Abstract
Purpose
We describe a novel neurobehavioral phenotype of autism spectrum disorder (ASD), intellectual disability, and/or attention-deficit/hyperactivity disorder (ADHD) associated with de novo or inherited deleterious variants in members of the RFX family of genes. RFX genes are evolutionarily conserved transcription factors that act as master regulators of central nervous system development and ciliogenesis.
Methods
We assembled a cohort of 38 individuals (from 33 unrelated families) with de novo variants in RFX3, RFX4, and RFX7. We describe their common clinical phenotypes and present bioinformatic analyses of expression patterns and downstream targets of these genes as they relate to other neurodevelopmental risk genes.
Results
These individuals share neurobehavioral features including ASD, intellectual disability, and/or ADHD; other frequent features include hypersensitivity to sensory stimuli and sleep problems. RFX3, RFX4, and RFX7 are strongly expressed in developing and adult human brain, and X-box binding motifs as well as RFX ChIP-seq peaks are enriched in the cis-regulatory regions of known ASD risk genes.
Conclusion
These results establish a likely role of deleterious variation in RFX3, RFX4, and RFX7 in cases of monogenic intellectual disability, ADHD and ASD, and position these genes as potentially critical transcriptional regulators of neurobiological pathways associated with neurodevelopmental disease pathogenesis.
Similar content being viewed by others
INTRODUCTION
Autism spectrum disorder (ASD), marked by deficits in social communication and the presence of restricted interests and repetitive behavior, is highly heritable and genetically heterogeneous, with de novo loss-of-function variants as known contributors to ASD risk.1 ASD is often comorbid with other neurodevelopmental diagnoses, including attention-deficit/hyperactivity disorder (ADHD). Emerging evidence also points to a role of de novo loss-of-function variants in ADHD.2
RFX3 is a member of the regulatory factor X (RFX) gene family that encodes transcription factors with a highly conserved DNA binding domain. RFX3 is expressed in several tissues including developing and adult brain, and other RFX family members (RFX1, 4, 5, and 7) are also highly expressed in brain tissue, with expression patterns of RFX1, 3, 4, and 7 clustering tightly.3
We report a series of 38 individuals from 33 families with deleterious, mostly de novo variants in three brain-expressed members of the RFX family: RFX3, RFX4, or RFX7. RFX3 was among 102 genes recently identified as statistically enriched for de novo variants in a large-scale analysis of trio exome data from individuals with ASD,4 but to date RFX4 and RFX7 have not been previously associated with human disease. Analysis of case clinical data reveals common features including intellectual disability (ID), ASD, and/or ADHD, delineating a novel neurobehavioral phenotype associated with RFX haploinsufficiency.
MATERIALS AND METHODS
Case ascertainment and data collection
We obtained phenotypic data from 15 unrelated individuals with loss-of-function variants in RFX3, 4 unrelated individuals with loss-of-function variants in RFX4, and 14 unrelated individuals with loss-of-function variants in RFX7. Individual case summaries for all individuals are provided (Supplemental Data). Variants arose de novo with the exception of four related individuals from the same nuclear family with the same heterozygous loss-of-function variant in RFX3, and three other related cases in RFX4 (homozygous for an inherited missense variant). Pedigree information and contributed photographs are shown in Fig. 1. Diagnoses of ASD were reported in the medical record, but not uniformly evaluated by standardized measures such as the Diagnostic and Statistical Manual, Fourth or Fifth Edition (DSM-IV and 5), Autism Diagnostic Observation Schedule (ADOS), or Autism Diagnostic Interview, Revised (ADI-R). Similarly, ID and ADHD diagnoses were accepted per clinician report and not always accompanied by standardized cognitive or behavioral testing measures.
Pedigrees and clinical photographs of individuals with variants in RFX3, RFX4, and RFX7. (a) RFX3, RFX4, and RFX7 case pedigrees. All pedigrees show de novo origin of variants except for RFX3-8a–d: a 33 year-old affected mother carrying the variant p.(Leu496Alafs*7) with transmission to three children, and pedigree RFX4-3a–c: three affected children homozygous for p.(Thr247Met). (b) Individuals with RFX3 variants. (c) Individuals with RFX4 variants. (d) Individuals with RFX7 variants.
Exome sequencing
Individuals included underwent exome sequencing on a clinical or research basis. Seven of the individuals were sequenced through GeneDx using genomic DNA from the proband or proband plus parents, captured using either the Clinical Research Exome kit (Agilent Technologies, Santa Clara, CA) or the IDT xGen Exome Research Panel v1.0, and sequenced on an Illumina system with 100 bp or greater paired-end reads. Reads were aligned to human genome build GRCh37/UCSC hg19, and variants were analyzed and interpreted as previously described using variant classification criteria publicly available on the GeneDx ClinVar submission page (see Web Resources). Two cases of RFX7 were sequenced through Ambry Genetics whose gene and variant classification process are available on the Ambry Genetics web page. The remainder of the individual’s exome sequencing was performed through the clinicians’ institutions or an external laboratory or research program (see Acknowledgments).
Variant analyses
Variant genomic coordinates are reported in relation to the Human December 2013 (GRCh38/hg38) Assembly. The reference messenger RNA (mRNA) and protein sequences used are RFX3 NM_134428.2, NP_602304.1; RFX4 NM_213594.2, NP_998759.1; and RFX7 NM_022841.5, NP_073752.5. The variant databases gnomAD v2.1.1 and v3 were examined for the presence of each variant.5 Predictions of the functional effects for all variants were assessed using MutationTaster, SIFT, PolyPhen-2, PROVEAN, LRT, and MutationAssessor, and the total number of algorithms out of six with a deleterious prediction is referred to as the Nonsynonymous Damaging score (NsynD) as previously described.6
Cell transfection and culture
Human RFX3 (NM_134428.2; Human RFX3 complementary DNA [cDNA]) was cloned into V5-tagged mammalian expression vectors using the Gateway cloning system (Thermo Fisher Scientific). Point mutations were introduced with the QuikChange Lighting Site-Directed Mutagenesis kit (Agilent Technologies) to incorporate variants from affected individuals. To quantify the expression level of exogenous RFX3, equal amounts of tagged-RFX3 expression vectors were transfected into HeLa cells using Lipofectamine 3000 (Thermo Fisher Scientific). The transfected cells were cultured for 48 hours before harvesting. Cell extracts were analyzed by immunoblotting, using antibodies raised against RFX3 (HPA035689, Sigma-Aldrich), V5 (R960-25, Thermo Fisher Scientific), or beta actin (ab6276, Abcam). Blots were scanned on a Li-Cor Odyssey imager (Li-Cor). Signal intensities were quantified using Image Studio Lite (Li-Cor). Each immunoblot analysis was replicated six times. One-way analysis of variance (ANOVA) by repeated measures was employed. Multiple comparison correction was performed by using Dunnett statistical testing.
KEGG pathway and ASD gene set overrepresentation analysis
ChIP-seq and eCLIP-seq narrowPeak bed files for RFX family members, CREBBP, EP300, FMR1, FXR1, and FXR2 were obtained from the ENCODE portal,7 and additional ChIP-seq data for RFX3_K562 were obtained from RegulomeDB8 (Table S6). Functional binding genes (1 kb upstream/downstream of transcriptional start site [TSS]) were annotated using ChIPseeker.9 ASD risk gene lists included 102 TADA genes from Satterstrom et al. and 253 ASD/ID genes from Coe et al. (Table S7).4,10] Differentially expressed genes (DEGs) in ASD brains were extracted from Velmeshev et al. excluding endothelial DEGs that could originate from vascular cells in the brain (Table S7).11 The SYSCILIA Gold Standard (SCGSv.1) was used as a gold standard of known ciliary genes in human.12 Customized Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was performed using clusterProfiler13 to determine the enrichment for KEGG pathways, ciliary genes, ASD risk gene sets, and ASD DEGs. Multiple testing correction was performed using Benjamini–Hochberg correction (Table S8). Annotations, statistical analyses, and plots were implemented in R.
Motif analysis
For motif occurrence analysis, FIMO was used to scan promoter sequences for individual occurrences of RFX motifs.14 For all analyses, motif models were obtained from the JASPAR 2020 database.15 Motifs searched for included RFX3 (MA0798.1), RFX4 (MA0799.1), and RFX7 (MA1554.1). Promoter sequences were defined as -1,000 base pairs and +500 base pairs relative to the transcription start site. Motif occurrences were classified as significant based on a reporting threshold of p value <0.00001 and q value (Benjamini) <0.10. For motif enrichment analysis, we used the HOMER findMotifs.pl and findMotifsGenome.pl scripts. Motifs were classified as enriched based on fold-enrichment >1.5 over randomly selected background sequences with matched GC% content, and q value (Benjamini) <0.01. Enhancer sequences associated with genes of interest were obtained from Enhancer Atlas 2.0.16 ASD risk gene lists were obtained as noted previously to include 102 TADA genes and 124 ASD/ID genes reaching exome-wide significance.4,10
RESULTS
Case series of individuals with de novo or inherited RFX3 variants
We identified and obtained clinical information from 18 individuals bearing loss-of-function variants in RFX3 via GeneMatcher.17 Genotypic information is provided in Table 1, clinical phenotypes are summarized in Table 2 and S1, and predicted variant impacts are summarized in Supplemental Table S2. A total of 15 distinct variants were identified: 2 frameshift variants, 2 canonical splice donor variants, 8 missense variants, 1 in-frame deletion, 1 42-kb deletion removing the last two exons of RFX3, and 1 227-kb deletion involving only RFX3. In one family, an affected parent transmitted a frameshift variant to three affected children; all other variants were de novo and novel (Fig. 1a).
There were 13 males and 5 females, with no sex-based differences in severity of phenotype. All individuals had neurodevelopmental delays, with formally recorded clinical diagnoses of ASD (72%) and ID of varying severity (borderline to moderate) or global developmental delay in young children (78%) and ADHD (56%) (Table 2 and Table S1). Many showed a distinct behavioral pattern marked by easy excitability/overstimulation, hypersensitivity to sensory (particularly auditory) stimuli, anxiety, emotional dysregulation and/or aggression (13/15 [87%] with specific behavioral information provided). Three individuals were reported to have seizures (17%). Some individuals had sleep difficulties (44%) including limited total duration of sleep, frequent awakenings, or early morning awakenings. Subtle nonspecific and nonrecurrent dysmorphisms were commonly reported (61%), including broad nasal bridge, high, arched palate, and hand and foot abnormalities (tapered fingers, widely spaced toes), but no consistent recognizable features were shared by all individuals (Fig. 1b). Both macrocephaly (six individuals) and microcephaly (two individuals) were reported (8/11 individuals [73%] with a head circumference measurement or percentile provided). Magnetic resonance imaging (MRI) of the brain was available for eight individuals, with reports of nonspecific findings in four, including white matter changes, uncal asymmetry, partially empty sella, or prominent ventricles. One individual had mild thinning of the corpus callosum (Table 2, individual RFX3-10). Five of seven individuals (71%) who were past the onset of puberty (ages 12–30 years) had reports of behavioral and/or cognitive worsening at the time of puberty/adolescence. Three had increased aggression specifically noted. Three were described as havior psychotic s/or psychotic symptoms, two were described as having hallucinations (one requiring psychiatric hospitalization) and another was described as having conversations with imaginary friends. Three were reported to have had decline in cognition, one in adolescence and another around 28 years of age.
Variants in additional RFX family genes are associated with similar neurodevelopmental phenotypes
Additional individuals were ascertained who harbored loss-of-function variants in other closely related genes of the RFX family. Fourteen individuals bearing de novo loss-of-function variants in RFX7 were identified (Tables 1 and 2), including four frameshift variants, five stop-gain variants, one in-frame deletion, and two missense variants (Table 1, Table S2). Slightly more males were identified than females (eight males, six females) without differences in phenotype based on sex. All individuals had language delay, and most had ID/global developmental delay (93%) (Table 2, Supplemental Table 1). While formal diagnoses of ASD (36%) and/or ADHD (29%) were less consistent, autistic features and/or significant behavioral challenges akin to those seen in RFX3 individuals were reported in the majority of cases, including excitability/overstimulation, sensitivity to sensory (particularly auditory) stimuli, a high pain threshold, emotional dysregulation, aggression, and anxiety (8/8, 100% of those with specific behavioral information provided). Abnormal head size (five individuals with microcephaly and three with macrocephaly) was noted in 7/11 (64%) that provided head circumference measurements. In 5/11 patients (45%) who had neuroimaging, MRI abnormalities were observed (Dandy–Walker malformation, cerebellar tonsillar herniation, an abnormality of the basal ganglia, and a fourth case with limited information but an “abnormal brain MRI” noted). Subtle clinical dysmorphisms were reported in 86% including abnormalities of the hands and feet such as widely spaced toes, syndactyly, or long tapered fingers (50%) (Table 2). Again, no consistent dysmorphisms were evident across individuals (Fig. 1d).
Six individuals with probable loss-of-function RFX4 variants were also identified (Tables 1 and 2). Three were individuals who harbored de novo RFX4 variants, including an in-frame deletion (RFX4 p.[Tyr639_Ser643del]), and two predicted damaging missense variants (RFX4 p.[Arg79Ser] and p.[Thr362Ala]) (Table 1, Supplemental Table S2). We also report a pedigree in which three additional related individuals (siblings) were homozygous for a missense variant in RFX4 (p.Thr247Met) altering a well-conserved threonine residue. Parents of these siblings were first cousins, each heterozygous for the variant and without any known neurobehavioral phenotype, and a heterozygous sibling was similarly reported as neurotypical; this pedigree therefore raises the possibility that the RFX4 phenotype may be associated with both monoallelic and biallelic inheritance as has been described for several other genetic conditions.18 Of these six individuals, three were female and three were male. All were noted to have ID or global developmental delay (100%) and most had documented ASD (83%). Four individuals were normocephalic, and one was microcephalic. Neuroimaging was performed in two and demonstrated asymmetric volume loss in one individual and absent pituitary gland in another individual with hypopituitarism. The latter individual also presented with cleft lip and palate. Seizures were described in two individuals (33%). No consistent dysmorphisms were evident (Fig. 1c).
RFX3, RFX4, and RFX7 variant analyses
In total, 33 distinct variants in RFX family members (15 RFX3, 4 RFX4, and 14 RFX7) were identified (Table 1). Excluding related individuals, each case involved a novel variant (e.g., there were no recurrent variants). RFX3, RFX4, and RFX7 each exhibit intolerance to loss-of-function variation in human population databases (gnomAD, pLI scores = 1.00). All variants were absent from gnomAD except for RFX7 p.Pro964_Thr965del, which is detected at a very low frequency in gnomAD v2.1.1 (AF 0.00007677) leading us to formally classify it as a variant of uncertain significance (VUS) (see Supplemental RFX7 Case Descriptions Individual 14 for further details). The fact that the majority of variants identified are predicted to cause outright protein truncation or gene deletion (20/33) strongly supports a loss-of-function/haploinsufficiency model. Of the 13 missense variants, 11 were predicted to be damaging by at least four of six algorithms (NsynD score ≥4) and two missense variants were predicted to be damaging by at least two algorithms (RFX4 p.[Thr247Met] and p.[Thr362 Ala]) (Table S2). All missense variants affect highly conserved amino acids (PhastCons vertebrate, mammalian, and primate scores ranging from 0.99 to 1.00) (Table S2).
RFX transcription factors are defined by a conserved, specialized winged-helix type DNA binding domain (DBD) that recognizes the X-box motif. In addition to the DBD, RFX3 and RFX4 have three known domains that are associated with dimerization (DD).3 RFX4 and RFX7 variants did not exhibit clustering to specific functional domains, but all of the nontruncating (missense or in-frame deletion) variants identified in RFX3 were found to be located in the DBD or one of the dimerization domains (Fig. 2a, Table 1). We engineered five of the nontruncating variants—p.(Glu195del), p.(Leu241Trp), p.(Phe383Ser), p.(Leu443Ile), and p.(Asp611Tyr)—into a V5-RFX3 heterologous expression vector for protein stability analyses in HeLa cells (Figure S1). The majority of these variants resulted in significant decreases in detectable RFX3 levels, consistent with a destabilizing impact on protein expression. Two missense variants, p.(Leu241Trp) and p.(Leu443Ile) (residing in the DNA binding domain or the second extended protein dimerization domain, respectively), did not appear to impact protein stability, raising the possibility that they might disrupt more specific functional interactions of RFX3 to be investigated more thoroughly in the future.
(a) Mapping of selected RFX variants to domains. Whole-gene deletion and intronic variants are not illustrated. RFX3 (NP_602304.1), RFX4 (NP_998759.1), RFX7 (NP_073752.5). (b) Missense variant deleteriousness scores for the currently reported variants (current) and prior reported variants (prior) in RFX3, 4, and 7. The distribution of MPC scores for missense variants reported in this study is significantly different from that of prior reported missense variants, Kolmogorov–Smirnov (K-S) test p value <0.05 (p value = 0.015). CADD Combined Annotation Dependent Depletion, MPC Missense badness, PolyPhen-2, and Constraint, NsynD Nonsynonymous Damaging score.
We examined 35 additional reported variants in RFX3, RFX4, and RFX7 from prior studies of de novo or inherited variants in ASD and neuropsychiatric conditions (Table S3, Figure S2).4,19,20,21,22 Missense variants from the literature tended to be of milder predicted deleteriousness than those reported here (Fig. 2b). Sixteen were in RFX3, including five de novo variants (four protein truncating and one missense variant predicted damaging by all six algorithms, NsynD6, supportive of likely deleteriousness), seven inherited variants (four copy-number variants [CNVs], one frameshift variant (p.[Pro408fs]), and three missense variants (p.[Thr151Ala], p.[Ala101Thr], p.[Arg615His]; NsynD scores 3–6), and four CNVs (all microdeletions) were reported for which parental inheritance was not established (Table S3). Among previously reported RFX7 variants, one was a de novo frameshift and one was an inherited frameshift variant. There were also six reported inherited missense variants (6/6 with NsynD >4), and two de novo missense variants that are likely benign. Finally, there were nine previously reported RFX4 variants, only one of which was de novo (a missense variant lacking strong evidence of pathogenicity), and eight inherited missense variants of varying predicted deleteriousness (NsynD scores 3–6).
RFX expression is enriched in human brain
RFX3, RFX4, and RFX7 have been reported to have relatively high expression in human fetal cortex.23 To determine whether specific cell types are affected by RFX haploinsufficiency, we examined single-cell transcriptomes from developing and adult human cortex (Fig. 3a–f, Figure S3A, B).11,24 In developing human cortex, RFX3 and RFX7 exhibited the strongest brain expression, with RFX3 most highly expressed in maturing excitatory upper enriched neurons, RFX4 most highly expressed in outer radial glia, and RFX7 most highly expressed in interneurons from the medial ganglionic eminence (Fig. 3a–c). We also examined RFX expression patterns in the adult human cortex (Fig. 3d–f, Figure S3A, B).11,25 Again, RFX3 and RFX7 exhibited the highest expression. RFX3 was most highly expressed in glutamatergic layer 2/3 neurons, followed by astrocytes. RFX7 was expressed in both inhibitory and excitatory neurons. RFX4 expression was much lower overall, but highest in astrocytes (Fig. 3f). These expression profiles suggest that RFX deleterious variants may lead to our observed neurodevelopmental phenotypes by altering early developmental cell fates or by impacting the function of upper-layer cortical neurons, astrocytes, and interneurons.
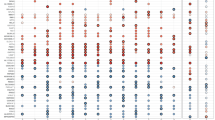
(a) Transcriptomic cell types in the prenatal human cortex identified by single-cell RNA-sequencing.24 (b) RFX3, 4, and 7 expression patterns in single cells of the prenatal human cortex. (c) Heatmap of RFX3, 4, and 7 expression levels among cell types in the prenatal human cortex. (d) Transcriptomic cell types in the postnatal human cortex identified by single-cell RNA-sequencing.11 (e) RFX3, 4, and 7 expression patterns in single cells of the postnatal human cortex. (f) Heatmap of RFX3, 4, and 7 expression levels among cell types in the postnatal human cortex. (g) The enrichment of KEGG pathways, ciliary genes, ASD risk gene sets, and ASD differentially expressed genes (DEGs) among RFX3 ChIP-seq binding targets. Pathways and ASD gene sets are ranked by their statistical significance (p.adjust values, Benjamini–Hochberg’s correction). Red arrows indicate ASD risk gene sets and ASD DEGs. X-axis shows the number of genes bound by RFX in their promoter regions. (h) Binding of RFX family transcription factors bind to X-box motif in promoter regions of ciliary and immunologic genes. Target gene lists obtained from Piasecki, Durand, Reith, Sugiaman-Trapman.3,38,39,40 Model of RFX gene dose-dependent regulation of genes. In tissues with higher expression of RFX genes, ASD genes are activated. Lower levels of RFX genes are sufficient to activate ciliary genes. ASD autism spectrum disorder, AST-FB fibrous astrocytes, AST-PP protoplasmic astrocytes, End endothelial, ExDp1 excitatory deep layer 1, ExDp2 excitatory deep layer 2, ExM-U maturing excitatory upper enriched, ExM maturing excitatory, ExN migrating excitatory, IN-PV parvalbumin interneurons, IN-SST somatostatin interneurons, IN-SV2C SVC2 expressing interneurons, IN-VIP VIP interneurons, InCGE interneuron CGE, InMGE interneuron MGE, IP intermediate progenitors, L2/3 layer 2/3 excitatory neurons, L4 layer 4 excitatory neurons, L5/6-CC layer 5/6 excitatory cortico-cortical projection neurons, L5/6 layer 5/6 excitatory neurons, Mic microglia, Neu-mat immature neurons, Neu-NRGN NRGN expressing neurons, OPC oligodendrocyte precursor cells, OPC oligodendrocyte precursor cells, oRG outer radial glia, Per pericyte, PgG2M cycling progenitors G2/M phase, PgS cycling progenitors S phase, vRG ventricular radial glia11.
RFX binding motifs are present in ASD risk gene cis-regulatory regions
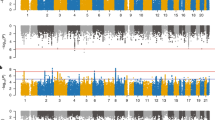
Dysregulated gene expression, especially in upper-layer cortical neurons, has been implicated in ASD pathogenesis.11,26 Given the expression of RFX3 in layer 2/3 neurons and the autistic features of individuals reported here, we considered whether RFX family genes might be important transcriptional regulators of ASD risk genes. RFX family transcription factors bind to a characteristic consensus motif called an X-box (GTHNYY AT RRNAAC)27 with individual family members having additional specificity for particular subsequences within this consensus. We therefore performed RFX3, 4, and 7 motif enrichment analysis in upstream regulatory sequences of 187 ASD risk genes (the union of 102 TADA genes from Satterstrom et al. and 124 genes meeting exome-wide significance from Coe et al.)4,10 and an additional set of 447 genes identified to be upregulated in ASD brains.11 We found enrichment of X-box motifs (q value <0.05) in human ESC-neuron specific enhancers for ASD risk genes (Table S5A). As a group, RFX3 and RFX4 motifs were particularly enriched (q value <0.005), while the RFX7 motif was not (q value 0.48). X-box, RFX3 and RFX4 motifs were similarly enriched in the enhancer regions of genes upregulated in ASD brains (Table S5B).11 Enrichment of RFX motifs in promoter regions of ASD risk genes and DEGs did not emerge (data not shown). Last, we analyzed available RFX ChIP-seq data from the ENCODE project (Table S6) to determine enrichment for KEGG pathways, ciliary genes, ASD risk gene sets, and ASD DEGs (Table S7). RFX functional binding genes from most ENCODE cell lines were significantly enriched in ASD risk genes and DEGs after multiple testing correction (p.adjust < 0.05; Benjamini–Hochberg’s correction; Fig. 3g, Figure S5, Table S8). Across cell lines, there was a positive correlation between enrichment in ASD genes and RFX expression levels in that cell type (Figure S6, Table S9), indicating that higher RFX expression levels may be required to engage ASD relevant targets.
Finally, single-gene analyses showed enrichment of RFX3 and RFX4 motifs in the promoters of five ASD-associated genes (FIMO p value <0.0001, q value <0.1): AP2S1, KDM6B, ANK2, NONO, and MYT1L (Figure S4B, D),14 and RFX3 ENCODE ChIP-seq data from HepG2 cells confirmed RFX3 binding peaks in the promoters of AP2S1, KDM6B, and NONO (Figure S4E–G). Notably, de novo loss-of-function variants in KDM6B (MIM 611577) cause a neurodevelopmental syndrome that has phenotypic overlap with RFX3 haploinsufficiency as described in this report, namely mild global delays, delayed speech, hypotonia, and features of ASD and ADHD, while loss-of-function variants in NONO (MIM 300084) and MYT1L (MIM 613084) are a cause of X-linked and autosomal dominant ID, respectively (Table S4). These cases support the model that RFX members may be transcriptional activators of a subset of ASD risk genes via actions at both enhancer and promoter sites.
DISCUSSION
Our results delineate a novel human neurobehavioral phenotype including ASD, ID, and/or ADHD due to deleterious variants in RFX family transcription factors. While presence of neuroimaging findings, seizures, and dysmorphisms varied between different RFX family members, the behavioral phenotypes of individuals with RFX3, RFX4, and RFX7 were strikingly similar, and often included sensory hypersensitivity and impulsivity. Like ID/DD and ASD more generally,28 individuals with RFX variants also exhibited a male bias.
This report complements accumulating statistical genetic evidence for RFX3 as an ASD risk gene,4,21 and extends these findings to the closely related RFX family members RFX4 and RFX7. Two-thirds of individuals with RFX3 variants in our series carried an ASD diagnosis, half had ADHD, and just over half of individuals had ID. Several individuals with RFX3 variants also exhibited postpubertal cognitive or behavioral regression sometimes accompanied by psychosis. RFX3 CNVs have been previously reported in schizophrenia.20,29 RFX3 also lies within the region of the chromosome 9p deletion syndrome (OMIM 158170), associated with developmental delay, ID, and ASD, although the size of the deletions in this syndrome make RFX3 unlikely to be the sole contributor.
This report also implicates both RFX4 and RFX7 as causes of human neurodevelopmental disorders. Individuals with RFX4 or RFX7 variants were somewhat more severely affected than those with RFX3 variants, with RFX7 less likely to be associated with ASD or ADHD, but showing almost uniform diagnoses of language delay and ID (92%). There were fewer individuals identified with RFX4 variants, but those identified had high rates of ASD and ID.
RFX family members have been previously known for their biological roles in cilia development. The RFX3 transcription factor activates core components necessary for development and maintenance of both motile and primary cilia,30,31,32 and biallelic Rfx3 knockout in mice results in situs inversus, hydrocephalus, and deficits in corpus callosum formation.32,33,34 This raises the question of whether the neurodevelopmental phenotypes reported here may be mechanistically related to cilia development—e.g., a hypomorphic human ciliopathy. The majority of described genetic ciliopathies are recessive, and therefore not due to haploinsufficiency, but some (e.g., Meckel syndrome, Joubert syndrome, Bardet–Biedl syndrome, oral–facial–digital syndrome type I) may be associated with neurodevelopmental abnormalities and/or brain malformations. On the other hand, the individuals described in this report lack systemic features of ciliopathies, suggesting the alternative hypothesis that RFX haploinsufficiency may directly dysregulate the expression of ASD risk genes while leaving ciliary genes intact (Fig. 3h). Future work, which may include analyzing cilia morphology and function in cellular or animal models of RFX haploinsufficiency, or characterizing transcriptome-wide effects of RFX gene disruption, may prove helpful in distinguishing these hypotheses.
Enriched expression of RFX3 in upper cortical layer neurons places this gene in cells that are involved in communication between regions of the cortex important for higher cognition and social behavior,35 raising the possibility that haploinsufficiency may disrupt either the developmental specification, synaptic connectivity, or electrophysiological function of this set of neurons. Projection neurons in this layer have been implicated in ASD by analyses of coexpression networks of autism genes,36,37 and superficial cortical neurons exhibit the strongest amount of differential gene expression in ASD brains compared with controls.11,26 Sun and colleagues in fact showed strong enrichment of RFX motifs in differentially acetylated peaks upregulated in ASD brains compared with controls.26 Future studies aimed at understanding the downstream targets of RFX family members in human brain may shed new light on pathways important to the molecular pathogenesis of ASD, ADHD, and ID.
WEB RESOURCES
Ensembl Variant Effect Predictor, https://uswest.ensembl.org/info/docs/tools/vep/index.html. Online Mendelian Inheritance in Man (OMIM), https://www.omim.org/. GeneDx ClinVar submission page, https://www.ncbi.nlm.nih.gov/clinvar/submitters/26957/. Ambry Genetics, https://ambrygen.com/clinician/our-scientific-excellence.
Data and code availability
All data referred to in this paper are either provided in the main text/supplementary material, or appropriately referenced (where derived from pre-existing, publicly accessible data sets).
References
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 485, 237–241 (2012).
Banaschewski, T., Becker, K., Scherag, S., Franke, B. & Coghill, D. Molecular genetics of attention-deficit/hyperactivity disorder: an overview. Eur. Child Adolesc. Psychiatry 19, 237–257 (2010).
Sugiaman-Trapman, D. et al. Characterization of the human RFX transcription factor family by regulatory and target gene analysis. BMC Genomics 19, 181, https://doi.org/10.1186/s12864-018-4564-6 (2018).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584, https://doi.org/10.1016/j.cell.2019.12.036 (2020).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443, https://doi.org/10.1038/s41586-020-2308-7 (2020).
Doan, R. N., Lim, E. T. & Rubeis, S. et al. Recessive gene disruptions in autism spectrum disorder. Nat. Genet. https://doi.org/10.1038/s41588-019-0433-8 (2019).
Davis, C. A. et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 46, D794–D801, https://doi.org/10.1093/nar/gkx1081 (2018).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797, https://doi.org/10.1101/gr.137323.112 (2012).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383, https://doi.org/10.1093/bioinformatics/btv145 (2015).
Coe, B. P. et al. Neurodevelopmental disease genes implicated by de novo mutation and copy number variation morbidity. Nat. Genet. 51, 106–116, https://doi.org/10.1038/s41588-018-0288-4 (2019).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689, https://doi.org/10.1126/science.aav8130 (2019).
van Dam, T., Wheway, G., Slaats, G. G., SYSCILIA Study Group, Huynen, M. A., Giles, R. H. The SYSCILIA gold standard (SCGSv1) of known ciliary components and its applications within a systems biology consortium. Cilia. https://doi.org/10.1186/2046-2530-2-7 (2013).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287, https://doi.org/10.1089/omi.2011.0118 (2012).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018, https://doi.org/10.1093/bioinformatics/btr064 (2011).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92, https://doi.org/10.1093/nar/gkz1001 (2020).
Gao, T. & Qian, J. EnhancerAtlas 2.0: an updated resource with enhancer annotation in 586 tissue/cell types across nine species. Nucleic Acids Res. 48, D58–D64, https://doi.org/10.1093/nar/gkz980 (2020).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 928–930, https://doi.org/10.1002/humu.22844 (2015).
Harel, T. et al. Monoallelic and biallelic variants in EMC1 identified in individuals with global developmental delay, hypotonia, scoliosis, and cerebellar atrophy. Am. J. Hum. Genet. 98, 562–570, https://doi.org/10.1016/j.ajhg.2016.01.011 (2016).
Sahoo, T. et al. Copy number variants of schizophrenia susceptibility loci are associated with a spectrum of speech and developmental delays and behavior problems. Genet. Med. 13, 868–880, https://doi.org/10.1097/GIM.0b013e3182217a06 (2011).
Walsh, T. et al. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 320, 539–543, https://doi.org/10.1126/science.1155174 (2008).
Li, J. et al. Targeted sequencing and functional analysis reveal brain-size-related genes and their networks in autism spectrum disorders. Mol. Psychiatry 22, 1282–1290, https://doi.org/10.1038/mp.2017.140 (2017).
Krumm, N. et al. Excess of rare, inherited truncating mutations in autism. Nat. Genet. 47, 582–588, https://doi.org/10.1038/ng.3303 (2015).
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489, https://doi.org/10.1038/nature10523 (2011).
Polioudakis, D. et al. A single-cell transcriptomic atlas of human neocortical development during mid-gestation. Neuron 103, 785–801, https://doi.org/10.1016/j.neuron.2019.06.011 (2019).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399, https://doi.org/10.1038/nature11405 (2012).
Sun, W. et al. Histone acetylome-wide association study of autism spectrum disorder. Cell 167, 1385–1397, https://doi.org/10.1016/j.cell.2016.10.031 (2016).
Efimenko, E. et al. Analysis of xbx genes in C. elegans. Development 132, 1923–1934, https://doi.org/10.1242/dev.01775 (2005).
Polyak, A., Rosenfeld, J. A., Girirajan, S. An assessment of sex bias in neurodevelopmental disorders. Genome Med. https://doi.org/10.1186/s13073-015-0216-5 (2015).
Sahoo, T. et al. Copy number variants of schizophrenia susceptibility loci are associated with a spectrum of speech and developmental delays and behavior problems. Genet. Med. 13, 868–880, https://doi.org/10.1097/GIM.0b013e3182217a06 (2011).
El Zein, L. et al. RFX3 governs growth and beating efficiency of motile cilia in mouse and controls the expression of genes involved in human ciliopathies. J. Cell Sci. 122, 3180–3189, https://doi.org/10.1242/jcs.048348 (2009).
Choksi, S. P., Lauter, G., Swoboda, P. & Roy, S. Switching on cilia: transcriptional networks regulating ciliogenesis. Development. 141, 1427–1441, https://doi.org/10.1242/dev.074666 (2014).
Bonnafe, E. et al. The transcription factor RFX3 directs nodal cilium development and left-right asymmetry specification. Mol. Cell. Biol. 24, 4417–4427, https://doi.org/10.1128/MCB.24.10.4417-4427.2004 (2004).
Benadiba, C. et al. The ciliogenic transcription factor RFX3 regulates early midline distribution of guidepost neurons required for corpus callosum development. PLoS Genet. 8, e1002606, https://doi.org/10.1371/journal.pgen.1002606 (2012).
Baas, D. et al. A deficiency in RFX3 causes hydrocephalus associated with abnormal differentiation of ependymal cells. Eur. J. Neurosci. 24, 1020–1030, https://doi.org/10.1111/j.1460-9568.2006.05002.x (2006).
Sorensen, S. A. et al. Correlated gene expression and target specificity demonstrate excitatory projection neuron diversity. Cereb. Cortex 25, 433–449, https://doi.org/10.1093/cercor/bht243 (2015).
Willsey, A. J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell. 155, 997, https://doi.org/10.1016/j.cell.2013.10.020 (2013).
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell. 155, 1008, https://doi.org/10.1016/j.cell.2013.10.031 (2013).
Durand, B. et al. RFXAP, a novel subunit of the RFX DNA binding complex is mutated in MHC class II deficiency. EMBO J. 16, 1045–1055, https://doi.org/10.1093/emboj/16.5.1045 (1997).
Reith, W., Siegrist, C. A., Durand, B., Barras, E. & Mach, B. Function of major histocompatibility complex class II promoters requires cooperative binding between factors RFX and NF-Y. Proc. Natl. Acad. Sci. U. S. A. 91, 554–558 (1994). https://www.ncbi.nlm.nih.gov/pmc/articles/PMC42987/.
Piasecki, B. P., Burghoorn, J. & Swoboda, P. Regulatory Factor X (RFX)-mediated transcriptional rewiring of ciliary genes in animals. Proc. Natl. Acad. Sci. U. S. A. 107, 12969–12974, https://doi.org/10.1073/pnas.0914241107 (2010).
Acknowledgements
We thank Billie Lianoglou (Fetal Treatment Center, University of California at San Francisco Fetal Treatment Center) for her contributions of an RFX4 variant of unclear significance (see Supplementary Information). We thank the genetic counselors at GeneDx (Kirsty McWalter and Erin Torti) and Ambry Genetics (Meghan Towne, Zoe Powis, and Deepali Shinde) for their contributions including facilitation of clinician communication. JL was supported by award T32GM007753 from the National Institute of General Medical Sciences. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health. BZ was supported by the Manton Center Pilot Project Award and Rare Disease Research Fellowship. JEP was supported in part by the National Human Genome Research Institute (NHGRI) and National Heart Lung and Blood Institute (NHBLI) and the Baylor-Hopkins Center for Mendelian Genomics (BHCMG, UM1 HG006542). KU, SAB, CF, and SR were supported by the French Ministry of Health and the Health Regional Agency from Bretagne, Pays de la Loire and Centre Val de Loire (HUGODIMS 2, 2017). The DDD study is supported by the Wellcome Trust and the UK Department of Health Innovation Challenge Fund [HICF-1009-003] and the Wellcome Trust Sanger Institute [grant number WT098051] (Nature 2015;519:223-8). TWY was supported by grant nos. NIH/NIMH R01MH113761, NICHD/NHGRI/NIH U19HD077671 and NIH/NICHD U24HD0938487, and by a SFARI Pilot Research Award.
Author information
Authors and Affiliations
Contributions
T.W.Y., H.K.H., and T.N. conceptualized the paper, collected case information from collaborators, conducted and managed all functional studies, drafted the initial manuscript, and edited and revised the manuscript. T.W.Y. provided research study oversight. J.L. designed and performed RFX motif analysis, analyzed published brain transcriptome and single-cell RNA-sequencing data, conducted variant analyses, contributed to the manuscript, and edited and revised the manuscript. B.Z. collected ChIP-seq data and performed overrepresentation analysis, contributed to the manuscript, and edited and revised the manuscript. N.A. and C.S.G. conducted functional studies of the impact of RFX3 variants on protein stability. A.S. performed RFX variant analyses. C.A.G. and V.S. coordinated research enrollment. L.H.R., R.P., T.G., B.B.A.d.V., M.E.H.S., K.L.I.v.G., E.v.B., C.M.L.V.-T., A.H., C.D.A., L.L.I., C.B., M.W., E.F., T.L.T., K.W.G., L.B., F.V., X.W., J.L.A., M.F., G.E.T., J.E.P., E.A., A.N., R.A., A. Rauch, P.B., C.R.F., M.J.L., M.K., G.L., A.L., A.P., K.K.P., L.E.W., K.A.A., J.B., C.S., J.M., C.P.B., G.P., P.G., M.B., S.K., M.N., I.G.R., M.Y.Z., C.K., A. Reis, M.I., K.U., S.A.-B., C.F., S.R., P.D.T., J.B., Y.W., G.Z., S.S., I.B., R.A.J., W.B.D., A.B., C.M., B.K., T.M., and L.L.C. contributed clinical case information and/or analyzed exome data. P.B.A. and A.H.B. supported research subject enrollment.
Corresponding author
Ethics declarations
Ethics declaration
This series was compiled via an international collaborative effort involving Boston Children’s Hospital, Kaiser Permanente, Lyon University Hospital, Nemours/A.I. DuPont Hospital for Children, University of Zurich, Johns Hopkins University, Peninsula Clinical Genetics at Royal Devon and Exeter NHS Foundation Trust, University Medical Centre Utrecht, Radboud University Medical Centre, Dell Children’s Medical Group, Washington University School of Medicine in St. Louis, University Medical Center Groningen, Ciphergene, SUNY at Buffalo School of Medicine, Oslo University Hospital, Baylor College of Medicine, Bambino Gesu Children’s Hospital, Odense University Hospital, Indiana University Health Neuroscience Center, Seattle Children’s Research Institute, Children’s Hospital of Philadelphia, Spectrum Health Helen DeVos Children’s Hospital, Sapienza University and San Camillo-Forlanini Hospital, CHU Nantes et Service de Génétique Médicale, University of Erlangen-Nuremberg, The Islamia University of Bahawalpur, Brest University Hospital, Children’s Minnesota, Universita Cattolica del Sacro Cuore, University Hospital Heidelberg, University of Leipzig Medical Center, University of Washington, Oslo University Hospital, APHP.Sorbonne Université, Département de Génétique, Groupe Hospitalier Pitié-Salpêtrière, Centre de Référence Déficiences Intellectuelles de Causes Rares, and Weill Cornell Medical College. Collaboration was facilitated by the online genetics/genomics resource GeneMatcher. Affected individuals were clinically assessed by at least one clinical geneticist from one of the participating centers. De-identified clinical data from collaborating institutions (collected with local institutional review board [IRB] approval or deemed exempt from IRB review as per local institutional policy) was shared for analysis and publication under a study protocol approved by the Boston Children’s Hospital IRB. Consent for the publication of full-face photographs was obtained from all appropriate individuals in Fig. 1.
Permissions
The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The data used for Figure S3C were obtained from the GTEx Portal, dbGaP accession number phs000424.v8.p2 on 05/01/2020.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Harris, H.K., Nakayama, T., Lai, J. et al. Disruption of RFX family transcription factors causes autism, attention-deficit/hyperactivity disorder, intellectual disability, and dysregulated behavior. Genet Med 23, 1028–1040 (2021). https://doi.org/10.1038/s41436-021-01114-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01114-z
This article is cited by
-
Primary cilia promote the differentiation of human neurons through the WNT signaling pathway
BMC Biology (2024)
-
RFX4 is an intrinsic factor for neuronal differentiation through induction of proneural genes POU3F2 and NEUROD1
Cellular and Molecular Life Sciences (2024)
-
Multi-omics analysis identifies RFX7 targets involved in tumor suppression and neuronal processes
Cell Death Discovery (2023)
-
Description of novel variants in consanguineous Pakistani families affected with intellectual disability
Genes & Genomics (2023)
-
Identification of RFX5 as prognostic biomarker and associated with immune infiltration in stomach adenocarcinoma
European Journal of Medical Research (2022)