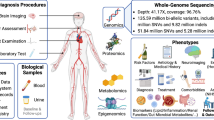
Abstract
This article updates our previous Stroke Gene Panels (SGP) from 2017. Online Mendelian Inheritance in Man and PubMed were searched. We divided detected genes into two SGP groups, SGP1: genes reported in at least one person with stroke and associated with one or more clinical subgroups: large artery atherosclerotic, large artery non-atherosclerotic (tortuosity, dolichoectasia, aneurysm, non-atherosclerotic dissection or occlusion), cerebral small vessel diseases, cardio-embolic (arrhythmia, heart defect, cardiomyopathy), coagulation dysfunctions (venous thrombosis, arterial thrombosis, bleeding tendency), intracerebral hemorrhage, vascular malformations (cavernoma, arteriovenous malformations) and metabolism disorders; and SGP2: genes related to diseases that may predispose to stroke. We identified 168 SGP1 genes, 70 of these were validated for clinical practice. We also detected 72 SGP2 genes. Nine genes were removed because of conflicting evidence. The number of genes increased from 168 to 240 during 4.5-years, reflecting a dynamic evolution and the need for regular updates for research and clinical use.
Similar content being viewed by others
Introduction
Monogenic conditions have an important contribution to stroke risk [1, 2] but they have been difficult to diagnose because of still incomplete knowledge on how monogenic mechanisms are related to disease, relatively expensive diagnostic methods, and because of the heterogenous and multifactorial nature of stroke. Many different monogenic conditions can cause or predispose for stroke [3]. The introduction of massively parallel sequencing methods such as whole exome and whole genome sequencing (WES, WGS) has led to the detection of more and more gene-disease associations [4, 5]. Likewise, WES and WGS have increasingly been used in clinical practice in the workup of patients with stroke where familial aggregation of stroke, the absence of classical risk factors, or young age suggest a high potential for discovering monogenic causes [6, 7]. For these purposes, a panel listing all known stroke genes is a valuable tool in research and clinical management.
The present article updates our previous comprehensive Stroke Gene Panel (SGP) publication [3] which was based on a literature search from August 2017. By using the same systematic methods to compile a new SGP 4.5-years later, we aimed to create an updated panel with all stroke-genes known to date. We also aimed to investigate how fast knowledge develops regarding the level of evidence of monogenic conditions related to stroke and cerebrovascular disease.
Methods
Systematic search
For the present SGP update, we used identical methods to identify genes as in our previously published SGP [3]. The systematic searches in Online Mendelian Inheritance in Man (OMIM) [8] and PubMed databases were conducted until February 2022. Genes reported to be associated with stroke were identified by using combinations of search terms: (stroke), (cerebrovascular), (cerebral OR intracerebral OR intracranial OR brain OR encephalic) AND (infarct OR infarction OR ischemia OR ischaemia), (ischemic OR ischaemic) AND (event OR stroke), (transitory OR transient) AND (event OR ischemic OR ischaemic), or (intracranial OR cerebral OR intracerebral OR encephalic OR brain) AND (haemorrhage OR haemorrhage OR bleeding OR hematoma).
Association with stroke (SGP1) or stroke-predisposing condition (SGP2)
Cerebrovascular conditions for whom the molecular basis of the disorder is known (OMIM phenomap key 3,4) [8] are shown in the updated SGPs. Genes on nuclear DNA containing at least one mutation where a causative role has been shown or postulated were included in the panels. We again compiled two subpanels SGP1 and SGP2. If the gene was reported to cause at least one well-documented stroke case (PubMed), it was included in SGP1. The OMIM search also retrieved genes for diseases that predispose to stroke but where no patients with stroke were reported in the literature; these genes were included in SGP2 when diseases caused by mutations in the gene were documented in at least one patient in publications from PubMed (Fig. 1). In cases where one gene was associated with several clinical phenotypes of stroke where the level of evidence differed, genes already included in SGP1 were not included again in SGP2.
OMIM Online Mendelian Inheritance in Man, SGP1 Stroke Gene Panel 1, SGP2 Stroke Gene Panel 2, LAA large artery atherosclerosis, LAN large artery non-atherosclerotic, SVD small vessel disease, CE cardioembolic, Coag coagulation, MB metabolic, ICH hemorrhage, VM vascular malformation, WMH white matter hyperintensities, BGC basal ganglia calcifications; * the clinical subcategory marked with “*” corresponds to SGP2 while the gene was mentioned in SGP1 because other clinical subcategories fulfill the definition of SGP1. Other characteristics as the presence of vasculitis, basal ganglia calcifications and white matter hyperintensities were evaluated only for the genes in SGP1. While our 2017 gene panel listed 14 genes in the mitochondrial DNA, we have not included genes on the mitochondrial genome in the 2022 update because determining the disease association for single genes on the mitochondrial DNA is very difficult. We recommend including the whole mitochondrial genome in genetic analyses of stroke patients and then evaluate the literature and database information on particular variants.
Each gene included in the SGPs was associated with one or more stroke-subtypes whenever this information could be found in published case reports. We amended the Causative Classification of Stroke (CCS)/Trial of Org 10172 in Acute Stroke Treatment classifications (TOAST) [9, 10] to better reflect pathobiology and thus—presumably—genetics. The following eight major stroke/cerebrovascular disease subtypes were used [3] (# indicates subtypes summarized as “other causes” in CCS) as well a separate category for intracerebral bleeding.
-
1.
large artery atherosclerotic (LAA)
-
2.
large artery nonatherosclerotic (LAN)#
-
3.
small vessel disease (SVD)
-
4.
cardio-embolic (CE- arrhythmia, morphologic cardiac defect, cardiomyopathy)
-
5.
coagulation defects (coag.- arterial thrombosis, venous thrombosis, bleeding)#
-
6.
vascular malformations (VM)#
-
7.
metabolic disorders (MB)#
-
8.
intracerebral bleeding (ICH)#
Additional characteristics of the main 8 categories of stroke in SGP1, regarding LAN-vasculitis, basal ganglia calcifications and white matter hyperintensities were also considered. We did not include age at stroke onset for each gene because the phenotypic variability regarding age at first stroke onset can be substantial.
Clinical versus research use
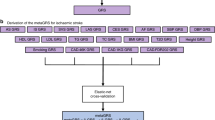
The genes included in SGP1 and SGP2 are not equally well documented to be disease-associated in the literature. We thus further evaluated if the evidence for each gene was sufficient to consider this gene for clinical genetic testing (marked “C”), or if the gene only should be considered in research (“R”). We define clinically useful stroke genes based on clinical and co-segregation criteria, in a way that would correspond to at least a moderate level of supportive evidence in established recommendations for clinical validity of gene-disease associations [11]. We based this evaluation on the number of published families where the clinical phenotype co-segregated with the gene variant (Fig. 2) and used the following criteria for considering genes suitable for clinical screening:
-
For autosomal dominant inheritance: genes where co-segregation of rare or very rare variants (minor allele frequency below 1% in the target population) have been related to disease in either:
-
(a)
two or more unrelated pedigrees, with at least one of the pedigrees containing 10 or more affected individuals, of whom at least 2 had to be third degree or more remote relatives of the proband, or
-
(b)
three or more unrelated smaller pedigrees with at least two affected individuals each.
-
For autosomal recessive inheritance: genes where co-segregation of variants (with a minor allele frequency below 2% in the target population) have been related to disease in:
-
(a)
at least three unrelated pedigrees, with at least two of them containing two or more individuals with the disease.
The pathogenicity of variants identified in these genes need then to be individually evaluated—as suggested in existing guidelines [12] by using existing databases containing regularly updated information on genetic variants.
Genes where conflicting information on disease-causing effect was reported were considered for research purpose only. The stroke-genes were identified through the same methods in 2017 and 2022, allowing a comparison between the number of stroke-genes.
Results
In total, we identified 168 SGP1 genes and 72 SGP2 genes. Compared with our earlier publication [3], 63 new stroke-genes were included in SGP1 (Supplemental Material). Among these, 52 were newly reported genes and 11 were genes previously included in SGP2 but now fulfilling the criteria for inclusion in SGP1 (Fig. 3). These 63 newly included genes in SGP1 were associated with the following phenotypes: 5 (LAA), 21 (LAN), 7 (SVD), 8 (CE), 10 (coagulopathies), 3 (vascular malformations), 11 (metabolic phenotype) and 16 (ICH) (Fig. 1). Re-evaluation showed that for three stroke-genes previously considered suitable for clinical screening, the relevance for stroke had become too inconsistent, and therefore they are now only recommended for research (CACNA1A, MYLK, MFN2), whereas the evidence for 10 other stroke-genes was strengthened and now fulfill our criteria for clinical testing. Twenty-eight new stroke-genes fulfilling the inclusion criteria for SPG2 (Supplemental Material) were identified (Fig. 3). One gene from SGP1 was now placed in SGP2 because of conflicting clinical evidence (FCGR2C) and nine genes from SGP2 were removed from SGP2 because conflicting evidence has emerged (ADIPOQ, CSA, CUL3, HCFC2, KLHL3, NR3C2, SAG, TBX20, THBD).
Discussion
As has been suggested for other disorders, the diagnostic yield of WES and WGS tests of stroke patients may increase if data are regularly re-evaluated and the gene panels are updated [6, 13] and used in combination with recent detailed information on the phenotype.
During the last 4.5 years additional monogenic causes for stroke have been identified at a rapid pace. By using the same search algorithm as in 2017, SGP1 increased by 60% of genes, and 30% more SGP1 stroke-genes now fulfill the criteria for genetic testing in clinical practice. Also, the number of genes in SGP2 increased. However, evolving knowledge also revealed that nine genes (5,4%), that were associated with monogenic stroke based on the evidence available in 2017, now no longer fulfill criteria for our SGPs (Fig. 3).
Our SGP contain clinical information on stroke subtypes. Besides the three established standardized TOAST/CCS subtypes of ischemic stroke [9, 10], we again used five additional subtypes (marked with a # sign in the Methods section) to delineate those “other causes” that frequently occur among the monogenic forms of stroke and that may represent distinct molecular and pathogenic mechanisms. Furthermore, in SGP1 we now also systematically included information on three other associated characteristics (large artery vasculitis, white matter hyperintensities, bilateral basal ganglia calcifications) to facilitate the correct interpretation of a genetic variant found in a stroke patient or family. Given the large spectrum of possible stroke mechanisms, an accurate matching between the stroke phenotype in the patient/family under investigation with the phenotype described in patients with pathogenic variants in the same gene increases the likelihood that the identified variant is truly disease-causing. As WES and WGS examine all genes simultaneously, false-positive “chance” findings are possible. Misinterpretation of such findings can be minimized when only considering genes for which the known clinical phenotype corresponds to the one in the patient under investigation [6, 14].
We compiled our panel by using a systematic and highly replicable approach that allowed us to accurately compare the number of stroke genes in 2017 with 2022. We are aware that this approach has missed genes that fulfill the criteria for SGP1 or SGP2 but that were not retrieved by our methodology. This includes some of the genes for moyamoya phenomenon, other vascular malformations, abnormalities of coagulation including CBL [15], DIAPH1 [16, 17], CHD4, CNOT3, and SETD5 [16]. This inherent difficulty in compiling gene panels is well known. Furthermore, too extensive panels may increase the yield of variants that are not relevant to the disease phenotype [18, 19].
While SGPs offer an evidence-based list of stroke-genes with specified level of evidence as clinical or research only, they do not offer specific guidance for variant interpretation. Complementary resources and available expert knowledge are needed to support clinicians in interpreting the variant pathogenicity [12] and the level of actionability [20].
Data availability
Data generated during this study can be found within the published article and its supplementary files.
References
Ilinca A, Kristoffersson U, Soller M, Lindgren AG. Familial aggregation of stroke amongst young patients in Lund Stroke Register. Eur J Neurol. 2016;23:401–7.
Jood K, Ladenvall C, Rosengren A, Blomstrand C, Jern C. Family history in ischemic stroke before 70 years of age: the Sahlgrenska Academy Study on Ischemic Stroke. Stroke 2005;36:1383–7.
Ilinca A, Samuelsson S, Piccinelli P, Soller M, Kristoffersson U, Lindgren AG. A stroke gene panel for whole-exome sequencing. Eur J Hum Genet. 2019;27:317–24.
Gorcenco S, Ilinca A, Almasoudi W, Kafantari E, Lindgren AG, Puschmann A. New generation genetic testing entering the clinic. Parkinsonism Relat Disord. 2020;73:72–84.
Fang F, Xu Z, Suo Y, Wang H, Cheng S, Li H, et al. Gene panel for Mendelian strokes. Stroke Vasc Neurol. 2020;5:416–21.
Ilinca A, Martinez-Majander N, Samuelsson S, Piccinelli P, Truvé K, Cole J, et al. Whole-Exome sequencing in 22 Young Ischemic stroke patients with familial clustering of stroke. Stroke 2020;51:1056–63.
Ilinca A, Englund E, Samuelsson S, Truvé K, Kafantari E, Martinez-Majander N, et al. MAP3K6 mutations in a neurovascular disease causing stroke, cognitive impairment, and tremor. Neurol Genet. 2021;7:e548.
McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University (Baltimore, MD). Online Mendelian Inheritance in Man, 2022. https://omim.org/.
Adams HP Jr., Bendixen BH, Kappelle LJ, Biller J, Love BB, Gordon DL et al. Classification of subtype of acute ischemic stroke. Definitions for use in a multicenter clinical trial. TOAST. Trial of Org 10172 in Acute Stroke Treatment. Stroke; 24:35–41.
Ay H, Benner T, Arsava EM, Furie KL, Singhal AB, Jensen MB, et al. A computerized algorithm for etiologic classification of ischemic stroke: The Causative Classification of Stroke system. Stroke 2007;38:2979–84.
Strande NT, Riggs ER, Buchanan AH, Ceyhan-Birsoy O, DiStefano M, Dwight SS, et al. Evaluating the clinical validity of gene-disease associations: an evidence-based framework developed by the clinical genome resource. Am J Hum Genet. 2017;100:895–906.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Won D, Kim SH, Kim B, Lee ST, Kang HC, Choi JR. Reanalysis of genomic sequencing results in a clinical laboratory: advantages and limitations. Front Neurol. 2020;11:612.
Matalonga L, Hernández-Ferrer C, Piscia D, Solve-RD SNV-indel working group, Schüle R, Synofzik M, et al. Solving patients with rare diseases through programmatic reanalysis of genome-phenome data. Eur J Hum Genet. 2021;29:1337–47.
Gannamani R, van der Veen S, van Egmond M, de Koning TJ, Tijssen MAJ. Challenges in clinicogenetic correlations: one phenotype - many genes. Mov Disord Clin Pr. 2021;8:311–21.
Hyakuna N, Muramatsu H, Higa T, Chinen Y, Wang X, Kojima S. Germline mutation of CBL is associated with moyamoya disease in a child with juvenile myelomonocytic leukemia and Noonan syndrome-like disorder. Pediatr Blood Cancer. 2015;62:542–4.
Kundishora AJ, Peters ST, Pinard A, Duran D, Panchagnula S, Barak T, et al. DIAPH1 variants in Non-East Asian patients with sporadic moyamoya disease. JAMA Neurol. 2021;78:993–1003.
Pinard A, Guey S, Guo D, Cecchi AC, Kharas N, Wallace S, et al. The pleiotropy associated with de novo variants in CHD4, CNOT3, and SETD5 extends to moyamoya angiopathy. Genet Med. 2020;22:427–31.
Angione K, Gibbons M, Demarest S. An objective method for evaluating next-generation sequencing panels. J Child Neurol. 2019;34:139–43.
Webber EM, Hunter JE, Biesecker LG, Buchanan AH, Clarke EV, Currey E, et al. ClinGen Resource. Evidence-based assessments of clinical actionability in the context of secondary findings: Updates from ClinGen’s Actionability Working Group. Hum Mutat. 2018;39:1677–85. https://search.clinicalgenome.org/kb/genes/HGNC:3603.
Funding
Disclosures This work was supported by grants from The Crafoord Foundation, The Segerfalk Foundation, Anna Lisa and Sven-Eric Lundgren’s foundation for medical research, Swedish Government (under the “Avtal om Läkarutbildning och Medicinsk Forskning”, ALF), Royal Physiographic Society of Lund. Multipark—a strategic research Environment at Lund University, The Swedish Research Council (2019–01757), The Swedish Heart and Lung Foundation, Region Skåne, Lund University, Skåne University Hospital, Sparbanksstiftelsen Färs och Frosta, Fremasons Lodge of Instruction Eos in Lund, and Bundy Academy foundation for medical research. Arne Lindgren is also supported by the CaNVAS project funded by NIH (1R01NS114045-01). Open access funding provided by Lund University.
Author information
Authors and Affiliations
Contributions
AI conceptualized the study, performed the database searches, collected and analyzed data, and wrote the manuscript; AP contributed to write the manuscript; AGL provided feedback on the report and obtained funding. All authors discussed the intellectual content and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Ethical approval was not necessary for this study based on information in existing databases and published reports.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ilinca, A., Puschmann, A., Putaala, J. et al. Updated Stroke Gene Panels: Rapid evolution of knowledge on monogenic causes of stroke. Eur J Hum Genet 31, 239–242 (2023). https://doi.org/10.1038/s41431-022-01207-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-022-01207-6
This article is cited by
-
A novel NODAL variant in a young embolic stroke patient with visceral heterotaxy
BMC Neurology (2024)
-
Genomic analysis of presumed perinatal stroke in Saudi Arabia reveals a strong monogenic contribution
Human Genetics (2024)
-
The value of exomes across the ages
European Journal of Human Genetics (2023)