Abstract
Accumulating evidence has indicated crucial roles for pseudogenes in human cancers. However, the roles played by pseudogenes in the pathogenesis of HCC, particularly HCC early recurrence, still incompletely elucidated. Herein, we identify a novel early recurrence related pseudogene RACGAP1P which was significantly upregulated in HCC and was associated with larger tumour size, advanced clinical stage, abnormal AFP level and shorter survival time. In vitro and in vivo experiments have shown that RACGAP1P is a prerequisite for the development of malignant characteristics of HCC cells, including cell growth and migration. Mechanistic investigations indicated that RACGAP1P elicits its oncogenic activity as a ceRNA to sequestrate miR-15-5p from its endogenous target RACGAP1, thereby leading to the upregulation of RACGAP1 and the activation of RhoA/ERK signalling. These results may provide new insights into the functional crosstalk of the pseudogene/miRNA/parent-gene genetic network during HCC early relapse and may contribute to improving the clinical intervention for this subset of HCC patients.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is one of most frequent cause of cancer-related mortality worldwide, with more than half a million deaths per annum1,2. Early HCC recurrence is predominantly responsible for dismal outcomes and is the major obstacle to improving HCC patients’ survival3,4. Therefore, molecular investigation of the early recurrence of HCC is the major challenge and there is a pressing need to develop new biomarkers to screen out HCC patients with the higher risk of recurrence, who would be extremely beneficial for the clinical intervention5.
In pursuit of the prediction of early recurrence for HCC patients, previous studies have highlighted valuable biomarkers such as Rac GTPase Activating Protein 1 (RACGAP1)6, Epithelial Cell Transforming 2 (ECT2)7, Protein Regulator Of Cytokinesis 1 (PRC1)8 et al., which are all protein-coding genes. Considering long non-coding RNAs (lncRNAs) make up the major component of human transcriptome, it is essential to determine whether lncRNAs may serve as diagnostic markers for HCC early relapse. Pseudogenes, defined as dysfunctional copies of protein-coding genes, represent an intriguing class of lncRNAs and were primitively considered as “genomic junk”, but their well preservation during the evolutionary pressures imposed by natural selection argues that they may possess important functions and that their dyregulation could contribute to the development of diseases9,10. Indeed, mounting evidence has suggested that pseudogenes are playing key roles in oncogenesis or tumour suppression11,12. In HCC, several lines of evidence have indicated that pseudogenes are aberrantly expressed, providing new insights into the pathogenesis of HCC13,14,15. However, few HCC early recurrence associated pseudogenes have been characterised thus far.
In the current study, we first identified a novel HCC early recurrence associated pseudogene RACGAP1P, of which the biological function and expression pattern in cancer have not been illuminated. We found that RACGAP1P was significantly upregulated in HCC tissues and markedly upregulated in the HCC samples with early recurrence. In addition, high expression of RACGAP1P in HCC was associated with poor prognosis. Mechanistically, RACGAP1P acts as an endogenous sponge for miR-15-5p to prevent its association with RACGAP1, leading to activation of the RhoA/ERK signalling axis and enhancing liver cancer cell proliferation and migration. Collectively, the results of this study show that RACGAP1P is an oncogenic regulator of HCC early recurrence, and may promote the development of effective recurrence-targeted therapy and improve the overall prognosis of HCC patients.
Materials and methods
Clinical specimens
Tumour and normal tissues from HCC patients that were histopathologically and clinically diagnosed, and did not receive neo-adjuvant treatment were obtained from the Affiliated Cancer Hospital and Institute of Guangzhou Medical University in Guangzhou. Written informed consent was obtained from each patient, and the study was approved by the Institute Research Ethics Committee at the Cancer Center.
Cell culture, proliferation assays and colony-formation assays
The human cancer cell lines were maintained in Dulbecco’s modified Eagle’s medium supplemented with 10% FBS at 37 °C and 5% CO2.
For the proliferation assays, 1000 cells/200 μl of medium were plated in a 96-well plate (Corning) in six copies and stained with MTS (Sigma, M2128) for 3.5 h determined at OD570 with a microplate reader every day for 6 days.
For the colony-formation assays, 500 cells/2 ml were seeded into a 6-well plate (Corning) in triplicates. The cells were washed with phosphate-buffered saline (PBS), fixed with methanol for 10 min at room temperature, and stained with 1% crystal violet for 20 min. The colonies were counted after 10 days.
Migration assay
For migration assays, 3 × 104 HCC cells in 200 μl of serum-free DMEM were seeded into 24-well plates upper chambers with an 8-mm pore size filter (Falcon), the lower chamber was filled with 600 μl of 10% FBS–DMEM. After 24 h of incubation, the cells on the lower surface of the filter were fixed and stained with 1% crystal violet. The number of migrated cells in five random optical fields (×100 magnification) from triplicate filters was averaged.
Animal experiments
All protocol was approved by the Institutional Animal Care and Use Committee of the Guangzhou medical University. For the tumourigenicity studies, Hep3B cells stably transfected with vector or RACGAP1P-3’UTR (1 × 106 cells/tumour in 150 mL DMEM with 25% matrigel) were subcutaneously injected into either side of flanks of the 5 female BALB/c athymic nude mice (3–4 weeks of age). Tumour diameters were measured in indicated day after tumours were established. Finally, the mice were euthanized, and the tumours were isolated and weighed, the tumour volume was calculated as volume = length × width2 × 0.52.
RNA extraction and real-time PCR
For real-time PCR, total RNA was extracted from cultured cell lines using Trizol reagent (Invitrogen) and subjected to reverse transcription using a cDNA Synthesis Kit (Thermo, K1622). Real-time PCR was performed using a SYBR FAST Universal qPCR Kit (KAPA, KK4602). The relative expression of the target mRNAs were calculated as two power values of ΔCt (the Ct of β-actin minus the Ct of the target cDNA). The sequences of the PCR primers used for amplification were as follows:
β-actin forward, 5′-AAGGTCATCC CTGAGCT GAA-3′;
β-actin reverse, 5′-TGACAAAGTG GTCGTTG AGG-3′;
RACGAP1 forward, 5′-GAAAGCAGAGACTGAGCGAAG-3′;
RACGAP1 reverse, 5′-GTTGAATGCTGCCAGATGTGT-3′.
RACGAP1P forward, 5′-GCACCTGTACTCTCTGCTCTAC-3′;
RACGAP1P reverse, 5′-TCCCAACAGTGACCAGAACA-3′.
DICER1 forward, 5′-TGGATAGTGGGATGTCACTGG-3′;
DICER1 reverse, 5′-CTCTGACCTTCCCGTCGTAAG-3′.
Small interfering RNA transfection
The negative control small interfering RNA (NC) and siRNA targeting human DICER1 are 5′-CAGCACCTTTACCCTTAGT-3′, were purchased from GenePharma, transient transfections of HCC cells were performed by using the Lipofectamine RNA iMAX Reagent (Invitrogen) protocol with 30 pmol siRNA in Opti-MEM Medium (Invitrogen) were mixed, and incubated at room temperature for 5 min, then the mixture was added to the cells.
MS2-RIP
The MS2bp-MS2bs-based RIP assay was performed according to previous reports16, with modifications for using the EZ-Magna RIP Kit (Millipore) in accordance with the manufacturer’s instructions.
Public expression profiles
Human pseudogene list is obtained from NCBI gene. Gene expression profiling datasets between HCC tumour tissues and normal tissues, including GSE84005, GSE76297, GSE6404, GSE54236, and GSE5975 were downloaded from GEO and TCGA. The expression data of HCC cancer cell lines was downloaded from CCLE.
Dual luciferase reporter assay
A dual luciferase reporter assay was performed as previously described with slight modifications. Briefly, the treated HCC cells were lysed, and the activities of the firefly and Renilla luciferases were analysed using a dual luciferase assay kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions. All reporter gene assays were performed in triplicate and were repeated twice. The results are expressed as the means ± SEM.
Plasmids
Luciferase reporter gene constructs containing the RACGAP1P and RACGAP1 3’ UTR region were first amplified by PCR with specific primers and then, the PCR products were cloned into pGL3-basic (Promega) digested by XbaI(NEB) using ClonExpress II One Step Cloning Kit (C112-02, Vazyme Biotech).
Full length or 3’UTR of RACGAP1P were amplified by PCR from DNA of Hep3B cells and were cloned into lentivirus vector pLVX-puro according to standard protocols. RACGAP1P and RACGAP1 3’UTR was also subcloned into the pMS2 vector for RIP analysis.
Immunoblotting
Immunoblotting was performed as described previously. The sources of the primary antibodies were as follows: anti-RACGAP1 (Proteintech, 1:1,000), anti-DICER1 (Proteintech, 1:1,000), and anti-β-actin (Proteintech, 1:1,000), p-ERK1/2 (CST, 1:1,000), total ERK (CST, 1:1,000), Active Rho Detection kit (#8820), anti-mouse and anti-rabbit peroxidase conjugated secondary antibodies were purchased from Cell Signalling Technology (Danvers, MA).
Statistical analysis
The two-tailed Student’s t-test was used to compare two independent groups of data. One-way analysis of variance (ANOVA) was used to analyze the significance among multi-groups. Chi-squared tests were applied to analyze the relationship between RACGAP1P expression and clinicopathological status. Pearson correlation analysis, Long-rank test were performed as indicated. P value < 0.05 was considered statistically significant in all cases. Asterisk (*) means p < 0.05, asterisks (**) means p < 0.01, asterisks (***) means p < 0.001. All statistical analyses were performed using the GraphPad Prism Software 5.0 or SPSS 16.0.
Results
Pseudogene RACGAP1P is consistently activated in early recurrent HCC and serves as a predictor of dismal outcome
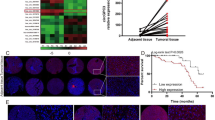
To evaluate the role of pseudogenes in HCC pathogenesis, we screened potential cancer-related pseudogenes, which were differently expressed between HCC samples and normal samples in 4 microarray datasets (GSE84005, GSE7629717, GSE6404118, GSE5423619) downloaded from Gene Expression Omnibus (GEO) and identified 7 candidates, of which 4 genes were upregulated and 3 genes were downregulated (Fig. 1a). We put our attention on RACGAP1P since its real gene, RACGAP1, was well-known to act as a oncogenic driver in HCC and importantly, played a key role in HCC early recurrence6. Similarly, assessment via data mining of RNA-seq data from The Cancer Genome Atlas (TCGA) database confirmed that RACGAP1P expression is significantly elevated in tumour tissues compared with non-tumour tissues. Of note, RACGAP1P expression was obviously upregulated in tumour from patients with early (<2 years) recurrent disease (HCC-R) compared with samples from patients with no obvious sign of recurrence within 4 years (HCC-NR) (Fig. 1b). The analogous pattern of RACGAP1P expression was further validated by QPCR (Fig. 1c). Moreover, in TCGA cohort, correlation analysis between RACGAP1P expression and clinicopathological characteristics shown that high level of RACGAP1P in primary tumours was significantly associated with the advanced T stage, clinical stage and abnormal AFP level (Table 1). Importantly, it was demonstrated that patients with high RACGAP1P expression had a significantly shorter overall (OS) and disease-free (DFS) duration compared with patients with low RACGAP1P expression (Fig. 1d, e). Taken together, these data sustain RACGAP1P as a novel prognostic factor in patients with HCC.
a Venn diagram analysis of altered pseudogenes profiling in GSE84005, GSE76297, GSE6404, and GSE54236 datasets. b The RACGAP1P mRNA expression were analysed in normal liver tissues (n = 47), non-recurrent tumours (NR-HCC, n = 22) and early recurrent tumours (R-HCC, n = 141) from TCGA. The highest RACGAP1P mRNA expression were shown in human hepatocellular carcinoma early recurrent tissues. c The RACGAP1P mRNA expression were tested in human hepatocellular carcinoma tissues by QPCR, including normal liver tissues (n = 25), non-recurrent tumours (NR-HCC, n = 25), early recurrent tumours (R-HCC, n = 41). d, e Kaplan–Meier curve shows that the overall survival (OS) and disease-free survival (DFS) rate were significantly lower in the high RACGAP1P group in TCGA cohort. ***P < 0.001, **P < 0.01, *P < 0.05. B&C, the p values were calculated by one-way ANOVA. D&E, the p values were calculated by long-rank test
Pseudogene RACGAP1P modulates RACGAP1 level in a ceRNA manner
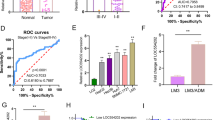
Recently, the competitive endogenous RNAs (ceRNA) hypothesis has been raised as a novel layer of gene expression modulation. The hypothesis proposed that RNA molecules share miRNA response elements (MREs) and sequester common microRNAs (miRNAs), thereby regulating expression between each other20,21,22. It is not difficult to speculate that pseudogenes may serve as ceRNAs for their parent genes, considering the high sequence similarity. To test the hypothesis that pseudogene RACGAP1P may regulate the expression of its parent gene as a ceRNA, we firstly assessed the correlation of RACGAP1P and RACGAP1 via mRNA expression data derived from the TCGA, GEO (GSE5975, GSE76297) and Cancer Cell Line Encyclopaedia (CCLE) and found the co-expression patterns between RACGAP1P and RACGAP1 in primary HCC tissues, as well as HCC cell lines (Fig. 2a, b). Then, we perform QPCR assay to validate the significantly positive correlation in HCC samples (R2 = 0.34, P = 0.0001; Fig. 2c). Next, to experimental examine whether RACGAP1P regulate the expression of RACGAP1, and whether the effect of the RACGAP1P on RACGAP1 expression was in a miRNA-dependent manner, we suppressed DICER1, a ribonuclease critical for miRNA biogenesis, in Hep3B and HCCLM3 cells using a specific siRNA (Fig. 2d, Supplementary Fig. 1a) and found that ectopic expression of RACGAP1P in wild type cells elevated the mRNA level of RACGAP1 but not in DICER1-KD cell lines (Fig. 2e). Similarly, RACGAP1 3’UTR luciferase reporter activity was significantly increased in DICER1-WT cells with RACGAP1P overexpression whereas these effects were abrogated in DICER1-KD cells (Fig. 2f). Thus, the RACGAP1 pseudogene-induced effects on the RACGAP1 level are 3’UTR and miRNA dependent in human HCC.
a, b Graphs showing a positive correlation between RACGAP1P and RACGAP1 in HCC using public gene expression datasets, including TCGA (n = 423, R2 = 0.43, p < 0.0001), GSE5975 (n = 238, R2 = 26, p < 0.0001), GSE76279 (n = 153, R2 = 0.29, p < 0.0001) and CCLE (n = 25, R2 = 0.19, p = 0.029). A, HCC clinical specimens, B, HCC cell lines. c The co-expression between RACGAP1P and RACGAP1 mRNA level was measured in 38 primary HCC tissues via QPCR (n = 38, R2 = 0.34, p = 0.0001). d The efficiency of RACGAP1P overexpression and DICER1 knockdown in HCC cell lines were determined by QPCR. e In Hep3B and HCCLM3 cells, RACGAP1P upregulated RACGAP1 mRNA levels in a DICER1 dependent manner. f RACGAP1P increased RACGAP1 luciferase reporter activity in a DICER1 dependent manner. Error bars represent means ± SD. ***P < 0.001, **P < 0.01, *P < 0.05, the p values were calculated by the Student’s t-test
Identification of the common miRNAs sharing between RACGAP1P and RACGAP1
The finding that the pseudogene RACGAP1 mediates its effect through 3’UTR and mature miRNAs suggests that it may function as a ceRNA. To further elucidate which of the miRNAs targeted both RACGAP1P and RACGAP1 to mediate the observed crosstalk, we overlaid the miRNAs predicted to target RACGAP1 and RACGAP1P in RegRNA2.023 and previously reported HCC tumour suppressive miRNAs24 and finally obtained 8 candidate miRNAs including miR-15a-5p, miR-23b-3p, miR-138-5p, miR-29c-3p, miR-125b-5p, miR-150-5p, miR-193b-3p, miR-320a (Fig. 3a). Then, Hep3B and HCCLM3 cells were delivered with these 8 potential miRNA mimics, respectively, and the expression of RACGAP1 was detected by western blot analysis. As shown in Fig. 3b, only miR-15a-5p was validated to significantly repress protein abundance of RACGAP1 in both cell lines. Therefore, miR-15a-5p was selected for further analysis and QPCR results revealed that miR-15a-5p overexpression strongly inhibited the mRNA level of RACGAP1 and RACGAP1P (Fig. 3c). Conversely, inhibition of miR-15a-5p using miRNA antagomirs could upregulated mRNA levels of both RACGAP1 and RACGAP1P (Fig. 3d).
a Eight candidate miRNAs were identified to be associated with RACGAP1P and RACGAP1 in HCC from Venn diagram analysis of three datasets, including tumour suppressor miRNAs dataset in HCC, the miRNAs datasets predicted to bind RACGAP1P and RACGAP1. b The 8 candidate miRNAs differentially regulated the RACGAP1 protein expression in Hep3B and HCCLM3 cells shown by WB. c QPCR analysis of RACGAP1P and RACGAP1 with miR-15a-5p mimics. d QPCR analysis of RACGAP1P and RACGAP1 with miR-15a-5p antagomics. e miR-15-5p seed sequence is shown along with the binding sites in RACGAP1P and RACGAP1 (shown by red box), and the mutated miR-15-5p binding sites (shown by star). f luciferase reporter assays verified the miR-15a-5p binding sites in 3’UTR of RACGAP1P and RACGAP1. g MS2-RIP strategy used to confirm endogenous miR-15a-5p binding to RACGAP1P and RACGAP1. Error bars represent means ± SD. ***P < 0.001, **P < 0.01, Student’s t-test
To further investigate whether miR-15a-5p could target both RACGAP1 and RACGAP1P, we performed the prediction of miR-15a-5p MREs for RACGAP1 and RACGAP1P transcripts and found two miR-15a-5p binding sites in RACGAP1 3’UTR and one in RACGAP1P 3’UTR (Fig. 3e). Then, we artificially introduced mutation of miR-15a-5p seed regions in luciferase reporters to disrupt the miRNA binding. The results of luciferase reporter assay demonstrated that overexpression of miR-15a-5p significantly decreased luciferase reporter activity of RACGAP1P-3’UTR and RACGAP1-3’UTR whereas the inhibition effect of miR-15a-5p was rescued on luciferase reporter with miR-15a-5p binding sites mutations (Fig. 3f). To validate the direct binding ability of the predicted miRNA response elements on these transcripts, we performed an MS2-RIP analysis using the 3’UTR regions of RACGAP1P and RACGAP1 as baits in Hep3B and HCCLM3 cells. Notably, miR-15a-5p was highly associated with the 3’UTRs of RACGAP1P and RACGAP1, and miR-21-5p as a control miRNA with no predicted MREs on RACGAP1P and RACGAP1 bound neither 3’UTR. However, the binding of miR-15a-5p on RACGAP1P and RACGAP1 3’UTR was negated in MREs mutation group (Fig. 3g). All together, these findings indicated that the ceRNA crosstalk between RACGAP1P and RACGAP1 may be mediated at least through miR-15a-5p in part.
RACGAP1P is an oncogenic driver in HCC dependent on miRNA
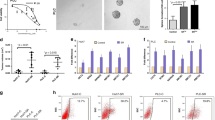
Considering increased RACGAP1 expression drives the tumourigenesis and progression of numerous types of cancer25,26,27,28, especially HCC, we determined whether aberrant control of RACGAP1 by RACGAP1P overexpression enhanced oncogenic transformation of HCC cells. Indeed, Stably exogenous expression of RACGAP1P 3’UTR in Hep3B and HCCLM3 resulted in a elevation of RACGAP1 protein, GTP-binding activity of RhoA, and ERK phosphorylation, indicating the activation of the RhoA/ERK pathway (Fig. 4a). These molecular observations were accompanied by growth promotion and a significant increase in the number of colonies, as well as high migration ability (Fig. 4b–d). Furthermore, to determine whether RACGAP1P affects tumour growth in vivo, we utilised a subcutaneous xenograft mouse model which were inoculated with Hep3B cells stably transfected with control vector or RACGAP1P 3’UTR. All of the mice developed tumours at the injection sites. However, the average size and weight of tumours generated by RACGAP1P 3’UTR-expressing cells were significantly larger than those generated from control cells (Fig. 4e–g). QPCR analysis confirmed that the average expression of RACGAP1P and RACGAP1 was much higher in RACGAP1P 3’UTR overexpressing tumour tissues compared with control tumours (Fig. 4h). Finally, immunoblotting showed that expression of RACGAP1 and phosphorylated ERK was higher in RACGAP1P 3’UTR-expressing tumours than control tumours (Fig. 4i). Importantly, the cancer cell proliferation and migration promotion effects were abrogated in DICER1-knock out HCC cells (Fig. 5a–d, Supplementary Fig. 1b), supporting the notion that the 3’UTR of RACGAP1P requires mature miRNAs for its function towards RACGAP1. Taken together, our results demonstrate that RACGAP1P, operating as a ceRNA, activates RACGAP1/RhoA/ERK pathway to favour malignant phenotypes of HCC (Fig. 5e).
a RACGAP1 expression, RhoA and ERK activation were measured by WB after RACGAP1P-3’UTR overexpression in Hep3B and HCCLM3 cells. b MTT assays shows that RACGAP1P-3’UTR overexpression promoted HCC cells growth. c Colony formation assays shows that RACGAP1P-3’UTR overexpression enhanced the proliferation of HCC cancer cells. Left: Representative images of colony formation. Right: Statistical results of colony number. d Transwell assays indicates that RACGAP1P-3’UTR overexpression augmented the migration ability of HCC cells. Left: Representative images of migrated cells. Right: statistical results of migrated cells. e–f Subcutaneous tumour growth in mice inoculated with RACGAP1P-3’UTR overexpression Hep3B cells or or control cells. e Photos of xenograft tumours, (f) growth curve of tumours, and (g) statistical analysis of tumour weight growth over 6 weeks. h QPCR reslults show that higher mRNA level of RACGAP1P and RACGAP1 in RACGAP1P-3’UTR overexpression xenograft tumours. i Western bloting indicates that the protein level of RACGAP1 and p-ERK were higher in RACGAP1P-3’UTR overexpression xenograft tumours. Error bars represent means ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, the Student’s t-test
a RACGAP1 expression, RhoA and ERK activation were detected by immunoblotting after RACGAP1P-3’UTR overexpression treatment in HCC cells with DICER1 knockout. b–d The cell growth and migration was inhibited in RACGAP1P overexpression cells with DICER1 depletion. b MTT assays. c Colony formation assays. d Transwell assays. e Diagram depicting the underlying mechanism of RACGAP1P contributing to HCC recurrence. RACGAP1P activates RhoA/ERK signalling axis by modulating RACGAP1P/miR-15a/RACGAP1 ceRNA network
RACGAP1P crosstalks with RACGAP1 and associates with poor prognosis in various human cancers
To figure out whether the ceRNA regulation between RACGAP1P and RACGAP1 is HCC specific or common in pan-cancers, the analysis of data from TCGA was performed and found the highly positive correlation between RACGAP1P and RACGAP1 in various types of cancer (Supplementary Fig. 2). In addition, exogenous overexpression of RACGAP1P 3’UTR in different human cancer cell lines could induce RACGAP1 expression showed by QPCR results (Fig. 6a). Moreover, the mRNA level of RACGAP1P was significantly upregulated in a great many human cancers compared with the normal tissues (Fig. 6b). Further, a high level of RACGAP1P indicated the shorter overall survival in pan-cancers, such as BRCA, LUAD, LGG, LAML, HNSC, PAAD (Fig. 6c). Thus, high RACGAP1P level in human cancers significantly correlated with poor outcome.
a QPCR analysis indicates that overexpression of RACGAP1P 3’UTR induced the expression of RACGAP1 in various human cancer cell lines. b The mRNA Level of RACGAP1P is significantly upregulated in 8 types of human cancer. c High expression of RACGAP1P was related to worse overall survival (OS) in 6 types of cancer. BLCA bladder urothelial carcinoma, BRCA breast invasive carcinoma, CESC cervical squamous cell carcinoma and endocervical adenocarcinoma, COAD colon adenocarcinoma, HNSC head and neck squamous cell carcinoma, KIRC kidney renal clear cell carcinoma, LUAD lung adenocarcinoma, LUSC lung squamous cell carcinoma, PAAD pancreatic adenocarcinoma, PRAD prostate adenocarcinoma, LGG low grade glima, AML acute myeloid leukaemia. Error bars represent means ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, the p values were calculated by Student’s t-test or long-rank test
Discussion
Early recurrence is the major obstacle in achieving long-term survival outcomes for the treatment of HCC via operative treatment. Our knowledge of the underlying cellular and molecular pathways driving HCC early relapse is rather finite. Previous studies have addressed the clinical implication of gene expression profiling in predicting the early recurrence of human HCC29. However, there is pretty limited information about ncRNA, especially pseudogenes involved in HCC early relapse. In the present study, we identified a novel liver cancer early recurrence associated lncRNA RACGAP1P, which is significantly upregulated in liver cancer tissues and mainly expressed much higher in recurrent HCC. Higher expression of RACGAP1P was correlated with late T stage, advanced clinical stage and abnormal AFP level. Moreover, increased RACGAP1P expression was associated with shorter OS and PFS of HCC patients. In vitro and in vivo assays demonstrated that ectogenic expression of RACGAP1P promoted cell proliferation and migration. These findings indicate that RACGAP1P has an oncogenic role in liver tumourigenesis and could be considered as a potential prognostic indicator for HCC.
RACGAP1, a member of GTPase activation protein family30, is the first gene reported to work as an independent informative biomarker for HCC early recurrent, as well as its silencing mainly affected genes in an interactome clinically relevant to early relapse. Given the significant effects of RACGAP1 on cancer recurrence, the regulatory network related to RACGAP1 urgently needed to be explored. Genetic or epigenetic alterations may directly lead to the aberrant expression of cancer-related genes. Notably, data analysing of the TCGA data found that the RACGAP1 locus undergoes copy number gains in 13.2% liver cancers and the methylation level in the promoter region is lower in cancer tissues than normal tissues and negatively associated with RACGAP1 mRNA level. However, the copy number and the methylation level were no obvious difference between recurrent and non-recurrent HCC (Supplementary Fig. 3). Therefore, the regulatory mechanisms of increasing expression of RACGAP1 in early recurrent HCC still remained to be elucidated. Herein, it was interesting to observe that the expression of pseudogene RACGAP1P significantly correlated with its cognate gene RACGAP1 expression and overexpression of RACGAP1P could increase the expression of RACGAP1 dependent on miRNA. Moreover, RACGAP1P is consistently upregulated in early recurrent HCC samples. All above suggest RACGAP1P may involve in the elevation of RACGAP1 during HCC early recurrence.
In recent years, ceRNA hypothesis provides a new crosstalk dimension between RNA moleculars that serve as miRNA sponges to regulate the level of the genes which harbour the common miRNA binding sites. Given the high sequence similarity between pseudogenes and their parent genes, pseudogenes are “perfect decoys” for their ancestral genes, critically impacting their parent protein-coding genes via the ceRNA network31,32. Based on the central of ceRNA hypothesis that miRNAs play as communication language, we identified miR-15a-5p, which is a well-known tumour suppressor in the pathogenesis of HCC, could target both RACGAP1P and RACGAP1. And the direct interaction between miR-15a-5p and these paired genes was confirmed by dual luciferase reporter assay and MS2-RIP assay, indicating that RACGAP1P regulates RACGAP1 expression at least partially through competitive binding with miR-15a-5p as a ceRNA. Similarly, the studies by the groups of Shen et al., Sharp et al. and Stoffel et al. focused on ceRNA regulation that is mediated by a single miRNA. Importantly, ceRNA pairs in general, and gene/pseudogene pairs in particular, share numerous miRNAs and it is reported that ceRNA crosstalk is enhanced when it is mediated by more miRNAs33,34. Therefore, more shared miRNAs that being present at crosstalk-favoring levels needed to be identified.
Pseudogenes could exert critical functions in biological processes and their dyregulation may conduce to the occurrence and progression of diseases12,35. Herein, our study establishes the RACGAP1P as a potent proto-oncogene that can activate RACGAP1/Rho/ERK axis to elicit malignant phenotypes, such as cell growth and migration. Mechanistically, RACGAP1P works as a ceRNA to compete with miR-15a-5p for binding to the 3’UTR region to relieve the inhibitory effects of miR-15a-5p on RACGAP1 expression, whereas MRE mutations impair the regulation of RACGAP1 by RCGAP1P. These findings provide strong evidence in support of a key role for RACGAP1P in the tumourigenesis of liver cancer. It is possible that RACGAP1P elicits its effects through more than one mechanism or pathway, so whether the oncogenic activity of RACGAP1P also requires additional ceRNA targets or non-ceRNA-related mechanisms will be the focus of future studies.
References
Ulahannan, S. V. et al. Earlier presentation and application of curative treatments in hepatocellular carcinoma. Hepatology 60, 1637–1644 (2014).
Li, W. et al. Accumulation of the mutations in basal core promoter of hepatitis B virus subgenotype C1 increase the risk of hepatocellular carcinoma in Southern China. Int. J Clin. Exp. Pathol. 6, 1076–1085 (2013).
Hoshida, Y. Risk of recurrence in hepatitis B-related hepatocellular carcinoma: impact of viral load in late recurrence. J. Hepatol. 51, 842–844 (2009).
El-Serag, H. B. Hepatocellular carcinoma. N. Engl. J. Med. 365, 1118–1127 (2011).
Feng, G. S. Conflicting roles of molecules in hepatocarcinogenesis: paradigm or paradox. Cancer Cell 21, 150–154 (2012).
Wang, S. M., Ooi, L. L. & Hui, K. M. Upregulation of Rac GTPase-activating protein 1 is significantly associated with the early recurrence of human hepatocellular carcinoma. Clin. Cancer Res. 17, 6040–6051 (2011).
Chen, J. et al. ECT2 regulates the Rho/ERK signalling axis to promote early recurrence in human hepatocellular carcinoma. J. Hepatol. 62, 1287–1295 (2015).
Chen, J. et al. The microtubule-associated protein PRC1 promotes early recurrence of hepatocellular carcinoma in association with the Wnt/beta-catenin signalling pathway. Gut 65, 1522–1534 (2016).
Balakirev, E. S. & Ayala, F. J. Pseudogenes: are they “junk” or functional DNA? Ann. Rev. Genet. 37, 123–151 (2003).
Li, W., Yang, W. & Wang, X. J. Pseudogenes: pseudo or real functional elements? J. Genet. Genomics 40, 171–177 (2013).
Han, L. et al. The Pan-Cancer analysis of pseudogene expression reveals biologically and clinically relevant tumour subtypes. Nat. Commun. 5, 3963 (2014).
Cooke, S. L. et al. Processed pseudogenes acquired somatically during cancer development. Nat. Commun. 5, 3644 (2014).
Wang, L. et al. Pseudogene OCT4-pg4 functions as a natural micro RNA sponge to regulate OCT4 expression by competing for miR-145 in hepatocellular carcinoma. Carcinogenesis 34, 1773–1781 (2013).
Peng, H. et al. Pseudogene INTS6P1 regulates its cognate gene INTS6 through competitive binding of miR-17-5p in hepatocellular carcinoma. Oncotarget 6, 5666–5677 (2015).
Kong, Y. et al. Pseudogene PDIA3P1 promotes cell proliferation, migration and invasion, and suppresses apoptosis in hepatocellular carcinoma by regulating the p53 pathway. Cancer Lett. 407, 76–83 (2017).
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3’ UTRs via Alu elements. Nature 470, 284–288 (2011).
Chaisaingmongkol, J. et al. Common molecular subtypes among asian hepatocellular carcinoma and cholangiocarcinoma. Cancer Cell 32, 57–70 e53 (2017).
Makowska, Z. et al. Gene expression analysis of biopsy samples reveals critical limitations of transcriptome-based molecular classifications of hepatocellular carcinoma. J. Pathol. Clin. Res. 2, 80–92 (2016).
Villa, E. et al. Neoangiogenesis-related genes are hallmarks of fast-growing hepatocellular carcinomas and worst survival. Results from a prospective study. Gut 65, 861–869 (2016).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Karreth, F. A. & Pandolfi, P. P. ceRNA cross-talk in cancer: when ce-bling rivalries go awry. Cancer Discov. 3, 1113–1121 (2013).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–352 (2014).
Chang, T. H. et al. An enhanced computational platform for investigating the roles of regulatory RNA and for identifying functional RNA motifs. BMC Bioinformatics 14(Suppl 2), S4 (2013).
Wang, D., Gu, J., Wang, T. & Ding, Z. OncomiRDB: a database for the experimentally verified oncogenic and tumour-suppressive microRNAs. Bioinformatics 30, 2237–2238 (2014).
Mi, S. et al. RNA-seq identification of RACGAP1 as a metastatic driver in uterine carcinosarcoma. Clin. Cancer Res. 22, 4676–4686 (2016).
Imaoka, H. et al. RacGAP1 expression, increasing tumour malignant potential, as a predictive biomarker for lymph node metastasis and poor prognosis in colorectal cancer. Carcinogenesis 36, 346–354 (2015).
Saigusa, S. et al. Clinical significance of RacGAP1 expression at the invasive front of gastric cancer. Gastric Cancer 18, 84–92 (2015).
Ke, H. L. et al. Expression of RACGAP1 in high grade meningiomas: a potential role in cancer progression. J. Neuro-oncol. 113, 327–332 (2013).
Wang, S. M., Ooi, L. L. & Hui, K. M. Identification and validation of a novel gene signature associated with the recurrence of human hepatocellular carcinoma. Clin. Cancer Res. 13, 6275–6283 (2007).
Toure, A. et al. MgcRacGAP, a new human GTPase-activating protein for Rac and Cdc42 similar to Drosophila rotundRacGAP gene product, is expressed in male germ cells. J. Biol. Chem. 273, 6019–6023 (1998).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Karreth, F. A. et al. The BRAF pseudogene functions as a competitive endogenous RNA and induces lymphoma in vivo. Cell 161, 319–332 (2015).
Bosson, A. D., Zamudio, J. R. & Sharp, P. A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 56, 347–359 (2014).
Denzler, R., Agarwal, V., Stefano, J., Bartel, D. P. & Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 54, 766–776 (2014).
Pink, R. C. & Carter, D. R. Pseudogenes as regulators of biological function. Essays Biochem. 54, 103–112 (2013).
Acknowledgements
This work was supported by grants from National Natural Science Foundation of China (No. 81702690), from Guangzhou Key Medical Discipline Construction Project fund, from Research fund for the Doctoral Program of Guangzhou Medical University (No. 2016C38). We apologise to all colleagues whose work could not be cited due to space constraints. We thank Prof. Yun-Fei Yuan for kindly providing HCC cancer cell lines.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Ciliberto
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, MY., Chen, DP., Qi, B. et al. Pseudogene RACGAP1P activates RACGAP1/Rho/ERK signalling axis as a competing endogenous RNA to promote hepatocellular carcinoma early recurrence. Cell Death Dis 10, 426 (2019). https://doi.org/10.1038/s41419-019-1666-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1666-2
This article is cited by
-
RACGAP1 promotes the progression and poor prognosis of lung adenocarcinoma through its effects on the cell cycle and tumor stemness
BMC Cancer (2024)
-
RHO GTPase family in hepatocellular carcinoma
Experimental Hematology & Oncology (2022)
-
EGF-induced nuclear translocation of SHCBP1 promotes bladder cancer progression through inhibiting RACGAP1-mediated RAC1 inactivation
Cell Death & Disease (2022)
-
A convolutional neural network model for survival prediction based on prognosis-related cascaded Wx feature selection
Laboratory Investigation (2022)
-
The emerging roles of non-coding competing endogenous RNA in hepatocellular carcinoma
Cancer Cell International (2020)