Abstract
Malaria is one of the deadliest infectious diseases in the world. Immune responses to Plasmodium falciparum malaria vary among individuals and between populations. Human genetic variation in immune system genes is likely to play a role in this heterogeneity. Natural killer (NK) cells produce inflammatory cytokines in response to malaria infection, kill intraerythrocytic Plasmodium falciparum parasites by cytolysis, and participate in the initiation and development of adaptive immune responses to plasmodial infection. These functions are modulated by interactions between killer-cell immunoglobulin-like receptors (KIRs) and human leukocyte antigens (HLAs). Therefore, variations in KIR and HLA genes can have a direct impact on NK cell functions. Understanding the role of KIRs and HLAs in immunity to malaria can help to better characterize antimalarial immune responses. In this review, we summarize the different KIRs and HLAs associated with immunity to malaria thus far.
Similar content being viewed by others
Introduction
Malaria represents one of the most serious infectious disease challenges in the world.1 Malaria burden is greatly affected by immunity,2,3 especially in populations with moderate to high-transmission intensity.4 Partial immunity to malaria develops over years of exposure.5 Although this partial immunity does not provide complete protection, it reduces the risk of parasitemia progressing to clinical illness and severe disease.6 This explains why most malaria deaths in high-transmission areas, such as much of sub-Saharan Africa, occur in children.7 Several studies have consistently shown that a number of human genetic variants are associated with protection against uncomplicated and severe P. falciparum malaria. Hemoglobin S heterozygous individuals are protected against uncomplicated and severe malaria.8,9 Hemoglobin C (HbAC) heterozygotes10,11 and α-thalassemia heterozygotes (_α/αα) and homozygotes (_α/_α) are protected against severe malaria.8 However, these well-characterized polymorphisms only partially explain the genetic variation in responses to malaria.12 It is important to identify additional human genetic variants that are associated with susceptibility or protection.
Genetic variants of human killer-cell immunoglobulin-like receptors (KIRs) and human leukocyte antigens (HLAs) are strongly associated with the risk of infectious diseases,13 autoimmune disorders,13,14,15 success in cell transplantation for the treatment of hematopoietic malignancies,16 certain cancers,17 and pregnancy outcomes.18 The KIR and HLA genes segregate independently on chromosomes 19 and 6, respectively; both gene families are highly diverse, with extensive allelic polymorphism.19 KIR and HLA genes are reported to be more polymorphic in African populations than in other populations.19 Evolutionary pressure from malaria pathogens may have partly driven the high KIR and HLA genetic diversity in Africa.20,21 The data regarding associations between KIR and HLA variants and malaria risk have been inconsistent, but since interactions between the genetically diverse KIR and HLA molecules modulate the functionality of the natural killer (NK) cell response to malaria infections, these genes remain good candidates for elucidating the role of immune cells in malaria.
Despite recent reports indicating improvement in the control of malaria in some populations and the potential for the elimination of malaria from many regions of the world, P. falciparum malaria still causes extensive morbidity and mortality, particularly in sub-Saharan Africa.22 In response to the persistent malaria burden, there have been increased efforts exerted in vector control using insecticides and malaria treatment and chemoprevention using antimalarial drugs.23 However, these approaches have faced challenges arising from both insecticide and drug resistance.24 Antimalarial drug discovery is challenging and costly,24 and parasite resistance develops easily.25 Given the limitations of insecticides and antimalarial drugs, a highly effective malaria vaccine would significantly contribute to malaria control.26 The major challenges to the development of vaccines against malaria include a failure to induce strong innate immune responses and a lack of potentiation and maintenance of adaptive immune responses.27
There have been efforts to develop malaria vaccines since the 1940s.28 Despite several promising candidates, an effective vaccine that provides long-lived protection against malaria has not been developed.29 One vaccine candidate, RTS,S/AS01, has recently been approved for pilot implementation trials in sub-Saharan Africa.30 However, RTS,S/AS01 offers only modest short-term protection,31,32 and the efficacy of this vaccine varies with the malaria transmission intensity.27 Other approaches are under study, but none have yet yielded a highly efficacious vaccine.32 A better understanding of the role of human genetic variation in heterogeneous immune responses to malaria infection may facilitate vaccine development. In this review, we provide a concise overview of the evidence for associations between KIR and HLA genetic variants and susceptibility to or protection against malaria.
Killer-cell immunoglobulin-like receptors
KIRs are a family of highly polymorphic type 1 transmembrane glycoproteins expressed on the surface of NK cells and some T cells33 that bind HLA class I molecules34 and regulate NK cell functions.35 KIRs are encoded by a set of highly polymorphic genes located within the leukocyte receptor complex on human chromosome 19q13.4.36 The KIRs are the second most genetically diverse family in the mammalian genome after HLA genes, and they differ between individuals at three main levels: copy number variation, allelic diversity and variation in the binding specificity of individual KIRs to HLA class I ligands.37
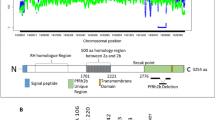
Sixteen KIR genes have been described to date, including genes that encode both inhibitory (KIR3DL1-3, KIR2DL1-3, and KIR2DL5) and activating (KIR3DS1 and KIR2DS1-5) receptors.38 KIR2DL4 is unique because it can trigger both activation and inhibition.39 KIR2DP1 and KIR3DP1 are pseudogenes that do not encode cell surface receptors.40 The nomenclature of KIR genes is based on structural and functional characteristics.41 Depending on whether KIRs have two or three extracellular immunoglobulin domains (D), they are designated as KIR2D or KIR3D.42 Functionally, KIRs with short (S) intracytoplasmic tails activate NK cells by pairing with the immunoreceptor tyrosine-based activation motif-containing adapter protein DAP12, while those with long (L) intracytoplasmic tails inhibit NK cell functions because they contain one or two immunoreceptor tyrosine-based inhibitory motifs that recruit the phosphatase SHP-1.43 Inhibitory KIRs, however, can also prime NK cells for functional competence if they bind to self HLA class I molecules, a process known as NK cell education.44 KIR genes with two or three extracellular immunoglobulin domains and short intracytoplasmic tails are designated as KIR2DS or KIR3DS, and specific genes are identified by a suffix (e.g., KIR2DS2, KIR2SD4, or KIR3DS1).45 KIR genes with two or three extracellular immunoglobulin domains and long intracytoplasmic tails are designated as KIR2DL or KIR3DL, and specific genes are identified by a suffix (e.g., KIR2DL1, KIR2DL2, KIR3DL1, or KIR3DL2).45 The human KIR genes are grouped into KIR A and KIR B haplotypes (Fig. 1). Haplotype A comprises a fixed number of 7 KIR genes, including 3 ‘framework’ genes present in all haplotypes (KIR3DL3, KIR2DL4, and KIR3DL2) and KIR2DL1, KIR2DL3, KIR3DL1, and KIR2DS4. KIR2DS4 is the only activating KIR in this haplotype; because it often carries a 22 bp deletion, haplotype A is thought to be mostly inhibitory. Approximately half of the individuals in any population studied to date exhibit haplotype A.46 Most diversity in haplotype A is conferred by allelic polymorphism.46 By definition, all other combinations of 4-16 KIR genes are classified as haplotype B, including many activating KIRs, such as KIR2DS1, KIR2DS2, KIR2DS5, and KIR3DS1, in addition to KIR2DL5.46 The diversity of haplotype B is conferred by gene content.46 However, within the centromeric (Cen) and telomeric (Tel) ends of the KIR locus, there are recombination hot spots, allowing crossover to occur and generate hybrid haplotypes such as CenA-TelB or CenB-TelA, which by definition belong to haplotype B.47 KIR genes exhibit strong linkage disequilibrium (LD) and are inherited together.36 Because of the tight LD, many disease association studies with KIR genes have actually analyzed the association with haplotypes because it is difficult to isolate the role of single KIR genes.48
KIR haplotypes. KIR haplotypes A and B are present in all populations worldwide. A recombination hotspot between KIR3DP1 and KIR2DL4 separates the centromeric region from the telomeric end of both types of haplotypes. The KIR A haplotype is mainly composed of inhibitory KIRs, except for KIR2DS4. Allelic polymorphism is very high in the KIR A haplotype (KIR3DL1, 3DL2, and 3DL3 exhibit >100 alleles, and 2DL1 and 2DL3 exhibit ~50 alleles). Haplotype B has several activating receptors, with variable numbers of genes (from 4 to 20 genes) and fewer allelic polymorphisms. Some KIR B haplotypes are composed of combinations of haplotypes A and B (CenA-TelB, CenB-TelA). The HLA epitopes bound by some KIRs are known and are indicated as C1, C2, or Bw4
A well-supported hypothesis is that KIR A specializes in fighting infectious pathogens,49 while KIR B is important for successful reproduction.50 How inhibitory receptors in haplotype A may protect against infectious diseases is unclear; however, there are at least two possibilities. The first is related to the degree of inhibition, in that weaker inhibition may be beneficial for successful immune responses.51 The second relates to NK cell education, a process that primes NK cell function through the binding of inhibitory receptors to self MHC molecules.52 The strength of KIR and HLA binding varies depending on the specific receptor-ligand pair as well as the affinity of the pairs53 and in turn impacts the regulation of NK cell activity.54 Heterogeneity in KIR gene content combined with allelic polymorphisms may lead to extensive haplotypic diversity and highly diverse NK cell populations within an individual.55 KIR diversity may in turn contribute to heterogeneous NK cell responses to P. falciparum malaria infection.36
Human leukocyte antigens
The HLA complex encodes the most polymorphic human genes, and it is usually thought that their diversity is driven by resistance to pathogens.56 The HLA complex is composed of genes on chromosome 6 that encode molecules that mediate antigen recognition and presentation as well as immunity against infectious pathogens including P. falciparum.57 There are two classes of polymorphic HLA molecules based on their structure and function, HLA class I (HLA-A, -B, and -C) and HLA class II (HLA-DR, -DQ, and -DP). HLA class I molecules are expressed on the surface of most nucleated cells. HLA class I and II molecules generally present antigenic peptides from infectious pathogens to CD8+ and CD4+ T cells, respectively.58 However, HLA class I molecules are also major ligands for KIRs,59 and as such, they play a role in the regulation of NK cell activity.59 KIR binding to HLA class I molecules involves four epitopes, A3/A11, Bw4, C1, and C2, found in some HLA-A allotypes, some HLA-B allotypes and all HLA-C allotypes (Fig. 1). The nonclassical HLA class I molecule HLA-G60 is expressed only on placental cells; HLA-G may play roles in reproduction61 and binds to KIR2DL4 and leukocyte Ig-like receptor B. The other nonclassical HLA class I molecule HLA-E60 presents peptides derived from other HLA class I molecules to the inhibitory CD94/NKG2A receptor on NK cells62 and some T cells. HLA class II molecules are confined to immunocompetent cells such as B lymphocytes, dendritic cells, macrophages, endothelial cells, and activated T-lymphocytes, but their expression may be induced on other cell types.63
Immunity to malaria
Acquired immunity to P. falciparum malaria infection is complex, requiring a balance of the control of parasite growth and inflammation, as the overproduction of inflammatory cytokines results in adverse pathological consequences.64 Innate and adaptive immunity are both essential for limiting P. falciparum parasite growth and the severity of malaria infection.3,65 Repeated exposure to P. falciparum results in the attenuation of inflammation66 and the generation of antimalarial antibodies that play an essential role in the control of blood-stage malaria infection.67,68 Antimalarial antibodies play an essential role in immunity to malaria in areas with a high P. falciparum malaria transmission intensity.69 Individuals living in malaria-endemic regions acquire immunity against severe malaria quickly, but immunity against symptomatic malaria and the control of parasitemia require years of repeated exposure to malaria parasites and remain incomplete.70,71,72
It has been observed that humans are protected against malaria by antibodies through two mechanisms. First, antibodies bind to circulating free malaria parasites and prevent them from invading red blood cells,73,74 and second, antibodies from malaria-immune individuals bind to P. falciparum proteins on the surface of infected red blood cells (iRBCs). This induces antibody-dependent cell-mediated cytotoxicity (ADCC) exerted by NK cells,73 phagocytosis by monocytes, and the engagement of effector cells via Fc-mediated interactions to mediate the clearance of opsonized red blood cells73
NK cells and innate immunity to P. falciparum malaria
In humans, NK cells are usually defined as CD3–CD56+ cells75 and can be further subdivided based on CD56 expression.76 Typically, CD56dim NK cells constitute the majority (about 90%) of peripheral blood NK cells,77 whereas CD56bright NK cells are more abundant in secondary lymphoid tissues.78 CD56dim NK cells express high levels of the low-affinity Fc receptor CD16, display heterogeneous expression of inhibitory KIRs for HLA class I, and express high levels of perforin.79 In contrast, CD56bright NK cells express no or low levels of CD16, exclusively express the inhibitory receptor CD94/NKG2A and show no KIR expression and present tenfold lower perforin expression than CD56dim NK cells.80 Therefore, the CD56dim and CD56bright NK cell subsets are considered to perform different functional roles (Fig. 2a). Another subset of NK cells referred to as adaptive or memory NK cells develops with age in people with certain infections. For example, in HCMV+ individuals, adaptive NK cells that can be found in blood are identified by the expression of the CD94/NKG2C receptor and the CD57 marker.81 Adaptive NK cells are differentiated from other types of NK cells by the absence of the transcription factor promyelocytic leukemia zinc finger (PLZF) and the signaling Fc receptor γ-chain (FcRγ), which are lost through epigenetic modifications.82
NK receptors and responses to malaria. NK cell responses to malaria may be either beneficial, if they target the parasite, or detrimental, if they contribute to immunopathology. The binding of HLA receptors (CD94/NKG2A or KIR) to self HLA molecules affects both how NK cells are primed for function in a steady state and how they function during an immune response. In the steady state, the binding of HLA molecules on hematopoietic cells by inhibitory NK cell receptors such as NKG2A (which binds HLA-E) and KIR (which binds HLA-A, -B, and -C) primes NK cells to make them functionally competent. The process of acquiring functional competence through the binding of inhibitory receptors to self HLA molecules on hematopoietic cells is referred to as NK cell education. The binding of the same inhibitory receptors to HLA molecules on potential target cells during an immune response, however, suppresses NK cell function. a For example, CD56bright NK cells that do not express KIR can be educated by NKG2A binding to HLA-E to produce IFN-γ during the immune response to malaria. On the other hand, CD56dim NK cells that express inhibitory KIRs (e.g., KIR2DL1 or KIR3DL1 in a KIR A haplotype) for self HLA (e.g., HLA-C2 or HLA-Bw4, respectively) found on hematopoietic cells are educated to recognize and kill infected red blood cells that do not express HLA molecules and therefore cannot suppress KIR2DL1+ and KIR3DL1+ CD56dim NK cells during immune responses to blood-stage malaria. CD56dim cells can also mediate ADCC because they express CD16, which binds antibodies through its Fc domain. b Liver NK cells are composed of different subsets, some of which express both KIR and NKG2A, and both of these receptors can educate liver NK cells by binding HLA-A, -B, -C, or -E on hematopoietic cells in a steady state. On the other hand, during liver-stage malaria, both receptors can suppress NK cell functions when NK cells interact with infected hepatocytes that express cognate HLA molecules
NK cells play a vital role in the innate immune response to falciparum malaria infections by the production of IFN-γ, the inhibition of parasite growth, and the cytotoxic killing of intraerythrocytic parasites.73 NK cells are the first cells in peripheral blood to produce IFN-γ in response to P. falciparum infection83 and participate in the initiation and development of adaptive immune responses.84 It has been shown that terminally differentiated CD56neg NK cells expand in children after chronic malaria exposure and in those diagnosed with endemic Burkitt lymphoma.85 NK cell activities can be triggered by three different but complementary pathways: cytokine activation,73 ADCC,73 and loss of inhibitory signaling due to the downregulation of HLA class I ligands, also known as missing self recognition.73 It was recently shown that CD56dim NK cells inhibit P. falciparum growth and kill P. falciparum within red blood cells flagged with IgG antibodies in individuals who live in malaria-endemic areas through ADCC (Fig. 2a). This is important during blood-stage P. falciparum malaria infection.73 Liver NK cells also express KIR, which may modulate the NK cell response to liver-stage malaria infection through interactions with HLA class I molecules expressed on the surface of hepatocytes (Fig. 2b).86
In a recent study in Mali, multiparameter flow cytometry showed an increase in the proportion of adaptive NK cells that were associated with lower parasitemia and protection against malaria infection.87 Indeed, these cells enhanced the ADCC response to P. falciparum-iRBCs in the presence of naturally acquired antibodies from malaria-resistant individuals.87 Malaria-susceptible individuals exhibiting a higher proportion of PLZF- and FcRγ-negative NK cells during the transmission season showed improved odds of getting protected against malaria infection during the subsequent season.87 Although the most prominent cytokines produced by NK cells are IFN-γ and tumor necrosis factor-α, they also secrete other cytokines, including IL-10, the growth factor GM-CSF, and the chemokines MIP-1α, MIP-1β, IL-8, and RANTES. It has been demonstrated that the production of the regulatory cytokine IL-10 by NK cells prevents overt pathology and death from cerebral malaria in mouse models.88
Some studies have demonstrated that individuals may exhibit a varying ability to elicit an innate immune response to malaria infection, with clear implications for disease manifestations.4 This variation may arise as a result of several factors, including the strength of the cytokine and costimulatory signals relayed to NK cells by accessory cells and differential NK cell maturation, depending on age and infection history. For example, human cytomegalovirus (HCMV) modulates NK cell activity by engaging inhibitory receptors without triggering activating surface receptors.89 HCMV also leads to the suppression of NK cell receptor recognition by the corresponding HLA class ligands.90 The heterogeneity of NK cell responses to plasmodial infection may be attributed to interactions between NK cell receptors and their HLA class I molecule ligands.83
NK cells and placental malaria
Placental malaria results from the sequestration of P. falciparum-infected erythrocytes within the intervillous spaces of the placenta and may cause serious pregnancy complications, such as abortion, stillbirth, and intrauterine growth restriction.91 The cytolysis of P. falciparum-iRBCs by peripheral NK cells may be important in defense against placental malaria.92,93 Previous studies have indeed shown that an increase in IFN-γ-expressing NK cells in the intervillous blood of the placenta at term is associated with a reduced risk of placental malaria.94
A distinct population of NK cells is the most abundant leukocyte population in the decidua, where early in pregnancy, these cells contribute to placentation and pregnancy outcomes by regulating trophoblast invasion and uterine vascular remodeling.95 Decidual NK cells may also provide first-line innate immune defense against infectious diseases, including placental malaria.96 Decidual NK cells mainly express inhibitory KIR2DL1, KIR2DL2, and KIR2DL3, activating KIR2DS1 and KIR2DL4, which may exhibit inhibitory or activating functions.96 KIR diversity may be driven by balanced selection between pregnancy success and defense against pathogens.50 Certain specific combinations of KIR and HLA genes affect the susceptibility to certain pregnancy complications.95 Other combinations may be associated with susceptibility to or protection against placental malaria. In a study of 688 placental malaria and HIV-coinfected individuals in Kenya, KIR BB homozygosity was associated with protection against placental malaria in HIV-negative pregnant women but with malaria susceptibility in HIV-positive pregnant women.96 This reverse association between KIR genes and placental malaria remained only in women with high CD4 cell counts and was not observed in those with low CD4 cell counts.
KIRs and immunity to P. falciparum malaria
The activity of NK cells in immunity against P. falciparum infection depends on a fine balance between the strength of the activating and inhibitory signals induced by KIR molecules.97 It has been clearly shown that KIR molecules regulate the NK cell-mediated production of IFN-γ in response to P. falciparum infection and consequently influence the pathogenesis and severity of malaria.98 For example, NK cells from individuals with KIR AB haplotypes produce stronger IFN-γ responses in vitro in response to P. falciparum infection compared with the responses observed in either KIR AA or KIR BB homozygous individuals.99
A study of 477 malaria cases in Thailand showed an association of the KIR2DL3 gene (which identifies haplotype A) and its ligand, HLA-C1, with cerebral malaria. Moreover, this receptor-ligand pair was found at a lower frequency in highly malaria-endemic populations, suggesting the existence of selection due to unhelpful NK cell responses, which may increase susceptibility to cerebral malaria.100 However, a comparison of the KIR genotypes of 321 children with severe and uncomplicated malaria with those of 314 control children showed that the Cen-AB haplotype was more frequent in malaria cases in the Gambia, with more activating KIR genes (KIR2DS2 and KIR2DS5) being found in very ill children, suggesting that exaggerated NK cell activation may contribute to the pathogenesis of severe malaria.97 Furthermore, parasitemia was higher in KIR AA homozygous individuals and more frequent in controls, suggesting that a more inhibitory A haplotype may not favor parasite clearance but may protect against severe malaria.97
In Southwest Nigeria, the KIR2DL5, KIR2DS3, and KIR2DS5 genes were observed to be overrepresented in individuals with asymptomatic P. falciparum parasitemia compared with those with either uncomplicated or severe malaria.101 Furthermore, the frequency of KIR2DS3 and KIR2DS5 was higher in individuals with uncomplicated malaria than in those with severe malaria.101 These results suggest a protective role of inhibitory KIR2DL5 and activating KIR2DS3 and KIR2DS5 genes against severe malaria. Heterozygosity at the KIR Cen region, Cen-AB2, has also been shown to be more frequent in asymptomatic controls compared with individuals with severe malaria.101 This implies that KIR heterozygous AB haplotypes may be associated with protection against severe malaria, and it has been suggested that this may be due to the greater proportion of educated NK cells in KIR AB heterozygous individuals. In Africans, the Cen region of the KIR locus, which encodes HLA-C receptors, is highly diverse, whereas the Tel region encoding Bw4-specific receptors (KIR3DL1) lacks diversity.102 These characteristics of KIR are consistent with the ongoing selection imposed by malaria pathogens in Africa.102
In northern India, individuals possessing the KIR2DS2, KIR2DL1, and KIR2DL3 genes were shown to be susceptible to severe malaria.103 In the same study, a combination of KIR2DS2-HLAC1, KIR2DL1-HLAC2, and KIR2DL3-HLAC1 was found to be associated with an increased risk of severe malaria.103 In a study of 77 Melanesian individuals with a history or symptoms of malaria in the highly malaria-endemic Solomon Islands, a higher frequency of KIR AB heterozygosity was found among 37 Plasmodium-positive individuals than among 40 negative individuals.104 The nondeleted allele of KIR2DS4 (*001) was also more frequent in malaria-positive individuals.104 Other studies have focused on the functional responses of lymphocytes from healthy donors to P. falciparum-iRBCs and found heterogeneity in donor NK cell IFN-γ production that was significantly associated with KIR genotypes,105 showing that KIR AB heterozygosity was associated with strong responses of CD56dim cells.106 It is difficult to draw a conclusive picture from the available evidence. While KIR AB heterozygosity may favor a larger number of educated CD56dim NK cells, which can respond more effectively to the infection, individuals with numerous activating KIRs may mount responses that are too vigorous and may be deleterious in infected individuals. Individual KIR genes as well as the KIRs and the corresponding HLA class I ligands that have been shown to be associated with susceptibility to or protection against severe malaria are summarized in Table 1.
HLA genetic variants and immunity to P. falciparum malaria
The unique variability of the genes that encode HLA molecules and their haplotypic composition, especially in native Africans, is proposed to have resulted from the need to fight multiple, frequent deadly infectious pathogens.107 HLA class I molecules are major ligands for KIRs, and as such, they play a role in the regulation of NK cell activity during malaria infection. HLA class I molecules also present malaria antigens to T cells and are therefore important in adaptive immunity to malaria, which is critical during liver-stage P. falciparum malaria infection. HLA class II molecules mediate the clearance of red blood cells infected with P. falciparum parasites through the stimulation of T helper cells.108 Some studies have clearly shown that HLA molecules influence antibody titers against malaria antigens, including glutamate-rich protein and merozoite surface antigens.109
Several studies have shown some HLA genetic variants to be associated with susceptibility to or protection against severe malaria. However, the results have been inconsistent (Table 2). The first study on the role of HLA in malaria described protection against cerebral malaria and severe malarial anemia conferred by the HLA class I allele HLA-Bw53 and the HLA class II haplotype HLA-DRBl*1302-DQAl*0102-DQBl*0501. Molecular analysis revealed that liver-stage antigen-I (LSA-1), a liver-stage-specific antigen of P. falciparum, was recognized by HLA-Bw53-positive individuals. However, these findings have not been confirmed in other malaria-endemic populations, and no significant associations of malaria antigens other than LSA-1 with HLA molecules have been discovered.
In studies of interactions between variable HLA molecules and polymorphic parasite factors, it has been demonstrated that the P. falciparum circumsporozoite protein binds to HLA-DR and HLA-DQ molecules in vitro as well as in animal models. Two HLA class II alleles, DRB1*04 and DPB1*1701, have been observed to be more frequent in severe malaria than in uncomplicated malaria. HLA-B49, -A1, -B27, and HLA-DRB1*0809 were shown to be strongly associated with severe malaria in a study conducted in Mumbai in India.110 HLA-A19 and HLA-DQB1*0203 were associated with protection against severe malaria in the same population.110 In a study conducted in Thailand in patients with severe cerebral malaria, HLA-B46 was significantly associated with the risk of cerebral malaria, while HLA-B56 and HLADR1*1001 were associated with protection against cerebral malaria.111 In Senegal, HLA-DR3 and HLA-DR10 were strongly associated with cerebral malaria.112 In the Gambia, HLA-B53 and HLA-DRB1*1302 alleles were found to be strongly associated with protection against severe malaria.113
Concluding remarks
It is clear that interactions between KIR and HLA molecules modulate the activity of NK cells. The role of NK cells in responses to malaria has been established, and it is therefore important to decipher how KIR and HLA immunogenetics regulate susceptibility to or protection against malaria to understand the heterogeneous responses of populations to malaria infection. This may in turn help understand malaria transmission and severity in specific populations and inform specific malaria control interventions.
Several studies have suggested that certain KIR and HLA variants are associated with malaria, but there are challenges at different levels to obtaining a comprehensive picture of how KIR and HLA variants contribute to susceptibility to or protection against malaria. Some challenges are related to population genetics, some to parasite genetics and others to the biology of KIR and HLA immunogenetics. First, some of the relevant studies were conducted with small sample sizes, highlighting the need to study larger populations in Africa and other malaria-endemic populations, particularly to accommodate allelic diversity. Different KIR and HLA variants may have been selected in different populations. Other infectious pathogens that are prevalent in certain malaria-endemic populations may also exert selective pressure on immune responses, thus shaping the diversity of KIR and HLA in those populations. Therefore, the role of other coinfections should be considered in studies involving KIR and malaria, especially in populations affected by many infectious pathogens. Different species of parasites may infect different populations, and unique antigenic drift may occur in parasites in different populations. How the binding between certain KIR and HLA molecules regulates NK cell function in the context of any given disease is still very challenging to determine. In the case of malaria, knowledge of the rich KIR allelic diversity in African populations and the binding affinities of these receptors to equally diverse HLA ligands are still being sought. KIRs can either inhibit or enhance NK cell effector functions. Combinations of KIR and HLA genes determine both NK cell education (the priming of effector function) and effector function itself. However, because inhibitory KIRs and other inhibitory receptors (e.g., CD94/NKG2A) contribute to education and effector function in opposite ways (i.e., they promote education but suppress function), it may be difficult to reconstruct how KIR and HLA immunogenetics determine the biology of NK cells during malaria infection. For example, inhibitory interactions may be necessary to educate NK cells to mediate ADCC required to eliminate iRBCs but may not be required or could even be counterproductive for the activation of IFN-γ production by NK cells upon interaction with HLA class I-expressing infected cells.
Finally, evolutionary pressure from P. falciparum malaria pathogens might have selected for certain KIR and HLA genetic variants that protect against severe malaria but increase the risk of other diseases, as has been well documented for sickle cell disease and other hemoglobinopathies. For example, the high frequency of certain KIR and HLA variants associated with pregnancy complications in sub-Saharan Africa114 may be the result of a trade-off for the selection of genetic variants that protect against malaria.
References
Ashley, E. A., Pyae Phyo, A. & Woodrow, C. J. Malaria. Lancet 391, 1608–1621 (2018).
Gomes, P. S., Bhardwaj, J., Rivera-Correa, J., Freire-De-Lima, C. G. & Morrot, A. Immune escape strategies of malaria parasites. Front. Microbiol. 7, 1617 (2016).
Gowda, D. C. & Wu, X. Parasite recognition and signaling mechanisms in innate immune responses to malaria. Front. Immunol. 9, 3006 (2018).
Doolan, D. L., Dobaño, C. & Baird, J. K. Acquired immunity to malaria. Clin. Microbiol. Rev. 22, 13 (2009).
Nhabomba, A. J. et al. Impact of age of first exposure to Plasmodium falciparum on antibody responses to malaria in children: a randomized, controlled trial in Mozambique. Malar. J. 13, 121 (2014).
Varo, R. et al. Adjunctive therapy for severe malaria: a review and critical appraisal. Malar. J. 17, 47 (2018).
Maitland, K. Severe malaria in African children - the need for continuing investment. N. Engl. J. Med. 375, 2416–2417 (2016).
Mpimbaza, A. et al. Associations between erythrocyte polymorphisms and s uncomplicated and severe malaria in Ugandan children: a case control study. PLoS ONE 13, e0203229–e0203229 (2018).
Goheen, M. M., Campino, S. & Cerami, C. The role of the red blood cell in host defence against falciparum malaria: an expanding repertoire of evolutionary alterations. Br. J. Haematol. 179, 543–56. (2017).
Agarwal, A. et al. Hemoglobin C associated with from severe malaria in the Dogon of Mali, a West African population with a low prevalence of hemoglobin S. Blood 96, 2358–2363 (2000).
Mockenhaupt, F. P. et al. Hemoglobin C and resistance to severe malaria in Ghanaian children. J. Infect. Dis. 190, 1006–1009 (2004).
Kwiatkowski, D. P. How malaria has affected the human genome and what human genetics can teach us about malaria. Am. J. Hum. Genet. 77, 171–192 (2005).
Norman, P. J. et al. Defining KIR and HLA class I genotypes at highest resolution via high-throughput sequencing. Am. J. Hum. Genet. 99, 375–391 (2016).
Matzaraki, V., Kumar, V., Wijmenga, C. & Zhernakova, A. The MHC locus and genetic susceptibility to autoimmune and infectious diseases. Genome Biol. 18, 76 (2017).
Bodis, G., Toth, V. & Schwarting, A. Role of human leukocyte antigens (HLA) in autoimmune diseases. Rheumatol. Ther. 5, 5–20 (2018).
Cooley, S., Parham, P. & Miller, J. S. Strategies to activate NK cells to prevent relapse and induce remission following hematopoietic stem cell transplantation. Blood 131, 1053 (2018).
Torres, A. R., Sweeten, T. L., Johnson, R. C., Odell, D., Westover, J. B., Bray-Ward, P. et al. Common genetic variants found in HLA and KIR immune genes in autism spectrum disorder. Front. Neurosci. 10, 463 (2016).
Colucci, F. The role of KIR and HLA interactions in pregnancy complications. Immunogenetics 69, 557–65. (2017).
Jiang, W. et al. qKAT: a high-throughput qPCR method for KIR gene copy number and haplotype determination. Genome Med. 8, 99 (2016).
Meyer, D. C., Aguiar, V. R., Bitarello, B. D. C., Brandt, D. Y. & Nunes, K. A genomic perspective on HLA evolution. Immunogenetics 70, 5–27 (2018).
Quintana-Murci, L. Understanding rare and common diseases in the context of human evolution. Genome Biol. 17, 225 (2016).
Dhiman, S. Are malaria elimination efforts on right track? An analysis of gains achieved and challenges ahead. Infect. Dis. Poverty 8, 14 (2019).
Burrows, J. N. et al. New developments in anti-malarial target candidate and product profiles. Malar. J. 16, 26 (2017).
Ashley, E. A. & Phyo, A. P. Drugs in development for malaria. Drugs 78, 861–79. (2018).
Sinha, S., Medhi, B. & Sehgal, R. Challenges of drug-resistant malaria. Parasite 21, 61 (2014).
Birkholtz, L.-M., Bornman, R., Focke, W., Mutero, C. & de Jager, C. Sustainable malaria control: transdisciplinary approaches for translational applications. Malar. J. 11, 431 (2012).
Ssemaganda, A. et al. Induction of Plasmodium-specific immune responses using liposome-based vaccines. Front. Immunol. 10, 135 (2019). -.
Coelho, C. H., Doritchamou, J. Y. A., Zaidi, I. & Duffy, P. E. Advances in malaria vaccine development: report from the 2017 malaria vaccine symposium. npj Vaccines. 2, 34 (2017).
Ouattara, A. & Laurens, M. B. Vaccines against malaria. Clin. Infect. Dis. 60, 930–936 (2014).
van den Berg, M., Ogutu, B., Sewankambo, N. K., Biller-Andorno, N. & Tanner, M. RTS,S malaria vaccine pilot studies: addressing the human realities in large-scale clinical trials. Trials 20, 316 (2019).
Gosling, R. & von Seidlein, L. The future of the RTS,S/AS01 malaria vaccine: an alternative development plan. PLoS Med. 13, e1001994 (2016).
Beeson, J. G. et al. Challenges and strategies for developing efficacious and long-lasting malaria vaccines. Sci. Transl. Med. 11, eaau1458 (2019).
Pallmer, K. & Oxenius, A. Recognition and regulation of T cells by NK cells. Front. Immunol. 7, 251 (2016).
Wroblewski, E. E., Parham, P. & Guethlein, L. A. Two to tango: co-evolution of hominid natural killer cell receptors and MHC. Front. Immunol. 10, 177 (2019).
Pende, D. et al. Killer Ig-like receptors (KIRs): their role in NK cell modulation and developments leading to their clinical exploitation. Front. Immunol. 10, 1179 (2019).
Middleton, D. & Gonzelez, F. The extensive polymorphism of KIR genes. Immunology 129, 8–19 (2010).
Parham, P. The genetic and evolutionary balances in human NK cell receptor diversity. Semin. Immunol. 20, 311–316 (2008).
Béziat, V., Hilton, H. G., Norman, P. J. & Traherne, J. A. Deciphering the killer-cell immunoglobulin-like receptor system at super-resolution for natural killer and T-cell biology. Immunology. 150, 248–264 (2017).
Faure, M. & Long, E. O. KIR2DL4 (CD158d), an NK cell-activating receptor with inhibitory potential. J. Immunol. 168, 6208 (2002).
Moffett, A., Chazara, O., Colucci, F. & Johnson, M. H. Variation of maternal KIR and fetal HLA-C genes in reproductive failure: too early for clinical intervention. Reprod. BioMedicine Online 33, 763–769 (2016).
Campbell, K. S. & Purdy, A. K. Structure/function of human killer cell immunoglobulin-like receptors: lessons from polymorphisms, evolution, crystal structures and mutations. Immunology 132, 315–325 (2011).
Moesta, A. K. & Parham, P. Diverse functionality among human NK cell receptors for the C1 epitope of HLA-C: KIR2DS2, KIR2DL2, and KIR2DL3. Front. Immunol. 3, 336 (2012). -.
Cheent, K. & Khakoo, S. I. Natural killer cells: integrating diversity with function. Immunology 126, 449–457 (2009).
Orr, M. T. & Lanier, L. L. Natural killer cell education and tolerance. Cell 142, 847–856 (2010).
Moesta, A. & Parham, P. Diverse functionality among human NK cell receptors for the C1 epitope of HLA-C: KIR2DS2, KIR2DL2, and KIR2DL3. Front Immunol. 3, 336 (2012).
Colucci, F. & Traherne, J. Killer-cell immunoglobulin-like receptors on the cusp of modern immunogenetics. Immunology 152, 556–561 (2017).
Jiang, W. et al. Copy number variation leads to considerable diversity for B but not A haplotypes of the human KIR genes encoding NK cell receptors. Genome Res. 22, 1845–1854 (2012).
Single, R. M., Martin, M. P., Meyer, D., Gao, X. & Carrington, M. Methods for assessing gene content diversity of KIR with examples from a global set of populations. Immunogenetics 60, 711–725 (2008).
Penman, B. S., Moffett, A., Chazara, O., Gupta, S. & Parham, P. Reproduction, infection and killer-cell immunoglobulin-like receptor haplotype evolution. Immunogenetics 68, 755–64. (2016).
Parham, P. & Moffett, A. Variable NK cell receptors and their MHC class I ligands in immunity, reproduction and human evolution. Nat. Rev. Immunol. 13, 133–144 (2013).
Khakoo, S. I. et al. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 305, 872–874 (2004).
Martin, M. P. et al. Innate partnership of HLA-B and KIR3DL1 subtypes against HIV-1. Nat. Genet. 39, 733–740 (2007).
Guethlein, L. A., Norman, P. J., Hilton, H. G. & Parham, P. Co-evolution of MHC class I and variable NK cell receptors in placental mammals. Immunological Rev. 267, 259–282 (2015).
Boudreau, J. E. & Hsu, K. C. Natural killer cell education and the response to infection and cancer therapy: stay tuned. Trends Immunol. 39, 222–239 (2018).
Korbel, D. S., Newman, K. C., Almeida, C. R., Davis, D. M. & Riley, E. M. Heterogeneous human NK cell responses to Plasmodium falciparum-infected erythrocytes. J. Immunol. 175, 7466 (2005).
Choo, S. Y. The HLA system: genetics, immunology, clinical testing, and clinical implications. Yonsei Med. J. 48, 11–23 (2007).
Mosaad, Y. M. Clinical role of human leukocyte antigen in health and disease. Scand. J. Immunol. 82, 283–306 (2015).
Rock, K. L., Reits, E. & Neefjes, J. Present yourself! by MHC class I and MHC class II molecules. Trends Immunol. 37, 724–737 (2016).
Hollenbach, J. A., Pando, M. J., Caillier, S. J., Gourraud, P. A. & Oksenberg, J. R. The killer immunoglobulin-like receptor KIR3DL1 in combination with HLA-Bw4 is protective against multiple sclerosis in African Americans. Genes Immun. 17, 199–202 (2016).
Carlini, F. et al. Association of HLA-A and non-classical HLA class I alleles. PLoS ONE 11, e0163570 (2016).
Durmanova, V., Homolova, M., Drobny, J., Shawkatová, I. & Buc, M. Role of HLA-G and other immune mechanisms in pregnancy. Central Eur. J. Biol. 8 (2013).
Rölle, A., Jäger, D. & Momburg, F. HLA-E peptide repertoire and dimorphism—centerpieces in the adaptive NK cell puzzle? Front. Immunol. 9, 2410 (2018).
Greenbaum, J. et al. Functional classification of class II human leukocyte antigen (HLA) molecules reveal seven different supertypes and a surprising degree of repertoire sharing across supertypes. Immunogenetics 63, 325–335 (2011).
Kumar, R., Ng, S. & Engwerda, C. The role of IL-10 in malaria: a double edged sword. Front. Immunol. 10, 229 (2019).
Liehl, P. et al. Innate immunity induced by Plasmodium liver infection inhibits malaria reinfections. Infect. Immun. 83, 1172 (2015).
Portugal, S. et al. Exposure-dependent control of malaria-induced inflammation in children. PLoS Pathog. 10, e1004079–e1004079 (2014).
Stanisic, D. I. et al. Acquisition of antibodies against Plasmodium falciparum merozoites and malaria immunity in young children and the influence of age, force of infection, and magnitude of response. Infect. Immun. 83, 646 (2015).
Cockburn, I. A. & Seder, R. A. Malaria prevention: from immunological concepts to effective vaccines and protective antibodies. Nat. Immunol. 19, 1199–1211 (2018).
Kusi, K. A. et al. Variations in the quality of malaria-specific antibodies with transmission intensity in a seasonal malaria transmission area of Northern Ghana. PLoS ONE 12, e0185303 (2017).
Ly, A. & Hansen, D. S. Development of B cell memory in malaria. Front. Immunol. 10, 559 (2019).
Farrington, L. et al. Both inflammatory and regulatory cytokine responses to malaria are blunted with increasing age in highly exposed children. Malar. J. 16, 499 (2017).
Valletta, J. J. & Recker, M. Identification of immune signatures predictive of clinical from malaria. PLoS computational Biol. 13, e1005812 (2017).
Arora, G. et al. NK cells inhibit Plasmodium falciparum growth in red blood cells via antibody-dependent cellular cytotoxicity. eLife 7, e36806 (2018).
Paul, A. S., Egan, E. S. & Duraisingh, M. T. Host-parasite interactions that guide red blood cell invasion by malaria parasites. Curr. Opin. Hematol. 22, 220–226 (2015).
Gharehbaghian, A., Donaldson, C., Newman, J., Bannister, G. & Bradley, B. A. The correlation between the percent of CD3- CD56+ cells and NK precursor function. Iran. J. Allergy Asthma Immunol. 5, 167–175 (2006).
Rebuli, M. E., Pawlak, E. A., Walsh, D., Martin, E. M. & Jaspers, I. Distinguishing human peripheral blood NK cells from CD56dimCD16dimCD69+CD103+ resident nasal mucosal lavage fluid cells. Sci. Rep. 8, 3394 (2018).
Amand, M. et al. Human CD56(dim)CD16(dim) cells as an individualized natural killer cell subset. Front. Immunol. 8, 699 (2017).
Michel, T. et al. Human CD56 NK cells: an update. J. Immunol. 196, 2923 (2016).
Fauriat, C., Long, E. O., Ljunggren, H.-G. & Bryceson, Y. T. Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood 115, 2167–2176 (2010).
Jacobs, R. et al. CD56bright cells differ in their KIR repertoire and cytotoxic features from CD56dim NK cells. Eur. J. Immunol. 31, 3121–3126 (2001).
Hammer, Q., Rückert, T., Dunst, J. & Romagnani, C. Adaptive natural killer cells integrate interleukin-18 during target-cell encounter. Front. Immunol. 8, 1976 (2018).
Schlums, H. et al. Cytomegalovirus infection drives adaptive epigenetic diversification of NK cells with altered signaling and effector function. Immunity 42, 443–456. (2015).
Wolf, A.-S., Sherratt, S. & Riley, E. M. NK cells: uncertain allies against malaria. Front. Immunol. 8, 212 (2017).
Obaid, A. et al. Model of the adaptive immune response system against HCV infection reveals potential immunomodulatory agents for combination therapy. Sci. Rep. 8, 8874 (2018).
Forconi, C. S. et al. Poorly cytotoxic terminally differentiated CD56negCD16pos NK cells accumulate in Kenyan children with Burkitt lymphomas. Blood Adv. 2, 1101–1114 (2018).
Liu, P., Chen, L. & Zhang, H. Natural killer cells in liver disease and hepatocellular carcinoma and the NK cell-based immunotherapy. J. Immunol. Res. 2018, 1206737 (2018).
Hart, G. T. et al. Adaptive NK cells in people exposed to Plasmodium falciparum correlate with from malaria. J. Exp. Med. 216, 1280–1290 (2019).
Burrack, K. S., Hart, G. T. & Hamilton, S. E. Contributions of natural killer cells to the immune response against Plasmodium. Malar. J. 18, 321 (2019).
Goodier, M. R., Jonjić, S., Riley, E. M. & Juranić Lisnić, V. CMV and natural killer cells: shaping the response to vaccination. Eur. J. Immunol. 48, 50–65 (2018).
Wilkinson, G. W. G. et al. Modulation of natural killer cells by human cytomegalovirus. J. Clin. Virol. 41, 206–212 (2008).
Sharma, L. & Shukla, G. Placental malaria: a new insight into the pathophysiology. Front. Med. 4, 117 (2017).
Oth, T. et al. Pathogen recognition by NK cells amplifies the pro-inflammatory cytokine production of monocyte-derived DC via IFN-γ. BMC Immunol. 19, 8 (2018).
Clark, S. E. et al. Bacterial manipulation of NK cell regulatory activity increases susceptibility to listeria monocytogenes Infection. PLoS Pathog. 12, e1005708 (2016).
Othoro, C. et al. Elevated gamma interferon-producing NK cells, CD45RO memory-like T cells, and CD4 T cells are associated with protection against malaria infection in pregnancy. Infect. Immun. 76, 1678–1685 (2008).
Moffett, A. & Colucci, F. Uterine NK cells: active regulators at the maternal-fetal interface. The. J. Clin. Investig. 124, 1872–1879 (2014).
Omosun, Y. O. et al. Differential association of gene content polymorphisms of killer cell immunoglobulin-like receptors with placental malaria in HIV- and HIV+ mothers. PLoS ONE 7, e38617–e38617 (2012).
Yindom, L. M. et al. Killer-cell immunoglobulin-like receptors and malaria caused by Plasmodium falciparum in The Gambia. Tissue Antigens 79, 104–113 (2012).
Korbel, D. S., Newman, K. C., Almeida, C. R., Davis, D. M. & Riley, E. M. Heterogeneous human NK cell responses to Plasmodium falciparum-infected erythrocytes. J. Immunol. 175, 7466–7473 (2005).
Wolf, A.-S., Sherratt, S. & Riley, E. M. NK cells: uncertain allies against malaria. Front Immunol. 8, 212 (2017).
Hirayasu, K. et al. Significant association of KIR2DL3-HLA-C1 combination with cerebral malaria and implications for co-evolution of KIR and HLA. PLoS Pathog. 8, e1002565–e1002565 (2012).
Olaniyan, S. A. et al. Killer-cell immunoglobulin-like receptors and falciparum malaria in southwest Nigeria. Hum. Immunol. 75, 816–821 (2014).
Norman, P. J. et al. Co-evolution of human leukocyte antigen (HLA) class I ligands with killer-cell immunoglobulin-like receptors (KIR) in a genetically diverse population of sub-Saharan Africans. PLoS Genet. 9, e1003938–e1003938 (2013).
Prakash, S., Ranjan, P., Ghoshal, U. & Agrawal, S. KIR-like activating natural killer cell receptors and their association with complicated malaria in north India. Acta Trop. 178, 55–60 (2018).
Taniguchi, M. & Kawabata, M. KIR3DL1/S1 genotypes and KIR2DS4 allelic variants in the AB KIR genotypes are associated with Plasmodium-positive individuals in malaria infection. Immunogenetics 61, 717–730 (2009).
Artavanis-Tsakonas, K. et al. Activation of a subset of human NK cells upon contact with Plasmodium falciparum-infected erythrocytes. J. Immunol. 171, 5396 (2003).
Korbel, D. S. et al. Killer Ig-like receptor (KIR) genotype predicts the capacity of human KIR-positive CD56dim NK cells to respond to pathogen-associated signals. J. Immunol. 182, 6426–6434 (2009).
Blackwell, J. M., Jamieson, S. E. & Burgner, D. HLA and infectious diseases. Clin. Microbiol. Rev. 22, 370–385 (2009).
Belachew, E. B. Immune response and evasion mechanisms of Plasmodium falciparum parasites. J. Immunol. Res 2018, 6529681 (2018).
Ghosh, K. Evolution and selection of human leukocyte antigen alleles by Plasmodium falciparum infection. Hum. Immunol. 69, 856–860 (2008).
Shankarkumar, U. et al. HLA associations in P. falciparum malaria patients from Mumbai, western India. Indian J. Malariol. 39, 76–82 (2002).
Hananantachai, H. et al. Polymorphisms of the HLA-B and HLA-DRB1 genes in Thai malaria patients. Jpn J. Infect. Dis. 58, 25–28 (2005).
Ndiaye, M. et al. Susceptibility to neuro-malaria and HLA-DR alleles in Senegal. Dakar Med. 43, 25–28 (1998).
Adrian, V. S. H. et al. Human leukocyte antigens and natural selection by malaria. Philos. Trans. 346, 379–385 (1994).
Nakimuli, A. et al. A KIR B centromeric region present in Africans but not Europeans protects pregnant women from pre-eclampsia. Proc. Natl Acad. Sci. USA 112, 845–850 (2015).
Hill, A. V. et al. Molecular analysis of the association of HLA-B53 and resistance to severe malaria. Nature 360, 434–439 (1992).
Hirayama, K. et al. Molecular analysis of HLA-B in the Malaysian aborigines. Tissue Antigens 48, 692–697 (1996).
Contu, L. et al. HLA-B35 frequency variations correlate with malaria infection in Sardinia. Tissue Antigens 52, 452–461 (1998).
Mehra, N. K., Jaini, R., Rajalingam, R., Balamurugan, A. & Kaur, G. Molecular diversity of HLA-A*02 in Asian Indians: predominance of A*0211. Tissue antigens 57, 502–507 (2001).
Hill, A. V. et al. Common west African HLA antigens are associated with from severe malaria. Nature 352, 595–600 (1991).
Busson, M. et al. HLA-DRB1 and DQB1 allele distribution in the Muong population exposed to malaria in Vietnam. Tissue Antigens 59, 470–474 (2002).
Acknowledgements
This work was supported through the DELTAS Africa Initiative (Grant no. 107743), which funded S.T. through a PhD fellowship award and A.N. through a group leader award. The DELTAS Africa Initiative is an independent funding scheme of the African Academy of Science (AAS) and the Alliance for Accelerating Excellence in Science in Africa and is supported by the New Partnership for Africa’s Development Planning and Coordinating Agency (NEPAD Agency) with funding from the Wellcome Trust (Grant no. 107743) and the UK government. F.C. is funded by Wellcome Trust grant 200841/Z/16/Z. The project received funding from the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant agreement No. 695551) for J. Traherne and J. Trowsdale. J.J. is a recipient of a fellowship from the Centre for Trophoblast Research. The views expressed in this publication are those of the authors and are not necessarily those of AAS, the NEPAD Agency, the Wellcome Trust or the UK government. Figure 2 was drawn using material from Servier Medical Art (https://smart.servier.com/).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
O.C. had started in a role as an employee of AstraZeneca, UK, at the time of manuscript preparation. Other authors have declared that no competing conflicts of interest exist.
Rights and permissions
About this article
Cite this article
Tukwasibwe, S., Nakimuli, A., Traherne, J. et al. Variations in killer-cell immunoglobulin-like receptor and human leukocyte antigen genes and immunity to malaria. Cell Mol Immunol 17, 799–806 (2020). https://doi.org/10.1038/s41423-020-0482-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-020-0482-z
Keywords
This article is cited by
-
Malaria-driven adaptation of MHC class I in wild bonobo populations
Nature Communications (2023)
-
Impact of high human genetic diversity in Africa on immunogenicity and efficacy of RTS,S/AS01 vaccine
Immunogenetics (2023)
-
Pre-referral intranasal artesunate powder for cerebral malaria: a proof-of-concept study
Malaria Journal (2022)
-
Placing human gene families into their evolutionary context
Human Genomics (2022)
-
Neighbor Correlated Graph Convolutional Network for multi-stage malaria parasite recognition
Multimedia Tools and Applications (2022)