Abstract
Purpose
Cardiovascular disease (CVD) is the leading cause of death in adults in the United States, yet the benefits of genetic testing are not universally accepted.
Methods
We developed the “HeartCare” panel of genes associated with CVD, evaluating high-penetrance Mendelian conditions, coronary artery disease (CAD) polygenic risk, LPA gene polymorphisms, and specific pharmacogenetic (PGx) variants. We enrolled 709 individuals from cardiology clinics at Baylor College of Medicine, and samples were analyzed in a CAP/CLIA-certified laboratory. Results were returned to the ordering physician and uploaded to the electronic medical record.
Results
Notably, 32% of patients had a genetic finding with clinical management implications, even after excluding PGx results, including 9% who were molecularly diagnosed with a Mendelian condition. Among surveyed physicians, 84% reported medical management changes based on these results, including specialist referrals, cardiac tests, and medication changes. LPA polymorphisms and high polygenic risk of CAD were found in 20% and 9% of patients, respectively, leading to diet, lifestyle, and other changes. Warfarin and simvastatin pharmacogenetic variants were present in roughly half of the cohort.
Conclusion
Our results support the use of genetic information in routine cardiovascular health management and provide a roadmap for accompanying research.
Similar content being viewed by others
INTRODUCTION
Cardiovascular disease (CVD) is the leading cause of mortality in the United States, leading to nearly 1 in every 4 deaths [1]. The role of inherited susceptibility to CVD is well established, from rare monogenic disorders to polygenic traits. More than half of the genes on the American College of Medical Genetics and Genomics (ACMG) secondary finding (SF) list are associated with cardiovascular phenotypes [2]. Although the utility of genetic testing in pediatrics is well established and precision medicine approaches can shift reactive care to stratified risk assessment with the potential to mitigate risk through adjustments of diet, lifestyle, or medications [3], few parallel applications exist in the adult clinical care system.
Among the many obstacles to widespread adoption of adult genetic testing is the perceived lack of clinical utility and availability of point-of-care genetic testing during physician–patient interactions. Rare alleles with large effect are often considered too infrequent or else only able to provide information superseded by family history. In selected families, individuals will be referred to genetic testing as an adjunct to standard care, reflecting a division between clinical genetics care and other service branches. Conversely, common alleles are regarded as only able to contribute minor risk compared to etiological factors. The latter has been somewhat addressed by the development of polygenic risk scores (PRS) based upon the summation of overall genotype risk and the recognition that extreme scores can approach the predictive value of single alleles with high penetrance. Nevertheless, genetic testing’s practicalities remain limiting, and the integration of a genetic screen in a standard CVD care program is not commonplace.
We, therefore, developed “HeartCare," a next-generation sequence–based DNA capture panel, to provide a genetic screen and to serve as the implementation tool to identify and address the practical issues involved in the delivery of genetic test results for CVD and related disease risk. The test consists of a 158 gene oligonucleotide capture-based panel, evaluating (1) Mendelian, actionable conditions including cardiomyopathies, aortopathies, arrhythmias, and dyslipidemias; (2) a coronary artery disease (CAD) PRS; (3) variants in the LPA gene encoding Lipoprotein(a) that are an independent risk factor for atherosclerotic CVD events; and (4) pharmacogenetic variants contributing to simvastatin-induced myopathy and warfarin metabolism.
A long-term aim of this program is to bring together the clinical translation of medicine and the accompanying gathering of detailed phenotype records from many patients, using an infrastructure that is compliant for research. The HeartCare genetic screen has enabled the identification of patients with elevated risk and offers a model for actionable return of results utilizing established clinical service delivery lines in adult cardiovascular health care. Further, HeartCare data management has enabled data mining, analysis, and follow-up in the well-characterized participant population.
MATERIALS AND METHODS
Participants
The study was approved by the Baylor College of Medicine (BCM) Institutional Review Board (protocol number: H-43884) and was open to all adult patients between the ages of 18 and 85 years at participating outpatient Baylor clinics (Table 1) at no cost to them, and insurance was not billed. Participants were informed of the study at the time of a clinical visit via posters and brochures in the waiting areas, study coordinators onsite, or physician referrals (workflow outlined in Fig. 1a). Interested patients were asked to give a blood or saliva sample and provide written consent to participate after undergoing pretest counseling. As the gene panel was offered to all patients, there were no required indications and there was no targeting of patient groups within a particular clinic. Family history for cardiac disease was also not required, but was collected if voluntarily provided by participants. Indications for testing and race/ethnicity were self-reported, the latter based on US Census–standardized terms. The HeartCare panel was added to the local electronic medical record (EMR) (Epic Systems Corporation) test menu for facile ordering and ease of incorporation of demographic and clinical data elements required to interpret results.
(a) Workflow from enrollment to return of results. Final report was added to local electronic medical record (EMR) for physician determination if any additional studies, medications, or referrals were required. (b) Example clinical report with four sections corresponding to (1) rare Mendelian conditions, (2) coronary artery disease polygenic risk score, (3) LPA gene risk variants, (4) pharmacogenetic variants related to simvastatin and warfarin. Subsequent pages conveyed additional information specific to each section, including guideline-based recommendations.
HeartCare Web Portal
The BCM Human Genome Sequencing Center (HGSC) developed the HeartCare Web Portal to receive test orders and facilitate data management for this project. The HeartCare Web Portal provides a secure, HIPAA compliant environment that can receive sensitive patient/participant information at enrollment and deliver finished reports when required. The portal, therefore, serves as both the entry point for receiving test orders from clinical providers and the mechanism for delivering test results back to the providers. The service generates a local identifier for each sample and uses that to track all genetic studies in a blinded fashion throughout the sequencing workflow. When reporting data back to participants, the HeartCare Portal allows the reuniting of personal data to the genetic data and transports that back to the referring physician and the patient’s medical record. The HeartCare Portal interfaced with the Epic EMR and is adaptable to similar other systems.
Panel design and content
The HeartCare genetic screen tests 158 genes related to inherited Mendelian cardiovascular conditions associated with increased risk of aortic aneurysms, cardiomyopathies, arrhythmias, and dyslipidemias (Table S1). The panel includes cardiac-related ACMG SF™ genes [2] and additional genes deemed actionable by the internal HeartCare group (“ACMG™, ACMG SF™, ACMG 59™, ACMG 56™, and related words and designs incorporating ACMG™, are trademarks of the American College of Medical Genetics and Genomics and may not be used without permission.”). For the latter, scientific literature, OMIM-morbid genes [4], and other laboratory panels were each reviewed to make a comprehensive list of genes linked to inherited CV phenotypes. We also included 50 single-nucleotide polymorphism (SNP) sites that had achieved genome-wide significance for association with CAD as part of a PRS as previously described by Khera et al. (Table S2) [5]. Two LPA risk alleles associated with high Lp(a) levels and a corresponding increased risk of CVD in European populations [6] were also analyzed (Table S3). Finally, specific Clinical Pharmacogenetics Implementation Consortium (CPIC)–curated gene variants in SLCO1B1 and CYP2C9/VKORC1 affecting the metabolism of simvastatin and warfarin, respectively, were also included (Table S4) [7, 8].
Sample collection and DNA isolation
Blood samples were collected in PAXgene Blood DNA tube (PreAnalytiX, Becton-Dickinson) or Hemogard K2 EDTA tubes (Becton-Dickinson). DNA was isolated with PAXgene blood DNA (PreAnalytix, QIAGEN) or Gentra Puregene Blood kits (QIAGEN), respectively. The saliva samples were collected in Oragene Dx OGR-500 tubes (DNA Genotek) and isolated with PrepIT L2P reagents (DNA Genotek). Extractions were done following manufacturer instruction in the HGSC CLIA-certified laboratory. DNA was quantified using Quant-iT PicoGreen dsDNA Assay kits (ThermoFisher) and its quality estimated by electrophoresis.
Capture, sequencing, and primary analysis
The assay was a NimbleGen 641 Kb oligonucleotide capture panel, including the 158 genes described above. An additional 5.8 Kb and 23.4 Kb of IDT lockdown probes were added to improve capture of low coverage regions and include pharmacogenetic and PRS variants, respectively. Each sample was tagged with a DNA barcode and capture pool contained a total of 35 samples, and 70 samples were run per lane. Average DNA sequence coverage across the captured region was ~400× with >99% of bases covered above 20× (Figs. S1, S2). Sequence alignment with BWA-MEM [9] and variant identification utilized Atlas and the Mercury Pipeline [10], followed by Sanger confirmation for reportable variants. Copy-number variant (CNV) calls were made via Atlas-CNV [11] and confirmed with multiplex ligation-dependent probe amplification (MRC-Holland). Sequencing was performed using the Illumina HiSeq 2500 platform in the BCM HGSC CAP/CLIA-certified clinical laboratory (HGSC-CL).
Variant interpretation and clinical reporting
A HeartCare Variant Interpretation Group was formed that included clinical geneticists and cardiologists with expertise in cardiovascular genetics. Variants from the 158 Mendelian genes were classified according to ACMG/AMP interpretation guidelines [12] with gene specific recommendations used if applicable. For variants of uncertain significance (VUS) potentially related to the patient’s phenotype, additional clinical and family information was requested from the ordering clinician to provide evidence to support reclassification. In the event that reclassification was not possible, referral to adult genetics specialists for additional evaluation, including family segregation studies, was made when appropriate. Thus, the final clinical report included pathogenic and likely pathogenic variants in accordance with ACMG/AMP guidelines that recommend variants of uncertain significance not be used in clinical decision making [12]. Such VUS remained available in the portal for clinician review. As per ACMG recommendations, reanalysis of primary data is ongoing and may also lead to reclassification as more evidence becomes available [13].
The PRS was calculated as previously described [5], and a positive result was returned if a patient was found to be above the 95th percentile of risk. Patients were informed of their high PRS test result by their physician, who subsequently counseled them on recommended lifestyle modifications American Heart Association (AHA) life’s simple seven [14]. A positive LPA result was reported if an LPA risk allele was detected. A disclaimer was present on the report describing the uncertain applicability of the PRS and LPA findings in individuals of non-European ancestries. For pharmacogenetic findings, star alleles were determined based on the variants detected and interpreted using CPIC guidelines [7, 8]. Specific recommendations related to that finding were also included based on accepted society guidelines.
Final reports (Fig. 1b) appeared in the HeartCare Portal, where the laboratory director approved them, followed by upload into the Epic EMR as a PDF document attached to the test order for physician review. Specific parts of the LPA results were also structured as HL7 V2 messages to Epic as a first step for clinical decision support of best practice advisories [15]. The HGSC-developed Neptune platform was used for managing the connection between the clinical laboratory and the EMR [16].
Return of results
Clinical reports were returned to the physician for review and determination if any changes to the care plan were indicated. Negative results and pharmacogenetic-only reports were sent directly to patients with a physician letter (mail or Epic MyChart). Positive results required an office visit or telephone follow-up for full disclosure. Complimentary genetic counseling services were available through ConsultageneTM (consultagene.org), and referral to adult genetics and other specialists for additional workup and family cascade testing was streamlined. Clinician surveys were conducted for positive cases to determine the usefulness of the result in managing a patient’s health and assess the overall benefit of the test.
Statistical analysis
The 95% confidence intervals (CI) were calculated for the reported diagnostic using the Wald method. The Pearson’s chi-squared test was used to evaluate statistical differences in medical management between groups.
RESULTS
From December 2018 through July 2020, a total of 878 patients were approached and 713 initially enrolled in the study, resulting in a refusal rate of ~19% (95% CI, 16–22%). The most common reason for declining participation was potential impact on life insurance. Four individuals were withdrawn either by request or death during the study, leaving 709 who completed analysis with the HeartCare gene panel. Race/ethnicity of participating patients was consistent with overall demographics within the participating clinics (Table 1). While patients frequently had multiple indications, the most common reasons for testing were dyslipidemia at 46% (n = 325; 95% CI, 42–50%), arrhythmia at 20% (n = 141; 95% CI, 17–23%), and cardiomyopathy at 12% (n = 87; 95% CI, 10–215%) (Table 1). Overall, a genetic finding with potential clinical implications was found in 32% of participants (n = 225; 95% CI, 28–35%) when considering rare, monogenic conditions, elevated polygenic risk, and LPA risk alleles alone (Fig. 2a). Among Mendelian conditions, the yield was highest among patients with a cardiomyopathy indication, where 13% (n = 11; 95% CI, 7–21%) harbored a likely pathogenic or pathogenic variant in a related gene (Table 1). This was followed by aortopathy, dyslipidemia, and arrhythmia indications, in which 12% (n = 4; 95% CI, 4–28%), 10% (n = 32; 95% CI, 7–14%), and 9% (n = 13; 95% CI, 5–15%) of such patients carried a disease-associated variant, respectively (Table 1). Overall, some 9% (n = 64; 95% CI, 7–11%) of patients were molecularly diagnosed with a cardiac-related Mendelian disorder (Table 2, Fig. 2b).
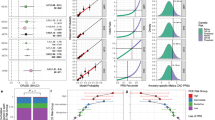
(a) HeartCare positive finding breakdown by category among the 709 study participants. LPA Lipoprotein(a) gene risk allele; LP/P likely pathogenic or pathogenic variants in 158 genes associated with rare, Mendelian conditions; PGx pharmacogenetic allele; PRS polygenic risk score. (b) Category breakdown among actionable findings in the patients testing positive (n = 64) for an actionable (pathogenic/likely pathogenic) finding in the 158 gene panel.
Twenty percent (n = 144; 95% CI, 18–23%) carried an LPA risk allele, and 9.3% (n = 66; 95% CI, 7–12%) belonged to the high-risk group according to the PRS (Fig. 2a). Nearly half of the cohort had a positive pharmacogenetic result, with 25% (n = 182; 95% CI, 46–53%) for simvastatin, 15% (n = 105) for warfarin, and 9% (n = 62) for both medications (Fig. 2a, Fig. S3). Notably, 30% and 9% of individuals prescribed either warfarin or simvastatin had a genetic variant impacting initial warfarin dosing or risk of simvastatin-induced myopathy, respectively (Fig. S4). Only 20% (n = 29) of patients had known Lp(a) levels prior to HeartCare (Fig. S5a). As of submission of the manuscript, Lp(a) levels had been measured in 59% (n = 85) of patients with a newly found LPA risk allele, with 82% (n = 70) found to have elevated levels and the corresponding risk for CVD (Fig. S5b). Reflecting the limited applicability of these known LPA alleles to non-European ancestries, Hispanic individuals made up two-thirds of cases (n = 10) with normal Lp(a) levels and a positive LPA risk allele despite representing only 10% of the entire cohort (Fig. S5c).
Clinician feedback was positive among those surveyed (n = 13) regarding the usefulness of HeartCare for the care of a subset of patients who tested positive on the 158 gene panel, PRS, or LPA risk allele (n = 122). Clinicians reported that the HeartCare results were beneficial to the patient overall (median 4 on a scale of 1 = “not at all beneficial” to 6 = “extremely beneficial”), with the 158 gene panel receiving the highest usefulness rating (median 5 on a scale of 1 = “not at all useful” to 6 = “extremely useful”) for managing a patient’s health (Table 3). In the majority (60.0%) of cases, the clinician agreed or strongly agreed that the patient’s HeartCare results improved overall clinical care beyond what clinical evaluation and nongenetic diagnostics alone could have achieved. Notably, clinicians reported recommending management changes for 84.4% (n = 103/122) of patients based on their HeartCare result (Table 3). Such interventions included but were not limited to medication changes, cascade testing, specialist referrals, and additional cardiac tests. Management changes were made more often in patients with a new diagnosis established via HeartCare (93.3%) than in those whose existing clinical diagnosis was confirmed (72.6%; p = 0.003).
DISCUSSION
In this study, we deployed a comprehensive molecular assay combining both rare, high-impact genetic variation and common susceptibility markers in the form of a PRS, LPA polymorphisms, and pharmacogenetic alleles. More than 30% of our cohort patients were found to have a positive finding with potential clinical impact, even after excluding pharmacogenetic findings present in nearly half of all patients. Physicians also found these results to benefit patient health and improve clinical care with 84% reporting medical management changes in those testing positive.
The study’s high overall yield is unheralded but reflects the manifest applicability of genetic testing to CVD in adults. In fact, cardiology has led the way in establishing the role of molecular testing in diagnosis and management, with numerous societies having published guidelines recommending genetic testing for patients with dyslipidemias, arrhythmias, cardiomyopathies, and thoracic aortic disease [17,18,19,20]. Nevertheless, uptake has been slow, due in part to lack of comprehensive testing coupled with challenges integrating it into the clinical workflow, both of which we addressed with HeartCare. Thus, our overall yield of 32% is an accurate reflection of heritable CVD among adults and represents an opportunity to individualize patient care.
Another major barrier to the adoption of genetic testing into clinical care is payer reimbursement policies that frequently consider such testing investigational and not accepted practice [21]. Although not attempted here, third party insurance reimbursement will be important for wider deployment of the HeartCare screen. To establish validity and utility of a diagnostic test, payers require evidence of an effect on medical outcomes and an equivalent benefit as established alternatives [22]. By tracking physician response in this study, we showed that incorporating both rare and common genetic variation affecting cardiovascular health does improve overall clinical management of the patient beyond standard of care. Additional economic analysis will be important in further directing reimbursement policy of payers by demonstrating cost-effectiveness of such testing.
The overall yield of 9% for identification of Mendelian disorders when testing CVD-related genes is comparable to results from studies performing comprehensive genetic testing for specific heritable cancers and chronic kidney disease (CKD). After heart disease, cancer is the second leading cause of death in the United States [1]. Recent studies involving individuals with breast and colorectal cancer have shown that 9.4% and 15.5% of cases, respectively, had a pathogenic or likely pathogenic variant in a high-penetrance cancer susceptibility gene [23, 24]. Similarly, CKD is also a major cause of morbidity and mortality and affects more than 10% of the population [25], with one study reporting a diagnostic yield of 9.3% in CKD patients undergoing exome sequencing (ES). In contrast, some studies have shown a yield as high as 30–40% for patients meeting strict criteria for diagnoses like hypertrophic cardiomyopathy [26]. Our study, on the other hand, was open to all cardiology clinic patients presenting with a variety of conditions and did not require any particular indications for inclusion. Thus, it offers a “real world” view of the yield expected in a typical cardiology care population.
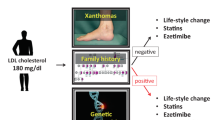
Half of the molecularly diagnosed Mendelian conditions in this study were lipid-related, the majority being familial hypercholesterolemia (FH). FH is highly treatable, with high-dose statin therapy and PCSK9 inhibitors leading to significant reductions in cardiac-related mortality [27, 28], and yet 9 in 10 cases go undiagnosed [29]. Many insurance payers require genetic testing results to cover PKSK9 inhibitors for FH patients but will not pay for the genetic tests [30]. We identified 25 patients in our cohort with molecularly confirmed FH, of which only a third were already on a PCSK9 inhibitor, indicating a significant potential opportunity for further LDL-C reducing therapy. We also identified several individuals with pathogenic variants in LPL, a gene associated with severe hypertriglyceridemia and pancreatitis. In addition to lifestyle and diet changes, these individuals are candidates for aggressive triglyceride-reducing therapy with a combination of fibrates, niacin, and/or fish oil with newer therapies under development using antisense and small interfering RNA (siRNA) approaches, which are effective in individuals with loss-of-function variants in LPL [31].
In 2018 the Heart Failure Society of America (HFSA) and ACMG published guidelines recommending genetic testing for patients with cardiomyopathies to assist in patient management [17]. More than 30% of our positive cases were cardiomyopathy-related and the diagnostic yield was highest for this indication (Table 1). One informative case included a 24-year-old female with a history of nonischemic postpartum cardiomyopathy. As part of her cardiology workup, the HeartCare panel identified a pathogenic multiexon deletion in BAG3, a gene associated with autosomal dominant dilated cardiomyopathy (DCM) [32]. Other loss-of-function variants in BAG3 have been reported in multigenerational families with DCM with some individuals requiring cardiac transplantation [32, 33]. This patient was subsequently referred to adult genetics specialists for additional genetic counseling and discussion of cascade testing for her siblings and children. With this molecular diagnosis and worsening cardiac function requiring a left ventricular assist device, she is currently undergoing evaluation for a heart transplant. Two African American males in the cohort with nonischemic cardiomyopathy were found to carry the TTR c.424G>A (p.Val142Ile) variant associated with hereditary transthyretin (hATTR) amyloidosis. This variant, present in up to 3.5% of the African American population, is an underrecognized cause of heart failure in this population, particularly those over age 60 [34]. This genetic finding is important as it may help predict the overall disease course, and it allows for the use of two genetic therapies, inotersen and patisiran, to ameliorate amyloid-associated polyneuropathy [35].
Genetic testing is also important in risk stratification and management of individuals with aortic aneurysm disease [20]. One such case involved a 33-year-old female originally thought to have hypermobile Ehlers–Danlos syndrome due to long standing joint hypermobility and dislocations. Baseline echocardiography revealed mild dilation of the aortic root at the sinus of Valsalva (4.0 ×3.6 cm), and on physical exam she had a pectus carinatum defect. Family history was significant for scoliosis in her mother and a maternal grandfather with a descending aortic aneurysm at age 65 years. The HeartCare panel identified a pathogenic variant in the TGFB2 gene, c.979C>T (p.Arg327Trp). Defects in TGFB2 are the cause of Loeys–Dietz syndrome 4 (LDS4), an autosomal dominant connective tissue disorder characterized by early-onset aneurysms and dissections of the aorta [36]. Though LDS has phenotypic overlap with Marfan syndrome, the vascular disease tends to be more aggressive in LDS compared to Marfan with aortic dissection occurring at smaller diameters and anywhere along the aorta. Consequently, surgical guidelines recommend prophylactic root replacement at smaller diameters compared to Marfan [36]. This patient is being followed closely by cardiothoracic surgery and was referred to adult genetics specialists for evaluation of other systemic features of LDS and additional discussion, including family cascade testing and the increased risk of dissection with pregnancy.
The LPA gene that encodes the apolipoprotein(a) component of the Lp(a) lipoprotein particle has received significant medical interest recently due to its association with atherosclerotic CVD. Increased levels of circulating Lp(a) lipoprotein are associated with an elevated risk of coronary disease and stroke [6, 37]. In particular, two common LPA polymorphisms (rs3798220 and rs10455872) are strongly associated with an increased level of Lp(a) lipoprotein and increase the risk of CAD by 42–57% in individuals of European ancestry [37]. The variable Kringle-IV (KIV-2) repeats within the LPA gene are also thought to contribute to circulating Lp(a) levels [38]. Current management for elevated Lp(a) includes lifestyle modifications as well as statin therapy. Clinical trials of an apo(a) antisense oligonucleotide (ASO) and siRNAs that reduce Lp(a) up to 80% offer hope for future reduction of cardiovascular events in those at risk [39]. Despite the well-established association with CVD and potentials for therapy, Lp(a) levels are rarely checked alongside standard lipid profiles and testing for the known risk alleles is not routinely done. Unsurprisingly, of the 144 patients with a LPA risk allele, only 20% had Lp(a) levels determined via a biochemical test before HeartCare. With this result, however, their Lp(a) levels are being followed and managed appropriately. Our study also demonstrates the need for additional studies of LPA genetics in diverse ethnicities since Lp(a) levels did not correlate well with the two alleles tested in individuals of non-European ancestry. Research into elucidating the number and role of LPA Kringle repeats in atherosclerotic disease in this cohort is ongoing.
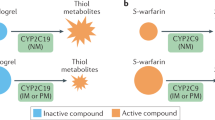
Recent recommendations focus on the utility of genetic testing for high-penetrance, monogenic conditions after careful phenotyping and collection of pedigree information [40]. However, within the precision medicine paradigm, pharmacogenetic results and PRS may offer an additional means to improve individual-level risk assessment and to better predict treatment response in both primary and secondary prevention for a broader population. We elected to test for pharmacogenetic variants associated with simvastatin and warfarin given the common use of these classes of medications in adults. Nearly 30% of all adults and 50% of those with atherosclerotic CVD in the United States were prescribed a statin medication as of 2013 [41]. And although newer anticoagulants that do not require monitoring or dose adjustment are gaining popularity, in 2017 more than 15 million prescriptions for warfarin were written in the United States according to the Medical Expenditure Panel Survey (MEPS), due in part to its effectiveness, low cost, and ability to quickly reverse. In our cohort, nearly half of patients had a positive pharmacogenetic finding related to these two medications. Among those currently prescribed either warfarin or simvastatin, 30% and 9% carried a genetic variant associated with altered metabolism, respectively. These individuals are being monitored for related complications (e.g., bleeding, myopathy) and family members have been encouraged to consider testing themselves. Additional pharmacogenetic markers affecting other cardiac-related medications (e.g., clopidogrel, beta blockers) may be another area of opportunity for this cohort in future studies.
PRS for CAD like the one used here have been proposed as a method to improve cardiovascular risk prediction by aggregating weighted associations between genetic variants and disease outcomes derived from genome-wide association studies (GWAS). Using a 50 SNP PRS that was state of the art at study onset, more than 9% of our cohort was found to be in the highest CVD risk group (top 5%). The clinical utility of CAD PRS has been investigated in a number of important areas, with a primary focus on applications to primary prevention in individuals without pre-existing CAD [42]. For example, studies have demonstrated the ability of a CAD for PRS to effectively stratify individuals according to their risk for CAD and to identify individuals with high risk for CAD who did not have high cholesterol levels or other traditional risk factors [43]. Additional work has provided evidence that genetic risk for CAD can be mitigated by statin therapy and a healthy lifestyle [43, 44]. PRS construction has now expanded to scores comprised of millions of SNPs with improved risk classification and the ability to identify individuals with risk for CAD equivalent to a monogenic trait [43]. Ultimately, further research is still needed to understand the proper role of PRS within a clinical context. Furthermore, research should be directed at the development of PRS that perform well in multiethnic populations and on defining target populations for genetic risk stratification.
In summary, we present here the compelling results of our HeartCare study, a comprehensive test targeting genes that influence risk for CVD. Impressively, the overall positive rate was 32%, reflecting the enriched patient population seen in ambulatory cardiology clinics where patients commonly present with arrhythmias, heart failure, dyslipidemias, and aortic aneurysms. Critical management changes resulted from these findings, from new pharmacologic therapies to surgical considerations. Our 9% rate for Mendelian cardiovascular conditions was consistent with other studies evaluating genetic causes of common diseases such as cancer and kidney disease. The primary findings have implications beyond the tested patients as close family members have up to 50% chance of sharing the same genetic risk, providing practitioners the opportunity to implement cascade testing of at-risk family members and offer preventive interventions. Clinicians were supportive of this test overall, finding it benefited patient health and improved clinical care. Data will continue to be reanalyzed as new interpretations become available, and we will leverage the research consent to perform genome sequencing to assess the added clinical utility of a broader assay. There is also opportunity to utilize phenotype data in the EMR to identify additional patients who may benefit from genetic testing, thus speeding up diagnosis and improving care even further [45]. In conclusion, our results demonstrate that comprehensive molecular testing can be routinely used in the ambulatory setting and that CVD is a prime model to demonstrate precision medicine’s utility.
Data availability
All reported variants have been submitted to ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/) under Human Genome Sequencing Center Clinical Lab, Baylor College of Medicine. Please contact the corresponding author for original data access requests.
References
Heron M, National Center for Health Statistics. Deaths: leading causes for 2018. 2021. https://doi.org/10.15620/cdc:104186
Miller DT, Lee K, Chung WK, Gordon AS, Herman GE, Klein TE, et al. ACMG SF v3.0 list for reporting of secondary findings in clinical exome and genome sequencing: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2021. Online ahead of print. https://doi.org/10.1038/s41436-021-01172-3
Collins FS, Varmus H. A new initiative on precision medicine. N Engl J Med. 2015;372:793–795.
Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015;43:D789–D798.
Khera AV, Emdin CA, Drake I, Natarajan P, Bick AG, Cook NR, et al. Genetic risk, adherence to a healthy lifestyle, and coronary disease. N Engl J Med. 2016;375:2349–2358.
Clarke R, Peden JF, Hopewell JC, Kyriakou T, Goel A, Heath SC, et al. Genetic variants associated with Lp(a) lipoprotein level and coronary disease. N Engl J Med. 2009;361:2518–2528.
Ramsey LB, Johnson SG, Caudle KE, Haidar CE, Voora D, Wilke RA, et al. The clinical pharmacogenetics implementation consortium guideline for SLCO1B1 and simvastatin-induced myopathy: 2014 update. Clin Pharmacol Ther. 2014;96:423–428.
Johnson JA, Caudle KE, Gong L, Whirl-Carrillo M, Stein CM, Scott SA, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) guideline for pharmacogenetics-guided warfarin dosing: 2017 update. Clin Pharmacol Ther. 2017;102:397–404.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26:589–595.
Reid JG, Carroll A, Veeraraghavan N, Dahdouli M, Sundquist A, English A, et al. Launching genomics into the cloud: deployment of Mercury, a next generation sequence analysis pipeline. BMC Bioinformatics. 2014;15:30.
Chiang T, Liu X, Wu T-J, Hu J, Sedlazeck FJ, White S, et al. Atlas-CNV: a validated approach to call single-exon CNVs in the eMERGESeq gene panel. Genet Med. 2019;21:2135–2144.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Deignan JL, Chung WK, Kearney HM, Monaghan KG, Rehder CW, Chao EC, et al. Points to consider in the reevaluation and reanalysis of genomic test results: a statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2019;21:1267–1270.
Lloyd-Jones DM, Hong Y, Labarthe D, Mozaffarian D, Appel LJ, Van Horn L, et al. Defining and setting national goals for cardiovascular health promotion and disease reduction: the American Heart Association’s strategic Impact Goal through 2020 and beyond. Circulation. 2010;121:586–613.
Murugan M, Babb LJ, Overby Taylor C, Rasmussen LV, Freimuth RR, Venner E, et al. Genomic considerations for FHIR®; eMERGE implementation lessons. J Biomed Inform. 2021;118:103795.
Venner E, Yi V, Murdock D, Kalla SE, Wu TJ, Sabo A, et al. Neptune: an environment for the delivery of genomic medicine. Genet Med. 2021. https://doi.org/10.1038/s41436-021-01230-w.
Hershberger RE, Givertz MM, Ho CY, Judge DP, Kantor PF, McBride KL, et al. Genetic evaluation of cardiomyopathy: a clinical practice resource of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2018;20:899–909.
Ackerman MJ, Priori SG, Willems S, Berul C, Brugada R, Calkins H, et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies: this document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Europace. 2011;13:1077–1109.
Khoury MJ, Bowen MS, Burke W, Coates RJ, Dowling NF, Evans JP, et al. Current Priorities for Public Health Practice in Addressing the Role of Human Genomics in Improving Population Health. Am J Prev Med. 2011;40:486–493.
Verhagen JMA, Kempers M, Cozijnsen L, Bouma BJ, Duijnhouwer AL, Post JG, et al. Expert consensus recommendations on the cardiogenetic care for patients with thoracic aortic disease and their first-degree relatives. Int J Cardiol. 2018;258:243–248.
Deverka PA, Kaufman D, McGuire AL. Overcoming the reimbursement barriers for clinical sequencing. JAMA. 2014;312:1857–1858.
Roundtable on Translating Genomic-Based Research for Health; Board on Health Sciences Policy; Institute of Medicine. Assessing Genomic Sequencing Information for Health Care Decision Making: Workshop Summary. Washington (DC): National Academies Press (US); 2014. 5, How Insurers Decide Whether to Pay for Testing. Available from: https://www.ncbi.nlm.nih.gov/books/NBK241339/.
Beitsch PD, Whitworth PW, Hughes K, Patel R, Rosen B, Compagnoni G, et al. Underdiagnosis of hereditary breast cancer: are genetic testing guidelines a tool or an obstacle? J Clin Oncol. 2019;37:453–460.
Uson PLS Jr, Riegert-Johnson D, Boardman L, et al. Germline cancer susceptibility gene testing in unselected patients with colorectal adenocarcinoma: a multicenter prospective study. Clin Gastroenterol Hepatol. 2021 Apr 20; S1542-3565(21)00447-X. doi: 10.1016/j.cgh.2021.04.013. Online ahead of print. https://doi.org/10.1016/j.cgh.2021.04.013
Groopman EE, Marasa M, Cameron-Christie S, Petrovski S, Aggarwal VS, Milo-Rasouly H, et al. Diagnostic utility of exome sequencing for kidney disease. N Engl J Med. 2019;380:142–151.
Hathaway J, Heliö K, Saarinen I, Tallila J, Seppälä EH, Tuupanen S, et al. Diagnostic yield of genetic testing in a heterogeneous cohort of 1376 HCM patients. BMC Cardiovasc Disord. 2021;21:126.
Humphries S, Cooper JA, Seed M, Capps N, Durrington PN, Jones B, et al. Coronary heart disease mortality in treated familial hypercholesterolaemia: update of the UK Simon Broome FH register. Atherosclerosis. 2018;275:e176 https://doi.org/10.1016/j.atherosclerosis.2018.06.535
Sabatine MS, Giugliano RP, Keech AC, Honarpour N, Wiviott SD, Murphy SA, et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N Engl J Med. 2017;376:1713–1722.
Goldberg AC, Hopkins PN, Toth PP, Ballantyne CM, Rader DJ, Robinson JG, et al. Familial hypercholesterolemia: screening, diagnosis and management of pediatric and adult patients: clinical guidance from the National Lipid Association Expert Panel on Familial Hypercholesterolemia. J Clin Lipidol. 2011;5:133–140.
Doshi JA, Puckett JT, Parmacek MS, Rader DJ. Prior authorization requirements for proprotein convertase subtilisin/kexin type 9 inhibitors across US private and public payers. Circ Cardiovasc Qual Outcomes. 2018;11:e003939.
Hussain A, Ballantyne CM. New approaches for the prevention and treatment of cardiovascular disease: focus on lipoproteins and inflammation. Annu Rev Med. 2021;72:431–446.
Norton N, Li D, Rieder MJ, Siegfried JD, Rampersaud E, Züchner S, et al. Genome-wide studies of copy number variation and exome sequencing identify rare variants in BAG3 as a cause of dilated cardiomyopathy. Am J Hum Genet. 2011;88:273–282.
Domínguez F, Cuenca S, Bilińska Z, Toro R, Villard E, Barriales-Villa R, et al. Dilated cardiomyopathy due to BLC2-associated athanogene 3 (BAG3) mutations. J Am Coll Cardiol. 2018;72:2471–2481.
Arvanitis M, Koch CM, Chan GG, Torres-Arancivia C, LaValley MP, Jacobson DR, et al. Identification of transthyretin cardiac amyloidosis using serum retinol-binding protein 4 and a clinical prediction model. JAMA Cardiol. 2017;2:305–313.
Gertz MA, Mauermann ML, Grogan M, Coelho T. Advances in the treatment of hereditary transthyretin amyloidosis: a review. Brain Behav. 2019;9:e01371.
MacCarrick G, Black JH, Bowdin S, El-Hamamsy I, Frischmeyer-Guerrerio PA, Guerrerio AL, et al. Loeys–Dietz syndrome: a primer for diagnosis and management. Genet Med. 2014;16:576–587.
Li Y, Luke MM, Shiffman D, Devlin JJ. Genetic variants in the apolipoprotein(a) gene and coronary heart disease. Circ Cardiovasc Genet. 2011;4:565–573.
Lanktree MB, Rajakumar C, Brunt JH, Koschinsky ML, Connelly PW, Hegele RA. Determination of lipoprotein(a) kringle repeat number from genomic DNA: copy number variation genotyping using qPCR. J Lipid Res. 2009;50:768–772.
Vogt A. Lipoprotein(a)-antisense therapy. Clin Res Cardiol Suppl. 2019;14:51–56.
Musunuru K, Hershberger RE, Day SM, Klinedinst NJ, Landstrom AP, Parikh VN, et al. Genetic testing for inherited cardiovascular diseases: a scientific statement from the American Heart Association. Circ Genom Precis Med. 2020;13:e000067.
Salami JA, Warraich H, Valero-Elizondo J, Spatz ES, Desai NR, Rana JS, et al. National trends in statin use and expenditures in the US adult population from 2002 to 2013: insights from the Medical Expenditure Panel Survey. JAMA Cardiol. 2017;2:56–65.
Lambert SA, Abraham G, Inouye M. Towards clinical utility of polygenic risk scores. Hum Mol Genet. 2019;28:R133–R142.
Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet. 2018;50:1219–1224.
Natarajan P, Young R, Stitziel NO, Padmanabhan S, Baber U, Mehran R, et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation. 2017;135:2091–2101.
Morley TJ, Han L, Castro VM, et al. Phenotypic signatures in clinical data enable systematic identification of patients for genetic testing. Nat Med. 2021;27:1097–1104.
Acknowledgements
The authors gratefully acknowledge the individuals who provided biological samples and data as part of the BCM HeartCare cohort. All DNA samples, phenotypic information, and all sequencing data were obtained with generous support from donors to the St. Luke’s Foundation. Dr. de Vries was supported by National Heart, Lung, and Blood Institute grant R01HL146860 and American Heart Association grant 18CDA34110116. We thank the HGSC Clinical Sequencing Team (full list in Supplemental material) and BCM Epic team for their work.
Author information
Authors and Affiliations
Contributions
Conceptualization: D.R.M, E.V., D.M.M, G.A.M., A.B., C.I.A., A.L.M., E.B., X.W., C.M.B., R.A.G. Data curation: E.V., M.M., J.H., M.C., M.G., V.Y. Formal analysis: D.R.M, E.V., D.M.M., T.D.H., V.C., P.S.d.V., A.S., S.L., Q.M., J.H., H.S., S.P. Funding acquisition: G.A.M., R.A.G. Investigation: D.R.M, E.V., D.M.M, X.J., A.H., A.M.A., A.C., M.G.C., R.M., E.B., X.W., C.M.B., R.A.G. Methodology: E.V., D.M.M; Project administration: G.A.M., M.C., C.L.K., M.G., V.K., Z.H., A.B., P.R.M. Supervision: C.M.B., R.A.G. Writing—original draft: D.R.M. Writing—review & editing: D.R.M., E.V., D.M.M., G.A.M., M.M., T.D.H., V.C., P.V., X.J., A.H., A.M.A., A.S., S.L., Q.M., J.H., M.C., V.Y., C.L.K., M.G., V.K., C.H., D.L.R., S.P., H.S., Z.A.H., A.B., P.R.M., L.L., A.B., C.I.A., A.C., M.G.C., R.M., A.L.M., E.B., X.W., C.M.B., R.A.G.
Corresponding author
Ethics declarations
Ethics Declaration
The BCM Institutional Review Boards approved the study (protocol number: H-43884). Written informed consent was obtained from all study participants.
Competing interests
C.M.B. has received grant/research support through his institution from Abbott Diagnostic, Akcea, Amgen, Esperion, Novartis, Regeneron, and Roche Diagnostic and consulting fees from Abbott Diagnostics, Akcea, Althera, Amarin, Amgen, Arrowhead, Astra Zeneca, Corvidia, Denka Seiken, Esperion, Gilead, Janssen, Matinas BioPharma Inc, New Amsterdam, Novartis, Novo Nordisk, Pfizer, Regeneron, Roche Diagnostic, and Sanofi-Synthelabo. D.R.M. has received consulting fees from Illumina. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Murdock, D.R., Venner, E., Muzny, D.M. et al. Genetic testing in ambulatory cardiology clinics reveals high rate of findings with clinical management implications. Genet Med 23, 2404–2414 (2021). https://doi.org/10.1038/s41436-021-01294-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01294-8
This article is cited by
-
The frequency of pathogenic variation in the All of Us cohort reveals ancestry-driven disparities
Communications Biology (2024)
-
Genetic Testing for Familial Hypercholesterolemia in Clinical Practice
Current Atherosclerosis Reports (2023)