Abstract
The linearmycin family of polyketides was originally classified as antifungal metabolites. However, in addition to antifungal activity, we previously found that linearmycins cause cellular lysis and colony degradation of the Gram-positive bacterium Bacillus subtilis. We recently showed that Streptomyces sp. strain Mg1 incorporates linearmycins into extracellular vesicles, which are capable of lysing B. subtilis. However, the mechanism of linearmycin-induced lysis was hitherto unexplored. Therefore, we sought to determine how linearmycin-laden vesicles cause lysis. In this study, we found that linearmycins inhibited the growth of all Gram-positive bacteria that we tested, but lysis was limited to some Bacillus species. Next, we found that linearmycin-induced lysis occurred even when cellular metabolism and growth were inhibited, which suggested that linearmycins possess the intrinsic capacity to lyse cells, unlike cell-wall targeting antibiotics. We showed that linearmycin exposure caused changes consistent with rapid depolarization of the B. subtilis cytoplasmic membrane, which was correlated with a loss of viability. Finally, using liposomes as in vitro membrane models, we demonstrated that linearmycins are capable of disrupting lipid bilayers without any other cellular components. Taken together, our results strongly indicate that the cytoplasmic membrane is the direct antibacterial target of linearmycins.
Similar content being viewed by others
Introduction
In natural environments, bacteria face challenge from competitors for nutrients, resources, and physical space [1]. To compete with neighboring organisms, bacteria use different competitive mechanisms including production of bioactive specialized metabolites (SMs) [2]. SMs function in diverse environments including marine sediments [3], insect-associated bacterial symbioses [4], and the human microbiome [5]. The chemical structures of SMs are as varied as the biological activities they possess. For example, some SMs function in nutrient sequestration [6] while others are signaling molecules [7]. However, molecules with antibiotic activity are currently the most prevalent and best understood class of SMs. This is both in part due to the ease of screening for producers of inhibitory activities and the anthropogenic use of antibiotics for clinical and industrial applications [8].
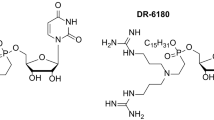
Although new SMs are continually discovered, how the idiosyncrasies of different microbial systems influence the activities of these metabolites is still relatively unknown. Laboratory interactions between soil bacteria including Bacillus, Myxococcus, Pseudomonas, and Streptomyces species are fruitful models for unraveling the activities of SMs in interspecies interactions [1]. A model system involving Bacillus subtilis and species of Streptomyces has uncovered many nuances in the role of antibiotic SMs in bacterial competition. In particular, when colonies of B. subtilis were cultured with Streptomyces sp. strain Mg1 on an agar surface, the B. subtilis extracellular matrix was degraded and the underlying cells were lysed [9]. Linearmycins were identified as the lytic molecules produced by Streptomyces sp. strain Mg1 [10, 11]. To demonstrate specificity, linearmycin B was isolated and shown to be an active lytic agent, similar to crude extracts and colonies of Streptomyces sp. strain Mg1 [10]. In addition, chalcomycin A, the only other known antibiotic produced by Streptomyces sp. strain Mg1, did not cause lysis of B. subtilis [9]. These results indicated that linearmycins are the sole lytic agents in biological assays with B. subtilis. Originally, the linearmycins were identified as a pair of antifungal metabolites: linearmycins A and B [12, 13] (Fig. 1a), but recently mass spectral molecular networking uncovered a large family of variants, all dependent upon a single type I polyketide synthase gene cluster [11]. In general the linearmycin variants share a polyketide backbone of ≥60 carbons that includes multiple conjugated double bonds (Fig. 1a). The linearmycins are structurally similar to antifungal SMs such as the linear ECO-02301, and cyclic amphotericin B and nystatin, all of which contain an extended carbon backbone and at least one polyene moiety. Like antifungal polyenes, purified linearmycins are predominantly insoluble in aqueous solutions. However, the linearmycins were found to be incorporated into extracellular vesicles produced by Streptomyces sp. strain Mg1, and the vesicles were demonstrated to deliver lytic concentrations of the metabolites to B. subtilis. Thus, extracellular vesicles appear to be the principal mode of delivery for linearmycins through aqueous environments [11].
Linearmycins lyse Bacillus subtilis independently of cell growth and metabolism. a The structure of linearmycin B. b–f Cultures of B. subtilis were grown to OD600 = 1, washed, and resuspended in (b) fresh medium, or fresh medium containing (c) 2.5 mM DCCD, d 200 µg/mL phleomycin (Phleo), e 1 µg/mL rifamycin (Rif), f or 250 µg/mL chloramphenicol and 100 µg/mL spectinomycin (Cm + Spc) with (black) or without (gray) 6 mAU s/µL of linearmycins. g Strains B. subtilis with mutations that affect production of CL (clsA), L-PG (mprF), PE and PS (pssA), and sporulenes (sqcH) were treated with linearmycins as above. Each data point is reported relative to the OD600 at 0 min. Each curve is representative of ≥2 biological replicates. The shading represents the standard deviation from technical triplicates. Note, the standard deviation for linearmycin-treated cultures was often smaller than the data points. For clarity, only the average values are shown in panel g
To address a mechanism of action for linearmycins, genetic screens for resistant mutants of B. subtilis were used to identify possible interaction partners. However, using multiple screening approaches, the only resistant mutants recovered had point mutations in the lnrJK (yfiJK) operon, which encodes a two-component signaling system [10]. The mutations activated LnrJK leading to increased expression of the lnrLMN (yfiLMN) operon, which encodes an ATP-binding cassette transporter that is necessary and sufficient for linearmycin resistance [10, 14]. As only gain-of-function mutations that exclusively activate expression of genes for a resistance efflux pump were identified, it is likely that linearmycins target a structure that is indispensable for cell viability. Purified linearmycin A was previously reported to inhibit growth of Escherichia coli and Staphylococcus aureus [13], but the mechanism of action is unknown. Because antifungal polyenes bind ergosterol in fungal cytoplasmic membranes [15, 16] and because linearmycins share structural similarity to antifungal polyenes, it was hypothesized that the essential structure that linearmycins target is the membrane.
In the present study, the antibacterial mechanism of linearmycin-induced lysis was investigated using extracellular vesicle preparations isolated from Streptomyces sp. strain Mg1. First, it was found that linearmycins inhibited the growth of Gram-positive bacteria, but lysis was limited to some Bacillus species. Next, it was shown that linearmycins likely do not require DNA replication, transcription, translation, or active metabolism in B. subtilis to cause lysis. Furthermore, consistent with a membrane-targeting mechanism, fluorescence dequenching of a membrane potentiometric probe was observed with linearmycin exposure in cultures of B. subtilis, suggesting that the cytoplasmic membrane was compromised. Finally, as a direct test of membrane activity, linearmycins lysed artificial membrane vesicles. Together, the data strongly indicated that linearmycins are membrane-targeting antibiotics.
Materials and methods
Bacterial strains and media
Strains of B. subtilis are listed in Table 1. Other strains are listed in Table 2. For general propagation B. subtilis was cultured in lysogeny broth (LB) [1% tryptone, 0.5% yeast extract, 0.5% sodium chloride]. For most assays, B. subtilis was cultured in MYM [0.4% maltose, 0.4% yeast extract, 0.4% malt extract]. For determining minimum inhibitory concentrations (MIC, see below), strains were cultured on 3% trypticase soy agar plates and inoculated into 3% trypticase soy liquid medium. All plates contained 1.5% agar.
Extracellular vesicle isolation and preparation
Extracellular vesicles were prepared from Streptomyces sp. strain Mg1 as previously described [11]. The linearmycin A content of each extracellular vesicle preparation was determined by integrating the 333 nm signal from high-performance liquid chromatography (HPLC) chromatograms [10]. Typically, the final linearmycin A concentration in assays was 5–20 mAU s/µL. As a negative control for lysis, equivalent preparations were obtained from Streptomyces sp. strain Mg1 ΔlnyI because this mutant strain does not produce linearmycins and is unable to lyse B. subtilis in co-culture. Furthermore, Streptomyces sp. strain Mg1 ΔlnyI has a defect in extracellular vesicle production, which results in production of ~27% of the number of vesicle-like particles as the wild type [11]. Therefore, treatment with the extracellular vesicle preparations from Streptomyces sp. strain Mg1 ∆lnyI controls for delivery vehicle and introduction of vesicles without linearmycin.
MIC determination
The MIC for linearmycins was determined using a standard broth microdilution assay. The MIC was defined as the lowest linearmycin concentration where there was no visible growth after overnight incubation. All MIC values were determined in duplicate with identical results.
Lysis assays
Overnight cultures of B. subtilis in LB were diluted to optical density at 600 nm (OD600) = 0.08 in 25 mL MYM. When the cultures reached OD600 = 1, aliquots of the culture were centrifuged at 21,130 × g for 10 min. The cell pellets were resuspended in an equal volume of fresh MYM. For testing the effects of different antibiotics and inhibitors on lysis, the fresh MYM contained 2.5 mM N,N′-dicyclohexylcarbodiimide (DCCD), 200 µg/mL phleomycin, 1 µg/mL rifamycin, or 250 µg/mL chloramphenicol and 100 µg/mL spectinomycin. 100 µL of the resuspended cells were mixed with 5 µL of linearmycin-containing extracellular vesicles in Greiner CELLSTAR® 96-well U bottom plates (Sigma) at a final linearmycin A concentration of 6 mAU s/µL. The plates were incubated at 30 °C in an Infinite M200PRO plate reader (Tecan) and the OD600 was measured every 5 min for 180 min. After each OD600 measurement, the plate was shaken for 85 s. Each assay was performed with at least two biological replicates and with technical triplicates. Given that the average cross-sectional surface area of a phospholipid is 64 Å2 [17], and the surface area of a B. subtilis cell is 7.6–14.2 µm2 (d = 0.87 µm, l = 2.3–4.7 µm) [18], the phospholipid concentration of 100 µL of an OD600 = 1 culture (~108 cells/mL) was calculated to be 4–7.4 µM. Therefore the final ratio of linearmycin A to phospholipid ranges from 0.81–1.5.
Membrane potential measurements
Overnight cultures of B. subtilis in LB were diluted to OD600 = 0.08 in 5 mL MYM with 1 µM of DiSC3(5) [19]. When the cultures reached OD600 = 0.1–0.2, 100 µL aliquots were transferred into Greiner CELLSTAR® black polystyrene 96-well flat bottom plates (Sigma) with 3 µL linearmycin-containing extracellular vesicles. The final linearmycin A concentrations are shown in Fig. 2 and range from 0.3–30 mAU s/µL. The fluorescence of DiSC3(5) was measured using a GloMax®-Multi+ Detection plate reader (Promega) with a red optical kit (excitation: 625 nm, emission: 660–720 nm) every min for 15 min. The assay was validated by treating B. subtilis with 0.5 µM gramicidin ABCD, which rapidly depolarizes membranes via pore formation (data not shown) [20].
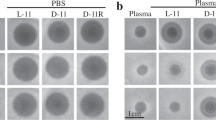
Linearmycins collapse the B. subtilis membrane potential. a Cultures of B. subtilis were grown to OD600 = 0.2 in the presence of DiSC3(5), a membrane potentiometric probe before the addition of linearmycin-containing extracellular vesicles. The change in DiSC3(5) fluorescence was measured over 15 min. No change in fluorescence is indicated by the dashed line. b After the membrane depolarization assay, aliquots of the cultures in a were plated to assess their viability. The data is representative of ≥2 biological replicates. The shading represents the standard deviation from technical triplicates. The linearmycin concentration in mAU s/µL is shown to the right of the traces and image
Large unilamellar vesicle (LUV) production
All phospholipids were purchased from Avanti Polar Lipids and stored in chloroform at 4 °C. Liposomes were produced using the standard film rehydration method [21, 22]. Briefly, 15:0 phosphatidylglycerol (PG), 15:0 phosphatidylethanolamine (PE), 14:0 cardiolipin (CL), and 18:1 1,2-dioleoyl phosphatidylserine (DOPS) were combined in a glass scintillation vial at molar ratios of 43:30:16:11 PG:PE:CL:DOPS, which mimics the phospholipid composition of the B. subtilis 168 cytoplasmic membrane [23, 24]. Using a gentle nitrogen stream, the bulk chloroform was evaporated. The resulting lipid film was stored in vacuo to remove trace chloroform. To form multilamellar vesicles (MLVs), the lipid film was hydrated with 1 mL of sodium phosphate buffer [10 mM sodium phosphate, 100 mM sodium chloride, pH 7.4], with 60 mM calcein. The MLVs were subjected to 20 cycles of freezing in liquid nitrogen followed by thawing in a 70 °C water bath. To form LUVs, the MLVs were sequentially extruded 11 passes each through 1 µm, 0.4 µm, and 0.1 µm pore size polycarbonate membranes (Whatman) using a Mini-Extruder (Avanti Polar Lipids) at ≥85 °C. LUVs were separated from free calcein by gel filtration using a Sephadex G-50 (GE Healthcare) column (2.5 × 17.5 cm). Fractions were collected in a 96-well plate. The absorbance at 450 and 750 nm of each fraction was measured with a GloMax®-Multi+ Detection plate reader (Promega) to determine which fractions contained calcein and LUVs, respectively. The LUV-containing fractions were pooled together. LUVs were stored at 4 °C under nitrogen and used within one week of their preparation.
Calcein leakage assays
Calcein leakage assays were performed as previously described [21, 22]. For each assay reaction, 120 µL of LUVs, 125 µL of sodium phosphate buffer, and 5 µL linearmycin-containing extracellular vesicles were combined. The final linearmycin A and phospholipid concentrations were 16 mAU s/µL and ~200 µM, respectively, for a ratio of 0.08, which is one order of magnitude lower than the concentration used in the above in vivo lysis assays. As a positive control for calcein leakage, LUVs were treated with 0.2% Triton X-100. After 1 h, leaked calcein was separated from intact LUVs by applying the assay mixtures onto illustra NAP-10 Sephadex G-25 columns (GE Healthcare) equilibrated with sodium phosphate buffer. 72 four drop fractions were collected into 96-well plates. The fractions were scanned using a GloMax®-Multi+ Detection plate reader with a blue optical kit (excitation: 450 nm, emission: 510–570 nm). The fractions from the Triton X-100-treated reaction and untreated LUVs were scanned identically to determine which fractions contained free calcein. The free calcein-containing fractions were pooled and the bulk fluorescence was measured as above. Calcein leakage relative to the positive control was quantified as follows: (Fcondition − Funtreated)/(Fpos − Funtreated), where Fcondition, Fpos, and Funtreated are measured fluorescence intensities from the experimental condition, Triton X-100-treated, and untreated LUVs, respectively. Therefore, relative leakage from Triton X-100-treated LUVs is 1 and relative leakage from untreated LUVs is 0. Each assay was triplicated and significance was determined using Welch’s two sample t test.
Dynamic light scattering (DLS) of LUVs
A Zetasizer Nano S (Malvern) equipped with a He-Ne 633 nm laser and an avalanche photodiode detector was used for DLS measurements. The scattering intensity was collected at an angle of 163°. The mean LUV size was determined from monodispersed samples using cumulant analysis.
Transmission electron microscopy of LUVs
For transmission electron microscopy, 5 µL of LUVs were adsorbed onto freshly glow-discharged carbon-coated Formvar grids. After briefly washing the grids with water, the LUVs were negatively stained with a 2% aqueous solution of ammonium molybdate. Images were captured using a JEOL 1200Ex transmission electron microscope with a bottom-mounted 3K × 3K, slowscan, lens-coupled CCD camera (SIA 15C).
Results
Linearmycins lyse Bacillus species and inhibit the growth of other Gram-positive bacteria
The mechanisms of linearmycin-induced growth inhibition and lysis have been hitherto unexplored due to the instability and insolubility of linearmycins in aqueous solution [10]. However, it was recently discovered that Streptomyces sp. strain Mg1 incorporates linearmycins into extracellular vesicles, which stabilize and solubilize the linearmycins [11]. Because extracellular vesicles are simple to isolate and provide a stable, soluble supply of linearmycins, vesicle preparations were used as the source of linearmycin to study the mechanism of action. Currently, the full composition of the extracellular vesicles is unknown. However, the linearmycins are the only known lytic agent produced by Streptomyces sp. strain Mg1, and disruption of linearmycin biosynthesis completely disrupts lytic activity from Streptomyces sp. strain Mg1 [10, 11]. Chalcomycin A, a macrolide antibiotic produced by Streptomyces sp. strain Mg1, was not associated with vesicles and did not have lytic activity [9, 11]. These observations, in addition to Streptomyces sp. strain Mg1 ∆lnyI producing a reduced number of vesicles and being unable to lyse B. subtilis, support the use of isolated vesicles to deliver the lytic activity specific to the linearmycins.
Streptomyces sp. strain Mg1 produces many variants of linearmycins, which are incorporated into extracellular vesicles [11]. As the linearmycin variants have different extinction coefficients, no single value encapsulates the total linearmycin concentration [11]. Therefore, preparations of extracellular vesicles were titrated against B. subtilis to determine a minimum lytic dose, which was correlated to the peak area of linearmycin A using a HPLC equipped with a diode array detector. Extracellular vesicle preparations with a final linearmycin A concentration between 6 and 12 mAU s/µL lyse B. subtilis cultures. Based upon HPLC estimates, 1 mAU s/µL corresponds to ~1 µM linearmycin A. Similar concentrations of linearmycins were used in all following experiments. There was no activity from equivalent fractions isolated from Streptomyces sp. strain Mg1 ΔlnyI. Therefore for simplicity, heretofore extracellular vesicles will be referred to as linearmycins and treatment with the ΔlnyI fractions will be referred to as mock treatment.
In qualitative agar plate assays, B. subtilis growth was inhibited and cells lysed by linearmycins (Table 2). Consistent with prior studies, linearmycins also inhibited the growth of Gram-positive S. aureus strains MW2 and RN4220, but no lysis was observed (Table 2). Previously, linearmycin A was reported to inhibit the growth of Gram-negative E. coli [13]. However, when using the E. coli K-12 strain MG1665, growth was not inhibited by linearmycins (Table 2), even to linearmycin A concentrations ~20× greater than the previously reported MIC. Additionally, linearmycins were inactive against Gram-negative Salmonella enterica serovar Typhimurium LT2 (Table 2), suggesting that the antibacterial activity of linearmycins may be limited to Gram-positive bacteria.
To assess a phylogenetic range of susceptibility, a number of Gram-positive bacterial species were tested for sensitivity to linearmycins. While S. aureus strains, Listeria innocua CLIP 11262 and Corynebacterium glutamicum ATCC 13032 were inhibited, Enterococcus faecalis ATCC 19433 and ATCC 51188 were only partially inhibited by linearmycins (Table 2), suggesting that sensitivity may have a limited phylogenetic range. Although all of the Gram-positive bacteria were at least partially inhibited by linearmycins, only B. subtilis was sensitive to colony lysis by linearmycins (Table 2). Given this limited susceptibility to lysis, other Bacillus species were tested for sensitivity to linearmycin. Bacillus licheniformis ATCC 10716, Bacillus velezensis (formerly Bacillus amyloliquefaciens) FZB42, and Bacillus megaterium PV361 were growth inhibited by linearmycins (Table 2). In addition, B. licheniformis and B. velezensis colonies were lysed by linearmycins. However, no lysis was evident on B. megaterium colonies, perhaps due to colony mucoidy (Table 2). Furthermore, there was no correlation between susceptibility to lysis and the MIC or presence of a homolog of the linearmycin resistance ATP-binding cassette transporter lnrLMN [14] (Table 2). All taken together, linearmycins are generally inhibitory towards Gram-positive bacteria, but lysis is restricted to species of Bacillus with the possible exception of B. megaterium.
Linearmycin-induced lysis is independent of active growth in B. subtilis
Given the limited taxonomic range for linearmycin-induced lysis (Table 2) and the breadth of knowledge on B. subtilis, this species was chosen as a model to uncover the mechanism of linearmycin-induced lysis. To test if lysis required active growth, such as in the case of peptidoglycan synthesis inhibitors, B. subtilis was treated with inhibitors of growth and metabolism. The following inhibitors were used: DCCD to inhibit ATPase activity [25], phleomycin to damage DNA [26, 27], rifamycin to inhibit transcription [28], and a combination of chloramphenicol and spectinomycin to block translation [29, 30]. Without the addition of any antibiotic treatment or linearmycin-containing extracellular vesicles, the final B. subtilis OD600 doubled over 180 min (Fig. 1b). In the absence of linearmycin-treatment, each inhibitor hindered the growth of B. subtilis (Fig. 1c–f). Under these conditions only DCCD and phleomycin were completely bacteriostatic (Fig. 1c, d). In the presence of rifamycin or a combination of chloramphenicol and spectinomycin, limited growth was observed as the final B. subtilis OD600 increased by 0.2 units (Fig. 1e, f). However, when B. subtilis was treated with linearmycins, the cells were lysed and the OD600 of all cultures decreased by 0.7 units, regardless of specific antibiotic treatment (Fig. 1b–f). Because linearmycins lyse cells in growth-inhibited cultures, these findings demonstrated that the lytic mechanism likely occurs independently of cell growth.
Prophages are not released from B. subtilis exposed to linearmycins
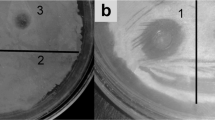
The observation that linearmycins likely lyse B. subtilis even when metabolism is inhibited suggested that linearmycins do not trigger a stress response that leads to lysis. However, it is possible that linearmycins could induce prophage to cause lysis [31]. For example, an endolysin encoded by the defective prophage PBSX [32] can degrade peptidoglycan and form holes in the cell wall. These holes lead to formation of membrane vesicle protrusions and eventual cell death [33]. In addition to PBSX, the B. subtilis genome harbors the prophage SPβ [34], and three prophage-like elements called prophage 1, prophage 3, and skin [35,36,37]. Though unlikely, to verify that prophages were not released as part of the lytic mechanism, lysates from linearmycin-treated cultures were filtered through 0.45 µm filters and tested against an engineered strain of B. subtilis 168 with all prophage elements deleted [37]. The supernatants of untreated B. subtilis cultures contained ~13 plaque forming units (PFU)/mL and the lysates from matched linearmycin-treated cultures contained ~103 PFU/mL, which was low and not significantly different (paired t(2) = 2.94, p = 0.10). No PFUs were observed from a media blank treated with linearmycins. Together, these results indicate that prophages are not involved in the lysis mechanism.
Linearmycins cause membrane depolarization
Linearmycins are structurally similar to antifungal polyenes, which are known to target fungal cytoplasmic membranes [38]. Although, there are contrasting models for the precise antifungal mechanism of polyenes [15, 16], it is known that polyenes interact with the fungal sterol ergosterol, which results in membrane depolarization and ultimately cell death [39]. To test if linearmycins depolarized the B. subtilis cytoplasmic membrane, the cyanine dye 3,3′-dipropylthiadicarbocyanine iodide [DiSC3(5)] was used as a membrane potentiometric probe. DiSC3(5) accumulates in polarized membranes where it forms self-quenching aggregates. When membranes become depolarized, DiSC3(5) is released into the medium, resulting in dequenching that can be measured fluorometrically. Immediately after linearmycin-treatment, a concentration-dependent increase in DiSC3(5) fluorescence signal was observed from the cultures. The mock-treated culture showed no change in fluorescence signal (Fig. 2a). To confirm cell death, culture aliquots were plated following the membrane depolarization assay. There were no viable colonies in the linearmycin-treated cultures, except for some anemic growth from the sample with the least concentrated linearmycins (3 mAU s/µL) (Fig. 2b), confirming the loss of cell viability following membrane depolarization (Fig. 2a).
A specific phospholipid composition is not required for lysis
The activity of linearmycins against growth inhibited cells (Fig. 1c–f), their ability to depolarize B. subtilis membranes (Fig. 2a), and their structural similarity to antifungal polyenes suggested that linearmycins may directly target membranes. The B. subtilis cytoplasmic membrane is composed of more than five classes of phospholipids including PG, PE, CL, phosphatidylserine (PS), and precursor phosphatidic acid [23, 24, 40]. To determine if linearmycin-induced lysis required the presence of specific phospholipid classes, mutants of B. subtilis that were unable to produce different classes of phospholipids were exposed to linearmycins. The requirement of PE and PS for lysis was tested by exposing a pssA deletion mutant [41, 42] to linearmycins. Similarly, the requirement of CL was tested by treating a deletion mutant of clsA, which is responsible for major production of CL [43]. In both cases, the clsA and pssA mutants lysed similarly to the wild-type strain (Fig. 1g). Because B. subtilis ΔpgsA mutants are not viable [44], the dependence of lysis on PG could not be determined. However, a mprF deletion mutant, which is unable to lysinylate PG and form L-PG [42], was still sensitive to lysis (Fig. 1g). Together, these results indicate that several major phospholipid classes were dispensable for lysis. Further, a specific composition of the B. subtilis cytoplasmic membrane was not required for linearmycin-induced lysis.
Antifungal polyenes require the fungal sterol ergosterol for their activity [38]. Strains of B. subtilis do not produce ergosterol, let alone any known sterols [45]. However, B. subtilis can produce sporulene, a heptaprenyl-derived pentacyclic C35-terpenoid [46, 47]. Though sporulenes and sterols have different biosynthetic origins, they are structurally similar molecules. To test if sporulenes were required for lysis, a sqcH mutant, which is unable to produce sporulenes, was treated with linearmycins. As above, the sqcH mutant lysed similarly to wild type B. subtilis (Fig. 1g).
Linearmycins disrupt artificial membrane vesicles
Previous results suggested that linearmycins target the membrane. To directly test for membrane activity, artificial LUVs were produced and treated with linearmycins. To mimic the B. subtilis membrane, the LUVs were produced using PG, PE, CL, and DOPS, a PS mimic. Each of these phospholipid classes comprise individually from 10% up to 43% of the total cytoplasmic membrane phospholipid content of B. subtilis [23, 24, 40]. Although any single phospholipid class was dispensable for lysis in our assays (Fig. 1g), their combination was used to mimic the B. subtilis cytoplasmic membrane composition. Note, B. subtilis primarily uses C15 branched-chain fatty acids in its cytoplasmic membrane [23, 24, 40]. However, C15 straight-chain fatty acids were used due to their commercial availability.
During production, LUVs were loaded with calcein, a self-quenching fluorescent dye, which allowed for the quantification of LUV leakage. When calcein is released from LUVs, the fluorescence signal is dequenched and can be measured fluorometrically. After treating LUVs with linearmycins, there was a significant difference in calcein leakage from linearmycin-treated LUVs (mean relative leakage = 1.19, SD = 0.39) and the mock-treated LUVs (mean relative leakage = −0.09, SD = 0.19); Welch’s t(2.89) = 5.065, p = 0.016 (Fig. 3a). DLS was used to determine the effect of linearmycin treatment on LUV size. The populations of untreated and mock-treated LUVs each had a single DLS peak with an average diameter of ~130 nm, indicating that there was no effect of reaction conditions on LUVs (Fig. 3b). In contrast, after linearmycin treatment, there was a shift from a single DLS peak to two peaks with average diameters of ~221 and ~1327 nm (Fig. 3b). Subsequently, the LUVs were analyzed using negative-staining transmission electron microscopy. As expected, the untreated LUVs were spherical particles with a diameter of ~150 nm (Fig. 3c). However, after linearmycin treatment, there were no intact LUVs. Instead, there were large aggregates of densely staining debris, which were otherwise absent in the untreated LUVs (Fig. 3d). The lysis of LUVs is consistent with the DLS results, which indicated direct targeting of membranes by linearmycins (Fig. 3c).
Linearmycins are active against membranes. a LUVs with B. subtilis phospholipid content were generated in vitro and loaded with calcein. The LUVs were treated with 16 mAU s/µL of linearmycins or were mock treated and the fluorescence of free calcein was monitored to measure liposome leakage. The background fluorescence of untreated LUVs was subtracted from each sample and leakage is reported relative to LUVs treated with 0.2% Triton X-100, as indicated by a value of 1.0. Each experiment was technically triplicated and shown as a dot. The black bar is the mean of the measurements and the upper and lower bounds of the boxes represent the standard deviation. The * indicates that leakage was significantly different at p < 0.05. b DLS measurements of the population size distribution of untreated LUVs and LUVs treated with linearmycin or the mock treatment. c, d Negatively stained transmission electron micrographs of (c) untreated LUVs and (d) linearmycin-treated LUVs. The scale bar is 0.2 µm
Discussion
The linearmycin family of polyene antibiotics has activity against both bacterial and fungal species, including opportunistic pathogens such as S. aureus and Candida albicans [13]. The antifungal mechanism of linearmycins may be consistent with the mechanism ascribed to well-established cyclic polyene antibiotics including amphotericin B and nystatin [38]. However, the antibacterial mechanism for linearmycins was hitherto unexplored. Because linearmycins cause cellular lysis and degradation of established B. subtilis colonies [9,10,11], the antibacterial mechanism of linearmycins was of interest both to further extend a well-established model of bacterial competition and to identify the bacterial target for potential clinical applications.
Using a combination of culture-based and in vitro approaches, the cytoplasmic membrane was determined to be the bacterial target of linearmycins. Linearmycins caused cell lysis in cultures of B. subtilis that were growth inhibited and metabolically inactivated by inhibitory molecules (Fig. 1b–f). The independence from cell growth indicates that cell division is not necessary for lethality, as is required for cell wall-targeting antibiotics such as β-lactams or glycopeptides [48]. Consistent with other membrane-targeting antibiotics, such as gramicidins [20], nisin [49], and other polyenes [39], exposure to linearmycins caused rapid depolarization of the B. subtilis cytoplasmic membrane (Fig. 2a). Intriguingly, membrane depolarization and cell death occur before the OD600 of the B. subtilis culture drops (Figs. 1, 2), which indicated that depolarization is not a consequence of cell lysis. This order of events suggested that linearmycins compromise the cytoplasmic membrane, which leads to cellular lysis. Further, the rapid loss in viability is inconsistent with a prophage-mediated lysis mechanism, as, for example, the prophage SPβ requires >60 min for induction [31]. Indeed, the lysates of B. subtilis cultures treated with linearmycins contained low titers of PFUs, which were insignificantly different from the supernatants of untreated cultures. Furthermore, because linearmycins disrupted artificial vesicles composed only of phospholipids (Fig. 3), interactions with peptidoglycan or membrane proteins are not required for lytic activity. Moreover, linearmycins are water-insoluble, aggregate in aqueous solution, and are associated with extracellular vesicles produced by Streptomyces sp. strain Mg1 [11], all suggesting that linearmycins are associated with membranes. The cytoplasmic membrane as the primary target of linearmycins is also consistent with previous efforts to identify resistant mutants. The only resistant mutants had gain-of-function mutations in the lnrJKLMN linearmycin resistance system [10]. As a target, the membrane is essential and no loss-of-function mutations would bestow linearmycin resistance. All taken together, these findings implicated the cytoplasmic membrane as the target of linearmycins.
The cytoplasmic membrane is a crucial component of living cells and is often a target for antibiotics [50,51,52]. However, deciphering the specific mechanisms through which membrane-targeting antibiotics function is not simple. Membrane-targeting antibiotics may affect bulk membrane properties including membrane curvature, fluidity, and lipid domains, and may introduce packing defects [53]. One challenge in determining specific mechanisms for membrane-targeting antibiotics is the overlapping physiological effects that occur when any of these bulk properties is perturbed. For example, the lipopeptide daptomycin kills B. subtilis by interfering with and rearranging fluid lipid domains, which inhibits cell wall biosynthesis by delocalizing MurG and PlsX and affects lipid clustering, leading to hydrophobic mismatch and membrane depolarization [54]. The morphology of B. subtilis colonies lysed by daptomycin resembles that of colonies lysed by linearmycin [10, 14]. However, because the artificial LUVs used in this study contained only phospholipids with fatty acid tails of similar length (C14 to C18), the potential formation of fluid lipid domains was minimized [55]. The lytic action of linearmycins on LUVs suggested that specific interactions with lipid domains are not required for lysis. Membrane-targeting antibiotics can also target specific membrane lipids or components [53]. In the classic model for antifungal polyene-mediated killing, polyenes accumulate in the membrane, sequester ergosterol, and form membrane-permeabilizing ion channels [38]. However, a recent model postulates that polyenes form extramembranous aggregates that act as ergosterol sponges and kill fungal cells without penetrating the membrane bilayer [16]. Linearmycins structurally resemble antifungal polyenes, which are lytically inactive against B. subtilis [10, 14]. Unlike fungal cells, B. subtilis membranes do not contain sterols and structurally similar sporulenes are not required for lytic activity against cells (Fig. 1g). The lack of a specific known target in the cytoplasmic membrane suggests that linearmycins function as general membrane targeting molecules, which use a substantively different mechanism than antifungal polyenes.
A speculative model for activity is that linearmycins anchor into the membrane by their non-polar polyene region. Indeed, the estimated length of the linearmycin polyene region alone is ~30 Å, while the thickness of a single leaflet of the phospholipid bilayer is ~20 Å. This indicates that the linearmycin polyene region alone can span at least one leaflet of the membrane bilayer. Between the two pentaenes, there are two hydroxy groups (Fig. 1a), which when inserted into the membrane may be placed in an electrostatically unfavorable hydrophobic environment. Perhaps, hydrogen bonding between hydroxyl groups on different molecules leads to linearmycin aggregation, causing pore formation and lysis. Alternatively, the hydroxyl groups could position linearmycin molecules in the membrane such that packing defects are introduced and compromise the membrane integrity [53]. Such effects could also work in combination with destabilizing distortions or strain on membranes due to the intercalation of the rigid polyene portion of linearmycins.
Although it is not currently known if the linearmycins are inserted into the phospholipid bilayer or are simply lumenal cargo of the extracellular vesicles of Streptomyces sp. strain Mg1, a tantalizing model is that fusion between vesicles and the target cell envelope acts as the delivery mechanism and leads to deposition of linearmycins into the membrane. Perhaps the association of linearmycins with extracellular lipid vesicles is favorable for Streptomyces sp. strain Mg1, yet the same molecules destabilize foreign B. subtilis membranes, leading to depolarization and cell lysis. Further studies will be required to determine how linearmycins disrupt membranes and what membrane or envelope features modulate specific susceptibility to lysis. Because linearmycins caused lysis of LUVs containing only phospholipid, a favorable model for relative susceptibility is that differences in cell envelope composition may determine linearmycins access to the plasma membrane target. Comparisons of the cell envelopes between bacterial species that are susceptible or resistant to lysis may uncover susceptibility and resistance factors in the cell envelope. By incorporating membrane active antibiotics into extracellular vesicles, Streptomyces sp. strain Mg1 is able to deliver otherwise insoluble molecules into the membranes of their competitors. These observations suggest that extracellular vesicles produced by bacteria and fungi may harbor new membrane active antibiotics to aid in competition with rising antibiotic resistance.
References
Stubbendieck RM, Vargas-Bautista C, Straight PD. Bacterial communities: interactions to scale. Front Microbiol. 2016;7:1–19.
Stubbendieck RM, Straight PD. Multifaceted interfaces of bacterial competition. J Bacteriol. 2016;198:00275–16.
Patin NV, et al. Effects of actinomycete secondary metabolites on sediment microbial communities. Appl Environ Microbiol. 2017;83:02676–16.
Van Arnam EB, et al. Selvamicin, an atypical antifungal polyene from two alternative genomic contexts. Proc Natl Acad Sci USA. 2016;113:12940–5.
Donia MS, et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell. 2014;158:1402–14.
Hider RC, Kong X. Chemistry and biology of siderophores. Nat Prod Rep. 2010;27:637–57.
Romero D, Traxler MF, López D, Kolter R. Antibiotics as signal molecules. Chem Rev. 2011;111:5492–505.
Davies J. Specialized microbial metabolites: functions and origins. J Antibiot (Tokyo). 2013;66:361–4.
Barger SR, et al. Imaging secondary metabolism of Streptomyces sp. Mg1 during cellular lysis and colony degradation of competing Bacillus subtilis. Antonie Van Leeuwenhoek. 2012;102:435–45.
Stubbendieck RM, Straight PD. Escape from lethal bacterial competition through coupled activation of antibiotic resistance and a mobilized subpopulation. PLoS Genet. 2015;11:e1005722.
Hoefler BC, et al. A link between linearmycin biosynthesis and extracellular vesicle genesis connects specialized metabolism and bacterial membrane physiology. Cell Chem Biol. 2017;24:1238 https://doi.org/10.1016/j.chembiol.2017.08.008. e7.
Sakuda S, Guce-Bigol U, Itoh M, Nishimura T, Yamada Y. Linearmycin A, a novel linear polyene antibiotic. Tetrahedron Lett. 1995;36:2777–80.
Sakuda S, Guce-Bigol U, Itoh M, Nishimura T, Yamada Y. Novel linear polyene antibiotics: linearmycins. J Chem Soc Perkin Trans 1996;2315–9, https://doi.org/10.1039/P19960002315.
Stubbendieck RM, Straight PD. Linearmycins activate a two-component signaling system involved in bacterial competition and biofilm morphology. J Bacteriol. 2017;199:e00186–17.
Gray KC, et al. Amphotericin primarily kills yeast by simply binding ergosterol. Proc Natl Acad Sci USA. 2012;109:2234–9.
Anderson TM, et al. Amphotericin forms an extramembranous and fungicidal sterol sponge. Nat Chem Biol. 2014;10:400–6.
van Meer G, Voelker DR, Feigenson GW. Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol. 2008;9:112–24.
Weart RB, et al. A metabolic sensor governing cell size in bacteria. Cell. 2007;130:335–47.
Te Winkel JD, Gray DA, Seistrup KH, Hamoen LW, Strahl H. Analysis of antimicrobial-triggered membrane depolarization using voltage sensitive dyes. Front Cell Dev Biol. 2016;4:29.
Kelkar DA, Chattopadhyay A. The gramicidin ion channel: a model membrane protein. Biochim Biophys Acta. 2007;1768:2011–25.
Erazo-Oliveras A, et al. The late endosome and its lipid BMP act as gateways for efficient cytosolic access of the delivery agent dfTAT and its macromolecular cargos. Cell Chem Biol. 2016;23:598–607.
Najjar K, Erazo-Oliveras A, Brock DJ, Wang T-Y, Pellois J-P. An l- to d-amino acid conversion in an endosomolytic analog of the cell-penetrating peptide TAT influences proteolytic stability, endocytic uptake, and endosomal escape. J Biol Chem. 2017;292:847–61.
Seydlová G, et al. Surfactin production enhances the level of cardiolipin in the cytoplasmic membrane of Bacillus subtilis. Biochim Biophys Acta. 2013;1828:2370–8.
Uttlová P, et al. Bacillus subtilis alters the proportion of major membrane phospholipids in response to surfactin exposure. Biochim Biophys Acta. 2016;1858:2965–71.
Serrahima-Zieger M, Monteil H, Luu B. Isolation and purification of dicyclohexylcarbodiimide-reactive proteolipid from Bacillus subtilis membrane. Biochim Biophys Acta. 1982;679:369–75.
Maeda K, Kosaka H, Yagishita K, Umezawa H. A new antibiotic, phleomycin. J Antibiot (Tokyo). 1956;9:82–5.
Sleigh MJ. The mechanism of DNA breakage by phleomycin in vitro. Nucleic Acids Res. 1976;3:891–901.
Campbell EA, et al. Structural mechanism for rifampicin inhibition of bacterial rna polymerase. Cell. 2001;104:901–12.
Wisseman CL, Smadel JE, Hahn FE, Hopps HE. Mode of action of chloramphenicol. I. Action of chloramphenicol on assimilation of ammonia and on synthesis of proteins and nucleic acids in Escherichia coli. J Bacteriol. 1954;67:662–73.
Davies J, Anderson P, Davis BD. Inhibition of protein synthesis by spectinomycin. Science. 1965;149:1096–8.
Warner FD, Kitos GA, Romano MP, Hemphill HE. Characterization of SPβ: a temperate bacteriophage from Bacillus subtilis 168M. Can J Microbiol. 1977;23:45–51.
Wood HE, Dawson MT, Devine KM, McConnell DJ. Characterization of PBSX, a defective prophage of Bacillus subtilis. J Bacteriol. 1990;172:2667–74.
Toyofuku M, et al. Prophage-triggered membrane vesicle formation through peptidoglycan damage in Bacillus subtilis. Nat Commun. 2017;8:481.
Lazarevic V, et al. Nucleotide sequence of the Bacillus subtilis temperate bacteriophage SPbetac2. Microbiology 1999;145(Pt 5):1055–67.
Takemaru K, Mizuno M, Sato T, Takeuchi M, Kobayashi, Y. Complete nucleotide sequence of a skin element excised by DNA rearrangement during sporulation in Bacillus subtilis. Microbiology 1995;141(Pt 2):323–7.
Mizuno M, et al. Systematic sequencing of the 283 kb 210 degrees-232 degrees region of the Bacillus subtilis genome containing the skin element and many sporulation genes. Microbiology 1996;142(Pt 1):3103–11.
Westers H, et al. Genome engineering reveals large dispensable regions in Bacillus subtilis. Mol Biol Evol. 2003;20:2076–90.
Odds FC, Brown AJP, Gow NAR. Antifungal agents: mechanisms of action. Trends Microbiol. 2003;11:272–9.
Brajtburg J, Powderly WG, Kobayashi GS, Medoff G. Amphotericin B: current understanding of mechanisms of action. Antimicrob Agents Chemother. 1990;34:183–8.
Salzberg LI, Helmann JD. Phenotypic and transcriptomic characterization of Bacillus subtilis mutants with grossly altered membrane composition. J Bacteriol. 2008;190:7797–807.
Matsumoto K, et al. Cloning, sequencing, and disruption of the Bacillus subtilis psd gene coding for phosphatidylserine decarboxylase. J Bacteriol. 1998;180:100–6.
Nishibori A, Kusaka J, Hara H, Umeda M, Matsumoto K. Phosphatidylethanolamine domains and localization of phospholipid synthases in Bacillus subtilis membranes. J Bacteriol. 2005;187:2163–74.
Kawai F, et al. Cardiolipin domains in Bacillus subtilis marburg membranes. J Bacteriol. 2004;186:1475–83.
Kobayashi K, et al. Essential Bacillus subtilis genes. Proc Natl Acad Sci USA. 2003;100:4678–83.
Vlamakis H, Chai Y, Beauregard P, Losick R, Kolter R. Sticking together: building a biofilm the Bacillus subtilis way. Nat Rev Microbiol. 2013;11:157–68.
Bosak T, Losick RM, Pearson A. A polycyclic terpenoid that alleviates oxidative stress. Proc Natl Acad Sci USA. 2008;105:6725–9.
Kontnik R, et al. Sporulenes, heptaprenyl metabolites from Bacillus subtilis spores. Org Lett. 2008;10:3551–4.
Kohanski MA, Dwyer DJ, Collins JJ. How antibiotics kill bacteria: from targets to networks. Nat Rev Microbiol. 2010;8: 423–35.
Ruhr E, Sahl HG. Mode of action of the peptide antibiotic nisin and influence on the membrane potential of whole cells and on cytoplasmic and artificial membrane vesicles. Antimicrob Agents Chemother. 1985;27:841–5.
Silhavy TJ, Kahne D, Walker S. The bacterial cell envelope. Cold Spring Harb Perspect Biol. 2010;2:a000414.
Rashid R, Veleba M, Kline KA. Focal targeting of the bacterial envelope by antimicrobial peptides. Front Cell Dev Biol. 2016;4:55.
Lihu Y, Breukink E. The membrane steps of bacterial cell wall synthesis as antibiotic targets. Antibiot (Basel, Switz). 2016;5:E28.
Epand RM, Walker C, Epand RF, Magarvey NA. Molecular mechanisms of membrane targeting antibiotics. Biochim Biophys Acta. 2016;1858:980–7.
Müller A, et al. Daptomycin inhibits cell envelope synthesis by interfering with fluid membrane microdomains. Proc Natl Acad Sci USA, https://doi.org/10.1073/pnas.1611173113 (2016).
Lehtonen JY, Holopainen JM, Kinnunen PK. Evidence for the formation of microdomains in liquid crystalline large unilamellar vesicles caused by hydrophobic mismatch of the constituent phospholipids. Biophys J. 1996;70:1753–60.
Acknowledgements
We thank David Forgacs, Abby Korn and Yicheng Xie (Texas A&M University [TAMU]) for assistance related to this project. We thank Min Woo Sung (TAMU Microscopy and Imaging Center) for transmission electron microscopy. We thank Ry Young (TAMU) for use of the plate reader. We thank Jan Maarten van Dijl (University of Groningen), John Helman (Cornell University), Jennifer Herman (TAMU), and Daniel Ziegler (Bacillus Genetic Stock Center) for strains. This work was supported by the National Science Foundation (NSF-CAREER Award MCB-1253215) to Paul D. Straight.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Stubbendieck, R.M., Brock, D.J., Pellois, JP. et al. Linearmycins are lytic membrane-targeting antibiotics. J Antibiot 71, 372–381 (2018). https://doi.org/10.1038/s41429-017-0005-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-017-0005-z