Abstract
Orkney and Shetland, the population isolates that make up the Northern Isles of Scotland, are of particular interest to multiple sclerosis (MS) research. While MS prevalence is high in Scotland, Orkney has the highest global prevalence, higher than more northerly Shetland. Many hypotheses for the excess of MS cases in Orkney have been investigated, including vitamin D deficiency and homozygosity: neither was found to cause the high prevalence of MS. It is possible that this excess prevalence may be explained through unique genetics. We used polygenic risk scores (PRS) to look at the contribution of common risk variants to MS. Analyses were conducted using ORCADES (97/2118 cases/controls), VIKING (15/2000 cases/controls) and Generation Scotland (30/8708 cases/controls) data sets. However, no evidence of a difference in MS-associated common variant frequencies was found between the three control populations, aside from HLA-DRB1*15:01 tag SNP rs9271069. This SNP had a significantly higher risk allele frequency in Orkney (0.23, p value = 8 × 10–13) and Shetland (0.21, p value = 2.3 × 10–6) than mainland Scotland (0.17). This difference in frequency is estimated to account for 6 (95% CI 3, 8) out of 150 observed excess cases per 100,000 individuals in Shetland and 9 (95% CI 8, 11) of the observed 257 excess cases per 100,000 individuals in Orkney, compared with mainland Scotland. Common variants therefore appear to account for little of the excess burden of MS in the Northern Isles of Scotland.
Similar content being viewed by others
Introduction
Multiple sclerosis (MS) is the most common neurological disability found in young adults in the Western world [1]. The disease is characterised by the inflammation and chronic degeneration of the central nervous system, a result of the destruction of the myelin sheath by the individual’s immune system [2]. There is no definitive explanation as to the reason for these immune attacks, with genetic susceptibility and environmental factors both contributing to MS risk [3, 4]. Although environmental factors (for example, vitamin D, smoking, Epstein–Barr viral exposure and body mass index (BMI)) are important for MS risk [5,6,7,8], MS has a material genetic component: broad-sense heritability has been estimated using twin, sibling and half-sibling data as 0.64 (with a 95% confidence interval (CI) of 0.36–0.76) [9]. Single-nucleotide polymorphism (SNP) heritability has been estimated at 0.19 (95% CI 0.18, 0.20) [10]. MS arises most frequently in genetically susceptible individuals who may have been exposed to risk-associated environmental factors or stochastic events [11]. A total of 233 genetic variants have been identified as associated with MS susceptibility, including 32 independent genetic effects within the major histocompatibility (MHC) region on chromosome 6 [10]. The majority of these variants have an odds ratio (OR) ranging from 1.05 to 1.20; however, the strongest association identified is with the HLA-DRB1*15:01 variant, with an OR of 2.92 [10, 12].
MS in the Northern Isles
The highest prevalence of MS in the world is found in the Orkney islands in the North of Scotland, where prevalence reaches 402 individuals per 100,000 [13]. The Shetland islands, 50 miles north of Orkney, have a similarly high rate of 295 individuals per 100,000 [13]. Mainland Scotland, for comparison, has a prevalence of 145 individuals per 100,000 [13]. The prevalence of MS within these island groups, in particular Orkney, is significantly higher than what would be expected for a population of that latitude. A number of previous studies have investigated the potential cause of this excess of MS prevalence. Vitamin D deficiency as a potential cause of MS was investigated by Weiss et al. in 2016; however, plasma 25-hydroxyvitamin D was found to be significantly higher in those on Orkney than in mainland Scotland (mean 35.3 compared to 31.7 nmol/L). Additionally, Orkney had a lower prevalence of severe plasma 25-hydroxyvitamin D deficiency (of 6.6% compared to 16.2% in mainland Scotland) [14]. Another study investigated homozygosity, the inheritance of identical haplotypes from both parents, in Orkney and Shetland [15]. Three measures of genome-wide homozygosity were generated for 88 MS patients and 178 matched controls and assessed for association with MS. However, no association was detected, and so consanguinity is not thought to be the principal cause of the excess MS prevalence. However, it is possible that this excess prevalence may be explained genetically through the Northern Isles having a higher proportion of common risk alleles. If the ancestors or founders of the islands by chance had higher frequencies of these risk alleles, this may cause additional cases of MS as they would be segregating at higher frequencies today. This idea was first suggested in 1981 by Compston, who implied that Orcadians in general may have higher frequencies of common risk variants [16].
Methods
Study participants
This research used 97 MS cases and 2118 controls from the Orkney Complex Disease Study and the Northern Isles Multiple Sclerosis Study (NIMS; collectively referred to as ORCADES) as a sample of the Orkney population, 15 MS cases and 2090 controls from the Viking Health Study Shetland (VIKING) as a sample of the Shetland population and 30 MS cases and 8708 controls from Generation Scotland (GS) as a sample of the mainland Scotland population. ORCADES, VIKING and GS are cross-sectional, family-based cohorts that were established to become platform resources for the study of complex disease in Scotland, while the NIMS was established with the specific aim of studying MS. Data collection and genotyping for ORCADES, VIKING and GS has been fully described in previous research papers but has been summarised along with genotype and sample quality control (QC) steps in Supplementary Table 1 [17,18,19]. A principal component plot of the three cohorts can also be found in Supplementary Fig. 1.
Selecting common risk variants for polygenic risk score (PRS) calculation
A total of 127 key MS SNPs (Supplementary Table 2) were compiled for PRS calculation, selected from the 2011 International Multiple Sclerosis Genetics Consortium (IMSGC) genome-wide association study (GWAS) [20] and the GWAS Catalogue [21,22,23,24,25,26,27,28,29,30]. The SNPs selected from the 2011 IMSGC GWAS comprised a group of 102 SNPs that were taken forward for replication analysis in the original study. The 2011 IMSGC GWAS was chosen in particular as this was the most recent large-scale GWAS that the NIMS cohort did not contribute towards. SNPs were included from the GWAS Catalogue if the disease trait was listed as “multiple sclerosis” and the SNPs originated from studies including only European individuals. Additionally, only studies up to 2012 were considered. Any SNPs originating from the 2011 IMSGC GWAS, or from a study that included any individuals from Orkney, were excluded from the GWAS Catalogue search. Only SNPs with a p value <1 × 10−3 were considered, and SNPs were only included if they were present in all three cohorts. Strict QC procedures were applied to ensure that PRS results produced from these SNPs were not biased or inaccurate and that subjects from the Northern Isles had not contributed to the underlying GWAS. These QC methods included removing SNPs without a reported OR or risk allele, using inverse variance meta-analyses to determine ORs and p values for duplicated SNPs, and clumping SNPs for linkage disequilibrium, using a cut-off threshold of r2 = 0.25 within a 200-kb window.
Calculating PRSs
PRSs were calculated for all individuals using the R package PRSice (v1.25), which used the HRC-imputed genotype dosage data for ORCADES, VIKING and GS as the target SNP set, along with SNP effect sizes from the original source (either the GWAS Catalogue or 2011 IMSGC GWAS). Risk scores were produced for the full SNP set (n = 127), the SNP set without HLA-DRB1*15:01 tag SNP rs9271069 (n = 126) and for rs9271069 alone.
Differentiating MS cases and controls
PRS were compared between cases and controls in each data set. The three data sets were first standardised by z-scoring the PRS to allow comparison between populations. Cases and controls were then compared within each data set by fitting a generalised linear model with Gaussian errors and an identity link function, using the R function glm. Age, sex and the first two principal components were included as covariates for all three populations. A meta-analysis for cases and a meta-analysis for controls were performed to determine an estimate for the overall case/control PRS.
Fitting a logistic regression model
Related individuals were removed before fitting the data to a logistic regression model using a genomic kinship coefficient threshold of 0.05. Following removal of related individuals, cases and control numbers in each data set were as follows: 80/645 ORCADES individuals, 14/642 VIKING individuals, and 29/ 8341 GS individuals. A logistic regression model was fitted separately to each data set using the R function glm, with MS disease status as the dependent variable, PRS as the independent variable and age, sex, principal component 1 and principal component 2 included as covariates. A null model (with only covariates) was also fitted to each data set.
Determining how much MS variance is explained by common risk variants
Logistic regression model results were used to assess how much variance in MS risk common risk variants could explain. Nagelkerke’s pseudo R2 value was calculated in R for each data set and SNP group (full SNP set, SNP set without rs9271069 and rs9271069 alone) using the model results.
Determining how successful common risk variants are in predicting MS
To determine whether common risk variants could be a predictor of an individual’s MS status, the model results were used to calculate a receiver operator characteristic curve (ROC) and the area under this curve (AUC). ROCs were plotted for each fitted model to assess how well the PRS with covariates predicted MS disease status. The AUC was also calculated to quantify the predictive ability of each model.
Comparing common variants between Orkney, Shetland and mainland Scotland
To assess the difference in common risk variants between each population, mean PRS between control individuals were compared between GS, ORCADES and VIKING using two-sample t tests. The frequency of each PRS SNP was calculated in individuals without MS and compared between GS, ORCADES and VIKING using a Pearson’s chi-squared test.
Determining how much risk is explained by common risk variants
An important aspect of this study was to estimate the contribution of these variants to the excess MS prevalence in the Northern Isles. This was done by determining the expected difference in prevalence due to the differences in frequencies of these variants between data sets. The expected difference in prevalence was then compared to the observed difference in prevalence seen in mainland Scotland, Orkney and Shetland. First, the difference in means between (a) GS and VIKING and (b) GS and ORCADES was calculated. Only control individuals were included in this calculation. The differences in means were then multiplied by the beta value produced from the meta-analysis of each of the data set’s models with covariates, to give the expected odds values. The expected odds values produced is the proportion of the odds that could be attributed to difference in mean PRS between populations. Thus, given the frequencies of the common risk variants that have been looked at in the PRS, these values would be the expected increase in MS risk for the Northern Isles populations, using GS (Glasgow/Dundee) as the baseline. This would reflect the genetic difference due to common risk variants. To determine how this expected difference in MS risk explained by common risk variants compared to the observed MS risk between populations, the expected differences in odds values were compared to the observed difference in odds values. Observed MS prevalence data were obtained from Visser et al. [13] using Aberdeen City for Mainland Scotland. ORs were calculated using contingency tables and converted into log of ORs for comparison with the expected values.
Results
Data summary
The demographic characteristics for study individuals can be found in Table 1.
Comparing PRS in MS cases and controls
MS cases have statistically significantly higher polygenic scores than MS controls using the full SNP set in each population (Fig. 1 and Supplementary Table 3, with p values: ORCADES, 5.63 × 10–9; GS, 7.6 × 10–4; VIKING, 2.82 × 10–2), validating the use of PRS. When rs9271069 is removed, only cases in ORCADES (p value = 1.8 × 10–5) and GS (p value = 5.67 × 10–3) are statistically different. When rs9271069 is considered alone, only ORCADES cases and controls are statistically different (p value = 1.02 × 10–5), with slight significance seen in GS (p value = 0.045). This pattern of significance reflects the number of cases found in each data set: ORCADES (significant in all three comparisons) has 97 cases, GS (significant in two comparisons) has 30 cases, and VIKING (significant in the main comparison only) has 15 cases.
How much variance PRS explain and their ability to predict MS
To determine how much MS variance is explained by common risk variants, a logistic regression model was fitted separately to each data set, with MS status as the dependent variable, PRS as the independent variable, and age, sex, PC 1 and PC2 as covariates. Nagelkerke’s pseudo R2 and AUC values were calculated using the model results to determine how much variance the risk scores explained and how successful they were in predicting MS.
The full model results, including R2 and AUC values, can be found in Table 2. The models fitted using the full SNP set (n = 127) with covariates resulted in the highest R2 values in each population (VIKING R2 = 0.075, ORCADES R2 = 0.070, GS R2 = 0.069). In comparison, the null model (containing only covariates) for each population resulted in an R2 value of 0.045 for VIKING, 0.019 for ORCADES and 0.038 for GS.
The models fitted using the full SNP set (n = 127) with covariates also resulted in the highest AUC values for each population: GS AUC = 0.77 (95% CI 0.68–0.85), VIKING AUC = 0.76 (0.65–0.87) and ORCADES AUC = 0.70 (0.65–0.76). In comparison, the null model (containing only covariates) for each population resulted in AUC values of 0.71 (0.61–0.81) for GS, 0.70 (0.59–0.81) for VIKING and 0.60 (0.54–0.66) for ORCADES. The ROC curves for each population and PRS can be found in Supplementary Fig. 1.
Do MS common risk variants differ in frequency between mainland Scotland and the Northern Isles?
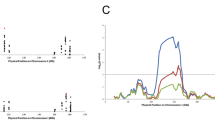
When we compared the control populations from each data set, a statistically significant difference between GS and both ORCADES and VIKING was seen for the full SNP set (Table 3). When rs9271069 was removed from the SNP set, there was no significant difference detectable between any of the control populations. When rs9271069 was looked at alone, a statistical difference was seen when comparing all populations. Supplementary Table 2 provides the risk allele frequency (RAF) of each PRS SNP, calculated using control individuals within each population, along with Pearson’s chi-squared results comparing RAF between populations. The SNP with the highest associated MS risk, HLA-DRB1*15:01 tag SNP rs9271069 (OR = 2.77), had a significantly higher frequency in Orkney controls (RAF = 0.23) and Shetland controls (RAF = 0.21) than mainland Scotland controls (RAF = 0.17: respective p values of 8 × 10–13 and 2.3 × 10–6).
How much of the excess MS cases in the Northern Isles is caused by common risk variants?
To determine the contribution of common risk variants to excess MS cases in the Northern Isles, a comparison was made between the calculated expected odds and the observed odds seen from MS prevalence data (Table 4). In Shetland, common risk variants account for 9 (95% CI 5, 14) out of 150 observed excess cases per 100,000 individuals. The majority of the expected odds is from the HLA-DRB1*15:01 tag SNP rs9271069, which contributes an equivalent of 6 cases (95% CI 3, 8) per 100,000 individuals. In Orkney, all the expected excess genetic odds are due to HLA-DRB1*15:01 tag SNP rs9271069, which accounts for 9 cases (95% CI 8, 11) of the observed 257 excess cases per 100,000 individuals. Considering the populations of Orkney (22,000) and Shetland (23,000), the differences in allele frequencies of common variants can thus only explain the equivalent of about two excess cases of MS in each archipelago.
Study limitations
A major limitation of this study was the use of the IMSGC 2011 GWAS as a key contributor to the SNP data on which the PRS calculation was based. It was preferable to use the most recent, large-scale GWAS for this study; however, the IMSGC 2011 GWAS is the most recent GWAS that did not include data generated from individuals in the NIMS. While using an older GWAS may result in some loss of power, we have included the most important common susceptibility alleles within the PRS calculation in this study. We also recognise that this PRS uses a limited number of strong-effect SNPs selected from multiple sources; while this does not bias our results, there are many ways to construct a PRS and this score could have been made in other ways by other researchers at other times.
The sample size was a further limitation to this study. Orkney and Shetland are both relatively small populations (approximately 22,000 individuals in Orkney and 23,000 individuals in Shetland) and will therefore yield small numbers of MS cases, despite the high rates of MS in the islands. However, the key analyses within this study were to determine: (i) whether common risk variants differed between the Northern Isles and mainland Scotland and (ii) how these variants, as a group, contribute to the excess burden of MS. Both of these analyses were conducted using only control individuals and so were not affected by the small number of cases.
Orkney and Shetland remain of key interest to MS: while it is not possible to increase the case numbers within ORCADES and VIKING, it was important to utilise these cohorts given the unique genetics of these population isolates.
Future work
The PRS used in this study included the strongest effect allele from the MHC region, HLA-DRB1*15:01. The MHC region is complex and includes other high-risk variants (which may be in linkage disequilibrium with the SNP included in this study). It is suggested that a future study using PRS for MS focus on the MHC region specifically.
Discussion
We found that common risk variants do not make a material contribution towards the higher rates of MS in the Northern Isles of Scotland. However, a small proportion of excess risk can be attributed to a tag SNP for HLA-DRB1*15:01, the major single genetic risk factor for MS (OR = 2.92). This SNP has a significantly higher frequency in the Northern Isles, particularly Orkney (RAF = 0.23), than mainland Scotland (RAF = 0.17, p value = 8 × 10–13). Frequencies for the tag SNP for HLA-DRB1*15:01 reported in other cohorts appears to be similar to that in mainland Scotland or lower: the ALFA Allele Frequency project reports the frequency within European populations as 0.14 (sample size 33,120) [31]. It is possible that the frequency of this allele is higher in the Northern Isles populations due to the founders of Orkney and Shetland having higher frequencies of the risk allele or it may have risen through genetic drift in the past thousand years. Regardless, the significantly higher frequency of this SNP in the Northern Isles in comparison to both mainland Scotland and the general European population suggests that it plays a modest role in the prevalence of MS in the Northern Isles, even if not the excess prevalence. We found that PRSs calculated with the 127 most strongly associated MS risk variants explained approximately 3–5% of variance. The 2019 IMSGC study, which analysed data from 47,429 MS cases and 68,374 control subjects, estimated the heritability attributable to all analysed common genetic variants at 19.2% [10]. Of this variance, 18% was explained by genome-wide significant variants. Therefore, the heritability of genome-wide significant variants was estimated at approximately 3.5%, which aligns with the estimate from this study. Additionally, the variance within this study is largely dominated by the increased frequency of the HLA-DRB1*15:01 SNP in ORCADES, offsetting any gain from including the additional SNPs with small (and uncertain) effects. The predictive capacity of the PRSs for MS status was calculated with an AUC of 0.77 (95% CI 0.68–0.85) in GS, 0.76 (0.65–0.87) in VIKING and 0.70 (0.65–0.76) in ORCADES. This is in line with the scores previously published in literature. A 2016 study using 452 MS cases with 103 common risk variants estimated the AUC to be 0.72 (95% CI 0.69, 0.75), which overlaps with the results produced here [32]. MS is a heterogenous disease, and while it is useful to see to what degree genetic risk scores could aid in prediction, additional, especially rare, variants or the incorporation of environmental factors into prediction models will be required to improve risk prediction. The cause of the excess MS prevalence in the Northern Isles remains mostly unexplained and is still a case for investigation: it can likely be attributed to several factors. One hypothesis is that a number of rare susceptibility variants of large effect are segregating in the islands. Much evidence has been presented in recent years to support the influence of rare variants in MS risk, particularly among family groups [33]. Within the Northern Isles, it is possible that there are one or more rare variants, drifted to higher frequency due to genetic simplification, possibly segregating within families, that are associated with MS. Smaller contributions will also be made from the high rates of overweight and obesity found in the islands. Orkney has the highest percentage of individuals who are classed as overweight and obese for any location in Scotland at 73% (95% CI 68, 78), while Shetland stands at 71% (95% CI 64, 77). For comparison, the Scottish average is 65% (95% CI 64, 66) [34]. Obesity is a known risk factor for MS; a Mendelian Randomisation study showed that an increase of 1 standard deviation in BMI increased the odds of developing MS by 41% (95% CI 20, 66) [8]. Additionally, interactive effects have been shown between HLA-DRB1*15:01 and obesity: individuals classed as obese who carry HLA-DRB1*15 but not HLA-A*02 have an OR of 16.2 (95% CI 7.5, 35.2) for developing MS [35]. Given that there are most likely multiple contributors to the excess rate of MS in the Northern Isles, there are numerous avenues for future research. The most obvious of these appears to be a targeted search for rare variants. Whole-genome sequence analysis has detected many Shetland-specific rare variants [36]. The similar strong differentiation of Orkney in genome-wide population genetic analyses [37] predicts a similar situation there; however, power will always be limited by the low absolute numbers of cases, overall and in any particular kindred. The future of MS research in the Northern Isles and beyond will lie in new methods to discover and explore the function of susceptibility variants and their interactions with their unique environment. GWAS summary statistics used to create the PRS and the raw PRS for each study population can be found within the Edinburgh DataShare (https://doi.org/10.7488/ds/2992).
References
Tremlett H, Rieckmann P. New perspectives in the natural history of multiple sclerosis. Neurology. 2010;74:2004–15.
Hauser SL, Oksenberg JR. The neurobiology of multiple sclerosis: genes, inflammation, and neurodegeneration. Neuron. 2006;52:61–76.
Dyment DA, Ebers GGC, Sadovnick AD. Genetics of multiple sclerosis. Lancet Neurol. 2004;3:104–10.
Marrie RA. Environmental risk factors in multiple sclerosis aetiology. Lancet Neurol. 2004;3:709–18.
Mokry LE, Ross S, Ahmad OS, Forgetta V, Smith GD, Leong A, et al. Vitamin D and risk of multiple sclerosis: a Mendelian Randomization study. PLoS Med. 2015;12:e1001866.
Riise T, Nortvedt MW, Ascherio A. Smoking is a risk factor for multiple sclerosis. Neurology. 2003;61:1122–4.
Holmøy T, Hestvik ALK. Multiple sclerosis: immunopathogenesis and controversies in defining the cause. Curr Opin Infect Dis. 2008;21:271–8.
Mokry LE, Ross S, Timpson NJ, Sawcer S, Smith GD, Richards JB. Obesity and multiple sclerosis: a Mendelian Randomization study. PLoS Med. 2016;13:e1002053.
Westerlind H, Ramanujam R, Uvehag D, Kuja-Halkola R, Boman M, Bottai M, et al. Modest familial risks for multiple sclerosis: a registry-based study of the population of Sweden. Brain. 2014;137:770.
International Multiple Sclerosis Genetics Consortium. Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science. 2019;365:eaav7188.
Goodin DS. The causal cascade to multiple sclerosis: a model for MS pathogenesis. PLoS One. 2009;4:e4565.
Patsopoulos NA, Barcellos LF, Hintzen RQ, Schaefer C, van Duijn CM, Noble JA, et al. Fine-mapping the genetic association of the major histocompatibility complex in multiple sclerosis: HLA and non-HLA effects. PLoS Genet. 2013;9:e1003926.
Visser EM, Wilde K, Wilson JF, Yong KK, Counsell CE. A new prevalence study of multiple sclerosis in Orkney, Shetland and Aberdeen city. J Neurol Neurosurg Psychiatry. 2012;83:719–24.
Weiss E, Zgaga L, Read S, Wild S, Dunlop MG, Campbell H, et al. Farming, foreign holidays, and vitamin D in Orkney. PLoS ONE. 2016;11:e0155633.
McWhirter RE, McQuillan R, Visser E, Counsell C, Wilson JF. Genome-wide homozygosity and multiple sclerosis in Orkney and Shetland Islanders. Eur J Hum Genet. 2012;20:198–202.
Compston A. Multiple sclerosis in the Orkneys. Lancet. 1981;318:98.
Smith BH, Campbell A, Linksted P, Fitzpatrick B, Jackson C, Kerr SM, et al. Cohort Profile: Generation Scotland: Scottish Family Health Study (GS:SFHS). The study, its participants and their potential for genetic research on health and illness. Int J Epidemiol. 2013;42:689–700.
McQuillan R, Leutenegger A-L, Abdel-Rahman R, Franklin CS, Pericic M, Barac-Lauc L, et al. Runs of homozygosity in European populations. Am J Hum Genet. 2008;83:359–72.
Kerr SM, Klaric L, Halachev M, Hayward C, Boutin TS, Meynert AM, et al. An actionable KCNH2 long QT syndrome variant detected by sequence and haplotype analysis in a population research cohort. Sci Rep. 2019;9:1–11.
Sawcer S, Hellenthal G, Pirinen M, Spencer CCA, Patsopoulos NA, Moutsianas L, et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature. 2011;476:214–9.
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H, et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014;42:D1001–6.
Hafler DA, Compston A, Sawcer S, Lander ES, Daly MJ, De Jager PL, et al. Risk alleles for multiple sclerosis identified by a genomewide study. N. Engl J Med. 2007;357:851–62.
Aulchenko YS, Hoppenbrouwers IA, Ramagopalan SV, Broer L, Jafari N, Hillert J, et al. Genetic variation in the KIF1B locus influences susceptibility to multiple sclerosis. Nat Genet. 2008;40:1402–3.
De Jager PL, Jia X, Wang J, de Bakker PIW, Ottoboni L, Aggarwal NT, et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat Genet. 2009;41:776–82.
Sanna S, Pitzalis M, Zoledziewska M, Zara I, Sidore C, Murru R, et al. Variants within the immunoregulatory CBLB gene are associated with multiple sclerosis. Nat Genet. 2010;42:495–7.
Nischwitz S, Cepok S, Kroner A, Wolf C, Knop M, Müller-Sarnowski F, et al. Evidence for VAV2 and ZNF433 as susceptibility genes for multiple sclerosis. J Neuroimmunol. 2010;227:162–6.
Briggs FBS, Shao X, Goldstein BA, Oksenberg JR, Barcellos LF, De Jager PL. Genome-wide association study of severity in multiple sclerosis. Genes Immun. 2011;12:615–25.
Patsopoulos NA, de Bakker PIW, Steering Committees of Studies Evaluating IFNβ-1b and a CCR1-Antagonist, ANZgene Consortium, GeneMSA, International Multiple Sclerosis Genetics Consortium, et al. Genome-wide meta-analysis identifies novel multiple sclerosis susceptibility loci. Ann Neurol. 2011;70:897–912.
Matesanz F, González-Pérez A, Lucas M, Sanna S, Gayán J, Urcelay E, et al. Genome-wide association study of multiple sclerosis confirms a novel locus at 5p13.1. PLoS One. 2012;7:e36140.
Bahlo M, Booth DR, Broadley SA, Brown MA, Foote SJ, Griffiths LR, et al. Genome-wide association study identifies new multiple sclerosis susceptibility loci on chromosomes 12 and 20. Nat Genet. 2009;41:824–8.
Phan L, Jin Y, Zhang H, Quiang W, Shekhtman E, Shao D, et al. ALFA: allele frequency aggregator. 2020. https://www.ncbi.nlm.nih.gov/snp/docs/gsr/alfa/.
Mescheriakova JY, Broer L, Wahedi S, Uitterlinden AG, van Duijn CM, Hintzen RQ. Burden of genetic risk variants in multiple sclerosis families in the Netherlands. Mult Scler J Exp Transl Clin. 2016;2:2055217316648721.
Harding KE, Robertson NP. New rare genetic variants in multiple sclerosis. J Neurol. 2019;266:278–80.
Scottish Government. Scottish health survey: results for local areas 2014 to 2017. 2018. https://www.gov.scot/publications/scottish-health-survey-results-local-areas-2014-2015-2016-2017/.
Hedström AK, Bomfim IL, Barcellos L, Gianfrancesco M, Schaefer C, Kockum I, et al. Interaction between adolescent obesity and HLA risk genes in the etiology of multiple sclerosis. Neurology. 2014;82:865–72.
Halachev M, Meynert A, Taylor MS, Vitart V, Kerr SM, Klaric L, et al. Increased ultra-rare variant load in an isolated Scottish population impacts exonic and regulatory regions. PLoS Genet. 2019;15:e1008480.
Gilbert E, O’Reilly S, Merrigan M, McGettigan D, Vitart V, Joshi PK, et al. The genetic landscape of Scotland and the Isles. Proc Natl Acad Sci USA. 2019;116:19064–70.
Acknowledgements
We would like to acknowledge the invaluable contributions of the research nurses in Orkney and Shetland, the administrative team in Edinburgh and the people of Orkney and Shetland and all individuals involved in Generation Scotland.
Funding
This work was supported by the Shetland and Orkney Multiple Sclerosis Research Project and the Medical Research Council. The Orkney Complex Disease Study (ORCADES) was supported by the Chief Scientist Office of the Scottish Government (CZB/4/276, CZB/4/710), a Royal Society URF to J.F.W., the MRC Human Genetics Unit quinquennial programme “QTL in Health and Disease”, Arthritis Research UK and the European Union framework programme 6 EUROSPAN project (contract no. LSHG-CT-2006–018947). The Viking Health Study – Shetland (VIKING) was supported by the MRC University Unit Programme Grant MC_UU_00007/10 “QTL in Health and Disease”. The Northern Isles Multiple Sclerosis Study (NIMS) was funded by the Multiple Sclerosis Society (UK) and the Volant Trust. Generation Scotland received core funding from the Chief Scientist Office of the Scottish Government Health Directorates (CZD/16/6) and the Scottish Funding Council (HR03006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barnes, C.L.K., Hayward, C., Porteous, D.J. et al. Contribution of common risk variants to multiple sclerosis in Orkney and Shetland. Eur J Hum Genet 29, 1701–1709 (2021). https://doi.org/10.1038/s41431-021-00914-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-021-00914-w
This article is cited by
-
Polygenic risk scores for autoimmune related diseases are significantly different in cancer exceptional responders
npj Precision Oncology (2024)
-
Epigenetic and proteomic signatures associate with clonal hematopoiesis expansion rate
Nature Aging (2024)
-
Investigating the role of common and rare variants in multiplex multiple sclerosis families reveals an increased burden of common risk variation
Scientific Reports (2022)