Abstract
Cancer is one of the most fatal diseases that threaten human health, whereas more than 90% mortality of cancer patients is caused by tumor metastasis, rather than the growth of primary tumors. Thus, how to effectively control or even reverse the migration of tumor cells is of great significance for cancer therapy. CtBP, a transcriptional cofactor displaying high expression in a variety of human cancers, has become one of the main targets for cancer prediction, diagnosis, and treatment. The roles of CtBP in promoting tumorigenesis have been well studied in vitro, mostly based on gain-of-function, while its physiological functions in tumor invasion and the underlying mechanism remain largely elusive. Snail (Sna) is a well-known transcription factor involved in epithelial-to-mesenchymal transition (EMT) and tumor invasion, yet the mechanism that regulates Sna activity has not been fully understood. Using Drosophila as a model organism, we found that depletion of CtBP or snail (sna) suppressed RasV12/lgl-/--triggered tumor growth and invasion, and disrupted cell polarity-induced invasive cell migration. In addition, loss of CtBP inhibits RasV12/Sna-induced tumor invasion and Sna-mediated invasive cell migration. Furthermore, both CtBP and Sna are physiologically required for developmental cell migration during thorax closure. Finally, Sna activates the JNK signaling and promotes JNK-dependent cell invasion. Given that CtBP physically interacts with Sna, our data suggest that CtBP and Sna may form a transcriptional complex that regulates JNK-dependent tumor invasion and cell migration in vivo.
Similar content being viewed by others
Introduction
Tumor metastasis is a major contributor to the high mortality rate of cancer and accounts for more than 90% of cancer-related fatalities in patients with palpable clinical traits1,2. Metastasis is a process of cancer cells disseminating from a primary lesion via lymphatic and/or blood circulations to distal organs, which involves a variety of cellular mechanisms3. These include invading through basement membranes (BM), escaping immune surveillance, modulating tissue microenvironment, and evolving resistance to therapeutic intervention4,5,6. Therefore, how to effectively control and reverse tumor invasion is of great significance to the clinical treatment of malignant cancer. Over the past decades, great progress has been made in exploring the mechanisms of tumor progression, in which numerous oncogenes, tumor suppressor genes, and multiple signaling pathways (e.g., Raf-MAPK, JNK, WNT, Hippo, Notch, JAK-STAT, and PI3K/AKT) have been implicated in tumor growth and invasion7,8,9,10. Among them, the oncogenic carboxy-terminal binding protein (CtBP) family members are the widely concerned because of their overexpression across an extensive spectrum of solid human tumors, including bladder, breast, ovarian, gastric, prostate, and sarcoma cancer, which have become the main targets for cancer prediction, diagnosis, and treatment11,12,13.
CtBP is a well-known and evolutionarily conserved transcriptional coregulator that was initially identified through its interaction with the human adenovirus E1A protein and plays a crucial role in regulating cell survival14,15. Although the invertebrate (nematodes, fruit flies, etc.) genome encodes a single CtBP protein, the vertebrate (mouse, rats, human, etc.) genome expresses two CtBP proteins (CtBP1 and CtBP2) that perform both unique and redundant functions11. Usually, CtBP binds to a variety of transcription repressors, such as Snail, Knirp, and Krüppel, through its N-terminal dehydrogenase PxDLS (Pro-x-Asp–Leu–Ser) motif, and recruits chromatin-modifying enzymes to form transcription repressor complex, which targets specific DNA promoter regions16,17. Extensive genetic and biochemical studies in model organisms have demonstrated that CtBP is indispensable for embryonic development and adult lifespan regulation. The homozygous mutation of mCtBP2 in mouse leads to developmental defects and embryonic death, while mCtBP1 homozygous deletion reduces their offsprings’ life span18. Conversely, loss of CtBP either by depletion or mutation triggered an extended life span in C. elegans19. In addition to its role in development, CtBP-mediated transcriptional coregulation plays important roles in a variety of diseases, especially cancer20,21,22. CtBP1/2 are highly expressed in several human cancer types, with their expression level correlating to the poor prognostic outcomes and aggressive tumor characteristics. In 2013, a comprehensive description of CtBP inhibition targets was achieved by the genome-wide analysis, in which the targets are mainly categorized into the following: genes that regulate DNA damage repair and genome stability, genes that regulate cell apoptosis and proliferation, and genes that regulate epithelial differentiation and impede epithelial-to-mesenchymal transition (EMT)23. Of note, signaling pathways related to these three categories' genes are generally dysregulated in cancer20. Besides, CtBP also functions as a coactivator to accelerate tumorigenesis by promoting cancer stem cell self-renewal11. However, despite many advances being made in elucidating the tumor-promoting effects of CtBP, most of the studies are based on gain of function, in vitro, or cell culture experiments, it remains poorly understood whether it could be the case in vivo. Whether endogenous CtBP is involved in cell invasion and EMT, or interacts with tumor-related signal pathways, as well as the potential molecular mechanisms, needs to be further explored.
Snail (Sna) belongs to the Sna transcriptional factor family, which was first identified in Drosophila as a critical regulator of mesoderm formation during embryonic development24. Snail family members have a conserved C-terminal DNA-binding domain containing four–six C2H2-type zinc fingers and a SNAG domain in N-terminus25. Generally, Sna acts as a transcriptional repressor through its SNAG domain to suppress the target gene expression26. Sna could also positively regulate transcription, and this functional switch of Sna may depend on its cofactor27,28. Sna has been extensively studied for its role in various biological processes, including embryonic development, cell fate decision, and cell differentiation24,29,30,31. In Drosophila, sna homozygous mutant embryos show defective in mesoderm formation24. Murine SNAI/SLUG interact with YAP/TAZ to control skeletal stem cell differentiation32. Besides, Sna is a well-known modulator of epithelial–mesenchymal transition (EMT) and tumor invasion33. SNAI is highly expressed in multiple types of cancer cell lines, and its expression correlates with increased metastasis potential. Sna family proteins regulate the transcription of a large number of genes essential for EMT and tumor progression34,35, for instance, SNAI directly represses the expression of epithelial markers like E-cadherin36, while upregulates that of mesenchymal markers like MMP2/9 to promote EMT37. Despite its well-established role in EMT and tumor invasion, the mechanism that regulates Sna activity in cancer progression has not been completely understood.
In this work, we found that Drosophila CtBP and Sna are physiologically required for RasV12/lgl-/- triggered tumor growth and invasion, and loss-of-cell polarity-induced invasive cell migration. We further showed that CtBP is indispensable for Sna-induced cell migration and tumor invasion. Moreover, Sna and CtBP regulate cell migration in thorax development. Finally, Sna activates the JNK signaling and promotes JNK-dependent cell migration. Taken together, these findings provide the in vivo evidences and the underlying mechanism for the role of CtBP in Sna-mediated cell migration and tumor invasion, and offer therapeutic strategies for clinical treatment of cancer and other related diseases.
Results and discussion
Loss-of-CtBP suppresses RasV12/lgl -/- induced tumor growth and invasion
In line with previous studies38,39, clones of GFP-marked wild-type cells mediated by eyeless (ey)-Flp/MARCM system were observed in the larval eye–antennal imaginal disks and the brain optic lobes (Fig. 1a and Supplementary Fig. 1a), but were not seen in the adjacent ventral nerve cord (VNC) of the central nervous system (Fig. 1i). While the GFP-labeled clones expressing activated Ras (RasV12) alone caused noticeable growth without invading to the VNC (Fig. 1b, j; Supplementary Fig. 1b), ectopic expression of RasV12 in lgl4 homozygous mutant (lgl-/-) mosaic clones resulted in massive tumor-like overgrowth (Fig. 1c; Supplementary Fig. 1c) and invasive metastasis to the VNC (Fig. 1k, q). Besides, the invasive tumor cells triggered an extended larval stage, which impeded the normal development of larvae into pupae, and animals died as bloated third instar larvae (Supplementary Fig. 1c, h). These phenotypes, mediated by the c-Jun N-terminal kinase (JNK) pathway40, were modestly enhanced by pucE69 heterozygosity and effectively blocked by the expression of Puckered (Puc) (Fig. 1e, f, m and n; Supplementary Fig. 1e, h), a phosphatase and inhibitor of JNK41. Consistently, the expression of puc-LacZ, a reporter of JNK pathway, was strongly upregulated in RasV12/lgl-/- tumor cells, compared with control or RasV12 clones (Supplementary Fig. 2c–e).
Fluorescent images showing Drosophila larval cephalic complexes (CC, a–h) and ventral nerve cords (VNC, i-p), the anterior is to the up in all panels. The CC (a) contains eye–antennal disks (EA), brain hemispheres (BH), and VNC (h). GFP-labeled mosaic clones were created in the EA. Compared with the control (a, h), RasV12/lgl-/- induced tumor overgrowth (c) and invasion to the VNC (j) were enhanced by pucE69 heterozygosity (e, m), and strongly suppressed by expressing Puc (f, n), or depleting CtBP (g, o, h and p), but not expressing LacZ (d, l). RasV12-overexpressing clones showed a visible growth advantage (b), but did not migrate to the VNC (j). Statistical analysis of the invasion percentage (q) as shown in figures a–h (a, 0.00%, n = 52; b, 0.00%, n = 55; c, 76.27%, n = 118; d, 72.58%, n = 124; e, 86.96%, n = 92; f, 5.09%, n = 118; g, 18.33%, n = 120; h, 35.19%, n = 108), respectively. Chi-squared test was applied to compute P-values, *P < 0.05, ***P < 0.001; ns, no significant difference. See the electronic supplementary material for detailed genotypes. Scale bar: 100 µm (a–p).
Using this RasV12/lgl-/- in vivo tumor model, we have conducted a genetic screen for regulators of tumor growth and invasion39,42,43,44,45. We found that the tumor overgrowth and metastasis phenotypes were strongly suppressed by RNA interference (RNAi)-mediated knockdown of CtBP (Fig. 1g, o, q; Supplementary Fig. 1f, h). To exclude the possibility of off-target effect from RNAi, we obtained from the Bloomington Drosophila stock center another independent CtBP RNAi line that targets a distinct region of the CtBP transcript46, and observed a similar, albeit moderate, suppression on tumor growth and invasion to VNC (Fig. 1h, p and q). Meanwhile, expression of CtBP RNAi in otherwise wild-type clones had no effect on the clonal growth (Supplementary Fig. 1g and Supplementary Fig. 3). qRT-PCR assay was performed to verify the knockdown efficiencies of the CtBP RNAi lines (Supplementary Fig. 2a). To exclude the possibility that the suppression effect of CtBP RNAi is a result of UAS titration, UAS-LacZ was included as a negative control (Fig. 1d, l and q and Supplementary Fig. 1d, h). Consistent with its role in RasV12/lgl-/--triggered tumor progression, we found that CtBP expression was increased in tumors (Supplementary Fig. 2b). Collectively, these results indicate that the transcriptional corepressor CtBP plays an essential role in RasV12/lgl-/- promoted tumor growth and invasion.
To examine whether loss-of-CtBP suppresses tumor invasion by increasing cell death or reducing cell proliferation, we checked cell death by cDcp-1 antibody staining and cell proliferation by Phospho-Histone H3 (PH3) staining in CtBP null mutant clones. Loss-of-CtBP did not cause enhanced cell death (Supplementary Fig. 2f–g) or reduced cell proliferation (Supplementary Fig. 4), suggesting that CtBP regulates tumor invasion independent of cell death and cell proliferation.
CtBP is required for disrupted cell polarity-induced cell invasion
To verify the physiological function of CtBP in cell invasion, we employed another well-established invasion model42,47. In the epithelia of Drosophila larval wing imaginal disks, knockdown of cell polarity genes, e.g., scrib, lgl, or dlg, driven by patched (ptc)-GAL4 in the anterior/posterior (A/P) compartment boundary, induced a JNK-dependent cell invasion phenotype43,48. Consistently, GFP-marked depletion-of-scrib resulted in broadscale cell migration toward the P compartment, coupled with upregulation of matrix metalloprotease 1 (MMP1) (Fig. 2a–a”, b–b”), which is essential for basement membrane degradation and a molecular feature of EMT49,50. We found that ptc > scrib-IR-triggered cell invasion and MMP1 activation were notably blocked by depletion of CtBP (Fig. 2d–d”, e–e”) or expression of Puc (Fig. 2f–f”), but remained unaffected by LacZ expression (Fig. 2c–c”). To quantify this phenotype, we counted the total number of migrating cells in the wing pouch region, and found that depletion of CtBP reduced the number by 86.48% (CtBP-IR32889) or 84.64% (CtBP-IR31334), which is comparable to that of Puc expression (92.85%), while LacZ served as a negative control (Fig. 2g). Taking these data together, we conclude that CtBP is required for cell polarity disruption-triggered cell invasion and MMP1 upregulation.
a–f Fluorescence micrographs of 3rd instar larval wing disks stained with anti-MMP1 antibody are shown, anterior is to the left and cells are marked with GFP expression. Compared with the control (a–a”), loss of scrib induced intensive cell migration and MMP1 upregulation (b–b”), which remained unchanged by expressing LacZ (c–c”), but was dramatically impeded by knockdown of CtBP (d–d”, e–e”), or expression of Puc (f–f”). g Column bar graph of the migrating cell number in a–g (n = 10 for each genotype; a, mean = 0.02; b, mean = 89.50; c, mean = 76.00; d, mean = 12.10; e, mean = 13.75; f, mean = 6.40), error bars indicate standard deviation. One-way ANOVA with Bonferroni multiple comparison test was used to compute P-values, ****P < 0.0001; ns, no significant difference. h Schematic diagram of the interaction between CtBP and Sna (generated by STRING 11.0 online analysis platform). Colored nodes: query proteins and first shell of interactors; filled nodes: 3D structure is known or predicted; rose red line: experimentally determined; dark-yellow line: textmining. The combined interaction score is 0.923. See the electronic supplementary material for detailed genotypes. Scale bar: 20 µm (a–f).
An evolutionary conserved role of Sna in tumor invasion
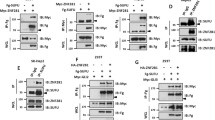
To uncover the mechanism by which CtBP modulates tumor invasion, we considered the transcription factor Snail (Sna) as a putative factor that cooperates with CtBP. First, Sna can interact with CtBP through its Pro-X-Asp–Leu–Ser-X-Lys (P-DLS-K) motif, and then recruit chromatin-modifying enzymes to form transcription repressors that bind to the promoter regions of target genes during development51,52. Second, the interaction score between Sna and CtBP, generated by the STRING 11.0 online analysis platform (http://string-db.org), is 0.923 with a high confidence (Fig. 2h). Third, previous studies reported that Sna is involved in controlling EMT during tumor progression, whose expression correlates with the tumor grade, nodal metastasis of multiple tumors, and indicates a poor outcome in patients with malignant tumor53.
Although the role of Sna in tumor invasion has been well studied in mammals, it remains unknown whether this function is conserved in Drosophila. To test this, we first examined the physiological role of Sna in the RasV12/lgl-/- tumor model. Based on previous work54, we selected from Vienna Drosophila RNAi center (VDRC) a sna RNAi line with a high knockdown efficiency, and found that depletion of sna significantly inhibited RasV12/lgl-/- triggered tumor growth and invasion (Fig. 3a–d and g), and 16.36% larvae successfully developed into pupal stage (Fig. 3e, f and h). Furthermore, ptc > scrib-IR-induced cell invasion and MMP1 upregulation were significantly suppressed by depleting sna (Fig. 3i–i”, j–j” and k). Finally, we investigated the ability of Sna to promote tumor invasion in Drosophila. While ectopic expression of Sna alone in the eye disks failed to induce any tumor-like growth and invasion (Fig. 4b, g and k), it is sufficient to promote the invasion of RasV12-expressing cells into the VNC (Fig. 4a, c, f, h and k). Thus, the critical role of Sna in tumor invasion has been conserved from fly to human.
(a–f, i–i” and j–j”) Fluorescence micrographs of larval CCs (a, b), VNCs (c, d), whole bodies (e, f) and wing discs (i–i”, j–j”) are shown. Compared with the LacZ controls (a, c, e and i–i’’), RasV12/lgl−/−-induced tumor growth and invasion, and ptc>scrib-IR-triggered cell migration and MMP1 expression were notable blocked by expressing a sna RNAi (b, d, f and j–j’’). Statistical analysis of the invasion percentage (g), the pupariation percentage (h) and the migrating cell number (k) as shown in figures a–b (a, 72.58%, n = 124; b, 22.81%, n = 114), e–f (e, 2.56%, n = 39; f, 16.36%, n = 55) and i–j (n = 10 for each genotype; i, mean = 76.00; j, mean = 19.00) were shown. Chi-squared test or two tailed unpaired t-test was applied to compute P-values, *P < 0.05, ****P < 0.0001. See the electronic supplementary material for detailed genotypes. Scale bar: 100 µm (a–f), 20 µm (i–j).
Fluorescent images of larval CCs (a–e) and VNCs (f–j). RasV12 clones showed an obvious growth advantage (a, f), while overexpression of Sna alone did not promote any tumor-like phenotype (b, g). Coexpression of Sna and RasV12 triggered moderate tumor overgrowth and VNC invasion (c, h), which was totally inhibited by knockdown of CtBP (d, e, i and j). k Column bar graph of the invasion percentage as shown in figures a–e (a, 0.00%, n = 55; b, 0.00%, n = 36; c, 31.11%, n = 45; d, 0.00%, n = 43; e, 0.00%, n = 27). Chi-squared test was applied to compute P-values, **P < 0.01. See the electronic supplementary material for detailed genotypes. Scale bar: 100 µm (a–j).
CtBP is essential for Sna-mediated tumor invasion and EMT
To dissect the role of CtBP in Sna-mediated tumor invasion, we employed the RasV12/Sna tumor model, and found that RasV12/Sna-triggered tumor invasion was remarkably impeded by CtBP knockdown (Fig. 4d, e and i–k), suggesting that CtBP is essential for RasV12/Sna-triggered tumor invasion.
Epithelial–mesenchymal transition (EMT) is a crucial step toward tumor metastasis, which endows cells with the capacity to break through basement membranes, resolve out cell–cell junctions, and migrate away from their initial site55,56. In mammals, Sna family members are regarded as the major transcription factors governing EMT53. To check whether this EMT-promoting function is conserved by Drosophila Sna, we overexpressed Sna along the A/P compartment boundary by ptc-GAL4. Consistently, we observed a conspicuous expansion of the GFP stripe, mostly notable in the dorsal region (Fig. 5b, u), with some GFP-positive cells migrating toward anterior (Fig. 5g), accompanied by EMT hallmarks, including MMP1 elevation (Fig. 5l, v) and β-integrin accumulation (Fig. 5q). These Sna-induced EMT features were significantly suppressed by knockdown of CtBP (Fig. 5c, d, h, i, m, n, r and s). However, ectopic expression of CtBP failed to produce any EMT-like phenotype (Fig. 5e, j, o and t). Thus, we conclude that CtBP is necessary, but not sufficient, for Sna-mediated EMT processes.
(a–t) Fluorescence micrographs of 3rd-instar larval wing disks are shown. Anterior is to the left and dorsal up. The individual channels detecting only GFP (green, a–e and f–j), only MMP1 (red, k–o), and only β-integrin signal (red, p–t). f–j, k–o and p–t are high magnification of the yellow-dotted boxed areas in a–e. ptc-GAL4 UAS-GFP is the control (a, f, k and p). Ectopic expression of Sna-induced cell migration (g, yellow arrows indicate the migrated cells), MMP1 upregulation (l), and β-integrin accumulation (q) was impeded by RNAi-mediated inactivation of CtBP (h, i, m, n, r and s). While expression of CtBP alone did not produce visible defects during larval stage (e, j, o and t). Statistical analysis of migrating cell number (u) and MMP1 + cell number (v), error bars indicate standard deviation. One-way ANOVA with Bonferroni multiple-comparison test was used to compute P-values, ****P < 0.0001; ns, no significant difference. See the electronic supplementary material for detailed genotypes. Scale bar: 40 µm (a–t).
Sna promotes cell invasion independent of cell death
Sna is also known to regulate cell death, while the role of cell death in Sna-induced cell invasion has not been explored. To this end, we performed cDcp-1 staining and found that ptc > scrib-IR-triggered cell death was partially suppressed by depletion of sna (Supplementary Fig. 5a–c). In addition, ectopic expression of Sna was sufficient to induce apoptosis, mostly in the dorsal region (Supplementary Fig. 5d), where invasion was observed (Fig. 2b and g). However, blocking apoptosis by overexpressing P35 did not affect Sna-induced cell invasion and MMP1 activation (Supplementary Fig. 6a–a”, c–c”). To further distinguish Sna-triggered cell invasion from apoptosis-induced proliferation (AiP), we expressed DroncDN to interfere the function of endogenous Dronc, which plays a key role in AiP. Blocking AiP had no effects on Sna-induced cell invasion and MMP1 upregulation (Supplementary Fig. 6b–b”). Collectively, these results suggest that Sna-triggered cell invasion is independent of cell death.
Sna and CtBP regulate cell migration in normal development
In Drosophila, thorax closure is another remarkable model to study epithelial cell migration in development57. To investigate whether CtBP and Sna regulate cell migration in normal development, we knocked down either gene by the thorax-specific pannier (pnr)-GAL4 driver. Intriguingly, we found that sna depletion resulted in a mild cleft phenotype in the thorax (Fig. 6b), which was enhanced by heterozygosity for CtBP mutation (Fig. 6c). Likewise, RNAi-mediated downregulation of CtBP induced a thorax cleft phenotype (Fig. 6f), which was exacerbated in heterozygous sna mutants (Fig. 6g). On the other hand, heterozygosity for sna or CtBP (Fig. 6d, h), or knockdown of an unrelated gene dFoxO (Fig. 6e), gave no distinguishable phenotype. Collectively, these evidences suggest that Sna and CtBP may function together to regulate cell migration in thorax development.
Light micrographs of Drosophila adult thoraxes are shown. Compared with the control (a), depletion of sna (b) or CtBP (f) induced thorax cleft in development. Mutating one copy of endogenous CtBP (d) or sna (h) produced no obvious phenotype, but synergistically aggravated the pnr > sna-IR (c) or pnr > CtBP-IR- (g) triggered thorax closure defects, respectively. Knockdown of dFoxO showed no obvious defects, which was used as a negative control (e). See the electronic supplementary material for detailed genotypes. Scale bar: 100 µm (a–h).
Sna promotes JNK pathway activation
The JNK pathway plays crucial roles in regulating cell migration and tumor invasion8,39. To investigate the mechanism that underlies Sna-induced EMT, we checked the activity of JNK signaling. Compared with the ptc > GFP control (Fig. 7a–a”, d), ectopic expression of Sna dramatically enhanced the expression of a puc-LacZ reporter by executing an antibody-staining (Fig. 7b–b”) or X-gal staining assay (Fig. 7e), which was abolished by expressing a dominant negative form of the Drosophila JNK ortholog Bsk (BskDN, Fig. 7c–c”, f). TRE-RFP, which carries multiple binding sites for the AP-1 (Jun/Fos) transcription complex, is another reporter of JNK signaling58. Compared with the control (Fig. 7g–g”, i–i”), expressing Sna was sufficient to upregulate TRE-RFP expression (Fig. 7h–h”), and induce JNK phosphorylation detected by a specific anti-pJNK antibody (Fig. 7j–j”). Of note, the puc-LacZ reporter is a LacZ-bearing P-element inserted into the second intron of puc, and hence, acts as a loss-of-function allele (also known as pucE69). Intriguingly, ptc > Sna-induced cell migration (Fig. 7j’) was significantly enhanced by loss-of-puc (Fig. 7b’), but suppressed by BskDN, suggesting that Sna promotes JNK-dependent cell migration in Drosophila.
a–c and g–j Merged fluorescence micrographs of Drosophila third-instar larval wing disks are shown. The individual channels detecting only GFP (green, a’–c’ and g’–j’), β-gal (red, a”–c”), RFP (g”, h”), or pJNK signal (red, i”–j”). (d–f) Light micrographs showing X-gal staining of the puc-LacZ reporter in wing disks. Compared with the controls (a, d, g and i), ectopic expression of Sna elevated the expression of puc-LacZ (b, e), TRE-RFP (h), and JNK phosphorylation (j). The increased puc transcription triggered by ptc > Sna is largely impeded by expressing BskDN (c, f). See the electronic supplementary material for detailed genotype. Scale bar: 40 µm (a–f).
Conclusions
Most cancer-related deaths are caused by secondary tumors formed through invasion, a rather complex and poorly understood process. With the multiple genetic tools and conserved tumor invasion machinery, Drosophila has been widely considered as an outstanding model organism to explore the invasion program43,44,48. The CtBP protein is a well-characterized and evolutionarily conserved transcriptional corepressor that plays crucial roles during development and oncogenesis. In this work, we identified CtBP as a novel regulator of RasV12/lgl-/- induced tumor growth and invasion. Besides, CtBP is also required for loss-of-cell polarity-triggered cell invasion in the wing disks, and developmental cell migration in thorax closure. Mechanistically, CtBP may interact with Sna to form a transcriptional complex that activates the JNK signaling and promotes JNK-dependent cell migration and tumor invasion. Yet, the contribution of CtBP and JNK in Sna-induced EMT needs to be verified in human cancers, which may provide additional drug targets and therapeutic strategies for clinical treatment of malignant tumors.
Materials and methods
Fly strains
Flies were kept on a cornmeal and agar medium at 25 °C according to standard protocols unless indicated. For producing the fluorescently labeled invasive tumors in the eye disks, the following strains were previously described38,39,44, including yw ey-Flp; tub-GAL80 FRT40A; act > y+>GAL4 UAS-GFP (40 A tester), lgl4 FRT40A UAS-RasV12 (40 A tester), and ey-Flp act > y+>GAL4 UAS-GFP and UAS-RasV12. Additional Drosophila strains used, including UAS-CtBP-IR (32889 and 31334), sna18 (3299), and UAS-dFoxO-IR (27656), were obtained from Bloomington Drosophila stock center. UAS-sna-IRV (6263) and UAS-scrib-IR (27424) were received from Vienna Drosophila RNAi center (VDRC). ptc-GAL459, UAS-Puc, UAS-BskDN, puc-LacZ60, UAS-LacZ, UAS-GFP, UAS-Sna74b 54, TRE-RFP58, pnr-GAL4, UAS-DroncDN, and UAS-P35 were previously described42,43,47,61. UAS-CtBP and CtBP87De-10 were kind gifts from Professor Ming Fang62,63. For all fly cross-experiments, healthy unmated male and female parents were randomly assigned to different groups. Double-blinded method was employed during the experiments.
CtBP mutant clones were generated in 3rd instar larval eye disk by using the following strains: ey-Flp act > y+ > Gal4 UAS-GFP; FRT82B tub-Gal80 (82B MARCM tester) and FRT82B CtBP87De-10.
For ptc > GFP + scrib-IR cell migration experiments, animals were reared at 25 °C for 2 days, then shifted to 29 °C for additional 3 days, and the wing disks were dissected from 3rd-instar larvae44. For ptc > GFP + Sna migration assays, as ectopic expression of Sna is too strong to cause lethality before reaching the third-instar larva stage, animals were maintained at 18 °C.
qRT-PCR
For RNAi-knockdown efficiency experiments, hs-Gal4 driver was used. Animals were raised at 25 °C, heat-shocked at 37 °C for 30 min, and recovered at 29 °C for 2 h before dissection.
Total RNAs were isolated from third-instar larval eye disk, and qRT-PCR was performed as previously described64. rp49 served as an internal control.
Primers used are provided:
rp49-FP: TACAGGCCCAAGATCGTGAA.
rp49-RP: TCTCCTTGCGCTTCTTGGA.
CtBP-FP: GTCATCTTCTACGATCCCTACCT.
CtBP-RP: GCAATCGGACTGGAAAAGCA.
Immunostaining
Dissected disks were fixed in 4% formaldehyde for 20 min. After several washes with 0.3% (v/v) PBST, disks were stained with primary antibodies at 4 °C overnight and then with secondary antibodies at room temperature for 2 h. The following antibodies were used: rabbit anti-cDcp-1 (1:100, Cell Signaling Technology, CST, Cat. #9578), rabbit anti-Phospho-Histone H3 (1:400, CST, Cat. #9701), mouse anti-MMP1 (1:200, Developmental Studies Hybridoma Bank, DSHB, Cat. #3A6B4), mouse anti-β-integrin (1:100, DSHB, Cat. #CF.6G11), mouse anti-β-Gal (1:500, DSHB, Cat. #40-1a), rabbit anti-phospho-JNK (1:200, Calbiochem, Cat. #559309), goat anti-mouse-Cy3 (1:1000, Life technologies, Cat. #A10521), and goat anti-Rabbit-Cyanine3 (1:1000, Life technologies, Cat. #A10520). Vectashield mounting media (Vector Laboratories, Cat. #H-1500) with DAPI (4,6-diamidino-2-phenylindole) was used for mounting.
X-gal staining
Wing disks were dissected from 3rd-instar larvae in PBST (1×PBS, pH 7.0, 0.1% Triton X-100) and stained for β-galactosidase activity as described65.
Statistics
All data were collected from at least three independent experiments. The results were presented as bar graphs created with GraphPad Prism 8.0.2. For statistical significance, one-way ANOVA with Bonferroni’s multiple-comparison test, chi-squared test or two-tailed unpaired t-test was applied. P value less than 0.05 was considered significant and center values as the mean. Error bars indicated standard deviation. ns means not significant, P ≥ 0.05; * is P < 0.05; ** is P < 0.01; *** is P < 0.001; **** is P < 0.0001. P values are included in the relevant figure legends.
References
Zeeshan, R. & Mutahir, Z. Cancer metastasis—tricks of the trade. Bosn. J. Basic Med. Sci. 17, 172–182 (2017).
Seyfried, T. N. & Huysentruyt, L. C. On the origin of cancer metastasis. Crit. Rev. Oncog. 18, 43–73 (2013).
Suhail, Y. et al. Systems biology of cancer metastasis. Cell Syst. 9, 109–127 (2019).
Fischer, K. R. et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 527, 472–476 (2015).
Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 16, 582–598 (2016).
Massague, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Ma, X. et al. Hippo signaling promotes JNK-dependent cell migration. Proc. Natl Acad. Sci. USA 114, 1934–1939 (2017).
Gerlach, S. U., Eichenlaub, T. & Herranz, H. Yorkie and JNK control tumorigenesis in Drosophila cells with cytokinesis failure. Cell Rep. 23, 1491–1503 (2018).
Anastas, J. N. & Moon, R. T. WNT signalling pathways as therapeutic targets in cancer. Nat. Rev. Cancer 13, 11–26 (2013).
Dcona, M. M., Morris, B. L., Ellis, K. C. & Grossman, S. R. CtBP—an emerging oncogene and novel small molecule drug target: advances in the understanding of its oncogenic action and identification of therapeutic inhibitors. Cancer Biol. Ther. 18, 379–391 (2017).
Byun, J. S. & Gardner, K. C-Terminal binding protein: a molecular link between metabolic imbalance and epigenetic regulation in breast cancer. Int. J. Cell Biol. 2013, 647975 (2013).
Zheng, X., Song, T., Dou, C., Jia, Y. & Liu, Q. CtBP2 is an independent prognostic marker that promotes GLI1 induced epithelial-mesenchymal transition in hepatocellular carcinoma. Oncotarget 6, 3752–3769 (2015).
Boyd, J. M. et al. A region in the C-terminus of adenovirus 2/5 E1a protein is required for association with a cellular phosphoprotein and important for the negative modulation of T24-ras mediated transformation, tumorigenesis and metastasis. EMBO J. 12, 469–478 (1993).
Chinnadurai, G. CtBP, an unconventional transcriptional corepressor in development and oncogenesis. Mol. Cell 9, 213–224 (2002).
Grooteclaes, M. L. & Frisch, S. M. Evidence for a function of CtBP in epithelial gene regulation and anoikis. Oncogene 19, 3823–3828 (2000).
van Vliet, J., Turner, J. & Crossley, M. Human Kruppel-like factor 8: a CACCC-box binding protein that associates with CtBP and represses transcription. Nucleic Acids Res. 28, 1955–1962 (2000).
Hildebrand, J. D. & Soriano, P. Overlapping and unique roles for C-terminal binding protein 1 (CtBP1) and CtBP2 during mouse development. Mol. Cell. Biol. 22, 5296–5307 (2002).
Reid, A. et al. The transcriptional repressor CTBP-1 functions in the nervous system of Caenorhabditis elegans to regulate lifespan. Exp. Gerontol. 60, 153–165 (2014).
Stankiewicz, T. R., Gray, J. J., Winter, A. N. & Linseman, D. A. C-terminal binding proteins: central players in development and disease. Biomol. Concepts 5, 489–511 (2014).
Saijo, K., Collier, J. G., Li, A. C., Katzenellenbogen, J. A. & Glass, C. K. An ADIOL-ERbeta-CtBP transrepression pathway negatively regulates microglia-mediated inflammation. Cell 145, 584–595 (2011).
Chinnadurai, G. The transcriptional corepressor CtBP: a foe of multiple tumor suppressors. Cancer Res. 69, 731–734 (2009).
Di, L. J. et al. Genome-wide profiles of CtBP link metabolism with genome stability and epithelial reprogramming in breast cancer. Nat. Commun. 4, 1449 (2013).
Grau, Y., Carteret, C. & Simpson, P. Mutations and chromosomal rearrangements affecting the expression of snail, a gene involved in embryonic patterning in DROSOPHILA MELANOGASTER. Genetics 108, 347–360 (1984).
Mauhin, V., Lutz, Y., Dennefeld, C. & Alberga, A. Definition of the DNA-binding site repertoire for the Drosophila transcription factor SNAIL. Nucleic Acids Res. 21, 3951–3957 (1993).
Chiang, C. & Ayyanathan, K. Snail/Gfi-1 (SNAG) family zinc finger proteins in transcription regulation, chromatin dynamics, cell signaling, development, and disease. Cytokine Growth Factor Rev. 24, 123–131 (2013).
Hemavathy, K., Hu, X., Ashraf, S. I., Small, S. J. & Ip, Y. T. The repressor function of snail is required for Drosophila gastrulation and is not replaceable by Escargot or Worniu. Dev. Biol. 269, 411–420 (2004).
Rembold, M. et al. A conserved role for Snail as a potentiator of active transcription. Genes Dev. 28, 167–181 (2014).
Ip, Y. T., Levine, M. & Bier, E. Neurogenic expression of snail is controlled by separable CNS and PNS promoter elements. Development 120, 199–207 (1994).
Ganguly, A., Jiang, J. & Ip, Y. T. Drosophila WntD is a target and an inhibitor of the Dorsal/Twist/Snail network in the gastrulating embryo. Development 132, 3419–3429 (2005).
Arefin, B., Parvin, F., Bahrampour, S., Stadler, C. B. & Thor, S. Drosophila neuroblast selection is gated by Notch, Snail, SoxB, and EMT gene interplay. Cell Rep. 29, 3636–3651 (2019).
Tang, Y., Feinberg, T., Keller, E. T., Li, X. Y. & Weiss, S. J. Snail/Slug binding interactions with YAP/TAZ control skeletal stem cell self-renewal and differentiation. Nat. Cell Biol. 18, 917–929 (2016).
Kaufhold, S. & Bonavida, B. Central role of Snail1 in the regulation of EMT and resistance in cancer: a target for therapeutic intervention. J. Exp. Clin. Cancer Res. 33, 62 (2014).
Yook, J. I. et al. A Wnt-Axin2-GSK3beta cascade regulates Snail1 activity in breast cancer cells. Nat. Cell Biol. 8, 1398–1406 (2006).
Muqbil, I., Wu, J., Aboukameel, A., Mohammad, R. M. & Azmi, A. S. Snail nuclear transport: the gateways regulating epithelial-to-mesenchymal transition? Semin. Cancer Biol. 27, 39–45 (2014).
Peinado, H., Ballestar, E., Esteller, M. & Cano, A. Snail mediates E-cadherin repression by the recruitment of the Sin3A/histone deacetylase 1 (HDAC1)/HDAC2 complex. Mol. Cell. Biol. 24, 306–319 (2004).
Qiao, B., Johnson, N. W. & Gao, J. Epithelial-mesenchymal transition in oral squamous cell carcinoma triggered by transforming growth factor-beta1 is Snail family-dependent and correlates with matrix metalloproteinase-2 and -9 expressions. Int J. Oncol. 37, 663–668 (2010).
Pagliarini, R. A. & Xu, T. A genetic screen in Drosophila for metastatic behavior. Science 302, 1227–1231 (2003).
Ma, X. et al. Myc suppresses tumor invasion and cell migration by inhibiting JNK signaling. Oncogene 36, 3159–3167 (2017).
Igaki, T., Pagliarini, R. A. & Xu, T. Loss of cell polarity drives tumor growth and invasion through JNK activation in Drosophila. Curr. Biol. 16, 1139–1146 (2006).
Martin-Blanco, E. et al. puckered encodes a phosphatase that mediates a feedback loop regulating JNK activity during dorsal closure in Drosophila. Genes Dev. 12, 557–570 (1998).
Ma, X. et al. Src42A modulates tumor invasion and cell death via Ben/dUev1a-mediated JNK activation in Drosophila. Cell Death Dis. 4, e864 (2013).
Ma, X. et al. dUev1a modulates TNF-JNK mediated tumor progression and cell death in Drosophila. Dev. Biol. 380, 211–221 (2013).
Zhang, S. et al. Wingless modulates activator protein-1-mediated tumor invasion. Oncogene 38, 3871–3885 (2019).
Guo, X. et al. Rox8 promotes microRNA-dependent yki messenger RNA decay. Proc. Natl Acad. Sci. USA 117, 30520–30530 (2020).
Worley, M. I., Alexander, L. A. & Hariharan, I. K. CtBP impedes JNK- and Upd/STAT-driven cell fate misspecifications in regenerating Drosophila imaginal discs. eLife 7, e30391 (2018).
Ma, X. et al. Bendless modulates JNK-mediated cell death and migration in Drosophila. Cell Death Differ. 21, 407–415 (2014).
Ma, X. et al. Rho1-Wnd signaling regulates loss-of-cell polarity-induced cell invasion in Drosophila. Oncogene 35, 846–855 (2016).
Uhlirova, M. & Bohmann, D. JNK- and Fos-regulated Mmp1 expression cooperates with Ras to induce invasive tumors in Drosophila. EMBO J. 25, 5294–5304 (2006).
Srivastava, A., Pastor-Pareja, J. C., Igaki, T., Pagliarini, R. & Xu, T. Basement membrane remodeling is essential for Drosophila disc eversion and tumor invasion. Proc. Natl Acad. Sci. USA 104, 2721–2726 (2007).
Nibu, Y., Zhang, H. & Levine, M. Interaction of short-range repressors with Drosophila CtBP in the embryo. Science 280, 101–104 (1998).
Nibu, Y. et al. dCtBP mediates transcriptional repression by Knirps, Kruppel and Snail in the Drosophila embryo. EMBO J. 17, 7009–7020 (1998).
Wang, Y., Shi, J., Chai, K., Ying, X. & Zhou, B. P. The role of Snail in EMT and tumorigenesis. Curr. Cancer Drug Targets 13, 963–972 (2013).
Wu, C. et al. Snail modulates JNK-mediated cell death in Drosophila. Cell Death Dis. 10, 893 (2019).
Kudo-Saito, C., Shirako, H., Takeuchi, T. & Kawakami, Y. Cancer metastasis is accelerated through immunosuppression during Snail-induced EMT of cancer cells. Cancer Cell 15, 195–206 (2009).
Jung, H. Y., Fattet, L. & Yang, J. Molecular pathways: linking tumor microenvironment to epithelial-mesenchymal transition in metastasis. Clin. Cancer Res. 21, 962–968 (2015).
Zeitlinger, J. & Bohmann, D. Thorax closure in Drosophila: involvement of Fos and the JNK pathway. Development 126, 3947–3956 (1999).
Chatterjee, N. & Bohmann, D. A versatile PhiC31 based reporter system for measuring AP-1 and Nrf2 signaling in Drosophila and in tissue culture. PloS ONE 7, e34063 (2012).
Wang, X. et al. FoxO mediates APP-induced AICD-dependent cell death. Cell Death Dis. 5, e1233 (2014).
Xue, L. et al. Tumor suppressor CYLD regulates JNK-induced cell death in Drosophila. Dev. Cell 13, 446–454 (2007).
Wu, C. et al. Toll pathway modulates TNF-induced JNK-dependent cell death in Drosophila. Open Biol. 5, 140171 (2015).
Fang, M. et al. C-terminal-binding protein directly activates and represses Wnt transcriptional targets in Drosophila. EMBO J. 25, 2735–2745 (2006).
Bi, C. et al. CtBP represses Dpp signaling as a dimer. Biochem. Biophys. Res. Commun. 495, 1980–1985 (2018).
Wang, M. C., Bohmann, D. & Jasper, H. JNK signaling confers tolerance to oxidative stress and extends lifespan in Drosophila. Dev. Cell 5, 811–816 (2003).
Xue, L. & Noll, M. Drosophila female sexual behavior induced by sterile males showing copulation complementation. Proc. Natl Acad. Sci. USA 97, 3272–3275 (2000).
Acknowledgements
We thank Bloomington Drosophila center, VDRC Drosophila center and Professor Ming Fang from Southeast University for fly stocks, and members of Xue Lab for discussion and critical comments. This work was supported by National Natural Science Foundation of China (31771595, 31970536) and Shanghai Committee of Science and Technology (09DZ2260100) to L. X., the Fundamental Research Funds for the Central Universities (22120180549) to W. L., and National Natural Science Foundation of China (31701244), Natural Science Fund of Hebei Province of China (C2021209013) and Science and Technology Project of Hebei Education Department (BJ2019040) to C. W.
Author information
Authors and Affiliations
Contributions
C.W., X.D., W.L. and L.X. conceived and designed the experiments. C.W., X.D., Z.L., Y.H., R.Z., and M.Z. conducted experiments. Q.X., H.C., C.J., X.L., G.L., and W.L. supervised/advised on the study. C.W., X.D., W.L., and L.X. analyzed the data and wrote the paper. All authors gave final approval for publication.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Rufini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, C., Ding, X., Li, Z. et al. CtBP modulates Snail-mediated tumor invasion in Drosophila. Cell Death Discov. 7, 202 (2021). https://doi.org/10.1038/s41420-021-00516-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-021-00516-x
This article is cited by
-
Slik maintains tissue homeostasis by preventing JNK-mediated apoptosis
Cell Division (2023)
-
dGLYAT modulates Gadd45-mediated JNK activation and cell invasion
Cell Division (2022)