Abstract
There are numerous approaches to improve the low transduction efficiency of retroviral vectors in two-dimensional (2D) cell culture substrates. However, the effect of a three-dimensional (3D) microenvironment, which better mimics in vivo conditions, is unknown. Cytocompatible hyaluronic acid (HA) hydrogels are a good candidate to study this issue. Here, photocrosslinkable HA hydrogels with an elastic modulus of 1.0–2.7 kPa were successfully prepared by varying the degree of methacrylation in the HA backbone. Culturing human adipose-derived stem cells (hASCs) in a 3D microenvironment significantly reduces the amount of time required for retroviral gene transduction compared with the conventional 2D method and maintains a high transduction efficiency. This acceleration of retroviral gene transduction correlates with the rate of cell-cycle synchronization. hASCs cultured in a 3D microenvironment have a shorter G1 phase and total cell-cycle length than hASCs cultured using the conventional 2D method. This cell-cycle regulation is dependent on expression of cyclin D1. In summary, prior culturing of hASCs in a 3D microenvironment accelerates retroviral gene transduction by regulating cyclin D1 expression and accelerating cell-cycle synchronization. We conclude that priming via culturing in a 3D microenvironment facilitates efficient and rapid retroviral gene transduction of hASCs without inducing apoptosis.
Similar content being viewed by others
Introduction
Mesenchymal stem cells (MSCs), which can undergo self-renewal and are multipotent, have been extensively studied as therapeutic agents for various disorders due to their regenerative properties1,2. According to a report by the National Institutes of Health in the United States, more than 770 clinical trials of MSCs are ongoing or have been completed worldwide as of April 2018 (http://www.clinicaltrials.gov). Genetic modifications of MSCs aimed at improving their therapeutic efficacies have been studied extensively. Current strategies for MSC-based therapies involve introducing genes that enhance the therapeutic potential of these cells using various non-viral and viral vectors3.
Widely used non-viral vector systems include cationic liposomes and polymers, nanoparticles, and electroporation. Transfection of MSCs with the forkhead box A2 (Foxa2) gene using the commercially available cationic lipid Lipofectamine 2000 enhances liver tissue regeneration4. Calcium phosphate nanoparticles coated with highly cationic glutamine-conjugated oligochitosan significantly enhance the transfection of MSCs with Noggin-targeting small interfering RNA and osteogenesis of these cells5. Transfection of SOX5/SOX6/SOX9 and BMP-2 genes via electroporation enhances chondrogenesis and osteogenesis of MSCs, respectively6,7. Although non-viral vectors are easy to use and do not elicit a marked immune response in the host, they have a low intracellular gene transfer efficiency and are not suitable for transfection of large populations of cells. Lentiviruses, adenoviruses, adeno-associated viruses, herpes virus, pox virus, human foamy virus, and retroviruses are currently used as viral gene delivery systems8,9. Among these, retroviral vectors, which were the first vectors to be employed for gene therapy, have been used in several clinical trials. Such clinical trials include gene therapy for severe combined immunodeficiency (SCID)-X1 disease using CD34+ hematopoietic stem cells transduced with murine leukemia virus-based retroviral vectors encoding the adenosine deaminase gene10. Human stem cells transduced with retroviral vectors have also been successfully used to treat patients with cancer and multiple congenital disorders9,11. Retroviral vectors stably integrate into the genome of host cells and are transmitted to all the progeny of these cells12. In addition, the target cell specificity of retroviral vectors can be easily altered by pseudotyping with heterologous envelope proteins13. However, only dividing cells are transduced with retroviruses. Consequently, the transduction efficiency of retroviral vectors in human stem cells is low, which is a major limitation for their use in stem-cell therapies14,15. Therefore, many researchers have sought to enhance the efficiency of retrovirus infection via various approaches, including cell-cycle regulation16,17,18. Unfortunately, cell-cycle regulation usually takes a long time and requires repeated stimulations, which may lead to loss of the regenerative capacity of the stem cells and decrease their viability15,16.
Numerous studies have demonstrated that environmental dimensionality plays an important role in cell behavior. Cells cultured in three-dimensional (3D) environments markedly differ from those cultured on conventional two-dimensional (2D) tissue culture plates. For instance, the phenotype and gene expression of articular chondrocytes are maintained upon culture in a 3D microenvironment but are lost upon culture in a 2D monolayer19. Cardiac fibroblasts remain relatively quiescent in the 3D tissue but are activated after being isolated from the heart and begin to proliferate and differentiate into myofibroblasts upon 2D culture20. Breast epithelial cancer cells do not produce physiologically relevant multicellular spheroidal structures when cultured on 2D tissue culture plates but form 3D breast cancer spheroids when cultured in a 3D environment21,22. Most studies investigating whether cell-cycle regulation enhances retroviral gene transduction have been performed in 2D environments. Ishihara’s group recently showed that 3D culture in hydrogels regulates the cell cycle. Briefly, murine MSCs encapsulated in hydrogels composed of a phospholipid polymer arrest at the G1 phase after 1–3 days, and this modulates their osteogenic differentiation23,24. However, it is unclear if culture in a 3D microenvironment regulates the cell cycle of human primary MSCs and, if so, whether such priming alters the retroviral gene transduction efficiency. Here, we report a novel strategy to regulate the cell cycle via culturing in a chemically defined 3D microenvironment, namely, a photocrosslinkable hyaluronic acid (HA) hydrogel and to evaluate the effect of this priming on retroviral gene transduction in human primary MSCs.
Materials and methods
Synthesis of photocrosslinkable polymers
Methacrylated HA (MAHA) was synthesized as previously reported25. Briefly, to prepare MAHA with a degree of substitution (DS) of 50%, methacrylic anhydride (8 mL, Sigma-Aldrich, St. Louis, MO) was added to a 1% w/v HA (Mw 500 kDa, Bioland, Korea) aqueous solution. The reaction mixture was allowed to react for 24 h with stirring at 4 °C after adjusting the solution pH to 8.0 using a 5 N NaOH solution. The reaction mixture was dialyzed against deionized water (DW) using a membrane with a cutoff molecular weight of 100 kDa for 3 days followed by lyophilization. 1H-NMR spectroscopy was used to determine the DS on HA. The DS was calculated from the ratio of the relative peak integrations of the methacrylate protons (peaks at ~6.1, 5.6, and 1.85 ppm) and methyl protons of HA (~1.9 ppm).
Methacrylated gelatin was prepared following the previously developed method26. Briefly, gelatin type B (4 g, Sigma-Aldrich, St. Louis, MO) was dissolved in phosphate-buffered saline (PBS, 10% w/v) at 60 °C and functionalized with methacrylic anhydride (3 mL) at pH 7.5 at 50 °C for 1 h. The reaction was then stopped by adding pre-warmed PBS. After dialysis against DW and lyophilization, the DS of methacrylated gelatin was measured as 90% using 1H-NMR spectroscopy.
Preparation of hydrogels
To prepare HA, gelatin, poly(ethylene glycol) (PEG) hydrogels, and polymer solutions of MAHA (0.5% w/v), methacrylated gelatin (7% w/v) and PEG diacrylate (Sigma-Aldrich, St. Louis, MO, 10% w/v) in PBS with Irgacure 2959 (final concentration 0.2% w/v) were prepared, and 40 μL of each polymer solution was placed in polydimethylsiloxane molds having a diameter of 5 mm and a depth of 2 mm and were exposed to UV light (365 nm, 60 mW/cm2, Sei Myung Vactron Co. LTD, Korea) for 10 s for MAHA and 50 s for methacrylated gelatin and PEG diacrylate.
Mechanical characterization of hydrogels
To characterize the mechanical properties of various types of hydrogel, we used a customized bulk-scale indenter consisting of a load cell (GS0-10, Transducer Techniques) and an automated stage (SM2-0803-3S and SZ-0604-3S, ST1) equipped with a microscope (AM4113, AnMo Electronics Corporation). A stainless steel spherical tip with a 2 mm diameter was attached to the load cell for indentation. The contact between the tip and the hydrogel was assumed to be Hertzian contact27. The maximum indentation depth was ~5% of the hydrogel thickness, and the indentation speed was 25 μm s−1. Both the applied force and the indentation depth were recorded at an acquisition rate of 10 Hz during the experiment. The effective modulus of the hydrogel was estimated by fitting a force-indentation curve to the Hertzian model given by equation (1)
where F is the applied force, E is the Young’s modulus of the hydrogel, ν is the Poisson’s ratio of the hydrogel, which is assumed to be 0.5, r is the radius of the indentation tip, and δ is the indentation depth.
Isolation and expansion of hASCs
hASCs were isolated from adipose tissues in the articular fat pads around the patients’ knee28 with approval of the Ethics Committee at CHA University (IRB No. 2014-07-096). In brief, the adipose tissues were washed three times with phosphate-buffered saline (PBS) containing 2% v/v penicillin/streptomycin and digested using 0.5 mg/mL of collagenase type II (Sigma-Aldrich, St. Louis, MO) in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone, Logan, UT) for 45 min at 37 °C with constant shaking. Digested tissues were collected in 50 mL conical tube, centrifuged (1000 × g, 10 min), and then supernatant fat was discarded, and the infranatant fluid was filtered through a cell strainer (pore size 40 μm). The filtered fluid was washed three times with DMEM, centrifuged (1000 × g, 10 min), and the cell pellet was resuspended with growth medium (DMEM supplemented with 20% v/v fetal bovine serum (FBS), and 1% v/v penicillin/streptomycin), and the cells were cultured on tissue culture plates at 37 °C in a 5% CO2 incubator. The culture medium was changed every 3 days.
Retroviral transduction
The day before transduction, Gp2 cells (Clontech) were seeded at a concentration of 3 × 106 cells per 100 mm dish. pMX-GFP (Addgene, Cambridge, MA) was transfected with retrovirus packaging vector VSV-G (Invitrogen) and Convoy transfection reagent (ACTGene Inc., Piscataway, NJ). A day after transfection, the medium was replaced with 10 mL of fresh media. After 48 h of transfection, the supernatants were collected, centrifuged at 1300 rpm for 3 min, and filtered with a 0.45 μm filter (Millipore). Then, retro-X concentrator was added at a volume equal to 1/3 of the media. The mixture was incubated at 4 °C overnight with constant inversion. After centrifugation at 1500 × g for 45 min, the concentrated pellet was resuspended with fresh hASC culture media. A day before infection, cell cycle regulated hASCs from the 2D and 3D systems were seeded into 6-well plates at a density of 3 × 104 cells per well. For infection, 2 mL of concentrated retrovirus and 1 μL mL−1 polybrene (final concentration 8 μg mL−1; Sigma-Aldrich) were added to the cells. After incubation for 24 h, the medium was replaced with 2 mL of fresh growth medium. Three days after, green fluorescent protein (GFP) fluorescence images of the infected cells were observed with a Nikon TE2000-U fluorescence microscope (Nikon, Japan).
Cell-cycle regulation on 2D tissue culture plate
To determine the cell-cycle regulation on 2D tissue culture polystyrene (TCPS) plates, we followed the conventional cell cycle regulating method of serum starvation followed by the addition of serum16,29,30. For cell cycle arrest at the G1 phase, hASCs were seeded at a density of 3 × 105 cells per well in a 6-well plate and cultured with growth medium for 24 h. Cells were washed with PBS, and serum-free medium (0.5% v/v FBS) was added. After serum starvation for 1, 3, 6, 12, 24, or 48 h, cells were passaged and released into cell cycle by the addition of growth medium.
3D primed cell-cycle regulation
For 3D primed cell cycle arrest, harvested hASCs were suspended in MAHA solution (final concentration 0.5% w/v) at a density of 2 × 106 cells mL−1, UV irradiated in the presence of Irgacure 2959 (final concentration 0.2% w/v), and cultured in growth medium at 37 °C in a 5% CO2 incubator. After 0.5–24 h of incubation, the cells were released from the hydrogels by treatment with hyaluronidase (2000 U mL−1 in growth media, Sigma-Aldrich) for 40 min at 37 °C. Released hASCs from the 3D physical cue were seeded on tissue culture plates and cultured with growth medium.
Cell-cycle analysis
At each time point, cells were collected and stained with propidium iodide (PI, Sigma-Aldrich). Briefly, cells were harvested by treatment with trypsin or hyaluronidase in the case of the 2D or 3D system, respectively. Cells were collected and washed with PBS by centrifugation at 1000 × g. Collected cells were fixed with cold 70% ethanol for at least 10 min and washed with PBS three times. Fixed cells were resuspended in 200 μL of PBS containing 100 μg mL−1 of RNase A (Thermo ScientificTM, Lithuania, EU) and 10 μL of PI solution (1 mg mL−1), and the cell suspensions were then held for 15 min in the dark at room temperature. The DNA content of individual cells was examined by flow cytometry (BD AccuriTM C6, BD Biosciences, San Jose, CA).
Apoptosis assay
FITC Annexin V and PI double labeling was used to determine the apoptosis-inducing effect of the cell cycle arrest at the G1 phase. Briefly, cells were harvested by treatment with trypsin 24 and 48 h after 2D arrest or with hyaluronidase 1 h after 3D primed arrest. Collected cells were resuspended in 100 μL of 1 × Annexin V binding buffer (BioLegend, San Diego, CA). Then, 5 μL of FITC Annexin V (100 μg mL−1, BioLegend, San Diego, CA) and 5 μL of PI solution (1 mg mL−1, Sigma-Aldrich) were added to the cells. Fifteen minutes after incubation in the dark at room temperature, the stained cells were washed with PBS three times. The percentage of apoptotic cells was measured by flow cytometry (BD AccuriTM C6, BD Biosciences).
Gene expression analysis
To confirm cyclin D1 gene expression, the total cellular RNA was extracted using TRizol (Ambion) reagent. PrimeScript RT Reagent Kit (Perfect Real Time, TAKARA) was used to synthesize complementary DNA from 1 μg of the total RNA. For quantification of gene expression level, qRT-PCR was performed using Power SYBER Green PCR Master Mix (Applied Biosystems) with a StepOnePlus Real-Time PCR System (Applies Biosystems). The PCR condition was 40 cycles at 95 °C for 15 min and 60 °C for 1 min, and the melting curve stage was 95 °C for 15 min and 60 °C for 1 min. The PCR primer sequences are shown in Supplementary Table S1. Target gene expression was normalized to the 18 s (housekeeping gene) gene for quantification.
Statistical analysis
All statistical analyses were implemented with Graphpad prism ver. 5.0 (Graphpad software, San Diego, CA). One-way ANOVAs with Tukey’s multiple comparison posttest were implemented to compare the samples. Statistical significance was set at p < 0.05.
Results and discussion
Characterization of hydrogels
Cells cultured in 3D environments have different biological and phenotypic characteristics than those cultured in 2D environments19,20,21,22. A recent study reported that encapsulation in a 3D microenvironment accelerates cellular reprogramming by inducing acquisition of a cell size and morphology similar to those of induced pluripotent stem cells and by promoting mesenchymal-to-epithelial transition and epigenetic plasticity31. In this study, we employed photocrosslinkable HA hydrogels to investigate whether priming of primary human adipose-derived stem cells (hASCs) via culturing in a 3D microenvironment (hereafter referred to as 3D priming) affects the retroviral gene transduction efficiency.
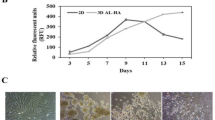
The chemical functionalities present in HA, such as carboxyl and hydroxyl groups, can be modified. We introduced methacrylate groups into the HA to provide photocrosslinkability by reacting hydroxyl groups with methacrylic anhydride (Fig. 1a). 1H-NMR confirmed that the degree of substitution (DS) of methacrylate groups onto HA was 50% (Fig. 1b). The MAHA solution (0.5% w/v in PBS) showed a rapid sol-to-gel transition after 10 s of UV irradiation in the presence of Irgacure 2959 (0.2% w/v), which was used as a photoinitiator (Fig. 1c). The elastic modulus of the hydrogels was 1.7 kPa. Radical polymerization was potentially cytotoxic32; therefore, the viability of hASCs in HA hydrogels was evaluated (Fig. 1d). hASCs had a high viability (>99%) and were uniformly distributed in the HA hydrogel after 1 day. In addition, the cell viability of the hASCs was maintained for 7 days of culture, indicating the additional ability of this hydrogel system to be a culture platform for cells (Supplementary Fig. S1).
Effect of 3D priming on retroviral gene transduction
To assess whether 3D priming accelerates retroviral gene transduction, we generated a green fluorescent protein (GFP)-expressing retroviral vectors and evaluated the transduction efficiency through a statistical analysis of the fluorescence microscope images and FACS data. We encapsulated hASCs for 1 h in HA hydrogels to provide an initial 3D physical cue (3D primed arrest) and then removed them from the hydrogels (3D primed release) for 1 h. The transduction efficiency of GFP was compared between cells exposed to 3D priming and those cultured using the conventional 2D method. In the latter method, hASCs were cultured in the absence of serum for 24 h (2D arrest) and then in the presence of serum for 24 h (2D release). As expected, the relative GFP expression significantly increased when hASCs were infected with the GFP-expressing retroviral vector following cell-cycle synchronization by the conventional 2D method (Fig. 2a, b, 3.3-fold higher than in the control group). While the conventional 2D method took 48 h, a comparable effect was induced via 3D priming in only 2 h (the percentage of GFP-positive cells was 4.1-fold higher than in the control group). By contrast, priming via culturing using the conventional 2D method for 2 h (1 h of 2D arrest followed by 1 h of 2D release) was ineffective (Supplementary Fig. S2). Quantitative analysis by FACS demonstrated that GFP-positive cells were 40.4% in the unsynchronized control group and significantly increased up to 56.8% and 63.1% of GFP-positive cells that were achieved through the conventional 2D method and 3D priming, respectively (Fig. 2c). These results suggest that spatial confinement of cells in the 3D microenvironment accelerates retroviral gene transduction.
Cell-cycle synchronization of hASCs in 2D and 3D microenvironments
Our previous experiments showed that 3D priming significantly accelerated retroviral gene transduction. Positive regulation of cell-cycle progression from the G1 to S phase (G1/S transition) increases the retroviral transduction efficiency16,17,18. Therefore, we investigated the relationship between the acceleration of retroviral gene transduction by 3D priming and the rate of cell-cycle synchronization. To this end, we compared the amount of time required for cell-cycle synchronization in 2D and 3D microenvironments (Fig. 3a). The cell cycle was synchronized by serum starvation (2D arrest) followed by serum addition (2D release), which is the most widely used method for cell-cycle synchronization in 2D environments33. We monitored the cell cycle of hASCs cultured in the absence or presence of serum. Consistent with previous reports16,33, PI staining showed that the percentage of hASCs in the G1 phase increased with the duration of serum starvation (Fig. 3b and Supplementary Fig. s3a) Specifically, 86.6% and 92.7% cells were in the G1 phase after serum starvation for 24 and 48 h, respectively. Serum starvation induces apoptosis of MSCs34; therefore, we monitored apoptosis of hASCs according to the duration of serum starvation. The percentage of apoptotic cells did not increased after serum starvation for 24 h but increased significantly after serum starvation for 48 h (Supplementary Fig. S4). We therefore performed serum starvation for 24 h in subsequent experiments to avoid the potential negative effects of prolonged nutrient deprivation.
a Schematic representation of the conventional 2D and 3D priming protocol. b Induction of G1 phase arrest following serum starvation upon 2D culture in a polystyrene tissue culture plate (2D arrest). c Release into the cell cycle following addition of serum upon 2D culture in a polystyrene tissue culture plate (2D release). d Induction of G1 phase arrest following encapsulation of cells in MAHA hydrogels (3D primed arrest). e Release into the cell cycle following removal of cells from hydrogels (3D primed release)
After serum starvation for 24 h, hASCs were released into the cell cycle by adding serum to the media. The addition of serum increased the percentages of hASCs in the S and G2/M phases. Cells had progressed from the G1 phase to the S phase 20 h after release into the cell cycle, and the percentages of cells in the S/G2/M phases peaked at 24 h (Fig. 3c and Supplementary Fig. s3b). The percentage of hASCs in the S phase decreased and that in the G2/M phases peaked 26 h after release, and then, cells re-entered the G1 phase.
We next evaluated the effect of 3D priming on cell-cycle regulation by encapsulating hASCs in HA hydrogels (3D primed arrest) and then removing them from the hydrogels (3D primed release). In total, 85.4% of hASCs were in the G1 phase after culture in HA hydrogels for 1 h (Fig. 3d and Supplementary Fig. s3c), which was 23 h less than the amount of time required to arrest cells in the G1 phase using the serum starvation method in a 2D microenvironment. The percentage of apoptotic cells after 1 h of culture in HA hydrogels did not significantly differ from that in the untreated control (Supplementary Fig. S4). After 1 h of 3D primed arrest of hASCs followed by 3D primed release, cells re-entered the S phase, and the percentage of cells in the S phase peaked at 1 h (Fig. 3e and Supplementary Fig. s3d), which was 23 h faster than the amount of time required using the serum addition method in a 2D microenvironment. In addition, acceleration of G1 arrest and G1/S transition by 3D priming shortened the cell-cycle length of hASCs.
This acceleration of G1 arrest by 3D priming might be due to a change in cellular morphology. hASCs in 3D microenvironments were spherical (Fig. 1d). Cellular morphology is closely related to the cell cycle. Round (poorly spread) primary human pulmonary capillary endothelial cells grown on a surface coated with a low density of fibronectin (25 ng cm−2) fail to progress through the G1 phase35. Mesenchymal-to-epithelial morphological changes of human cancer cells upon ectopic expression of microRNA-147 also induce cell cycle arrest at the G1 phase36.
These results suggest that cell-cycle regulation via 3D priming using hydrogels is a fast and efficient tool to enhance the retroviral gene transduction efficiency in primary hASCs in comparison with the conventional 2D method.
Mechanism by which 3D priming synchronizes the cell cycle
We next sought to elucidate the mechanism that underlies the significant acceleration of cell-cycle synchronization by 3D priming in comparison with the conventional 2D method. Cyclin D1 expression is the rate-determining step for G1/S transition in various rat fibroblast lines37. Decreased and increased expression of cyclin D1 is associated with arrest in the G1 phase and accelerated entry into the S phase, respectively38,39. Cyclin D binds to and activates cyclin dependent kinase 4 and 6 (Ckd4/6), which is essential for transition into the S phase, cell proliferation, and efficient transduction40. Here, cyclin D1 gene expression was measured after G1 arrest and release into the cell cycle in the 2D and 3D microenvironments to determine whether cell-cycle synchronization by 3D priming regulates cyclin D1 expression (Fig. 4a). Cyclin D1 gene expression was significantly downregulated by 24 h of 2D arrest and 1 h of 3D primed arrest, which induced synchronization of hASCs in the G1 phase (Fig. 4b). By contrast, cyclin D1 gene expression was significantly upregulated when hASCs were released into the cell cycle by culture in the presence of serum for 24 h in the 2D system (2D release) and by removal from the 3D microenvironment and culture for 1 h (3D primed release). Specifically, cyclin D1 expression was decreased to 2.95- and 27.4-fold after 2D arrest and 3D primed arrest, respectively, but was increased to 2.20- and 3.86-fold 2D release and 3D primed release, respectively. These results indicate that expression of cyclin D1 is closely associated with cell-cycle regulation induced by 3D priming (Fig. 4c), as reported in previous studies of cells cultured in 2D environments37,38,39,40.
a Schematic representation of the optimized conventional 2D and 3D priming protocol. b Regulation of Cyclin D1 gene expression upon cell cycle arrest and release. ###p < 0.001, ####p < 0.0001 compared with the untreated control and **p < 0.01, ****p < 0.0001. c Proposed mechanism of accelerated cell-cycle regulation via 3D priming
Effects of the elastic modulus and hydrogel material on cell-cycle synchronization
To investigate the effect of microenvironmental stiffness on G1 arrest and re-entry into the cell cycle, we prepared MAHA with methacrylate DS of 30%, 50%, and 100% (termed DS30, DS50, and DS100, respectively) by varying the amount of methacrylic anhydride and the reaction duration (Fig. 5a). The elastic moduli of DS30, DS50, and DS100 HA hydrogels were 1.0, 1.7, and 2.7 kPa, respectively (Fig. 5b). hASCs encapsulated in DS30, DS50, and DS100 HA hydrogels showed similar cell-cycle synchronization patterns. Specifically, 76.9%, 84.8%, and 77.8% of hASCs arrested in the G1 phase after culture in DS30, DS50, and DS100 HA hydrogels for 1 h, respectively (Fig. 5c and Supplementary Fig. S5). In addition, re-entry of hASCs into the cell cycle did not significantly differ after they were removed from these various hydrogels (Fig. 5d and Supplementary Fig. S5). Our results indicate that HA hydrogels with a compressive elastic modulus ranging from 1.0 to 2.7 kPa induced cell-cycle synchronization of hASCs comparably, which is in contrast to a previous report using other type of cells and other hydrogel systems23.
a 1H-NMR (in D2O) of MAHA with different DS. b Elastic modulus of HA hydrogels with different DS (n = 14–18; ****p < 0.0001). c Arrest of hASCs in the G1 phase upon culture in HA hydrogels with different DS. d Progression of hASCs into the S phase over time following removal from HA hydrogels with different DS
To investigate the effect of hydrogel materials on cell-cycle regulation, we prepared cytocompatible gelatin and PEG hydrogels with the same elastic modulus as the DS50 HA hydrogel. The percentage of hASCs arrested in the G1 phase did not significantly differ between these hydrogels (Supplementary Fig. S6). Next, we compared re-entry into the cell cycle between hASCs removed from gelatin hydrogels and those removed from HA hydrogels (Supplementary Fig. S7). Cells had progressed from the G1 phase to the S phase 1 h after removal from gelatin hydrogels, and the percentages of cells in the S and G2/M phases peaked at 3 h. This was 2 h slower than observed following removal from HA hydrogels but much faster than observed with the conventional 2D method. Our results suggest that cell-cycle regulation via 3D priming of hASCs is independent of changes in the materials of hydrogels, such as gelatin and PEG, and the change in the elastic modulus of the HA hydrogels in the determined range from 1.0 to 2.7 kPa.
Conclusions
To the best of our knowledge, our findings are the first evidence that 3D priming accelerates retroviral gene transduction. Our results show that 3D priming by encapsulating hASCs in HA hydrogels reduces the amount of time required for retroviral gene transduction in comparison with the conventional 2D method. This effect closely correlated with the acceleration of cell-cycle regulation (G1 arrest and G1/S transition) by 3D priming. hASCs arrested in the G1 phase within 1 h when encapsulated in photocrosslinkable HA hydrogels in the absence of any additional chemical or physical stimuli. Upon removal from HA hydrogels, hASCs progressed more rapidly from the G1 to S phase than cells cultured in the presence of serum following serum starvation using the conventional 2D method. In addition, cell-cycle regulation via 3D priming was independent of the hydrogel stiffness and material and was closely related to cyclin D1 expression. This study describes a new method to accelerate retroviral gene transduction while maintaining a high efficiency, which is important for stem-cell therapy.
References
Boregowda, S. V. & Phinney, D. G. Therapeutic applications of mesenchymal stem cells: current outlook. BioDrugs 26, 201–208 (2012).
Larsen, S. & Lewis, I. D. Potential therapeutic applications of mesenchymal stromal cells. Pathology 43, 592–604 (2011).
Oggu, G. S. et al. Gene delivery approaches for mesenchymal stem cell therapy: strategies to increase efficiency and specificity. Stem Cell Rev. Rep. 13, 725–740 (2017).
Cho, J. W., Lee, C. Y. & Ko, Y. Therapeutic potential of mesenchymal stem cells overexpressing human forkhead box A2 gene in the regeneration of damaged liver tissues. J. Gastroenterol. Hepatol. 27, 1362–1370 (2012).
Choi, B. et al. Glutamine-chitosan modified calcium phosphate nanoparticles for efficient siRNA delivery and osteogenic differentiation. J. Mater. Chem. B 3, 6448–6455 (2015).
Kim, H. J. & Im, G. I. Electroporation-mediated transfer of SOX trio genes (SOX-5, SOX-6, and SOX-9) to enhance the chondrogenesis of mesenchymal stem cells. Stem Cells Dev. 20, 2103–2114 (2011).
Ferreira, E. et al. Sustained and promoter dependent bone morphogenetic protein expression by rat mesenchymal stem cells after BMP-2 transgene electrotransfer. Eur. Cells Mater. 24, 18–28 (2012).
Huang, Y. et al. Development of viral vectors for gene therapy for chronic pain. Pain. Res. Treat. 2011, 968218 (2011).
Nayerossadat, N., Maedeh, T. & Ali, P. A. Viral and nonviral delivery systems for gene delivery. Adv. Biomed. Res. 1, 27 (2012).
Cavazzana-Calvo, M. et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288, 669–672 (2000).
Havenga, M., Hoogerbrugge, P., Valerio, D. & van Es, H. H. Retroviral stem cell gene therapy. Stem Cells 15, 162–179 (1997).
Varmus, H. E. Form and function of retroviral proviruses. Science 216, 812–820 (1982).
Russell, S. J. & Cosset, F. L. Modifying the host range properties of retroviral vectors. J. Gene Med. 1, 300–311 (1999).
Miller, D. G., Adam, M. A. & Miller, A. D. Gene transfer by retrovirus vectors occurs only in cells that are actively replicating at the time of infection. Mol. Cell. Biol. 10, 4239–4242 (1990).
Trobridge, G. & Russell, D. W. Cell cycle requirements for transduction by foamy virus vectors compared to those of oncovirus and lentivirus vectors. J. Virol. 78, 2327–2335 (2004).
Chen, M. et al. Serum starvation induced cell cycle synchronization facilitates human somatic cells reprogramming. PLoS One 7, e28203 (2012).
Liu, H., Hung, Y., Wissink, S. D. & Verfaillie, C. M. Improved retroviral transduction of hematopoietic progenitors by combining methods to enhance virus-cell interaction. Leukemia 14, 307–311 (2000).
Kwon, Y. J. & Peng, C. A. Impact of cell growth morphology on retroviral transduction: effect of contact inhibition. Biotechnol. Prog. 17, 240–246 (2001).
Choi, B. G. et al. In situ thermal gelling polypeptide for chondrocytes 3D culture. Biomaterials 31, 9266–9272 (2010).
Zuppinger, C. 3D culture for cardiac cells. Biochim. Biophys. Acta 1863, 1873–1881 (2016).
Debnath, J. & Brugge, J. S. Modelling glandular epithelial cancers in three-dimensional cultures. Nat. Rev. Cancer 5, 675–688 (2005).
Baker, A. E. G., Tam, R. Y. & Shoichet, M. S. Independently tuning the biochemical and mechanical properties of 3D hyaluronan-based hydrogels with oxime and Diels-Alder chemistry to culture breast cancer spheroids. Biomacromolecules 18, 4373–4384 (2017).
Oda, H., Konno, T. & Ishihara, K. The use of the mechanical microenvironment of phospholipid polymer hydrogels to control cell behavior. Biomaterials 34, 5891–5896 (2013).
Oda, H., Konno, T. & Ishihara, K. Efficient differentiation of stem cells encapsulated in a cytocompatible phospholipid polymer hydrogel with tunable physical properties. Biomaterials 56, 86–91 (2015).
Smeds, K. A. et al. Photocrosslinkable polysaccharides for in situ hydrogel formation. J. Biomed. Mater. Res. 54, 115–121 (2001).
Van Den Bulcke, A. I. et al. Structural and rheological properties of methacrylamide modified gelatin hydrogels. Biomacromolecules 1, 31–38 (2000).
Hertz, H., Jones, D. E. & Schott, G. A. Miscellaneous Papers. (Macmillan and co., 1896).
Arai, Y. et al. Enhancement of matrix metalloproteinase-2 (MMP-2) as a potential chondrogenic marker during chondrogenic differentiation of human adipose-derived stem cells. Int. J. Mol. Sci. 17, 963 (2016).
Attardi, L. D., de Vries, A. & Jacks, T. Activation of the p53-dependent G1 checkpoint response in mouse embryo fibroblasts depends on the specific DNA damage inducer. Oncogene 23, 973–980 (2004).
Masutomi, K. et al. Telomerase maintains telomere structure in normal human cells. Cell 114, 241–253 (2003).
Caiazzo, M. et al. Defined three-dimensional microenvironments boost induction of pluripotency. Nat. Mater. 15, 344–352 (2016).
Williams, C. G., Malik, A. N., Kim, T. K., Manson, P. N. & Elisseeff, J. H. Variable cytocompatibility of six cell lines with photoinitiators used for polymerizing hydrogels and cell encapsulation. Biomaterials 26, 1211–1218 (2005).
Kues, W. A. et al. Cell cycle synchronization of porcine fetal fibroblasts: effects of serum deprivation and reversible cell cycle inhibitors. Biol. Reprod. 62, 412–419 (2000).
Zhu, W., Chen, J., Cong, X., Hu, S. & Chen, X. Hypoxia and serum deprivation-induced apoptosis in mesenchymal stem cells. Stem Cells 24, 416–425 (2006).
Sui, H., Christopher, S. C., Donald, E. I. & Richard, O. H. Control of cyclin D1, p27Kip1, and cell cycle progression in human capillary endothelial cells by cell shape and cytoskeletal tension. Mol. Biol. Cell 9, 3179–3193 (1998).
Lee, C. G., McCarthy, S., Gruidl, M., Timme, C. & Yeatman, T. J. MicroRNA-147 induces a mesenchymal-to-epithelial transition (MET) and reverses EGFR inhibitor resistance. PLoS One 9, e84597 (2014).
Resnitzky, D. & Reed, S. I. Different roles for cyclins D1 and E in regulation of the G1-to-S transition. Mol. Cell. Biol. 15, 3463–3469 (1995).
Canales, J. et al. Helicobacter pylori induced phosphatidylinositol-3-OH kinase/mTOR activation increases hypoxia inducible factor-1α to promote loss of cyclin D1 and G0/G1 cell cycle arrest in human gastric. Cells Front. Cell. Infect. Microbiol. 7, 92 (2017).
Resnitzky, D., Gossen, M., Bujard, H. & Reed, S. I. Acceleration of the G1/S phase transition by expression of cyclins D1 and E with an inducible system. Mol. Cell. Biol. 14, 1669–1679 (1994).
Thornton, T. M. & Rincon, M. Non-classical p38 map kinase functions: cell cycle checkpoints and survival. Int. J. Biol. Sci. 5, 44–51 (2009).
Acknowledgements
This study was supported by the National Research Foundation of Korea (NRF) Grants funded by MSIP (NRF-2016R1A2A1A05004987) and MEST (NRF-2014R1A6A3A04055123).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, Y., Arai, Y., Ahn, J. et al. Three-dimensional microenvironmental priming of human mesenchymal stem cells in hydrogels facilitates efficient and rapid retroviral gene transduction via accelerated cell cycle synchronization. NPG Asia Mater 11, 27 (2019). https://doi.org/10.1038/s41427-019-0127-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41427-019-0127-9