Abstract
Vanitaracin A, an anti-hepatitis B virus polyketide, has been previously isolated from Talaromyces sp. In the present study, we searched for novel compounds in the culture broth obtained from a vanitaracin A-producing fungus under various conditions. Three novel compounds (vanitaracin C, vanitaraphilone A, and 2-hydroxy-4-(hydroxymethyl)-6-methylbenzaldehyde) were isolated, and their structures were determined using spectroscopic methods (1D/2D NMR and MS). In addition, the antiviral spectrum of vanitaracin A was examined by measuring its antiviral activities against rabies virus, Borna disease virus 1, and bovine leukemia virus. This compound exhibited antiviral activity against bovine leukemia virus, which is the causative agent of enzootic bovine leukosis. The anti-bovine leukemia virus effects of other compounds isolated from the vanitaracin A-producing fungus, namely, vanitaracins B and C, vanitaraphilone A, and 2-hydroxy-4-(hydroxymethyl)-6-methylbenzaldehyde, were also evaluated. Vanitaracin B, vanitaraphilone A and 2-hydroxy-4-(hydroxymethyl)-6-methylbenzaldehyde were also found to exhibit activity against bovine leukemia virus. These findings reveal the broad-spectrum antiviral activity of the vanitaracin scaffold and suggest several candidates for the development of anti-bovine leukemia virus drugs.
Similar content being viewed by others
Introduction
Among antivirals isolated from natural sources, several exhibit activities against multiple virus types by targeting host cellular pathways that are exploited by different viruses [1, 2]. For example, fungus-derived cyclosporine A inhibits the replication of hepatitis C virus (HCV), influenza virus, and corona virus by binding to cyclophilin A2. Lipid-lowering statins, such as lovastatin isolated from the fungus Aspergillus terreus, inhibit HMG-CoA reductase and attenuate the replication of some enveloped viruses [1]. Ivermectin, an antiparasitic drug that inhibits the replication of HIV-1 and dengue virus by blocking importin α/β-mediated nuclear import [3], has also recently been found to inhibit the replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [4]. Such host-targeting antivirals (HTAs) are generally expected to possess a much higher genetic barrier to drug resistance than direct-acting antivirals (DAAs) [2]. Furthermore, HTAs with broad-spectrum antiviral activities may provide treatment options for rapidly growing infectious diseases caused by new viral pathogens [1, 2, 5, 6].
In addition to viral infectious diseases that are a serious threat to human health, animal viral infectious diseases, including zoonosis such as rabies, are also a major concern in pet and livestock animals. Previously, by screening for antiviral compounds against human or animal viruses from the culture broth of fungi, we found anti-hepatitis B virus (HBV) [7], anti-HCV [8,9,10], and anti-bovine leukemia virus (BLV) compounds [11]. HBV and HCV cause chronic hepatitis, liver cirrhosis, and hepatocellular carcinoma. In contrast, BLV is a retrovirus that causes enzootic bovine leukosis, a lethal infectious disease of cattle. BLV infection also reduces lifetime milk production, reproductive efficiency, and lifespan, thus causing major economic losses in the cattle industry [12,13,14,15]. In addition, BLV is closely related to human T-cell leukemia virus (HTLV), a human retrovirus that is the etiological agent of adult T-cell leukemia.
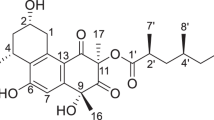
Previously, we isolated vanitaracin A (1, Fig. 1), an anti-HBV tricyclic polyketide, from a fungus of the genus Talaromyces [7, 16]. Compound 1 inhibits viral entry by binding to sodium taurocholate cotransporting polypeptide, an HBV entry receptor on the cell surface [17]. This compound, which is an HTA, represents a novel class of anti-HBV agents with a different mechanism of action than existing DAAs, such as nucleos(t)ide analogs. In addition, the structure of 1 seems to be uncommon, comprising a tricyclic skeleton, which, to the best of our knowledge, is only found in penijanthinone A and dothideomycetide A [18, 19].
In the present study, we cultured 1-producing fungus of the genus Talaromyces under various conditions to isolate novel vanitaracin derivatives. The structures of the obtained compounds were determined using spectroscopic methods (1D/2D NMR, MS, and IR spectroscopy), and their anti-HBV activities were assessed. Furthermore, 1 was subjected to antiviral assays against zoonotic viruses, such as rabies virus, Borna disease virus 1, and BLV to examine its antiviral spectrum. In addition to anti-HBV activity, 1 was found to be effective against BLV, suggesting that this compound has broad-spectrum antiviral activity. We also found that various compounds isolated from the 1-producing fungus exhibited anti-BLV activity. Our data suggest that vanitaracins have potential for controlling BLV infection and that the vanitaracin scaffold shows potent broad-spectrum antiviral activity.
Results and discussion
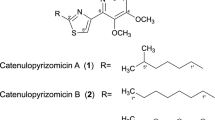
Our previous study demonstrated that the cultivation of a fungus of the genus Talaromyces in potato dextrose broth (PDB) yielded 1, vanitaracin B (2), 3,5-dihydroxy-2-(2-(2-hydroxy-6-methylphenyl)-2-oxoethyl)-4-methylbenzaldehyde, 7-hydroxy-5-methyl-2-(2-oxobutyl)-4H-chromen-4-one, and 2,7-dihydroxy-5-methyl-2-(2-oxobutyl)chroman-4-one [7]. To isolate further novel compounds related to 1, we cultured the fungus under various conditions. First, procaine, a DNA methyltransferase inhibitor, which reportedly enhances secondary metabolite production in fungi [20], was added to the PDB culture medium. The crude extract from this fungal culture broth was subjected to TLC-guided fractionation using silica gel chromatography and HPLC. Under these culture conditions, 1 was obtained together with unknown compound 3. The molecular formula of compound 3 (C24H34O6) was determined using HRMS (FAB). 1H NMR spectroscopy suggested that the structure of compound 3 was similar to that of 2, except for the presence of a methine proton signal (δ 5.36) and the absence of a ketone signal at C-10 (Fig. 1 and Table 1). Based on the correlation between H2-9 and H-10 in the 1H–1H COSY spectrum of 3, the methine proton signal was assigned to H-10, which was supported by HMBC correlations of H3-16 with C-10, C-11, and C-12 (Fig. 2). The consecutive 1H–1H COSY correlations of H-2′–H-8′, and the HMBC correlations of H-3′ and H-7′ with C-1′ (δ 176.5) revealed the presence of a 2,4-dimethylhexanoate unit, which is also present in 1 and 2. The HMBC correlation of H-10 with the ester carbon (δ 176.5) for 3 suggested that the 2,4-dimethyl hexanoate side chain was connected to C-10, whereas this side chain was connected to C-11 in the structures of 1 and 2. The differences in the molecular formulas of 2 and 3 indicated the presence of a hydroxy group at the C-11 position in 3. The remaining proton and carbon signals were assigned using 1H–1H COSY, HMQC, and HMBC experiments (Table 1). Thus, the structure of 3 was determined, as shown in Fig. 1, and this compound was named vanitaracin C. A NOESY correlation between H-2 and H-4 was observed, which indicated a syn relationship between OH-2 and H3-15 (Fig. 2). The NOESY correlation between H3-16 and H-9 (δ 3.34) suggested that these two protons were on the same face of the ring. Cross peaks were observed between H-9 (δ 3.34) and H-10 but not between H-9 (δ 3.09) and H-10, indicating that H-10 was also on the same face. Thus, the relative configuration of the tricyclic moiety was determined, as shown in Fig. 2.
The relative configurations of the 2,4-dimethyl hexanoate moiety in 2 and 3 were determined using Schmidt’s method, which we previously applied to determine the configuration of the moiety in 1 (refs. 16, 21). The differences in the chemical shifts (Δδ) of the C-3′ geminal protons of compounds 2 and 3 were calculated to be 0.64 and 0.60 ppm, respectively (Figure S17). These Δδ values were all greater than 0.4 ppm, suggesting a syn relationship between C-2′ and C-4′ in 2 and 3, similar to that in 1 (refs. 16).
Because the conformational characteristics between the tricyclic skeleton and the side chain of 3 is still unclear, we employed computational methods to predict the absolute configuration of the stereogenic centers in 3. Based on the data described above, we set four possible diastereomers, (10S,11S,2′S,4′S)-3a, (10S,11S,2′R,4′R)-3b, (10R,11R,2′S,4′S)-3c, and (10R,11R,2′R,4′R)-3d, which were analyzed using the DP4 analysis program [22, 23] (Figure S18). Initially, all the diastereomers were submitted to a conformational search using the MMFF94s conformer research algorithm. Thereafter, all the conformers with relative differences within 5 kcal/mol were optimized via DFT calculations at the B3LYP/6-31G(d,p) level based on the solvent effect for PCM (CHCl3) in the Gaussian 16 package [24]. The optimized conformers were identified using the GIAO method at the mPW1PW91/6-31G(d,p) level, and the NMR spectra of the resultant conformers within 5 kcal/mol were averaged based on the Boltzmann populations to give the estimated chemical shifts. The 1H and 13C chemical shifts of all diastereomers with the statistical DP4+ value resulted in agreement with (10S,11S,2′S,4′S)-3a, which suggested that compound 3 has the structure of 3a or its enantiomer. Furthermore, we calculated CD spectra of each diastereomer using TD-DFT at the cam-B3LYP/6-311++G(2d,p) level. The calculated ECD spectrum of 3a matched the experimental data well as shown in Figure S19. Thus, the absolute and relative stereo configurations were determined as (2S,4R,10S,11S,2′S,4′S)-3a.
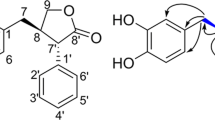
Next, we cultured the fungus in a malt extract broth. Purification of the crude extract via silica gel chromatography yielded 1 as well as unknown compounds 4 and 5. The molecular formula of compound 4 (C25H32O7) was determined by HRMS (FAB). The 1H NMR spectrum implied that the scaffold of 4 differed from that of vanitaracin. The 13C NMR and HMQC spectra revealed the presence of 25 carbons, including 3 ketone carbons, 1 ester carbon, 4 quaternary carbons, 8 methine carbons, 4 methylene carbons, and 5 methyl carbons (Table 2). The 1H–1H COSY and HMBC correlations revealed the presence of a 2,4-dimethylhexanoate unit, which is present in all the vanitaracins (Fig. 3). The 1H and 13C NMR data, the HMBC correlations of the methyl protons (H-17) with ketone carbons C-12 (δ 192.9) and C-14 (δ 193.0) as well as oxygenated quaternary carbon C-13 (δ 83.9), and the HMBC correlations of olefinic protons H-9, H-11, and H-16 with C-8, C-10, C-14, and C-15 indicated the presence of an azaphilone skeleton. The presence of a 3-hydroxy-5-methylcyclohexanone moiety was established based on the consecutive 1H–1H COSY correlations for H-2–H-7 and the HMBC correlation of H-2 with C-1 (δ 202.3). The HMBC correlation between H-9 and C-6 indicated that the cyclohexanone moiety was connected to the azaphilone skeleton. Thus, the planer structure of 4 was determined, as shown in Fig. 3, and this compound was named vanitaraphilone A. The relative configuration of the 3-hydroxy-5-methylcyclohexanone moiety in 4 was determined based on NOESY correlations (Fig. 3). The NOESY correlation between H-3 and H-5 suggested a syn relationship between OH-3 and H3-7. The relative configurations of C-5 and C-6 were defined based on the typical trans-diaxial coupling constant (JH-5/H-6 = 12.2 Hz), as supported by the NOESY correlation between H-2α (δ 2.44) and H-6. Compound 4 is structurally related to azaphilones, cohaerin B isolated from the stromata of the xylariaceous ascomycete Hypoxylon cohaerens [25], and penicilone A isolated from the marine-derived fungus Penicillium janthinellum HK1‑6 (ref. 26). The difference between the chemical shifts (Δδ) of the C-3′ geminal protons of 4 was 0.64 ppm (Figure S17), suggesting a syn relationship between C-2′ and C-4′ (ref. 21). Given that the stereochemistry of the C-13 positions of several azaphilones, including cohaerins and penicilones, have been established via ECD spectroscopy [25, 26], we measured ECD spectrum of 4. Compound 4 showed a negative Cotton effect at 354 nm, which clearly indicated (R)-configuration for C-13 (Figure S20). However, due to shortage of materials, the absolute configurations of other stereogenic centers in 4 were not determined.
The molecular formula of compound 5 (C9H10O3) was determined using HRMS (FAB). The 1H NMR and 13C NMR data suggested the presence of one aldehyde carbon, six aromatic carbons, one oxymethylene carbon, and one methyl carbon. All the proton and carbon signals were assigned based on the HMBC correlations of H-7 with C-1 and C-2, H2-8 with C-3, C-4, and C-5, and H3-9 with C-1, C-5, and C-6 (Figure S21). Thus, compound 5 was determined to be 2-hydroxy-4-(hydroxymethyl)-6-methylbenzaldehyde (Fig. 1).
None of the newly identified compounds (3–5) exhibited anti-HBV activity (Figures S22 and S23). Our previous study demonstrated that 2 has much lower anti-HBV activity than 1 (ref. 7). These observations suggested that the overall structure of 1, especially the methyl and hydroxy groups at the C-9 position, is essential for anti-HBV activity. Additionally, we examined the antiviral spectrum of 1 by evaluating its antiviral activities against zoonotic viruses such as rabies virus, Borna disease virus 1, and BLV. Compound 1 had no significant effect on rabies virus or Borna disease virus 1 (Figure S24), but displayed dose-dependent anti-BLV activity without cytotoxicity (Fig. 4a, b). Compound 1 inhibited syncytia formation, a typical cellular morphology caused by BLV infection. We also evaluated the anti-BLV activities of all the other metabolites obtained from a 1-producing fungus. At a concentration of 10 µM, 2 and 5 exhibited significant anti-BLV activities. However, vanitaracin C (3) and vanitaraphilone A (4) did not show any such behavior (Fig. 5a and b). Compound 4 only exhibited significant anti-BLV activity at a concentration of 30 µM (Fig. 5c). Our results also indicated that at concentrations in the range 30–100 µM, 3 significantly decreased the viability of CC81 cells. Further, 4 showed cytotoxicity at high concentration (50 and 100 µM) (Figure S25).
Cytotoxicity and anti-BLV activity of 1. a Cell viability of CC81 cells treated with various concentrations of 1. b Anti-BLV activity measured via syncytium assays. CC81 cells were cultivated with FLK-BLV supernatant supplemented with various concentrations of 1. The counts of the infectious virus are shown as relative syncytium-forming units (SFU). Data are presented as mean ± SE (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001: ANOVA with post-hoc Tukey test
Anti-BLV activities of compounds 2–5. CC81 cells were treated with FLK-BLV supernatant containing BLV and compounds 2–5 (10 μM) (a, b) and compound 4 at 30 μM (c). Cells containing more than five nuclei were defined as syncytia. *P < 0.05: ANOVA with post-hoc Tukey test (a) and Student’s t-test (b, c). Data are presented mean ± SE (n = 3)
In conclusion, the cultivation of 1-producing fungus in PDB containing DNA methyltransferase inhibitor or malt extract broth revealed three novel metabolites. The structures of vanitaracin derivative 3, azaphilone derivative 4, and benzaldehyde derivative 5 were established based on spectroscopic data. Notably, compounds 1, 2, 4, and 5 showed anti-BLV activity. Although BLV infection is prevalent worldwide and causes significant economic losses, no anti-BLV drugs have yet been developed. Thus, compounds 1, 2, 4, and 5 could be useful for controlling BLV infection. In future, we will evaluate the antiviral activities of these compounds against HTLV, a human retrovirus that is closely related to BLV, and other viruses. Zoonotic diseases caused by the emergence of new viral pathogens, which are a serious global public health problem, could potentially be treated using such broad-spectrum antiviral small molecules [5]. Among the antiviral compounds that we previously identified via chemical library screening, only 1 exhibited both anti-HBV and anti-BLV activities. This observation indicated that the vanitaracin scaffold shows potent broad-spectrum antiviral activity. Further studies on the antiviral spectrum of vanitaracins and their mechanisms of action are currently underway.
Materials and methods
General experimental procedures
Optical rotations were recorded using a JASCO P-2200 digital polarimeter (Jasco Corp., Tokyo, Japan) at room temperature. UV spectra were obtained using a UVmini-1240 spectrophotometer (Shimadzu Corp., Kyoto, Japan). ECD spectra were recorded in MeOH at a concentration of 2.0 × 10−4 M and at 23 °C using a JASCO J-725 CD spectrometer with 10-mm path-length cuvettes.
IR spectra were recorded using a JASCO FT/IR-4600 spectrophotometer (Jasco Corp.). 1H and 13C NMR spectra were recorded in CDCl3 using a Bruker 400 MHz spectrometer (Avance DRX-400 or Avance III-400; Bruker, Billerica, MA, USA) with TMS and CDCl3 as internal references for 1H and 13C NMR measurements, respectively. Chemical shifts are expressed in δ (ppm) relative to the TMS or residual solvent resonance, and coupling constants (J) are expressed in Hz. Mass spectra were obtained using a JEOL mass spectrometer (JMS-700; JEOL, Tokyo, Japan). Analytical TLC was performed on precoated silica gel 60 F254 plates (Merck, Darmstadt, Germany). Silica gel 60N (Kanto Chemical, Tokyo, Japan) was used for silica gel column chromatography.
Extraction and purification of compound 3
The vanitaracin A-producing fungus [7] was cultured in 4 l of PDB containing 0.5 mM procaine hydrochloride under static conditions at room temperature in the dark for 39 days. The culture broth was extracted using CH2Cl2, and the organic layer was evaporated in vacuo to obtain a crude extract (179 mg). This crude extract was then separated via silica gel column chromatography using CHCl3–MeOH (100:0–95:5) to obtain fractions 1–6. Fraction 4 was further separated via silica gel column chromatography using toluene–EtOAc (20:1–1:1) to give fractions 4-1–4-5. Fraction 4-5 was then purified via HPLC (Shiseido Capcell Pak C18, 5 μm, 20 × 250 mm, with 50%–100% MeOH–H2O gradient elution and at a flow rate of 5 ml min−1) to afford compound 3 (1.4 mg).
Vanitaracin C (3)
Pale pink oil; [α]26D + 59 (c 0.070, CHCl3); UV λMeOH max nm (ε) 284 (11,800), 235 (13,500), 215 (14,000); ECD (c 2.0 × 10−4, MeOH) Δε (nm) −0.68 (280), +0.29 (234); IR νmax (film) cm−1 3425, 2960, 2926, 1726, 1712, 1666, 1583, 1259, 1121; HRMS (FAB) m/z 441.2255 [M + Na]+ (calcd for C24H34O6Na, 441.2253); 13C and 1H data, see Table 1.
Extraction and purification of compounds 4 and 5
The fungal strain was cultured in 10 l of malt extract broth under static conditions at room temperature in the dark for 21 days. The culture broth was extracted using CH2Cl2, and the organic layer was evaporated in vacuo to obtain a crude extract (220 mg). This crude extract was separated via silica gel column chromatography using CHCl3–MeOH (99:1–95:5) to obtain fractions 1–12. Fraction 8 was separated by silica gel column chromatography using hexane–EtOAc (4:1–0:1) to yield compound 4 (2.8 mg). Fraction 5 was separated by silica gel column chromatography using hexane–EtOAc (4:1–0:1) to yield compound 5 (0.4 mg).
Vanitaraphilone A (4)
Brown oil; [α]22D –163 (c 0.065, CHCl3); UV λMeOHmax nm (ε) 331 (29,200), 219 (23,500); ECD (c 2.0 × 10−4, MeOH) Δε (nm) −0.34 (354), +0.21 (272), +0.20 (240), −0.23 (220); IR νmax (film) cm−1 3431, 3020, 2966, 1718, 1641, 1216; HRMS (FAB) m/z 445.2225 [M + H]+ (calcd for C25H33O7, 445.2226); 13C and 1H data, see Table 2.
2-Hydroxy-4-(hydroxymethyl)-6-methylbenzaldehyde (5)
Pale yellow oil; UV λMeOH max nm (ε) 338 (1,100), 271 (3,800), 216 (5,400); IR νmax (KBr) cm−1 3335, 2928, 1644, 1067; HRMS (FAB) m/z 167.0708 [M + H]+ (calcd for C9H11O3, 167.0708); 1H NMR (400 MHz, CDCl3) δ 11.97 (1H, s, OH-2), 10.29 (1H, s, H-7), 6.82 (1H, s, H-3), 6.73 (1H, s, H-5), 4.68 (2H, s, H-8), 2.61 (3H, s, H-9); 13C NMR (100 MHz, CDCl3) δ 194.8 (C-7), 163.6 (C-2), 151.2 (C-4), 142.4 (C-6), 119.6 (C-5), 117.7 (C-1), 113.3 (C-3), 64.5 (C-8), 18.2 (C-9).
Anti-BLV assay
To evaluate the anti-BLV activities of the compounds, a syncytium assay was conducted as previously described [11]. In brief, CC81 cells were cultivated in growth medium containing supernatant from persistently BLV-infected FLK-BLV cells and the compound of interest or DMSO, supplemented with 100 ng ml−1 polybrene. When the cells reached confluence, the formation of syncytia were visualized by Giemsa staining and light microscopy. Cells containing more than five nuclei were defined as syncytia.
References
Ianevski A, et al. Novel activities of safe-in-human broad-spectrum antiviral agents. Antivir Res. 2018;154:174–82.
Ji X, Li Z. Medicinal chemistry strategies toward host targeting antiviral agents. Med Res Rev. 2020;40:1519–57.
Wagstaff KM, Sivakumaran H, Heaton SM, Harrich D, Jans DA. Ivermectin is a specific inhibitor of Importin Α/Β-mediated nuclear import able to inhibit replication of HIV-1 and Dengue Virus. Biochem J. 2012;443:851–6.
Caly L, Druce JD, Catton MG, Jans DA, Wagstaff KM. The FDA-approved drug Ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antivir Res. 2020;178:104787.
Adamson CS et al. Antiviral drug discovery: preparing for the next pandemic. Chem Soc Rev. 2021;5:3647–55.
Andersen PI, et al. Discovery and development of safe-in-man broad-spectrum antiviral agents. Int J Infect Dis. 2020;93:268–76.
Matsunaga H, et al. Isolation and structure of Vanitaracin A, a novel Anti-Hepatitis B virus compound from Talaromyces Sp. Bioorg Med Chem Lett. 2015;25:4325–8.
Nakajima S, et al. Specific inhibition of Hepatitis C virus entry into host hepatocytes by fungi-derived Sulochrin and its derivatives. Biochem Biophys Res Commun. 2013;440:515–20.
Nishikori S, et al. Anti-Hepatitis C virus natural product from a fungus, Penicillium Herquei. J Nat Prod. 2016;79:442–6.
Nakajima S, et al. Fungus-derived Neoechinulin B as a novel antagonist of Liver X receptor, identified by chemical genetics using a Hepatitis C virus cell culture system. J Virol. 2016;90:9058–74.
Murakami H, et al. Specific antiviral effect of Violaceoid E on Bovine Leukemia Virus. Virology. 2021;562:1–8.
Polat M, Takeshima SN, Aida Y. Epidemiology and genetic diversity of Bovine Leukemia Virus. Virol J. 2017;14:209–4.
Nekouei O, VanLeeuwen J, Stryhn H, Kelton D, Keefe G. Lifetime effects of infection with bovine leukemia virus on longevity and milk production of dairy cows. Prev Vet Med. 2016;133:1–9.
Brenner J, Van-Haam M, Savir D, Trainin Z. The implication of BLV infection in the productivity, reproductive capacity and survival rate of a dairy cow. Vet Immunol Immunopathol. 1989;22:299–305.
Schwartz I, Levy D. Pathobiology of bovine leukemia virus. Vet Res. 1994;25:521–36.
Kamisuki S, et al. Determining the absolute configuration of Vanitaracin A, an anti-Hepatitis B virus agent. J Antibiot. 2022;75:92–7.
Kaneko M, et al. A Novel Tricyclic Polyketide, Vanitaracin A, specifically inhibits the entry of Hepatitis B and D viruses by targeting sodium taurocholate cotransporting polypeptide. J Virol. 2015;89:11945–53.
Chen M, et al. NaBr-induced production of brominated azaphilones and related tricyclic polyketides by the marine-derived Fungus Penicillium Janthinellum HK1-6. J Nat Prod. 2019;82:368–74.
Senadeera SP, Wiyakrutta S, Mahidol C, Ruchirawat S, Kittakoop P. A novel Tricyclic Polyketide and its biosynthetic precursor azaphilone derivatives from the endophytic fungus Dothideomycete Sp. Org Biomol Chem. 2012;10:7220–6.
Asai T, et al. Tenuipyrone, a novel skeletal polyketide from the entomopathogenic fungus, Isaria tenuipes, cultivated in the presence of epigenetic modifiers. Org Lett. 2012;14:513–5.
Schmidt Y, Breit B. Direct assignment of the relative configuration in 1,3,n-Methyl-branched carbon chains by 1H NMR Spectroscopy. Org Lett. 2010;12:2218–21.
Grimblat N, Gavín JA, Hernández Daranas A, Sarotti AM. Combining the power of J coupling and DP4 analysis on stereochemical assignments: the J-DP4 Methods. Org Lett. 2019;21:4003–7.
Grimblat N, Zanardi MM, Sarotti AM. Beyond DP4: an improved probability for the stereochemical assignment of isomeric compounds using quantum chemical calculations of NMR Shifts. J Org Chem. 2015;80:12526–34.
Frisch MJ, et al. Gaussian 16, rev. C.01; Wallingford, CT, 2016.
Quang DN, et al. Cohaerins A and B, Azaphilones from the Fungus Hypoxylon Cohaerens, and comparison of HPLC-based metabolite profiles in Hypoxylon Sect. Annulata. Phytochemistry. 2005;66:797–809.
Chen M, Shen N, Chen Z, Zhang F, Chen Y. Penicilones A–D, anti-MRSA azaphilones from the marine-derived fungus Penicillium janthinellum HK1-6. J Nat Prod. 2017;80:1081–6.
Acknowledgements
We acknowledge support from Mr. Yoshihisa Sei and the Materials Analysis Division, Open Facility Center, Tokyo Institute of Technology for NMR analysis, and Mr. Makoto Roppongi and center for Instrumental Analysis, Utsunomiya University for CD analysis. We also thank Dr. Arata Yajima (Tokyo University of Agriculture) for his encouragement and experimental support. This work was supported by the Ministry of Education, Culture, Sports, Science and Technology-Supported Program for the Private University Research Branding Project (2016–2020), grants-in-aid from the Japan Society for the Promotion of Science (KAKENHI 18K05343, 20H03499, 21K05299, and 22K05467), the Program for Basic and Clinical Research on Hepatitis (JP22fk0310504, JP22fk0310511, JP21fk0108589, JP22jm0210068, JP20wm0325007, and JP20fk0210036) from the Japan Agency for Medical Research and Development (AMED), and the Center for Human and Animal Symbiosis Science, Azabu University. The computation was performed by the Research Center for Computational Science, Okazaki, Japan (Project: 22-IMS-C103).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kamisuki, S., Shibasaki, H., Murakami, H. et al. Isolation, structural determination, and antiviral activities of metabolites from vanitaracin A-producing Talaromyces sp.. J Antibiot 76, 75–82 (2023). https://doi.org/10.1038/s41429-022-00585-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-022-00585-9