Abstract
Purpose
Balanced reciprocal translocation carriers are at increased risk of producing gametes with unbalanced forms of the translocation leading to miscarriage, fetal anomalies, and birth defects. We sought to determine if genome-wide cell-free DNA based noninvasive prenatal screening (gw-NIPS) could provide an alternative to prenatal diagnosis for carriers of these chromosomal rearrangements.
Methods
This pilot series comprises a retrospective analysis of gw-NIPS and clinical outcome data from 42 singleton pregnancies where one parent carried a balanced reciprocal translocation. Gw-NIPS was performed between August 2015 and March 2018. Inclusion criteria required at least one translocation segment to be ≥15 Mb in size.
Results
Forty samples (95%) returned an informative result; 7 pregnancies (17.5%) were high risk for an unbalanced translocation and confirmed after diagnostic testing. The remaining 33 informative samples were low risk and confirmed after diagnostic testing or normal newborn physical exam. Test sensitivity of 100% (95% confidence interval [CI]: 64.6–100%) and specificity of 100% (95% CI: 89.6–100%) were observed for this pilot series.
Conclusion
We demonstrate that gw-NIPS is a potential option for a majority of reciprocal translocation carriers. Further confirmation of this methodology could lead to adoption of this noninvasive alternative.
Similar content being viewed by others
INTRODUCTION
Noninvasive prenatal screening (NIPS) for trisomies 13, 18, and 21 has been widely adopted into clinical practice. Since 2011, NIPS offerings have expanded to include screening for sex chromosome aneuploidies,1,2 microdeletion syndromes,3,4 rare autosomal aneuploidies,5,6 and genome-wide subchromosomal changes.7,8,9 Several studies have reported cases where genome-wide analysis of cell-free DNA (cfDNA) has detected a terminal deletion together with a terminal duplication.10,11,12,13,14 This pattern suggests an unbalanced translocation, which may be inherited from a parent carrying a balanced form of the translocation.
Balanced reciprocal translocations are the most commonly reported structural chromosome rearrangement, carried by approximately 1 in 500 people.15 Carriers are at risk of producing gametes with unbalanced forms of the rearrangement, following malsegregation of the translocation at meiosis.16 Carriers of reciprocal translocations can have reproductive histories that may include infertility, recurrent miscarriage, fetal anomalies, and chromosomally abnormal offspring.15,17 For reciprocal translocations where the translocated segments are very large, the chance of a viable unbalanced embryo surviving to the time of prenatal diagnosis using either chorionic villus sampling (CVS) or amniocentesis, is low. Furthermore, couples with poor reproductive histories, including those who may have had preimplantation genetic testing (PGT), can be hesitant to undergo an invasive prenatal procedure, despite the low risk of miscarriage.18,19 While an assessment of unbalanced reciprocal translocations has been performed in the in vitro fertilization preimplantation setting, to date no noninvasive methods of genetic testing performed during pregnancy have been described. Using maternal plasma cfDNA to screen for an unbalanced translocation offers the possibility of a highly sensitive and specific noninvasive testing option for these couples. We present our experience of offering screening to known reciprocal translocation carrier couples using genome-wide cell-free DNA based noninvasive prenatal screening (gw-NIPS). The use of gw-NIPS for screening of other partial chromosomal imbalances is beyond the scope of this paper and is not considered further.
MATERIALS AND METHODS
Ethics statement
Genetic testing was performed in accordance with the approved ethical guidelines of the Melbourne Children’s Campus, Royal Children’s Hospital, Melbourne, Australia. All participants having gw-NIPS for known parental translocations provided written informed consent following pretest genetic counseling. Permission to collect pregnancy outcome data was obtained as part of routine Victorian Clinical Genetics Services (VCGS) informed written consent obtained at the time of maternal blood collection for gw-NIPS.
Sample cohort
Between August 2015 and March 2018, blood samples for gw-NIPS were collected from 42 singleton pregnancies of at least 10–11 weeks gestational age where one parent was a known carrier of a balanced reciprocal translocation. Samples were accepted for testing if the following inclusion criteria were met: (1) a parental karyotype was available documenting the translocation in its balanced form—this sometimes included a microarray report documenting an unbalanced form of the known parental translocation; (2) all possible unbalanced translocation segregation products contained at least one segment with a genomic imbalance of ≥15 Mb; and (3) pregnancies were confirmed as singleton by ultrasound, without evidence for co-twin demise. The mode of ascertainment of the familial translocation was recorded when available (e.g., miscarriage, prenatal diagnosis, previous chromosomally abnormal live birth).
Couples having translocation analysis were referred for gw-NIPS by the VCGS, hospital-based clinical genetics services, and specialist private ultrasound or obstetric medical practices offering NIPS. These services provided pretest genetic counseling and obtained informed written consent prior to testing. Couples seeking additional information and/or support in decision-making were offered an opportunity to speak to a genetic counselor at VCGS. Pretest genetic counseling included a discussion of alternative testing options such as prenatal diagnosis. Couples who had a live born child with an unbalanced translocation were counseled to consider a diagnostic procedure (CVS or amniocentesis). This was due to the high risk of recurrence of approximately 20% for another ongoing unbalanced translocation pregnancy.20,21 NIPS was still available to these couples if the inclusion criteria were met. Gw-NIPS was carried out at Victorian Clinical Genetics Services, Melbourne, Australia, which is accredited by the National Association of Testing Authorities, Australia and the Royal College of Pathologists of Australasia for gw-NIPS. All samples meeting the inclusion criteria between August 2015 and March 2018 were included.
Calculation of genomic size of translocation segments
The genomic size of translocation segments was estimated from the karyotype translocation breakpoint data, using the University of California–Santa Cruz (UCSC) Human Genome Browser22 (GRCh37/hg19 assembly). A maximum and minimum segment size was calculated based on the chromosome band involved and the calculations independently checked by trained analysts (N.J.F., M.D.P., and O.G.). All theoretical unbalanced segregations were considered in the calculations. Microarray analysis reports from a previous pregnancy or child with an unbalanced translocation were preferred, as these documented the exact size of the genomic imbalances.
Sample processing
Maternal blood was collected into a single Cell-Free DNA BCT® tube (Streck, Omaha, NE, USA) and accepted when received within five days from sample collection. Plasma was isolated by double centrifugation at 4 °C (1600 g for 10 minutes followed by 16,000 g for 10 minutes after transfer of the plasma portion to a fresh tube). CfDNA was extracted from 0.9 mL plasma using QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Sequencing libraries were prepared using TruSeq Nano LT kit (Illumina, San Diego, CA, USA) according to the manufacturer’s protocol. Indexed libraries from 15 samples were normalized to 10 nM and then pooled equivolume. From sample 20, double the volume of the normalized library was added to the final pool to further increase the sequence count. Multiplex sequencing was carried out on NextSeq500 (Illumina) in high output mode to produce 36 bp single-end reads. Samples for translocation analysis were pooled and sequenced with samples for standard cfDNA screening. An average of 36 ± 10.9 million unique reads per sample were generated for translocation data analysis.
Data analysis
Screening for chromosomal aneuploidies (13, 18, 21, X, Y) using standard NIPS was based on the Illumina Verifi® test (Illumina).23 Analysis for subchromosomal changes associated with known parental translocations was performed using WISECONDOR algorithm v2.0.0. WISECONDOR is an algorithm to assess shallow genome-wide sequencing to detect imbalances of 10–20 Mb at fetal fractions of ~5% and a sequencing depth of 10–12 million reads.6,24 Subchromosomal deletions or duplications were called only in the regions of the parental translocation when the z-score was ≥3, using the sliding window approach.24 The minimum sliding window called is 10 Mb in size. The actual genomic size of the subchromosomal change may be smaller than the sliding window called and is influenced by the z-score of individual 1-Mb bins within the sliding window. For each sample the WISECONDOR output plot was also manually curated and assessed for evidence of aberrations not called by the sliding window method in the region of the parental translocation. Samples were reported as high risk or low risk for an unbalanced translocation. Confirmatory diagnostic testing was recommended for all high risk test results. Ultrasound surveillance and/or prenatal diagnosis was available for low risk results.
Estimating fetal fraction
Fetal fraction was estimated using SeqFF, a multivariate model containing two regression models.25 For male fetuses, fetal fraction was also estimated based on reads mapped to the Y chromosome. A minimum fetal fraction of 5% was accepted for pregnancies returning normal test results. There was no fetal fraction cutoff for high risk results if an unbalanced translocation was detected.
Cytogenetic and clinical outcome studies
Pre- and postnatal confirmatory testing options were available to couples and were taken up at their discretion. Uptake of these options varied depending on the personal preferences of each couple. For all high risk gw-NIPS results, confirmatory diagnostic testing was recommended. For low risk results, diagnostic testing was available and ultrasound surveillance was recommended. Samples for prenatal diagnosis were obtained via CVS or amniocentesis. Postnatal samples for study included umbilical cord, newborn blood, and saliva. Diagnostic testing was performed using single-nucleotide polymorphism (SNP) microarray (HumanCytoSNP-12 or Infinium CoreExome-24, Illumina Inc., or CytoScan 750 K, Affymetrix, Santa Clara, CA, USA), or by using conventional karyotyping.
Pregnancy outcome data were obtained through records of pre- or postnatal diagnostic testing, or by a member of the VCGS NIPS genetics counseling team confirming with the patient’s referring practitioner, or with the patient themselves, that there had been a normal newborn physical exam following a low risk gw-NIPS result. Newborn physical exams were performed by an experienced obstetrician or pediatrician. A normal newborn physical exam was considered to support a low risk NIPS result. The type of pregnancy outcome data collected for each case is listed in Tables 1 and 2.
Statistical analyses
Performance characteristics of the test (sensitivity and specificity) were calculated. Confidence intervals (CIs) are two-sided 95% CIs based on the Wilson score method. Statistical analyses were performed using the R statistical package (software version 3.6.1; R Foundation for Statistical Computing, Vienna, Austria).
RESULTS
Fifty reciprocal translocations were assessed over the time period for suitability; not all translocation carrier couples were pregnant at the time of assessment. Three reciprocal translocations (6%) had at least one unbalanced segregation product where both genomic imbalances were <15 Mb in size. These translocations were excluded from testing and the couples were offered prenatal diagnosis instead. Maternal plasma cfDNA was tested in 42 singleton pregnancies from 39 reciprocal translocation carriers (Tables 1 and 2). The majority of samples tested (25/42) were referred by clinical genetics services, with 10 samples referred by specialist obstetrics practices and 7 from specialist ultrasonography groups. Three women had testing in two pregnancies. Blood was collected at a mean gestational age of 12.3 ± 2.5 weeks (median 11.6 weeks, range 10.0–21.9 weeks). Mean maternal age was 33.0 ± 5.1 years (median 33.7 years, range 18.5–43.2 years). The mean fetal fraction was 8.7 ± 3.5% (median 7.7%, range 4.0–24.1%).
Two samples with normal results (samples 2 and 12) were excluded due to fetal fractions less than 5% (4% and 4.2% respectively). Manual curation of these cases did not identify any evidence of an unbalanced rearrangement in the region of the parental translocation. Follow-up investigations were concordant with the manual curation findings; the first pregnancy (sample 2) ended in a spontaneous miscarriage at 15 weeks and molecular karyotyping showed a normal result, while the second pregnancy (sample 12) continued without prenatal diagnosis and had a normal newborn physical exam (Table 2).
Seven of 40 samples (17.5%) that met the inclusion criteria were high risk for an unbalanced form of the parental reciprocal translocation (Table 1, Figs. 1 and 2). The remaining 33 samples (82.5%) showed no evidence of subchromosomal changes in the regions of the parental reciprocal translocation (Table 2). Confirmatory diagnostic testing was available for all seven unbalanced translocations. Chromosomal microarray identified subchromosomal changes ranging in size from 0.73 to 66.7 Mb. Eight unbalanced segments were >15 Mb in size, and all were detected using the WISECONDOR sliding window method at fetal fractions ranging from 4.8% to 9.4% (Table 1). Subchromosomal changes <10 Mb were not called by the sliding window method, the largest segment size being 7.2 Mb.
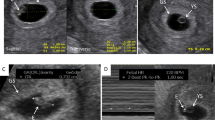
Translocation sample 13; the unbalanced 3:1 derivative chromosome 22 from the recurrent reciprocal translocation t(11;22)(q23.3q11.2) is demonstrated. (a) Far left: the WISECONDOR plot for chromosome 11. The x-axis indicates z-score, the vertical red line plots z-score using a sliding window, the purple bars indicate bins called by the algorithm at 11qter. SNP microarray analysis of saliva for chromosome 11 is shown next to the WISECONDOR plot. Blue dots are genotyping calls plotted as B-allele frequency (BAF); upper horizontal axis. Deviation of BAF across 11qter together with an increase in logR (sliding window of smoothed logR represented by red line) is consistent with an 18.2-Mb duplication of 11q23.3q25. The duplication called on WISECONDOR is concordant with the duplication called using chromosomal microarray. Right: WISECONDOR plot for chromosome 22 next to the SNP microarray analysis. The 4.2-Mb duplication of 22q11.1q11.21 is not called by the WISECONDOR sliding window method. (b) Partial karyotype; chromosome 11 homologs are at left. At right, two chromosome 22 homologs and a derivative chromosome 22 (black arrow) result in trisomy for 11q23.3q25 and trisomy for 22q11.1q11.21.
Translocation sample 17; the unbalanced derivative chromosome 15 (adjacent-1 malsegregation) from the maternal reciprocal translocation t(3;15)(p23;q26.1) is demonstrated. (a) Far left: WISECONDOR plot for chromosome 3. SNP microarray analysis of products of conception for chromosome 3 is shown next to the WISECONDOR plot. The SNP microarray data indicate a duplication of 3p26.3p24.1, which is concordant with the duplication called on WISECONDOR. Right: WISECONDOR plot for chromosome 15 next to the SNP microarray analysis; the 4.8-Mb deletion of 15q26.2q26.3 is not detected by the WISECONDOR sliding window method. (b) Partial karyotype, chromosome 3 homologs are at left. At right, two chromosome 15 homologs with one derivative chromosome 15 (black arrow) results in trisomy for 3p26.3p24.1 and monosomy for 15q26.2q26.3. Refer to Fig. 1 for interpretation of WISECONDOR and SNP microarray data.
One gw-NIPS sample (33) was identified with a more complex unbalanced rearrangement than expected from the documented parental karyotype. This sample showed an apparent 29-Mb interstitial duplication involving the long arm of chromosome 7, while the expected large deletion (>20 Mb) on the long arm of chromosome 10 was not present. Instead, evidence for a small interstitial deletion (<5 Mb) near the chromosome 10q translocation breakpoint was observed. This pattern suggested a complex maternal deletion insertion event, rather than a reciprocal translocation. The chromosomal imbalance predicted by the gw-NIPS finding was subsequently confirmed after microarray analysis of a chorionic villus sample. A repeat analysis of the maternal karyotype confirmed the complex insertional rearrangement.
The majority of couples (31/33) who received an informative low risk NIPS result declined confirmatory prenatal diagnostic testing. A normal chromosomal microarray was obtained in the two low risk pregnancies where prenatal diagnosis was elected (samples 3 and 24). Of 31 low risk pregnancies without prenatal diagnosis, 28 cases were assessed as phenotypically normal at birth. Of the remaining three pregnancies, one case (31) resulted in fetal demise at 19 weeks gestation and chromosomal microarray on fetal tissue showed no genomic imbalance. One case (18) showed microcephaly and mild dysmorphic features at birth. A normal chromosomal microarray was obtained from blood, confirming the low risk NIPS result. The final case (36) had left positional talipes and a large fontanel noted at birth. Genetic testing was not performed. These clinical findings were not considered to be associated with the parental translocation.
One low risk case (25) required three collections to return an informative result for the translocation analysis. The first two collections had fetal fractions of less than 5%. The final collection, taken at 20 weeks of gestation, had a fetal fraction of 6.5% and returned a low risk result. The patient’s body mass index (BMI) was 43.7 and she had declined prenatal diagnosis.
Our data indicate a test sensitivity of 100% (95% CI: 64.6–100%) and specificity of 100% (95% CI: 89.6–100%) for the detection of an inherited unbalanced translocation from a known carrier when our inclusion and test quality criteria were met.
DISCUSSION
Our data from this pilot series show that gw-NIPS can potentially be employed as a sensitive and specific method for detecting unbalanced translocations in pregnancies from known reciprocal translocation carriers. Unbalanced translocations were detected in 7 of 40 pregnancies (17.5%) meeting inclusion criteria, all of which were confirmed after diagnostic testing.
Balanced reciprocal translocation carriers have previously had few options for genetic screening beyond preimplantation genetic testing and cytogenetic prenatal diagnosis. While screening has been performed in the preimplantation setting,26,27 no noninvasive genetic testing options during pregnancy have been available. As gw-NIPS allows the interrogation across the genome for subchromosomal deletions and duplications, application of this noninvasive methodology to screen for unbalanced translocations is an attractive option for these couples. Similar to balanced reciprocal translocations, balanced inversion carriers are at risk of producing gametes containing recombinant chromosomes. This method could also be applied to screen a pregnancy where one parent carries an inversion, as well as pregnancies at risk for other known inherited structural chromosomal imbalances, such as interchromosomal insertions. However, care is needed when accepting these types of referrals for gw-NIPS as more complex rearrangements are possible. For example, Ardalan et al.28 described an intrachromosomal insertion initially misclassified as a pericentric inversion. If a rearrangement has been misclassified, this might lead to an unexpected recombinant outcome below the screening resolution of the assay.
Guidelines for preimplantation genetic testing recommend that patients should understand that a misdiagnosis is possible and prenatal diagnosis should be offered to all women who become pregnant following PGT.29 There is limited literature on patient choices around prenatal diagnosis following preimplantation genetic testing for structural chromosome rearrangements (PGT-SR). When PGT was used to screen for aneuploidy (PGT-A), one study reported only 5 of 68 couples opted for prenatal diagnostic testing.30 Our experience indicates that gw-NIPS could be an acceptable alternative to prenatal diagnosis for couples who have had PGT-SR.
The majority of couples who received a low risk result elected to continue their pregnancies without prenatal diagnosis. Many couples in this group had reproductive histories that included recurrent early miscarriages and were uncomfortable with the notion of an invasive procedure for a pregnancy that was still viable at 11–12 weeks gestational age. While many of these couples were at a relatively low risk of having a pregnancy with an unbalanced translocation at the time of NIPS due to the very large genomic imbalances expected, their low risk NIPS result provided additional reassurance, as well as allowing for concurrent screening of other chromosome anomalies by NIPS, including the common autosomal trisomies. There was one high risk test result where the pregnancy was continued without prenatal diagnosis (case 13). The unbalanced translocation, causing Emanuel syndrome, was confirmed after birth. The couple had a history of recurrent early pregnancy loss and were committed to continuing the pregnancy.
Our strict assessment protocol applied to each translocation was intentionally conservative, considering the smallest possible segment size for each translocation breakpoint. Conventional chromosome analysis has limited resolution, is subjective in the assignment of translocation breakpoints, and translocations can be misclassified. This was demonstrated by case 33, where an insertional rearrangement had been misclassified as a reciprocal translocation. This subjectivity needs to be considered in the assessment of the genomic size of the translocation segregation products, and for this reason a microarray assessment involving an unbalanced translocation is preferred, as the precise genomic size of the translocation products can be determined.
Our approach allowed for sensitive screening using the laboratory’s standard whole genome sequencing workflow. Test sensitivity is determined by a combination of sequence count, fetal fraction, the size of the genomic imbalance, and the bioinformatics algorithm being used to generate each result.24,31,32,33,34 The sequencing depth was increased to an average of 36 M reads by doubling the volume of the individual library in the sample pool, prior to sequencing. Fetal fraction is also a key metric influencing the ability to detect subchromosomal changes in cfDNA.7,35 A minimum fetal fraction cutoff of 5% was applied to maintain high sensitivity in the 15–20 Mb range, as previously described using the WISECONDOR algorithm,24 and employed in conjunction with an increase in sequencing depth. Finally, prior knowledge of the genomic location of the inherited imbalances is expected to increase the performance of the assay compared with screening genome-wide for de novo or unknown inherited imbalances.36 Our data demonstrates this approach can reliably detect known unbalanced segregants with imbalances ≥15 Mb. Manual curation and assessment of the WISECONDOR output data provided an additional level of scrutiny in cases called as low and high risk.
This approach did not generate a result at first collection in three cases (2, 12, 24). Over the period of the study a repeat collection policy for cases not meeting the fetal fraction requirement was implemented. Samples 2 and 12 were collected prior to the repeat collection policy implementation and a repeat collection was not requested. A repeat collection was requested for sample 24 and an informative result was returned after the third collection. The high maternal BMI of 43.7 likely contributed to the persistent low fetal fraction.37 Repeat collections in low fetal fraction samples could reduce the test failure rate; however, consideration needs to be given to the timing of repeat collections and test results; ideally the patient should have access to early prenatal diagnosis, should a high risk result be returned.
A previous report noted the failure of NIPS to detect an unbalanced translocation from a known carrier that caused Wolf–Hirschhorn (4p-) syndrome.38 In this case NIPS was requested with a microdeletion screening panel that included 4p-. The testing laboratory was not informed of the parental translocation and a false negative result for 4p- was returned. This case demonstrates the importance of communicating clinical history to the testing laboratory to ensure the methodology employed is sufficiently sensitive for the purpose of the test. We manage this aspect of our testing by requiring presubmission and assessment of translocation karyotype data before a sample is accepted for testing.
In our experience, gw-NIPS at 11–12 weeks gestational age is ideal for translocation carrier couples. Test results are available within 3–5 days, which allows for timely prenatal diagnosis using CVS when test results are returned as high risk. Inherited, unbalanced translocations are unlikely to be mosaic and absent from the cytotrophoblast, thus confined placental mosaicism will not affect the accuracy of NIPS results in this application.36 Identification of an inherited unbalanced translocation in this setting might potentially be considered diagnostic, but we have recommended prenatal diagnosis until sufficient outcome data are collated. One potential cause of a false positive result might be an undiagnosed co-twin demise having the unbalanced translocation, with a chromosomally normal or balanced translocation live twin. For this reason we require a prior ultrasound, which is also used to confirm gestational age. In the absence of ultrasound, an undiagnosed dichorionic twin pregnancy might also increase the risk for a false negative result, as the effective fetal fraction per twin is halved.
Although our data are promising, several limitations should be noted. The majority of outcomes from low risk pregnancies relied on normal newborn physical exams. Although we cannot exclude the possibility that a baby with a chromosome imbalance might have had a normal newborn examination, it is notable that most of the imbalances are large (minimum size of 20 Mb) and these would be expected to cause obvious physical and neurological abnormalities in the newborn period. Moreover, none of these offspring have subsequently come to the attention of our statewide clinical and laboratory genetics service. Gw-NIPS showed 100% concordance with cytogenetic testing and pregnancy outcomes, however, we acknowledge the wide CIs of sensitivity and specificity due to limited sample size. More data are needed to narrow these intervals. A small proportion of samples will not yield an informative result or will be contraindicated for testing using our method. In this series, 3/50 (6.0%) of couples seeking gw-NIPS were excluded on the basis of chromosomal segmental size limitations. Of samples tested, 2/42 (4.7%) did not produce a result for the translocation due to low fetal fraction. Increasing sequencing depth further could reduce this rate but would be associated with an increased test cost. During pretest counseling, couples should be made aware that a decision to elect NIPS instead of diagnostic testing may mean that other clinically significant chromosome conditions could be overlooked. Lastly, misclassification of a parental chromosome rearrangement might lead to an unexpected recombinant imbalance below the screening resolution of the assay. Despite these limitations, approximately 90% of reciprocal translocations couples seeking testing using our gw-NIPS benefited from this noninvasive test option.
Screening pregnancies of known translocation carriers is a clinically relevant and practical application for gw-NIPS. As discussed by Srebniak et al.,36 while carrying a balanced translocation or inversion is typically an indication for diagnostic testing, couples may decline invasive prenatal diagnosis because of the miscarriage risk. Our experience indicates there is strong demand from couples seeking a noninvasive testing option. We show that gw-NIPS can potentially provide a sensitive and specific screening option for couples where one partner carries a balanced reciprocal translocation. Further confirmation of this methodology could lead to adoption of this noninvasive alternative.
References
Mazloom AR, Dzakula Z, Oeth P, et al. Noninvasive prenatal detection of sex chromosomal aneuploidies by sequencing circulating cell-free DNA from maternal plasma. Prenat Diagn. 2013;33:591–597.
Bianchi DW, Parsa S, Bhatt S, et al. Fetal sex chromosome testing by maternal plasma DNA sequencing: clinical laboratory experience and biology. Obstet Gynecol. 2015;125:375–382.
Wapner RJ, Babiarz JE, Levy B, et al. Expanding the scope of noninvasive prenatal testing: detection of fetal microdeletion syndromes. Am J Obstet Gynecol. 2015;212:332.e1–332.e9.
Gross SJ, Stosic M, McDonald-McGinn DM, et al. Clinical experience with single-nucleotide polymorphism-based noninvasive prenatal screening for 22q11.2 deletion syndrome. Ultrasound Obstet Gynecol. 2016;47:177–183.
Pertile MD, Halks-Miller M, Flowers N. et al. Rare autosomal trisomies, revealed by maternal plasma DNA sequencing, suggest increased risk of feto-placental disease. Sci Transl Med. 2017;9:eaan1240.
Van Opstal D, van Maarle MC, Lichtenbelt K, et al. Origin and clinical relevance of chromosomal aberrations other than the common trisomies detected by genome-wide NIPS: results of the TRIDENT study. Genet Med. 2018;20:480–485.
Zhao C, Tynan J, Ehrich M, et al. Detection of fetal subchromosomal abnormalities by sequencing circulating cell-free DNA from maternal plasma. Clin Chem. 2015;61:608–616.
van der Meij KRM, Sistermans EA, Macville MVE, et al. TRIDENT-2: national implementation of genome-wide noninvasive prenatal testing as a first-tier screening test in the Netherlands. Am J Hum Genet. 2019;105:1091–1101.
Lefkowitz RB, Tynan JA, Liu T, et al. Clinical validation of a noninvasive prenatal test for genomewide detection of fetal copy number variants. Am J Obstet Gynecol. 2016;215:227.e1–227.e16.
Sahoo T, Hovanes K, Strecker MN, Dzidic N, Commander S, Travis MK. Expanding noninvasive prenatal testing to include microdeletions and segmental aneuploidy: cause for concern? Genet Med. 2016;18:275–276.
Ehrich M, Tynan J, Mazloom A, et al. Genome-wide cfDNA screening: clinical laboratory experience with the first 10,000 cases. Genet Med. 2017;19:1332–1337.
Mei J, Wang H, Zhan L. 10p15.3p13 duplication inherited from paternal balance translocation (46,XY,t(5;10)(q35.1;p13)) identified on noninvasive prenatal testing. J Obstet Gynaecol Res. 2017;43:1076–1079.
Liu H, Gao Y, Hu Z, et al. Performance evaluation of NIPT in detection of chromosomal copy number variants using low-coverage whole-genome sequencing of plasma DNA. PLoS One. 2016;11:e0159233.
Fiorentino F, Bono S, Pizzuti F, et al. The clinical utility of genome-wide non invasive prenatal screening. Prenat Diagn. 2017;37:593–601.
Gardner RJM, Amor DJ. Gardner and Sutherland’s chromosome abnormalities and genetic counseling. 5th edition. Oxford: Oxford University Press; 2018.
Mackie Ogilvie C, Scriven PN. Meiotic outcomes in reciprocal translocation carriers ascertained in 3-day human embryos. Eur J Hum Genet. 2002;10:801–806.
Stern C, Pertile M, Norris H, Hale L, Baker HW. Chromosome translocations in couples with in-vitro fertilization implantation failure. Hum Reprod. 1999;14:2097–2101.
Kukulu K, Buldukoglu K, Keser I, et al. Psychological effects of amniocentesis on women and their spouses: importance of the testing period and genetic counseling. J Psychosom Obstet Gynaecol. 2006;27:9–15.
Akolekar R, Beta J, Picciarelli G, Ogilvie C, D’Antonio F. Procedure-related risk of miscarriage following amniocentesis and chorionic villus sampling: a systematic review and meta-analysis. Ultrasound Obstet Gynecol. 2015;45:16–26.
Daniel A, Hook EB, Wulf G. Risks of unbalanced progeny at amniocentesis to carriers of chromosome rearrangements—data from United States and Canadian laboratories. Am J Med Genet. 1989;33:14–53.
Youings S, Ellis K, Ennis S, Barber J, Jacobs P. A study of reciprocal translocations and inversions detected by light microscopy with special reference to origin, segregation, and recurrent abnormalities. Am J Med Genet A. 2004;126A:46–60.
Kent WJ, Sugnet CW, Furey TS, et al. The human genome browser at UCSC. Genome Res. 2002;12:996–1006.
Bianchi DW, Platt LD, Goldberg JD, et al. Genome-wide fetal aneuploidy detection by maternal plasma DNA sequencing. Obstet and Gynecol. 2012;119:890–901.
Straver R, Sistermans EA, Holstege H, Visser A, Oudejans CB, Reinders MJ. WISECONDOR: detection of fetal aberrations from shallow sequencing maternal plasma based on a within-sample comparison scheme. Nucleic Acids Res. 2014;42:e31.
Kim SK, Hannum G, Geis J, et al. Determination of fetal DNA fraction from the plasma of pregnant women using sequence read counts. Prenat Diagn. 2015;35:810–815.
Tan YQ, Tan K, Zhang SP, et al. Single-nucleotide polymorphism microarray-based preimplantation genetic diagnosis is likely to improve the clinical outcome for translocation carriers. Hum Reprod. 2013;28:2581–2592.
Tobler KJ, Brezina PR, Benner AT, Du L, Xu X, Kearns WG. Two different microarray technologies for preimplantation genetic diagnosis and screening, due to reciprocal translocation imbalances, demonstrate equivalent euploidy and clinical pregnancy rates. J Assist Reprod Genet. 2014;31:843–850.
Ardalan A, Prieur M, Choiset A, Turleau C, Goutieres F, Girard-Orgeolet S. Intrachromosomal insertion mimicking a pericentric inversion: molecular cytogenetic characterization of a three break rearrangement of chromosome 20. Am J Med Genet A. 2005;138A:288–293.
Harton G, Braude P, Lashwood A, et al. ESHRE PGD consortium best practice guidelines for organization of a PGD centre for PGD/preimplantation genetic screening. Hum Reprod. 2011;26:14–24.
Kimelman D, Confino R, Confino E, Shulman LP, Zhang JX, Pavone ME. Do patients who achieve pregnancy using IVF-PGS do the recommended genetic diagnostic testing in pregnancy? J Assist Reprod Genet. 2018;35:1881–1885.
Lo KK, Karampetsou E, Boustred C, et al. Limited clinical utility of noninvasive prenatal testing for subchromosomal abnormalities. Am J Hum Genet. 2016;98:34–44.
Srinivasan A, Bianchi DW, Huang H, Sehnert AJ, Rava RP. Noninvasive detection of fetal subchromosome abnormalities via deep sequencing of maternal plasma. Am J Hum Genet. 2013;92:167–176.
Benn P, Cuckle H. Theoretical performance of noninvasive prenatal testing for chromosome imbalances using counting of cell-free DNA fragments in maternal plasma. Prenat Diagn. 2014;34:778–783.
Chen S, Lau TK, Zhang C, et al. A method for noninvasive detection of fetal large deletions/duplications by low coverage massively parallel sequencing. Prenat Diagn. 2013;33:584–590.
Helgeson J, Wardrop J, Boomer T, et al. Clinical outcome of subchromosomal events detected by whole-genome noninvasive prenatal testing. Prenat Diagn. 2015;35:999–1004.
Srebniak MI, Vogel I, Van Opstal D. Is carriership of a balanced translocation or inversion an indication for noninvasive prenatal testing?. Expert Rev Mol Diagn. 2018;18:1–3.
Ashoor G, Poon L, Syngelaki A, Mosimann B, Nicolaides KH. Fetal fraction in maternal plasma cell-free DNA at 11-13 weeks’ gestation: effect of maternal and fetal factors. Fetal Diagn Ther. 2012;31:237–243.
Qi Z, Madaan S, Chetty S, Yu J, Wiita AP. False negative fetal cell free DNA screening for microdeletion syndromes in the presence of an unbalanced translocation involving monosomy 4p. Prenat Diagn. 2017;37:420–422.
Acknowledgements
We acknowledge the expert technical assistance and additional analytical support provided by R. Manser, I. Burns, S. Baeffel, T. Harrington, and A. Tsegay.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Flowers, N.J., Burgess, T., Giouzeppos, O. et al. Genome-wide noninvasive prenatal screening for carriers of balanced reciprocal translocations. Genet Med 22, 1944–1955 (2020). https://doi.org/10.1038/s41436-020-0930-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0930-2
Keywords
This article is cited by
-
What proportion of couples with a history of recurrent pregnancy loss and with a balanced rearrangement in one parent can potentially be identified through cell-free DNA genotyping?
Molecular Cytogenetics (2023)
-
Inherited unbalanced reciprocal translocation with 3q duplication and 5p deletion in a foetus revealed by cell-free foetal DNA (cffDNA) testing: a case report
European Journal of Medical Research (2021)