Abstract
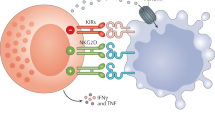
NK cells play important roles in innate defenses against viruses and in the control of tumor growth and metastasis. The regulation/induction of NK cell function is mediated by an array of activating or inhibitory surface receptors. In humans, major activating receptors involved in target cell killing are the natural cytotoxicity receptors (NCRs) and NKG2D. Activating receptors recognize ligands that are overexpressed or expressed de novo upon cell stress, viral infection, or tumor transformation. The HLA-class I-specific inhibitory receptors, including KIRs recognizing HLA-class I allotypic determinants and CD94/NKG2A recognizing the class-Ib HLA-E, constitute a fail-safe mechanism to avoid unwanted NK-mediated damage to healthy cells. Other receptors such as PD-1, primarily expressed by activated T lymphocytes, are important inhibitory checkpoints of immune responses that ensure T-cell tolerance. PD-1 also may be expressed by NK cells in cancer patients. Since PD-1 ligand (PD-L1) may be expressed by different tumors, PD-1/PD-L1 interactions inactivate both T and NK cells. Thus, the reliable evaluation of PD-L1 expression in tumors has become a major issue to select patients who may benefit from therapy with mAbs disrupting PD-1/PD-L1 interactions. Recently, NKG2A was revealed to be an important checkpoint controlling both NK and T-cell activation. Since most tumors express HLA-E, mAbs targeting NKG2A has been used alone or in combination with other therapeutic mAbs targeting PD-1 or tumor antigens (e.g., EGFR), with encouraging results. The translational value of NK cells and their receptors is evidenced by the extraordinary therapeutic success of haploidentical HSCT to cure otherwise fatal high-risk leukemias.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Kiessling, R. et al. Killer cells: a functional comparison between natural, immune T-cell and antibody-dependent in vitro systems. J. Exp. Med. 143, 772–780 (1976).
Herberman, R. B. et al. Fc receptors on mouse effector cells mediating natural cytotoxicity against tumor cells. J. Immunol. 119, 322–326 (1977).
Karre, K., Ljunggren, H. G., Piontek, G. & Kiessling, R. Selective rejection of H-2-deficient lymphoma variants suggests alternative immune defence strategy. Nature 319, 675–678 (1986).
Ljunggren, H. G. & Karre, K. In search of the ‘missing self’: MHC molecules and NK cell recognition. Immunol. Today 11, 237–244 (1990).
Moretta, A. et al. Receptors for HLA class-I molecules in human natural killer cells. Annu. Rev. Immunol. 14, 619–648 (1996).
Moretta, A., Pantaleo, G., Moretta, L., Cerottini, J. C. & Mingari, M. C. Direct demonstration of the clonogenic potential of every human peripheral blood T cell. Clonal analysis of HLA-DR expression and cytolytic activity. J. Exp. Med. 157, 743–754 (1983).
Moretta, A., Pantaleo, G., Moretta, L., Mingari, M. C. & Cerottini, J. C. Quantitative assessment of the pool size and subset distribution of cytolytic T lymphocytes within human resting or alloactivated peripheral blood T cell populations. J. Exp. Med. 158, 571–585 (1983).
Moretta, A. et al. A novel surface antigen expressed by a subset of human CD3- CD16+natural killer cells. Role in cell activation and regulation of cytolytic function. J. Exp. Med. 171, 695–714 (1990).
Moretta, A. et al. Identification of four subsets of human CD3-CD16+natural killer (NK) cells by the expression of clonally distributed functional surface molecules: correlation between subset assignment of NK clones and ability to mediate specific alloantigen recognition. J. Exp. Med. 172, 1589–1598 (1990).
Moretta, A. et al. P58 molecules as putative receptors for major histocompatibility complex (MHC) class I molecules in human natural killer (NK) cells. Anti-p58 antibodies reconstitute lysis of MHC class I-protected cells in NK clones displaying different specificities. J. Exp. Med. 178, 597–604 (1993).
Ciccone, E. et al. Evidence of a natural killer (NK) cell repertoire for (allo) antigen recognition: definition of five distinct NK-determined allospecificities in humans. J. Exp. Med. 175, 709–718 (1992).
Colonna, M. & Samaridis, J. Cloning of immunoglobulin-superfamily members associated with HLA-C and HLA-B recognition by human natural killer cells. Science 268, 405–408 (1995).
Wagtmann, N. et al. Molecular clones of the p58 NK cell receptor reveal immunoglobulin-related molecules with diversity in both the extra- and intracellular domains. Immunity 2, 439–449 (1995).
Braud, V. M. et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 391, 795–799 (1998).
Andre, P. et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing Both T and NK cells. Cell 175, 1731–1743 e1713 (2018).
Mingari, M. C., Pietra, G. & Moretta, L. Immune checkpoint inhibitors: Anti-NKG2A antibodies on board. Trends Immunol. 40, 83–85 (2019).
Anfossi, N. et al. Human NK cell education by inhibitory receptors for MHC class I. Immunity 25, 331–342 (2006).
Kim, S. et al. Licensing of natural killer cells by host major histocompatibility complex class I molecules. Nature 436, 709–713 (2005).
Joncker, N. T., Fernandez, N. C., Treiner, E., Vivier, E. & Raulet, D. H. NK cell responsiveness is tuned commensurate with the number of inhibitory receptors for self-MHC class I: the rheostat model. J. Immunol. 182, 4572–4580 (2009).
Sivori, S. et al. p46, a novel natural killer cell-specific surface molecule that mediates cell activation. J. Exp. Med. 186, 1129–1136 (1997).
Pessino, A. et al. Molecular cloning of NKp46: a novel member of the immunoglobulin superfamily involved in triggering of natural cytotoxicity. J. Exp. Med. 188, 953–960 (1998).
Sivori, S. et al. NKp46 is the major triggering receptor involved in the natural cytotoxicity of fresh or cultured human NK cells. Correlation between surface density of NKp46 and natural cytotoxicity against autologous, allogeneic or xenogeneic target cells. Eur. J. Immunol. 29, 1656–1666 (1999).
Vitale, M. et al. NKp44, a novel triggering surface molecule specifically expressed by activated natural killer cells, is involved in non-major histocompatibility complex-restricted tumor cell lysis. J. Exp. Med. 187, 2065–2072 (1998).
Pende, D. et al. Identification and molecular characterization of NKp30, a novel triggering receptor involved in natural cytotoxicity mediated by human natural killer cells. J. Exp. Med. 190, 1505–1516 (1999).
Moretta, A. et al. Activating receptors and coreceptors involved in human natural killer cell-mediated cytolysis. Annu. Rev. Immunol. 19, 197–223 (2001).
Pazina, T., Shemesh, A., Brusilovsky, M., Porgador, A. & Campbell, K. S. Regulation of the functions of natural cytotoxicity receptors by interactions with diverse ligands and alterations in splice variant expression. Front. Immunol. 8, 369 (2017).
Raffaghello, L. et al. Downregulation and/or release of NKG2D ligands as immune evasion strategy of human neuroblastoma. Neoplasia 6, 558–568 (2004).
Moretta, L. et al. Different checkpoints in human NK-cell activation. Trends Immunol. 25, 670–676 (2004).
Caligiuri, M. A. Human natural killer cells. Blood 112, 461–469 (2008).
Moretta, L. Dissecting CD56dim human NK cells. Blood 116, 3689–3691 (2010).
Hammer, Q. et al. Peptide-specific recognition of human cytomegalovirus strains controls adaptive natural killer cells. Nat. Immunol. 19, 453–463 (2018).
Della Chiesa, M., Sivori, S., Carlomagno, S., Moretta, L. & Moretta, A. Activating KIRs and NKG2C in viral infections: Toward NK cell memory? Front. Immunol. 6, 573 (2015).
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
Vivier, E., van de Pavert, S. A., Cooper, M. D. & Belz, G. T. The evolution of innate lymphoid cells. Nat. Immunol. 17, 790–794 (2016).
Luetke-Eversloh, M., Killig, M. & Romagnani, C. Signatures of human NK cell development and terminal differentiation. Front. Immunol. 4, 499 (2013).
Montaldo, E. et al. Human innate lymphoid cells. Immunol. Lett. 179, 2–8 (2016).
Guia, S., Fenis, A., Vivier, E. & Narni-Mancinelli, E. Activating and inhibitory receptors expressed on innate lymphoid cells. Semin. Immunopathol. 40, 331–341 (2018).
Soriani, A., Stabile, H., Gismondi, A., Santoni, A. & Bernardini, G. Chemokine regulation of innate lymphoid cell tissue distribution and function. Cytokine Growth Factor Rev. 42, 47–55 (2018).
Mingari, M. C. et al. Interleukin-15-induced maturation of human natural killer cells from early thymic precursors: selective expression of CD94/NKG2-A as the only HLA class I-specific inhibitory receptor. Eur. J. Immunol. 27, 1374–1380 (1997).
Yu, J., Freud, A. G. & Caligiuri, M. A. Location and cellular stages of natural killer cell development. Trends Immunol. 34, 573–582 (2013).
Montaldo, E. et al. Human RORgammat(+)CD34(+) cells are lineage-specified progenitors of group 3 RORgammat(+) innate lymphoid cells. Immunity 41, 988–1000 (2014).
Vacca, P. et al. CD34+hematopoietic precursors are present in human decidua and differentiate into natural killer cells upon interaction with stromal cells. Proc. Natl Acad. Sci. USA 108, 2402–2407 (2011).
Poggi, A. et al. Extrathymic differentiation of T lymphocytes and natural killer cells from human embryonic liver precursors. Proc. Natl Acad. Sci. USA 90, 4465–4469 (1993).
Vacca, P., Moretta, L., Moretta, A. & Mingari, M. C. Origin, phenotype and function of human natural killer cells in pregnancy. Trends Immunol. 32, 517–523 (2011).
Parham, P. & Guethlein, L. A. Genetics of natural killer cells in human health, disease, and survival. Annu. Rev. Immunol. 36, 519–548 (2018).
Moretta, A. et al. Existence of both inhibitory (p58) and activatory (p50) receptors for HLA-C molecules in human natural killer cells. J. Exp. Med. 182, 875–884 (1995).
Biassoni, R. et al. The human leukocyte antigen (HLA)-C-specific “activatory” or “inhibitory” natural killer cell receptors display highly homologous extracellular domains but differ in their transmembrane and intracytoplasmic portions. J. Exp. Med. 183, 645–650 (1996).
Chewning, J. H., Gudme, C. N., Hsu, K. C., Selvakumar, A. & Dupont, B. KIR2DS1-positive NK cells mediate alloresponse against the C2 HLA-KIR ligand group in vitro. J. Immunol. 179, 854–868 (2007).
Stewart, C. A. et al. Recognition of peptide-MHC class I complexes by activating killer immunoglobulin-like receptors. Proc. Natl Acad. Sci. USA 102, 13224–13229 (2005).
Graef, T. et al. KIR2DS4 is a product of gene conversion with KIR3DL2 that introduced specificity for HLA-A*11 while diminishing avidity for HLA-C. J. Exp. Med. 206, 2557–2572 (2009).
Burian, A. et al. HLA-F and MHC-I Open Conformers Bind Natural Killer Cell Ig-Like Receptor KIR3DS1. PLoS ONE 11, e0163297 (2016).
Garcia-Beltran, W. F. et al. Open conformers of HLA-F are high-affinity ligands of the activating NK-cell receptor KIR3DS1. Nat. Immunol. 17, 1067–1074 (2016).
Carlomagno, S. et al. KIR3DS1-mediated recognition of HLA-*B51: modulation of KIR3DS1 responsiveness by self HLA-B allotypes and effect on NK cell licensing. Front. Immunol. 8, 581 (2017).
Naiyer, M. M. et al. KIR2DS2 recognizes conserved peptides derived from viral helicases in the context of HLA-C. Sci. Immunol. 2, pii: eaal5296 (2017). https://doi.org/10.1126/sciimmunol.aal5296.
Parham, P. MHC class I molecules and KIRs in human history, health and survival. Nat. Rev. Immunol. 5, 201–214 (2005).
Vitale, M. et al. The leukocyte Ig-like receptor (LIR)-1 for the cytomegalovirus UL18 protein displays a broad specificity for different HLA class I alleles: analysis of LIR-1+NK cell clones. Int. Immunol. 11, 29–35 (1999).
Colonna, M. et al. A common inhibitory receptor for major histocompatibility complex class I molecules on human lymphoid and myelomonocytic cells. J. Exp. Med. 186, 1809–1818 (1997).
Cosman, D. et al. A novel immunoglobulin superfamily receptor for cellular and viral MHC class I molecules. Immunity 7, 273–282 (1997).
Ponte, M. et al. Inhibitory receptors sensing HLA-G1 molecules in pregnancy: decidua-associated natural killer cells express LIR-1 and CD94/NKG2A and acquire p49, an HLA-G1-specific receptor. Proc. Natl Acad. Sci. USA 96, 5674–5679 (1999).
Muccio, L. et al. Analysis of memory-like natural killer cells in human cytomegalovirus-infected children undergoing alphabeta+T and B cell-depleted hematopoietic stem cell transplantation for hematological malignancies. Haematologica 101, 371–381 (2016).
Vales-Gomez, M., Reyburn, H. T., Erskine, R. A., Lopez-Botet, M. & Strominger, J. L. Kinetics and peptide dependency of the binding of the inhibitory NK receptor CD94/NKG2-A and the activating receptor CD94/NKG2-C to HLA-E. EMBO J. 18, 4250–4260 (1999).
Heatley, S. L. et al. Polymorphism in human cytomegalovirus UL40 impacts on recognition of human leukocyte antigen-E (HLA-E) by natural killer cells. J. Biol. Chem. 288, 8679–8690 (2013).
Della Chiesa, M. et al. Features of Memory-Like and PD-1(+) Human NK Cell Subsets. Front. Immunol. 7, 351 (2016).
Guma, M. et al. Expansion of CD94/NKG2C+NK cells in response to human cytomegalovirus-infected fibroblasts. Blood 107, 3624–3631 (2006).
Lopez-Verges, S. et al. Expansion of a unique CD57+NKG2Chi natural killer cell subset during acute human cytomegalovirus infection. Proc. Natl Acad. Sci. USA 108, 14725–14732 (2011).
Tomasello, E. et al. Gene structure, expression pattern, and biological activity of mouse killer cell activating receptor-associated protein (KARAP)/DAP-12. J. Biol. Chem. 273, 34115–34119 (1998).
Campbell, K. S., Yusa, S., Kikuchi-Maki, A. & Catina, T. L. NKp44 triggers NK cell activation through DAP12 association that is not influenced by a putative cytoplasmic inhibitory sequence. J. Immunol. 172, 899–906 (2004).
Muccio, L. et al. Late development of Fc epsilon R gamma(neg) adaptive natural killer cells upon human cytomegalovirus reactivation in umbilical cord blood transplantation recipients. Front. Immunol. 9, 1050 (2018).
Arnon, T. I., Markel, G. & Mandelboim, O. Tumor and viral recognition by natural killer cells receptors. Semin. Cancer Biol. 16, 348–358 (2006).
Arnon, T. I. et al. Inhibition of the NKp30 activating receptor by pp65 of human cytomegalovirus. Nat. Immunol. 6, 515–523 (2005).
Pogge von Strandmann, E. et al. Human leukocyte antigen-B-associated transcript 3 is released from tumor cells and engages the NKp30 receptor on natural killer cells. Immunity 27, 965–974 (2007).
Reiners, K. S. et al. Soluble ligands for NK cell receptors promote evasion of chronic lymphocytic leukemia cells from NK cell anti-tumor activity. Blood 121, 3658–3665 (2013).
Baychelier, F. et al. Identification of a cellular ligand for the natural cytotoxicity receptor NKp44. Blood 122, 2935–2942 (2013).
Rosental, B. et al. Proliferating cell nuclear antigen is a novel inhibitory ligand for the natural cytotoxicity receptor NKp44. J. Immunol. 187, 5693–5702 (2011).
Brandt, C. S. et al. The B7 family member B7-H6 is a tumor cell ligand for the activating natural killer cell receptor NKp30 in humans. J. Exp. Med. 206, 1495–1503 (2009).
Textor, S. et al. The proto-oncogene Myc drives expression of the NK cell-activating NKp30 ligand B7-H6 in tumor cells. Oncoimmunology 5, e1116674 (2016).
Matta, J. et al. Induction of B7-H6, a ligand for the natural killer cell-activating receptor NKp30, in inflammatory conditions. Blood 122, 394–404 (2013).
Narni-Mancinelli, E. et al. Complement factor P is a ligand for the natural killer cell-activating receptor NKp46. Sci. Immunol. 2, pii: eaam9628 (2017). https://doi.org/10.1126/sciimmunol.aam9628.
Gaggero, S. et al. Nidogen-1 is a novel extracellular ligand for the NKp44 activating receptor. Oncoimmunology 7, e1470730 (2018).
Schlecker, E. et al. Metalloprotease-mediated tumor cell shedding of B7-H6, the ligand of the natural killer cell-activating receptor NKp30. Cancer Res. 74, 3429–3440 (2014).
Pesce, S. et al. B7-H6-mediated downregulation of NKp30 in NK cells contributes to ovarian carcinoma immune escape. Oncoimmunology 4, e1001224 (2015).
De Maria, A. et al. The impaired NK cell cytolytic function in viremic HIV-1 infection is associated with a reduced surface expression of natural cytotoxicity receptors (NKp46, NKp30 and NKp44). Eur. J. Immunol. 33, 2410–2418 (2003).
Bauer, S. et al. Activation of NK cells and T cells by NKG2D, a receptor for stress-inducible MICA. Science 285, 727–729 (1999).
Raulet, D. H., Gasser, S., Gowen, B. G., Deng, W. & Jung, H. Regulation of ligands for the NKG2D activating receptor. Annu. Rev. Immunol. 31, 413–441 (2013).
Lanier, L. L. NKG2D receptor and its ligands in host defense. Cancer Immunol. Res 3, 575–582 (2015).
Sivori, S. et al. 2B4 functions as a co-receptor in human NK cell activation. Eur. J. Immunol. 30, 787–793 (2000).
Bottino, C. et al. NTB-A [correction of GNTB-A], a novel SH2D1A-associated surface molecule contributing to the inability of natural killer cells to kill Epstein-Barr virus-infected B cells in X-linked lymphoproliferative disease. J. Exp. Med. 194, 235–246 (2001).
Shibuya, A. et al. DNAM-1, a novel adhesion molecule involved in the cytolytic function of T lymphocytes. Immunity 4, 573–581 (1996).
Marcenaro, E. et al. CD59 is physically and functionally associated with natural cytotoxicity receptors and activates human NK cell-mediated cytotoxicity. Eur. J. Immunol. 33, 3367–3376 (2003).
Vitale, M. et al. Identification of NKp80, a novel triggering molecule expressed by human NK cells. Eur. J. Immunol. 31, 233–242 (2001).
Sivori, S. et al. CpG and double-stranded RNA trigger human NK cells by Toll-like receptors: induction of cytokine release and cytotoxicity against tumors and dendritic cells. Proc. Natl Acad. Sci. USA 101, 10116–10121 (2004).
Sivori, S., Carlomagno, S., Moretta, L. & Moretta, A. Comparison of different CpG oligodeoxynucleotide classes for their capability to stimulate human NK cells. Eur. J. Immunol. 36, 961–967 (2006).
Sivori, S. et al. Heterogeneity of TLR3 mRNA transcripts and responsiveness to poly (I:C) in human NK cells derived from different donors. Int. Immunol. 19, 1341–1348 (2007).
Sivori, S. et al. A novel KIR-associated function: evidence that CpG DNA uptake and shuttling to early endosomes is mediated by KIR3DL2. Blood 116, 1637–1647 (2010).
Ochoa, M. C. et al. Antibody-dependent cell cytotoxicity: immunotherapy strategies enhancing effector NK cells. Immunol. Cell Biol. 95, 347–355 (2017).
Vivier, E. et al. Innate or adaptive immunity? The example of natural killer cells. Science 331, 44–49 (2011).
Chiossone, L., Vienne, M., Kerdiles, Y. M. & Vivier, E. Natural killer cell immunotherapies against cancer: checkpoint inhibitors and more. Semin. Immunol. 31, 55–63 (2017).
Munari, E. et al. PD-L1 expression comparison between primary and relapsed non-small cell lung carcinoma using whole sections and clone SP263. Oncotarget 9, 30465–30471 (2018).
Munari, E. et al. PD-L1 Expression heterogeneity in non-small cell lung cancer: Defining criteria for harmonization between biopsy specimens and whole sections. J. Thorac. Oncol. 13, 1113–1120 (2018).
Ishida, Y., Agata, Y., Shibahara, K. & Honjo, T. Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. EMBO J. 11, 3887–3895 (1992).
Nishimura, H., Nose, M., Hiai, H., Minato, N. & Honjo, T. Development of lupus-like autoimmune diseases by disruption of the PD-1 gene encoding an ITIM motif-carrying immunoreceptor. Immunity 11, 141–151 (1999).
Okazaki, T., Chikuma, S., Iwai, Y., Fagarasan, S. & Honjo, T. A rheostat for immune responses: the unique properties of PD-1 and their advantages for clinical application. Nat. Immunol. 14, 1212–1218 (2013).
Huang, A. C. et al. T-cell invigoration to tumour burden ratio associated with anti-PD-1 response. Nature 545, 60–65 (2017).
Pesce, S. et al. Identification of a subset of human natural killer cells expressing high levels of programmed death 1: A phenotypic and functional characterization. J. Allergy Clin. Immunol. 139, 335–346 (2017).
Beldi-Ferchiou, A. et al. PD-1 mediates functional exhaustion of activated NK cells in patients with Kaposi sarcoma. Oncotarget 7, 72961–72977 (2016).
Iraolagoitia, X. L. et al. NK cells restrain spontaneous antitumor CD8+T cell priming through PD-1/PD-L1 interactions with dendritic cells. J. Immunol. 197, 953–961 (2016).
Vari, F. et al. Immune evasion via PD-1/PD-L1 on NK cells and monocyte/macrophages is more prominent in Hodgkin lymphoma than DLBCL. Blood 131, 1809–1819 (2018).
Quatrini, L. et al. Endogenous glucocorticoids control host resistance to viral infection through the tissue-specific regulation of PD-1 expression on NK cells. Nat. Immunol. 19, 954–962 (2018).
Hsu, J. et al. Contribution of NK cells to immunotherapy mediated by PD-1/PD-L1 blockade. J. Clin. Invest. 128, 4654–4668 (2018).
Bottino, C. et al. Identification of PVR (CD155) and Nectin-2 (CD112) as cell surface ligands for the human DNAM-1 (CD226) activating molecule. J. Exp. Med. 198, 557–567 (2003).
Dougall, W. C., Kurtulus, S., Smyth, M. J. & Anderson, A. C. TIGIT and CD96: new checkpoint receptor targets for cancer immunotherapy. Immunol. Rev. 276, 112–120 (2017).
Zhou, X. M. et al. Intrinsic expression of immune checkpoint molecule TIGIT could help tumor growth in vivo by suppressing the function of NK and CD8(+) T Cells. Front. Immunol. 9, 2821 (2018).
Zhang, Q. et al. Blockade of the checkpoint receptor TIGIT prevents NK cell exhaustion and elicits potent anti-tumor immunity. Nat. Immunol. 19, 723–732 (2018).
Hung, A. L. et al. TIGIT and PD-1 dual checkpoint blockade enhances antitumor immunity and survival in GBM. Oncoimmunology 7, e1466769 (2018).
Huntington, N. D. et al. NK cell maturation and peripheral homeostasis is associated with KLRG1 up-regulation. J. Immunol. 178, 4764–4770 (2007).
Huang, Y. et al. IL-25-responsive, lineage-negative KLRG1(hi) cells are multipotential ‘inflammatory’ type 2 innate lymphoid cells. Nat. Immunol. 16, 161–169 (2015).
Ito, M. et al. Killer cell lectin-like receptor G1 binds three members of the classical cadherin family to inhibit NK cell cytotoxicity. J. Exp. Med. 203, 289–295 (2006).
Salimi, M. et al. A role for IL-25 and IL-33-driven type-2 innate lymphoid cells in atopic dermatitis. J. Exp. Med. 210, 2939–2950 (2013).
Simoni, Y. et al. Human innate lymphoid cell subsets possess tissue-type based heterogeneity in phenotype and frequency. Immunity 46, 148–161 (2017).
Carrega, P. et al. NCR(+)ILC3 concentrate in human lung cancer and associate with intratumoral lymphoid structures. Nat. Commun. 6, 8280 (2015).
Baixeras, E. et al. Characterization of the lymphocyte activation gene 3-encoded protein. A new ligand for human leukocyte antigen class II antigens. J. Exp. Med. 176, 327–337 (1992).
Lichtenegger, F. S. et al. Targeting LAG-3 and PD-1 to enhance T cell activation by antigen-presenting cells. Front. Immunol. 9, 385 (2018).
Woo, S. R. et al. Immune inhibitory molecules LAG-3 and PD-1 synergistically regulate T-cell function to promote tumoral immune escape. Cancer Res. 72, 917–927 (2012).
Miyazaki, T., Dierich, A., Benoist, C. & Mathis, D. LAG-3 is not responsible for selecting T helper cells in CD4-deficient mice. Int. Immunol. 8, 725–729 (1996).
Wada, J. & Kanwar, Y. S. Identification and characterization of galectin-9, a novel beta-galactoside-binding mammalian lectin. J. Biol. Chem. 272, 6078–6086 (1997).
DeKruyff, R. H. et al. T cell/transmembrane, Ig, and mucin-3 allelic variants differentially recognize phosphatidylserine and mediate phagocytosis of apoptotic cells. J. Immunol. 184, 1918–1930 (2010).
Chiba, S. et al. Tumor-infiltrating DCs suppress nucleic acid-mediated innate immune responses through interactions between the receptor TIM-3 and the alarmin HMGB1. Nat. Immunol. 13, 832–842 (2012).
Huang, Y. H. et al. CEACAM1 regulates TIM-3-mediated tolerance and exhaustion. Nature 517, 386–390 (2015).
Gao, X. et al. TIM-3 expression characterizes regulatory T cells in tumor tissues and is associated with lung cancer progression. PLoS ONE 7, e30676 (2012).
Lu, X. et al. Tumor antigen-specific CD8(+) T cells are negatively regulated by PD-1 and Tim-3 in human gastric cancer. Cell Immunol. 313, 43–51 (2017).
Shayan, G. et al. Adaptive resistance to anti-PD1 therapy by Tim-3 upregulation is mediated by the PI3K-Akt pathway in head and neck cancer. Oncoimmunology 6, e1261779 (2017).
Fourcade, J. et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+T cell dysfunction in melanoma patients. J. Exp. Med. 207, 2175–2186 (2010).
da Silva, I. P. et al. Reversal of NK-cell exhaustion in advanced melanoma by Tim-3 blockade. Cancer Immunol. Res 2, 410–422 (2014).
Xu, L. et al. Increased Tim-3 expression in peripheral NK cells predicts a poorer prognosis and Tim-3 blockade improves NK cell-mediated cytotoxicity in human lung adenocarcinoma. Int. Immunopharmacol. 29, 635–641 (2015).
Das, M., Zhu, C. & Kuchroo, V. K. Tim-3 and its role in regulating anti-tumor immunity. Immunol. Rev. 276, 97–111 (2017).
Thomassen, E., Renshaw, B. R. & Sims, J. E. Identification and characterization of SIGIRR, a molecule representing a novel subtype of the IL-1R superfamily. Cytokine 11, 389–399 (1999).
Mantovani, A., Locati, M., Polentarutti, N., Vecchi, A. & Garlanda, C. Extracellular and intracellular decoys in the tuning of inflammatory cytokines and Toll-like receptors: the new entry TIR8/SIGIRR. J. Leukoc. Biol. 75, 738–742 (2004).
Molgora, M. et al. IL-1R8 is a checkpoint in NK cells regulating anti-tumour and anti-viral activity. Nature 551, 110–114 (2017).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Gong, J., Chehrazi-Raffle, A., Reddi, S. & Salgia, R. Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: a comprehensive review of registration trials and future considerations. J. Immunother. Cancer 6, 8 (2018).
Migden, M. R. et al. PD-1 blockade with cemiplimab in advanced cutaneous squamous-cell carcinoma. N. Engl. J. Med. 379, 341–351 (2018).
Buttner, R. et al. Programmed death-ligand 1 immunohistochemistry testing: A review of analytical assays and clinical implementation in non-small-cell lung cancer. J. Clin. Oncol.: Off. J. Am. Soc. Clin. Oncol. 35, 3867–3876 (2017).
Munari, E. et al. PD-L1 Assays 22C3 and SP263 are not interchangeable in non-small cell lung cancer when considering clinically relevant cutoffs: An interclone evaluation by differently trained pathologists. Am. J. Surg. Pathol. 42, 1384–1389 (2018).
Munari, E. et al. PD-L1 expression heterogeneity in non-small cell lung cancer: evaluation of small biopsies reliability. Oncotarget 8, 90123–90131 (2017).
Munari, E. et al. Expression of programmed cell death ligand 1 in non-small cell lung cancer: Comparison between cytologic smears, core biopsies, and whole sections using the SP263 Assay. Cancer Cytopathol. 127, 52–61 (2018).
Pietra, G. et al. Human natural killer cells: news in the therapy of solid tumors and high-risk leukemias. Cancer Immunol. Immunother. 65, 465–476 (2016).
Kim, N. & Kim, H. S. Targeting checkpoint receptors and molecules for therapeutic modulation of natural killer cells. Front. Immunol. 9, 2041 (2018).
Sivori, S. et al. Inhibitory CD94 molecules identified by the Z199 monoclonal antibody recognize different HLA-class I molecules. Transplant. Proc. 28, 3199–3203 (1996).
Pende, D. et al. HLA-G recognition by human natural killer cells. Involvement of CD94 both as inhibitory and as activating receptor complex. Eur. J. Immunol. 27, 1875–1880 (1997).
Morandi, F., Rizzo, R., Fainardi, E., Rouas-Freiss, N. & Pistoia, V. Recent advances in our understanding of HLA-G biology: Lessons from a wide spectrum of human diseases. J. Immunol. Res. 2016, 4326495 (2016).
Vitale, M., Cantoni, C., Pietra, G., Mingari, M. C. & Moretta, L. Effect of tumor cells and tumor microenvironment on NK-cell function. Eur. J. Immunol. 44, 1582–1592 (2014).
Mingari, M. C. et al. Cytolytic T lymphocytes displaying natural killer (NK)-like activity: expression of NK-related functional receptors for HLA class I molecules (p58 and CD94) and inhibitory effect on the TCR-mediated target cell lysis or lymphokine production. Int. Immunol. 7, 697–703 (1995).
Mingari, M. C. et al. Human CD8 + T lymphocyte subsets that express HLA class I-specific inhibitory receptors represent oligoclonally or monoclonally expanded cell populations. Proc. Natl Acad. Sci. USA 93, 12433–12438 (1996).
Mingari, M. C. et al. HLA class I-specific inhibitory receptors in human T lymphocytes: interleukin 15-induced expression of CD94/NKG2A in superantigen- or alloantigen-activated CD8+T cells. Proc. Natl Acad. Sci. USA 95, 1172–1177 (1998).
Bertone, S. et al. Transforming growth factor-beta-induced expression of CD94/NKG2A inhibitory receptors in human T lymphocytes. Eur. J. Immunol. 29, 23–29 (1999).
Ruggeri, L. et al. Effectiveness of donor natural killer cell alloreactivity in mismatched hematopoietic transplants. Science 295, 2097–2100 (2002).
Pende, D. et al. Anti-leukemia activity of alloreactive NK cells in KIR ligand-mismatched haploidentical HSCT for pediatric patients: evaluation of the functional role of activating KIR and redefinition of inhibitory KIR specificity. Blood 113, 3119–3129 (2009).
Pende, D. et al. Analysis of the receptor-ligand interactions in the natural killer-mediated lysis of freshly isolated myeloid or lymphoblastic leukemias: evidence for the involvement of the Poliovirus receptor (CD155) and Nectin-2 (CD112). Blood 105, 2066–2073 (2005).
Pistoia, V. et al. Human gammadelta T-Cells: From surface receptors to the therapy of high-risk leukemias. Front. Immunol. 9, 984 (2018).
Locatelli, F. et al. NK cells mediate a crucial graft-versus-leukemia effect in haploidentical-HSCT to Cure High-risk Acute Leukemia. Trends Immunol. 39, 577–590 (2018).
Moretta, F. et al. The generation of human innate lymphoid cells is influenced by the source of hematopoietic stem cells and by the use of G-CSF. Eur. J. Immunol. 46, 1271–1278 (2016).
Balsamo, M. et al. Melanoma cells become resistant to NK-cell-mediated killing when exposed to NK-cell numbers compatible with NK-cell infiltration in the tumor. Eur. J. Immunol. 42, 1833–1842 (2012).
Huergo-Zapico, L. et al. NK-cell editing mediates epithelial-to-mesenchymal transition via phenotypic and proteomic changes in melanoma cell lines. Cancer Res. 78, 3913–3925 (2018).
Rajagopalan, S. & Long, E. O. KIR2DL4 (CD158d): An activation receptor for HLA-G. Front Immunol 3, 258 (2012). https://doi.org/10.3389/fimmu.2012.00258.
Acknowledgements
This study is dedicated to Alessandro Moretta, who sadly passed away on February 17, 2018. His seminal discoveries of inhibitory and activating NK receptors made it possible to understand how NK cells function. Many of the issues described in this review are based on his pioneering studies. We greatly miss his scientific insight and even more his humanity and smile. Supported by grants awarded by Associazione Italiana per la Ricerca sul Cancro (AIRC)-Special Program Metastatic disease: the key unmet need in oncology 5 per mille 2018 Id. 21147 (S.S. and L.M.), AIRC IG2017 Id. 20312 (S.S.), AIRC IG2017 Id.19920 (L.M.), RC-2018 OPBG (P.V. and L.M.); 5 × 1000 Italian Ministry of Health 2015 (M.C.M); and Ministero della Salute RF-2013, GR-2013-02356568 (P.V.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Sivori, S., Vacca, P., Del Zotto, G. et al. Human NK cells: surface receptors, inhibitory checkpoints, and translational applications. Cell Mol Immunol 16, 430–441 (2019). https://doi.org/10.1038/s41423-019-0206-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-019-0206-4
Keywords
This article is cited by
-
CXCR6-positive circulating mucosal-associated invariant T cells can identify patients with non-small cell lung cancer responding to anti-PD-1 immunotherapy
Journal of Experimental & Clinical Cancer Research (2024)
-
The causality between CD8+NKT cells and CD16−CD56 on NK cells with hepatocellular carcinoma: a Mendelian randomization study
Infectious Agents and Cancer (2024)
-
Unraveling the dynamic mechanisms of natural killer cells in viral infections: insights and implications
Virology Journal (2024)
-
BET inhibitors drive Natural Killer activation in non-small cell lung cancer via BRD4 and SMAD3
Nature Communications (2024)
-
Inhibition of USP7 enhances CD8+ T cell activity in liver cancer by suppressing PRDM1-mediated FGL1 upregulation
Acta Pharmacologica Sinica (2024)