Abstract
Wolfram syndrome (WS) is a rare neurodegenerative disease, the main pathological hallmarks of which associate with diabetes, optic atrophy, and deafness. Other symptoms may be identified in some but not all patients. Prognosis is poor, with death occurring around 35 years of age. To date, no treatment is available. WS was first described as a mitochondriopathy. However, the localization of the protein on the endoplasmic reticulum (ER) membrane challenged this hypothesis. ER contacts mitochondria to ensure effective Ca2+ transfer, lipids transfer, and apoptosis within stabilized and functionalized microdomains, termed “mitochondria-associated ER membranes” (MAMs). Two types of WS are characterized so far and Wolfram syndrome type 2 is due to mutation in CISD2, a protein mostly expressed in MAMs. The aim of the present review is to collect evidences showing that WS is indeed a mitochondriopathy, with established MAM dysfunction, and thus share commonalities with several neurodegenerative diseases, including Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis, as well as metabolic diseases, such as diabetes.
Similar content being viewed by others
Facts
-
Wolfram syndrome is a rare neurodegenerative disease.
-
Wolfram syndrome symptoms looks like mitochondriopathy.
-
MAMs are key players in neurodegenerative diseases.
-
Two types of Wolfram syndrome are described.
-
WFS1, responsible for Wolfram syndrome type 1, is a transmembrane protein that regulates Ca2+ homeostasis.
-
CISD2, responsible for Wolfram syndrome type 2, is involved in Ca2+ homeostasis through MAMs.
Open questions
-
How an ER protein (WFS1) may have an essential role in mitochondrial physiology?
-
What are the interacting partners of CISD2 and WFS1 in MAMs?
-
Do WFS1 and CISD2 share a common signaling pathway?
-
Does MAM dysregulation share common pathways in neurodegenerative diseases?
Physiopathology of the Wolfram syndrome (WS): WS1, WS2, and WS-like syndrome
The WS is a rare multi-systemic genetic disease characterized by devastating clinical symptoms (Table 1). WS generally associates with diabetes insipidus, diabetes mellitus, optic atrophy, and deafness—the disease being accordingly known as DIDMOAD1. It can also provoke ataxia and other neurological symptoms2, renal and vesical dysfunctions3, and psychiatric outcomes4. The prognosis of the syndrome is poor as most patients die prematurely with severe neurological disabilities, including bulbar dysfunction and organic brain syndrome5. The natural history of WS shows diabetes mellitus during the first decade of life together with progressive optic atrophy. Deafness, neuropathic bladder, and diabetes insipidus appear during the second decade. The median age of death for patients is around 35 years and death occurs usually from respiratory failure, as a result of brain stem atrophy, or from complications of urinary tract atony5.
Clinically, patients with WS have benefited, up to now, essentially from symptomatic or substitutive therapies targeting the diabetes mellitus or diabetes insipidus. However, identification of pathological molecular mechanisms has stimulated new approaches, and two clinical trials are currently initiated. They both target discrete endpoints of WFS1 deficiency, directly associated with cell death. First, Valproate is tested and expected to oppose the downregulation of p21cip (T. Barrett, personal communication). Indeed, Gharanei et al.6 analyzed WFS1 role in secretory granules from human neuroblastoma cells and showed that cell cycle assays showed reduced p21cip protein levels in WFS1-depleted cells6. Moreover, an inverse association was measured between p21cip expression and apoptosis6. Second, the ryanodine receptor antagonist Dantrolene (ClinicalTrials.gov Identifier: NCT02829268; F. Urano, personal communication) is expected to counteract calcium leakage from the endoplasmic reticulum (ER).
WS is an autosomal-recessive genetic disease and the causative gene is WFS1, encoding for the Wolframin (WFS1) protein7,8. WFS1 is involved in the regulation of ER calcium homeostasis9. ER serves as a cellular calcium store and quality control system for identifying abnormally conformed proteins and targeting them to degradation. In case of pathological accumulation of aberrant proteins, the ER initiates a stress response, termed unfolded protein response (UPR)10. Pancreatic β-cell death and neuronal cell dysfunction in WS are indeed considered to be due to high levels of ER stress in affected cells11,12,13. WFS1 is therefore a component of UPR and its deficiency, due to chronic ER stress, leads to apoptosis in neuronal and pancreatic β-cells.
Other genetic disorders can be related to wolframin mutations. Mutations in WFS1 are not only found in WS with its autosomal-recessive inheritance but also in a variety of autosomal-dominant conditions. DFNA6/14/38 (OMIM #600965) is characterized by non-syndromic low-frequency hearing loss14,15,16,17,18,19,20,21,22. The Wolfram-like syndrome (OMIM #614296) is characterized by progressive hearing loss, optic atrophy, and/or impaired glucose regulation23,24,25,26,27. An example of Wolfram-like syndrome is a condition driven by the E864K missense mutation in exon-8 (c.2590G→A). First reported in 200623, Wolfram-like syndrome provokes a low-frequency sensorineural hearing loss, optic atrophy, and diabetes. Deafness presents a juvenile onset, but optic atrophy can appear at later ages. Some of these patients develop psychiatric complications as well23,28,29,30. Furthermore, WFS1 mutations are also responsible for rare cases of non-syndromic autosomal-dominant diabetes31,32, autosomal-dominant diabetes, and congenital hearing loss30 or autosomal-dominant congenital cataract33. Finally, as reported by Grenier et al.34, some patients with isolated autosomal-recessive non-syndromic optic atrophy have bi-allelic mutations in WFS1, like WS patients. In conclusion, recessive or dominant mutations in WFS1 consistently lead to neuronal and/or endocrine dysfunctions.
Wolfram syndrome type 2 (WS2, OMIM #604928) is a disorder caused by mutations in the CISD2 gene. It encodes for miner 1 ER-membrane-localized zinc finger protein that regulates UPR, Ca2+ homeostasis, and autophagy35. In WS2, symptoms other than the characteristic optic atrophy are a high-frequency sensorineural hearing loss and diabetes mellitus, with an early onset and autosomal-recessive inheritance as observed in WS1. However, patients do not develop diabetes insipidus29. Other dysfunctions are also present but varying from one patient to another.
Both WS1 and WS2 syndromes, even being not directly related to mitochondrial malfunction, are caused by imbalance of Ca2+ homeostasis originating from the ER and therefore incorporate a secondary mitochondrial aspect.
ER stress in physiological and pathological conditions
When the ER is stressed, it triggers the UPR adaptive response. UPR will lead to overexpression of specific ER proteins—including protein disulfide isomerase, lectin, and oxydoreductase—that prevent accumulation of stress-induced unfolded proteins and restore ER homeostasis. Three ER-resident transmembrane proteins function as stress sensors: RNA-activated protein kinase-like endoplasmic reticular kinase (PERK), activating transcription factor 6 (ATF6), and inositol-requiring kinase 1 (IRE1). Their activations transduce the unfolded protein stress signal across ER membrane and lead to UPR activation36. Activation of the PERK pathway leads to attenuation of general protein translation by phosphorylation of the α subunit of eukaryotic translation initiation factor 2 (eIF2α)37. Phosphorylated eIF2α can selectively enhance the translation of mRNAs containing inhibitory upstream open reading frames in their 5′ untranslated region, such as ATF438. In addition, under ER stress, ATF6 acts as an active transcription factor by translocating to the Golgi complex, where it is cleaved by site-1 and site-2 proteases39. The active cleaved form of ATF6 then translocates into the nucleus and binds to the promoter of UPR-inducible genes, resulting in an upregulation of proteins, the role of which is to adjust ER protein folding, including ER chaperones and X-box-binding protein-1 (XBP-1)40. IRE1 acts as an endoribonuclease and its activation facilitates the unconventional splicing of XBP-1 mRNA and subsequent translation of an active transcription factor36,40. This latter promotes the expression of ER-resident chaperones, which facilitate protein folding in the ER36,40. If these adaptive coordinated responses can not eliminate inappropriately folded proteins during prolonged and severe ER stress, the UPR elicits a pro-apoptotic pathway triggering apoptotic cell death41.
ER stress is implicated in numerous pathologies. It is involved, for instance, in cancer41; in diabetes42, in cardiomyopathy43, and in neurological disorders44,45. In this review, we will focus on neurological disorders. In Alzheimer’s diseases (AD), the expression level of BiP is increased in the hippocampus and temporal cortex of patients46,47. Moreover, phosphorylation of IRE1 in AD brain tissues48 and PERK and its main target eIF2α have been detected in hippocampal structure where they colocalized with abnormal hyperphosphorylated Tau, a hallmark of AD49. In Parkinson’s disease (PD), an increased expression level of BiP was alo shown in postmortem nigral dopaminergic neurons50. Moreover, α-synuclein aggregation activated the UPR-related activating transcription factor 4/cAMP-responsive element-2. These findings suggest that activation of the UPR pathway in the PD brain is associated with α-synuclein accumulation. In amyotrophic lateral sclerosis (ALS), the expression level of the three major components of the UPR, PERK, IRE1, and ATF6 is increased in the spinal cord of patients51,52,53,54. Finally, in Huntington’s disease (HD), mutant huntingtin affects the normal function of ER-associated degradation (ERAD) system in PC12 cells55. Impaired ERAD leads to accumulation of misfolded proteins in the ER56. In WS, WFS1 inhibits the UPR by targeting ATF6 for degradation by the proteasome in vitro10. In the retina, WFS1 deficiency leads to an increased in the protein expression level of BiP, PDI, and IRE111. ER stress-mediated cell death may be triggered by ER membrane permeabilization. In brain tissues from WFS1 knockout (KO) mice, more ER proteins were found in the cytosol, suggesting an ER permeabilization13. Finally, dominant mutation of WFS1 induced the expression of ER stress response12. Taken together, these data highlighted the essential role of the ER stress and UPR in most of the neurodegenerative disorders and suggested that these debilitating pathologies may share common physiopathological signaling pathways.
Interestingly, a substantial number of proteins involved in UPR are localized in mitochondria-associated ER membranes (MAMs)57. Mitofusin 2 (MFN2), a dynamin-like GTPase localized in the outer mitochondrial and ER membranes, modulates ER homeostasis since its deficiency leads to ER stress in vitro and in vivo58,59,60. Some ER chaperones involved in UPR are enriched in MAMs. The sigma-1 protein (S1R), for instance, binds BiP and inositol 1,4,5-trisphosphate receptor channel (IP3R)61. Upon ER Ca2+ depletion or via ligand stimulation, S1R dissociates from IP3R, leading to a prolonged Ca2+ signaling in to mitochondria via IP3R62,63. At the integrated level, S1R has been shown to be implicated in neuroprotection and neuroplasticity61,64. In addition, Calnexin, a type I integral membrane protein that helps in folding newly synthesized proteins is essential in mitigating ER stress65. Finally, two major proteins involved in UPR, PERK66, and IRE164,67 are enriched in MAMs. A more detailed description of the role of these proteins in MAM physiology is presented below.
MAMs: structure and function
Mitochondria are the powerhouse of cells in the organism. They play essential function in generating energetic metabolism, Ca2+ homeostasis, lipid synthesis, and apoptosis. To achieve these functions properly, mitochondria need to be spatially and temporally controlled. Mitochondria could make contact with different organelles in the cell, including peroxisomes, lysosomes, or the ER68. Mitochondria interact with peroxisomes to assure β-oxidation69, to eliminate reactive oxygen species70, to insure peroxisome membrane dynamics71,72, and to cooperate in viral combat73,74. Close contacts between mitochondria and lysosomes are necessary for autophagy75. Finally, mitochondria interaction with the ER is involved in lipid homeostasis76, UPR57, and Ca2+ transfer between the two organelles77.
Interaction domains between mitochondria and ER, called MAMs78,79, are dynamic structures sequestering more than a thousand different proteins80,81 that are necessary for structurally stabilizing MAMs and for the functional dialog between ER and mitochondria. Table 2 summarizes the most important proteins involved in MAM biology.
Proteins that play a role in MAMs' structure
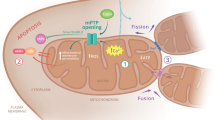
In MAMs, the distance between ER and mitochondria should be maintained between 10 nm and 30 nm, in order to allow efficient protein interactions and focused Ca2+ exchange82. Some proteins are involved in the tethering—by increasing contact site formation— or spacing—by increasing the distance between ER and mitochondria—of ER and mitochondrial membranes. One of the most characterized protein involved in MAMs' formation is MFN2. MFN2 homo-dimerizes or hetero-dimerizes with MFN1, another dynamin-like GTPase of the outer mitochondrial membrane, bridging ER and mitochondria83,84,85,86,87 (Fig. 1). The exact function of MFN2 as both a tether and spacer is still a matter of debate since both roles have been demonstrated in different experiments. For instance, downregulation or ablation of MFN2 provoked a decrease83,84 or an increase85,86,87 in ER–mitochondria contact sites. The mitochondrial ubiquitin protein ligase (MITOL) also regulates mitochondrial dynamics. Interestingly, MITOL binds to and regulates MFN2 in the mitochondria but not in the ER88. MITOL-induced ubiquitination leads to oligomerization of MFN2 and to the tethering of MAMs.
Close interaction between ER and mitochondria are necessary for a plethora of function. This peculiar microdomain is called mitochondrial-associated membranes (MAMs). The structure of the MAMs is tightly controlled by the interaction of MFN2/MFN1/2, FIS1/BAP31, PTPIP51/VAPB, and EMERIN-FATE1-MITOFILIN. The truncated form of SERCA1, S1T, PDZD8, TpM, and PERK may also participate in MAM tethering. MITOL and PACS2 influence MAM's structure by interacting with MFN2 and BAP31, respectively. The apposition of ER to mitochondria allows the passage of Ca2+ from the ER lumen to the mitochondria through the tripartite complex, IP3R (the ER IP3-sensitive Ca2+ channel), GRP75 (a cytoplasmic chaperone), and VDAC (the OMM Ca2+ channel). This transfer may be modulated by S1R, BiP, calnexin, and PML, for instance. The entrance of Ca2+ into the mitochondrial matrix occurs via MCU (the mitochondrial calcium uniporter). The Ca2+ is necessary for the correct function of the TCA cycle and for the respiratory complexes. Some proteins involved in neurodegenerative diseases are expressed in MAMs, such as HTT, α-synuclein, APOE4, and PS1-2
MAMs' structural stability is also permitted by direct association of vesicle-associated membrane protein-associated protein B (VAPB), on the ER membrane, and protein tyrosine phosphatase interacting protein 51 (PTPIP51), on the outer mitochondrial membrane (Fig. 1). VAPB–PTPIP51 interaction fosters ER–mitochondria contact sites to regulate Ca2+ homeostasis89 and autophagy90. This interaction has been shown to be specifically disrupted in ALS91 and PD92, outlining the essential role of MAMs in neurodegenerative diseases (NDs). A last complex of proteins potentially involved in MAMs' tethering is the bridge between integral ER membrane protein (Bap31) and mitochondrial fission protein 1 (Fis1) in the outer mitochondrial membrane (Fig. 1). Fis1 triggers an apoptotic signal from mitochondria to the ER by interacting with Bap31 and provoking its cleavage into the pro-apoptotic p20Bap31 fragment93. Moreover, another signaling protein, phosphofurin acidic cluster sorting protein 2 (PACS2), known to regulate ER–mitochondria communication, ER homeostasis, and apoptosis, may control the apposition of mitochondria along the ER (Fig. 1). PACS2 downregulation increased the distance between ER and mitochondria and triggered BAP31-dependent mitochondria fragmentation and uncoupling from the ER94. In contrast, PACS2 overexpression has been suggested to be responsible for increased contacts between ER and mitochondria in hippocampal neurons from a mouse model of AD95.
Chami et al.96 described the particular role of a truncated form of sarco/endoplasmic reticulum Ca2+-ATPase type 1 (SERCA1), called S1T, in mitochondrial dynamics (Fig. 1). Normal SERCA1 protein contains 10 transmembrane domains, whereas S1T contains only transmembrane domains 1–4 and is not able to pump Ca2+. S1T favored ER Ca2+ depletion due to increased Ca2+ leak, increased the number of ER-mitochondria contact sites, decreased the distance between ER and mitochondria, and inhibited mitochondrial dynamics. Taken together, the data suggested that S1T is a MAM protein that controls tethering of ER to mitochondria in a Ca2+-dependent manner96. The exact mechanism by which S1T modulates the tethering of ER to mitochondria is not fully understood, but it is tempting to speculate that S1T interacts with a not yet identified outer mitochondrial membrane protein that would efficiently impact the distance between ER and mitochondria. Therefore, S1T might be considered as a novel MAM structural protein. In opposition to S1T, overexpression of fetal and adult testis-expressed transcript protein homolog increased the distance between ER and mitochondria97, by interacting with Mitofilin, on the mitochondrial side, and Emerin, on the ER side (Fig. 1). Increased MAM thickness reduced mitochondrial Ca2+ uptake and induced apoptosis97.
In yeast, contact between ER and mitochondria is controlled by a macrocomplex named ERMES98 and no functional ortholog of any ERMES proteins have been identified in mammals. Very recently, Hirabayashi et al.99 identified PDZD8 as a novel ER-resident protein expressed at the ER–mitochondria interface (Fig. 1). PDZD8 contains an SMP domain functionally orthologous to the SMP domain of yeast Mmm1, a component of ERMES. They generated PDZD8-KO cells and determined that the number and the size of the contact were highly reduced in PDZD8-KO cells. This decrease is associated with a reduced Ca2+ transfer from the ER to mitochondria.
Cerqua et al.100 showed that trichoplein/mitostatin (TpM) is expressed in MAMs and that is essential for the ER–mitochondria tethering. TpM is a keratin-binding protein that colocalizes with mitochondria101 (Fig. 1). The protein is downregulated in various cancer-derived cells and in solid tumors. Indeed, when TpM is downregulated by short hairpin RNA (shRNA), the tethering is increased, whereas when TpM is overexpressed, the tethering is decreased100. Moreover, mitochondrial morphology is dependent on the expression level of TpM, with a higher proportion of elongated mitochondria when TpM is downregulated.
Finally, PERK, a key player in the UPR102 is also localized to the MAMs66 (Fig. 1). Verfaillie et al.66 demonstrated that PERK−/− mouse embryonic fibroblasts (MEFs) showed altered ER morphology and Ca2+ signaling as well as decreased ER–mitochondria contact sites. Indeed, in PERK−/− MEFs, the fraction of mitochondria overlapping ER is decreased. Interestingly, overexpression of a PERK dead mutant restored the contact sites, whereas overexpression of a truncated C-ter cytoplasmic PERK did not. These data showed that cytoplasmic domain of PERK is essential for the ER–mitochondria tethering but not its kinase activity.
Proteins that play a role in MAMs' function
One of the most important role of MAMs is therefore to allow direct Ca2+ transfer between ER and mitochondria and this is mainly allowed by the ER transmembrane IP3R (Fig. 1). The ER is the major Ca2+ storage organelle within the cell103, with a steady-state Ca2+ concentration in the ER, [Ca2+]er, of approximately 1 mM. At resting state, Ca2+ concentration in the cytosol, [Ca2+]c, is maintained at 100 nM. Ca2+ efflux from the ER contributes rapidly and efficiently to a rising in [Ca2+]c. The juxtaposition, in close contacts, of ER and mitochondria allows focused Ca2+ entry into the mitochondria. A dynamic transfer should be tightly regulated in order to avoid Ca2+ overload and consequent adverse effect triggering apoptosis104,105. Under physiological conditions, Ca2+ originating from the ER accumulates into the mitochondrial matrix and modulates Ca2+-sensitive dehydrogenases of the tricarboxylic acid cycle106 and metabolite carriers107, stimulating oxidative metabolism. After being released by the ER, Ca2+ is taken up by the mitochondria through the outer mitochondrial transmembrane voltage-dependent anion channel (VDAC). Among the three isoforms108, VDAC1 is physically linked to IP3R through the Hsp70 family chaperone GRP75, optimizing Ca2+ transfer from IP3R to mitochondria (Fig. 1). Indeed, downregulation of GRP75 impaired IP3R-mediated Ca2+ transfer into mitochondria109. The complex is, however, is regulated by several partner proteins.
The promyelocytic leukemia (PML) tumor suppressor is a modulator of apoptosis110. PML is primarily localized in the nucleus but Giorgi et al.111 detected a fraction of the protein in MAMs (Fig. 1). Since MAM is the site of Ca2+ transfer between ER and mitochondria, they measured Ca2+ concentration in ER, cytoplasm, and mitochondria and they showed a decrease in all compartments. To determine whether these anomalies were due to the fraction of PML expressed in the MAMs, they overexpressed a chimeric PML targeted to the outer surface of the ER. Using this approach, they elegantly demonstrated that the ER-expressed PML is necessary for a normal Ca2+ transfer between ER and mitochondria111.
MAM dysfunction is a common trait in neurodegenerative pathologies
Recently, numerous evidences accumulated suggesting that MAM dysfunction contributes to the neurodegenerative processes in AD, PD, ALS, or HD112,113,114. In AD, both presenilin-1 and presenilin-2—the two major components of the γ-secretase complex that processes amyloid precursor protein (APP) to release amyloid-β proteins (Aβ) and that can be mutated in familial forms of AD—are present in MAMs115 (Fig. 1). MAMs are a site of production of Aβ and this is consistent with the localization of presenilins in these regions116,117,118. Moreover, mutations of presenilins are a cause of familial forms of AD with early onset and mutant presenilins are catalytic loss-of-function mutants119. Both loss of presenilins and expression of mutant presenilins have been shown to affect ER–mitochondria associations and related functions116. Moreover, MAM are particularly sensitive to the neurodegenerative process since treatment of neurons with Aβ affects ER–mitochondria contacts; alterations of ER–mitochondria association and function are seen in APP transgenic mouse models; and small interfering RNA knockdown of MAM proteins (S1R, phosphofurin acidic cluster sorting protein-2) results in neurodegeneration while MAM proteins are upregulated in AD mouse models95. Finally, the ε4 allele of apolipoprotein E—ApoE4, the main genetic risk factor for AD—upregulates MAM activity120.
In PD, the neurodegenerative process affecting dopaminergic neurons from the nigro-striatal pathway is characterized by accumulation of pathological α-synuclein protein. A subpopulation of α-synuclein resides at the MAM121 (Fig. 1) and mutations in α-synuclein cause an alteration in the regulation of MAM function121.
In ALS, an hyper-phosphorylated, ubiquitinated, and cleaved form of transactive response DNA-binding protein 43 kDa (TDP-43) is the major pathological protein in frontotemporal dementia and ALS122. Pathological TDP-43 induces activation of glycogen synthase kinase-3β and perturbs ER–mitochondria associations by impacting VAPB–PTPIP51 bridges91 (Fig. 1). TDP-43 downregulates MFN levels in Drosophila (J.C. Lievens, personal communication) and mouse models. Decreased MFN1/MFN2 levels are also reported in ALS patient biopsies and in a mouse model expressing wild-type TDP-43123,124. Moreover, a mutation of the MAM protein S1R may be responsible for familial ALS cases125,126 (Fig. 1). Loss of S1R leads to motor neuron degeneration in vitro127.
Alterations of ER–mitochondria associations may also occur in HD, but further research is required to provide stronger evidence. For instance, upregulation of striatal S1R was reported in YAC HD mice and HD patients (Ryskamp et al., Neurobiol Dis 2017), but it is unclear whether these alterations are causal mechanisms or compensatory regulations.
However, evidences are clearly accumulating showing that pathological proteins, responsible for the toxicity observed in neurodegenerative pathologies, particularly accumulate within MAM and that the concomitant/subsequent MAM alterations observed participate in the resulting toxicity.
Could WS2 also be a MAM-related pathology?
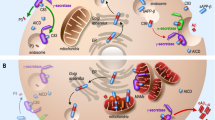
CDGSH iron-sulfur domain-containing protein 2 (CISD2, also known as Miner1, NAF-1, ERIS) was initially described as the cause of WS2 in 2007128. CISD2 is localized in the ER membrane and colocalizes with calnexin, a well-known ER chaperone128 (Fig. 2). Remarkably, ER chaperones have emerged as important proteins for MAM functions. ER chaperones are important for the folding of newly imported polypeptides129, and during the past decade, it has been shown that some of them are enriched in the MAMs. For example, S1R61, BiP61, and Calnexin130 are associated with MAM Ca2+ handling proteins to adjust Ca2+ import to or exit from the ER in order to control apoptosis and mitochondrial metabolism (see ref. 131 for a review). Surprisingly, CISD2 did not interact with WFS1128. Resting [Ca2+]c were not different between a cell line derived from an affected patient and a cell line derived from a control. In contrast, when stimulated by thapsigargin, a SERCA inhibitor, Ca2+ release was more significantly increased in the affected cell line than in the unaffected cell line128. This inhibition induces a depletion of the ER Ca2+ store thus giving an indirect measure of the ER Ca2+ content. The ER Ca2+ content in lymphoblastoid WS2 patient therefore appeared higher than that in control. This elevated [Ca2+]er might be responsible for the degeneration of β-cells and neurons since ER Ca2+ overload increases the cell susceptibility to apoptosis77,132. Similar results were obtained in fibroblasts from WS2 patients133. In addition, the number of ER–mitochondrial contacts was increased in patient fibroblasts compared to controls, as visualized using transmission electron microscopy (TEM). This observation was confirmed in living cell by analyzing the colocalization between ER, using the GFP Sec61b marker, and mitochondria, using MitoTracker133. Finally, even if no ultrastructural abnormalities could be observed in mitochondria preparations from patients, both the average length and volume of mitochondrial fragments were increased in fibroblasts from patients. The more fused and elongated mitochondrial network was associated, in a galactose medium used to force cells to rely predominantly on OXPHOS for ATP production, with a respiratory chain defect in complexes I and II of the mitochondrial respiratory chain133.
In 2009, the group of Tsai134 generated a mutant mice in which the expression of Cisd2 was abolished to study the role of Cisd2 in development and physiopathology. The mice showed a shortened lifespan probably due to a premature aging phenotype. Using TEM, they observed that the phenotype was linked to mitochondrial degeneration and autophagy. Interestingly, in contrast to the data from Amr et al.128, the expression of Cisd2 was measured in the outer mitochondrial membrane and not in the ER134. Remarkably, lack of Cisd2 in mice led to respiratory chain dysfunction, suggesting that WS2 is finally a mitochondria-related disorder134. On the contrary, mice overexpressing Cisd2 showed delayed aging and restored mitochondrial complex functionality135. Taken together, these studies demonstrated an essential role of CISD2 in mitochondrial normal function.
Cisd2 was identified as a B-cell lymphoma 2 (Bcl-2) interacting protein to regulate autophagy136, confirming the observation by Tsai’s group in their mutant mice. Bcl-2 is a well-known antiapoptotic protein that regulates the outer membrane permeabilization137. In addition to its mitochondrial localization, Bcl-2 also localized to the ER membrane (Fig. 2). This ER localization seems necessary for the inhibition of autophagy138. Indeed, autophagy, which is a major intracellular process for the degradation and recycling of proteins and cytoplasmic damaged organelles, is inhibited when Bcl-2 binds to Beclin 1138. Cisd2 binds Bcl-2 at the ER and is required for Bcl-2 to inhibit Beclin 1-mediated autophagy136. In addition, Cisd2 interacts with IP3R. This interaction seems to intervene in the depressed levels of ER Ca2+ stores following elevated Bcl-2 (Bcl-2b5) at the ER136. To extend these findings, Ca2+-sensitive ER-targeted aequorins were used to directly measure changes in luminal [Ca2+]er. The results confirmed that Bcl-2b5 required Cisd2 in order to reduce ER Ca2+ stores139. Notably, Bcl-2 interacts also with IP3R to inhibit Ca2+ release140. Taken together, all these data suggest that Cisd2, IP3R, and Bcl-2 form a macrocomplex to regulate Ca2+ signaling and MAMs' physiology141,142.
The conflicting localization of Cisd2, either in ER or outer mitochondrial membrane, was resolved by Murphy’s group in 201335. After a subcellular fractionation of ER, mitochondria, and MAM fractions from the rat liver, they observed that Cisd2 was most abundant in ER-enriched fraction and not detectable in purified mitochondria. The protein was also abundant in MAM fraction35. To address the impact of Cisd2 loss on Ca2+ homeostasis and mitochondrial activity, they used Cisd2 KO mouse embryonic cells (MEFs). Interestingly, after treatment with histamine, ER Ca2+ release was higher in Cisd2 KO than in wild-type MEFs35. Consequently, mitochondrial Ca2+ uptake was greater in Cisd2 KO than in wild-type MEFs. They concluded that Cisd2 is a key determinant in regulating not only ER but also mitochondrial Ca2+ homeostasis35. The increase of mitochondrial Ca2+ loading in Cisd2 KO cells was followed by a higher oxygen consumption rate for both maximally stimulated and basal measure conditions.
Loss of function of Cisd2 leads to neurons and β-cells death, but the exact mechanism is not fully understood. It has recently been shown that downregulation of Cisd2 in mouse neuronal NSC34 cells as well as in induced pluripotent stem cells from WS patients triggers cell death by overactivation of the calcium-dependent proapoptotic protease calpain-2. This activation seems to be due to the increase of the [Ca2+]c143. Surprisingly, the potent inhibitor of the ryanodine receptors Dantrolene, supposedly able to decrease the Ca2+ leakage from the ER to the cytosol, failed to block cell death provoked by Cisd2 knockdown143. These observations therefore suggested that Cisd2 does not directly affect ER Ca2+ homeostasis.
Cisd2 has been shown to regulate the differentiation and functioning of adipocytes. Indeed, Cisd2 deficiency increase cytosolic Ca2+ and impairs the Ca2+ buffering capability of mitochondria144. This increase is supposed to impair the in vitro differentiation of primary MEFs into adipocytes. This defect would be due to the lack interaction of Cisd2 with GTPase of the immune-associated nucleotide binding protein 5 (Gimap5) in MAMs (Fig. 2). Indeed, together, these proteins regulate mitochondrial Ca2+ influx and the maintenance of intracellular Ca2+ homeostasis. Moreover, Cisd2 deficiency activates calcineurin, which then acts as a negative regulatory effect of white adipogenesis144. Loss of function of Cisd2 is not only responsible for adipocyte differentiation but also for osteogenic differentiation145. This alteration of the osteogenic differentiation is also due to an increase in the cytosolic Ca2+ concentration.
Is MAMs' dysfunction playing a role in WS1 pathology?
The first evidence of a potential functional role of WFS1 in MAMs came from the observations that WFS1 is present in MAM fraction from human fibroblasts146, mouse brain samples81, and huh7 cells80 (Fig. 3). Moreover, reconstitution of wolframin from oocyte membranes into planar lipid bilayers was able to induce a large IP3-dependent cation-selective ion channel, blocked by Mg2+ or Ca2+147. IP3 was able to activate channels in the fused bilayers similarly as channel components induced by wolframin expression. These observations were strengthened by a recent work by Cagalinec et al. 148. Using Wfs1 downregulation or KO models, the authors described that Wfs1 deficiency in neurons led to dramatic changes in mitochondrial dynamics, with inhibited mitochondrial fusion, altered mitochondrial trafficking, and increased autophagy. Moreover, lack of Wfs1 induced ER stress, IP3R dysfunction, and disturbed [Ca2+]c homeostasis148.
Finally, WFS1 appears to be a negative regulator of SERCA2b expression in the ER (Fig. 3). Zatyka et al.149 observed that SERCA2b expression was elevated in several Wfs1-depleted cells models and primary islets. They demonstrated a novel interaction between Wfs1 and SERCA2b by co-immunoprecipitation in COS7 cells and with endogenous proteins in human neuroblastoma cells149. Using MG-132 proteasome inhibitor, they concluded that WFS1 targets SERCA2b to the proteasome for degradation. Since SERCA2b is expressed in MAMs and is a well-known effector of ER Ca2+ uptake150, Wfs1 may be a novel MAM physiological effector essential for Ca2+ homeostasis. In contrast, Morikawa et al.151 described a reduced mRNA level of SERCA2b in HEK-293 cells transfected with mutant WFS1 cDNA compared to HEK-293 cells transfected with wild-type WFS1 cDNA. This elevation of [Ca2+]cyto is associated with an increase of the mRNA level of CCAAT-enhancer-binding protein homologous protein, leading to ER stress-induced cell apoptosis152. In another study, Hara et al.153 demonstrated that downregulation of WFS1 via shRNA induced an increase in [Ca2+]cyto in β-cell. They proposed that such an increase may activate calpain-2 that will lead to β-cell death. Since no information on the protein expression level of SERCA2b was provided, more experiments are needed in order to clarify the real impact of the absence of WFS1 on SERCA2b expression and activity.
Conclusions
The aim of this review was to integrate WS as a novel neurodegenerative MAMpathy together with AD, PD, HD, and ALS112,113,114. Indeed, CISD2 has been shown to play a role in ER–mitochondria Ca2+ signaling and regulation of autophagy and CISD2 deficient leads to ER stress and apoptosis. In addition, WFS1 regulate ER Ca2+ homeostasis by controlling the expression level of SERCA2b and WFS1 deficiency leads to ER stress and cell death. Since the majority of the case of NDs is sporadic and since WS is a rare genetic disorder, WS may be useful for the understanding of MAMs in a broader context. Finally, either in classical ND or in WS, there is a defect in MAMs and the presence of ER stress. It should be interesting to determine whether these two phenomena are tightly linked or are two independent mechanisms responsible for the pathology.
References
Barrett, T. G., Bundey, S. E. & Macleod, A. F. Neurodegeneration and diabetes: UK nationwide study of Wolfram (DIDMOAD) syndrome. Lancet 346, 1458–1463 (1995).
Grosse Aldenhovel, H. B., Gallenkamp, U. & Sulemana, C. A. Juvenile onset diabetes mellitus, central diabetes insipidus and optic atrophy (Wolfram syndrome)--neurological findings and prognostic implications. Neuropediatrics 22, 103–106 (1991).
Tekgul, S., Oge, O., Simsek, E., Yordam, N. & Kendi, S. Urological manifestations of the Wolfram syndrome: observations in 14 patients. J. Urol. 161, 616–617 (1999).
Swift, R. G., Perkins, D. O., Chase, C. L., Sadler, D. B. & Swift, M. Psychiatric disorders in 36 families with Wolfram syndrome. Am. J. Psychiatry 148, 775–779 (1991).
Kinsley, B. T., Swift, M., Dumont, R. H. & Swift, R. G. Morbidity and mortality in the Wolfram syndrome. Diabetes Care 18, 1566–1570 (1995).
Gharanei, S. et al. Vacuolar-type H+-ATPase V1A subunit is a molecular partner of Wolfram syndrome 1 (WFS1) protein, which regulates its expression and stability. Hum. Mol. Genet. 22, 203–217 (2013).
Inoue, H. et al. A gene encoding a transmembrane protein is mutated in patients with diabetes mellitus and optic atrophy (Wolfram syndrome). Nat. Genet. 20, 143–148 (1998).
Strom, T. M. et al. Diabetes insipidus, diabetes mellitus, optic atrophy and deafness (DIDMOAD) caused by mutations in a novel gene (wolframin) coding for a predicted transmembrane protein. Hum. Mol. Genet. 7, 2021–2028 (1998).
Fonseca, S. G. et al. WFS1 is a novel component of the unfolded protein response and maintains homeostasis of the endoplasmic reticulum in pancreatic beta-cells. J. Biol. Chem. 280, 39609–39615 (2005).
Fonseca, S. G. et al. Wolfram syndrome 1 gene negatively regulates ER stress signaling in rodent and human cells. J. Clin. Investig. 120, 744–755 (2010).
Bonnet Wersinger, D. et al. Impairment of visual function and retinal ER stress activation in Wfs1-deficient mice. PLoS ONE 9, e97222 (2014).
De Franco, E. et al. Dominant ER stress-inducing WFS1 mutations underlie a genetic syndrome of neonatal/infancy-onset diabetes, congenital sensorineural deafness, and congenital cataracts. Diabetes 66, 2044–2053 (2017).
Kanekura, K. et al. IRE1 prevents endoplasmic reticulum membrane permeabilization and cell death under pathological conditions. Sci. Signal. 8, ra62 (2015).
Bai, X. et al. Identification of a novel missense mutation in the WFS1 gene as a cause of autosomal dominant nonsyndromic sensorineural hearing loss in all-frequencies. Am. J. Med. Genet. A 164A, 3052–3060 (2014).
Bramhall, N. F., Kallman, J. C., Verrall, A. M. & Street, V. A. A novel WFS1 mutation in a family with dominant low frequency sensorineural hearing loss with normal VEMP and EcochG findings. BMC Med. Genet. 9, 48 (2008).
Chaussenot, A. et al. Mutation update and uncommon phenotypes in a French cohort of 96 patients with WFS1-related disorders. Clin. Genet. 87, 430–439 (2015).
Goncalves, A. C. et al. WFS1 and non-syndromic low-frequency sensorineural hearing loss: a novel mutation in a Portuguese case. Gene 538, 288–291 (2014).
Kunz, J. et al. Identification of a novel mutation in WFS1 in a family affected by low-frequency hearing impairment. Mutat. Res. 525, 121–124 (2003).
Lesperance, M. M., Hall, J. W. 3rd, San Agustin, T. B. & Leal, S. M. Mutations in the Wolfram syndrome type 1 gene (WFS1) define a clinical entity of dominant low-frequency sensorineural hearing loss. Arch. Otolaryngol. Head. Neck Surg. 129, 411–420 (2003).
Noguchi, Y. et al. A mutation in Wolfram syndrome type 1 gene in a Japanese family with autosomal dominant low-frequency sensorineural hearing loss. Acta Otolaryngol. 125, 1189–1194 (2005).
Sun, Y. et al. Identification of two novel missense WFS1 mutations, H696Y and R703H, in patients with non-syndromic low-frequency sensorineural hearing loss. J. Genet. Genom. 38, 71–76 (2011).
Young, T. L. et al. Non-syndromic progressive hearing loss DFNA38 is caused by heterozygous missense mutation in the Wolfram syndrome gene WFS1. Hum. Mol. Genet. 10, 2509–2514 (2001).
Eiberg, H. et al. Autosomal dominant optic atrophy associated with hearing impairment and impaired glucose regulation caused by a missense mutation in the WFS1 gene. J. Med. Genet. 43, 435–440 (2006).
Fujikawa, T., Noguchi, Y., Ito, T., Takahashi, M. & Kitamura, K. Additional heterozygous 2507A>C mutation of WFS1 in progressive hearing loss at lower frequencies. Laryngoscope 120, 166–171 (2010).
Gurtler, N. et al. Two families with nonsyndromic low-frequency hearing loss harbor novel mutations in Wolfram syndrome gene 1. J. Mol. Med. (Berl.) 83, 553–560 (2005).
Hogewind, B. F. et al. Autosomal dominant optic neuropathy and sensorineual hearing loss associated with a novel mutation of WFS1. Mol. Vis. 16, 26–35 (2010).
Rendtorff, N. D. et al. Identification of p.A684V missense mutation in the WFS1 gene as a frequent cause of autosomal dominant optic atrophy and hearing impairment. Am. J. Med. Genet. A 155A, 1298–1313 (2011).
Fukuoka, H., Kanda, Y., Ohta, S. & Usami, S. Mutations in the WFS1 gene are a frequent cause of autosomal dominant nonsyndromic low-frequency hearing loss in Japanese. J. Hum. Genet. 52, 510–515 (2007).
Rigoli, L. & Di Bella, C. Wolfram syndrome 1 and Wolfram syndrome 2. Curr. Opin. Pediatr. 24, 512–517 (2012).
Valero, R., Bannwarth, S., Roman, S., Paquis-Flucklinger, V. & Vialettes, B. Autosomal dominant transmission of diabetes and congenital hearing impairment secondary to a missense mutation in the WFS1 gene. Diabet. Med. 25, 657–661 (2008).
Bonnycastle, L. L. et al. Autosomal dominant diabetes arising from a Wolfram syndrome 1 mutation. Diabetes 62, 3943–3950 (2013).
Zalloua, P. A. et al. WFS1 mutations are frequent monogenic causes of juvenile-onset diabetes mellitus in Lebanon. Hum. Mol. Genet. 17, 4012–4021 (2008).
Berry, V. et al. Wolfram gene (WFS1) mutation causes autosomal dominant congenital nuclear cataract in humans. Eur. J. Hum. Genet. 21, 1356–1360 (2013).
Grenier, J. et al. WFS1 in optic neuropathies: mutation findings in nonsyndromic optic atrophy and assessment of clinical severity. Ophthalmology 123, 1989–1998 (2016).
Wiley, S. E. et al. Wolfram Syndrome protein, Miner1, regulates sulphydryl redox status, the unfolded protein response, and Ca2+homeostasis. EMBO Mol. Med. 5, 904–918 (2013).
Schroder, M. & Kaufman, R. J. ER stress and the unfolded protein response. Mutat. Res. 569, 29–63 (2005).
Schroder, M. & Kaufman, R. J. Divergent roles of IRE1alpha and PERK in the unfolded protein response. Curr. Mol. Med. 6, 5–36 (2006).
Lu, P. D., Harding, H. P. & Ron, D. Translation reinitiation at alternative open reading frames regulates gene expression in an integrated stress response. J. Cell Biol. 167, 27–33 (2004).
Ye, J. et al. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 6, 1355–1364 (2000).
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891 (2001).
Urra, H., Dufey, E., Lisbona, F., Rojas-Rivera, D. & Hetz, C. When ER stress reaches a dead end. Biochim. Biophys. Acta 1833, 3507–3517 (2013).
Cnop, M., Toivonen, S., Igoillo-Esteve, M. & Salpea, P. Endoplasmic reticulum stress and eIF2alpha phosphorylation: the Achilles heel of pancreatic beta cells. Mol. Metab. 6, 1024–1039 (2017).
Groenendyk, J., Agellon, L. B. & Michalak, M. Coping with endoplasmic reticulum stress in the cardiovascular system. Annu. Rev. Physiol. 75, 49–67 (2013).
Hetz, C. & Saxena, S. ER stress and the unfolded protein response in neurodegeneration. Nat. Rev. Neurol. 13, 477–491 (2017).
Mollereau, B. et al. Adaptive preconditioning in neurological diseases - therapeutic insights from proteostatic perturbations. Brain Res. 1648, 603–616 (2016).
Hoozemans, J. J. et al. The unfolded protein response is activated in Alzheimer’s disease. Acta Neuropathol. 110, 165–172 (2005).
Stutzbach, L. D. et al. The unfolded protein response is activated in disease-affected brain regions in progressive supranuclear palsy and Alzheimer’s disease. Acta Neuropathol. Commun. 1, 31 (2013).
Duran-Aniotz, C. et al. IRE1 signaling exacerbates Alzheimer’s disease pathogenesis. Acta Neuropathol. 134, 489–506 (2017).
Hoozemans, J. J. et al. The unfolded protein response is activated in pretangle neurons in Alzheimer’s disease hippocampus. Am. J. Pathol. 174, 1241–1251 (2009).
Bellucci, A. et al. Induction of the unfolded protein response by alpha-synuclein in experimental models of Parkinson’s disease. J. Neurochem. 116, 588–605 (2011).
Atkin, J. D. et al. Endoplasmic reticulum stress and induction of the unfolded protein response in human sporadic amyotrophic lateral sclerosis. Neurobiol. Dis. 30, 400–407 (2008).
Hetz, C. et al. XBP-1 deficiency in the nervous system protects against amyotrophic lateral sclerosis by increasing autophagy. Genes Dev. 23, 2294–2306 (2009).
Ito, Y. et al. Involvement of CHOP, an ER-stress apoptotic mediator, in both human sporadic ALS and ALS model mice. Neurobiol. Dis. 36, 470–476 (2009).
Sasaki, S. Endoplasmic reticulum stress in motor neurons of the spinal cord in sporadic amyotrophic lateral sclerosis. J. Neuropathol. Exp. Neurol. 69, 346–355 (2010).
Duennwald, M. L. & Lindquist, S. Impaired ERAD and ER stress are early and specific events in polyglutamine toxicity. Genes Dev. 22, 3308–3319 (2008).
Lajoie, P. & Snapp, E. L. Changes in BiP availability reveal hypersensitivity to acute endoplasmic reticulum stress in cells expressing mutant huntingtin. J. Cell Sci. 124, 3332–3343 (2011).
Carreras-Sureda, A., Pihan, P. & Hetz, C. The unfolded protein response: at the intersection between endoplasmic reticulum function and mitochondrial bioenergetics. Front. Oncol. 7, 55 (2017).
Munoz, J. P. et al. Mfn2 modulates the UPR and mitochondrial function via repression of PERK. EMBO J. 32, 2348–2361 (2013).
Ngoh, G. A., Papanicolaou, K. N. & Walsh, K. Loss of mitofusin 2 promotes endoplasmic reticulum stress. J. Biol. Chem. 287, 20321–20332 (2012).
Schneeberger, M. et al. Mitofusin 2 in POMC neurons connects ER stress with leptin resistance and energy imbalance. Cell 155, 172–187 (2013).
Hayashi, T. & Su, T. P. Sigma-1 receptor chaperones at the ER-mitochondrion interface regulate Ca(2+) signaling and cell survival. Cell 131, 596–610 (2007).
Su, T. P., Hayashi, T., Maurice, T., Buch, S. & Ruoho, A. E. The sigma-1 receptor chaperone as an inter-organelle signaling modulator. Trends Pharmacol. Sci. 31, 557–566 (2010).
Hayashi, T., Maurice, T. & Su, T. P. Ca(2+) signaling via sigma(1)-receptors: novel regulatory mechanism affecting intracellular Ca(2+) concentration. J. Pharmacol. Exp. Ther. 293, 788–798 (2000).
Mori, T., Hayashi, T., Hayashi, E. & Su, T. P. Sigma-1 receptor chaperone at the ER-mitochondrion interface mediates the mitochondrion-ER-nucleus signaling for cellular survival. PLoS ONE 8, e76941 (2013).
Truettner, J. S., Hu, K., Liu, C. L., Dietrich, W. D. & Hu, B. Subcellular stress response and induction of molecular chaperones and folding proteins after transient global ischemia in rats. Brain Res. 1249, 9–18 (2009).
Verfaillie, T. et al. PERK is required at the ER-mitochondrial contact sites to convey apoptosis after ROS-based ER stress. Cell Death Differ. 19, 1880–1891 (2012).
Son, S. M., Byun, J., Roh, S. E., Kim, S. J. & Mook-Jung, I. Reduced IRE1alpha mediates apoptotic cell death by disrupting calcium homeostasis via the InsP3 receptor. Cell Death Dis. 5, e1188 (2014).
Schrader, M., Godinho, L. F., Costello, J. L. & Islinger, M. The different facets of organelle interplay-an overview of organelle interactions. Front. Cell Dev. Biol. 3, 56 (2015).
Hunt, M. C., Tillander, V. & Alexson, S. E. Regulation of peroxisomal lipid metabolism: the role of acyl-CoA and coenzyme A metabolizing enzymes. Biochimie 98, 45–55 (2014).
Fransen, M., Nordgren, M., Wang, B., Apanasets, O. & Van Veldhoven, P. P. Aging, age-related diseases and peroxisomes. Subcell. Biochem. 69, 45–65 (2013).
Delille, H. K., Alves, R. & Schrader, M. Biogenesis of peroxisomes and mitochondria: linked by division. Histochem. Cell Biol. 131, 441–446 (2009).
Schrader, M. & Yoon, Y. Mitochondria and peroxisomes: are the ‘big brother’ and the ‘little sister’ closer than assumed? Bioessays 29, 1105–1114 (2007).
Dixit, E. et al. Peroxisomes are signaling platforms for antiviral innate immunity. Cell 141, 668–681 (2010).
Medzhitov, R. & Horng, T. Transcriptional control of the inflammatory response. Nat. Rev. Immunol. 9, 692–703 (2009).
Anding, A. L. & Baehrecke, E. H. Cleaning house: selective autophagy of organelles. Dev. Cell 41, 10–22 (2017).
Murley, A. & Nunnari, J. The emerging network of mitochondria-organelle contacts. Mol. Cell 61, 648–653 (2016).
Marchi, S. et al. Mitochondrial and endoplasmic reticulum calcium homeostasis and cell death. Cell Calcium 69, 62–72 (2017).
Giorgi, C. et al. Mitochondria-associated membranes: composition, molecular mechanisms, and physiopathological implications. Antioxid. Redox Signal. 22, 995–1019 (2015).
Ruby, J. R., Dyer, R. F. & Skalko, R. G. Continuities between mitochondria and endoplasmic reticulum in the mammalian ovary. Z. Zellforsch. Mikrosk. Anat. 97, 30–37 (1969).
Horner, S. M., Wilkins, C., Badil, S., Iskarpatyoti, J. & Gale, M. Jr. Proteomic analysis of mitochondrial-associated ER membranes (MAM) during RNA virus infection reveals dynamic changes in protein and organelle trafficking. PLoS ONE 10, e0117963 (2015).
Poston, C. N., Krishnan, S. C. & Bazemore-Walker, C. R. In-depth proteomic analysis of mammalian mitochondria-associated membranes (MAM). J. Proteom. 79, 219–230 (2013).
Csordas, G. et al. Imaging interorganelle contacts and local calcium dynamics at the ER-mitochondrial interface. Mol. Cell 39, 121–132 (2010).
de Brito, O. M. & Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456, 605–610 (2008).
Naon, D. et al. Critical reappraisal confirms that Mitofusin 2 is an endoplasmic reticulum-mitochondria tether. Proc. Natl. Acad. Sci. USA 113, 11249–11254 (2016).
Cosson, P., Marchetti, A., Ravazzola, M. & Orci, L. Mitofusin-2 independent juxtaposition of endoplasmic reticulum and mitochondria: an ultrastructural study. PLoS ONE 7, e46293 (2012).
Filadi, R. et al. Mitofusin 2 ablation increases endoplasmic reticulum-mitochondria coupling. Proc. Natl. Acad. Sci. USA 112, E2174–E2181 (2015).
Leal, N. S. et al. Mitofusin-2 knockdown increases ER-mitochondria contact and decreases amyloid beta-peptide production. J. Cell. Mol. Med. 20, 1686–1695 (2016).
Sugiura, A. et al. MITOL regulates endoplasmic reticulum-mitochondria contacts via Mitofusin2. Mol. Cell 51, 20–34 (2013).
De Vos, K. J. et al. VAPB interacts with the mitochondrial protein PTPIP51 to regulate calcium homeostasis. Hum. Mol. Genet. 21, 1299–1311 (2012).
Gomez-Suaga, P. et al. The ER-mitochondria tethering complex VAPB-PTPIP51 regulates autophagy. Curr. Biol. 27, 371–385 (2017).
Stoica, R. et al. ER-mitochondria associations are regulated by the VAPB-PTPIP51 interaction and are disrupted by ALS/FTD-associated TDP-43. Nat. Commun. 5, 3996 (2014).
Paillusson, S. et al. alpha-Synuclein binds to the ER-mitochondria tethering protein VAPB to disrupt Ca2+homeostasis and mitochondrial ATP production. Acta Neuropathol. 134, 129–149 (2017).
Iwasawa, R., Mahul-Mellier, A. L., Datler, C., Pazarentzos, E. & Grimm, S. Fis1 and Bap31 bridge the mitochondria-ER interface to establish a platform for apoptosis induction. EMBO J. 30, 556–568 (2011).
Simmen, T. et al. PACS-2 controls endoplasmic reticulum-mitochondria communication and Bid-mediated apoptosis. EMBO J. 24, 717–729 (2005).
Hedskog, L. et al. Modulation of the endoplasmic reticulum-mitochondria interface in Alzheimer’s disease and related models. Proc. Natl. Acad. Sci. USA 110, 7916–7921 (2013).
Chami, M. et al. Role of SERCA1 truncated isoform in the proapoptotic calcium transfer from ER to mitochondria during ER stress. Mol. Cell 32, 641–651 (2008).
Doghman-Bouguerra, M. et al. FATE1 antagonizes calcium- and drug-induced apoptosis by uncoupling ER and mitochondria. EMBO Rep. 17, 1264–1280 (2016).
Herrera-Cruz, M. S. & Simmen, T. Of yeast, mice and men: MAMs come in two flavors. Biol. Direct 12, 3 (2017).
Hirabayashi, Y. et al. ER-mitochondria tethering by PDZD8 regulates Ca(2+) dynamics in mammalian neurons. Science 358, 623–630 (2017).
Cerqua, C. et al. Trichoplein/mitostatin regulates endoplasmic reticulum-mitochondria juxtaposition. EMBO Rep. 11, 854–860 (2010).
Vecchione, A. et al. MITOSTATIN, a putative tumor suppressor on chromosome 12q24.1, is downregulated in human bladder and breast cancer. Oncogene 28, 257–269 (2009).
Hetz, C. & Papa, F. R. The unfolded protein response and cell fate control. Mol. Cell 69, 169–181 (2018).
Somlyo, A. P. Cell physiology: cellular site of calcium regulation. Nature 309, 516–517 (1984).
Pinton, P. et al. The Ca2+concentration of the endoplasmic reticulum is a key determinant of ceramide-induced apoptosis: significance for the molecular mechanism of Bcl-2 action. EMBO J. 20, 2690–2701 (2001).
Scorrano, L. et al. BAX and BAK regulation of endoplasmic reticulum Ca2+: a control point for apoptosis. Science 300, 135–139 (2003).
Williams, G. S., Boyman, L. & Lederer, W. J. Mitochondrial calcium and the regulation of metabolism in the heart. J. Mol. Cell. Cardiol. 78, 35–45 (2015).
McCormack, J. G., Halestrap, A. P. & Denton, R. M. Role of calcium ions in regulation of mammalian intramitochondrial metabolism. Physiol. Rev. 70, 391–425 (1990).
Messina, A., Reina, S., Guarino, F. & De Pinto, V. VDAC isoforms in mammals. Biochim. Biophys. Acta 1818, 1466–1476 (2012).
Szabadkai, G. et al. Chaperone-mediated coupling of endoplasmic reticulum and mitochondrial Ca2+channels. J. Cell Biol. 175, 901–911 (2006).
Salomoni, P. & Pandolfi, P. P. The role of PML in tumor suppression. Cell 108, 165–170 (2002).
Giorgi, C. et al. PML regulates apoptosis at endoplasmic reticulum by modulating calcium release. Science 330, 1247–1251 (2010).
Joshi, A. U., Kornfeld, O. S. & Mochly-Rosen, D. The entangled ER-mitochondrial axis as a potential therapeutic strategy in neurodegeneration: a tangled duo unchained. Cell Calcium 60, 218–234 (2016).
Krols, M. et al. Mitochondria-associated membranes as hubs for neurodegeneration. Acta Neuropathol. 131, 505–523 (2016).
Paillusson, S. et al. There’s something wrong with my MAM; the ER-mitochondria axis and neurodegenerative diseases. Trends Neurosci. 39, 146–157 (2016).
Area-Gomez, E. et al. Upregulated function of mitochondria-associated ER membranes in Alzheimer disease. EMBO J. 31, 4106–4123 (2012).
Area-Gomez, E. et al. Presenilins are enriched in endoplasmic reticulum membranes associated with mitochondria. Am. J. Pathol. 175, 1810–1816 (2009).
Schreiner, B., Hedskog, L., Wiehager, B. & Ankarcrona, M. Amyloid-beta peptides are generated in mitochondria-associated endoplasmic reticulum membranes. J. Alzheimers Dis. 43, 369–374 (2015).
Sepulveda-Falla, D. et al. Familial Alzheimer’s disease-associated presenilin-1 alters cerebellar activity and calcium homeostasis. J. Clin. Investig. 124, 1552–1567 (2014).
De Strooper, B. Loss-of-function presenilin mutations in Alzheimer disease. Talking point on the role of presenilin mutations in Alzheimer disease. EMBO Rep. 8, 141–146 (2007).
Tambini, M. D. et al. ApoE4 upregulates the activity of mitochondria-associated ER membranes. EMBO Rep. 17, 27–36 (2016).
Guardia-Laguarta, C. et al. alpha-Synuclein is localized to mitochondria-associated ER membranes. J. Neurosci. 34, 249–259 (2014).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Russell, A. P. et al. Disruption of skeletal muscle mitochondrial network genes and miRNAs in amyotrophic lateral sclerosis. Neurobiol. Dis. 49, 107–117 (2013).
Xu, Y. F. et al. Wild-type human TDP-43 expression causes TDP-43 phosphorylation, mitochondrial aggregation, motor deficits, and early mortality in transgenic mice. J. Neurosci. 30, 10851–10859 (2010).
Al-Saif, A., Al-Mohanna, F. & Bohlega, S. A mutation in sigma-1 receptor causes juvenile amyotrophic lateral sclerosis. Ann. Neurol. 70, 913–919 (2011).
Tagashira, H., Shinoda, Y., Shioda, N. & Fukunaga, K. Methyl pyruvate rescues mitochondrial damage caused by SIGMAR1 mutation related to amyotrophic lateral sclerosis. Biochim. Biophys. Acta 1840, 3320–3334 (2014).
Bernard-Marissal, N., Medard, J. J., Azzedine, H. & Chrast, R. Dysfunction in endoplasmic reticulum-mitochondria crosstalk underlies SIGMAR1 loss of function mediated motor neuron degeneration. Brain 138, 875–890 (2015).
Amr, S. et al. A homozygous mutation in a novel zinc-finger protein, ERIS, is responsible for Wolfram syndrome 2. Am. J. Hum. Genet. 81, 673–683 (2007).
Braakman, I. & Bulleid, N. J. Protein folding and modification in the mammalian endoplasmic reticulum. Annu. Rev. Biochem. 80, 71–99 (2011).
Lynes, E. M. et al. Palmitoylated TMX and calnexin target to the mitochondria-associated membrane. EMBO J. 31, 457–470 (2012).
Gutierrez, T. & Simmen, T. Endoplasmic reticulum chaperones tweak the mitochondrial calcium rheostat to control metabolism and cell death. Cell Calcium, 70, 64–75 (2018).
Marchi, S. et al. Endoplasmic reticulum-mitochondria communication through Ca(2+) signaling: the importance of mitochondria-associated membranes (MAMs). Adv. Exp. Med. Biol. 997, 49–67 (2017).
Rouzier, C. et al. A novel CISD2 mutation associated with a classical Wolfram syndrome phenotype alters Ca2+homeostasis and ER-mitochondria interactions. Hum. Mol. Genet. 26, 1786 (2017).
Chen, Y. F. et al. Cisd2 deficiency drives premature aging and causes mitochondria-mediated defects in mice. Genes Dev. 23, 1183–1194 (2009).
Wu, C. Y. et al. A persistent level of Cisd2 extends healthy lifespan and delays aging in mice. Hum. Mol. Genet. 21, 3956–3968 (2012).
Chang, N. C., Nguyen, M., Germain, M. & Shore, G. C. Antagonism of Beclin 1-dependent autophagy by BCL-2 at the endoplasmic reticulum requires NAF-1. EMBO J. 29, 606–618 (2010).
Kale, J., Osterlund, E. J. & Andrews, D. W. BCL-2 family proteins: changing partners in the dance towards death. Cell Death Differ. 25, 65–80 (2018).
Pattingre, S. et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122, 927–939 (2005).
Chang, N. C. et al. Bcl-2-associated autophagy regulator Naf-1 required for maintenance of skeletal muscle. Hum. Mol. Genet. 21, 2277–2287 (2012).
Chen, R. et al. Bcl-2 functionally interacts with inositol 1,4,5-trisphosphate receptors to regulate calcium release from the ER in response to inositol 1,4,5-trisphosphate. J. Cell Biol. 166, 193–203 (2004).
Vervliet, T., Parys, J. B. & Bultynck, G. Bcl-2 proteins and calcium signaling: complexity beneath the surface. Oncogene 35, 5079–5092 (2016).
Vervliet, T. et al. Modulation of Ca2+signaling by anti-apoptotic B-Cell lymphoma 2 proteins at the endoplasmic reticulum-mitochondrial Interface. Front. Oncol. 7, 75 (2017).
Lu, S. et al. A calcium-dependent protease as a potential therapeutic target for Wolfram syndrome. Proc. Natl. Acad. Sci. USA 111, E5292–E5301 (2014).
Wang, C. H. et al. Cisd2 modulates the differentiation and functioning of adipocytes by regulating intracellular Ca2+homeostasis. Hum. Mol. Genet. 23, 4770–4785 (2014).
Tsai, P. H. et al. Dysregulation of mitochondrial functions and osteogenic differentiation in Cisd2-deficient murine induced pluripotent stem cells. Stem Cells Dev. 24, 2561–2576 (2015).
Zhang, A. et al. Quantitative proteomic analyses of human cytomegalovirus-induced restructuring of endoplasmic reticulum-mitochondrial contacts at late times of infection. Mol. Cell Proteom. 10, M111 009936 (2011).
Osman, A. A. et al. Wolframin expression induces novel ion channel activity in endoplasmic reticulum membranes and increases intracellular calcium. J. Biol. Chem. 278, 52755–52762 (2003).
Cagalinec, M. et al. Role of mitochondrial dynamics in neuronal development: mechanism for Wolfram syndrome. PLoS Biol. 14, e1002511 (2016).
Zatyka, M. et al. Sarco(endo)plasmic reticulum ATPase is a molecular partner of Wolfram syndrome 1 protein, which negatively regulates its expression. Hum. Mol. Genet. 24, 814–827 (2015).
Lynes, E. M. et al. Palmitoylation is the switch that assigns calnexin to quality control or ER Ca2+signaling. J. Cell Sci. 126, 3893–3903 (2013).
Morikawa, S., Tajima, T., Nakamura, A., Ishizu, K. & Ariga, T. A novel heterozygous mutation of the WFS1 gene leading to constitutive endoplasmic reticulum stress is the cause of Wolfram syndrome. Pediatr. Diabetes 18, 934–941 (2017).
Nishitoh, H. CHOP is a multifunctional transcription factor in the ER stress response. J. Biochem. 151, 217–219 (2012).
Hara, T. et al. Calcium efflux from the endoplasmic reticulum leads to beta-cell death. Endocrinology 155, 758–768 (2014).
Urano, F. Wolfram syndrome: diagnosis, management, and treatment. Curr. Diab Rep. 16, 6 (2016).
Varnai, P., Balla, A., Hunyady, L. & Balla, T. Targeted expression of the inositol 1,4,5-triphosphate receptor (IP3R) ligand-binding domain releases Ca2+via endogenous IP3R channels. Proc. Natl. Acad. Sci. USA 102, 7859–7864 (2005).
Takei, D. et al. WFS1 protein modulates the free Ca(2+) concentration in the endoplasmic reticulum. FEBS Lett. 580, 5635–5640 (2006).
Myhill, N. et al. The subcellular distribution of calnexin is mediated by PACS-2. Mol. Biol. Cell 19, 2777–2788 (2008).
Pinton, P., Giorgi, C. & Pandolfi, P. P. The role of PML in the control of apoptotic cell fate: a new key player at ER-mitochondria sites. Cell Death Differ. 18, 1450–1456 (2011).
Harding, H. P., Zhang, Y. & Ron, D. Protein translation and folding are coupled by an endoplasmic-reticulum-resident kinase. Nature 397, 271–274 (1999).
Bergeron, J. J., Brenner, M. B., Thomas, D. Y. & Williams, D. B. Calnexin: a membrane-bound chaperone of the endoplasmic reticulum. Trends Biochem. Sci. 19, 124–128 (1994).
Cali, T., Ottolini, D., Negro, A. & Brini, M. alpha-Synuclein controls mitochondrial calcium homeostasis by enhancing endoplasmic reticulum-mitochondria interactions. J. Biol. Chem. 287, 17914–17929 (2012).
Atwal, R. S. et al. Huntingtin has a membrane association signal that can modulate huntingtin aggregation, nuclear entry and toxicity. Hum. Mol. Genet. 16, 2600–2615 (2007).
Reijonen, S., Putkonen, N., Norremolle, A., Lindholm, D. & Korhonen, L. Inhibition of endoplasmic reticulum stress counteracts neuronal cell death and protein aggregation caused by N-terminal mutant huntingtin proteins. Exp. Cell Res. 314, 950–960 (2008).
Acknowledgements
This work was supported by external ressources from “Institut National de la Santé et de la Recherche Médicale” (INSERM), Université de Montpellier, and Association Syndrome de Wolfram (Grand-Champ, France) and grants from the Agence Nationale pour la Recherche (ANR-12-JSV1-0008-01), Fondation pour la Recherche Médicale, and Fondation de France.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Pinton
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Delprat, B., Maurice, T. & Delettre, C. Wolfram syndrome: MAMs’ connection?. Cell Death Dis 9, 364 (2018). https://doi.org/10.1038/s41419-018-0406-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0406-3
This article is cited by
-
A deep phenotyping study in mouse and iPSC models to understand the role of oligodendroglia in optic neuropathy in Wolfram syndrome
Acta Neuropathologica Communications (2024)
-
ER calcium depletion as a key driver for impaired ER-to-mitochondria calcium transfer and mitochondrial dysfunction in Wolfram syndrome
Nature Communications (2024)
-
LDLs take a shortcut to mitochondria
Nature Cell Biology (2023)
-
Targeting Sigma Receptors for the Treatment of Neurodegenerative and Neurodevelopmental Disorders
CNS Drugs (2023)
-
Multiomic analysis on human cell model of wolfram syndrome reveals changes in mitochondrial morphology and function
Cell Communication and Signaling (2021)