Abstract
As the most primordial signaling pathway in animal physiology, the Hippo pathway and innate immunity play crucial roles not only in sensing cellular conditions or infections, but also in various metabolite homeostasis and tumorigenesis. However, the correlation between cellular homeostasis and antiviral defense is not well understood. The core kinase LATS1/2, could either enhance or inhibit the anti-tumor immunity in different cellular contexts. In this study, we found that LATS2 can interact with PQBP1, the co-factor of cGAS, thus enhanced the cGAS-STING mediated innate immune response to HIV-1 challenge. LATS2 was observed to upregulate type-I interferon (IFN-I) and cytokines in response to HIV-1 reverse-transcribed DNA and inhibited HIV-1 infection. Due to the involvement of PQBP1, the function of LATS2 in regulating cGAS activity is not relying on the downstream YAP/TAZ as that in the canonical Hippo pathway. The related kinase activity of LATS2 was verified, and the potential phosphorylation site of PQBP1 was identified. Our study established a novel connection between Hippo signaling and innate immunity, thus may provide new potential intervention target on antiviral therapeutics.
Similar content being viewed by others
Introduction
The Hippo signaling pathway was first uncovered as a key mechanism to control tissue growth in Drosophila [1,2,3,4,5,6,7,8,9,10] and it is highly conserved from unicellular organisms to mammals and human beings [11,12,13,14,15,16]. Central to the mammalian Hippo pathway is a Hippo kinase cascade composed of upstream kinase MST1/2 and downstream kinase LATS1/2 as well as their respective adapters SAV1 and MOB1. Upon various stimuli, activated MST1/2 phosphorylates and activates LATS1/2, in turn active LATS1/2 phosphorylates and inactivates the effector YAP/TAZ, resulting in turn-off of the transcriptional output of target genes in a transcription factor TEAD1/2/3/4 (TEADs)-dependent manner [17]. Intensive research has demonstrated that the canonical Hippo pathway, characterized by signaling input through the Hippo kinase cascade ultimately impinging on the downstream transcriptional complex YAP/TAZ/TEADs, functions not only in growth control and tumorigenesis during animal development and regeneration, but also in non-developmental processes such as innate immunity against Gram-positive bacterium infection in Drosophila [18].
Beyond the important roles of Hippo signaling in growth control and homeostasis, components of the Hippo pathway also play indispensable roles in diverse processes including immune response through the non-canonical Hippo pathway, which is usually recognized as the coordination between the Hippo kinase cascade and transcriptional complex YAP/TAZ/TEADs is not necessary during signal transduction. Except the canonical function in controlling cell replication, MST1/2 has been associated with both innate and adaptive immunity. MST1/2 can phosphorylate TBK1 and IRF3, thus upregulate the innate immune response [19, 20]. MST1/2 can also inhibit immune response and attenuate the inflammation in RIG-I/MAVS-mediated RNA virus infection [21]. MST1/2 is also important in T-cell development, including T-cell maturation, migration, and differentiation [22,23,24,25]. In addition, LATS1/2 can either enhance or reduce immune response by upregulate the expression of IFNβ or suppress the secretion of immunity-inducing nucleic acids-containing exosomes [26, 27]. Furthermore, YAP/TAZ can interact with TBK1 and IRF3, inhibiting the activation of TBK1 and blocking both RNA and DNA virus-induced immune response [27].
As the component connecting upstream MST1/2 and downstream YAP/TAZ, LATS1/2 is indispensable for the appropriate signaling in Hippo pathway. Unlike MST1/2, which has multiple substrates and could lead to various downstream signaling [28,29,30,31], LATS1/2 specifically targets YAP/TAZ, and almost all the phenotypes observed in the lats1/2 knockout are mediated via YAP/TAZ [32]. A parallel pathway to MST1/2 has also been observed, in which MAP4Ks are used to phosphorylate LATS1/2 [33, 34]. All these observations emphasize the central role of LATS1/2 in the Hippo pathways, both canonical pathway in cell growth and non-canonical pathway in immune response.
PQBP1, a protein predominantly expressed in lymphoid and myeloid cells, was initially identified in the pre-mRNA splicing by interacting with splicing-related factors [35]. The mutation within pqbp1 gene causes X-linked mental retardation disorders, such as the Renpenning, Sutherland-Haan, Hamel, Porteous, and Golabi-Ito-Hall syndromes [36,37,38]. Recently, the role of PQBP1 in innate immune response was gradually revealed [39, 40]. By forming a complex with cGAS, PQBP1 can decide the intensity and specificity of cGAS on DNA recognization. PQBP1 might contribute to the recognization of viral DNA pattern and stabilize the DNA–cGAS interaction, thus enhance the binding of DNA with cGAS and define the DNA sequence spectrum that can induce the enzymatic activity of cGAS. The specific role of PQBP1 in retrovirus-, especially HIV-1-induced immune response has been well established [39], despite the plausibility that PQBP1 involvement in other viral infection is also implicated.
In this report, during investigating the innate immune response induced by HIV-1 infection, we found LATS2, but not LATS1, can influence the HIV-1 replication by interacting with PQBP1, the co-factor of cGAS [39]. The influence of LATS2 on HIV-1 replication is independent of downstream YAP/TAZ, but dependent on the kinase activity of LATS2. With this regard, PQBP1 is the substrate of LATS2, and the phosphorylation of PQBP1 leads to a more stable PQBP1–cGAS complex and stronger immune response upon HIV-1 infection. Accordingly, the HIV-1 infection also elicits this LATS2-PQBP1-cGAS-STING pathway in the primary human MDM (myeloid-derived macrophage) cells. With these findings, a novel regulatory mechanism of cellular innate immunity against HIV-1 infection by Hippo component is uncovered.
Results
PQBP1 interacts with LATS2
In the exploration of PQBP1 interacting proteins by an immunoprecipitation followed by mass-spectrometry, serendipitously we found PQBP1 can interact with LATS2 (Table S1), a central component of Hippo pathway. PQBP1 is known as a co-factor of cGAS, thus we interrogated the interaction between PQBP1 and LATS2. From co-immunoprecipitate experiment in 293T cells, LATS2, but not its paralog LATS1, interacts with PQBP1 (Fig. 1a), implying a specific function of LATS2 in PQBP1-mediated cellular processes. Furthermore, we established pqbp1 and lats2 knockout (sgPQBP1 and sgLATS2) THP1 cells, a monocyte cell line that is widely used for study of innate immunity, and further confirmed their specific interaction (Fig. S1 and Fig. 1b, c). To dissect which region is responsible for the interaction, we generated series of expression fragments of both proteins according to their domain demarcation. From the co-immunoprecipitate results, the protein-binding domain of LATS2 and the WW domain (WWD) of PQBP1 are accountable for their interaction (Fig. 1d, e). Interestingly, their interaction is enhanced upon pseudovirus HIV-1 infection (Fig. 1f), indicating their interaction is physiologically relevant to the cellular context. The location of PQBP1 in HeLa cells and THP1 cells, either represented with GFP, or shown with immunostaining for endogenous PQBP1, confirmed its colocalization with LATS2 in cytoplasm (Fig. S2). Moreover, colocalization of LATS2 and PQBP1 was also observed in HIV-1 infected macrophage-like cells (THP1-PMA), despite a strong nuclear localization of PQBP1 can always be observed (Fig. 1g).
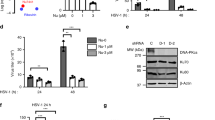
a Association of exogenously expressed LATS2 and PQBP1 in HEK293T cells. HEK293T cells (4 × 106) were transfected with PQBP1-Flag and Myc-LATS1/2. Co-immunoprecipitation was performed with anti-Myc beads followed by the immunoblot analysis. Determination of endogenous LATS2-PQBP1 association in THP1 cells. Immunoprecipitation was performed by using anti-PQBP1, anti-LATS2, or control IgG in PQBP1 knockout (b) or LATS2 knockout (c) cell lines. d, e Mapping of interacting domains between LATS2 and PQBP1 by GFP-trap co-immunoprecipitation. Co-immunoprecipitation was performed with GFP-trap beads followed by immunoblotting with the indicated antibodies. The schematic diagrams of LATS2 or PQBP1 truncated mutants were shown on the upper panels (For LATS2: LCD1 LATS conserved domain 1, 1–160 aa; LCD2 LATS conserved domain 2, 403–463 aa: PBD protein-binding domain, 618–720 aa: KD kinase domain, 667–1088 aa. For PQBP1: WWD WW domain, 49–81 aa; PRD Pro-rich domain, 104–163 aa; CTD, C-terminal domain, 191–265 aa. “+” indicates as interaction, “−” as no interaction, and “+/−” as weak interaction). f The association of endogenous LATS2-PQBP1 was reinforced by HIV-1 challenge. THP1 cells were untreated or infected with HIV-1 for the indicated times. Co-immunoprecipitation was performed with control IgG or anti-PQBP1 beads followed by the immunoblot analysis. g Colocalization of LATS2 and endogenous PQBP1. THP1 cells were infected with LATS2-mCherry for 24 h, then treated with PMA and HIV-1/VPX for another 20 h. Endogenous PQBP1 was analyzed by immunostaining with anti-PQBP1 primary antibody and FITC-labeled secondary antibody. The red line on the zoom panel indicates the plot profile analysis using ImageJ software. Scale bar, 10 μm.
LATS2 inhibits HIV-1 replication
The interaction between LATS2 and PQBP1 is enhanced upon HIV-1 infection, therefore we speculate that LATS2 might be involved in the process of HIV-1 infection. The THP1 cells, a monocyte cell line widely used for the study of HIV-1 infection, were used to investigate the influence of LATS2 on HIV-1 infection. Both the LATS2 overexpression (LATS2-OE) and lats2 knockout (lats2-KO) THP1 cell lines were established (Fig. 2a, e and Fig. S1c, d). Judged by either luciferase activity or Pr55Gag protein levels, the HIV-1 replication was reduced in LATS2 overexpression cells (Fig. 2b–d). Accordingly, the HIV-1 replication was enhanced in lats2 knockout cells (Fig. 2f–h). Using the shRNA-mediated knockdown of LATS2, the effect of LATS2 on HIV-1 infection was also confirmed in peripheral blood mononuclear cells (PBMCs) purified from healthy human blood (Fig. S3 and Fig. 2i, j). A consecutive viral replication with live HIV-1 virus was also measured in the lats2-KO THP1 cells. The live virus infection on these cells confirmed that LATS2 has an inhibitory effect on HIV-1 replication (Fig. 2k) as well as the effect of PQBP1 and cGAS (Fig. S4). During viral infection, multiple steps constitute the viral life cycle, which includes viral entry, disassembly, reverse transcription, nuclear import, integration, mRNA transcription and translation, viral assembly and release, and the efficiency of each step can be measured by the products generated at every step. To examine the point at which LATS2 affects HIV infection, we identified that LATS2 deficiency increased 2-LTR circles, but not the early- and intermediate-stage products of reverse transcription (RU5, U3/U5, GAG) (Fig. 2i). The observation supports that the effect of LATS2 on HIV infection is partly controlled by ISGs, such as MX2, which inhibits HIV-1 cDNA nuclear import [41, 42].
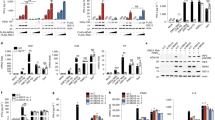
a The expression level of LATS2 in a stable LATS2-overexpressing THP1 cell line. Immunoblot analysis was performed to detect the protein levels of LATS2 and GAPDH. b–d LATS2-overexpressing THP1 cells were infected with a single-cycle VSV-G pseudotyped HIV-1 (with luciferase reporter). Luciferase activity (b) and the protein levels of HIV-1 Pr55Gag (c) were determined at indicated time points. Densitometry analysis was performed for Pr55Gag by Image J (d). e LATS2 knockout THP1 cell lines were generated by CRISPR-Cas9/sgRNA. The efficiency was determined by western blot. f–h Same as (b) except the infection was done in LATS2 knockout cells. sg Con sg control; sg #1 sgLATS2 #1; sg #2 sgLATS2 #2. i–j Knockdown efficiency of LATS2 in human PBMCs expressing LATS2 shRNA compared with a scramble control (i). Primary human PBMCs were infected with LATS2-shRNA lentivirus for 24 h, followed by infection with VSV-G pseudotyped HIV-1. The infection of HIV-1 was reflected by luciferase activity (j). k Multiple-round virus replication in LATS2 knockout THP1 cells. Supernatants were harvested at the indicated time points, and HIV-1 particle release was quantified by p24 ELISA. l Real-time PCR analysis of HIV-1 reverse transcription as previously described by Mbisa et al. [41]. LATS2 knockout THP1 cells were infected with VSV-G pseudotyped HIV-1. Total DNA was extracted at indicated time points, then HIV-1 DNA was quantified by amplifying indicated target sequences. Data shown were representative of three independent experiments. Error bars indicate SD. ***P < 0.001.
LATS2 upregulates the cellular innate immune response upon HIV-1 infection
The inhibition of LATS2 on HIV-1 replication occurs in a stage after DNA fragments generation by reverse-transcription (Fig. 2i, Fig. S5a), which was required for HIV-1 to activate the innate immune response [39, 43]. Moreover, LATS2 can strongly interact with PQBP1, and faintly with cGAS and STING (Fig. S5b). Thus, these results prompt us to investigate if the innate immune system is influenced by LATS2 during HIV-1 infection. The THP1 cell lines established above were induced into macrophage by PMA treatment, and HIV-1 in combined with VPX was used to infect these cells, a procedure that is widely used in studying the relationship between HIV-1 infection and innate immunity (Fig. 3a) [44]. The co-expression of VPX could inhibit the dNTPase activity of SAMHD1 in THP1 cells and thus allow quantitative DNA fragments to be reversely transcribed by HIV-1. Compared to sole pseudotyped HIV-1 infection, co-infection of HIV-1 and VPX-expressing retrovirus led to a lower, but accountable infection rate on these cells as implied with yellow fluorescence encoded within HIV-1 backbone (Fig. S6a, b). Upon HIV-1/VPX infection, the significant upregulation of interferon β (IFNβ) and interferon-stimulated genes 54 (ISG54) at mRNA levels, two reporters widely used to measure the magnitude of innate immunity, indicates a significant activation on cellular immune response (Fig. 3b) [39]. Using this system, the effect of LATS2 on activation of innate immunity was interrogated. In agreement with the influence of LATS2 on HIV-1 replication (Fig. 2), the increase of LATS2 enhanced the cellular innate immune response, whereas the decrease of LATS2 reduced the immune response (Fig. 3c, d). These observations support a direct link between HIV-1 inhibition and innate immunity upregulation due to the presence of LATS2.
a The sketch map to illustrate the infection of HIV-1 in macrophage. PMA-treated THP1 (PMA-THP1) or human macrophage are non-permissive to HIV-1 infection due to expression of SAMHD1, which converts intracellular dNTPs to deoxynucleosides (dN) and inorganic triphosphates (PPP). However, this restriction is circumvented by VPX, which targets SAMHD1 for degradation by proteasome. b Gene expression was measured as fold-induction over untreated cells with VPX or HIV-1/VPX in PMA-THP1. c LATS2 positively regulates the innate immune response upon HIV-1 infection. LATS2-overexpressing THP1 cells were PMA-differentiated and infected with HIV-1/VPX for 20 h. The relative level of IFNβ, ISG54, MX2, and CXCL10 were measured by qPCR. d LATS2 deficiency inhibits HIV-1-triggered signaling. Same as (b) except the infection was done in LATS2 knockout cells. e, f Knockdown of LATS2 decreases HIV-1-triggered signaling in human primary macrophages. The silencing efficiency was determined by western blotting (e). Monocyte-derived macrophages from healthy human blood were infected with LATS2-shRNA lentivirus for 24 h, followed by infection with HIV-1/VPX for another 20 h before qPCR analysis (f).
To confirm this conclusion further, the macrophages differentiated from PBMCs mediated by PMA were subjected to LATS2 knockdown and HIV-1/VPX infection. The HIV-1/VPX-induced IFNβ and interferon-stimulated genes (ISGs), ISG54 and CXCL10, were all reduced upon LATS2 knockdown with lats2-specific shRNA, suggesting LATS2 may also enhance the innate immune responses in primary immune cells (Fig. 3e, f).
LATS2 regulates antiviral signaling at the upstream of STING
The host cells can activate antiviral cGAS-STING-TBK1/IRF3 signaling during HIV-1 infection. By recognizing reversely transcribed DNA fragments of HIV-1, cGAS can synthesize cGAMP [45], the critical molecule used to activate downstream STING (also named as MITA, ERIS, MPYS, TMEM173) [46,47,48,49]. STING can undergo oligomerization, translocation, and post-translational modification upon cGAMP binding, thus recruits TBK1 and IRF3, resulting in IRF3 phosphorylation, dimerization, and nuclear translocation [50]. To address which step is influenced by LATS2 in this signaling pathway, first, we examined the effects of deletion of LTAS2 on IFN-I activation mediated by delivering chemical cGAMP, which would activate STING directly and increase the mRNA levels of immune responsive genes (IFNβ, ISG54, and CXCL10) (Fig. 4a). The lats2-KO cells have comparable immune response with wild-type cells (Fig. 4b), indicating LATS2 regulates HIV-1-triggered signaling at the upstream of STING. Second, the endogenously synthesized cGAMP was extracted from same amount of virus-infected lats2-KO and wild-type cells and was used to activate downstream monitor cells (Fig. 4c). The significant difference on immune response by extracted cGAMP from cells (Fig. 4d) and the measurement of cGAMP levels (Fig. 4e) indicated that the synthesized cGAMP in lats2-KO was dramatically reduced. Third, the HIV-1-triggered IRF3 phosphorylation, dimerization, and nuclear translocation was impaired in the lats2-KO cells compared with that in the control cells (Fig. 4f, g). Consistently, the HIV-1-triggered IRF3 activation was enhanced by overexpression of LATS2 (Fig. S6c). These findings support LATS2 acts on upstream of STING, with the plausibility on cGAMP synthesis step by cGAS.
a, b A schematic diagram illustrating the activation of downstream signaling of STING by cGAMP (a). cGAMP (400 ng) was delivered to PFO-permeabilized LATS2 knockout THP1 cells and incubated for 4 h before qPCR analysis (b). c, d A schematic diagram illustrates the measurement of cGAMP generated in the HIV-1 infected lats2 knockout THP1 cells (c). LATS2-deficient and control THP1 cells were infected with HIV-1 for 20 h. Cell extracts containing cGAMP were delivered to PFO-permeabilized THP1 cells and incubated for 6 h before qPCR analysis (d). e cGAMP in the cell extracts was measured by ELISA. f, g Effects of LATS2 on the HIV-1-induced IRF3 phosphorylation, dimerization, and nuclear translocation. LATS2-deficient THP1 cell lines were untreated or infected with HIV-1 for 20 h before IRF3 dimerization analysis and immunoblot analysis (f). The densitometry of p-IRF3 was calculated in (g). Data shown were representative of three independent experiments. Error bars indicate SD. ns no significance; ***P < 0.001.
LATS2 phosphorylates PQBP1
The PQBP1-binding ability (Fig. 1) and the influence on cGAS (Fig. 4) indicate that LATS2 might act on immune response through its regulation on PQBP1. To address this, first we checked if the kinase activity of LATS2 is necessary for this function. A LATS2 mutant with point mutation on critical residue for kinase activity (LATS2-KA, loss-of-activity LATS2 with K697A mutation) was used for comparison [51, 52]. As shown in Fig. 5a, b, only wild-type LATS2, but not LATS2-KA, can enhance immune response during HIV-1 infection, implying the kinase activity is indispensable for LATS2 function here. Second, we interrogated if PQBP1 could be phosphorylated by LATS2. An anti-phosphoserine antibody was used to detect the phosphorylated PQBP1 in lats2-KO cells. A weak band corresponding to the phosphorylated PQBP1 could be observed in wide-type, but not in last2-KO cells (Fig. 5c). To clarify this further, an in vitro phosphorylation assay was used with purified proteins. As shown in Fig. 5d, PQBP1 can be phosphorylated by LATS2, but not LATS2-KA or LATS1. The plausible phosphorylation sites of PQBP1 were also interrogated. The phosphorylation of Ser94 and Ser247 in PQBP1 have once been observed in a large-scale proteomics study [53, 54], so these two sites were under consideration with priority. We also included Ser208, a site implied by an online program used for predicting potential phosphorylation sites (NetPhos, http://www.cbs.dtu.dk/services/NetPhos/). Both the non-phosphorylated and phosphorylation-mimic forms (Ser to Ala and Ser to Asp) were prepared. In the in vitro phosphorylation assay, S247A PQBP1 has the most significant decrease on phosphorylation level (Fig. 5e). The mutants were then used to rescue the PQBP1 function in a pqbp1-KO THP1 cell line. Following HIV-1 infection, only Ser247 mutants showed a significant alteration compared to wild-type PQBP1 on immune response, which were downregulated by PQBP1-S247A and upregulated by PQBP1-S247D (Fig. 5f, g). Based on these observations, we can conclude that LATS2 regulates immune response by phosphorylating PQBP1, and the S247 is the most plausible phosphorylation site, despite we cannot exclude the involvement of other phosphorylation sites at this stage. Furthermore, the distribution of PQBP1 within cells was not disturbed by these site mutations (Fig. S7), indicating the influence of LATS2 on PQBP1 was independent on the PQBP1 localization.
a, b The kinase activity is necessary for LATS2 function in HIV-1-triggered signaling. THP1 cells were infected with LATS2 or LATS2-KA (LATS2 with K697A mutation) retrovirus for 24 h and differentiated by treating with 50 ng/ml PMA. Then the cells were infected with HIV-1/VPX for 20 h before qPCR analysis (b). Immunoblot analysis was performed to detect the protein levels of LATS2 or LATS2-KA (a). c Serine phosphorylation of endogenous PQBP1 was detected in HIV-1 infected THP1 cells. LATS2-deficient and control THP1 cells were infected with HIV-1 for 24 h, then the cell lysates were analyzed by co-immunoprecipitation using an anti-PQBP1 antibody and immunoblotting using an anti-phospho-Ser antibody. d The PQBP1 phosphorylation was detected in the presence of LATS2 in the in vitro kinase assay. Myc-LATS1, Myc-LATS2, and kinase-dead LATS2-KA were purified from transfected HEK293T cells with anti-Myc antibody, while 6×His-PQBP1 was expressed and purified from E. coli BL21 (DE3) with Ni-NTA beads. In vitro kinase assay was performed in the presence of ATP-γS, and the phosphorylation of PQBP1 was detected by immunoblotting with anti‐thiophosphate ester antibody. e Similar in vitro kinase assays were done as (d) except the substrates were replaced by PQBP1 mutants. f, g S247 is the plausible phosphorylation site in PQBP1 challenged by HIV-1. PQBP1-deficient THP1 cells were infected with retroviruses expressing PQBP1 or PQBP1 mutants (PAM sites were mutated to avoid cleaving by Cas9) for 24 h. Then the cells were differentiated into macrophage by PMA for another 24 h and infected with HIV-1/VPX for additional 20 h before qPCR analysis. Data shown were representative of three independent experiments. Error bars indicate SD. ns no significance; ***P < 0.001.
The influence of LATS2 on cGAS is mediated by PQBP1
From above observations, we established the connection among LATS2, PQBP1, and cGAS. To solidify the signaling pathway of LATS2-PQBP1-cGAS, we investigated their interaction further. First, by using the pqbp1 knockout and cgas knockout cells, we confirmed the function of LATS2 during HIV-1 infection was mediated by PQBP1/cGAS. In pqbp1 knockout and cgas knockout cells, LATS2 had no effect on antiviral signaling of the cells (Fig. 6a, b and Fig. S8a, b). As a kinase, LATS2 was not directly associate with HIV-1 nucleic acids (Fig. S8c, d). In the co-immunoprecipitate experiment between cGAS and PQBP1, the co-expression of LATS2 increased the interaction between cGAS and PQBP1 in a dose-dependent manner (Fig. 6c left). Consistently, knockdown of LATS2 attenuated the interaction between cGAS and PQBP1 during HIV-1 infection (Fig. 6c right). In addition, S247D mutation, which was shown to increase the immune response of PQBP1 (Fig. 5g), also increased the binding ability with cGAS (Fig. 6d). Furthermore, LATS2 and cGAS had no significant colocalization in untreated HeLa cells, but their colocalization was enhanced dramatically upon co-expression of PQBP1 or infection of HIV-1, accompanying with a striking increase of cGAS puncta within the cells (Fig. 6e). This observation was confirmed in THP1 cells (Fig. 6f). All these observations suggest LATS2 regulates immune response by phosphorylating PQBP1, which leads to the alteration of the stability of PQBP1/cGAS complex and/or PQBP1/cGAS/DNA triple complex.
a, b LATS2 regulates HIV-triggered signaling in a PQBP1-dependent manner. PQBP1-deficient and control THP1 cells were transduced by retrovirus expressing LATS2 for 24 h and differentiated by PMA before the cells were infected with HIV-1/VPX for 20 h. The protein level of LATS2 was confirmed by immunoblot (a). RT-PCR was performed to detect the indicated gene expression (b). c LATS2 strengthens assembly of the PQBP1-cGAS complexes. HEK293T cells (4 × 106) were co-transfected with PQBP1-Flag and HA-cGAS, together with increasing amounts of Myc-LATS2, or together with LATS2 shRNA. Co-immunoprecipitation and immunoblotting were performed with the indicated antibodies. d S247D mutation of PQBP1 increases the binding ability with cGAS. HEK293T cells were transfected with HA-cGAS and PQBP1-GFP or its mutants for 48 h before GFP-trap co-immunoprecipitation and immunoblot analysis with the indicated antibodies. e LATS2 and cGAS co-localize at foci in response to overexpression of PQBP1 or HIV-1 infection. cGAS-GFP and LATS2-mCherry constructs were exogenously expressed in HeLa cells for 24 h, then either co-expression with PQBP1 or infection with HIV-1 for another 20 h before observation. Fluorescence was imaged by a confocal laser scanning microscope. Scale bar, 10 μm. f Colocalization of LATS2 and endogenous cGAS in HIV-1 infected THP1 cell. THP1 cells were transduced by retrovirus expressing LATS2-mCherry for 24 h before infection with HIV-1 for another 20 h. Endogenous cGAS was analyzed by immunostaining with anti-cGAS primary antibody and FITC-labeled secondary antibody. The red line on the zoom panel indicates the plot profile analysis using ImageJ software. Scale bar, 10 μm. Data were representative of three independent experiments. Error bars indicate SD. ns no significance; ***P < 0.001.
LATS2 enhances cGAS oligomerization
The cGAMP synthase cGAS undergoes oligomerization upon activation, which contributes to the increase of cGAS activity following recognization of extrinsic DNA [55, 56]. By co-expression of FLAG-tagged and HA-tagged cGAS, we investigated the oligomerization of cGAS with or without HIV-1 infection. In both cases, the absence of LATS2 reduced the oligomerization of cGAS (Fig. 7a). The oligomerization of cGAS was also detected by running a semi-denaturing detergent agarose gel electrophoresis (SDD-AGE) [56, 57]. The endogenous cGAS in wide-type THP1 cells formed significant oligomer upon HIV-1 infection, increasing with infection time (Fig. 7b). In contrast, no obvious cGAS oligomer could be observed in THP1 lats2-KO cells (Fig. 7b), indicating a critical role of LATS2 in enhancing cGAS oligomerization.
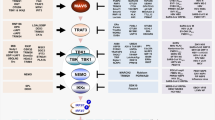
a Effects of LATS2-deficiency on self-association of cGAS. LATS2-deficient and control HEK293T cells were transfected with HA-cGAS and Flag-cGAS for 24 h and infected with HIV-1 for another 20 h. Co-immunoprecipitation was performed with anti-HA beads followed by immunoblotting analysis with the indicated antibodies. b Effects of LATS2-deficiency on oligomerization of cGAS triggered by HIV-1 infection in THP1. LATS2-deficient and control THP1 cells were untreated or infected with HIV-1 for the indicated times. Cell lysates were fractionated with SDD-AGE and detected by immunoblotting to measure the oligomerization of cGAS. c A proposed model of LATS2-mediated regulation of innate antiviral signaling to inhibit HIV-1 infection. In unstimulated cells, co-activators YAP/TAZ accumulate in the nucleus, and induce the pro-growth process by interacting with transcription factor TEADs. When cells are stimulated under stress, multiple upstream signals regulate the phosphorylation of LATS1/LATS2 kinases, thereby triggering YAP/TAZ phosphorylation. Phosphorylation of YAP/TAZ leads to their cytoplasmic retention and/or degradation by recruiting 14-3-3 proteins. Upon infection with HIV-1, cGAS and PQBP1 sense reverse-transcribed HIV cDNA and elicit the IFN response by producing cGAMP. Meanwhile, LATS2 interacts with PQBP1 and triggers its phosphorylation, subsequently enhances oligomerization of cGAS in response to HIV-1 infection, increasing the production of cGAMP. Ultimately, the second messenger cGAMP triggers robust innate immune responses via the STING-TBK1-IRF3 axis.
In summary, these results support a pivotal role of LATS2 in controlling the cGAS activity by regulating the phosphorylation of the cGAS co-factor PQBP1. A schematic model was illustrated in Fig. 7c. In normal growth condition, LATS2 regulates the cell proliferation by affecting the phosphorylation of downstream transcription activator YAP/TAZ as suggested in canonical Hippo pathway. Upon HIV-1 infection, LATS2 phosphorylates PQBP1, thus enhances the activity of PQBP1/cGAS complex and augments the intensity of cellular immune response, therefore regulates the balance between the cell growth and the cell immunity.
Discussion
The cells confront various stressful environmental conditions during proliferation, within which Hippo pathway plays critical roles in deciding the cell fate. Upon pathogenic infection, the cells have to decide to continue on growth or switch to immune response induced by infection for the better opportunity of cell survival, therefore, it is not surprising that the Hippo pathway is also involved in the activation of cellular immune responses. Nearly every component of Hippo pathway, including MST, LATS, and YAP/TAZ have been implicated in the immune response, which involves both innate and adaptive immunity process [18, 58, 59]. As the core component of the Hippo pathway, the function of LATS2 has been well recognized in controlling cell growth, whereas the underlying mechanism of its roles in innate immunity has not been clearly elucidated. In this study, we identified a specific function of LATS2 in affecting HIV-infection-induced cGAS-STING-TBK1 immune axis, which is achieved by regulating the phosphorylation of cGAS co-factor PQBP1. By impact on IFN-I and cytokines induction, LATS2 contributes to the cellular anti-HIV immunity directly. HIV-1 replication can be inhibited by type-I interferon, and the expression of a number of gene products with anti-HIV-1 activity is induced by type-I interferon [43, 60]. HIV infection led to the induction of IFNβ and the chemokine CXCL10 [45]. HIV-1 elicits an IRF3-dependent innate immune response, which resulted in IRF3-dependent ISG54 induction [39]. MX2 protein was reported as an inhibitor of HIV-1 infection and as a major effector of IFN-mediated resistance to HIV-1 infection [60]. MX2 inhibits HIV-1 cDNA nuclear import, a process that is partly controlled by ISGs [43].
PQBP1 is originally recognized as the critical component of RNA spliceosome, a function in consistent with its largely nuclear localization and close association with many splicing factors, including the U5 snRNP-specific 15kD protein (U5-15kD) and WBP11/NpwBP/SIPP1 [35, 61]. Recently, PQBP1 was identified as the co-factor of cGAS [39]. By association with cGAS, PQBP1 enhances the cGAS enzymatic activity and the specificity of DNA recognition. At the beginning, the co-factor role of PQBP1 was thought as retrovirus-specific, but the latter studies revealed that the PQBP1–cGAS complex might response to various extrinsic DNA. In this study, we discovered that the recognization on HIV-1 DNA by PQBP1-cGAS was controlled by Hippo kinase LATS2. The LATS2-PQBP1-cGAS axis might not be limited to HIV-1, because the cellular immune response to HSV-1 infection has also significant difference when LATS2 was knocked out (Fig. S9). Despite the largely nuclear localization of PQBP1, the colocalization among these proteins in cytoplasm could be observed (Fig. S2, Figs. 1g and 6e, f), which was strongly enhanced upon HIV-1 infection.
The kinase activity is indispensable for LATS2 function in mediating immune response (Fig. 5b), accordingly, we sought to interrogate if PQBP1 was phosphorylated by LATS2, and which residue was modified upon viral infection. Our study referred to the Ser247 as the major phosphorylation site by LATS2. Previously the N-terminal WWD, but not the C-terminal part of PQBP1 was indicated in cGAS interaction [39]. The WWD of PQBP1 was also responsible for the interaction with LATS2 (Fig. 1f). From these observations, we speculate that the modification on Ser247 located within C-terminal part might enhance the binding between PQBP1 and cGAS by disclosing the N-terminal WWD of PQBP1, thus leading to an augment on immune response.
The function of LATS2 on immunity was once reported. LATS1/2 can either enhance or reduce immune response by upregulate IFNβ or suppress the secretion of immunity-inducing nucleic acids-containing exosomes, which is mediated by phosphorylation on YAP/TAZ [26]. Furthermore, YAP/TAZ can interact with TBK1 and IRF3 directly, inhibiting activation of TBK1 and blocking both RNA and DNA virus-induced immune response [27]. In our study, LATS2, but not LATS1, affects the interaction between PQBP1 and cGAS, thus regulates the cGAS activity, which is upstream of STING and TBK1/IRF3, thus independent of YAP/TAZ. The infection of HIV-1 has no significant effect on amounts of either total YAP/TAZ or their phosphorylated forms, further supporting an independent role of LATS2 on HIV-1 infection irrelevant to its activity on YAP/TAZ (Fig. S10).
As the core component of Hippo pathway in controlling cell proliferation, the new function of LATS2 we identified here uncovers a new function of LATS2 in innate immunity, thus establishes a direct link between the cell growth and immune response via LATS2. In nutrient-rich and growth-prone condition, LATS2 is inactivated and YAP/TAZ keep active to promote cell proliferation. Under stressful condition, LATS2 is activated. The activated LATS2, in one hand, as reported in canonical Hippo pathway, phosphorylates YAP/TAZ, enhances their cytosolic retention and degradation, thus leads to cell growth arrest; in the other hand, as revealed in this study, phosphorylates PQBP1 and promotes the cGAS activity and immune activation, thus switches the cells to fight against the infection. In either way, the LATS2 activation primes the cells for the unfavorable environment and benefits the cell survival. The imbalance between cell growth and cell immune response plays central roles in various cancers and immune diseases. Our study on LATS2-PQBP1-cGAS provides a new intervention target for fighting against these diseases.
Materials and methods
Cell culture and viral infection
HEK293T, HeLa, and THP1 cells were obtained from American Type Culture Collection. HEK293T and HeLa cells were cultured in DMEM (Corning) supplemented with 10% FBS (ExCell Bio). HEK293T and HeLa cells were transfected with plasmids using polyethylenimine (PEI, Polysciences) when cells were 60% confluent. The human monocyte-like THP1 cells were maintained in RPMI-1640 medium (Corning) supplemented with 10% FBS, and were differentiated into macrophage by treating with 50 ng/ml PMA (Phorbol myristate acetate) for 24 h, followed by culturing in completed RPMI-1640 medium for 24 h. Primary monocyte-derived macrophages (MDMs) were prepared by differentiation of monocytes from human PBMCs. PBMCs were isolated from buffy coats derived from fresh, healthy donor blood from Tianjin Second People’s Hospital (Tianjin, China) using human peripheral blood lymphocyte separation medium (Beijing Solarbio Science & Technology) as previously described [62]. Experiments involving human materials were approved by the academic committee of Nankai University and human bloods were collected after signing written informed consent by all donors. Primary human CD14+ monocytes were isolated by incubating with CD14+ Micro beads (STEMCELL Technologies) according to the manufacturer’s guidelines. The isolated CD14+ monocytes were subsequently differentiated into M1-like MDMs in IMDM medium (Gibco) supplemented with 10% FBS, 100 U/ml granulocyte-macrophage colony-stimulating factor (PeproTech) for 5 days. Primary MDMs were treated with either LATS2-shRNA lentivirus or scramble shRNA lentivirus for 24 h, then infected with VSV-G pseudotyped HIV-1 virus and VLP-VPX for 20 h as described in the literature [39, 63, 64]. Herpes simplex virus-1 (HSV-1) was kindly provided by Dr. Youjia Cao (Nankai University, Tianjin, China) and propagated in Vero cells. The titer of HSV-1 was measured by plaque assay. THP1 cells (1.2 × 106) were infected with HSV-1 (MOI = 5) for 6 h, and subjected to RNA extraction.
Reagents, antibodies, and constructs
HIV‐1 p24 antigen capture ELISA kit was produced by the Biomedical Engineering Center of Hebei Medical University (Shijiazhuang, China). 6×His-tagged Perfingolysin O (PFO) was expressed in Escherichia coli BL21 (DE3) strain (Novagen) and was purified by Ni-NTA Sepharose beads (GE Healthcare) as previously described [62]. The 6×His tag was cleaved by TEV protease and the mixture was loaded onto a Superdex 200 column (GE Healthcare) for further purification. Luciferase assay kit (Promega); Polybrene (Millipore); PMA (Sigma-Aldrich); 2′3′-cGAMP (InvivoGen); phosphatase inhibitor cocktail (Roche); puromycin (Solarbio) were purchased from the indicated manufacturers. Mouse monoclonal antibodies against enhanced GFP (eGFP; BGI), HA (Sigma-Aldrich), Flag (Sigma-Aldrich), Myc (Utibody), IRF3 (Santa Cruz Biotechnology), GAPDH (Utibody); rabbit antibodies against LATS2 (Proteintech Group), PQBP1 (Abcam), cGAS (Cell Signaling Technology), Thiophosphate ester γS (Abcam), Phosphoserine (Abcam), were purchased from the indicated manufacturers. HRP-conjugated anti-mouse and anti-rabbit IgG antibodies were purchased from Immunoway Biotechnology and Abclonal Technology, respectively.
Expression plasmids for FLAG- or GFP-tagged PQBP1 and its truncation mutants were constructed by standard molecular cloning techniques. PQBP1 mutants (S94A, S94D, S208A, S208D, S247A, and S247D) and CRISPR-Cas9 resistant mutants were generated by site-directed mutagenesis as described previously [62]. Expression plasmids for HA-, or Myc-tagged LATS2 and its truncation mutants were previously described [65]. cGAS and PQBP1 were constructed into a C-terminally eGFP-tagged eukaryotic expression vector pEGFP-N1. LATS2 was fused with a mCherry tag at the C terminus in eukaryotic expression pCherry vector. The human LATS2-shRNA constructs were generated using pLKO.1 vector (digested by AgeI and EcoRI). A scrambled, non-target shRNA plasmid SHC002 (Sigma-Aldrich) was used as negative control. The HIV-1 packaging plasmid NLENY1-ES-IRES (HIV-1 wild-type ΔEnv, encoding a YFP reporter) was kindly provided by Dr. David Levy (University of Alabama, Birmingham) [66]. The single-round HIV packaging plasmid pNL4.3-Luc E-R+ (ΔEnv, encoding firefly luciferase), pVSV-G, pQCXIN, pCMV-MLV-gag-pol, lentiCRISPR, and psPAX2 were gifts from Dr. Wentao Qiao (Nankai University).
Virus production and infection
The single-round HIV-1 virus NLENY1-ES-IRES or pNL4.3-Luc E-R+ were pseudotyped with VSV-G. HEK293T cells (~4.2 × 106) were seeded in 100 mm dishes 20 h before transfection. 4.8 µg NLENY1-ES-IRES or pNL4.3-Luc E-R+ and 1.2 µg pVSV-G were transfected into the cells by PEI (pH 7, 1 μg/ml) (the ratio of PEI to plasmid is 4). The medium was replaced with fresh one 6 h after transfection and the supernatants were harvested at 50 h by centrifugation for 10 min at 3000 rpm. The virus was filtered through 0.22 µm filters and incubated with lenti-virus concentration reagent (Biomega) at 4 °C overnight. The virus-containing pellet was obtained by centrifugation for 30 min at 3000 rpm. The pelleted particles were resuspended in 1 mL completed culture medium and stored in two aliquots at −80 °C. For the production of live virus, HEK293T cells were transfected with the HIV-1 proviral construct pNL4-3 using polyethyleneimine. After 48 h, the supernatant was harvested and the virus titer was determined according to the p24 antigen level in the supernatant.
The lentiviruses containing LATS2 shRNA were produced in HEK293T cells which were transfected with 5 µg psPAX2, 5 µg pLKO.1-LATS2 shRNA, and 2.5 µg pVSV-G. The VSV-G pseudotyped retrovirus was produced by Moloney murine leukemia virus-based retroviral vector system. The retrovirus was produced similarly to HIV-1 virus in HEK293T cells, which were transfected with 5 µg pCMV-MLV-gag-pol, 5 µg retroviral vector, and 2.5 µg pVSV-G. The supernatants were filtered and concentrated as described above.
Co-immunoprecipitation and immunoblot analysis
Cell lysates were prepared by sonication in Triton X-100 lysis buffer (1% Triton X-100; 20 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1 mM PMSF) and centrifuged at 12,000 rpm for 8 min at 4 °C. The lysates were incubated with GFP-Trap beads (for GFP-Trap co-immunoprecipitation; Chromotek) or the indicated antibody and protein A SefinoseTM Resin (for co-immunoprecipitation; Sangon Biotech) at 4 °C overnight. The beads were washed with ice-cold Triton X-100 lysis buffer four times and analyzed by immunoblotting with the indicated antibodies. Procedures of immunoblot analysis were done as previously described [62, 67].
Semi-denaturing detergent agarose gel electrophoresis (SDD-AGE)
The procedures of SDD-AGE were carried out as described in the literature with a minor modification [57, 68]. THP1 cells were lysed in NP-40 lysis buffer (20 mM Tris-HCl pH 8.0, 137 mM NaCl, 10% glycerol, 1% NP-40, 1 mM EDTA, and 1 mM PMSF) for 30 min on ice. Cell lysates were prepared by slight sonication and freeze-thaw three times in liquid nitrogen. Cell lysates were mixed with sample buffer (0.5× TBE, 10% glycerol, 2% SDS, and 0.0025% bromophenol blue) and loaded on a vertical 2% agarose gel. Proteins samples were separated in the running buffer (1× TBE and 0.1% SDS) with a constant voltage of 100 V at 4 °C, and transferred to a nitrocellulose membrane (Millipore) for immunoblotting analysis.
IRF3 dimerization assay
The procedures of native gel electrophoresis for detection of IRF3 dimerization were described previously [69,70,71]. THP1 cells (1.2 × 106) were infected with or without HIV-1 for 20 h in the presence of 8 µg/ml of polybrene. Cells were collected and lysed in native lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 1 mM EDTA and 1 mM PMSF) for 1 h on ice. Cell lysates were mixed with 5× sample buffer (62.5 mM Tris-HCl pH 6.8, 15% glycerol, and 1% sodium deoxycholate) and separated in the running buffer without SDS for 100 min at 20 mA on ice. The proteins bands were transferred onto a PVDF membrane and analyzed by immunoblotting with IRF3 antibody.
Real-time RT-PCR
Macrophage differentiated from THP1 and primary MDMs were uninfected or infected with VSV-G pseudotyped HIV-1 virus and VLP-Vpx for 20 h in the presence of polybrene by spinoculation (1000 rpm, 90 min). Cell lysates were prepared by Biozol reagent (Biomega) according to the manufacturer’s guidelines and mixed with chloroform followed with centrifugation (12,000 × g, 15 min, 4 °C). Total RNA was isolated by isopropanol from aqueous upper phase by centrifugation for 15 min at 12,000 rpm and eluted with RNase-free ddH2O. RNA samples (2 µg) were reverse transcribed using the 5× All-In-One RT MasterMix (Applied Biological Materials). Real-time RT-PCR was performed with TB Green Premix (Takara). The relative abundance of the indicated mRNA was normalized to the housekeeping gene GAPDH.
Real-time PCR analysis of HIV-1 reverse transcripts
THP1 cells were infected with VSV-G pseudotyped HIV-1 virus for the indicated time periods in the presence of polybrene (8 µg/ml). Cells were collected and gently rinsed twice with PBS. Total DNA was extracted by QIAamp DNA Blood Mini Kit (QIAGEN) according to the manufacturer’s protocol. HIV-1 reverse transcripts were quantified by real-time PCR as described in the literature [41, 42]. The different stages of HIV-1 reverse transcription were analyzed by comparing the products of reverse transcription as time progressing. The cellular HIV receptor gene CCR5 was used for normalization.
Generation of stable or knockout cell lines
For knockout experiments, the small guide RNA (sgRNA) sequence to target gene was taken from a lentiCRISPR-sgRNA library designed by Dr. Feng Zhang’s lab [72]. The sgRNA oligos were inserted into lentiCRISPR vector containing Cas9 and puromycin. Lentiviruses were produced in HEK293T cells transfected with 5 µg psPAX2, 5 µg lentiCRISPR-sgRNA, and 2.5 µg pVSV-G. For LATS2 overexpression experiments, THP1 cells were transduced with retroviral vector pQCXIN. The procedures of concentration and infection were described as above. Cells were selected with puromycin (2 mg/ml) after 48 h infection and subjected to efficiency detection by western blotting at day 10. LATS2-KO HEK293T cells were obtained by transiently transfected with 2 µg of pLentiCRISPR-LATS2-sgRNA #2 and cultured in presence of puromycin (2 mg/ml) after 48 h transfection. Monoclonal cell lines were selected by limited dilution method in 96-well plates and confirmed by sequencing and immunoblotting.
Measurement of cGAMP activity
The cGAMP activity assay was adapted from a protocol described in the literature [69, 73]. LATS2-KO THP1 cells were infected with HIV-1 for 20 h in the presence of polybrene and then homogenized by slight sonication in the hypotonic buffer (10 mM Tris-HCl pH 7.4, 10 mM KCl, and 1.5 mM MgCl2). Cytosolic extracts were “squeezed out” by centrifugation at 100,000 rpm for 10 min and heated at 95 °C for 5 min. After centrifugation at 12,000 rpm for 5 min, the heat-resistant supernatant containing cGAMP was mixed with wild-type THP1 in reaction buffer (2 mM ATP, 1 U/μl Benzonase and 1.5 ng/μl PFO) and incubated at 30 °C for 1 h. Then, the cells were cultured with completed RPMI-1640 medium for another 6 h before real-time RT-PCR analysis. The concentration of cGAMP in the cell extracts was also measured by a 2′3′-cGAMP ELISA kit (Cayman Chemical).
In vitro kinase assay
LATS2 in vitro kinase assay was performed as described in the literature [74, 75]. The recombinant 6×His-PQBP1 and its mutants were purified from E. coli BL21 (DE3) as described above. The kinase LATS2 was immunoprecipitated from HEK293T cells transfected with LATS2 eukaryotic expression plasmid. After 48 h, the cells were harvested and lysed in ice-cold Triton X-100 lysis buffer with inhibitor (phosphatase inhibitor cocktail 1/2 and complete protease inhibitor cocktail). 6×His-PQBP1 (1 µg) was incubated with immunoprecipitated LATS2 in the presence of ATP-γS (100 ng, Sigma-Aldrich) in 30 µl kinase buffer (25 mM Tris-HCl, pH 7.5, 5 mM β-glycerophosphate, 2 mM DTT, 0.1 mM Na3VO4 and 10 mM MgCl2) for 30 min with gentle rocking. The reaction was stopped by adding in 0.1 mM EDTA, then the thiophospholyation site was alkylated by pNitrobenzyl mesylate (PNBM, 2.5 mM; Abcam) for 2 h with mild shaking. The integration of γS on PQBP1 was detected by western blotting with thiophosphate ester antibody (Abcam).
Confocal microscopy
HeLa or THP1 cells were seeded on coverslips in 12-well plate. The cells were transfected with fluorescent plasmids or infected with indicated retrovirus as in the figures, then fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.1% Triton X-100 for 10 min and blocked in 3% BSA for 2 h at room temperature. For immunofluorescence assay, cell slides were sequentially incubated with indicated primary antibodies and fluorescent-labeled secondary antibodies for 2 h at room temperature. Then cells were stained with DAPI in mounting medium and images were acquired using confocal microscope (Leica TCS SP5).
Quantification and statistical analysis
Statistical analysis was performed using Student’s unpaired t-test or analysis of variance with GraphPad Prism 8.0 (Graph Pad Software). The data were presented as the mean ± SD from at least three independent experiments. Differences were considered significant at ∗p < 0.05, and reported by asterisks in the figures and figure legends.
The oligoes used in this study were summarized in Table S2.
References
Harvey KF, Pfleger CM, Hariharan IK. The Drosophila Mst ortholog, hippo, restricts growth and cell proliferation and promotes apoptosis. Cell. 2003;114:457–67.
Jia J, Zhang W, Wang B, Trinko R, Jiang J. The Drosophila Ste20 family kinase dMST functions as a tumor suppressor by restricting cell proliferation and promoting apoptosis. Genes Dev. 2003;17:2514–9.
Pantalacci S, Tapon N, Leopold P. The Salvador partner Hippo promotes apoptosis and cell-cycle exit in Drosophila. Nat Cell Biol. 2003;5:921–7.
Udan RS, Kango-Singh M, Nolo R, Tao C, Halder G. Hippo promotes proliferation arrest and apoptosis in the Salvador/Warts pathway. Nat Cell Biol. 2003;5:914–20.
Wu S, Huang J, Dong J, Pan D. hippo encodes a Ste-20 family protein kinase that restricts cell proliferation and promotes apoptosis in conjunction with salvador and warts. Cell. 2003;114:445–56.
Huang J, Wu S, Barrera J, Matthews K, Pan D. The Hippo signaling pathway coordinately regulates cell proliferation and apoptosis by inactivating Yorkie, the Drosophila Homolog of YAP. Cell. 2005;122:421–34.
Lai ZC, Wei X, Shimizu T, Ramos E, Rohrbaugh M, Nikolaidis N, et al. Control of cell proliferation and apoptosis by mob as tumor suppressor, mats. Cell. 2005;120:675–85.
Goulev Y, Fauny JD, Gonzalez-Marti B, Flagiello D, Silber J, Zider A. SCALLOPED interacts with YORKIE, the nuclear effector of the hippo tumor-suppressor pathway in Drosophila. Curr Biol. 2008;18:435–41.
Wu S, Liu Y, Zheng Y, Dong J, Pan D. The TEAD/TEF family protein Scalloped mediates transcriptional output of the Hippo growth-regulatory pathway. Dev Cell. 2008;14:388–98.
Zhang L, Ren F, Zhang Q, Chen Y, Wang B, Jiang J. The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev Cell. 2008;14:377–87.
Overholtzer M, Zhang J, Smolen GA, Muir B, Li W, Sgroi DC, et al. Transforming properties of YAP, a candidate oncogene on the chromosome 11q22 amplicon. Proc Natl Acad Sci USA. 2006;103:12405–10.
Dong JX, Feldmann G, Huang JB, Wu S, Zhang NL, Comerford SA, et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell. 2007;130:1120–33.
Ota M, Sasaki H. Mammalian Tead proteins regulate cell proliferation and contact inhibition as transcriptional mediators of Hippo signaling. Development. 2008;135:4059–69.
Zhao B, Wei X, Li W, Udan RS, Yang Q, Kim J, et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Gene Dev. 2007;21:2747–61.
Zhao B, Ye X, Yu J, Li L, Li W, Li S, et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 2008;22:1962–71.
Sebe-Pedros A, Zheng Y, Ruiz-Trillo I, Pan D. Premetazoan origin of the hippo signaling pathway. Cell Rep. 2012;1:13–20.
Zheng Y, Pan D. The Hippo signaling pathway in development and disease. Dev Cell. 2019;50:264–82.
Liu B, Zheng YG, Yin F, Yu JZ, Silverman N, Pan DJ. Toll receptor-mediated Hippo signaling controls innate immunity in Drosophila. Cell. 2016;164:406–19.
Boro M, Singh V, Balaji KN. Mycobacterium tuberculosis-triggered Hippo pathway orchestrates CXCL1/2 expression to modulate host immune responses. Sci Rep. 2016;6:37695.
Wang S, Xie F, Chu F, Zhang ZK, Yang B, Dai T, et al. YAP antagonizes innate antiviral immunity and is targeted for lysosomal degradation through IKK epsilon-mediated phosphorylation. Nat Immunol. 2017;18:733–43.
Meng FS, Zhou RY, Wu SY, Zhang Q, Jin QH, Zhou Y, et al. Mst1 shuts off cytosolic antiviral defense through IRF3 phosphorylation. Gene Dev. 2016;30:1086–100.
Nehme NT, Schmid JP, Debeurme F, Andre-Schmutz I, Lim A, Nitschke P, et al. MST1 mutations in autosomal recessive primary immunodeficiency characterized by defective naive T-cell survival. Blood. 2012;119:3458–68.
Abdollahpour H, Appaswamy G, Beier R, Schaffer AA, Gertz EM, Kreipe HH, et al. The phenotype of human STK4 deficiency. Blood. 2011;118:316–7.
Zhou DW, Medoff BD, Chen LF, Li LQ, Zhang XF, Praskova M, et al. The Nore1B/Mst1 complex restrains antigen receptor-induced proliferation of naive T cells. Proc Natl Acad Sci USA. 2008;105:20321–6.
Dong YL, Du XR, Ye J, Han M, Xu T, Zhuang Y, et al. A cell-intrinsic role for Mst1 in regulating thymocyte egress. J Immunol. 2009;183:3865–72.
Moroishi T, Hayashi T, Pan WW, Fujita Y, Holt MV, Qin J, et al. The Hippo pathway kinases LATS1/2 suppress cancer immunity. Cell. 2016;167:1525–39. e1517.
Zhang Q, Meng FS, Chen SS, Plouffe SW, Wu SY, Liu SD, et al. Hippo signalling governs cytosolic nucleic acid sensing through YAP/TAZ-mediated TBK1 blockade. Nat Cell Biol. 2017;19:362–74.
Geng J, Sun XF, Wang P, Zhang SH, Wang XZ, Wu HT, et al. Kinases Mst1 and Mst2 positively regulate phagocytic induction of reactive oxygen species and bactericidal activity. Nat Immunol. 2015;16:1142–52.
Thompson BJ, Sahai E. MST kinases in development and disease. J Cell Biol. 2015;210:871–82.
Lehtinen MK, Yuan ZQ, Boag PR, Yang Y, Villen J, Becker EBE, et al. A conserved MST-FOXO signaling pathway mediates oxidative-stress responses and extends life span. Cell. 2006;125:987–1001.
Wilkinson DS, Hansen M. LC3 is a novel substrate for the mammalian Hippo kinases, STK3/STK4. Autophagy. 2015;11:856–7.
Zanconato F, Cordenonsi M, Piccolo S. YAP/TAZ at the roots of cancer. Cancer Cell. 2016;29:783–803.
Meng ZP, Moroishi T, Mottier-Pavie V, Plouffe SW, Hansen CG, Hong AW, et al. MAP4K family kinases act in parallel to MST1/2 to activate LATS1/2 in the Hippo pathway. Nat. Commun. 2015;6:8357.
Meng ZP, Moroishi T, Guan KL. Mechanisms of Hippo pathway regulation. Gene Dev. 2016;30:1–17.
Waragai M, Junn E, Kajikawa M, Takeuchi S, Kanazawa I, Shibata M, et al. PQBP-1/Npw38, a nuclear protein binding to the polyglutamine tract, interacts with US-15kD/dim1p via the carboxyl-terminal domain. Biochem Bioph Res Commun. 2000;273:592–5.
Germanaud D, Rossi M, Bussy G, Gerard D, Hertz-Pannier L, Blanchet P, et al. The Renpenning syndrome spectrum: new clinical insights supported by 13 new PQBP1-mutated males. Clin Genet. 2011;79:225–35.
Kalscheuer VM, Freude K, Musante L, Jensen LR, Yntema HG, Gecz J, et al. Mutations in the polyglutamine binding protein 1 gene cause X-linked mental retardation. Nat Genet. 2003;35:313–5.
Stevenson RE, Bennett CW, Abidi F, Kleefstra T, Porteous M, Simensen RJ, et al. Renpenning syndrome comes into focus. Am J Med Genet A. 2005;134a:415–21.
Yoh SM, Schneider M, Seifried J, Soonthornvacharin S, Akleh RE, Olivieri KC, et al. PQBP1 Is a proximal sensor of the cGAS-dependent innate response to HIV-1. Cell. 2015;161:1293–305.
Shannon JL, Murphy MS, Kantheti U, Burnett JM, Hahn MG, Dorrity TJ, et al. Polyglutamine binding protein 1 (PQBP1) inhibits innate immune responses to cytosolic DNA. Mol Immunol. 2018;99:182–90.
Mbisa JL, Delviks-Frankenberry KA, Thomas JA, Gorelick RJ, Pathak VK. Real-time PCR analysis of HIV-1 replication post-entry events. Methods Mol Biol. 2009; 485:55–72.
Guo HT, Koenig R, Deng M, Riess M, Mo JY, Zhang L, et al. NLRX1 sequesters STING to negatively regulate the interferon response, thereby facilitating the replication of HIV-1 and DNA viruses. Cell Host Microbe. 2016;19:515–28.
Kane M, Yadav SS, Bitzegeio J, Kutluay SB, Zang T, Wilson SJ, et al. MX2 is an interferon-induced inhibitor of HIV-1 infection. Nature. 2013;502:563–6.
St Gelais C, Wu L. SAMHD1: a new insight into HIV-1 restriction in myeloid cells. Retrovirology. 2011;8:55.
Gao D, Wu J, Wu Y-T, Du F, Aroh C, Yan N, et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science. 2013;341:903.
Ishikawa H, Barber GN. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature. 2008;455:674–8.
Zhong B, Yang Y, Li S, Wang YY, Li Y, Diao F, et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity. 2008;29:538–50.
Sun W, Li Y, Chen L, Chen H, You F, Zhou X, et al. ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc Natl Acad Sci USA. 2009;106:8653–8.
Jin L, Hill KK, Filak H, Mogan J, Knowles H, Zhang B, et al. MPYS is required for IFN response factor 3 activation and type I IFN production in the response of cultured phagocytes to bacterial second messengers cyclic-di-AMP and cyclic-di-GMP. J Immunol. 2011;187:2595–601.
Liu S, Cai X, Wu J, Cong Q, Chen X, Li T, et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science. 2015;347:aaa2630.
Li YF, Pei J, Xia H, Ke HN, Wang HY, Tao WF. Lats2, a putative tumor suppressor, inhibits G1/S transition. Oncogene. 2003;22:4398–405.
Yu T, Bachman J, Lai ZC. Mutation analysis of large tumor suppressor genes LATS1 and LATS2 supports a tumor suppressor role in human cancer. Protein Cell. 2015;6:6–11.
Olsen JV, Vermeulen M, Santamaria A, Kumar C, Miller ML, Jensen LJ, et al. Quantitative phosphoproteomics reveals widespread full phosphorylation site occupancy during mitosis. Sci Signal. 2010;3:ra3.
Dephoure N, Zhou C, Villen J, Beausoleil SA, Bakalarski CE, Elledge SJ, et al. A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci USA. 2008;105:10762–7.
Li X, Shu C, Yi GH, Chaton CT, Shelton CL, Diao JS, et al. Cyclic GMP-AMP synthase is activated by double-stranded dna-induced oligomerization. Immunity. 2013;39:1019–31.
Lian H, Wei J, Zang R, Ye W, Yang Q, Zhang XN, et al. ZCCHC3 is a co-sensor of cGAS for dsDNA recognition in innate immune response. Nat Commun. 2018;9:3349.
Alberti S, Halfmann R, King O, Kapila A, Lindquist S. A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell. 2009;137:146–58.
Taha Z, van Rensburg HJJ, Yang XL. The hippo pathway: immunity and cancer. Cancers. 2018;10:94.
Yamauchi T, Moroishi T. Hippo pathway in mammalian adaptive immune system. Cells. 2019;8:398.
Goujon C, Moncorge O, Bauby H, Doyle T, Ward CC, Schaller T, et al. Human MX2 is an interferon-induced post-entry inhibitor of HIV-1 infection. Nature. 2013;502:559–62.
Wang QQ, Moore MJ, Adelmant G, Marto JA, Silver PA. PQBP1, a factor linked to intellectual disability, affects alternative splicing associated with neurite outgrowth. Gene Dev. 2013;27:615–26.
He TS, Ji W, Zhang J, Lu J, Liu X. ALG-2 couples T cell activation and apoptosis by regulating proteasome activity and influencing MCL1 stability. Cell Death Dis. 2020;11:5.
Manel N, Hogstad B, Wang Y, Levy DE, Unutmaz D, Littman DR. A cryptic sensor for HIV-1 activates antiviral innate immunity in dendritic cells. Nature. 2010;467:214–7.
Laguette N, Sobhian B, Casartelli N, Ringeard M, Chable-Bessia C, Segeral E, et al. SAMHD1 is the dendritic- and myeloid-cell-specific HIV-1 restriction factor counteracted by Vpx. Nature. 2011;474:654–U132.
Ma B, Chen Y, Chen L, Cheng HC, Mu CL, Li J, et al. Hypoxia regulates Hippo signalling through the SIAH2 ubiquitin E3 ligase. Nat Cell Biol. 2015;17:95–+.
Levy DN, Aldrovandi GM, Kutsch O, Shaw GM. Dynamics of HIV-1 recombination in its natural target cells. Proc Natl Acad Sci. USA. 2004;101:4204–9.
He TS, Xie T, Li J, Yang YX, Li C, Wang W, et al. THO complex subunit 7 homolog negatively regulates cellular antiviral response against RNA viruses by targeting TBK1. Viruses. 2019;11:158.
Lian H, Wei J, Zang R, Ye W, Yang Q, Zhang XN, et al. ZCCHC3 is a co-sensor of cGAS for dsDNA recognition in innate immune response. Nat Commun. 2018;9:3349.
Gao DX, Wu JX, Wu YT, Du FH, Aroh C, Yan N, et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science. 2013;341:903–6.
He TS, Chen T, Wang DD, Xu LG. HAUS8 regulates RLRVISA antiviral signaling positively by targeting VISA. Mol Med Rep. 2018;18:2458–66.
Robitaille AC, Mariani MK, Fortin A, Grandvaux N. A high resolution method to monitor phosphorylation-dependent activation of IRF3. J Vis Exp. 2016;107:e53723.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343:84–7.
Wu JX, Sun LJ, Chen X, Du FH, Shi HP, Chen C, et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science. 2013;339:826–30.
Allen JJ, Li M, Brinkworth CS, Paulson JL, Wang D, Hubner A, et al. A semisynthetic epitope for kinase substrates. Nat Methods. 2007;4:511–6.
Guo JP, Dai XP, Laurent B, Zheng NN, Gan WJ, Zhang J, et al. AKT methylation by SETDB1 promotes AKT kinase activity and oncogenic functions. Nat Cell Biol. 2019;21:226–37.
Acknowledgements
We thank Dr. David Levy (University of Alabama at Birmingham), Dr. Wentao Qiao (Nankai University), and Dr. Min Wei (Nankai University) for providing retrovirus plasmids. We also thank Dr. Youjia Cao (Nankai University) for providing HSV-1. This project is supported by the National Natural Science Foundation of China (31870730, 31671513, and 82002136).
Author information
Authors and Affiliations
Contributions
SW and XL conceived the project. TSH and XL designed the experiments. TSH, LD, JZ, JZ, GW, and EW performed the experiments. HX provided essential materials. TSH and XL analyzed the data and wrote the paper. WZ, SW, and XL revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Our studies involving human materials were approved by the academic committee of Nankai University. Human blood samples were collected after signing written informed consent by all donors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Piacentini
Rights and permissions
About this article
Cite this article
He, TS., Dang, L., Zhang, J. et al. The Hippo signaling component LATS2 enhances innate immunity to inhibit HIV-1 infection through PQBP1-cGAS pathway. Cell Death Differ 29, 192–205 (2022). https://doi.org/10.1038/s41418-021-00849-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-021-00849-1
This article is cited by
-
E3 ubiquitin ligase RNF128 negatively regulates the IL-3/STAT5 signaling pathway by facilitating K27-linked polyubiquitination of IL-3Rα
Cell Communication and Signaling (2024)
-
The Hippo signalling pathway and its implications in human health and diseases
Signal Transduction and Targeted Therapy (2022)