Abstract
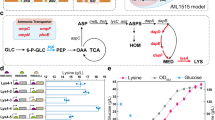
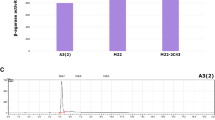
Streptomyces albulus NBRC14147 produces a homopoly(amino acid), ε-poly-l-lysine (ε-PL). Due to its antibiotic activity, thermostability, biodegradability, and non-toxicity to humans, ε-PL is used as a food preservative. In this study, homology searches of diaminopimelate (DAP) pathway genes (dapB and dapE), in an S. albulus genome database, were shown to encode predicted enzymes using dapB or dapE in Escherichia coli strain complementation assays. We observed that dapB and dapE transcriptional levels were weak during ε-PL production stages. Therefore, we strengthened this expression using an ermE constitutive promoter. Engineered strains generated faster growth and ε-PL production rates when compared with the control strain. Moreover, maximum ε-PL yields in S. albulus, where dapB was constitutively expressed, were approximately 14% higher when compared with the control strain. These findings showed that enhanced lysine biosynthetic gene expression generated faster and higher ε-PL production levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Shima S, Sakai H. Polylysine produced by streptomyces. Agric Biol Chem. 1977;41:1807–9.
Shima S, Sakai H. Poly-l-lysine produced by streptomyces. Part iii. chemical studies. Agric Biol Chem. 1981;45:2503–8.
Shima S, Sakai H. Poly-l-lysine produced by streptomyces. Part ii. taxonomy and fermentation studies. Agric Biol Chem. 1981;45:2497–502.
Shima S, Fukuhara Y, Sakai H. Inactivation of bacteriophages by e-poly-l-lysine produced by streptomyces. Agric Biol Chem. 1982;46:1917–9.
Shima S, Matsuoka H, Iwamoto T, Sakai H. Antimicrobial action of E-poly-L-lysine. J Antibiotics. 1984;37:1449–55.
Hamano Y, et al. ε-Poly-L-lysine producer, Streptomyces albulus, has feedback-inhibition resistant aspartokinase. Appl Microbiol Biotechnol. 2007;76:873–82.
Xu Z, et al. Recent advances in the biotechnological production of microbial poly(ɛ-l-lysine) and understanding of its biosynthetic mechanism. Appl Microbiol Biotechnol. 2016;100:6619–30.
Wang L, et al. Epsilon-poly-L-lysine: recent advances in biomanufacturing and applications. Front Bioeng Biotechnol. 2021;9:1–21.
Takeuchi Y, et al. First direct evidence for direct cell-membrane penetrations of polycationic homopoly(amino acid)s produced by bacteria. Commun Biol. 2022;5:1132.
Yamanaka K, Maruyama C, Takagi H, Hamano Y. ε-poly-L-lysine dispersity is controlled by a highly unusual nonribosomal peptide synthetase. Nat Chem Biol. 2008;4:766–72.
Yamanaka K, Hamano Y. Draft genome sequence of the most traditional ε-Poly-L-Lysine Producer, Streptomyces albulus NBRC14147. Microbiol Resour Announc. 2019;8:e01515–18.
Hiraki J, Hatakeyama M, Morita H, Izumi Y. Improved ε-Poly-L-Lysine production of an S-(2-aminoethyl)-L-cysteine resistant mutant of streptomyces albulus. Seibutsu Kogaku. 1998;76:487–93.
Li S, et al. Genome shuffling enhanced ε-Poly-l-Lysine production by improving glucose tolerance of streptomyces graminearus. Appl Biochem Biotechnol. 2012;166:414–23.
Li S, et al. Combining genome shuffling and interspecific hybridization among streptomyces improved ε-Poly-l-Lysine production. Appl Biochem Biotechnol. 2013;169:338–50.
Ochi K. From microbial differentiation to ribosome engineering. Biosci Biotechnol Biochem. 2007;71:1373–86.
Zhang K, et al. Combinatorial effect of ARTP mutagenesis and ribosome engineering on an industrial strain of streptomyces albus S12 for enhanced biosynthesis of salinomycin. Front Bioeng Biotechnol. 2019;7:1–13.
Li W, et al. Effects of amino acids and overexpression of dapA Gene on the production of ε-Poly-L-lysine by Streptomyces diastatochromogenes Strains. Curr Microbiol. 2021;78:2640–7.
Xu D, et al. Enhancement of ε-poly-l-lysine production by overexpressing the ammonium transporter gene in Streptomyces albulus PD-1. Bioprocess Biosyst Eng. 2018;41:1337–45.
Hashimoto M, et al. Cell size and nucleoid organization of engineered Escherichia coli cells with a reduced genome. Mol Microbiol. 2005;55:137–49.
Kato JI, Hashimoto M. Construction of consecutive deletions of the Escherichia coli chromosome. Mol Syst Biol. 2007;3:1–7.
Yamanaka K, et al. Mechanism of ε-poly-L-lysine production and accumulation revealed by identification and analysis of an ε-poly-L-lysine-degrading enzyme. Appl Environ Microbiol. 2010;76:5669–75.
Kahar P, Iwata T, Hiraki J, Park EY, Okabe M. Enhancement of epsilon-polylysine production by Streptomyces albulus strain 410 using pH control. J Biosci Bioeng. 2001;91:190–4.
Chen XS, et al. Enhancement of ε-poly-l-lysine production coupled with precursor l-lysine feeding in glucose-glycerol co-fermentation by streptomyces sp. M-Z18. Bioprocess Biosyst Eng. 2013;36:1843–9.
Hamano Y, et al. Development of gene delivery systems for the ε-poly-L-lysine producer, Streptomyces albulus. J Biosci Bioeng. 2005;99:636–41.
Dairi T, Nakano T, Aisaka K, Katsumata R, Hasegawa M. Cloning and nucleotide sequence of the gene responsible for chlorination of tetracycline. Biosci Biotechnol Biochem. 1995;59:1099–106.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–60.
Pertea M, et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33:290–5.
Robinson MD, McCarthy DJ, Smyth G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–40.
Itzhaki RF. Colorimetric method for estimating polylysine and polyarginine. Anal Biochem. 1972;50:569–74.
Acknowledgements
The authors would like to thank Enago (www.enago.jp) for the English language review.
Funding
This research was supported by JSPS KAKENHI grants 20K21284 (Y.H.), 20H02918 (Y.H.), 20K15437 (F.H.), and 22K05390 (F.H.), and by a JSPS KAKENHI Grant-in-Aid for Transformative Research Areas (A) 22H05122 (C.M.).
Author information
Authors and Affiliations
Contributions
FH, KY, TO, CM, and YH designed experiments. CM and YH performed the experiments for RT-PCR and the construction of S. albulus transformants. FH and KA conducted complementation assays and fermentation experiments on S. albulus transformants. KY performed RNA-Seq analysis. FH, KY and YH wrote the manuscript. YH led the study.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hasebe, F., Adachi, K., Yamanaka, K. et al. Constitutive and high gene expression in the diaminopimelate pathway accelerates ε-poly-l-lysine production in Streptomyces albulus. J Antibiot 76, 522–531 (2023). https://doi.org/10.1038/s41429-023-00636-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-023-00636-9