Abstract
Purpose
We investigated the value of transcriptome sequencing (RNAseq) in ascertaining the consequence of DNA variants on RNA transcripts to improve the diagnostic rate from exome or genome sequencing for undiagnosed Mendelian diseases spanning a wide spectrum of clinical indications.
Methods
From 234 subjects referred to the Undiagnosed Diseases Network, University of California–Los Angeles clinical site between July 2014 and August 2018, 113 were enrolled for high likelihood of having rare undiagnosed, suspected genetic conditions despite thorough prior clinical evaluation. Exome or genome sequencing and RNAseq were performed, and RNAseq data was integrated with genome sequencing data for DNA variant interpretation genome-wide.
Results
The molecular diagnostic rate by exome or genome sequencing was 31%. Integration of RNAseq with genome sequencing resulted in an additional seven cases with clear diagnosis of a known genetic disease. Thus, the overall molecular diagnostic rate was 38%, and 18% of all genetic diagnoses returned required RNAseq to determine variant causality.
Conclusion
In this rare disease cohort with a wide spectrum of undiagnosed, suspected genetic conditions, RNAseq analysis increased the molecular diagnostic rate above that possible with genome sequencing analysis alone even without availability of the most appropriate tissue type to assess.
Similar content being viewed by others
INTRODUCTION
Clinical exome sequencing has become a routine test for the diagnosis of the extraordinarily heterogeneous set of rare Mendelian diseases. The diagnostic rate across diverse clinical laboratories has remained consistently around 30%.1,2 Exome sequencing–negative cases likely remain unsolved because the causal variant(s) reside in a gene not yet associated with disease or is a type not readily detectable by exome sequencing.3,4 Such variants include structural variants (SVs), repeat expansions, and deep intronic variants. In theory, genome sequencing has the potential to capture most of the variants missed by exome sequencing. However, predicting the consequence of novel or rare noncoding variation is not always possible using genome sequencing alone. Therefore, the increase in diagnostic rate attributed to genome sequencing relative to exome sequencing is generally modest, largely due to detection of SVs, which, thus far, resolved about 3–7% of all such undiagnosed cases.5,6,7
Transcriptome sequencing (RNAseq) is commonly used for assessing differential expression in case–control studies and is a powerful tool to identify alternative splicing.8,9,10 Recent studies demonstrate RNAseq increases diagnostic yield when applied to specific cohorts of exome sequencing–negative patients with well-defined disease types, including muscle disease11,12 or disorders of mitochondrial dysfunction,13 for which relevant tissues (muscle and fibroblast, respectively) are easily obtainable. However, the diagnostic rates from these studies do not extrapolate well for the majority of cases commonly referred for clinical exome or genome sequencing, which are highly enriched for neurodevelopmental diseases.1,2,14
As one of the clinical sites from the Undiagnosed Diseases Network (UDN) phase I program, a nationwide effort funded by the National Institutes of Health (NIH) established to characterize undiagnosed or previously unrecognized diseases, we pioneered applying RNAseq systematically to genome-negative cases with the goal of delivering a more comprehensive genetic testing method for identifying the molecular diagnosis for patients with a wide spectrum of presumed rare Mendelian diseases that have remained undiagnosed despite extensive testing. Our approach relies on first identifying all potentially causal DNA variants from genome sequencing data, coupled with a comprehensive search for alteration of messenger RNA (mRNA). Here, we report analysis from 138 affected participants and unaffected family members who were enrolled at the University of California–Los Angeles (UCLA) clinical site and determined the relative contribution of RNAseq to the diagnostic rate when integrated with genome sequencing in this cohort.
MATERIALS AND METHODS
Study population
The study was approved by the National Human Genome Research Institute (NHGRI) central Institutional Review Board (IRB; registration number 00000014).15 From 234 referrals, a total of 113 probands were accepted and evaluated at the UCLA clinical site during the first phase of UDN (July 2014 to August 2018). Twenty-five probands had similarly affected family members enrolled. Informed consent was obtained from all subjects participating in the study.
DNA sequencing and analysis
The UCLA clinical site took a sequencing-first approach, where accepted cases first underwent exome or genome sequencing. Of the 113 cases enrolled with UDN clinical and genetic evaluation completed in phase I of the study, 29 cases underwent exome sequencing and 77 underwent genome sequencing as they had prior uninformative exome sequencing (exome-negative). Seven cases had prior sequencing and the data were obtained and reanalyzed in the UDN. If parental samples were available, both parents were sequenced simultaneously with the proband on the same platform.
Genomic DNA was extracted at the UCLA Molecular Pathology Laboratory from whole-blood sample collected in EDTA tubes. For three patients, additional genomic DNA was extracted from cultured skin fibroblast cells (fibroblast) or muscle biopsies (muscle) to search for somatic mosaicism (Supplementary Materials and Methods). Exome and genome sequencing were performed at the UDN sequencing cores at the average depth of coverage of >150× and >50×, respectively. Analysis was performed by a custom-built pipeline developed at UCLA (Supplementary Materials and Methods).
RNA sequencing and analysis
Total RNA was extracted from whole blood, fibroblast, muscle, or bone marrow following standard protocols (Supplementary Materials and Methods). Library preparation and sequencing were performed at the UCLA sequencing core facilities to generate 50~100 million 69~150 bp paired-end reads. Analysis was performed by a custom-built pipeline developed at UCLA (Supplementary Materials and Methods).
Variant interpretation for exome and genome sequencing
For single-nucleotide variants (SNVs) and small insertions and deletions (indels), common variants with minor allele frequency >1% in public databases were removed, as these are unlikely to be causal for a rare genetic disease. Remaining rare variants were categorized by their predicted consequence at the protein level, genic location, and inheritance pattern (Supplementary Materials and Methods). All variants were interpreted in the context of the patient’s phenotype.
Variant interpretation for genome sequencing and RNAseq
RNAseq data was used to determine the functional consequence at the transcript level for all rare synonymous, splice region, untranslated region (UTR), and deep intronic variants. RNAseq data for the entire family was loaded into Integrative Genomics Viewer (IGV)16 and each variant position was inspected manually to search for splicing abnormalities creating a novel splice isoform. Only variants with at least five RNAseq reads spanning their genomic positions were considered. If no splicing abnormalities were observed spanning the two flanking exons, it was determined that there were no splicing abnormalities resulting from the genomic variant. Additionally, to rule out the possibility of the abnormal transcript undergoing nonsense-mediated decay (NMD), the presence of biallelic expression of the gene was assessed by examining allelic ratios for other heterozygous variants (determined by exome or genome sequencing) present in the same transcript. We also confirmed that there was no evidence of differential expression level, especially if there was no heterozygous coding variant in the transcript to assess the allelic ratio. If the variant was inherited from a parent and/or shared with other family members, the RNAseq data from those could be used to observe the splicing and/or expression abnormalities from the same DNA variant and assess NMD. Splicing abnormalities were classified as exon skipping, inclusion of intronic pseudoexon, exon extension, exon retraction, and intron retention. When a splicing abnormality was observed, the genomic location of the new junction was searched in the splice junction database that was populated by importing the splice junction table output from STAR17 for 338 blood, 317 fibroblast, and 44 muscle RNAseq samples available from this and other internal studies. Junctions were separated into known (annotated according to GENCODE v19) or novel splice junctions. For each novel junction, we calculated (1) the number of individuals in which it was observed, and (2) the coverage ratio, defined as the number of spliced reads aligning to the novel junction divided by the sum of this and the number of reads aligning to an annotated junction with a common donor/acceptor site. We used this information to determine both the rarity of a novel splice junction, in terms of both occurrence and quantitative effect. Even if a novel event was observed in multiple individuals within the database, if the coverage ratio was a significant outlier in the patient, it was considered a rare event and evaluated for disease relevance.
Genomic data board review
The clinical relevance of all rare variants was evaluated at the UCLA-UDN genomic data board meeting consisting of the entire UCLA clinical site team.
Variant classification and reporting
Variant classification was done as previously described2 and a molecular diagnosis was given only when the variants were classified as likely pathogenic or pathogenic according to current American College of Medical Genetics and Genomics (ACMG) sequence interpretation guidelines18 and consistent with the established mode of inheritance for a given gene and associated disease(s). All genomic variants were confirmed by Sanger sequencing at the UDN sequencing cores or UCLA Orphan Diseases Testing Center. All likely pathogenic splicing abnormalities were confirmed by reverse transcription polymerase chain reaction (RT-PCR) and/or complementary DNA (cDNA) sequencing (Table S1).
Clinical evaluation
Following sequence interpretation, patients and their family members were invited to UCLA for one to five days for the UDN clinical evaluation. Based on reported phenotype and identified variants of interest from sequencing, the UDN team utilized clinical laboratory studies, specialist consultations, and/or procedures to further detail the phenotype and identify diagnoses during their visit. Thirteen cases were deemed not appropriate for genetic analysis and not used in this study because their clinical indication at the time of the UDN clinical evaluation was not consistent with the information provided in the prior medical notes acquired before the enrollment.
Diagnostic rate calculation and statistical analysis
A 95% confidence interval (CI) for the diagnostic rate, calculated as proportions assuming independence among participants, was calculated using the Wald method as implemented by QuickCalc.19 The diagnostic rate for RNAseq was calculated by counting the number of cases that were only diagnosable by integrating RNAseq data with genome sequencing data. Cases for which RNAseq was used as supporting evidence by allowing to observe the exact consequence at the transcript level were not included.
RESULTS
Study population characteristics
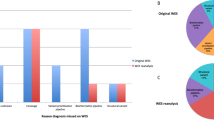
Of 100 probands deemed appropriate for genetic analysis, 79% (79/100) were ≤18 years of age at the time of enrollment (Table 1). Ninety percent (71/79) of children and 62% (13/21) of adults were sequenced as a trio with both parents available. There was a higher proportion of males among children (male: 48/79 [61%; 95% CI, 50–71%]) than adults (male: 9/21 [43%; 95% CI, 24–63%]). Consistent with the statistics from the UDN at large,14 the most common primary symptoms were neurologic (47/100; 47%) and musculoskeletal or orthopedic (21/100; 21%). The most prevalent clinical indications were developmental delay overall (61/100; 61%) and among children (58/79; 73%), and muscle weakness (13/21; 62%) among adults.
Diagnostic rate from exome and genome sequencing without RNAseq
Of 100 probands who received comprehensive genetic analysis after UDN clinical evaluation, 31 cases were diagnosed by exome or genome sequencing alone (Fig. 1, Table 2). Twenty-three cases were diagnosed with SNVs or small indels within protein coding sequence, essential splice site locations, or a recurrent deep intronic location (exome = 9/26; genome = 14/74): 8 of the 14 genome cases were undiagnosed by prior exome sequencing because a novel disease gene was discovered after UDN enrollment. Of the remaining 60 genome cases, 8 cases were diagnosed by SVs (13%; 95% CI, 7–24%): 1 case with mixed triploidy detected in fibroblast but not in blood, 1 with a repeat expansion, and 6 cases with SVs (deletions) of sizes 500 bp–200 kb. Of all 31 cases diagnosed by exome or genome sequencing, 61% (19/31) had de novo variants, 13% (4/31) had compound heterozygous variants, and 16% (5/31) had homozygous variants (Table S2).
Molecular diagnostic rate in the 113 Undiagnosed Diseases Network–University of California–Los Angeles (UDN-UCLA) clinical site cohort enrolled between July 2014 and August 2018. aDetermined not appropriate for UDN genetic study after clinical evaluation. bSingle-nucleotide variant (SNV)/small indel variants within coding exons (includes essential splice site (+/−2 bp) variants) that are predicted to be nonsynonymous or loss-of-function: of the 23 probands, 9 were diagnosed with exome sequencing (35%; includes 1 proband who was diagnosed with a recurrent deep intronic pathogenic variant in COL6A1) and 14 with genome sequencing (19%). cExome-negative cases were removed from further analysis for this study due to the lack of DNA sequencing data in the noncoding genomic region. dSV: Structural variants affecting coding exons (includes mixed triploidy and repeat expansion). eVariants that are synonymous or in untranslated region (UTR). fVariants within +/−3 to +/−10 bp from the exon–intron boundaries. ES exome sequencing, GS genome sequencing.
Diagnostic rate from genome sequencing with RNAseq
RNAseq was performed on 48 families (91 samples, Fig. S1) who were genome sequencing–negative after analysis of coding SNVs, small indels, and SVs (Fig. 1) and an additional 284 samples to use as controls. By integrating RNAseq data with genome sequencing data, we were able to diagnose an additional seven cases (7/48; 15%; 95% CI, 7–27%, Table 2), increasing the overall molecular diagnostic rate to 38% (38/100; 95% CI, 29–48%). The genomic variants identified in these seven cases were splice region SNVs (+/−3 bp to +/−10 bp region, n = 1), synonymous variants (n = 2), and deep intronic SNVs or SVs (n = 4) that caused alterations in RNA splicing: exon skipping (n = 2), inclusion of intronic pseudoexon (n = 2), and intronic retention (n = 3) (Table 3). Critically, in all seven cases, the clinical significance of these DNA variants could not be determined without the use of RNAseq.
Illustrative cases
A 2-year-old female with severe hypotonia, global developmental delay with cerebral atrophy, hypomyelination, and central volume loss of the cerebrum underwent trio genome sequencing and trio RNAseq from blood. A paternally inherited known pathogenic frameshift variant, previously identified by exome sequencing, was confirmed in SEPSECS (OMIM 613009), associated with autosomal recessive (AR) pontocerebellar hypoplasia, type 2d (PCH2d, OMIM 613811).20 No pathogenic maternally inherited coding variant was observed by analysis of SNVs, indels, or SVs. However, there were maternally inherited rare synonymous (42 bp downstream from a splice acceptor site) and deep intronic variants in SEPSECS of unknown significance. From RNAseq, the 130-bp exon 7 containing the synonymous variant (p.Leu282=) was skipped in about half of the mRNAs leading to an unexpected frameshift and identification of the pathogenic allele (Fig. 2a, top). There were no transcripts with the synonymous variant, suggesting that exon skipping was occurring in most of the maternally inherited transcripts. Interestingly, the novel splice junction that joins exon 6 and exon 8 was detected in 25/695 additional unrelated samples without this variant, but at a much lower ratio compared with our proband and mother, suggesting that exon 7 may be susceptible to low-level skipping (Fig. 2a, bottom). Conventionally, synonymous variants would not be considered for pathogenicity unless already proven to be pathogenic but transcript-level analysis is necessary to determine if more synonymous variants are indeed disease-causing as loss-of-function (LoF) variants.21 In this case, of 21 synonymous variants in AR disease genes that are expressed in blood or muscle, only this one in SEPSECS resulted in an observable splicing abnormality.
Sashimi plots and noncanonical junction coverage ratio plots. a SEPSECS exon skipping. b LMNA intron retention. c SLC25A46 intron retention and intronic pseudoexon inclusion. For the sashimi plots, the exon coverage and the splice junctions for the family members carrying the genomic variant are in red, for the family members not carrying the genomic variant are in blue, and for the unrelated individuals not carrying the genomic variant are in gray. Canonical exons and the genomic variants are shown above the respective sashimi plots with the exon number and the transcription direction indicated. For the noncanonical junction coverage ratio plots, the x-axis is the total number of reads at the junction (sum of canonical and noncanonical junctions) and the y-axis is the noncanonical junction coverage ratio (noncanonical junction coverage/total junction coverage). Family members carrying the noncanonical junctions are noted by P (proband), M (mother), or F (father) in respective color of the tissues observed and unrelated individuals carrying the noncanonical junctions are noted in dots in respective color of the tissues observed (blue: blood; yellow: fibroblast; red: muscle). Below each plot is a violin plot showing the coverage distribution from all samples at each junction for different tissues (FB fibroblast, SM muscle, WB blood).
A 7-year-old male with progressive muscle weakness and elevated creatine kinase (highest at 1200) with muscle biopsy showing myopathic changes with mild to moderate myofiber size variation was enrolled with extensive prior negative genetic testing. Trio genome sequencing and trio RNAseq from blood and fibroblast were performed. Clinical evaluation of all coding variation was negative, so RNAseq was utilized to evaluate all rare noncoding variation. Within the known muscle disease genes (Table S3), compiled from commercially available gene panels for muscle diseases and Human Gene Mutation Database (HGMD)22,23 search, there were 2 de novo heterozygous, 8 homozygous, and 31 hemizygous deep intronic variants. Additionally, there were 86 inherited deep intronic heterozygous variants in five autosomal recessive disease genes that were a good phenotypic match, with muscle weakness and myopathy reported as part of the reported phenotypic spectrum, and with one heterozygous coding variant in trans. Of 127 noncoding variants, 86 had sufficient coverage to check for splicing abnormalities in blood or fibroblast. In both tissues, intron retention was observed surrounding a de novo 26-bp deep intronic deletion in LMNA (OMIM 150330). All of intron 6, which contained the 26-bp deletion (66 bp), was included in the transcript, predicted to insert 22 amino acids. LMNA is associated with multiple muscle diseases that are consistent with the patient’s phenotype,24,25 but the patient had some unique features, such as features of progeria and lipodystrophy, making suspicion of an LMNA-associated muscle disease difficult for the referring physician. There is no similar in-frame insertion reported in the literature and this intron retention was not observed in any other sample within our internal database (Fig. 2b).
A 3-year-old male with global developmental delay with regression, seizures, and optic atrophy with negative extensive genetic workup was enrolled and underwent trio genome sequencing and trio RNAseq from whole-blood samples. There were no clinically significant variants that could explain the phenotype within the coding exons. Of 14 de novo, 15 homozygous, 36 hemizygous deep intronic variants necessitating evaluation with RNAseq, 18 were well expressed in blood, but no splicing abnormalities were detected. In addition, there were 64 inherited deep intronic heterozygous variants in 3 autosomal recessive disease genes that were a good phenotypic match, expressed in blood and with one heterozygous coding variant in trans. Compound heterozygous variants in SLC25A46 (OMIM 610826) were observed: a maternally inherited heterozygous missense variant and a paternally inherited heterozygous deletion located within intron 3, removing 114 bp of an Alu-repeat sequence. RNAseq showed presence of two different splicing abnormalities affecting more than half of total transcripts: (1) intron retention between 3’-end of exon 3 and 5’ end of the deletion and (2) inclusion of intronic pseudoexon upstream of the deleted region (Fig. 2c). These changes are predicted to disrupt the transcript by adding 1626 bp and 139 bp, respectively. SLC25A46 is associated with autosomal recessive neuropathy, hereditary motor and sensory, type VIB (OMIM 616505) with phenotypic variability from congenital lethal pontocerebellar hypoplasia to a milder form of optic atrophy with later-onset sensory neuropathy depending on the variant type and the degree of protein instability.26,27 The proband’s phenotype fit the more severe pontocerebellar hypoplasia form.
DISCUSSION
Here we report how RNAseq can be integrated with genome sequencing to interpret both exonic and intronic SNVs and SVs and significantly increase the molecular diagnostic rate for a wide spectrum of rare Mendelian diseases. To our knowledge, this is the first cohort study that has systematically applied RNAseq from multiple, readily available patient tissue sources, including fibroblast, muscle, and blood, to interpret genomic variants in a general patient population referred for clinical genetic testing, without preselection of cases by the primary symptoms or with a priori knowledge of the most appropriate tissue source for RNAseq. In our cohort of 48 consecutive cases that remained unsolved by genome sequencing alone, we were able to identify pathogenic DNA variants in 7 cases (15%; 95% CI, 7–27%) by observing the impact on mRNA as determined by rare, abnormal transcriptional events consistent with Mendelian inheritance, achieving an overall diagnostic rate of 38%. This increase is greater than the published diagnostic yield obtained by genome sequencing–SV5,6 or chromosomal microarray (CMA) alone,28 highlighting the significance of transcriptome sequencing in improving the diagnostic rate for rare Mendelian diseases. Even after removing the two cases that had strong a priori candidate genes (COL6A1 and DMD), the diagnostic rate augmentation due to RNAseq when combined with genome sequencing was 11% (5/46, 95% CI, 4–23%).
Although our cohort size is small, the contribution of RNAseq to the diagnostic rate of 33% achieved in the muscle disease cohort (4 of 12 cases, 95% CI: 14–61%, Table 2) was consistent with prior diagnostic rates11,12 in similar patient populations that had muscle tissue on which to perform RNAseq, suggesting that the lower overall diagnostic rate in nonmuscle diseases within our cohort is due to use of blood or fibroblasts for observing gene expression. Of these four cases, because of the limited availability of tissue, two were diagnosed from muscle and two from fibroblast and blood (Table 3), which obviated the need to request muscle biopsy for diagnosis of a child. We note though that all four cases would have been diagnosed by muscle alone, three by fibroblast alone, and two by blood alone. Similarly for the neurology disease cohort, the three cases diagnosed (diagnostic rate 12%, 95% CI: 3–31%, Table 2) were from two blood and one fibroblast (Table 3) but all three would have been diagnosed by fibroblast alone while two would have been diagnosed by blood alone. This is consistent with blood being the least informative tissue due to lower expression of genes involved in syndromic genetic disorders referred for clinical exome or genome sequencing11 (Supplementary Materials and Methods, Fig. S2) and is in line with a recent study that investigated the utility of RNAseq solely from blood to identify rare disease genes, reporting a 7.5% diagnostic rate (including three cases that, by our criteria, could have been diagnosed by exome or genome sequencing alone).29 We expect that of the remaining 41 undiagnosed cases, another few cases may receive diagnoses if we had access to muscle for the 3 muscle disease cases and fibroblast for the 7 neurology disease cases and accessing the most affected tissue or implementing transdifferentiation protocols to induce gene expression could further improve diagnostic yield. Other possible reasons for the remaining undiagnosed cases include the causal variant residing in a gene or noncoding RNA not yet associated with disease, not being readily detectable by short-read sequencing, or resulting in a more subtle isoform switch only in specific tissues that are not accessible.
As opposed to other studies that utilized RNAseq to search globally for an outlier in the transcriptome,11,13,29 our approach initiated from observed rare genomic variants. We surveyed each genomic variant to determine if there were any evidence of altered transcription. For each case, there were hundreds of rare genomic variants that required evaluation, most of which were benign. Even though all seven of the splice-altering DNA variants discovered in our cohort were detected with a positive score by at least one of the three splice-altering variant predictors we assessed, the algorithms were not highly specific, with all three generating high scores for the vast majority of DNA variants with no observation of splicing abnormality from RNAseq. In total, only 15 of 347 predicted splice-altering DNA variants (4.3%) had any support from RNAseq data (Fig. S3). This indicates that the prediction methods can be useful for prioritizing variants but more high-throughput and sensitive methods with global search options remain essential. We find that the biggest analytical challenge lies in differentiating the splicing change that is characteristic of a pathogenic DNA variant from natural variation in splicing and noisy data, particularly within genes at low expression levels. Consistent with previous reports,13 for pathogenic transcript-altering variants, we frequently observed the exact same noncanonical splice junctions in normal individuals (4/7 splicing abnormalities), albeit at much lower levels. This suggests that most pathogenic splicing alterations occur naturally at low levels within the transcriptome and are further induced in the presence of rare genomic variants. Thus, we cannot remove consideration of splicing abnormalities in affected individuals just because the same abnormalities are observed in normal individuals. We propose that a noncanonical junction with minimum four independent reads at a (noncanonical junction coverage/total junction coverage) ratio of 0.2 or greater be used as a metric to differentiate a pathogenic splicing abnormality caused by a DNA variant from natural variation and/or noise. Generating independent libraries and sequencing data from the same or different tissues from the patient or from the carrier parents is particularly useful to augment evidence. For recessive disorders, a confirmatory testing showing that there are no or almost no normal transcripts present by generating full length or fully phased transcripts could be done if RNAseq data is not informative. However, since it is possible that residual normal transcripts can be present and modify the phenotype, RNAseq data must be interpreted in the context of a DNA variant and phenotypic information.
Finally, we note that RNAseq allows the determination of the exact consequence to the mRNA of essential splice site variants, which are otherwise indeterminant. Missense, synonymous, and loss-of-function (LoF) variants within the coding exons can affect splicing or expression level. For example, from exome or genome sequencing analysis, a missense variant can be ruled out despite the consistent phenotype because only LoF variants are reported to be disease-causing for the gene. However, if a variant predicted to be missense is affecting splicing, it is likely pathogenic and if the gene is expressed in one of the accessible tissues, RNAseq would be crucial for the diagnosis. Also, RNAseq can clarify if a LoF variant is undergoing or escaping NMD or if there is evidence of allele-specific expression that could be caused by imprinting, NMD, or a variant in the regulatory region that decreased the expression of one allele. Of the 38 diagnosed cases, 8 of them had 9 pathogenic or likely pathogenic variants within or spanning the essential splice sites or spanning the untranslated region (UTR). Of the nine variants, five (55.6%) were in genes that are expressed in blood and/or fibroblast, allowing us to check the consequence at the transcript level (Supplementary Materials and Methods).
In this heterogeneous cohort of consecutive subjects with undiagnosed, suspected genetic conditions, RNAseq was essential in the evaluation of the potential pathogenic effect of 18% (7 of 38 cases, 95% CI: 8.9–33.7%) of all genomic variants that were ultimately deemed to be pathogenic from trio exome or genome sequencing. Thus, RNAseq data provided key information on the consequence of some DNA variants at the transcript level, leading to an increased diagnostic rate. As next-generation sequencing costs continue to decrease, genome sequencing, augmented with RNAseq, will emerge as a more comprehensive method of genome-wide genetic testing for complex, heterogeneous undiagnosed cases. An area of weakness is the inability to observe many genes of interest due to lack of expression in accessible tissues. Thus, an important parallel effort should be to improve ex vivo transdifferentiation of accessible cells (i.e., skin fibroblasts or blood mononuclear cells) to specific cell types to improve observation of genes more broadly by RNAseq. Further, improvements in the comprehensive search for modification of splicing and expression across the genome are needed to further improve diagnostic sensitivity.
References
Yang Y, Muzny DM, Xia F, et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA. 2014;312:1870–1879.
Lee H, Deignan JL, Dorrani N, et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA. 2014;312:1880–1887.
Skinner D, Raspberry KA, King M. The nuanced negative: meanings of a negative diagnostic result in clinical exome sequencing. Sociol Health Illn. 2016;38:1303–1317.
Biesecker LG, Shianna KV, Mullikin JC. Exome sequencing: the expert view. Genome Biol. 2011;12:128.
Lionel AC, Costain G, Monfared N, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2018;20:435–443.
Stavropoulos DJ, Merico D, Jobling R, et al. Whole genome sequencing expands diagnostic utility and improves clinical management in pediatric medicine. NPJ Genom Med. 2016;1:15012.
Belkadi A, Bolze A, Itan Y, et al. Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants. Proc Natl Acad Sci U S A. 2015;112:5473–5478.
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10:57–63.
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008;40:1413–1415.
Sultan M, Schulz MH, Richard H, et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science. 2008;321:956–960.
Gonorazky HD, Naumenko S, Ramani AK, et al. Expanding the boundaries of RNA sequencing as a diagnostic tool for rare Mendelian disease. Am J Hum Genet. 2019;104:466–483.
Cummings BB, Marshall JL, Tukiainen T, et al. Improving genetic diagnosis in Mendelian disease with transcriptome sequencing. Sci Transl Med. 2017;9:386.
Kremer LS, Bader DM, Mertes C, et al. Genetic diagnosis of Mendelian disorders via RNA sequencing. Nat Commun. 2017;8:15824.
Splinter K, Adams DR, Bacino CA, et al. Effect of genetic diagnosis on patients with previously undiagnosed disease. N Engl J Med. 2018;379:2131–2139.
Splinter K, Hull SC, Holm IA, et al. Implementing the single institutional review board model: lessons from the Undiagnosed Diseases Network. Clin Transl Sci. 2018;11:28–31.
Thorvaldsdottir H, Robinson JT, Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 2013;14:178–192.
Dobin A, Davis CA, Schlesinger F, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
GraphPad. QuickCalcs. https://www.graphpad.com/quickcalcs/confInterval2/. Accessed 29 March 2019.
Agamy O, Ben Zeev B, Lev D, et al. Mutations disrupting selenocysteine formation cause progressive cerebello-cerebral atrophy. Am J Hum Genet. 2010;87:538–544.
Hunt RC, Simhadri VL, Iandoli M, Sauna ZE, Kimchi-Sarfaty C. Exposing synonymous mutations. Trends Genet. 2014;30:308–321.
Stenson PD, Ball EV, Mort M, et al. Human Gene Mutation Database (HGMD): 2003 update. Hum Mutat. 2003;21:577–581.
Stenson PD, Mort M, Ball EV, Shaw K, Phillips A, Cooper DN. The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum Genet. 2014;133:1–9.
Worman HJ, Bonne G. “Laminopathies”: a wide spectrum of human diseases. Exp Cell Res. 2007;313:2121–2133.
Benedetti S, Menditto I, Degano M, et al. Phenotypic clustering of lamin A/C mutations in neuromuscular patients. Neurology. 2007;69:1285–1292.
Abrams AJ, Fontanesi F, Tan NBL, et al. Insights into the genotype-phenotype correlation and molecular function of SLC25A46. Hum Mutat. 2018;39:1995–2007.
Wan J, Steffen J, Yourshaw M, et al. Loss of function of SLC25A46 causes lethal congenital pontocerebellar hypoplasia. Brain. 2016;139:2877–2890.
Clark MM, Stark Z, Farnaes L, et al. Meta-analysis of the diagnostic and clinical utility of genome and exome sequencing and chromosomal microarray in children with suspected genetic diseases. NPJ Genom Med. 2018;3:16.
Fresard L, Smail C, Ferraro NM, et al. Identification of rare-disease genes using blood transcriptome sequencing and large control cohorts. Nat Med. 2019;25:911–919.
Acknowledgements
Supported by awards from the National Institutes of Health (NIH) Common Fund, through the Office of Strategic Coordination and the Office of the NIH Director: U01HG007703 to the University of California–Los Angeles, U01HG007942 to Baylor College of Medicine, and U01HG007943 to HudsonAlpha Institute for Biotechnology. J.D.W. is supported by the UCLA Intercampus Medical Genetics Training Program, US Department of Health and Human Services (USHHS) Ruth L. Kirschstein Institutional National Research Service Award T32GM008243. This research and cores used are supported by NIH National Center for Advancing Translational Science (NCATS) UCLA Clinical and Translational Science Institute (CTSI) grant number UL1TR001881. B.L.F. is supported by NIH R01NS082094. S.N.-R. is supported by the NIH Training Grant in Genomic Analysis and Interpretation T32HG002536.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lee, H., Huang, A.Y., Wang, Lk. et al. Diagnostic utility of transcriptome sequencing for rare Mendelian diseases. Genet Med 22, 490–499 (2020). https://doi.org/10.1038/s41436-019-0672-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0672-1
Keywords
This article is cited by
-
The first case of intellectual disability caused by novel compound heterozygosity for NUDT2 variants
BMC Pediatrics (2024)
-
RNA sequencing and target long-read sequencing reveal an intronic transposon insertion causing aberrant splicing
Journal of Human Genetics (2024)
-
Genomes in clinical care
npj Genomic Medicine (2024)
-
Next-generation sequencing and bioinformatics in rare movement disorders
Nature Reviews Neurology (2024)
-
The expanding diagnostic toolbox for rare genetic diseases
Nature Reviews Genetics (2024)