Abstract
Neurodegenerative diseases, a subset of age-driven diseases, have been known to exhibit increased oxidative stress. The resultant increase in reactive oxygen species (ROS) has long been viewed as a detrimental byproduct of many cellular processes. Despite this, therapeutic approaches using antioxidants were deemed unsuccessful in circumventing neurodegenerative diseases. In recent times, it is widely accepted that these toxic by-products could act as secondary messengers, such as hydrogen peroxide (H2O2), to drive important signaling pathways. Notably, mitochondria are considered one of the major producers of ROS, especially in the production of mitochondrial H2O2. As a secondary messenger, cellular H2O2 can initiate redox signaling through oxidative post-translational modifications (oxPTMs) on the thiol group of the amino acid cysteine. With the current consensus that cellular ROS could drive important biological signaling pathways through redox signaling, researchers have started to investigate the role of cellular ROS in the pathogenesis of neurodegenerative diseases. Moreover, mitochondrial dysfunction has been linked to various neurodegenerative diseases, and recent studies have started to focus on the implications of mitochondrial ROS from dysfunctional mitochondria on the dysregulation of redox signaling. Henceforth, in this review, we will focus our attention on the redox signaling of mitochondrial ROS, particularly on mitochondrial H2O2, and its potential implications with neurodegenerative diseases.
Similar content being viewed by others
Facts
-
Mitochondria are one of the major producers of ROS in the cellular environment.
-
Mitochondria possess an extensive antioxidant defense to maintain redox homeostasis.
-
ROS, such as hydrogen peroxide, are increasingly viewed as important redox signaling molecules.
-
Mitochondrial dysfunction is implicated in many neurodegenerative diseases.
-
Increasing oxidative stress through aging is associated with many neurodegenerative diseases.
Open questions
-
Do the mitochondria produce ROS, such as hydrogen peroxide, to mediate cellular redox signaling?
-
Is there a central role for mitochondria in the cellular redox signaling cascade?
-
Does mitochondrial dysfunction disrupt the cellular redox signaling landscape?
-
Is the disruption to the cellular redox signaling landscape that is mediated by mitochondrial dysfunction a trigger of neurodegenerative diseases, or is it a secondary effect from the manifestation of neurodegenerative diseases?
Introduction
Growing evidence from various studies highlights the importance of mitochondria in disease manifestation, especially in aging and neurodegenerative diseases1,2. A long-standing viewpoint is the contribution of mitochondria to oxidative stress during aging: the decline of mitochondrial function leads to redox imbalance, which results in an increased production of reactive oxygen species (ROS) and the loss of cellular antioxidant defense; the subsequent overload of ROS induces mitochondrial dysfunction and triggers the apoptotic cascade. This scenario has been well documented in various neurodegenerative diseases3. Moreover, ROS dysregulation in microglia (resident brain macrophages that protect against brain damage) strongly correlates to increased oxidative stress and, consequently, neuronal death4,5. However, ROS do more than inducing cell death, they also contribute to cellular signaling.
ROS are chemical entities possessing a radical nature; they trigger chemical reactions that could lead to deleterious biochemical changes to the cellular environment6. Detailed documentation of the radical processes triggered by ROS has been well-reviewed elsewhere7. Recently, the effects of ROS have been shown to extend beyond their destructive nature8, as various species of ROS target cellular signaling by acting as important secondary messengers9. Notably, a review by Sies highlights superoxide (O2•−) and hydrogen peroxide (H2O2) as secondary messengers in various signaling pathways10, though H2O2 is considered more important owing to its higher stability and diffusive nature.
Evidently, ROS, particularly mitochondrial ROS (mtROS), contribute to neurodegenerative disease onset and progression via both redox imbalance and cellular signaling. Thus, this review will present a more detailed and up-to-date summary of mtROS-mediated signaling pathways. Specifically, we will feature how H2O2, as a secondary messenger, can induce cysteine-centered oxidative post-translational modifications (oxPTMs); these protein oxPTMs drive downstream signaling pathways and affect cellular processes. In addition, a summary of known cellular signaling pathways that are targeted by mtH2O2 will be discussed, in the context of neurodegenerative diseases.
Intracellular sources of hydrogen peroxide production and their regulation
The intracellular balance of ROS is paramount for cell survival and is maintained by reducing excess cellular ROS via antioxidants11. There are two types of ROS sources within the cell: first, ROS are produced from intracellular biological processes such as OXPHOS and protein disulfide bridge formation12. Second, ROS can be produced by external cues such as xenobiotics, microbial invasion, and immune system-derived cytokines13; here, ROS production acts to break down foreign entities or induce a downstream signaling pathway. Hence, despite their near-constant production, ROS must be maintained at safe levels.
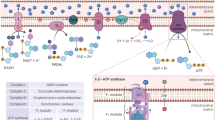
The three main producers of ROS are mitochondria, along with the endoplasmic reticulum (ER), and peroxisome (Fig. 1). In the mitochondrial electron transport chain (ETC), mtROS are produced from the one-electron transfer from O2 to the respective electron donors and acceptors, thus generating mtO2•− 14. Owing to mtO2•− instability, the presence of nearby antioxidant enzymes—superoxide dismutase 1 (SOD1) and 2 (SOD2)—convert mtO2•− to the more stable mtH2O215. mtH2O2 can be further reduced to H2O by other mitochondrial antioxidant enzymes, such as catalase, glutathione peroxidases (GPXs; isoform 1 and 4), and peroxiredoxins (PRXs; isoform 3 and 5)16. Alternatively, mtH2O2 may exit the mitochondria via inner mitochondrial membrane (IMM) channels, such as aquaporins, and diffuse through the porous outer mitochondrial membrane (OMM) into the cytoplasm for potential redox signaling17,18,19.
A In the ER, the major source of ER-ROS production is from the process of oxidative protein folding for disulfide bond formation. This process is governed by the enzyme ER oxidoreductin 1 (ERO1). NADPH oxidase 4 (NOX4) is also involved in the production of ER-ROS, mainly ER-H2O2. B In the peroxisomes, the major producer of peroxisomal ROS is the process of beta-oxidation of fatty acids. The main player involved is the enzyme, Acyl-CoA oxidase (ACOX). Numerous other enzymes in the peroxisomes also contribute to the production of peroxisomal ROS: xanthine oxidase (XO); d-amino-acid oxidase (DAO); d-aspartate oxidase (DDO); l-pipecolic acid oxidase (PIPOX); l-α-hydroxyacid oxidase (HAO); polyamine oxidase (PAOX). C The main producers of mitochondrial ROS in the mitochondria are from the activity of the electron transport chain (ETC) during oxidative phosphorylation (OXPHOS). The mitochondrial superoxide (O2•−) produced by the ETC is immediately converted to mitochondrial hydrogen peroxide (H2O2) by superoxide dismutase 1 and 2 (SOD1 and SOD2). In general, cellular H2O2 is one of the major and common forms of cellular ROS. Its role as a secondary messenger and its production in these three organelles has raised a suggestion of a potential redox signaling complex among them, highlighting the importance of cellular H2O2 in cellular redox communications. Created with BioRender.com.
ER-ROS are produced mainly during protein synthesis12 and contribute the greatest to cellular ROS levels20. ER-ROS, predominantly as ER-H2O2, is produced by ER oxidoreductin 1 (ERO1, with α and β subunits) during the disulfide bond formation in oxidative protein folding21. In addition, ER-H2O2 is produced by NADPH oxidase 4 (NOX4), member of the NOX family of membrane-bound NADPH oxidases. NOX4 is the only NOX isoform to directly produce H2O2, and it has been documented to have many functions, ranging from mediating O2 sensing to cell proliferation and differentiation22,23. Besides the ER, NOX4 localizes to the nucleus, plasma membrane, and mitochondria24,25,26, though its multiple cellular localization is a source of debate27,28. In addition, a detailed review by Chen et al.27 describes the multiple roles of NOX4 and highlights H2O2 as an important secondary messenger. Excessive ER-H2O2 in the ER is prevented by the activities of various antioxidant enzymes, such as GPXs (isoform 7 and 8) and PRX429.
Peroxisomal ROS, mainly H2O2, is produced by numerous oxidative enzymes (Fig. 1). For example, a review by Antonenkov et al.30 presents fatty acid beta-oxidation by acyl-CoA oxidases as one of the major sources of peroxisomal H2O2. Akin to the mitochondria and ER, peroxisomes contain antioxidant enzymes such as catalase, SOD1 and PRX5, to maintain the balance of cellular oxidative levels30.
The presence of multiple cellular H2O2 sources and antioxidant enzymes demonstrates that cellular H2O2 functions beyond toxicity; it is an important secondary messenger that modulates and drives vital downstream signaling pathways, as emphasized in a review by Yoboue et al.31. The next section of this review will underscore how cellular H2O2 can influence signaling pathways by modifying the reactive thiol side chains of the amino acid cysteine.
Hydrogen peroxide-driven oxPTMs
The complexity of the cellular proteome arises from the presence of post-translational modifications (PTMs), of which numerous types exist, including the well-studied phosphorylation32. Commonly, these PTMs modulate protein activity by inducing conformational changes. They may stabilize the protein to avoid degradation or induce the opposite effect. Growing evidence has shown cellular H2O2 to be a key secondary messenger that can elicit oxPTMs and affect biological processes like cell proliferation33.
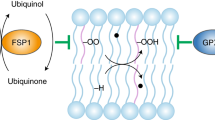
The reactive state of ROS can induce chemical modification, altering protein conformation and activity, as observed in other PTMs34. Moreover, like phosphorylation, oxPTMs are reversible, demonstrating the potential influence that H2O2 has on the cellular signaling network35. Cellular H2O2 targets mainly cysteine residues due to their thiol side chains that can be oxidized36. Cellular H2O2 and ROS, along with cellular reactive molecules such as reactive nitrogen species and nitric oxide, can cause the cysteine amino acid to adopt different oxidative states, including S-OH, S-O2H, S-O3H, and S-NO37. Fig. 2 summarizes the complete types of cysteine oxidative state, and the processes that lead to their formation. Such diversity highlights the variation of redox signaling that exists in the cell.
In the presence of cellular hydrogen peroxide (H2O2), the free thiol will undergo sulfenylation to form sulfenic acid, which in recent times has been considered an important intermediate in redox signaling. In the presence of increasing oxidative stress, the elevated levels of cellular H2O2 would lead to the sulfenic acid undergoing sulfinylation to form irreversible sulfinic acid. However, the presence of SRXN1 could convert the sulfinic acid back to sulfenic acid. With even higher concentrations of cellular H2O2, sulfinic acid can undergo further reaction via sulfonylation to form sulfonic acid, which is the final irreversible form of thiol. This highlights the detrimental effects of increasing oxidative stress in cells, which could lead to formation of the irreversible sulfonic acid and the loss of function of important proteins in the cell. Another important thiol oxPTM is the formation of disulfide bonds, both intermolecularly and intramolecularly. The formation of the disulfide bonds is known to potentially activate as well as inhibit the function of the target proteins. Another notable oxPTM is the S-glutathionylation of proteins in the presence of glutathione molecules. Other modifications of free thiol include the sulfhydration by H2S, which is an important modification in the central nervous system, and S-nitrosylation that is induced by reaction nitrogen species in the cell. Created with BioRender.com.
Cysteine oxPTMs include disulfide bonds, where both intermolecular and intramolecular bond formations can occur. Disulfide bonds can occur between thiyl radicals (RS•) of two closely placed independent free thiols on the same or separate protein molecule38. Alternatively, ROS may convert S-OH groups to RS•, thereafter forming disulfide bonds with other thiolates39. Disulfide bond formation could lead to changes to protein conformation and function40; for instance, the α-subunit of ATP synthase can form a disulfide bridge with the γ-subunit between Cys294 and Cys103 to inhibit ATP production41.
Another form of oxPTM is sulfenylation (S-OH), which was previously classified as a toxic byproduct from cellular H2O2 and O2•− reactions; however, researchers in the last decade have identified it as an important intermediate in redox signaling pathways39. The reactive and unstable nature of S-OH enables its conversion into other types of oxPTMs in the presence of reduced glutathione (GSH). S-glutathionylation, as reviewed by Zhang et al.42, is a reversible process that depends on the presence of cellular antioxidants; it thus protects the target protein by preventing irreversible oxPTMs39. In addition, S-OH can react with free thiols of a target protein to form disulfide bonds. Such plasticity in protein modification demonstrates how oxPTMs can help drive redox signaling.
In extremely high levels of H2O2, oxPTMs can become an irreversible process, as the oxidative state of cysteine shifts beyond sulfenylation to sulfinylation and eventually, to sulfonylation (Fig. 2). Although sulfonylation is classified as irreversible, it is possible to reverse sulfinylation via sulfiredoxin-1; this process forms the reversible sulfenic acid form of cysteine43,44. The progression of cysteine to its irreversible form illustrates the importance of cellular ROS levels: Low levels of cellular ROS would initiate oxPTMs, whereas excessive levels of cellular ROS would irreversibly oxidize proteins and cause deleterious consequences. A review by Chung et al.45 delineates the capability of cysteine-based oxPTMs in redox signaling and physiological processes, particularly in the highly oxidized cardiovascular environment.
The ability of cellular H2O2 to induce oxPTMs is a key component in redox signaling33, especially when cysteine residues form only 2% of the cellular proteome but is involved in the most PTMs46. Hence, understanding how H2O2 influences the biological signaling pathways in various diseases facilitates potential therapeutic interventions, particularly in neurodegenerative diseases, due to their close association to increased oxidative stress. In the next section of this review, we will highlight biological mechanisms that can be influenced by cellular H2O2, especially those associated with the mitochondria.
Mitochondria-associated biological processes regulated by potential mitochondrial hydrogen peroxide-mediated redox signaling
Besides producing ATP during OXPHOS, mitochondria also play an important role in triggering and regulating apoptosis, as documented by Orrenius et al.47. Lower levels of ROS were shown to activate cellular survival responses, while higher levels of ROS activate death processes48. In the intrinsic apoptotic pathway, mitochondrial outer membrane permeabilization—a process regulated by the B-cell lymphoma-2 (BCL-2) protein family, such as the Bcl-2-associated X protein (BAX)—releases numerous proapoptotic proteins from the mitochondrial intermembrane space, including cytochrome c (Cyt c)49. Cyt c can trigger rapid oligomerization of apoptotic protease-activating factor-1, which recruits caspase-9 into the apoptosome to cleave and activate downstream effectors like caspase-350. Moreover, Cyt c release triggers the production of mtROS, suggesting a role of mtROS in redox signaling51. H2O2-mediated oxPTM of BAX can induce protein disulfide dimerization and translocation to the OMM52; this was seen in human colon adenocarcinoma cells upon Cys62 modification of BAX by H2O253. In addition, apoptotic caspase activity can be redox-regulated by oxPTM, with procaspase-9, procaspase-3, and caspase-3 susceptible to S-glutathiolation54. As detailed by Benhar55, ROS such as H2O2 can contribute to oxPTM of proteins involved in the apoptotic pathway. However, whether these events are regulated by mtH2O2 remain unknown. Neurodegenerative diseases, such as Alzheimer’s disease (AD), are associated with the accumulation of misfolded proteins, like β-amyloids, that can cause neuronal death via oxidative stress56. However, it remains debatable whether ROS mainly causes cell death or could contribute to redox signaling of important survival pathways.
The mitochondrial enzyme pyruvate dehydrogenase (PDH) catalyzes pyruvate to enable its entry into the Krebs cycle; however, PDH is also a major contributor of mtH2O2 production in various tissues, approximately four times more than Complex I of the ETC57, thus suggesting a role of PDH in redox signal modulation. A recent study done by O’Brien et al.58 demonstrated that as mtH2O2 increases, so does oxidation of GSH to glutathione (GSSG). The increase of GSSG leads to PDH S-glutathionylation, causing a reduction in mtH2O2/O2•− production (Fig. 3E). This observation suggests that PDH can control the changing mitochondrial redox status and regulate the emission of mtH2O2. Interestingly, when reverse electron transfer occurs in the ETC under stress conditions, S-glutathionylation led to an increased production of mtH2O2/O2•− by PDH, potentially leading to mitochondrial dysfunction and cellular oxidative stress. This bi-directional effect of PDH S-glutathionylation on mtROS production may have implications in oxidative stressed-related diseases such as neurodegenerative diseases.
A Calcium (Ca2+) signaling communicated between the endoplasmic reticulum (ER) and the mitochondria (Mito) via the inositol 1,4,5-trisphosphate receptor (IP3R), sarco-/endoplasmic reticulum Ca2+-ATPase (SERCA) Ca2+ channels, and the voltage-dependent anion channel (VDAC); MAM: Mitochondria-associated membrane. B Ca2+ uptake by mitochondrial calcium uniport (MCU) after S-glutathionylation of Cys97. C mtH2O2 efflux from mitochondrial matrix swelling and reduced intermembrane space. D Cell death induced by the opening of the mitochondrial permeability transition pore (mPTP) via the tetramerization of p66Shc; PKCβ: Protein Kinase C β; Pin1: Prolyl Isomerase 1. E Increasing oxidized glutathione (GSSG) to reduced glutathione (GSH) ratio could reduce mtROS production by pyruvate dehydrogenase (PDH) and reduce the release of mtH2O2. F Parkin-dependent mitophagy mediated by mtROS; OMM: Outer mitochondrial membrane. G Increased levels of cellular ROS have been known to cause Parkin sulfinylation and inhibition by intracellular H2O2. H Increased heat production by sulfenylated uncoupler protein 1 (UCP1) during hypothermia. I Activation of mitofusion from the disulfide bond formation of mitofusin 1 (MFN1) and mitofusin 2 (MFN2). This disulfide bond formation could further S-glutathionate MNF2 to activate mitofusion. Created with BioRender.com.
Another study of mtH2O2-mediated biological process occurred in brown adipose tissue (BAT) by Chouchani et al.59. The authors found that cold-induced BAT thermogenesis was triggered by an increase in mtROS. Conversely, mtROS diminution in BAT using antioxidants mitoQ and N-acetylcysteine (NAC) led to hypothermia during cold exposure. The authors identified Cys253 sulfenylation on uncoupling protein 1 (UCP1), which is part of the mitochondrial respiratory system (Fig. 3H); this oxPTM sensitizes UCP1 to adrenergic activation in BAT, increasing UCP1-mediated thermogenic respiration and ATP expenditure, which then stimulates increased heat production. Contrarily, the scavenging of mtROS during cold exposure abrogated UCP1-mediated thermogenesis. This study affirms the significance of mtROS level in redox signaling. Although the authors did not mention mtH2O2 in the oxPTM of UCP1, the accepted role of H2O2 in sulfenylation implies its involvement here60. These examples highlight the influence of mtH2O2 as a secondary messenger in cellular pathways.
The development of neurodegenerative diseases may stem from mitochondrial loss of function mediated by excessive production of ROS; however, antioxidant treatment to combat neurodegenerative diseases received disappointing results, potentially due to the loss of these mtH2O2-driven biological processes required for cellular recovery61. Moreover, studies have also shown that increased mtROS did not negatively impact longevity, suggesting an important role of mtROS in age-related diseases such as neurodegenerative diseases62.
Cellular survival necessitates mitochondrial quality control (MQC); mitochondrial dysfunction can lead to cell death upon the accumulation of many apoptotic mediators in the mitochondria, as suggested in AD63. Hence, to safeguard mitochondrial integrity, mitochondrial dynamics (fission and fusion) exist, and their dysregulation could lead to neurodegenerative diseases64. Through mitophagy, dysfunctional mitochondria can be removed from the cell. Interestingly, there is evidence showing the involvement of mtROS in the process of mitochondrial fusion (mitofusion). A study conducted by Thaher et al.65 identified mitofusin 2 (MFN2) to undergo oxPTM and form disulfide bonds with adjacent MFN1 to induce mitofusion (Fig. 3I). The authors found that the Cys684 residue in MFN2 is key in this redox-regulated mitofusion and suggested that S-glutathionylation of Cys684 could also play a role in MFN2-mediated MQC in response to the changes in the intracellular redox environment. Accordingly, Mailloux and Treberg66 proposed the existence of a mitochondrial metabolism-linked redox signal that is indirectly mediated by mtH2O2 via the glutathione redox buffering system. Moreover, our lab recently demonstrated that mtROS could induce Parkin/PINK1-dependent mitophagy during mitochondrial stress67. We demonstrated that the increased mtROS from VDAC1 overexpression led to Parkin recruitment to the mitochondria (Fig. 3F), and this effect was abrogated by NAC and catalase. Although we did not further probe for oxPTMs, our results suggest that Parkin/PINK1-mediated mitophagy could be redox-sensitive, and there could be some form of redox signaling involved in the mitochondrial turnover mechanism. Interestingly, the cysteine-rich regions of Parkin can be sulfonylated by cellular H2O2, leading to the loss of its E3 ligase activity (Fig. 3G), which may be a contributing factor in Lewy bodies formation in Parkinson’s disease (PD)68. As such, these studies show that mitochondrial redox signaling might play a role in MQC, whose dysfunction has been implicated in the pathogenesis of PD.
Redox communication between the ER and the mitochondria
Apart from the redox signaling system, mitochondria also mediate calcium (Ca2+) signaling, which is involved in many diverse physiological processes such as muscle contraction and neuronal excitation69. Ca2+ imbalance can trigger cell death via apoptosis70. The exchange of Ca2+ between the ER and mitochondria occurs at the mitochondria-associated membrane (MAM) (Fig. 3A)71. A study by Booth et al.72 showed that the presence of ER-H2O2 at the MAM can lead to the inactivation of sarco-/ER Ca2+-ATPase (SERCA) and activation of inositol 1,4,5-trisphosphate receptor (IP3R). These events induce an influx of Ca2+ into the mitochondria, thereafter causing an influx of K+ and H2O into the mitochondrial matrix (MM). Consequently, MM swelling occurs (Fig. 3C), reducing the volume of the mitochondrial intermembrane space (IMS) and releasing mtH2O2 into the interface region between the MAM and the mitochondria. mtH2O2-containing redox nanodomains are formed; however, their function is not known, according to the authors. Additionally, mitochondrial Ca2+ uniporter (MCU) can be S-glutathionylated at Cys97 by mtH2O2 to increase Ca2+ uptake (Fig. 3B). Thus, these examples highlight the potential redox signal exchange between the ER-MAM and the mitochondria.
Despite being largely localized in the cytoplasm, recent studies have identified p66Shc to be associated to the MAM73. Oxidative stress induces p66Shc phosphorylation by protein kinase C β (PKCβ), triggering p66Shc interaction with prolyl isomerase 1 (Pin1) and subsequent mitochondrial translocation (Fig. 3D)74. A study by Gertz et al.75 highlighted the presence of a dimer-tetramer shift of p66Shc, where tetrameric p66Shc could trigger apoptosis by generating mtH2O2 and opening the mitochondrial permeability transition pore (mPTP). Gertz and Steegborn76 suggested that tetramerization increases p66Shc affinity to mPTP, and the local production of mtH2O2 opens mPTP to trigger apoptosis. They also remarked that tetrameric p66Shc could, via an unknown process, increase mtH2O2 levels enough to trigger mPTP opening and apoptosis. Therefore, a possible question is whether a mechanism to control the amount of mtH2O2 produced by the p66Shc tetramer exists. Indeed, Gertz et al.75 demonstrated that thioredoxin and glutathione can abrogate the apoptosis triggered by tetrameric p66Shc. These results highlight the important interplay between mitochondrial antioxidant system and the level of mtROS, simultaneously raising the question on how cells determine the level of cellular ROS required for redox signaling transduction.
The above redox signaling between the ER-MAM and mitochondria illustrates the existence of inter-organelle redox signaling regulation. Notably, there are growing studies to suggest MAM might play a role in the pathogenesis of neurodegenerative diseases, as reviewed by Rodríguez-Arribas et al.77. Considering the potential existence of the redox nanodomains, MAM’s association in neurodegenerative and age-related diseases might be of interest to determine whether this pocket of mitochondrial-derived redox signaling might influence the pathogenesis of neurodegenerative diseases.
Mitochondrial hydrogen peroxide and their potential role in the pathogenesis of neurodegenerative diseases
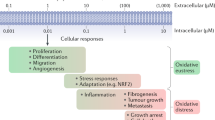
The brain consumes nearly a quarter of the body’s total intake of glucose and O278,79, with 20% of the total O2 uptake driving ATP-producing OXPHOS. The brain’s high metabolic level produces high levels of ROS, making brain tissue the most vulnerable tissue to oxidative damage80. Indeed, the mitochondrial and free-radical theories of aging attribute the brain’s high cellular ROS levels to oxidative damage and aging80,81; however, recent and growing evidence shows cellular ROS as essential signaling molecules, especially with cellular H2O2-associated oxPTMs.
This paradigm shift is derived mainly from model-based organism studies with invertebrates such as Caenorhabditis elegans and yeast. For example, a study by Schulz et al.82 demonstrated that glucose deprivation increases OXPHOS and mtROS, and delays C. elegans aging. Similarly, a recent study by Wang et al.83 showed that lifespan extension conferred by the increased mtROS was diminished by glucose via reducing cellular ROS levels. Other studies echo this concept by identifying the increase in mtROS as a common aftermath of numerous conserved longevity-promoting interventions, thus originating the term “mitohormesis”84,85. Mitohormesis is the process of mitochondrial adaption from increased mtROS exposure, activating stress resistance mechanisms to improve anti-aging effects85. As the brain consumes a high volume of glucose for energy production, the detrimental effect of glucose on aging and neurodegenerative diseases could stem from mitochondrial dysfunction, as highlighted by Cheng et al.86. The disruption of redox balance and glucose regulation in dysfunctional mitochondria could lead to the development of neurodegenerative diseases. In addition, a recent study by Smith et al.87 demonstrated that the GSH redox pathway can control mitochondrial shape in axons of Drosophila. Collectively, these observations accentuate the importance of “healthy” mitochondria in preventing age-related diseases like neurodegenerative diseases.
ROS formation (mainly O2•−) increases with increasing O2 concentration88,89,90,91. However, studies have revealed that ROS levels are either constant or elevated with declining O2 levels92,93. Moreover, aging and neurodegenerative diseases are closely linked to hypoxia94. Despite the hypoxic decline of O2 levels in aging, O2 to the brain is not completely depleted, but the state of chronic hypoxia increases oxidative stress and trigger apoptosis and neurodegeneration94. In a review, Waypa et al.95 explored the possibility that mtROS are O2 sensors during hypoxia. Previous studies have shown intracellular H2O2 levels reversibly increase during hypoxia96,97,98,99; inhibiting mitochondrial complex III-mediated ROS production during hypoxia impeded the stabilization of hypoxia inducible factor-1 subunit alpha (HIF-1α) and affected the cellular response to hypoxia100,101. These observations emphasize the importance of functional mitochondria, failure of which could explain the manifestation of neurodegenerative diseases.
A review by Shadel and Horvath102 illustrated many important mtROS signaling pathways in reversing aging processes: particularly, the relationship between UCP2 and mtROS in controlling energy metabolism of the brain. During starvation, the increase in mtROS production from lipid oxidation can promote UCP2 activity to preserve low ROS levels while maintaining energy production103. In the well-fed state, glucose OXPHOS increases mtROS production104, which drives leptin and insulin to exert the systemic equivalent of satiety103,105,106,107,108. Importantly, disruption to such mitochondrial redox signaling would impair these physiological processes;103,106,108,109 this highlights the importance of mtROS and may explain the near-zero success rates in antioxidant therapies for neurodegenerative diseases61. Effective neurodegenerative disease treatment thus lies in redox homeostasis rather than the absolute amount of cellular ROS. The solution should restore redox balance and redox signaling, maintaining the correct ROS type, location in the signaling pathway, and amounts.
A hallmark of aging is the accumulation of damaged mitochondria in the brain; particularly, ETC activity is disrupted as aging progresses110,111,112,113,114,115. Decreased Complex IV activity increases mtROS production, whereas reduced Complex I activity affects cellular differentiation. Interestingly, mtROS produced by Complex I enters the MM, whereas mtROS by Complex III enters the IMS and is involved in site-specific redox signaling (such as HIF-1α signaling, as previously highlighted)116. These differences imply site-specific mitochondrial redox signaling pathways exist, which are disrupted by the accumulation of damaged mitochondria and subsequent elevation of mtROS levels.
Two types of ROS exist: low-reactive ROS (such as O2•− and H2O2), and high-reactive ROS (such as hydroxyl radicals and peroxynitrites, produced by H2O2 undergoing Fenton’s Haber–Weiss reactions)117,118. Although low-reactive ROS act as secondary messengers, high-reactive ROS contribute to oxidative damage and cell death. A study by Forster et al.119 showed oxidative stress to correlate with loss of cognitive ability in rodents, whereas neuronal cell death initiates brain aging and triggers neurodegenerative diseases120. However, mtROS should not be synonymized with oxidative stress and neuronal death, as we continuously emphasize the role of mtROS in redox signaling.
The aforementioned discussions, along with a review by Angelova and Abramov121, point out mitochondrial dysfunction as a pathological cue to the progression of various neurodegenerative diseases. Although damaged mitochondria are usually removed by quality control processes such as mitophagy and proteasomal degradation122, aging could reduce the efficiency and quality of these corrective processes, causing gradual accumulation of “unhealthy” mitochondria. Studies have shown that increasing the efficiency of proteasomal activities and mitophagy could increase longevity of various model organisms including worms, flies, and mice123,124,125,126. Moreover, Chen et al.127 highlights the importance of mitophagy in the pathology of neurodegenerative diseases. Although there is no concrete evidence yet to suggest that maintaining a pool of healthy mitochondria is key to reducing the negative aspects of aging, the manifestation of neurodegenerative diseases through such a scenario might still hold true. A review by Stefanatos and Sanz110 suggests that specific mtROS, like mtH2O2, are produced in specific locations and amounts to drive site-specific redox signaling in healthy mitochondria; conversely, “unhealthy” mitochondria produce ROS of the wrong type, location, or amount, which could derail redox signaling. This is highlighted in our previous study showing mitophagy activation by mtROS, suggesting that understanding the complexity of redox signaling in the brain, and how they are alternated in neurodegenerative diseases are beginning to be of importance to further our understanding of their pathogenesis and identify correct therapeutic interventions.
The studies highlighted thus far demonstrate the importance of mtROS in redox signaling, though the type of ROS is rarely specified. A review by Mailloux128 explains that mtO2•− must be removed swiftly to prevent the deactivation of Fe-S cluster-containing proteins in the mitochondria; the rapid conversion of mtO2•− to mtH2O2 maintains its concentration at ρM range14. Therefore, the dominant mtROS in the matrix is mtH2O2. Apart from mtO2•−, mitochondria also have an extensive system of removing mtH2O2; in addition, this system acts as a sink for cellular H2O2, as suggested by Mailloux. A study by Dey et al.129 highlights the importance of the mitochondrial antioxidant defenses, wherein the loss of the mitochondrial antioxidant defenses led to the dysregulation of the cytosolic H2O2 redox environment. This study emphasizes the role of mitochondria in governing the cellular levels of H2O2, thus demonstrating the mitochondria’s important role in modulating cellular H2O2 signals. Therefore, it is important to further our understanding in the events how dysregulation of mtH2O2 and cytosolic H2O2 in neurodegenerative diseases could lead to their pathologies.
Conclusion and future perspectives of mitochondrial and ER-derived hydrogen peroxide in neurodegenerative diseases
Although current therapeutic approaches against neurodegenerative diseases lack success, continuous research has enhanced understanding of such diseases. Mitochondrial dysfunction has been identified as an important hallmark, and ROS as important secondary messengers. Furthermore, a key aspect of neurodegenerative diseases pathogenesis has been suggested: the potential and paramount role of redox signaling.
We mentioned earlier that the presence of communication between the MAM at the ER and mitochondrial OMM interface was identified in recent studies, and that H2O2 could affect the Ca2+ exchange between ER and mitochondria (Fig. 4A). Ca2+ signaling has been implicated in various neurodegenerative diseases, where their protein aggregates can target various Ca2+ channels and affect their Ca2+ flux130. For instance, α-synuclein (α-syn), the major aggregated protein in PD pathology, reportedly associates with MAM131. However, the pathological implication of this association has not been found. As Ca2+ is known to play a role in the ER during protein folding132, and aberrant protein aggregation characterizes AD and PD, disrupted Ca2+ signaling may promote disease-associated protein misfolding and aggregation. Furthermore, previous studies showed Ca2+ to partake in cellular quality control such as autophagy133, further tying the dysregulation of Ca2+ signaling to the accumulation of protein aggregates. Therefore, understanding the relationship of Ca2+ and redox signaling could help better understand their roles in neurodegenerative diseases.
A Investigate how the dysregulation of redox-regulated calcium (Ca2+) signaling between the endoplasmic reticulum (ER) and the mitochondria (Mito) could contribute to the pathogenesis of neurodegenerative diseases; MAM: mitochondria-associated membrane. B Understand how the release of redox signals such as mitochondrial hydrogen peroxide (H2O2) into the MAM-mitochondrial interface could affect ER-mitochondria tethering and dysregulate mitochondrial dynamics that could contribute to the pathogenesis of neurodegenerative diseases. C Understand how site-specific redox signaling in the mitochondria could be mediated by the levels of intracellular ROS and determine the presence of H2O2 pool that can act as a reservoir of secondary messengers to trigger different signaling pathways throughout the whole cells. D Identify the underlying redox regulation of mitochondrial quality control and its significance to the pathogenesis of neurodegenerative diseases. Created with BioRender.com.
Besides Ca2+ and redox signaling, another area of interest is the potential role of mitochondria tethering to the MAM, which can modulate mitochondrial morphology. As highlighted previously, oxPTMs of MFN2 by H2O2 can initiate mitofusion (Fig. 4B). Notably, we highlighted that mtH2O2 release to the MAM-mitochondrial interface during MM swelling (Fig. 3C) could generate a pool of H2O2 as secondary messengers for redox signaling. If this pool exists, it could affect various signaling pathways like Ca2+ signaling. Furthermore, this pool of mtH2O2 could affect mitochondria tethering to the MAM, thus affecting mitochondrial morphology and recycling, including the clearance of “unhealthy” mitochondria and generation of “incorrect” ROS types. Therefore, whether α-syn association with MAM disrupts mitochondrial turnover and homeostasis in PD could be of interest to study.
As suggested previously, the existence of site-specific redox signaling by mtROS could trigger different processes. Some of these known site-specific ROS-driven processes (like cell growth and differentiation) are involved in nucleus signaling, suggesting mtROS secretion to the MAM-mitochondrial interface to create a ROS pool, as mentioned above (Fig. 4C). Transmission of the redox signal could occur: first, from the mitochondria to the ER then to the nucleus; secondly, mitochondria tethering to an ER region near the nucleus; or thirdly, mitochondria tethering directly to the nucleus. Interestingly, a recent study by Pak et al.134 debunked the existence of H2O2 efflux from the mitochondria to the cytosol despite identifying the existence of intracellular H2O2 gradient; however, the experiments were done using K562 lymphoblast cells instead of a normal cell line. Therefore, replicating the experiments in cell lines such as neurons will give insight into the existence of a mtH2O2 pool in the MAM-mitochondrial interface and its role in neurodegenerative diseases.
The “inaccurate” redox signal could also affect cellular quality controls, such as proteasomal degradation of misfolded or aggregated proteins, or organellar quality control, such as mitophagy (Fig. 4D). In the latter case, our previous study showed that ROS could trigger mitophagy directly, demonstrating that mitophagy could also be a redox-sensitive process. Further research on the regulation of cellular quality control by redox signaling would aid the identification of diagnostic and therapeutic targets in neurodegenerative diseases.
Evidently, the involvement of ROS in aging and neurodegenerative diseases must be further investigated to elucidate disease pathogenesis. One approach is to identify proteins that undergo cysteine oxPTMs and map the neuronal redox signaling pathways. Through this, disruptions to redox signaling pathways in neurodegenerative diseases can be identified. Various methods to identify proteins with oxPTMs were reviewed elsewhere42; these assays use thiol-labeling chemicals such as iodoacetamide135 to identify proteins with thiol groups oxidized by intracellular redox signaling. Coupled with an appropriate reporter such as biotin, the oxPTMed proteins can be identified after pulldown and mass spectrometry (MS). Most of these methods operate under the assumption that the assay will not react to oxidized thiols of target proteins. However, a major disadvantage of these approaches is the lack of quantification. For instance, a 10-fold change in oxPTMs may influence little if it curtails 0.1–1% of the intact protein level, compared with a 10-fold change representing 10–100% of the intact protein level. Therefore, quantitative proteomics has been used to address this issue via the use of labels such as isotope-coded affinity tag (ICAT).
ICAT is similar to the above approaches but utilizes isotopic tags to differentially label samples. The mass difference of the isotopic tags is detected and quantified using MS, thus determining the level of oxPTMed cysteine of the target protein. This approach has been used for over a decade, and Topf et al.136 demonstrated recently on its use for their study to identify redox switches modulating the global protein translation in yeast triggered by mtROS. However, ICAT detects only the site of oxPTM, not the type. In addition, S-OH is considered a key redox switch in the field of oxPTMs, thus methods to identify such intermediates were developed137,138. Most approaches thus far utilize chemical derivatization or trapping to induce reactions with S-OH, forming a stable thioether adduct139,140,141 that can then be quantified using MS or via a fluorescent or biotinylated tag142,143,144,145.
Besides the identification of oxPTMed proteins, identifying the localization of cellular ROS, particularly mitochondrial and ER-H2O2, can give greater insight into redox signaling. As highlighted in a methods paper by Oparka et al.146, the ROS-reactive fluorescent probe 2´,7´-dichlorodihydrofluorecein diacetate (H2DCFDA) and its derivatives have been widely used as a H2O2-specific probe in intact cells. However, H2DCFDA is not specific to H2O2. A H2O2-specific biosensor, termed HyPer, was subsequently developed from circularly permutated YFP inserted into the H2O2-sensitive OxyR-regulatory domain from Escherichia coli147,148,149. As HyPer is both H2O2 and pH-sensitive, a parallel control using SypHer is required, as it is only pH-sensitive but possesses the same spectral properties150,151,152. Furthermore, these H2O2-biosensors are insensitive to smaller changes to H2O2 concentration near the basal oxidant level134. Fortunately, Pak et al.134, in a recent study, developed a new HyPer variant, termed HyPer7, using circularly permutated GFP inserted into the OxyR-regulatory domain from Neisseria meningitidis. This H2O2-biosensor is ultrasensitive, ultrafast, and pH-stable. Using this probe, the authors identified the existence of H2O2 gradients within the cell and evaluated the H2O2 movement between the mitochondria and cytosol.
Despite the improving technology, much remains lacking for reliable and robust methodology in redox signaling, including the absolute quantification of oxidized proteins that can be mapped onto complex biological systems. Before such a system is in place, we will continue to rely on current methods, preferably utilizing multiple approaches to improve the accuracy of our research data on identifying the redox signaling of neurodegenerative diseases.
References
Grimm, A. & Eckert, A. Brain aging and neurodegeneration: from a mitochondrial point of view. J. Neurochem. 143, 418–431 (2017).
Selvaraji, S., Poh, L., Natarajan, V., Mallilankaraman, K. & Arumugam, T. V. Negative conditioning of mitochondrial dysfunction in age-related neurodegenerative diseases. Cond. Med. 2, 30–39 (2019).
Akbar, M. et al. Mitochondrial dysfunction and cell death in neurodegenerative diseases through nitroxidative stress. Brain Res. 1637, 34–55 (2016).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Tönnies, E. & Trushina, E. Oxidative stress, synaptic dysfunction, and Alzheimer’s disease. J. Alzheimer’s Dis. 57, 1105–1121 (2017).
Ayala, A., Muñoz, M. F. & Argüelles, S. Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell Longev. 2014, 360438 (2014).
Pham-Huy, L. A., He, H. & Pham-Huy, C. Free radicals, antioxidants in disease and health. Int J. Biomed. Sci. 4, 89–96 (2008).
Zuo, L., Zhou, T., Pannell, B. K., Ziegler, A. C. & Best, T. M. Biological and physiological role of reactive oxygen species—the good, the bad and the ugly. Acta Physiol. (Oxf.) 214, 329–348 (2015).
Zhang, J. et al. ROS and ROS-mediated cellular signaling. Oxid. Med. Cell Longev. 2016, 4350965 (2016).
Sies, H. Role of metabolic H2O2 generation: redox signaling and oxidative stress. J. Biol. Chem. 289, 8735–8741 (2014).
Nordberg, J. & Arnér, E. S. Reactive oxygen species, antioxidants, and the mammalian thioredoxin system. Free Radic. Biol. Med. 31, 1287–1312 (2001).
Tu, B. P. & Weissman, J. S. Oxidative protein folding in eukaryotes: mechanisms and consequences. J. Cell Biol. 164, 341–346 (2004).
Finkel, T. Signal transduction by reactive oxygen species. J. Cell Biol. 194, 7–15 (2011).
Murphy, M. P. How mitochondria produce reactive oxygen species. Biochem. J. 417, 1–13 (2009).
Fukai, T. & Ushio-Fukai, M. Superoxide dismutases: role in redox signaling, vascular function, and diseases. Antioxid. Redox Signal. 15, 1583–1606 (2011).
Handy, D. E. & Loscalzo, J. Redox regulation of mitochondrial function. Antioxid. Redox Signal. 16, 1323–1367 (2012).
Bienert, G. P. & Chaumont, F. Aquaporin-facilitated transmembrane diffusion of hydrogen peroxide. Biochim. Biophys. Acta Gen. Subj. 1840, 1596–1604 (2014).
Malinouski, M., Zhou, Y., Belousov, V. V., Hatfield, D. L. & Gladyshev, V. N. Hydrogen peroxide probes directed to different cellular compartments. PLoS ONE 6, e14564 (2011).
Sena, L. A. & Chandel, N. S. Physiological roles of mitochondrial reactive oxygen species. Mol. Cell 48, 158–167 (2012).
Giulivi, C., Boveris, A. & Cadenas, E. in Reactive Oxygen Species in Biological Systems: An Interdisciplinary Approach (eds. Daniel L. Gilbert & Carol A. Colton) 77–102 (Springer US, Boston, MA, 2002).
Ramming, T. et al. A PDI-catalyzed thiol-disulfide switch regulates the production of hydrogen peroxide by human Ero1. Free Radic. Biol. Med. 83, 361–372 (2015).
Guo, S. & Chen, X. The human Nox4: gene, structure, physiological function and pathological significance. J. Drug Target 23, 888–896 (2015).
Serrander, L. et al. NOX4 activity is determined by mRNA levels and reveals a unique pattern of ROS generation. Biochem. J. 406, 105–114 (2007).
Block, K., Gorin, Y. & Abboud, H. E. Subcellular localization of Nox4 and regulation in diabetes. Proc. Natl Acad. Sci. USA 106, 14385–14390 (2009).
Kuroda, J. et al. The superoxide-producing NAD(P)H oxidase Nox4 in the nucleus of human vascular endothelial cells. Genes Cells 10, 1139–1151 (2005).
Sies, H. & Jones, D. P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell Biol. 21, 363–383 (2020).
Chen, F., Haigh, S., Barman, S. & Fulton, D. J. R. From form to function: the role of Nox4 in the cardiovascular system. Front. Physiol. 3, 412 (2012).
Graham, K. A. et al. NADPH oxidase 4 is an oncoprotein localized to mitochondria. Cancer Biol. Ther. 10, 223–231 (2010).
Delaunay-Moisan, A. & Appenzeller-Herzog, C. The antioxidant machinery of the endoplasmic reticulum: protection and signaling. Free Radic. Biol. Med. 83, 341–351 (2015).
Antonenkov, V. D., Grunau, S., Ohlmeier, S. & Hiltunen, J. K. Peroxisomes are oxidative organelles. Antioxid. Redox Signal. 13, 525–537 (2010).
Yoboue, E. D., Sitia, R. & Simmen, T. Redox crosstalk at endoplasmic reticulum (ER) membrane contact sites (MCS) uses toxic waste to deliver messages. Cell Death Dis. 9, 331 (2018).
Mann, M. & Jensen, O. N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21, 255–261 (2003).
Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: oxidative eustress. Redox Biol. 11, 613–619 (2017).
Wani, R., Nagata, A. & Murray, B. W. Protein redox chemistry: Post-translational cysteine modifications that regulate signal transduction and drug pharmacology. Front. Pharmacol. 5, 224 (2014).
Denu, J. M. & Tanner, K. G. Specific and reversible inactivation of protein tyrosine phosphatases by hydrogen peroxide: evidence for a sulfenic acid intermediate and implications for redox regulation. Biochemistry 37, 5633–5642 (1998).
Wang, Y., Yang, J. & Yi, J. Redox sensing by proteins: oxidative modifications on cysteines and the consequent events. Antioxid. Redox Signal. 16, 649–657 (2012).
Brandes, N., Schmitt, S. & Jakob, U. Thiol-based redox switches in eukaryotic proteins. Antioxid. Redox Signal. 11, 997–1014 (2009).
Bonifacic, M. & Asmus, K. D. Adduct formation and absolute rate constants in the displacement reaction of thiyl radicals with disulfides. J. Phys. Chem. 88, 6286–6290 (1984).
Roos, G. & Messens, J. Protein sulfenic acid formation: from cellular damage to redox regulation. Free Radic. Biol. Med. 51, 314–326 (2011).
Hogg, P. J. Disulfide bonds as switches for protein function. Trends Biochem. Sci. 28, 210–214 (2003).
Wang, S. B. et al. Redox regulation of mitochondrial ATP synthase: Implications for cardiac resynchronization therapy. Circ. Res. 109, 750–757 (2011).
Zhang, J., Ye, Z. W., Singh, S., Townsend, D. M. & Tew, K. D. An evolving understanding of the S-glutathionylation cycle in pathways of redox regulation. Free Radic. Biol. Med. 120, 204–216 (2018).
Hyun, A. W. et al. Reduction of cysteine sulfinic acid by sulfiredoxin is specific to 2-Cys peroxiredoxins. J. Biol. Chem. 280, 3125–3128 (2005).
Lowther, W. T. & Haynes, A. C. Reduction of cysteine sulfinic acid in eukaryotic, typical 2-Cys peroxiredoxins by sulfiredoxin. Antioxid. Redox Signal. 15, 99–109 (2011).
Chung, H. S., Wang, S. B., Venkatraman, V., Murray, C. I. & Van Eyk, J. E. Cysteine oxidative posttranslational modifications: emerging regulation in the cardiovascular system. Circ. Res. 112, 382–392 (2013).
Miseta, A. & Csutora, P. Relationship between the occurrence of cysteine in proteins and the complexity of organisms. Mol. Biol. Evol. 17, 1232–1239 (2000).
Orrenius, S., Gogvadze, V. & Zhivotovsky, B. Calcium and mitochondria in the regulation of cell death. Biochem. Biophys. Res. Commun. 460, 72–81 (2015).
Kaminskyy, V. O. & Zhivotovsky, B. Free radicals in cross talk between autophagy and apoptosis. Antioxid. Redox Signal. 21, 86–102 (2014).
Wang, X. The expanding role of mitochondria in apoptosis. Genes Dev. 15, 2922–2933 (2001).
Taylor, R. C., Cullen, S. P. & Martin, S. J. Apoptosis: controlled demolition at the cellular level. Nat. Rev. Mol. Cell Biol. 9, 231–241 (2008).
Düssmann, H., Kögel, D., Rehm, M. & Prehn, J. H. Mitochondrial membrane permeabilization and superoxide production during apoptosis. A single-cell analysis. J. Biol. Chem. 278, 12645–12649 (2003).
D’Alessio, M. et al. Oxidative Bax dimerization promotes its translocation to mitochondria independently of apoptosis. FASEB J. 19, 1504–1506 (2005).
Nie, C. et al. Cysteine 62 of Bax is critical for its conformational activation and its proapoptotic activity in response to H2O2-induced apoptosis. J. Biol. Chem. 283, 15359–15369 (2008).
Circu, M. L. & Aw, T. Y. Glutathione and modulation of cell apoptosis. Biochim. Biophys. Acta 1823, 1767–1777 (2012).
Benhar, M. Oxidants, antioxidants and thiol redox switches in the control of regulated cell death Ppathways. Antioxid. (Basel) 9, 309 (2020).
Okouchi, M., Ekshyyan, O., Maracine, M. & Aw, T. Y. Neuronal apoptosis in neurodegeneration. Antioxid. Redox Signal. 9, 1059–1096 (2007).
Quinlan, C. L. et al. The 2-oxoacid dehydrogenase complexes in mitochondria can produce superoxide/hydrogen peroxide at much higher rates than complex I. J. Biol. Chem. 289, 8312–8325 (2014).
O’Brien, M., Chalker, J., Slade, L., Gardiner, D. & Mailloux, R. J. Protein S-glutathionylation alters superoxide/hydrogen peroxide emission from pyruvate dehydrogenase complex. Free Radic. Biol. Med. 106, 302–314 (2017).
Chouchani, E. T. et al. Mitochondrial ROS regulate thermogenic energy expenditure and sulfenylation of UCP1. Nature 532, 112–116 (2016).
Finkel, T. From sulfenylation to sulfhydration: what a thiolate needs to tolerate. Sci. Signal 5, pe10 (2012).
Sanz, A., Pamplona, R. & Barja, G. Is the mitochondrial free radical theory of aging intact? Antioxid. Redox Signal. 8, 582–599 (2006).
Sanz, A. Mitochondrial reactive oxygen species: Do they extend or shorten animal lifespan? Biochim. Biophys. Acta 1857, 1116–1126 (2016).
Eckert, A. et al. Mitochondrial dysfunction, apoptotic cell death, and Alzheimer’s disease. Biochem. Pharmacol. 66, 1627–1634 (2003).
Chen, H. & Chan, D. C. Mitochondrial dynamics-fusion, fission, movement, and mitophagy-in neurodegenerative diseases. Hum. Mol. Genet. 18, R169–R176 (2009).
Thaher, O. et al. The thiol switch C684 in Mitofusin-2 mediates redox-induced alterations of mitochondrial shape and respiration. Neurochem. Int. 117, 167–173 (2018).
Mailloux, R. J. & Treberg, J. R. Protein S-glutathionlyation links energy metabolism to redox signaling in mitochondria. Redox Biol. 8, 110–118 (2016).
Xiao, B. et al. Reactive oxygen species trigger Parkin/PINK1 pathway–dependent mitophagy by inducing mitochondrial recruitment of Parkin. J. Biol. Chem. 292, 16697–16708 (2017).
Meng, F. et al. Oxidation of the cysteine-rich regions of parkin perturbs its E3 ligase activity and contributes to protein aggregation. Mol. Neurodegener. 6, 34 (2011).
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007).
Pinton, P., Romagnoli, A., Rizzuto, R. & Giorgi, C. Ca2+ signaling, mitochondria and cell death. Curr. Mol. Med. 8, 119–130 (2008).
Szabadkai, G. et al. Chaperone-mediated coupling of endoplasmic reticulum and mitochondrial Ca2+ channels. J. Cell Biol. 175, 901–911 (2006).
Booth, D. M., Enyedi, B., Geiszt, M., Várnai, P. & Hajnóczky, G. Redox nanodomains are induced by and control calcium signaling at the ER-mitochondrial interface. Mol. Cell 63, 240–248 (2016).
Lebiedzinska, M., Duszynski, J., Rizzuto, R., Pinton, P. & Wieckowski, M. R. Age-related changes in levels of p66Shc and serine 36-phosphorylated p66Shc in organs and mouse tissues. Arch. Biochem. Biophys. 486, 73–80 (2009).
Pinton, P. et al. Protein kinase C beta and prolyl isomerase 1 regulate mitochondrial effects of the life-span determinant p66Shc. Science 315, 659–663 (2007).
Gertz, M., Fischer, F., Wolters, D. & Steegborn, C. Activation of the lifespan regulator p66Shc through reversible disulfide bond formation. Proc. Natl Acad. Sci. USA 105, 5705–5709 (2008).
Gertz, M. & Steegborn, C. The Lifespan-regulator p66Shc in mitochondria: redox enzyme or redox sensor? Antioxid. Redox Signal. 13, 1417–1428 (2010).
Rodríguez-Arribas, M. et al. Mitochondria-associated membranes (MAMs): overview and its role in Parkinson’s disease. Mol. Neurobiol. 54, 6287–6303 (2017).
Erbsloh, F., Bernsmeier, A. & Hillesheim, H. The glucose consumption of the brain & its dependence on the liver. Arch. Psychiatr. Nervenkr Z. Gesamt. Neurol. Psychiatr. 196, 611–626 (1958).
Mergenthaler, P., Lindauer, U., Dienel, G. A. & Meisel, A. Sugar for the brain: the role of glucose in physiological and pathological brain function. Trends Neurosci. 36, 587–597 (2013).
Gadoth, N. & Göbel, H. H. Oxidative Stress and Free Radical Damage in Neurology. (Humana Press, New York, 2011).
Barja, G. Updating the mitochondrial free radical theory of aging: an integrated view, key aspects, and confounding concepts. Antioxid. Redox Signal. 19, 1420–1445 (2013).
Schulz, T. J. et al. Glucose restriction extends Caenorhabditis elegans life span by inducing miochondrial respiration and increasing oxidative stress. Cell Metab. 6, 280–293 (2007).
Wang, C., Zhang, Y., Li, F. & Wei, Y. Conserved roles of glucose in suppressing reactive oxygen species-induced cell death and animal survival. Aging (Albany N. Y.) 11, 5726–5743 (2019).
Ristow, M. & Zarse, K. How increased oxidative stress promotes longevity and metabolic health: The concept of mitochondrial hormesis (mitohormesis). Exp. Gerontol. 45, 410–418 (2010).
Yun, J. & Finkel, T. Mitohormesis. Cell Metab. 19, 757–766 (2014).
Cheng, H. et al. Mitochondrial dysfunction plays a key role in the development of neurodegenerative diseases in diabetes. Am. J. Physiol. Endocrinol. Metab. 318, E750–E764 (2020).
Smith, G. A. et al. Glutathione S-transferase regulates mitochondrial populations in axons through increased glutathione oxidation. Neuron 103, 52–65.e56 (2019).
Turrens, J. F. Superoxide production by the mitochondrial respiratory chain. Biosci. Rep. 17, 3–8 (1997).
Genova, M. L. et al. The site of production of superoxide radical in mitochondrial Complex I is not a bound ubisemiquinone but presumably iron–sulfur cluster N2. FEBS Lett. 505, 364–368 (2001).
Alvarez, S., Valdez, L. B., Zaobornyj, T. & Boveris, A. Oxygen dependence of mitochondrial nitric oxide synthase activity. Biochem. Biophys. Res. Commun. 305, 771–775 (2003).
Zorov, D. B., Juhaszova, M. & Sollott, S. J. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol. Rev. 94, 909–950 (2014).
Chandel, N. S. et al. Mitochondrial reactive oxygen species trigger hypoxia-induced transcription. Proc. Natl Acad. Sci. USA 95, 11715–11720 (1998).
Waypa, G. B. & Schumacker, P. T. O2 sensing in hypoxic pulmonary vasoconstriction: the mitochondrial door re-opens. Respir. Physiol. Neurobiol. 132, 81–91 (2002).
Hota, K. B., Hota, S. K. & Singh, S. B. in Brain Aging and Therapeutic Interventions (eds Mahendra K. Thakur & Suresh I. S. Rattan) 177–189 (Springer Netherlands, Dordrecht, 2012).
Waypa, G. B., Smith, K. A. & Schumacker, P. T. O2 sensing, mitochondria and ROS signaling: the fog is lifting. Mol. Asp. Med. 47-48, 76–89 (2016).
Waypa, G. B. et al. Mitochondrial reactive oxygen species trigger calcium increases during hypoxia in pulmonary arterial myocytes. Circ. Res. 91, 719–726 (2002).
Waypa, G. B. et al. Increases in mitochondrial reactive oxygen species trigger hypoxia-induced calcium responses in pulmonary artery smooth muscle cells. Circ. Res. 99, 970–978 (2006).
Waypa, G. B. et al. Hypoxia triggers subcellular compartmental redox signaling in vascular smooth muscle cells. Circ. Res. 106, 526–535 (2010).
Waypa, G. B. et al. Superoxide generated at mitochondrial complex III triggers acute responses to hypoxia in the pulmonary circulation. Am. J. Respir. Crit. Care Med. 187, 424–432 (2013).
Cho, Y. S. et al. Functional inhibition of UQCRB suppresses angiogenesis in zebrafish. Biochem. Biophys. Res. Commun. 433, 396–400 (2013).
Orr, A. L. et al. Suppressors of superoxide production from mitochondrial complex III. Nat. Chem. Biol. 11, 834–836 (2015).
Shadel, G. S. & Horvath, T. L. Mitochondrial ROS signaling in organismal homeostasis. Cell 163, 560–569 (2015).
Andrews, Z. B. et al. UCP2 mediates ghrelin’s action on NPY/AgRP neurons by lowering free radicals. Nature 454, 846–851 (2008).
Parton, L. E. et al. Glucose sensing by POMC neurons regulates glucose homeostasis and is impaired in obesity. Nature 449, 228–232 (2007).
Benani, A. et al. Role for mitochondrial reactive oxygen species in brain lipid sensing: redox regulation of food intake. Diabetes 56, 152–160 (2007).
Diano, S. et al. Peroxisome proliferation-associated control of reactive oxygen species sets melanocortin tone and feeding in diet-induced obesity. Nat. Med. 17, 1121–1128 (2011).
Leloup, C. et al. Mitochondrial reactive oxygen species are required for hypothalamic glucose sensing. Diabetes 55, 2084–2090 (2006).
Long, L., Toda, C., Jeong, J. K., Horvath, T. L. & Diano, S. PPARγ ablation sensitizes proopiomelanocortin neurons to leptin during high-fat feeding. J. Clin. Investig. 124, 4017–4027 (2014).
Horvath, T. L., Andrews, Z. B. & Diano, S. Fuel utilization by hypothalamic neurons: roles for ROS. Trends Endocrinol. Metab. 20, 78–87 (2009).
Stefanatos, R. & Sanz, A. The role of mitochondrial ROS in the aging brain. FEBS Lett. 592, 743–758 (2018).
Cabré, R. et al. Sixty years old is the breakpoint of human frontal cortex aging. Free Radic. Biol. Med. 103, 14–22 (2017).
Ferguson, M., Mockett, R. J., Shen, Y., Orr, W. C. & Sohal, R. S. Age-associated decline in mitochondrial respiration and electron transport in Drosophila melanogaster. Biochem. J. 390, 501–511 (2005).
Navarro, A. & Boveris, A. Rat brain and liver mitochondria develop oxidative stress and lose enzymatic activities on aging. Am. J. Physiol. Regul. Integr. Comp. Physiol. 287, R1244–R1249 (2004).
Rygiel, K. A., Grady, J. P. & Turnbull, D. M. Respiratory chain deficiency in aged spinal motor neurons. Neurobiol. Aging 35, 2230–2238 (2014).
Scialò, F. et al. Mitochondrial ROS produced via reverse electron transport extend animal lifespan. Cell Metab. 23, 725–734 (2016).
Muller, F. L., Liu, Y. & Van Remmen, H. Complex III releases superoxide to both sides of the inner mitochondrial membrane. J. Biol. Chem. 279, 49064–49073 (2004).
Kirsch, M. & De Groot, H. Formation of peroxynitrite from reaction of nitroxyl anion with molecular oxygen. J. Biol. Chem. 277, 13379–13388 (2002).
Vásquez-Vivar, J., Kalyanaraman, B. & Kennedy, M. C. Mitochondrial aconitase is a source of hydroxyl radical. An electron spin resonance investigation. J. Biol. Chem. 275, 14064–14069 (2000).
Forster, M. J. et al. Age-related losses of cognitive function and motor skills in mice are associated with oxidative protein damage in the brain. Proc. Natl Acad. Sci. USA 93, 4765–4769 (1996).
Chi, H., Chang, H. Y. & Sang, T. K. Neuronal cell death mechanisms in major neurodegenerative diseases. Int. J. Mol. Sci. 19, 3082 (2018).
Angelova, P. R. & Abramov, A. Y. Role of mitochondrial ROS in the brain: from physiology to neurodegeneration. FEBS Lett. 592, 692–702 (2018).
Wang, K. & Klionsky, D. J. Mitochondria removal by autophagy. Autophagy 7, 297–300 (2011).
Chondrogianni, N., Georgila, K., Kourtis, N. & Tavernarakis, N. & Gonos Efstathios, S. Enhanced proteasome degradation extends Caenorhabditis elegans lifespan and alleviates aggregation-related pathologies. Free Radic. Biol. Med. 75, S18 (2014).
Lapierre, L. R. et al. The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in Caenorhabditis elegans. Nat. Commun. 4, 2267–2267 (2013).
Pyo, J.-O. et al. Overexpression of Atg5 in mice activates autophagy and extends lifespan. Nat. Commun. 4, 2300–2300 (2013).
Simonsen, A. et al. Promoting basal levels of autophagy in the nervous system enhances longevity and oxidant resistance in adult Drosophila. Autophagy 4, 176–184 (2008).
Chen, G., Kroemer, G. & Kepp, O. Mitophagy: an emerging role in aging and age-associated diseases. Front. Cell Dev. Biol. 8, 200 (2020).
Mailloux, R. J. Mitochondrial antioxidants and the maintenance of cellular hydrogen peroxide levels. Oxid. Med. Cell Longev. 2018, 7857251 (2018).
Dey, S., Sidor, A. & O’Rourke, B. Compartment-specific control of reactive oxygen species scavenging by antioxidant pathway enzymes. J. Biol. Chem. 291, 11185–11197 (2016).
Pchitskaya, E., Popugaeva, E. & Bezprozvanny, I. Calcium signaling and molecular mechanisms underlying neurodegenerative diseases. Cell Calcium 70, 87–94 (2018).
Paillusson, S. et al. α-Synuclein binds to the ER–mitochondria tethering protein VAPB to disrupt Ca2+ homeostasis and mitochondrial ATP production. Acta Neuropathol. 134, 129–149 (2017).
Michalak, M., Robert Parker, J. M. & Opas, M. Ca2+ signaling and calcium binding chaperones of the endoplasmic reticulum. Cell Calcium 32, 269–278 (2002).
Cárdenas, C. & Foskett, J. K. Mitochondrial Ca2+ signals in autophagy. Cell Calcium 52, 44–51 (2012).
Pak, V. V. et al. Ultrasensitive genetically encoded indicator for hydrogen peroxide identifies roles for the oxidant in cell migration and mitochondrial function. Cell Metab. 31, 642–653.e646 (2020).
Eaton, P. Protein thiol oxidation in health and disease: techniques for measuring disulfides and related modifications in complex protein mixtures. Free Radic. Biol. Med. 40, 1889–1899 (2006).
Topf, U. et al. Quantitative proteomics identifies redox switches for global translation modulation by mitochondrially produced reactive oxygen species. Nat. Commun. 9, 324–324 (2018).
Akter, S. et al. Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. Nat. Chem. Biol. 14, 995–1004 (2018).
Li, R. & Kast, J. Biotin switch assays for quantitation of reversible cysteine oxidation. Meth. Enzymol. 585, 269–284 (2017).
Devarie-Baez, N. O., Lopez, E. I. S. & Furdui, C. M. Biological chemistry and functionality of protein sulfenic acids and related thiol modifications. Free Radic. Res. 50, 172–194 (2016).
Poole, L. B. The basics of thiols and cysteines in redox biology and chemistry. Free Radic. Biol. Med. 80, 148–157 (2015).
Yang, J., Carroll, K. S. & Liebler, D. C. The expanding landscape of the thiol redox proteome. Mol. Cell Proteom. 15, 1–11 (2016).
Furdui, C. M. & Poole, L. B. Chemical approaches to detect and analyze protein sulfenic acids. Mass Spectrom. Rev. 33, 126–146 (2014).
Gupta, V., Paritala, H. & Carroll, K. S. Reactivity, selectivity, and stability in sulfenic acid detection: a comparative study of nucleophilic and electrophilic probes. Bioconjug. Chem. 27, 1411–1418 (2016).
Lo Conte, M., Lin, J., Wilson, M. A. & Carroll, K. S. A chemical approach for the detection of protein sulfinylation. ACS Chem. Biol. 10, 1825–1830 (2015).
Yang, J. et al. Global, in situ, site-specific analysis of protein S-sulfenylation. Nat. Protoc. 10, 1022–1037 (2015).
Oparka, M. et al. Quantifying ROS levels using CM-H2DCFDA and HyPer. Methods 109, 3–11 (2016).
Belousov, V. V. et al. Genetically encoded fluorescent indicator for intracellular hydrogen peroxide. Nat. Methods 3, 281–286 (2006).
Baird, G. S., Zacharias, D. A. & Tsien, R. Y. Circular permutation and receptor insertion within green fluorescent proteins. Proc. Natl Acad. Sci. U. S. A. 96, 11241–11246 (1999).
Choi, H. et al. Structural basis of the redox switch in the OxyR transcription factor. Cell 105, 103–113 (2001).
Forkink, M. et al. Complex I and complex III inhibition specifically increase cytosolic hydrogen peroxide levels without inducing oxidative stress in HEK293 cells. Redox Biol. 6, 607–616 (2015).
Poburko, D., Santo-Domingo, J. & Demaurex, N. Dynamic regulation of the mitochondrial proton gradient during cytosolic calcium elevations. J. Biol. Chem. 286, 11672–11684 (2011).
Santo-Domingo, J., Giacomello, M., Poburko, D., Scorrano, L. & Demaurex, N. OPA1 promotes pH flashes that spread between contiguous mitochondria without matrix protein exchange. EMBO J. 32, 1927–1940 (2013).
Acknowledgements
We thank all members of Y-C.L. laboratory for their valuable discussion during the writing of this review article. In addition, we would like to extend our sincere thanks to Ms. Deryn En-Jie Teoh for her help in editing this review article. This work is financially supported by the Ministry of Education (MOE), Singapore, Tier 2 (MOE-T2- 1-131) and Tier1 (A51-114; NUSMed SoM-FoS Seed Fund) grants awarded to Y-C.L. Y.M.L. is a MOE Research Scholarship Fellow.
Author information
Authors and Affiliations
Contributions
Y.M.L. and Y-C.L. identified and outlined the scope of this review article. Y.M.L., W.H., and Y-C.L. drafted the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Agostini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, Y.M., He, W. & Liou, YC. The redox language in neurodegenerative diseases: oxidative post-translational modifications by hydrogen peroxide. Cell Death Dis 12, 58 (2021). https://doi.org/10.1038/s41419-020-03355-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-03355-3
This article is cited by
-
Oxidative stress and the role of redox signalling in chronic kidney disease
Nature Reviews Nephrology (2024)
-
All-optical spatiotemporal mapping of ROS dynamics across mitochondrial microdomains in situ
Nature Communications (2023)
-
A selective fluorescent turn-on probe for imaging and sensing of hydrogen peroxide in living cells
Analytical and Bioanalytical Chemistry (2023)
-
Boronates as hydrogen peroxide–reactive warheads in the design of detection probes, prodrugs, and nanomedicines used in tumors and other diseases
Drug Delivery and Translational Research (2023)
-
Rotenone-induced oxidative stress in THP-1 cells: biphasic effects of baicalin
Molecular Biology Reports (2023)