust results for “Highways”, with ƞp2 effect sizes ranging from 0.0154–0.019.
Our patch-based simulations (Appendix F) showed that, when Wolbachia spread was modelled with no barrier effects, simulated infection frequencies in patches separated by highways were overestimated relative to the empirical infection frequencies in Schmidt et al. (2017), with an average overestimation across treatments of 6.5%. By comparison, no overestimation of infection frequencies was observed in patches not separated by highways. Overall, simulation of the Wolbachia invasion progress south of Bruce Highway indicated that barrier strength corresponded to an added 30–35 m of separation.
Estimates of long-distance movement
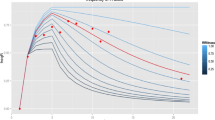
We detected 31 putative full-sibling groups that contained 43 full-sibling pairs, 41 of which were found within single ovitraps. Two pairs were spread between different plots (Fig. 2): one was split between CNW and CNE (239 m separation, k = 0.203); the other between CNW and PPS (1312 m separation, k = 0.235). Standard maximum-likelihood simulations gave strong support to both pairs being full-siblings (both P < 0.001). However, conservative simulations were unable to reject the hypotheses that either pair represented half-siblings (P < 0.001 and P < 0.02, respectively), but rejected them as unrelated (both P = 1). Nevertheless, the putative full-siblings from traps 1312 m apart had a k-score 25.3% higher than the full-sibling/half-sibling category boundary.
Pairwise relatedness (k) across geographic distance. Pairs of 0.0938 < k < 0.1875 are most likely half-sibs, those of k > 0.1875 are most likely full-sibs. Unrelated pairs of k < 0.04688 are not shown. Most related pairs were found within the same trap, but separation distances of up to 1312 m were observed
Among individuals sampled from within the same trap, we found a mixture of putative full-siblings, half-siblings and unrelated individuals. Of the 27 traps in which two individuals were sequenced, eight (29.63%) contained unrelated individuals, thirteen (48.15%) contained full-siblings and six (22.22%) contained half-siblings. Of the 20 traps where three individuals were sequenced, three (15%) contained no related individuals, five (25%) contained only full-siblings, four (20%) contained a single full-sibling pair, and one (5%) contained a single half-sibling pair.
Loss of Wolbachia
Triplicate PCR runs for wMel detection indicated that 60 of the 161 mosquitoes were infected with Wolbachia. We detected ten Wolbachia-infected matrilineages, containing 21 individuals. Importantly, we recorded a single case of infection loss in CNW. The sample from this matrilineage consisted of a pair of putative full-siblings (k = 0.376), one of which carried the infection (titre = 6.15) and one of which did not. Maximum-likelihood simulation confirmed that this pair were full-siblings (P < 0.001) and rejected the alternative hypotheses (both P = 1). We calculated a tentative probability of infection loss among offspring within infected matrilineages, giving a likelihood of loss of one in 21 (µ = 0.048). However, the 95% binomial confidence intervals around this estimate were very large (0.001, 0.238), and additional observations of infection loss will be necessary to make predictions about the impact of Wolbachia transmission loss.
Our finding of infection loss runs counter to previous estimates of µ = 0 among laboratory-reared Ae. aegypti (Hoffmann et al. 2014; Walker et al. 2011). However, individuals assayed in this study were oviposited in the field, suggesting that field conditions could be more conducive to infection loss. We compared Wolbachia infection titres among infected mosquitoes in our semi-field-reared Cairns sample and those of a field-reared sample from the neighbouring town of Gordonvale (see Appendix G), which had been subjected to releases in 2011 (cf. Hoffmann et al. 2011). The Gordonvale sample had titres that were almost three times higher than Cairns on average, but also three times as dispersed (x̅ = 18.4, σ = 10.8; compared with x̅ = 6.6, σ = 3.8 in Cairns). One individual from Gordonvale had a titre of only one Wolbachia gene copy for every three Aedes gene copies, while the lowest titre in the Cairns sample was more than three times higher than this. The highest titre in Gordonvale was 56 Wolbachia gene copies for each Aedes gene copy, more than double the highest recorded in Cairns.
Discussion
Our study produced three main findings: (i) highways exert a modest but statistically significant influence on Ae. aegypti genetic structure; (ii) adult Ae. aegypti can travel > 1 km; and (iii) Ae. aegypti in Cairns lose the wMel infection at some non-zero frequency (μ > 0). Each phenomenon is expected to have a slowing effect on the spatial spread of a wMel invasion (Barton and Turelli 2011; Turelli and Barton 2017). Because (ii) and (iii) were based on single observations, we cannot confidently predict their frequency among Ae. aegypti in Cairns, and thus it is unclear to what degree these are likely to affect spread. However, field observations of slow spread of wMel through Cairns (Schmidt et al. 2017; Turelli and Barton 2017) are congruent with some long-distance movement and occasional infection loss.
Highways act as dispersal barriers
While previous studies suggested that roads could be barriers to Ae. aegypti dispersal (Hemme et al. 2010; Russell et al. 2005), we provide the first evidence for such an effect through explicit hypothesis testing within a landscape genetics framework. We detected a small but statistically significant barrier effect of highways, corresponding to 1.7% of dbRDA variance in genetic distance between individuals. Our simulations of Wolbachia invasion progress south of Bruce Highway (Appendix F) showed that barrier strength corresponded to an added 30–35 m of separation, which could be investigated further with Ae. aegypti population modelling tools such as Skeeter Buster (Magori et al. 2009). The wMel invasion observed at Parramatta Park was slow (100–200 m per year: Schmidt et al. 2017), with the infection frequencies at the wave front being only slightly above the critical frequency \(\widehat p\) ≈ 0.35 (Turelli and Barton 2017). Therefore, even a small added “cost” to cross the highway could slow the invasion considerably (Barton and Turelli 2011). If the restrictive effect of highways on dispersal increase with highway width and traffic levels, then many urban highways in cities earmarked for future wMel releases would likely be effective barriers to spread.
On the other hand, the restrictive effects of highways on dispersal can strengthen the invasion within the area they enclose. The habitat patches along these highway boundaries will have increased infection frequencies relative to patches in regions that are not subdivided, and could fortify the local invasion from the counteracting effect of uninfected immigrants. Following the 2013 Wolbachia releases in Cairns, the largest and most successfully invaded release site at Edge Hill/Whitfield recorded an influx of uninfected Ae. aegypti at the start of the 2014/2015 wet season (Schmidt et al. 2017). This site was deemed particularly vulnerable to such reinvasion, due to its greater connectivity to surrounding uninfected regions and very low dry-season mosquito density. Ideally, future release sites in areas with low dry-season densities should be positioned adjacent to dispersal barriers such as highways, as this may help reduce the threat of reinvasion.
Long-distance movement of Aedes aegypti adults
Analyses of local kinship patterns have provided several useful inferences regarding Ae. aegypti ecology. The pair of probable full-siblings collected from ovitraps 1312 m apart most likely indicates movement of a single female between oviposition events, and represents either the extreme end of the flight range in Ae. aegypti or some combination of active and passive dispersal. Previous maximum dispersal estimates of gravid Ae. aegypti females have been less than a kilometre in a single gonotrophic (egg producing) cycle (Honório et al. 2003; Reiter et al. 1995). In our study, 7 days passed between deployment and sampling of the two ovitraps 1312 m apart, which is long enough for an additional unobserved egg deposition somewhere between them (Christophers 1960). If we assume this distance to reflect active dispersal, an additional gonotrophic cycle is likely. The length of a single such cycle is rarely more than a few days, particularly when temperatures exceed 30 °C (Christophers 1960) as they did during our sampling. If the distance was crossed in a single gonotrophic cycle, the average daily speed would have been an order of magnitude greater than the previous estimates of average female flight speed near Cairns of 17–25 m/day (Muir and Kay 1998). Therefore, passive, human-assisted transport is likely to have occurred. Adults of the related mosquito Ae. albopictus have been found in cars stopped at major roads in Barcelona, Spain (Eritja et al. 2017).
Using a continuous coefficient of relationship such as k to assign pairs of individuals to discrete kinship categories can be problematic when scores are close to the critical cutoff values. Pairing this method with conservative maximum-likelihood estimation resolved some of the clearer distinctions (i.e., the full-siblings exhibiting Wolbachia loss) but not others (i.e., the putative full-siblings exhibiting long-distance movement). However, the k-scores of putative full-siblings and putative half-siblings were clearly separated from each other relative to the variability in k-scores within each category (Fig. 2), and the putative full-siblings from traps 1312 m apart had a k-score 25.3% higher than the category boundary, suggesting that this pair was much more likely to represent full-siblings than half-siblings.
We consider the distribution of k-scores within kinship categories to reflect the power of genome-wide SNPs for inferring relationships (Blouin 2003). Inferring dispersal from relatedness also avoids potential biases resulting from laboratory-raised Ae. aegypti used in MRR studies failing to develop experience in local conditions that will inform their future oviposition choices (Kaur et al. 2003; Ruktanonchai et al. 2015). Our findings are broadly consistent with the results of several MRR studies showing potential for long-distance movement in Ae. aegypti imagoes (Honório et al. 2003; Reiter et al. 1995), and suggest that this method provides an alternative to MRR for studying dispersal.
Infection loss in Cairns Aedes aegypti
We found the first evidence in support of intergenerational infection loss (µ > 0) in the Ae. aegypti/wMel system, albeit from a single occurrence. A comparable loss of µ = 0.033 was recorded for the wAlbA infection in field-collected Ae. albopictus (Kittayapong et al. 2002), though this concerned a natural Wolbachia infection rather than a transinfection. Transmission of wMel in Ae. aegypti has previously been estimated as 100% in the field, but this was based on assaying eggs that were oviposited in the laboratory (Hoffmann et al. 2014). By comparison, samples in this study were eclosed from eggs that might have spent days in the field before collection, potentially exposing them to stressors such as high-heat fluctuations, which have been observed to affect titre of wMel transmission in laboratory populations (Ross et al. 2017). Fluctuating high temperatures could be the reason for the very low titres recorded for some Gordonvale individuals (Appendix G), though this could also be due a reduction in titre with age, which has been recorded for wAlbA in Ae. albopictus males (Tortosa et al. 2010). The inclusion of blood-fed females in the Gordonvale sample may explain the occurrence of some very high wMel titres, which can double in blood-fed Ae. aegypti (Frentiu et al. 2014).
The wMel strain successfully invaded Ae. aegypti populations in Gordonvale and Yorkey’s Knob, two quasi-isolated release sites near Cairns, in 2011 (Hoffmann et al. 2011) and these areas have maintained infection frequencies close to 95% for years without reaching fixation (Hoffmann et al. 2014). Under the assumptions of µ = 0, high migration rates of uninfected gravid females of 0.03 into Gordonvale and 0.06 into Yorkey’s Knob would be required to maintain these infection frequencies (Hoffmann et al. 2014). Alternatively, the failure to reach fixation in these areas could be due to a combination of migration and infection loss, which would mean that migration rates of uninfected, gravid females may be lower than 0.06.
Conclusions
Our study has provided empirical evidence for three processes predicted to slow down the spatial spread of wMel in Ae. aegypti, using a landscape genomics analytical framework and molecular assays of Wolbachia infection. This approach could be extended to other host/Wolbachia systems that are increasingly considered for the biocontrol of disease vectors and pests. Non-perfect maternal transmission of wMel in Ae. aegypti may not occur in other Wolbachia strains such as wAlbB, whose density shows greater constancy under fluctuating high temperatures (Ross et al. 2017). Also, it is as yet unclear whether the observed transmission failure of wMel occurs at a high enough frequency to affect invasion dynamics, and a more extensive field test of transmission fidelity will be necessary to derive an accurate estimate of μ. On the other hand, regardless of the Wolbachia strain deployed, dispersal barriers and higher incidences of long-range dispersal are potential challenges for any Wolbachia invasion strategy requiring spatial spread (Turelli and Barton 2017).
Data archiving
Aligned sequences for Aedes aegypti used in this study are available from NCBI SRA, retrievable with accession number SRP118883. Geographical coordinates for each mosquito are also available.
References
Barton N, Turelli M (2011) Spatial waves of advance with bistable dynamics: cytoplasmic and genetic analogues of Allee effects. Am Nat 178(3):E48–E75
Blouin MS (2003) DNA-based methods for pedigree reconstruction and kinship analysis in natural populations. Trends Ecol Evol 18(10):503–511
Bogoch II, Brady OJ, Kraemer M, German M, Creatore MI, Kulkarni MA et al. (2016) Anticipating the international spread of Zika virus from Brazil. Lancet 387(10016):335–336
Brown S, Severson D, Smith L, Knudson D (2001) Integration of the Aedes aegypti mosquito genetic linkage and physical maps. Genetics 157(3):1299–1305
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22(11):3124–3140
Cho K, Dupuis J (2009) Handling linkage disequilibrium in qualitative trait linkage analysis using dense SNPs: a two-step strategy. BMC Genet 10(1):44
Christophers S (1960) Aëdes aegypti (L.) the yellow fever mosquito: its life history, bionomics and structure. Cambridge University Press, London
Clancy DJ, Hoffmann AA (1998) Environmental effects on cytoplasmic incompatibility and bacterial load in Wolbachia-infected Drosophila simulans. Entomol Exp Appl 86(1):13–24
Clark JS, Fastie C, Hurtt G, Jackson ST, Johnson C, King GA et al. (1998) Reid’s paradox of rapid plant migration: dispersal theory and interpretation of paleoecological records. Biosci 48(1):13–24
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA et al. (2011) The variant call format and VCFtools. Bioinformatics 27(15):2156–2158
Eritja R, Palmer J, RB, Roiz D, Sanpera-Calbet I, Bartumeus F. (2017). Population Association of America 2017 Annual Meeting: Chicago
Ferguson NM, Kien DTH, Clapham H, Aguas R, Trung VT, Chau TNB et al. (2015) Modeling the impact on virus transmission of Wolbachia-mediated blocking of dengue virus infection of Aedes aegypti. Sci Transl Med 7(279):279ra237
Focks DA, Brenner RJ, Hayes J, Daniels E (2000) Transmission thresholds for dengue in terms of Aedes aegypti pupae per person with discussion of their utility in source reduction efforts. Am J Trop Med Hyg 62(1):11–18
Frentiu FD, Zakir T, Walker T, Popovici J, Pyke AT, van den Hurk A et al. (2014) Limited dengue virus replication in field-collected Aedes aegypti mosquitoes infected with Wolbachia. PLoS Negl Trop Dis 8(2):e2688
Goldberg CS, Waits LP (2010) Quantification and reduction of bias from sampling larvae to infer population and landscape genetic structure. Mol Ecol Resour 10(2):304–313
Guillaume F, Rougemont J (2006) Nemo: an evolutionary and population genetics programming framework. Bioinformatics 22(20):2556–2557
Hancock PA, White VL, Callahan AG, Godfray CH, Hoffmann AA, Ritchie SA (2016) Density-dependent population dynamics in Aedes aegypti slow the spread of wMel. Wolbachia J Appl Ecol 53(3):785–793
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2(4):618–620
Harrington LC, Scott TW, Lerdthusnee K, Coleman RC, Costero A, Clark GG et al. (2005) Dispersal of the dengue vector Aedes aegypti within and between rural communities. Am J Trop Med Hyg 72(2):209–220
Hemme RR, Thomas CL, Chadee DD, Severson DW (2010) Influence of urban landscapes on population dynamics in a short-distance migrant mosquito: evidence for the dengue vector Aedes aegypti. PLoS Negl Trop Dis 4(3):e634
Hoffmann A, Montgomery B, Popovici J, Iturbe-Ormaetxe I, Johnson P, Muzzi F et al. (2011) Successful establishment of Wolbachia in Aedes populations to suppress dengue transmission. Nature 476(7361):454–457
Hoffmann A, Turelli M (1997) Cytoplasmic incompatibility in insects. In: ON SL, AA H, JH W (eds) Influential passengers: inherited microorganisms and arthropod reproduction. Oxford University Press, Oxford, p 42–80
Hoffmann AA, Iturbe-Ormaetxe I, Callahan AG, Phillips BL, Billington K, Axford JK et al. (2014) Stability of the wMel Wolbachia infection following invasion into Aedes aegypti populations. PLoS Negl Trop Dis 8(9):e3115
Hoffmann AA, Turelli M, Harshman LG (1990) Factors affecting the distribution of cytoplasmic incompatibility in Drosophila simulans. Genetics 126(4):933–948
Honório NA, Silva WdC, Leite PJ, Gonçalves JM, Lounibos LP, Lourenço-de-Oliveira R (2003) Dispersal of Aedes aegypti and Aedes albopictus (Diptera: Culicidae) in an urban endemic dengue area in the State of Rio de Janeiro, Brazil. Mem Inst Oswaldo Cruz 98(2):191–198
Iacchei M, Ben-Horin T, Selkoe KA, Bird CE, García-Rodríguez FJ, Toonen RJ (2013) Combined analyses of kinship and FST suggest potential drivers of chaotic genetic patchiness in high gene-flow populations. Mol Ecol 22(13):3476–3494
Ikeda T, Ishikawa H, Sasaki T (2003) Infection density of Wolbachia and level of cytoplasmic incompatibility in the Mediterranean flour moth, Ephestia kuehniella. J Invertebr Pathol 84(1):1–5
Kaur J, Lai Y, Giger A (2003) Learning and memory in the mosquito Aedes aegypti shown by conditioning against oviposition deterrence. Med Vet Entomol 17(4):457–460
Kierepka E, Latch E (2015) Performance of partial statistics in individual-based landscape genetics. Mol Ecol Resour 15(3):512–525
Kierepka E, Latch E (2016) Fine-scale landscape genetics of the American badger (Taxidea taxus): disentangling landscape effects and sampling artifacts in a poorly understood species. Heredity 116(1):33–43
Kittayapong P, Baisley KJ, Sharpe RG, Baimai V, O’Neill SL (2002) Maternal transmission efficiency of Wolbachia superinfections in Aedes albopictus populations in Thailand. Am J Trop Med Hyg 66(1):103–107
Landguth EL, Cushman S (2010) CDPOP: a spatially explicit cost distance population genetics program. Mol Ecol Resour 10(1):156–161
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25
Lee SF, White VL, Weeks AR, Hoffmann AA, Endersby NM (2012) High-throughput PCR assays to monitor Wolbachia infection in the dengue mosquito (Aedes aegypti) and Drosophila simulans. Appl Environ Microbiol 78(13):4740–4743
Legendre P, Anderson MJ (1999) Distance-based redundancy analysis: testing multispecies responses in multifactorial ecological experiments. Ecol Monogr 69(1):1–24
Legendre P, Fortin MJ (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol Ecol Resour 10(5):831–844
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae) Am J Bot 82:1420–1425
Maciel-De-Freitas R, Codeco CT, Lourenco-De-Oliveira R (2007) Daily survival rates and dispersal of Aedes aegypti females in Rio de Janeiro, Brazil. Am J Trop Med Hyg 76(4):659–665
Magori K, Legros M, Puente ME, Focks DA, Scott TW, Lloyd AL et al. (2009) Skeeter Buster: a stochastic, spatially explicit modeling tool for studying Aedes aegypti population replacement and population suppression strategies. PLoS Negl Trop Dis 3(9):e508
McGraw EA, O’Neill SL (2013) Beyond insecticides: new thinking on an ancient problem. Nat Rev Microbiol 11(3):181–193
Muir LE, Kay BH (1998) Aedes aegypti survival and dispersal estimated by mark-release-recapture in northern Australia. Am J Trop Med Hyg 58(3):277–282
Nene V, Wortman JR, Lawson D, Haas B, Kodira C, Tu ZJ et al. (2007) Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316(5832):1718–1723
Oksanen J, Kindt R, Legendre P, O’Hara B, Stevens MHH, Oksanen MJ et al. (2007) The vegan package. Community Ecol Package 10:631–637
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7(5):e37135
Rašić G, Endersby-Harshman N, Tantowijoyo W, Goundar A, White V, Yang Q et al. (2015a) Aedes aegypti has spatially structured and seasonally stable populations in Yogyakarta, Indonesia. Parasit Vectors 8(1):610
Rašić G, Filipović I, Weeks AR, Hoffmann AA (2014) Genome-wide SNPs lead to strong signals of geographic structure and relatedness patterns in the major arbovirus vector, Aedes aegypti. BMC Genom 15:1
Rašić G, Schama R, Powell R, Maciel-de Freitas R, Endersby-Harshman NM, Filipović I et al. (2015b) Contrasting genetic structure between mitochondrial and nuclear markers in the dengue fever mosquito from Rio de Janeiro: implications for vector control. Evol Appl 8(9):901–915
Reiter P (2007) Oviposition, dispersal, and survival in Aedes aegypti: implications for the efficacy of control strategies. Vector Borne Zoonotic Dis 7(2):261–273
Reiter P, Amador MA, Anderson RA, Clark GG (1995) Dispersal of Aedes aegypti in an urban area after blood feeding as demonstrated by rubidium-marked eggs. Am J Trop Med Hyg 52(2):177–179
Ritchie SA (2001) Effect of some animal feeds and oviposition substrates on Aedes oviposition in ovitraps in Cairns, Australia. J Am Mosq Control Assoc. Mosq News 17(3):206–208
Ross PA, Endersby NM, Hoffmann AA (2016) Costs of three Wolbachia infections on the survival of Aedes aegypti larvae under starvation conditions. PLoS Negl Trop Dis 10(1):e0004320
Ross PA, Wiwatanaratanabutr I, Axford JK, White VL, Endersby-Harshman NM, Hoffmann AA (2017) Wolbachia infections in Aedes aegypti differ markedly in their response to cyclical heat stress. PLoS Pathog 13(1):e1006006
Rousset F (2000) Genetic differentiation between individuals. J Evol Biol 13(1):58–62
Ruktanonchai N, Lounibos L, Smith D, Allan S (2015) Experience-and age-mediated oviposition behaviour in the yellow fever mosquito Stegomyia aegypti ( = Aedes aegypti). Med Vet Entomol 29(3):255–262
Russell RC, Webb C, Williams C, Ritchie S (2005) Mark–release–recapture study to measure dispersal of the mosquito Aedes aegypti in Cairns, Queensland, Australia. Med Vet Entomol 19(4):451–457
Schmidt TL, Barton NH, Rašić G, Turley AP, Montgomery BL, Iturbe-Ormaetxe I et al. (2017) Local introduction and heterogeneous spatial spread of dengue-suppressing Wolbachia through an urban population of Aedes aegypti. PLoS Biol 15(5):e2001894
Suh E, Dobson SL (2013) Reduced competitiveness of Wolbachia infected Aedes aegypti larvae in intra- and inter-specific immature interactions. J Invertebr Pathol 114(2):173–177
Tabachnick BG, Fidell LS (1996) Using multivariate statistics, 3rd edn. Harper Collins, New York, NY
Tortosa P, Charlat S, Labbé P, Dehecq J-S, Barré H, Weill M (2010) Wolbachia age-sex-specific density in Aedes albopictus: a host evolutionary response to cytoplasmic incompatibility? PLoS ONE 5(3):e9700
Turelli M (2010) Cytoplasmic incompatibility in populations with overlapping generations. Evolution 64(1):232–241
Turelli M, Barton NH (2017) Deploying dengue-suppressing Wolbachia: robust models predict slow but effective spatial spread in Aedes aegypti. Theor Popul Biol 115:45–60
Ulrich JN, Beier JC, Devine GJ, Hugo LE (2016) Heat sensitivity of wMel Wolbachia during Aedes aegypti development. PLoS Negl Trop Dis 10(7):e0004873
Walker T, Johnson P, Moreira L, Iturbe-Ormaetxe I, Frentiu F, McMeniman C et al. (2011) The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature 476(7361):450–453
Weaver SC, Lecuit M (2015) Chikungunya virus and the global spread of a mosquito-borne disease. N Engl J Med 372(13):1231–1239
Xi Z, Khoo CC, Dobson SL (2005) Wolbachia establishment and invasion in an Aedes aegypti laboratory population. Science 310(5746):326–328
Acknowledgements
We would like to thank Eliminate Dengue Cairns, particularly field officer Angela Caird, for assistance with ovitrap deployment. We thank the Cairns householders participating in the study for granting permission to deploy ovitraps on their property, as well as Scott Ritchie and Christopher Paton from the Centre for Biosecurity in Tropical Infectious Diseases, James Cook University, for assisting with the processing of field collections. The National Health and Medical Research Council provided funding for this research through a Programme grant and Fellowship grant to A.A. Hoffmann.
Author contributions
AA Hoffmann, G Rašić, and TL Schmidt conceived of and designed the study. TL Schmidt, G Rašić, and I Filipović collected and dried the samples. TL Schmidt performed the laboratory work and conducted the analyses, with assistance from G Rašić, and computational support from I Filipović. TL Schmidt wrote the manuscript with assistance from AA Hoffmann and G Rašić. All authors gave final approval for this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Schmidt, T.L., Filipović, I., Hoffmann, A.A. et al. Fine-scale landscape genomics helps explain the slow spatial spread of Wolbachia through the Aedes aegypti population in Cairns, Australia. Heredity 120, 386–395 (2018). https://doi.org/10.1038/s41437-017-0039-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-017-0039-9
This article is cited by
-
Aedes aegypti microbiome composition covaries with the density of Wolbachia infection
Microbiome (2023)
-
Urban population structure and dispersal of an Australian mosquito (Aedes notoscriptus) involved in disease transmission
Heredity (2023)
-
Genomic databanks provide robust assessment of invasive mosquito movement pathways and cryptic establishment
Biological Invasions (2023)
-
Remnants of horizontal transfers of Wolbachia genes in a Wolbachia-free woodwasp
BMC Ecology and Evolution (2022)
-
Bridging landscape ecology and urban science to respond to the rising threat of mosquito-borne diseases
Nature Ecology & Evolution (2022)