Abstract
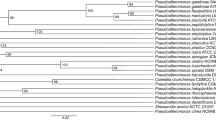
To further exploit secondary metabolic potential of a minor actinomycete genus Phytohabitans within the family Micromonosporaceae, metabolite profiling by HPLC-UV analysis, combined with 16S rDNA sequence-based phylotyping were attempted on seven Phytohabitans strains available at the public culture collection. The strains were grouped into three clades and each exhibited unique and distinct metabolite profiles, which were highly conserved among strains within the same clade. These results were consistent with previous observations on two other actinomycetes genera, reconfirming species-specificity of secondary metabolite production, which were conventionally thought to be strain-specific. A strain RD003215, belonging to the P. suffuscus clade, produced multiple metabolites, some of which were presumed to be naphthoquinones. Liquid fermentation followed by chromatographic separation of the broth extract led to the discovery of three new pyranonaphthoquinones, designated habipyranoquinones A–C (1–3), and one new isatin derivative, (R)-N-methyl-3-hydroxy-5,6-dimethoxyoxindole (4), along with three known synthetic compounds, 6,8-dihydroxydehydro-α-lapachone (5), N-methyl-5,6-dimethoxyisatin (6), and 5,6-dimethoxyisatin (7). Structures of 1–4 were unequivocally elucidated by NMR, MS, and CD spectral analysis, with assistance of density functional theory-based NMR chemical shift prediction and ECD spectral calculation. Compound 2 displayed antibacterial activity against Kocuria rhizophila and Staphylococcus aureus with MIC 50 µg/mL and cytotoxicity against P388 murine leukemia cells with an IC50 value of 34 µM. Compounds 1 and 4 also showed cytotoxicity against P388 cells with IC50 values of 29 and 14 µM, respectively.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Demain AL. Importance of microbial natural products and the need to revitalize their discovery. J Ind Microbiol Biotechnol. 2014;41:185–201.
Komaki H, et al. Diversity of nonribosomal peptide synthetase and polyketide synthase gene clusters among taxonomically close Streptomyces strains. Sci Rep. 2018;8:6888.
Zdouc MM, et al. Planomonospora: a metabolomics perspective on an underexplored actinobacteria genus. J Nat Prod. 2021;84:204–19.
List of prokaryotic names with standing in nomenclature. http://www.bacterio.net. accessed 17 Nov 2022.
Dictionary of Natural Products 31.1 Chemical Search. https://dnp.chemnetbase.com/faces/chemical/ChemicalSearch.xhtml. accessed 10 Dec 2022.
Inahashi Y, Matsumoto A, Hirofumi D, Omura S, Takahashi Y. Phytohabitans suffuscus gen. nov., sp. nov., an actinomycete of the family Micromonosporaceae isolated from plant roots. Int J Syst Evol Microbiol. 2010;60:2652–8.
Uchida R, Yokota S, Tomoda H. Structure elucidation of meroterpenoid habiterpenol, a novel abrogator of bleomycin-induced G2 arrest in Jurkat cells, produced by Phytohabitans suffuscus 3787_5. J Antibiot. 2014;67:783–6.
Saito S, Ye X, Zhou T, Nakajima-Shimada J, Tashiro E, Triningsih DW, Harunari E, Oku N, Igarashi Y. Phytohabitols A–C, δ-lactone-terminated polyketides from an actinomycete of the genus Phytohabitans. J Nat Prod. 2022;85:1697–703.
Harunari E, Mae S, Fukaya K, Tashiro E, Urabe D, Igarashi Y. Bisprenyl naphthoquinone and chlorinated calcimycin congener bearing thiazole ring from an actinomycete of the genus Phytohabitans. J Antibiot. 2022;75:542–51.
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019;47:W81–7.
List of available RD strains on domestically derived strains. https://www.nite.go.jp/nbrc/cultures/rd/available_rd_list.html. accessed 11 Nov 2021.
Jensen PR, Williams PG, Oh D, Zeigler L, Fenical W. Species-specific secondary metabolite production in marine actinomycetes of the genus Salinispora. Appl Environ Microbiol. 2007;73:1146–52.
Snyder SA, Tang Z, Gupta R. Enantioselective total synthesis of (–)-napyradiomycin A1 via asymmetric chlorination of an isolated olefin. J Am Chem Soc. 2009;131:5744–5.
Taylor A. The synthesis of the dimethoxyisatins. J Chem Res Synop. 1980;10:347.
Wang Z, Wang C, Sun Y, Zhang N, Liu Z, Liu J. A novel strategy to the synthesis of 4-anilinoquinazoline derivatives. Tetrahedron. 2014;70:906–13.
Villemin D, Hammadi M, Hachemi M. Supported metalated phthalocyanine as catalyst for oxidation by molecular oxygen. Synthesis of quinones and carbonyl compounds. Synth Commun. 2002;32:1501–15.
Charan RD, Schlingmann G, Bernan VS, Feng X, Carter GT. Fumaquinone, a new prenylated naphthoquinone from Streptomyces fumanus. J Antibiot. 2005;58:271–4.
Peña-López M M, Montserrat Martínez MM, Sarandeses LA, Sestelo JP. A versatile synthesis of fumaquinone. J Org Chem. 2010;75:5337–9.
Lodewyk MW, Siebert MR, Tantillo DJ. Computational prediction of 1H and 13C chemical shifts: a useful tool for natural product, mechanistic, and synthetic organic chemistry. Chem Rev. 2012;112:1839–62.
Grimblat N, Sarotti AM. Computational chemistry to the rescue: modern toolboxes for the assignment of complex molecules by GIAO NMR calculations. Chem Eur J. 2016;22:12246–61.
Shen X, Wang X, Huang T, Deng Z, Lin S. Naphthoquinone-based meroterpenoids from marine-derived Streptomyces sp. B9173. Biomolecules. 2020;10:1187.
Isogai S, Nishiyama M, Kuzuyama T. Identification of 8-amino-2,5,7-trihydroxynaphthalene-1,4-dione, a novel intermediate in the biosynthesis of Streptomyces meroterpenoids. Bioorg Med Chem Lett. 2012;22:5823–6.
Nugroho AE, Morita H. Circular dichroism calculation for natural products. J Nat Med. 2014;68:1–10.
Pescitelli G, Bruhn T. Good computational practice in the assignment of absolute configurations by TDDFT calculations of ECD spectra. Chirality. 2016;28:466–474.
Moore RE, Singh H, Scheuer PJ. A pyranonaphthazarin pigment from the sea urchin Echinothrix diadema. Tetrahedron Lett. 1968;43:4581–3.
Donnelly DMX, O’Reilly J. 6-Methyldehydro-α-lapachone from Fomes annosus. Phytochemistry. 1980;19:277–9.
Matsumoto T, Ichihara A, Yanagiya M, Yuzawa T, Sannai A, Oikawa H, Sakamura S, Eugster CH. Two new syntheses of the pyranojuglone pigment α-caryopterone. Helv Chim Acta. 1985;68:2324–31.
Lima CSA, de Amorim ELC, Nascimento SC, de Araújo CF, Agra MF, Barbosa-Filho JM, Silva MS, Da-Cunha EVL, Vieira IJC, Braz-Filho R. Cytotoxic pyranonaphthoquinones from Melloa quadrivalvis (Bignoniaceae). Nat Prod Res. 2005;19:217–22.
Takao T, Kitatani F, Watanabe N, Yagi A, Sakata K. A simple screening method for antioxidants and isolation of several antioxidants produced by marine bacteria from fish and shellfish. Biosci Biotech Biochem. 1994;58:1780–3.
Shaaban KA, Shaaban M, Nair V, Schuhmann I, Win HY, Lei L, Dittrich B, Helmke E, Schüffler A, Laatsch H. Structure elucidation and synthesis of hydroxylated isatins from Streptomycetes. Z Naturforsch. 2016;71:1191–8.
Bérdy J. Bioactive microbial metabolites. J Antibiot. 2005;58:1–26.
Saito S, Indo K, Oku N, Komaki H, Kawasaki M, Igarashi Y. Unsaturated fatty acids and a prenylated tryptophan derivative from a rare actinomycete of the genus Couchioplanes. Beilstein J Org Chem. 2021;17:2939–49.
Liu C, Zhang Z, Fukaya K, Urabe D, Harunari E, Oku N, Igarashi Y, Catellatolactams A–C. plant growth-promoting ansamacrolactams from a rare actinomycete of the genus Catellatospora. J Nat Prod. 2022;85:1993–9.
Lu S, Harunari E, Oku N, Igarashi Y, Trehangelin E. a bisacyl trehalose with plant growth promoting activity from a rare actinomycete Polymorphospora sp. RD064483. J Antibiot. 2022;75:296–300.
Saito S, Oku N, Igarashi Y. Mycetoindole, an N-acyl dehydrotryptophan with plant growth inhibitory activity from an actinomycete of the genus Actinomycetospora. J Antibiot. 2022;75:44–7.
MacroModel, Schrödinger, LLC: New York, NY, 2020. https://www.schrodinger.com/products/macromodel
Gaussian 16, Revision C.01; Gaussian, Inc.: Wallingford, CT, 2016. https://gaussian.com/gaussian16/
Yoon H, et al. Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol. 2017;67:1613–7.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9.
Karim MR, Harunari E, Oku N, Akasaka K, Igarashi Y. Bulbimidazoles A−C, antimicrobial and cytotoxic alkanoyl imidazoles from a marine gammaproteobacterium Microbulbifer species. J Nat Prod. 2020;83:1295–9.
Acknowledgements
The authors thank Dr. Shun Saito (currently at Keio University) for providing the 16S rRNA gene sequences of strains RD002984 and RD003215.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Triningsih, D.W., Harunari, E., Fukaya, K. et al. Species-specific secondary metabolism by actinomycetes of the genus Phytohabitans and discovery of new pyranonaphthoquinones and isatin derivatives. J Antibiot 76, 249–259 (2023). https://doi.org/10.1038/s41429-023-00605-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-023-00605-2
This article is cited by
-
Development of a drug discovery approach from microbes with a special focus on isolation sources and taxonomy
The Journal of Antibiotics (2023)