Abstract
Purpose
To expand the recent description of a new neurodevelopmental syndrome related to alterations in CDK19.
Methods
Individuals were identified through international collaboration. Functional studies included autophosphorylation assays for CDK19 Gly28Arg and Tyr32His variants and in vivo zebrafish assays of the CDK19G28R and CDK19Y32H.
Results
We describe 11 unrelated individuals (age range: 9 months to 14 years) with de novo missense variants mapped to the kinase domain of CDK19, including two recurrent changes at residues Tyr32 and Gly28. In vitro autophosphorylation and substrate phosphorylation assays revealed that kinase activity of protein was lower for p.Gly28Arg and higher for p.Tyr32His substitutions compared with that of the wild-type protein. Injection of CDK19 messenger RNA (mRNA) with either the Tyr32His or the Gly28Arg variants using in vivo zebrafish model significantly increased fraction of embryos with morphological abnormalities. Overall, the phenotype of the now 14 individuals with CDK19-related disorder includes universal developmental delay and facial dysmorphism, hypotonia (79%), seizures (64%), ophthalmologic anomalies (64%), and autism/autistic traits (56%).
Conclusion
CDK19 de novo missense variants are responsible for a novel neurodevelopmental disorder. Both kinase assay and zebrafish experiments showed that the pathogenetic mechanism may be more diverse than previously thought.
Similar content being viewed by others
INTRODUCTION
The multisubunit Mediator complex is a conserved multiprotein interface between gene-specific transcription factors and RNA polymerase II that influences nearly all stages of transcription in programs that control diverse physiological processes, including cell growth and homeostasis, development, and differentiation.1,2 Regulation of the Mediator complex activity is partly achieved by the reversible association of a four-subunit kinase module (CDK8 module) involved in transcriptional repression and activation.1,3,4,5 This module consists of CDK8, cyclin C, Mediator complex subunit 12 (MED12), and Mediator complex subunit 13 (MED13).6 In vertebrates, the kinase module is more complex due to separate gene duplication events that resulted in the origin of paralogs for three of the four components: MED12L for MED12, MED13L for MED13, and CDK19 for CDK8, respectively.7 In turn, these paralogs are mutually exclusive of each other, though not exclusive of the other kinase module members.8
Over the last few years, germline variants in different CDK8 module subunits including CDK8, MED12, MED13, MED12L, and MED13L have been found in individuals with various neurodevelopmental disorders.3,4,9,10,11,12 The only other component of the kinase module with catalytic activity and CDK8 paralog (77% shared sequence identity, 97% identity in the kinase domain), CDK19, is a gene that comprises 13 exons; it is located at chromosomal region 6q21 and encodes a major isoform of 502 amino acids.2,13 Previously, the only report of an individual with an alteration involving CDK19 was that of an adult female with intellectual disability (along with bilateral congenital retinal folds, microcephaly, and café au lait spots) with a de novo pericentric inversion in chromosome 6 that resulted in CDK19 haploinsufficiency.14
Recently, three individuals with syndromic intellectual disability were shown to have de novo nonsynonymous variants in the CDK19 gene (Thr196Ala in 2 individuals and Tyr32His in another) and the functional relevance of these variants was evaluated by a Drosophila model system although the Drosophila genome does not have a direct counterpart of human CDK19 and the closest homologue is the Cdk8 gene.15 While biallelic disruption of the Cdk8 locus in the Drosophila was lethal, forced expression of wild-type human CDK19 rescued embryonic lethality whereas forced expression of the mutant CDK19 (Thr196Ala or Tyr32His) failed to rescue the embryonic lethal phenotype.15 The authors suspected that these variants represented hypomorphic alleles because these alleles failed to rescue embryonic lethality.
Phenotypic and mutational spectra of CDK19-related disorder (early infantile epileptic encephalopathy 87, OMIM 618916) should be further delineated and the evaluation of the variant alleles using model organisms whose genome carries both CDK8 and CDK19 homolog will be helpful to better understand the underlying pathogenetic mechanisms. We herein provide data showing that pathogenic de novo missense variants in CDK19 are associated with a neurodevelopmental disorder that solidifies the recently proposed concept of mediator kinase modulopathies. We functionally validated recurrent nonsynonymous variants in CDK19 using kinase assay and zebrafish as a vertebrate disease model.
MATERIALS AND METHODS
Genomic studies and variant interpretation
Blood samples were collected from each participating individual and after genomic DNA was extracted, all individuals underwent trio exome sequencing (ES) according to standard procedures. Variant nomenclature is according to Human Genome Variation Society (HGVS) guidelines with reference transcript NM_015076.4. In silico predictions for interpretation of the variants reported in this study were performed using an integrated genetic database for coding variants.16 Pathogenicity of missense variants was ascribed based on the American College of Medical Genetics and Genomics (ACMG) variant interpretation guidelines.17 To assess changes in protein conformation, protein 3D structures of two different CDK19 missense variants, p.Gly28Arg and p.Tyr32His, were obtained using the Have Our Protein Explained (HOPE) project analysis program based on the structure of the homologous human PDB 5XS2.18 Predictions of protein stability were obtained by using I-mutant2.0 based on the structure homologous to the human PDB 5HBE under stable conditions (temperature = 25 ˚C and pH = 7).19
Preparation of recombinant CDK19 kinase modules for substrate phosphorylation assays
The complementary DNA (cDNA) encoding each human CDK19, Cyclin C, MED12, or MED13 was amplified by reverse transcription polymerase chain reaction (RT-PCR) from a HeLa cDNA library and cloned into a modified pFastBac vector (ThermoFisher). The constructs for CDK19 and MED13 contained an N-terminal Flag-tag and a C-terminal His-tag, respectively. The sequence of each construct was verified by Sanger sequencing. The baculoviruses of each gene were obtained by using the Bac-to-Bac baculovirus expression system (ThermoFisher) and were coinfected into Sf9 cells at 28 °C for seven days. The coinfected Sf9 cells were then subjected to affinity purification using anti-Flag affinity M2 resins (ThermoFisher) followed by size fractionation (glycerol gradient centrifugation) to obtain near stoichiometric complexes. To evaluate the purity of the obtained complexes, the complexes were resolved by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and fluorescent-stained with OrioleTM (Bio-Rad) (Figure S1).
Substrate phosphorylation assays for CDK19 Gly28Arg and Tyr32His variants
A plasmid for glutathione S-transferase (GST)-tagged recombinant C-terminal domain (CTD) of mouse RNA polymerase II (Pol II) large subunit was prepared as described.20 A cDNA encoding the C-terminal region of human STAT1 (510-751amino acids) was amplified by RT-PCR and cloned into the pGEX6P-1 bacterial expression vector (Merck). The sequence of the construct was verified by Sanger sequencing. Each GST-tagged recombinant protein was expressed and purified as previously described21 and used as substrates for the following phosphorylation assay. Substrate phosphorylation assays were performed by incubation of around 5 ng of purified CDK19 kinase module and GST-fusion protein (5 ng GST-CTD or 100 ng GST-STAT1) in kinase buffer (20 mM Tris-HCl (pH 7.5), 20% glycerol, 100 mM KCl, 5 mM MgCl2, 1 mM DTT, 20 μg/ml BSA, 50 μM ATP) at 30 °C for three hours. The reaction products were resolved by SDS-PAGE and analyzed by immunoblotting using phosphorylation-specific anti-CTD antibody (3E10 for Ser5P, Ascenion) and phospho-Ser727 specific anti-STAT1 antibody (Cell signaling 9177). The proteins were illuminated using ECL Plus Western Blotting Substrate (ThermoFisher) and visualized with LAS4000mini (GE Healthcare).
Autophosphorylation assays for CDK19 Gly28Arg and Tyr32His variants
Autophosphorylation activity of the CDK19 carrying Gly28Arg and Tyr32His variants identified in reported individuals were evaluated in vitro. Human embryonic kidney 293 (HEK293) cells were transfected with the expression plasmids encoding Flag-tagged CDK19 protein or its mutants (Gly28Arg and Tyr32His) with Cyclin C expression plasmids. Each fusion protein was expressed and immunoprecipitated with anti-Flag antibody. After purification, the protein was used as substrate for the following phosphorylation assay. Aliquots of immunoprecipitated CDK19 proteins were incubated for 90 minutes with or without 50 μM ATP in 40 μL of kinase buffer containing 40 mM Tris (pH7.5), 100 μM BSA, and 20 mM MgCl2 at 25 °C, and the ADP produced by NLK autophosphorylation was measured using a ADP-Glo® Kinase Assay kit (Promega). Immunoprecipitates from cells transfected with empty vector were used for measurement of background activity. Amount of immunoprecipitated CDK19 proteins were confirmed by immunoblotting with anti-Flag antibody.
In vivo zebrafish assays of the CDK19Y32H and CDK19G28R variants
Human CDK19-WT mRNA and mutant mRNA were injected into zebrafish one-cell stage embryos and morphological features were evaluated at the 5-day postfertilization (dpf) stage of the embryos. Zebrafish Cdk19 protein shares 88% homology with human CDK19 protein, and the amino acid sequences flanking the variants are highly conserved. We cloned the cDNA of wild-type human CDK19 (a gift from Dr. Conaway, Addgene 24762) into the pCS2p+Flag vector in-frame of the Flag-tag. The Tyr32His (CDK19Y32H) and Gly28Arg (CDK19G28R) variants identified in the patients’ genomic DNA sequences were then introduced into human CDK19 genomic sequence using a QuikChange Site-Directed Mutagenesis kit (Agilent). Next, capped mRNA was synthesized using a SP6 mMESSAGE mMACHINE kit (Ambion, Austin, TX) and purified on Micro Bio-Spin columns (Bio-Rad, Hercules, CA). Capped mRNAs (CDK19-WT and CDK19Y32H) were injected into one-cell stage embryos (500 pg) and the embryos were evaluated for morphologic abnormalities at the 5-dpf stage. The same amount of EGFP mRNA was injected as a negative control. To characterize endogenous cdk19 knockout in zebrafish, morpholino antisense oligonucleotides (MOs) were used (Figure S2). The MOs blocking splicing were obtained from Gene Tools. The sequences of MOs were as follows: cdk8, 5’-AGCTTGATGTACTTACTGCTATCTC-3’; cdk19, 5’-TAGCATTGGAGTTACTCACAGCGAT-3’. For all injections, 5–10 pg of MOs were injected at the one-cell stage. Rate of abnormalities between groups was compared using Fisher’s exact t-test.
RESULTS
Through international collaboration and aided by the web-based tool GeneMatcher,22 we identified seven distinct de novo missense CDK19 variants affecting five different residues with strong molecular evidence for pathogenicity in 11 unrelated individuals with neurodevelopmental disorders. The variants were identified by trio ES from research and clinical cohorts.
With a constraint z score of 3.56 for missense variants and a probability of loss-of-function intolerance (pLI) of 1.00 for CDK19 (o/e = 0.03), the gnomAD database suggests a selection against coding variation in this gene.23 The six novel variants identified in this cohort in CDK19 were absent from ExAC and gnomAD23 and were found de novo in all individuals (Fig. 1). All variants were located within the kinase domain (between amino acids 21 and 335), affected evolutionarily conserved amino acids, and met criteria for classification as likely pathogenic/pathogenic according to American College of Medical Genetics and Genomics (ACMG) criteria (Table S1).17 One variant was recurrently observed: p.Gly28Arg was identified in four unrelated individuals while another individual had a substitution at the same position: p.Gly28Ala. Three additional unrelated individuals also had substitutions in the same residue: p.Tyr32Cys and p.Tyr32His.
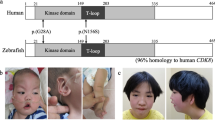
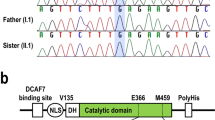
(a) Cartoon representation of the human CDK19 (based on UniProt: Q9BWU1) showing the location of the protein kinase domain (21–335; blue box), Gly-rich loop (27–35; pink box) ATP binding domain, and activation segment (173-203, green box). Novel variants from this report are represented in bold and italics. Four variants were located within the ATP binding domain and three more within the activation segment of the CDK19 kinase domain. Note that the p.Gly28Arg and the p.Tyr32His substitutions were identified in four unrelated subjects each (one individual with the p.Tyr32His had been previously reported). Images were constructed using Illustrator for Biosequence (IBS1.0.1)27. (b) Homology model of CDK19 (Structure ID: NP_055891.1_3) illustrating two recurrent missense variants. Green: Gly28Arg, red: Tyr32His variant, light blue: ATP binding site. Bottom left and right. Close-up of the two variants with protein colored in gray and side chains of both the wild-type and the mutant residue in green and red, respectively.
The Gly28 and Tyr32 residues are located within the Glycine-rich loop ATP binding pocket (amino acids 27–35); the protein stability is predicted to be decreased and could affect the interaction with other molecules (Fig. 1).24 The additional substitutions reported here, p.Phe197Leu, p.Trp198Cys, and p.Arg200Trp (along with the previously reported p.Thr196Ala) affect conserved amino acids within the activation segment of CDK19.
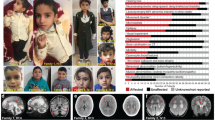
We assessed the phenotypes of the 11 subjects here described along with three individuals previously reported in the literature (eight males; age 0.75–14.5 years at the last clinical assessment) with CDK19 substitutions (summarized in Table 1, additional information in Table S2). Most individuals had unremarkable perinatal histories, were born at or near term, and had normal anthropometric measurements at birth. Infantile hypotonia (79%) leading to early gross motor developmental delay and gait/balance abnormalities later in life (43%) was often the first manifestation. Moderate to severe developmental delay was often seen and typically with associated profound language delay (8/11 individuals 3 years or older were nonverbal). Behavioral symptoms were often described with autism/autistic traits reported in five individuals. The head circumference was normal except in one subject, who had significant microcephaly (-4.0 SD at 13 years of age). Nondistinctive brain anomalies were seen in 57% of individuals with brain atrophy being the most common in three individuals. Other findings included thinning of the corpus callosum, delayed myelination, and ventriculomegaly (Figure S3, Table S2). Seizures were present in 64% including four individuals with infantile spasms (current antiepileptic management listed in Table S2). Ophthalmological abnormalities were frequent (64%) including strabismus (n = 7), refractive errors (n = 3), optic nerve atrophy (n = 1), and/or cortical blindness (n = 1). Variable abnormalities in the extremities were present in eight subjects. Of interest, edema of hands and feet was present in three individuals (Figure S3). Facial dysmorphism was identified in all subjects often with features described as coarse (Fig. 2, Table S2).
Common features included full lips, coarsening of face, thick alea nasi, hypertelorism, low nasal bridge (full description in Table S2). (a) Individual 1 at 4 years of age. (b) Individual 4 at 11 years of age. (c) Individual 5 at 10 years of age. (d) Individual 6 at 6 years of age. (e) Individual 8 at 3 years of age. (f) Individual 9 at 12 years of age. (g) Individual 10 at 11 years of age.
In vitro kinase assay of human CDK19 with amino acid substitutions
We evaluated the functional relevance of representative amino acid substitutions p.Gly28Arg and p.Tyr32His, which have now been identified in multiple individuals. Kinase activities of the wild-type and mutant CDK19 proteins were evaluated both by substrate phosphorylation assay and autophosphorylation assay (Fig. 3, Figure S1). The kinase activity of protein with p.Gly28Arg substitution was significantly lower than that of the wild-type protein. Substrate phosphorylation assay showed that CDK19 protein with the p.Gly28Arg substitution almost lost its ability to phosphorylate CTD or STAT1 (Fig. 3a–b).25 Autophosphorylation assay also showed that CDK19 protein with p.Gly28Arg substitution had significantly lower kinase activity than that of the wild-type protein (Fig. 3c).
(a,b) Substrate phosphorylation assays were performed using the equal amounts of each purified recombinant CDK19 kinase module (wild-type [wt], Gly28Arg [G28R], or Tyr32His [Y32H]) and either recombinant Pol II-CTD (a) or STAT1 (b) as a substrate (30 °C, 3 hours). The reaction products were analyzed by immunoblotting using indicated phospho-specific antibodies for evaluating phosphorylation levels and anti-GST antibody (Pol II-CTD) or anti-STAT1 antibody (STAT1) for evaluating the total amount of the substrates (top row). Each assay was performed multiple times and one representative result is shown. Phosphorylation degree (bar charts on the bottom) was evaluated by quantification of the intensity of immunoblotting signal using ImageJ software. Error bars indicate the standard error of the mean (SEM). The asterisks represent statistically significant differences between wt and the indicated variant (Fisher’s exact t-test, **p < 0.01, ***p < 0.001). Note that the Gly28Arg mutant almost completely loses the ability to phosphorylate both the CTD (a) and STAT1 (b), although this mutant still can form the complex with other subunits (cyclin C, MED12, and MED13) as seen in Fig. S1A. The Tyr32His variant, on the other hand, tends to show higher kinase activity than wild-type CDK19 (a,b). (c) Autophosphorylation activity rates of wild-type CDK19, Gly28Arg (G28R)-mutated CDK19, and Tyr32His (Y32H)-mutated CDK19 using HEK293 cells. Note that kinase activity of Gly28Arg (G28R) mutant tended to be lower than that of wild-type CDK19. Kinase activity of Tyr32His (Y32H) mutant, was (significantly) higher than that of wild-type CDK19. The values shown are the average of the values observed in one representative experiment in which each transfection was performed in triplicate. Error bars indicate the standard error of the mean (SEM). The imunoprecipitated CDK19 proteins were analyzed by immunoblotting (bottom panel) showing stable levels.
In contrast to the p.Gly28Arg substitution, kinase activity of protein with p.Tyr32His substitution was significantly higher than that of the wild-type protein. Substrate phosphorylation assay showed that CDK19 protein with the Tyr32His substitution had an enhanced ability to phosphorylate CTD or STAT1 (Fig. 3a–b).25 Autophosphorylation assay also showed that CDK19 protein with p.Tyr32His substitution had significantly higher kinase activity than that of the wild-type protein (Fig. 3c).
In vivo functional assays of human CDK19 containing Tyr32His and Gly28Arg substitutions
Because CDK19 functions as a subunit of multimers, the results of the in vitro kinase assay do not always reflect the pathogenetic mechanism in vivo. We developed a zebrafish model in which an extra amount of human CDK19 protein with and without the p.Tyr32His or the p.Gly28Arg variants were expressed in addition to intact zebrafish Cdk19 proteins. Injection of wild-type human CDK19 mRNA induced a shortened body axis, hypoplastic anterior structures, and an aberrant tail in approximately 17% of the 5-dpf stage embryos (Fig. 4a). By comparison, injection of CDK19 mRNA with the Tyr32His variant and the Gly28Arg variant markedly increased the fraction of embryos with these morphological changes to 45% (p = 0.0001) and 35% (p = 0.01), respectively. The ratio of dead fish was also significantly higher in those with mutated CDK19 injection than in those with wild-type CDK19 injection or controls (Fig. 4a). We then compared the resulting phenotype from the injection of the mutant human CDK19 mRNA to the endogenous cdk19 knockout in zebrafish using MOs. Double knockdown of cdk8 and cdk19 (cdk8/19) showed anatomical defects, especially in the head region (Fig. 4b).
(a) Zebrafish with abnormal phenotypes at 5 dpf after injection of wild-type, Tyr32His, and Gly28Arg human CDK19 messenger RNA (mRNA). The ratios of fish with abnormal phenotypes (shortened body axis, hypoplastic anterior structures, and an aberrant tail) and dead fish were significantly higher in fish injected with the Tyr32His or Gly28Arg mutants than in those without injection, those injected with enhanced green fluorescent protein (EGFP) or in those injected with wild-type CDK19. (b) The ratio of fish with abnormal phenotypes (head abnormality) was significantly higher in fishes injected with both cdk19 and cdk8 morpholino antisense oligonucleotides (MOs) compared with no injection or single mutants. The asterisks represent statistically significant differences between wt and the indicated variant (Fisher’s exact t-test, **p < 0.01, ***p < 0.001).
DISCUSSION
With 77% shared sequence identity (97% identity in the kinase domain), CDK19 is a paralog of the bona fide Mediator subunit CDK8 and together they associate with C-type cyclins as part of the Mediator complex.2,13 There was mounting prior compelling evidence of the potential role of CDK19 and neurodevelopmental phenotypes. First, CDK8 and CDK19 interact with Mediator as part of the kinase module and subunits within this module are related to syndromic forms of intellectual disability.12,25,26 Second, CDK19 is highly expressed in both fetal brain and fetal retina.14 Third, the most structurally and functionally similar molecule, Cdk8, is important for regulation of dendritic development and has now been directly linked to a syndromic neurodevelopmental disorder.2,3,14 Lastly, an adult female with intellectual disability with a de novo pericentric inversion in chromosome 6 that resulted in CDK19 haploinsufficiency was previously reported.14 To further characterize the role of CDK19 in development, we studied the phenotype of endogenous cdk19 knockout in zebrafish using cdk19 morpholino antisense oligonucleotides. Double knockdown of cdk8 and cdk19 (cdk8/19) showed anatomical defects, especially in the head region suggesting and endogenous role of cdk8/19 in craniofacial and neuronal development. The similarities seen in the zebrafish model regardless of the mutation correlate with the shared common phenotypic features seen in the described individuals with these variants including intellectual disability, craniofacial dysmorphism, and epilepsy.
The importance of CDK19 in human neurodevelopment was recently highlighted by the report of three individuals with global developmental delay, hypotonia, facial dysmorphisms, and epilepsy found to have de novo missense variants in CDK19.15 While haploinsufficiency (i.e., loss of function) of CDK19, according to the prior Drosophila experiments, was interpreted as the potential mechanism linking this gene and the resulting phenotype for the two previously reported variants (including p.Tyr32His here reported as well),15 our results using both kinase assay and zebrafish experiments showed that the pathogenetic mechanism may be more diverse and both loss-of-function and gain-of-function mechanisms are in action. Kinase assays showed that the p.Gly28Arg and p.Tyr32His variants resulted in decreased and increased kinase activity, respectively. Overexpression of either human CDK19Y32H or CDK19G28R proteins in zebrafish, however, resulted in similar severe structural abnormalities suggesting a potential disruption of the normal allele’s function by other mechanisms such as a dominant-negative effect. Given the severity of the resulting phenotype, we were unable to determine whether the effects were specific or not to the overexpression experiments. Additional rescue or other experiments would be needed in the future to further elucidate the pathogenicity of these CDK19 variants.
The overall frequency and impact of CDK19 alterations in the context of seizures and developmental delay appears to be small. Screening for coding pathogenic variants and deletions of CDK19 in 100 sporadic patients with intellectual disability with negative fragile X testing did not reveal any abnormalities.14 There are no reported missense substitutions in the gnomAD database for the CDK19 canonical transcript (ENST00000368911) affecting either the Glycine-rich loop or the activation segment. Three of the individuals with CDK19 variants included in this cohort were diagnosed through ES performed at a single clinical laboratory that, as of June 2020, has identified a total of 3 cases in a total of 34,609 individuals who underwent ES for the evaluation of intellectual disability/developmental delay (0.01%), suggesting a low frequency.
In summary, we present 11 additional individuals with de novo missense variants in CDK19 bringing the total of described individuals to 14. The eight total different protein substitutions are located in either the Glycine-rich loop or activation segment of the CDK19 kinase domain with Gly28 (five individuals) and Tyr32 (four individuals) emerging as potential hotspots. The phenotype of the CDK19-related disorder includes universal developmental delay, often severe and with profound speech compromise. Other common features include high frequency of seizures, hypotonia, ophthalmologic anomalies, balance abnormalities, and autism/autistic traits. From the ubiquitous description of facial dysmorphism in these individuals, coarse facial features seem to be often recognized. The presence of edema in hands and feet could also be a suggestive feature that will be elucidated as more cases are described. Further investigation is needed to determine possible subgrouping among individuals with CDK19-related disorder and to further delineate this emerging condition.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Please see ClinVar SCV001424134.1, SCV001424133.1, SCV001424135.1, SCV001428436, SCV001428437, SCV001428438, SCV001428439, SCV001428440, and SCV001428441.
References
Malumbres, M. Cyclin-dependent kinases. Genome Biol. 15, 122 (2014).
Spaeth, J. M., Kim, N. H. & Boyer, T. G. Mediator and human disease. Semin. Cell Dev. Biol. 22, 776–787 (2011).
Calpena, E. et al. De novo missense substitutions in the gene encoding CDK8, a regulator of the mediator complex, cause a syndromic developmental disorder. Am. J. Hum. Genet. 104, 709–720 (2019).
Nizon, M. et al. Variants in MED12L, encoding a subunit of the mediator kinase module, are responsible for intellectual disability associated with transcriptional defect. Genet. Med. 21, 2713–2722 (2019).
Furumoto, T. et al. A kinase subunit of the human mediator complex, CDK8, positively regulates transcriptional activation. Genes Cells 12, 119–132 (2007).
Dannappel, M. V., Sooraj, D., Loh, J. J. & Firestein, R. Molecular and in vivo functions of the CDK8 and CDK19 kinase modules. Front. Cell. Dev. Biol. 6, 171 (2018).
Allen, B. L. & Taatjes, D. J. The Mediator complex: a central integrator of transcription. Nat. Rev. Mol. Cell. Biol. 16, 155–166 (2015).
Daniels, D. et al. Mutual exclusivity of MED12/MED12L, MED13/13L, and CDK8/19 paralogs revealed within the CDK-Mediator kinase module. J Proteomics Bioinform. S2:004. https://doi.org/10.4172/jpb.S2-004.
Adegbola, A. et al. Redefining the MED13L syndrome. Eur J Hum. Genet. 23, 1308–1317 (2015).
Graham, J. M. Jr. & Schwartz, C. E. MED12 related disorders. Am. J. Med. Genet. A. 161A, 2734–2740 (2013).
Snijders Blok, L. et al. De novo mutations in MED13, a component of the Mediator complex, are associated with a novel neurodevelopmental disorder. Hum. Genet. 137, 375–388 (2018).
Poot, M. Mutations in mediator complex genes CDK8, MED12, MED13, and MEDL13 mediate overlapping developmental syndromes. Mol. Syndromol. 10, 239–242 (2020).
Poss, Z. C., Ebmeier, C. C. & Taatjes, D. J. The Mediator complex and transcription regulation. Crit. Rev. Biochem. Mol. Biol. 48, 575–608 (2013).
Mukhopadhyay, A. et al. CDK19 is disrupted in a female patient with bilateral congenital retinal folds, microcephaly and mild mental retardation. Hum. Genet. 128, 281–291 (2010).
Chung, H. L. et al. De novo variants in CDK19 are associated with a syndrome involving intellectual disability and epileptic encephalopathy. Am. J. Hum. Genet. 106, 717–725 (2020).
Li, J. et al. VarCards: an integrated genetic and clinical database for coding variants in the human genome. Nucleic Acids Res. 46, D1039–D1048 (2018).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Venselaar, H., Te Beek, T. A., Kuipers, R. K., Hekkelman, M. L. & Vriend, G. Protein structure analysis of mutations causing inheritable diseases. An e-Science approach with life scientist friendly interfaces. BMC Bioinformatics 11, 548 (2010).
Capriotti, E., Fariselli, P. & Casadio, R. I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res. 33, W306–310 (2005).
Hirose, Y. & Manley, J. L. RNA polymerase II is an essential mRNA polyadenylation factor. Nature. 395, 93–96 (1998).
Hirose, Y., Iwamoto, Y., Sakuraba, K., Yunokuchi, I., Harada, F. & Ohkuma, Y. Human phosphorylated CTD-interacting protein, PCIF1, negatively modulates gene expression by RNA polymerase II. Biochem. Biophys. Res. Commun. 369, 449–455 (2008).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 928–930 (2015).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 536, 285–291 (2016).
Niknafs, N. et al. MuPIT interactive: webserver for mapping variant positions to annotated, interactive 3D structures. Hum. Genet. 132, 1235–1243 (2013).
Tsutsui, T., Fukasawa, R., Tanaka, A., Hirose, Y. & Ohkuma, Y. Identification of target genes for the CDK subunits of the Mediator complex. Genes Cells 16, 1208–1218 (2011).
Audetat, K. A., Galbraith, M.D., & Odell, A. T. et al. A kinase-independent role for cyclin-dependent kinase 19 in p53 response. Mol. Cell. Biol. 37, e00626-16 (2017).
Liu, W. et al. IBS: an illustrator for the presentation and visualization of biological sequences. Bioinformatics. 31, 3359–3361 (2015).
Acknowledgements
Part of this work was supported by the Ministry of Education, Culture, Sports, Science and Technology (MEXT KAKENHI; grant numbers 24118003 and 25131704 to Y.O. and grant numbers 20K21502 and 20H05365 to T.I.), the Japanese Society for the Promotion of Science (JSPS) (KAKENHI; grant numbers 20570162 and 17K07282 to Y.H.), and Japan Agency for Medical Research and Development (JP18ek0109288h0002 and JP19ek0109288h003 to T.I.).
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.A.Z., K. Kosaki. Formal analysis: Y.A.Z. Investigation: K.A., M.O., S.H., S.I., Y.K., Y.O., Y.H., T.I., K. Kosaki. Resources: Y.A.Z, T.U., A-E. L., T.B., K. Kloth, N.E., D.H., M.H., K.A., D.V., C.L.E.-Z., M.J. G.S., R.E.S., M. Morrow, A.S.-V., J.P., R.R., M. Muona, A-K.A., K.P., J.L., J. G-A., T.K., Writing—original draft: Y.A.Z., Y.H., T.I., K. Kosaki. Writing—review & editing: Y.A.Z., Y.H., T.I., K. Kosaki.
Corresponding author
Ethics declarations
Ethics declaration
Approval for the review and reporting of these cases was given by the Institutional Review Boards of the University of Arkansas for Medical Sciences and of the Helsinki University Central Hospital and written informed consent was obtained from the individuals’ parents for genetic testing and the scientific use of photo documentation. Approval for review and reporting these cases was given by all the Japanese medical institutions: Keio University School of Medicine, National Center for Child Health and Development, Osaka Women’s and Children’s Hospital, and Central Hospital, Aichi Human Service Center. Zebrafish wild-type strain (AB) was raised and maintained under standard conditions. All experimental animal care was performed in accordance with institutional and national guidelines and regulations. The study protocol was approved by the Institutional Animal Care and Use Committee of Osaka University, RIMD Permit R02-04.
Competing interests
M.J.G.S., M. Morrow, and R.E.S. are employees of GeneDx, Inc. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zarate, Y.A., Uehara, T., Abe, K. et al. CDK19-related disorder results from both loss-of-function and gain-of-function de novo missense variants. Genet Med 23, 1050–1057 (2021). https://doi.org/10.1038/s41436-020-01091-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-01091-9
This article is cited by
-
Cdk8/CDK19 promotes mitochondrial fission through Drp1 phosphorylation and can phenotypically suppress pink1 deficiency in Drosophila
Nature Communications (2024)
-
Six years’ accomplishment of the Initiative on Rare and Undiagnosed Diseases: nationwide project in Japan to discover causes, mechanisms, and cures
Journal of Human Genetics (2022)