Abstract
Germinal center (GC) B cells are among the fastest replicating cells in our body, dividing every 4–8 h. DNA replication errors are intrinsically toxic to cells. How GC B cells exert control over the DNA damage response while introducing mutations in their antibody genes is poorly understood. Here, we show that the DNA damage response regulator Checkpoint kinase 1 (CHK1) is essential for GC B cell survival. Remarkably, effective antibody-mediated immunity relies on optimal CHK1 dosage. Chemical CHK1 inhibition or loss of one Chk1 allele impairs the survival of class-switched cells and curbs the amplitude of antibody production. Mechanistically, active B cell receptor signaling wires the outcome of CHK1-inhibition towards BIM-dependent apoptosis, whereas T cell help favors temporary cell cycle arrest. Our results predict that therapeutic CHK1 inhibition in cancer patients may prove potent in killing B cell lymphoma and leukemia cells addicted to B cell receptor signaling, but will most likely dampen humoral immunity.
Similar content being viewed by others
Introduction
Mature B cells predominate in a quiescent state until engaged by antigen via their B cell antigen receptor (BCR). Upon infection or vaccination, B cells undergo T cell-dependent clonal expansion in secondary lymphoid tissues, a process termed the germinal center (GC) reaction [1,2,3].
GC B cells repeatedly traffic between two distinct compartments [4,5,6]. In the dark zone (DZ) GC centroblasts divide and introduce point mutations in their BCR gene variable region. Mutagenesis is initiated by activation-induced cytidine deaminase (AID) and completed by error-prone DNA repair [7, 8]. This somatic hypermutation (SHM) generates a pool of cells carrying related BCRs with improved affinities to the activating antigen. In addition, conversion of AID-initiated lesions into double-strand breaks (DSB) permits class switch recombination (CSR), exchanging the BCR gene constant region. In the light zone (LZ), GC centrocytes are positively selected through BCR:antigen binding, providing a competitive advantage to B cells carrying a high-affinity BCR for stable interaction with cognate follicular helper T cells [4, 5]. T cell-derived signals enhance survival and provide contextual cues to the nature of the immunogen through receptor ligands (e.g. CD40–CD40L) or cytokines (e.g. IL-4, IL-21) [9]. T cell help promotes proliferation at maximum speed [10], initiates CSR, and allows GC-exit and differentiation into memory B cells or antibody-secreting cells (ASCs), i.e. plasmablasts/plasma cells [11, 12].
On average, GC B cells spend ~4 h in the S-phase and ~1.5 h in the G2–M phases, dividing once every 4–8 h [4, 6, 13]. DNA replication is intrinsically error prone, with replication fork progression being susceptible to impediments in DNA, interference with the transcription machinery, or impaired nucleotide supply. Replication fork stalling exposes single-stranded DNA, which poses a risk for collapse into DSB. Impairments in replication fork progression/stability are commonly referred to as replication stress [14]. AID-initiated on- or off-target DSBs may persist into the S-phase, increase replication stress, and accidentally cause mutations or translocations [15]. However, many recurrent mutations in GC B cell lymphomas are not associated with AID activity [16], implying mutational potency of replication stress.
Cellular mechanisms that recognize replication stress and support DNA repair or the removal of irreversibly damaged cells are likely critical for GC homeostasis and a barrier to transformation. Early in the DNA damage response, Ataxia telangiectasia mutated-related kinase (ATR) is recruited to and activated at junctions of single- and double-stranded DNA [17]. ATR phosphorylates its major substrate kinase CHK1 at Ser-317 and Ser-345. Active CHK1 targets the cyclin-dependent kinase activators CDC25A/C for degradation or nuclear export, promoting cell cycle arrest in the S-phase or at the G2–M boundary. Furthermore, CHK1 activates WEE1 to decelerate G2-progression. CHK1-mediated cell cycle arrest provides time for DNA repair, while also promoting replication fork stabilization and restart [14]. Failed restoration of faithful S-phase progression mostly culminates in apoptosis.
In GC B cells, the transcriptional repressor BCL6 suppresses DNA damage signaling through dampening ATR, CHK1, p53, and p21 [18,19,20,21]. These observations have led to the assumption that GC B cells are programmed to “tolerate” DNA damage. Per contra, survival in the context of rapid proliferation and recurrent mutagenesis requires efficient DNA damage control. Accordingly, T cell help was shown to transiently rescue ATR and CHK1 from transcriptional repression when GC B cells enter the proliferative DZ [18].
To elucidate the role of CHK1 in this delicate balance, we generated a mouse model of conditional CHK1 ablation in GC B cells, and report a vital role for this checkpoint kinase in GC maintenance.
Results
CHK1-mediated survival of in vitro-activated B cells depends on the nature of the mitogen
The rapid proliferation of B cells upon activation may create a strong dependence on ATR-CHK1 activity for survival. To differentiate the effects of key mitogenic signals on CHK1 activity, murine splenic B cells were cultivated in vitro in the presence of F(ab)2 αIgM-fragments ± IL-4 to stimulate BCR-activity, or αCD40 ± IL-4 ± IL-21 to mimic T cell help.
All stimuli led to proliferation (Fig. 1a) and substantial ATR phosphorylation in actively cycling (S–G2–M) cells (Fig. 1b), suggesting replication stress. Whereas CHK1 mRNA was robustly induced by signals mimicking T cell help, CHK1 protein accumulated without a notable increase in transcription when BCR-signaling was triggered, indicating post-translational control (Fig. 1c, d). The extent of CHK1 phosphorylation (Ser-345), a proxy for its activity, was also more pronounced when subjecting cells to signals jointly mimicking T cell help (Fig. 1d).
CHK1-mediated survival of in vitro-activated B cells depends on the nature of the mitogen. a Murine splenic wild-type B cells were loaded with eFluor450 cell proliferation dye and stimulated with the indicated mitogens inducing BCR-signaling or mimicking T cell help, respectively. After 72 h, cells were analyzed for proliferation by flow cytometry as indicated by the division-dependent loss of the proliferation dye over time. Filled gray peaks represent un-stimulated (non-dividing) cells that were cultured for 72 h. Data are representative for three mice per genotype. b Contour plots depict intracellular flow cytometry staining for DNA content by TO-PRO-3 and for pATR within the S–G2–M population of the cultures described in a. Data are representative for three mice per genotype. c qRT-PCR analysis of CHK1 mRNA expression in wild-type B cells directly after isolation (naïve ex vivo) or after 48 h of cultivation without (naïve in vitro) or with mitogenic stimulation as indicated. The data are presented as means ± SD and represent 3–6 mice per cell population. d Immunoblot analysis for CHK1 and pCHK1 in wild-type B cells directly after isolation (naïve ex vivo) or after 48 h of cultivation without (naïve in vitro) or with mitogenic stimulation as indicated. Western blot is representative of four independent experiments. e Splenic wild-type B cells were left untreated or stimulated with the indicated mitogens. After 48 h, the cells were treated with vehicle or graded doses of either of the two CHK1-inhibitors PF-477736 (25, 100, 400, 1600, and 6400 nM) or CHIR-124 (0.38, 1.5, 6.25, 25, 100, and 400 nM), and analyzed 24 h later for Annexin V−/TO-PRO-3− viable cells by flow cytometry. Survival is depicted normalized to the survival of the vehicle-treated culture, and hence termed “survival (% of control)”. Data are cumulative from three experiments with three mice per concentration, and presented as log(inhibitor) vs. response curves
Curiously, 48 h cultivation of B cells without stimulation caused CHK1 mRNA and protein up-regulation as compared to ex vivo-analyzed B cells (Fig. 1c, d). Therefore, CHK1 activity may not always be linked to cell proliferation but might also control the survival of resting cells.
Next, we addressed to what extent the survival of mitogen-stimulated B cells depends on CHK1 function. Pharmacological CHK1 inhibitors (CHK1i), several of which have been approved for clinical trials, result in excessive DNA damage and trigger apoptosis [22]. Hence, we cultivated splenic B cells for 72 h in the absence (naïve) or presence of various mitogens, and treated theses cells after 48 h with graded doses of the CHK1 inhibitors P-477736 and CHIR-124. Generally, cell viability was reduced in a dose-dependent manner (Fig. 1e, Supplementary Figure 1). However, whereas signals mimicking T cell help did not render B cells significantly more susceptible to CHK1-inhibition as compared to non-stimulated naïve B cells, BCR-ligation profoundly sensitized B cells to CHK1i. Altogether, these data suggest that ATR-CHK1 signaling is commonly induced in proliferating B cells. However, the magnitude of the apoptotic response to CHK1i depends on the mitogenic stimulus, and is not a consequence of cell proliferation per se.
Mitogenic signals differentially instruct CHK1 function
The finding that the magnitude of CHK1i-induced apoptosis is mitogen-dependent prompted us to ask whether differences in cell division rates might underlie this phenomenon. Surprisingly, despite the average number of cell divisions differing between mitogens, proliferation speed and sensitivity to CHK1i-induced apoptosis did not correlate (Fig. 1a–e, Fig. 2a, left panel, Supplementary Figure 2a, b, left panel).
Mitogenic signals differentially instruct CHK1 function. a Murine splenic wild-type B cells were loaded with eFluor450 cell proliferation dye and stimulated with the indicated mitogens. After 48 h, cells were treated with vehicle or with the indicated doses of PF-477736 and CHIR-124 for 24 h and analyzed by flow cytometry for the loss of the proliferation dye (left panel) within Annexin V−/TO-PRO-3− viable cells. Furthermore, murine splenic wild-type B cells were stimulated with the indicated mitogens, and after 48 h treated with vehicle or with the indicated doses of PF-477736 and CHIR-124 for 24 h, fixed and stained with DAPI for cell cycle analysis (middle panel), or fixed and stained with a phospho-Histone H3 antibody in combination with DAPI (right panel). Data are cumulative from three experiments, with each symbol representing one wild-type mouse (n = 3–6 for all data points), and shown as mean ± SEM. **p < 0.01, ***p < 0.005, ****p < 0.0001 (one-way ANOVA followed by Tukey post-hoc test) as compared to the respective vehicle-treated control
BCR-ligation sensitized towards immediate apoptosis already at the lowest nanomolar CHK1i concentration tested (Fig. 1e, Supplementary Figure 1), precluding analysis at higher concentrations. For the lowest inhibitor concentrations used, no signs of cell cycle arrest were observed upon BCR-ligation, suggesting that these cells stand no chance to arrest and quickly die (Fig. 2a, Supplementary Figure 2b, Supplementary Figure 2c). In sharp contrast, mitogenic signals mimicking T cell help did not significantly sensitize to apoptosis above the levels observed in non-cycling B cells in vitro (Fig. 1e, Supplementary Figure 1), and wired the cellular response towards S–G2 cell cycle arrest, as indicated by a pronounced reduction of mitotic phospho-Histone 3+ cells already at the lowest CHK1i concentration (Fig. 2a, Supplementary Figure 2b, central/right panel). Moderately increased CHK1i-dosage enforced this S–G2 arrest.
In cancer cells, CHK1i trigger the accumulation of single-stranded DNA, excessive origin firing, replication fork stalling, DSB, and consequently apoptosis [22]. Therefore, we speculated that CHK1i-treated proliferating B cells should similarly present with increased DNA damage. Histone 2A is phosphorylated by ATR at S-139 (γ-H2AX) only seconds after DSB formation. Performing intracellular flow cytometry, we found that both, BCR-ligation and signals that mimic joint T cell help caused excessive accumulation of DSB while naïve cultivated B cells did not show such an increase (Fig. 3a, b, Supplementary Figure 3a). Enhanced accumulation of γ-H2AX+ cells upon CHK1i in cultures driven by signals mimicking T cell help in the absence of increased cell death is indicative of increased tolerance to genotoxic damage, and in line with the observed S–G2 arrest in Fig. 2.
Mitogenically stimulated B cells accumulate genotoxic damage that is further increased upon CHK1i in a stimulus-dependent manner. a Murine splenic wild-type B cells were stimulated with the indicated mitogens. After 48 h, cells were treated with vehicle or with the indicated doses of PF-477736 and CHIR-124 for 24 h, fixed and stained with a γ-H2AX antibody in combination with TO-PRO-3. Data are cumulative from three experiments, with each symbol representing one wild-type mouse (n = 3–6 for all data points), and shown as mean ± SEM. b Representative contour plots of the analysis shown in a. **p < 0.01, ***p < 0.005, ****p < 0.0001 (one-way ANOVA followed by Tukey post-hoc test) as compared to the respective vehicle-treated control
Although AID may contribute to the DNA damage observed during T cell help mimicking stimulation, AID is not up-regulated by BCR-signaling and hence does not contribute to the genotoxic stress that sensitizes these cells to CHK1i upon BCR-ligation (Supplementary Figure 3b).
In summary, BCR-signaling generates a specific vulnerability to CHK1i-induced apoptosis whereas T cell mimicking signals allow B cells to respond with cell cycle arrest, potentially to facilitate AID-mediated mutagenesis.
BCR-ligation primes activated B cells for BIM-induced apoptosis upon CHK1 inhibition
The pro-apoptotic BCL2-interacting mediator of cell death (BIM) is key for developmental apoptosis in B cells [23,24,25], and has been reported to aid the elimination of B cell progenitors carrying damaged DNA [26,27,28]. BCL2-family proteins oppose each other either by initiating (BH3-only proteins such as BIM) or inhibiting (e.g. BCL2, BCLXL, MCL1, BCL2a1) mitochondrial apoptosis through complex protein interactions [29].
Despite comparable apoptosis rates of wild-type B cells stimulated with BCR-ligation or signals mimicking T cell help (Supplementary Figure 1), BIM protein levels reached a maximum in cells undergoing BCR-ligation (Fig. 4a). BCR-ligation led to a general reprogramming of the cellular composition of anti-apoptotic BCL2-proteins, with BCL2 being repressed and BCLXL as well as BCL2a1 becoming induced. MCL1 levels were comparable between stimuli. Based on current literature [30], we speculated that active BCR-signaling may prime B cells for apoptosis via increasing BIM levels. Cells primed for death are exquisitely sensitive to apoptotic triggers. Hence we wondered whether apoptotic priming upon BCR-ligation may sensitize B cells to BIM-mediated apoptosis upon CHK1 inhibition. To test this hypothesis, we subjected B cells isolated from Bim−/− mice to CHK1i. BIM-deficiency substantially rescued the synergistic lethality of low-dose CHK1i and BCR-ligation while having no further protective impact on cells receiving CHK1i in combination with stimuli mimicking T cell help (Fig. 4b, Supplementary Figure 4a). Increased survival of BIM-deficient cells did not cause changes in total or phosphorylated CHK1 levels (Supplementary Figure 4b). Thus, we asked whether these aberrantly surviving cells would undergo cell cycle arrest, comparable to cells receiving signals mimicking T cell help. However, we did not observe signs of S–G2 arrest upon BCR-ligation in BIM-deficient cells (Fig. 4c, Supplementary Figure 4c).
BCR-ligation primes activated B cells for BIM-induced apoptosis upon CHK1 inhibition. a Immunoblot analysis for BCL2-proteins in wild-type B cells directly after isolation (naïve ex vivo) or after 48 h of cultivation with mitogenic stimuli as indicated. Western blot is representative of two independent experiments. b Splenic wild-type or Bim−/− B cells were stimulated with the indicated mitogens. After 48 h, the cells were vehicle-treated or treated with low-dose CHK1i as indicated, and analyzed 24 h later for Annexin V−/TO-PRO-3− viable cells by flow cytometry. Survival is depicted normalized to the survival of the vehicle-treated culture, and termed “survival (% of control)”. Data are cumulative from three experiments (n = 3 for all data points) and shown as mean ± SEM. c Splenic Bim−/− B cells were left untreated or stimulated with the indicated mitogens. After 48 h, cells were treated with vehicle or with the indicated doses of PF-477736 and CHIR-124 for 24 h, fixed and stained with DAPI for cell cycle analysis. Data are cumulative from three experiments (n = 3 for all data points) and shown as mean ± SEM. *p < 0.05, **p < 0.01, ****p < 0.0001 (one-way ANOVA followed by Tukey post-hoc test)
In summary, CHK1i-mediated cell death upon BCR-ligation largely depends on BIM. In vivo, CHK1 function may counteract GC B cell apoptosis initiated by strong BCR-ligation, potentially promoting the survival of autoreactive cells.
CHK1 function supports BCL2-mediated cell survival during CSR
GC B cell proliferation and S-phase entry are essentially involved in both, AID-mediated CSR and the differentiation into ASCs [31, 32]. Consequently, class-switched B cells and ASCs accumulate stochastically with each division cycle [33]. We used crosslinking of CD40 under saturating IL-4 or IL-4/IL-21 concentrations in vitro to induce isotype-switching to IgG1, or the differentiation into mitotically active CD138+ ASCs. Under these conditions, low-dose CHK1i led to cell cycle arrest and only moderate apoptosis in the bulk culture (Fig. 1e, Supplementary Figure 1).
When compared to vehicle-treated controls, already low nanomolar concentrations of CHK1i caused a severe and specific reduction in IgG1+ cells (Fig. 5a, b, Supplementary Figure 5a). Quantifying the fraction of IgG1+ cells relative to each cell division cycle, we could exclude that the loss of IgG1+ cells is a consequence of CHK1i-induced cell cycle arrest (Fig. 5b, Supplementary Figure 5b). Instead, we noted a specific loss of IgG1+ cells on a per-division basis throughout consecutive division cycles. Of note, BIM-deficiency rescued this specific loss of IgG1+ cells upon low-dose CHK1i (Fig. 5c). These results suggest that CHK1 safeguards B cells in the presence of physiological DSB generated during CSR. In contrast to CSR, plasmacytic differentiation was not significantly impaired upon CHK1i (Fig. 5d, Supplementary Figure 5c).
CHK1 function supports BCL2-mediated cell survival during class switch recombination. a Wild-type B cells were loaded with cell proliferation dye eF450, stimulated with αCD40/IL-4/IL-21, treated after 48 h with the indicated doses of CHK1i and analyzed 24 h later for proliferation as indicated by the division-dependent loss of the proliferation dye, and class switch recombination to IgG1. Representative dot plots and histograms of triplicate samples are shown for each condition. Numbers adjacent to gates depict % of the corresponding population within total viable (Annexin V−/TO-PRO-3−) cells in culture. Numbers in histograms depict how many times cells had divided. b Upper graph depicts the fraction of wild-type viable (Annexin V−/TO-PRO-3−) IgG1+ cells within the culture under the indicated doses of CHK1i or vehicle treatment. Lower graph depicts the fraction of wild-type viable (Annexin V−/TO-PRO-3−) IgG1+ cells within each division cycle under the indicated doses of CHK1i or vehicle treatment. Data are cumulative from three experiments, and shown as mean ± SD. c Bar graph depicts the fraction of Bim−/− viable (Annexin V−/TO-PRO-3−) IgG1+ cells within the culture under graded doses of CHK1i. Data are cumulative from three experiments, and shown as mean ± SD. d Wild-type B cells were loaded with cell proliferation dye, stimulated with αCD40/IL-4/IL-21, treated after 72 h with the indicated doses of CHK1i, and analyzed 24 h later for proliferation as indicated by the division-dependent loss of the proliferation dye and plasmacytic differentiation to CD138+ cells. Bar graph depicts the fraction of CD138+ viable (Annexin V−/TO-PRO-3−) cells within the culture under graded doses of CHK1i. Data are cumulative from three experiments, and shown as mean ± SD. *p < 0.05, **p < 0.01 (one-way ANOVA followed by Tukey post-hoc test) as compared to the respective vehicle-treated control
Altogether, these results suggest that CHK1 facilitates GC B cell survival during CSR independent of its cell cycle arrest function, but relies on preventing mitochondrial apoptosis.
CHK1 expression levels tune the GC reaction
Given our in vitro data, we speculated that GC B cell expansion in vivo might depend on CHK1 activity. This question gained importance in the light of a wealth of clinical trials that use CHK1i to fight cancer [34]. Cell sorting of wild-type bone marrow and splenic B cell subsets followed by qRT-PCR showed a robust increase in CHK1 mRNA and protein in GC B cells as compared to naive follicular B cells, with no differences in CHK1 mRNA detectable between the DZ and LZ compartments (Fig. 6a). Phosphorylated CHK1 is indicative of active ATR-CHK1 signaling in GC B cells (Fig. 6b).
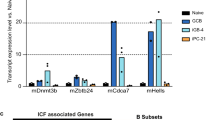
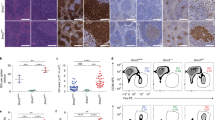
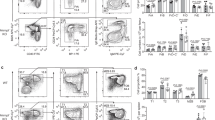
CHK1 expression levels tune the GC reaction. a qRT-PCR analysis of CHK1 mRNA expression in FACS-sorted wild-type developing B cells in the bone marrow (pro B cells, large pre B cells, small pre B cells, immature IgM+ B cells), immature splenic B cells (T (transitional) 1 B cells, T2 B cells, T3 B cells), mature splenic B cells (Fo (follicular) B cells, MZ (marginal zone) B cells), and splenic antigen-experienced B cells (GC B cells, plasmablasts/plasma cells (PC), DZ centroblasts, LZ centrocytes). GC B cells, PC, DZ, and LZ GC B cells were from wild-type mice that had been challenged with sheep RBCs 14 days prior FACS-sorting, all other cell subsets were sorted from unchallenged wild-type mice. Data are means ± SD for three mice per cell population. b Immunoblot analysis for CHK1, pCHK1, and caspase-3 expression in wild-type naïve (Fo) follicular and GC B cells FACS-sorted from spleens 14 days post immunization with sheep RBCs. Cells from six mice were pooled per lane. c Mice of the indicated genotypes were immunized at the age of 8–10 weeks with NP-CGG and analyzed 14 days later. Flow cytometry analysis of splenic single-cell suspensions identified the fractions of B cells (B220+CD19+), GC B cells (B220+CD19+CD138−CD95hiCD38lo/−) and NP+IgG1+ GC B cells within total splenocytes. Data are cumulative from two experiments with 5–6 mice per genotype, with each symbol representing one mouse, and shown as mean ± SEM. d Representative dot plots for the data shown in c. Numbers adjacent to regions indicate the % corresponding population within the respective parent population, and red arrows indicate the gating strategy. e Spleen sections from the indicated genotypes 14 days post immunization with NP-CGG were analyzed for Ki67 (proliferating cells) and for the ability of cells to bind PNA (characteristic of GC B cells). The brown precipitate indicates positive cells. Specimens were counterstained with hematoxylin. One representative mouse out of three per group is shown. Bars: 200 μm. f qRT-PCR analysis for CHK1, BCL6, and AID mRNA expression in FACS-sorted Fo (naïve) and GC splenic B cells of the indicated genotypes 10 days post immunization with NP-CGG. Data are shown as mean ± SD for three mice per group. g Flow cytometry analysis of Peyer’s patch single-cell suspensions in non-challenged 10-week-old mice identified the fractions of B cells (B220+CD19+), GC B cells (B220+CD19+CD95hiCD38lo/−), and class-switched IgG1+ or IgA+ B cells within total Peyer’s patch cells. Data are cumulative from two experiments with 3–6 mice per genotype, with each symbol representing one mouse, and shown as mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.0001 (one-way ANOVA followed by Tukey post-hoc test)
To directly address a potential functional role of CHK1 in the maintenance of acutely induced GCs in vivo we took advantage of a conditional mouse model that allows Chk1 ablation in established GC B cells, during the phase of clonal expansion (Cγ1-cre; [35]). We immunized Cγ1-cre;Chk1+/+ (henceforth referred to as Cγ1-cre), Cγ1-cre;Chk1F/+, and Cγ1-cre;Chk1F/F mice with the T cell-dependent model antigen 4-hydroxy-3-nitrophenylacetyl (NP)-conjugated chicken gammaglobulin (CGG) adsorbed to alum. The fractions of GC B cells and NP-responsive IgG1+ GC B cells were indistinguishable between Cγ1-cre;Chk1F/+ and Cγ1-cre control mice 14 days after immunization (Fig. 6c, d), including the compartmentalization into DZ and LZ (Supplementary Figure 6a). Although one Chk1 allele sufficed to maintain normal-sized induced GCs, homozygous deletion resulted in a near-complete loss of GC B cells. Consistent with the flow cytometry data, structural analysis by immunofluorescence and immunohistochemistry showed that Cγ1-cre;Chk1F/+ GCs were indistinguishable from Cγ1-cre GCs (Fig. 6e, Supplementary Figure 6b), whereas GCs in Cγ1-cre;Chk1F/F mice were rarely detected by PNA and Ki67 staining (Fig. 6e). Of note, Chk1 deletion in Cγ1-cre;Chk1F/+ GC B cells was efficient by day 10 post immunization, and a reduction of CHK1 mRNA levels by half did not lead to an overall deregulation of BCL6 or AID mRNA (Fig. 6f). CHK1 expression, albeit reduced, could be detected in the few remaining GC B cells isolated from Cγ1-cre;Chk1F/F mice, indicating that these cells had escaped Chk1 deletion.
Next, we analyzed GCs in unchallenged mice. Chronic stimulation by a variety of endogenous microbe or food antigens promotes continuous GCs in the gut-associated lymphoid tissues, such as Peyer’s patches. In contrast to our findings in spleens from acutely challenged mice, the fraction of Peyer’s patch GC B cells was reduced by half in Cγ1-cre;Chk1F/+ mice (Fig. 6g). In line with the in vitro observation that CHK1i prevent the appearance of class-switched cells (Fig. 5a, b), Peyer’s patch and splenic IgG1+ and IgA+ cells were reduced by two-third in Cγ1-cre;Chk1F/+ mice (Fig. 6g, Supplementary Figure 6c).
Our data indicate that full CHK1 dosage is critical for the appearance and/or maintenance of class-switched B cells. This becomes particularly apparent when Cγ1-cre;Chk1F/+ B cells are placed in culture, where mitogenic stimulation with either stimulus largely prevented cell proliferation (Supplementary Figure 7a). After 72 h, such cultures contained mainly cells that had not divided and escaped Cre-mediated Chk1 deletion (Supplementary Figure 7b).
Altogether, Chk1 heterozygosity suffices to promote the survival of acutely induced GC B cells, but impedes chronic GCs in Peyer’s patches and generally limits the fraction of class-switched cells.
Full CHK1 expression is required for optimal humoral immunity
As CHK1i are used clinically it is of relevance to understand their effects on antibody maturation and production. During the GC reaction, SHM gradually increases the affinity of serum antibodies and CSR endows these antibodies with new biological functions.
To directly address whether Chk1 heterozygosity negatively affects these processes in vivo, we analyzed NP-specific IgG1 14 days upon immunization with NP-CGG using enzyme-linked immunosorbent assay (ELISA). The fraction of IgG1+ NP-binding GC B cells was comparable between Cγ1-cre and Cγ1-cre;Chk1F/+ mice, but these cells were absent in Cγ1-cre;Chk1F/F mice (Fig. 6c, d). Consequently, NP-specific IgG1 serum titers were absent in Cγ1-cre;Chk1F/F mice (Fig. 7a). Remarkably, however, titers of NP-binding IgG1 were reduced by half in Cγ1-cre;Chk1F/+ mice (Fig. 7a) despite the normal GC B cell percentage (Fig. 6c). In contrast, affinity maturation, as determined by the ratio of high-affinity (NP1.7) to total (NP18) α-NP IgG1, remained unaffected by the loss of one Chk1 allele (Fig. 7b). These results were further supported by enzyme-linked immuno spot assay (ELISPOT) analysis that revealed a general reduction but no alteration in the ratio of high-affinity to total α-NP IgG1 ASCs in the spleens of Cγ1-cre;Chk1F/+ mice (Fig. 7c, d).
Full CHK1 expression is required for optimal humoral immunity. a Serum total (α-NP18) and high-affinity (α-NP1.7) α-NP antibody titers of the IgG1 isotype before (day 0) or 14 days after immunization with NP-CGG. b Antibody affinity maturation of NP-specific IgG1 14 days after immunization. The maturation index is defined as the ratio of α-NP18 and α-NP1.7 antibody titers of the mice shown in a. c Representative picture of ELISPOT analysis of splenic total or high-affinity α-NP IgG1-secreting cells at day 14 after immunization with NP-CGG. Bar: 3 mm. d Quantification of ELISPOT analysis of splenic total or high-affinity α-NP IgG1-secreting cells at day 14 after immunization with NP-CGG. Data are shown as mean ± SEM (a, b, d) and are representative of two independent experiments with 3–6 mice per group. *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.0001 (one-way ANOVA followed by Tukey post-hoc test)
Collectively, our findings indicate that full CHK1 dosage is critical for optimal antibody titers, but probably not required for the selection of high-affinity B cells in acute GCs. Our observations suggest a role for CHK1-mediated cell survival after affinity maturation, during or after the initiation of terminal differentiation of GC B cells into memory B cells and ASCs.
Discussion
The motivation for the present study was sparked by our discovery that Chk1 ablation is toxic during B cell development [36]. However, reduction of Chk1 dosage by half satisfies progenitor B cell survival.
Here, we explored the role of CHK1 in GC B cell homeostasis using a conditional mouse model where CHK1 is specifically ablated in established GC B cells [35]. Bi-allelic elimination of Chk1 is incompatible with GC maintenance, galvanizing an essential role for CHK1 in humoral immunity. Although ablation of one Chk1 allele upon acute challenge resulted in phenotypically normal GCs, the fraction of chronic GCs in Peyer’s patches of unchallenged Cγ1-cre;Chk1F/+ mice was reduced by 50%. Whereas acutely induced GCs usually collapse within 1 month, gut microbiota-induced GCs remain active for longer [37]. Chk1 heterozygosity might sensitize GC B cells with a prolonged GC dwell time to apoptosis upon cumulative mutagenesis by AID and/or replication-associated DNA damage [38, 39]. We also noticed a general reduction of IgG1+ and IgA+ cells in unchallenged mice, the majority likely representing memory B cells. Memory B cells can re-enter GCs to further diversify their BCR [40, 41], and it seems plausible that repeated passaging through GCs reduces the number of Chk1+/− memory B cells due to cumulative DNA damage.
Results in DT40 B cells in vitro suggested that CHK1 activity limits affinity maturation [42]. In our ELISA and ELISPOT experiments, affinity maturation appears not detectably affected in Cγ1-cre;Chk1F/+ mice upon challenge. However, more extensive SHM, e.g. in chronic GCs, may require full CHK1 dosage.
Although Chk1 heterozygosity did not limit SHM and the maintenance of acutely induced GCs, it is yet remarkable that the amplitude of antigen-specific humoral immunity, i.e. the quantity of α-NP IgG1, was reduced by 50%. This phenotype is a consequence of reduced α-NP IgG1-secreting ASC numbers. However, plasmacytic differentiation into ASCs per se appeared unperturbed. In contrast, already low-dose CHK1i specifically diminished the appearance of IgG1+ cells in vitro, independent of CHK1 cell cycle arrest functions. Although AID activity is restricted to the G1 cell cycle phase [43], DSB resolution during CSR requires S-phase entry [32, 43] and hence is likely to be controlled by ATR-CHK1. Our results suggest that CHK1 promotes cell survival in the presence of CSR-initiated DSBs that are carried into the S-phase.
Given this pronounced in vitro phenotype, it appears surprising that NP-CGG induced Chk1+/− GCs contain normal fractions of IgG1+ cells, and that NP-specific IgG1 levels are only reduced by two-fold. In contrast to in vitro-proliferating B cells that are continuously exposed to the same mitogen, BCR-signaling and T cell help are mutually exclusive in GCs. It is therefore conceivable that CHK1 dependency oscillates in vivo, and that short-term dampening of CHK1 safeguards S–G2–M DZ GC B cells carrying AID-initiated lesions [18, 19]. This might come at the cost of replication-associated bystander DNA lesions.
BCR-signaling during positive selection in the LZ is restricted to the G1 cell cycle phase. However, a small fraction of G2–M DZ B cells undergoes active BCR-signaling [44], and it is tempting to speculate that BCR-signaling in combination with physiological CHK1-repression represents a checkpoint that allows the elimination of genotoxically damaged G2–M DZ cells before entering the LZ.
In vitro, BCR-ligation sensitized cells to immediate apoptosis upon low-dose CHK1i whereas T cell-mimicking conditions favored S–G2 cell cycle arrest. Both types of stimulation led to pronounced replication stress and similar CHK1 levels. Yet, CHK1 activity was more strongly induced upon joint T cell help mimicking stimuli. Differential activities of cytoplasmic and nuclear CHK1, the former needed for cell cycle arrest and the latter for survival, have been reported [45] and it will be interesting to see whether cellular distribution of CHK1 plays a role for GC B cell survival.
The differential apoptotic response to CHK1i correlated with a mitogen-specific rewiring of the BCL2-family network, with BCR-ligation priming B cells to BIM-induced apoptosis in the presence of CHK1i. Curiously, BCR-stimulated Bim−/− B cells failed to arrest in S–G2 upon CHK1 inhibition despite being protected from apoptosis, further substantiating a direct involvement of BCR-signaling in preventing cell cycle arrest in this context. BCR hyperactivation drives BIM-dependent apoptosis [23, 46, 47], and BIM is essential for developmentally programmed apoptosis of GC B cells, memory B cells, and ASCs [48]. Hence, one may expect that Bim-deficiency rescues humoral immunity in Cγ1-cre;Chk1F/F mice. However, a functional rescue appears unlikely given that BCL2-overexpressing CHK1-deficient B cell progenitors fail to expand but mount a cell cycle arrest response when cell death is blocked [36]. Whether BIM-loss suffices to rescue the phenotypes observed in Cγ1-cre;Chk1F/+ mice remains to be addressed experimentally.
As a side note, cultivation of un-stimulated B cells caused the up-regulation of CHK1, a phenomenon that might explain why naïve B cells undergo cell death upon CHK1i in vitro despite being quiescent [36]. This suggests that CHK1 may exert a pro-survival function that is unrelated to its role in S-phase progression.
Owing to its essential nature, human tumors rarely lose but often overexpress CHK1 [22]. Therefore, pharmacological inhibition of CHK1 is a promising strategy to treat tumors with high endogenous replication stress, and selective inhibitors are undergoing clinical trials [34]. Although systemic CHK1 inhibition might be tolerated by quiescent cells [49] and Chk1 heterozygosity in mice leaves most cell types unperturbed [50], our data predict that the synergistic cytotoxicity of CHK1i and BCR-signaling will come at a cost for immunity upon infection or vaccination, defining a caveat of therapies targeting this kinase. However, the therapeutic benefit of CHK1i may outweigh the detrimental effects on humoral immunity.
B cell lymphomas are particularly sensitive to CHK1i, presumably due to substantial endogenous replication stress [51,52,53]. Our observation that CHK1i synergizes with active BCR-signaling in killing B cells at low nanomolar concentrations reveals a therapeutic option for leukemias or lymphomas whose survival relies on BCR-signaling [54], such as CLL or ABC-type DLBCL. Exploiting the lethal relationship between BCR-signaling and CHK1i may be of particular value for patients relapsing from BCR-signaling inhibitors that are characterized by limited salvage options. Finally, understanding the relation between active BCR-signaling and CHK1i will be essential for the rational manipulation of CHK1 in malignancy.
Materials and methods
Mice
The Cγ1-cre [35], Chk1F [55], and Bim- [56] alleles have been described and were generated on or backcrossed to the C57BL/6 genetic background for more than 10 generations. For all experiments, male and female littermate mice were used indiscriminately. For T cell-dependent immunization, 8- to 10-week-old mice were injected intraperitoneally with 1 × 109 sheep RBCs (Labor Dr. Merck, Ochsenhausen, Germany, E-400) in phosphate-buffered saline (PBS) or with 100 μg alum-precipitated NP15-CGG (Biosearch Technologies, Hoddeston, UK, N-5055B-5). Mice were analyzed at day 10 or 14 post immunization. Animal procedures were approved by the official review boards (BMWF: 66-011/0106-WF/3b/2015).
DNA preparation and genotyping
Mouse ear snips were digested in 100 μl hot shot reagent (25 mM NaOH, 0.2 mM disodium EDTA (pH 12.0)) for 30 min at 95 °C and subsequently placed on ice. One hundred microliters of neutralization reagent (40 mM Tris-HCl (pH 5.0)) were added and the DNA was stored at 4 °C. For genotyping the Taq Plus Master Mix (Vazyme Biotech, Nanjing, China, P212-03) or the PCR Taq Plus Master Mix (Vazyme biotech P211) were used as per the manufacturer’s instructions.
Preparation of alum-precipitated NP-CGG
To determine the murine humoral immune response to a model antigen, the hapten-carrier conjugate NP-CGG was used. For every immunization, a 10% Alum solution was freshly prepared (KAl(SO4)2 (Sigma, Saint Louis, MO, USA, 31242) in PBS). About 1 mg/ml NP15-CGG in PBS (Biosearch Technologies N-5055B-5) was mixed with the same volume 10% Alum solution, and the pH was adjusted to 6.5–7.0 by adding 10 M NaOH (pH indicator; Macherey-Nagel, Düren, Germany, 92118). The precipitate was pelleted at 5000 r.p.m. for 15 s, washed three times with PBS, and finally resuspended in PBS to reach the initial volume of the NP-CGG/Alum mixture. Mice were immunized with 100 µg NP-CGG/mouse (200 µl/mouse).
Preparation of single-cell suspensions and cell counting
Staining buffer (PBS with 3% fetal calf serum (Thermo Fisher Scientific, Waltham, MA, USA, 10270-106), pH 7.2) was used for all cell preparation steps. Single-cell suspensions were prepared by forcing murine spleen and Peyer’s patches through 70 µM mesh filters (Corning, Cambridge, MA, USA, 352350), or flushing both femurs and tibiae using a 23 G needle. Erythrocyte depletion was achieved by incubating cells for 3 min in 1 ml lysis buffer (155 mM NH4Cl, 10 mM KHCO3, 0.1 mM EDTA; pH 7.5) on ice. Cells were washed and filtered through a 50 µM cup filcon (BD Biosciences, San Diego, CA, USA, 340632). Generally, cell numbers were determined using a hemocytometer (Neubauer) and trypan blue exclusion.
Culture of primary B cells
For in vitro B cell cultivation, B cells were enriched from splenic single-cell suspensions by immunomagnetic enrichment using MagniSort Streptavidin Negative Selection Beads (Thermo Fisher Scientific MSNB-6002-74) and biotinylated antibodies against Ter119 (BioLegend, San Diego, CA, USA, 116204, 1:100), NK1.1 (BioLegend 108704, 1:100), Gr-1 (BioLegend 108404, 1:100), CD11b (BioLegend 101204, 1:100), and TCRβ (BioLegend 109204, 1:50), as per the manufacturer’s instructions.
Primary B cells were cultured in B cell medium at a density of 1 × 106/ml in Dulbecco's modified Eagle's medium (DMEM) (Sigma, Saint Louis, USA, WHMISDZB) supplemented with 10% (v/v) FBS (Gibco, Grand Island, NY, USA, 10270-106), 2 mM l-glutamine (Sigma G7513), 10 mM Hepes (LONZA, Basel, Switzerland, BE17-737E), 1 mM sodium pyruvate (Gibco 13360-039), 1× non-essential amino acids (Gibco 11140-035), 100 U/ml penicillin and 100 μg/ml streptomycin (Sigma 0781), and 50 µM β-mercaptoethanol (Sigma M3148). Cells were cultured at 37 °C in a humidified atmosphere containing 5% CO2.
Mitogenic stimuli were used as follows: 1 μg/ml α-CD40 (Thermo Fisher Scientific 16-0402-85), 10 ng/ml IL-4 (Peprotech, Rocky Hill, NJ, USA, 214-14), 10 ng/ml IL-21 (Peprotech 210-21), 13 μg/ml α-IgM F(ab′)2 fragments (Jackson ImmunoResearch, West Grove, PA, USA, 115-006-020).
PF-477736 (Selleckchem, Munich, Germany S2904) and CHIR-124 (Selleckchem S2683), both dissolved in dimethyl sulfoxide (DMSO) (Sigma D5879), were added to cultures at the indicated concentrations after 48 h of mitogenic stimulation, and cells were analyzed 24 h later. Vehicle controls shown refer to the highest DMSO concentration used in each inhibitor experiment.
To determine cell proliferation in vitro, B cells were labeled with 10 μM Cell Proliferation Dye eFluor450 (Thermo Fisher Scientific 65-0842-90) as per the manufacturer’s instructions and placed in culture in the absence or presence of mitogens.
Flow cytometry
For flow cytometry analysis of ex vivo isolated splenic and Peyer’s patch cells, 2 × 106 cells in single-cell suspension were used per staining. For all staining procedures, 96-well plates were used until otherwise stated, and cells were commonly washed with 200 μl of staining buffer by centrifugation at 2 000 r.p.m. for 2 min. During all preparation and staining steps, cells were kept on ice or at 4 °C.
To block non-specific antibody-binding, cells were pre-incubated with 1 μg/ml of αCD16/32 Fc-Block (BioLegend 101310) in 30 μl staining buffer for 10 min, washed, and stained for 20 min with 30 μl antibody cocktails. The following fluorochrome-labeled antibodies were used: αB220-PerCP/Cy5.5 (BioLegend 103236, 1:300), αB220-BV510 (BioLegend 103247, 1:300), αCD19-BV605 (BioLegend 115540, 1:300), αCD19-BV421 (BioLegend 115538, 1:300), αAA4.1-PE/Cy7 (BioLegend 136507, 1:400), αAA4.1-APC (BioLegend, 136510 1:400), αIgM F(ab′)2-FITC (Jackson ImmunoResearch 115-096-072, 1:300), αCD25-PE (BioLegend 102007, 1:400), αcKit-APC (BioLegend 135108, 1:300), αCD1d-PE (Thermo Fisher Scientifics 12-0011-82, 1:500), αCD23-PE/Cy7 (BioLegend 101614, 1:400), αCD38-FITC (BioLegend 102705, 1:300), αCD38-APC (BioLegend 102712, 1:400), αCD38-eFluor450 (Thermo Fisher Scientific 48-0381-80, 1:300), αCD95-PE (BD Biosciences 554258, 1:300), αCD95-PE/Cy7 (BD Biosciences 557653, 1:800), αCXCR4-APC (BioLegend 146508, 1:100), αCD86-PE/Cy7 (BioLegend 105014, 1:100), αIgG1-FITC (BD Biosciences 553443, 1:200), αIgG1-PE (BD Biosciences 550083, 1:200), αIgA-PE (Thermo Fisher Scientific 12-4204-82, 1:400), CD138-PE (BioLegend 142504, 1:300), and CD138-BV510 (BioLegend 142521, 1:200). Total NP-binding B cells were detected using NP24-PE (Biosearch Technologies N-5070-1, 1:500). Cells were resuspended in 200 μl of staining buffer prior acquisition. Alternatively, apoptosis was assessed by resuspending cells in 200 μl 1× Annexin V-binding buffer (BioLegend 422201) containing Annexin V-FITC (BioLegend 640945, 1:800) and 10 nM TO-PRO-3 (Invitrogen, Carlsbad, CA, USA, T3605) 10 min prior flow cytometry analysis.
For intracellular γ-H2AX or phospho-ATR staining, 1 × 106 cells were cell-surface stained as described above, fixed for 20 min in fixation buffer (BD Biosciences 554655) and permeabilized with 1× permeabilization buffer (BD Biosciences 554723) as per the manufacturer’s instructions. Cells were incubated with γ-H2AX (Cell Signaling, Beverley, MA, USA, 2577, 1:100) or pATR Ser-428 (Cell Signaling 2853, 1:100) antibody in 50 μl 1× permeabilization buffer for 30–60 min. Cells were washed twice with 1× permeabilization buffer and incubated for 30 min with Alexa Fluor 488 goat α-rabbit (Invitrogen A11034, 1:1000) in 50 μl 1× permeabilization buffer. Cells were washed twice with 1× permeabilization buffer, resuspended in 100 μl PBS containing 250 μg/ml RNase A (Sigma R5500), and incubated for 30 min at 37 °C. Finally, 50 μl of 3 μM TO-PRO-3 (Invitrogen T3605) in PBS was added, and cells were acquired on LOW mode.
For intracellular phospho-Histone 3 staining and DNA content analysis, 1 × 106 cells were fixed in 1 ml ice-cold 70% ethanol in FACS-tubes for a minimum of 16 h at −20 °C. Prior antibody staining, cells were washed twice adding 3 ml PBS (2000 r.p.m., 5 min) to remove ethanol, transferred in 96-well plates, and subsequently permeabilized with 200 μl PBS containing 0.25% Triton X-100 (Sigma T8787) for 20 min. Cells were washed with PBS and incubated for 30–60 min with a phospho-Histone H3 Ser-10 (Cell Signaling 9701, 1:200) antibody in 50 μl staining buffer. Cells were washed twice with PBS/0.25% Triton X-100 and incubated for another 30 min with Alexa Fluor 488 goat α-rabbit (Invitrogen A11034, 1:1000) in 50 μl staining buffer. Cells were washed twice with PBS, resuspended in 100 μl PBS containing 250 μg/ml RNase A, and incubated for 30 min at 37 °C. Finally, 50 μl 10 μg/ml DAPI (Sigma D9542) in PBS was added, and cells were acquired on LOW mode.
Data were acquired on an LSRII cytometer (BD Biosciences) and analyzed by FlowJo software (Tree Star, Ashland, OR, USA). Non-singlet events were typically excluded from analyses based on characteristics of FSC-H/FSC-W, SSC-H/SSC-W, in case of DNA staining in addition with DAPI 405-A/405-H or TO-PRO-3 APC-A/APC-H.
Cell sorting
For cell sorting, bone marrow or splenic single-cell suspensions were pre-incubated with 1 μg/ml of αCD16/32 Fc-Block in 500 μl staining buffer for 10 min, washed, and stained for 20 min with fluorochrome-labeled monoclonal antibodies in a volume of 500 μl staining buffer, as described above. The sorted cell subsets were defined as follows: bone marrow: pro B cells (B220loCD19+AA4.1+IgM−CD25−ckit+), large pre B cells (B220loCD19+AA4.1+IgM−CD25+ckit−FSChi), small pre B cells (B220loCD19+AA4.1+IgM−CD25+ckit−FSClo), and immature IgM+ B cells (B220loCD19+AA4.1+IgM+).
Spleen: T1 B cells (CD19+B220+AA4.1+CD23−IgMhi), T2 B cells (CD19+B220+AA4.1+CD23+IgMhi), T3 B cells (B220+CD19+AA4.1+CD23+IgMint/lo), Fo B cells (CD19+B220+AA4.1−CD1d+IgM+), MZ B cells (CD19+B220+AA4.1−CD1dhiIgMhi), plasma cells/plasmablasts (B220−/loCD138hi), GC B cells (CD19+B220+CD138−FashiCD38lo/−), DZ GC B cells (CD19+B220+CD138−FashiCD38lo/−CXCR4hiCD86lo), and LZ GC B cells (CD19+B220+CD138−FashiCD38lo/−CXCR4loCD86hi).
Cell sorting was carried out on a FACS Aria III (BD Biosciences). Non-singlet events were typically excluded from analyses based on characteristics of FSC-H/FSC-W and SSC-H/SSC-W.
Quantitative real-time-PCR
RNA from snap-frozen cell pellets of in vitro-cultivated B cells was isolated using Trizol reagent (Thermo Fisher Scientific 15596026) as per the manufacturer’s instructions. DNA was removed using the RQ1 RNAse-free DNase (Promega, Madison, WI, USA, M610A), and RNA was retrieved using GlycoBlue Coprecipitant (Thermo Fisher Scientific AM9515) as per the manufacturer’s instructions. RNA from snap-frozen FACS-sorted cell pellets was isolated using the RNeasy Mini Kit (QIAGEN, Chatsworth, CA, USA, 74104), including an optional DNA digestion step using the RNase-free DNase Set (QIAGEN 79254). First-strand cDNA was generated from 100 ng of total RNA with a 260/280 ratio of >1.8 using the iScript cDNA Synthesis Kit (Bio-Rad, Hercules, CA, USA, 170-8891) as per the manufacturer’s instructions. cDNA was amplified using the AceQ qPCR SYBR Green Master Mix (Vazyme Q111-02) as per the manufacturer’s instructions on a StepOnePlus Real-time PCR system (Applied Biosystems, Foster City, CA, USA). Melt curve analysis was performed for every run to verify the presence of one PCR product. The expression of individual mRNAs was normalized and expressed as % relative to the endogenous reference gene Hprt with the following formula: fold induction = 2(−ΔCt), where ΔCt = Ct(target)−Ct(HPRT). The following primers were used: HPRT F: GTCATGCCGACCCGCAGTC, HPRT R: GTCCTTCCATAATAGTCCATGAGGAATAAAC; CHK1 F: CTCCATCAGCAA GGATCACC, CHK1 R: ACGGTTTCTTCACTGGAACC; AID F: GGACTTCGGCCACCTTC, AID R: CATCTCAGAAACTCAGCCACG; BCL6 F: GCCCACGTTCCCGGAGGAGA, BCL6 R: CGTCTGCAGCGTGTGCCTCT.
Immunoblotting
Snap-frozen cell pellets from FACS-sorted or in vitro-cultivated B cells were lysed for 30 min in ice-cold whole-cell extraction buffer (25 mM Hepes (pH 7.9), 300 mM NaCl, 1.5 mM MgCl2, 0.2 mM EDTA (pH 8.0), 0.5% Triton X-100) completed with protease inhibitors (Roche, Basel, Switzerland, 11873580001) and 2 mM dithiothreitol before use. Protein concentration was quantified using Bradford Assay (Bio-Rad, Munich, Germany, 500-0006). Protein extracts were denatured by the addition of sample buffer (50 mM Tris-HCl, 2% SDS, 10% glycerol, 0.1% bromphenolblue, 100 mM β-mercaptoethanol), boiled for 5 min, resolved by SDS-polyacrylamide gel electrophoresis (Tris-glycine 12% gel, 50 µg/lane) and transferred to a nitrocellulose membrane (GE Healthcare, Chicago, IL, USA, 10600004) for 75 min at 100 V (Bio-Rad, Mini Trans-Blot system). Membranes were blocked with 5% skim milk in PBS-T (PBS with 0.1% Tween) and probed with primary (overnight) and secondary (2 h) antibodies. The following primary antibodies were diluted in 5% skim milk in PBS-T: α-HSP90 (Santa Cruz, Dallas, TX, USA, 13119, 1:200), α-CHK1 (Cell Signaling 2360, 1:1000), α-Caspase-3 (Cell Signaling 9665, 1:1000), α-BIM (ENZO ALX-804-527, 1:500), α-MCL1 (Rockland 600-401-394 S, 1:500), α-BCL2 (BioLegend 633502, 1:500), α-BCLXL (Cell Signaling 2764, 1:500), α-BCL2a1 (clone 6D6 [57], 1:500). α-pCHK1 Ser-345 (Cell Signaling 2345, 1:500) was diluted in 5% BSA in PBS-T. The following horseradish peroxidase (HRP)-conjugated secondary antibodies were probed in 5% skim milk in PBS-T: α-rabbit Ig/HRP (Dako P0448, 1:10,000), α-mouse Ig/HRP (Dako P0161, 1:10,000), and α-rat IgG/HRP (Cell Signaling 7077, 1:10,000). Secondary antibodies were detected using ECL (Advansta, K-12049-D50), and the chemiluminescent signal was visualized using X-ray films (GE Healthcare 28906837; AGFA CURIX HT1.000G PLUS).
Enzyme-linked immunosorbent assay (ELISA)
For NP-specific serum IgG1 titers 14 days after immunization with NP-CGG, 50 μg/μl NP conjugated to BSA (Biosearch Technologies) in the ratios 18:1 and 1.7:1 was diluted in PBS and coated overnight at 4 °C onto 96-well enzyme-linked immunosorbent assay plates (Sigma CLS3590). Plates were washed three times with wash buffer (PBS containing 0.05% TWEEN-20), blocked with 100 μl/well 1% BSA in PBS for 4 h at room temperature and washed three more times with wash buffer. The optimal serum dilution was tested upfront to ensure absorbance readings in the linear range. Sera diluted 1:102,400 in PBS were incubated at 4 °C overnight in 100 μl/well. Plates were washed three times with wash buffer and incubated with 100 μl/well of HRP-conjugated anti-mouse IgG1 (Southern Biotech, Birmingham, AL, USA, 1070-05, 1:5000 in 1% BSA in PBS) for 4 h at room temperature. For detection of plate-bound IgG1-HRP, 100 μl of ABTS substrate solution ((200 μl ABTS (Stock: 15 mg/ml in a.d.), 10 ml citrate-phosphate buffer (574 mg citric acid monohydrate in 50 ml a.d.), 10 μl H2O2) was added to each well and incubated for 20 min. Absorbance of the colored product was measured at 405 nm using a microplate reader (Tecan Sunrise, Männedorf, Switzerland).
Enzyme-linked immuno spot assay (ELISPOT)
The frequency of NP-specific IgG1 ASCs within total splenocytes 14 days after immunization with NP-CGG was determined as described [58]. Briefly, 1 × 106 splenocytes/well were incubated in B cell medium overnight at 37 °C on 96-well MultiScreen-HA filter plates (Merck Millipore, Billerica, MA, USA, MAIPS4510) pre-coated with 20 μg/ml NP18-BSA. Spots were visualized with an HRP-conjugated goat α-mouse IgG1 (Southern Biotech 1070-05, 1:1000), and the color was developed by the addition of 2.5 mg/ml 3-amino-9-ethyl carbazole (Sigma A5754) in 0.05 M sodium acetate (pH 5.0) and H2O2 (1:1000). Spots were counted using a CTL ImmunoSpot analyzer (CTL Europe, Bonn, Germany).
Immunohistochemistry
IHC was performed on 7 μm sections of murine spleens that were formalin-fixed (4% paraformaldehyde in PBS; 24 h at room temperature) and embedded in paraffin wax according to standard procedures. Sections were deparaffinized and rehydrated in the following order: 3× xylol for 5, 7, and 10 min; 2× 100% ethanol for 10 min; 2× 96% ethanol for 10 min; 1× 70% ethanol for 10 min; 1× 50% ethanol for 10 min; 2× deionized H2O for 5 min. For Ki67 staining, heated citrate antigen retrieval (10 mM Tri-sodium citrate dehydrate in deionized H2O, pH 6.0) was performed 3 × 5 min in the microwave (450 W), and the slides were cooled down for 30 min. For PNA staining, antigen retrieval was performed in TE buffer (10 mM Tris-Base, 1 mM EDTA deionized H2O, pH 8.0) 3 × 5 min in the microwave (450 W), and the slides were cooled down for 30 min. Slides were placed in H2O 3× for 5 min. Endogenous peroxidases were blocked (3% H2O2 in deionized H2O), the slides washed 2 × 5 min in H2O, placed for 5 min in TBST (Tris-buffered saline pH 7.6, 0.1% TWEEN-20), and blocked for unspecific antigen binding with 100 μl 5% goat serum in TBST for 1 h. Sections were stained as follows:
Ki67: Sections were incubated with 50 μl αKi-67 in TBST (Cell signaling 12202, 1:400) overnight at 4 °C in a humidified chamber. Slides were washed 3 × 10 min in TBST, and sections were incubated with 50 μl biotinylated goat αrabbit (Jackson ImmunoResearch 111-065-144, 1:100) in TBST for 2 h at room temperature in a humidified chamber.
PNA: Sections were incubated with 50 μl biotinylated PNA (Sigma L6135, 1:250) in TBST overnight at 4 °C in a humidified chamber.
Slides were washed 3 × 10 min in TBST, and biotinylated antibodies were labeled with the ABC Peroxidase Standard Staining Kit (Vector Laboratories PK-6100) according to the manufacturer’s instructions. HRP-labeled avidin complexes were visualized with a Peroxidase Substrate Kit (Vector Laboratories, Burlingame, CA, USA, SK-4105) as per the manufacturer’s instructions. Sections were counterstained for <10 s with hematoxylin (Merck Millipore, 109249). Sections were imaged on a Zeiss Axioplan 2 imaging with the ×10 objective (Zeiss) and an Axiocam 305 color camera (Zeiss, Oberkochen, Germany), and pictures were processed with the Zeiss ZEN blue 2.5 lite software (Zeiss).
Immunofluorescence
Four μm cryosections of spleens from NP-CGG immunized mice were fixed with 1% paraformaldehyde in PBS for 10 min at room temperature. Sections were blocked with 2% BSA in PBS for 30 min at room temperature followed by endogenous biotin and avidin blocking according to the manufacturer’s instructions (Vector Labs, SP-2002). Samples were stained with mouse monoclonal antibodies αCD3-APC (Thermo Fisher Scientific 17-0031-82, 1:50) and αB220-PE (Thermo Fisher Scientific 12-0425-82, 1:40), and a biotin-labeled PNA (Vector Labs B-1085, 1:50) in 1% BSA in PBS for 45 min at room temperature. Subsequently, spleens were incubated with Streptavidin-A488 (Invitrogen S32354, 1:1000) for 45 min at room temperature. After washing, slides were mounted with Diamond Antifade medium (Invitrogen P36965) and images were acquired on a Leica DMi8 inverted microscope (Leica, Wetzlar, Germany) with ×10 and ×20 objectives, and processed using Fiji software [59].
Quantification and statistical analysis
Results are shown as mean and standard deviation (SD), or as standard error of the mean (SEM) as stated in the respective figure legends. Graphs were plotted and statistical analysis was performed with GraphPad Prism 7 software (San Diego, CA, USA) using one-way ANOVA followed by Tukey’s post-hoc test when comparing multiple groups. Dose–response curves depict log(inhibitor) vs. response (variable slope). Sample sizes were empirically determined and the exact number of biological repetitions (n) is stated in the respective figure legends. Every experiment was performed at least twice. No specific randomization or blinding protocol was used for experimental grouping. Differences between groups were considered statistically significant when P < 0.05. In figures, asterisks stand for: *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001.
References
Mesin L, Ersching J, Victora GD. Germinal center B cell dynamics. Immunity. 2016;45:471–82.
De Silva NS, Klein U. Dynamics of B cells in germinal centres. Nat Rev Immunol. 2015;15:137–48.
Shlomchik MJ, Weisel F. Germinal center selection and the development of memory B and plasma cells. Immunol Rev. 2012;247:52–63.
Victora GD, Schwickert TA, Fooksman DR, Kamphorst AO, Meyer-Hermann M, Dustin ML, et al. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell. 2010;143:592–605.
Gitlin AD, Shulman Z, Nussenzweig MC. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature. 2014;509:637–40.
Allen CDC, Okada T, Tang HL, Cyster JG. Imaging of germinal center selection events during affinity maturation. Science. 2007;315:528–31.
Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 2000;102:553–63.
Di Noia JM, Neuberger MS. Molecular mechanisms of antibody somatic hypermutation. Annu Rev Biochem. 2007;76:1–22.
Fazilleau N, Mark L, McHeyzer-Williams LJ, McHeyzer-Williams MG. Follicular helper T cells: lineage and location. Immunity. 2009;30:324–35.
Gitlin AD, Mayer CT, Oliveira TY, Shulman Z, Jones MJK, Koren A, et al. Humoral immunity. T cell help controls the speed of the cell cycle in germinal center B cells. Science. 2015;349:643–46.
Wortis HH, Teutsch M, Higer M, Zheng J, Parker DC. B-cell activation by crosslinking of surface IgM or ligation of CD40 involves alternative signal pathways and results in different B-cell phenotypes. Proc Natl Acad Sci USA. 1995;92:3348–52.
Otipoby KL, Waisman A, Derudder E, Srinivasan L, Franklin A, Rajewsky K. The B-cell antigen receptor integrates adaptive and innate immune signals. Proc Natl Acad Sci USA. 2015;112:12145–50.
Hauser AE, Junt T, Mempel TR, Sneddon MW, Kleinstein SH, Henrickson SE, et al. Definition of germinal-center B cell migration in vivo reveals predominant intrazonal circulation patterns. Immunity. 2007;26:655–67.
Kotsantis P, Petermann E, Boulton SJ. Mechanisms of oncogene-induced replication stress: jigsaw falling into place. Cancer Discov. 2018;8:537–55.
Daniel JA, Nussenzweig A. The AID-induced DNA damage response in chromatin. Mol Cell. 2013;50:309–21.
Robbiani DF, Nussenzweig MC. Chromosome translocation, B cell lymphoma, and activation-induced cytidine deaminase. Annu Rev Pathol. 2013;8:79–103.
Saldivar JC, Cortez D, Cimprich KA. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat Rev Mol Cell Biol. 2017;18:622–36.
Ranuncolo SM, Polo JM, Dierov J, Singer M, Kuo T, Greally J, et al. Bcl-6 mediates the germinal center B cell phenotype and lymphomagenesis through transcriptional repression of the DNA-damage sensor ATR. Nat Immunol. 2007;8:705–14.
Ranuncolo SM, Polo JM, Melnick A. BCL6 represses CHEK1 and suppresses DNA damage pathways in normal and malignant B-cells. Blood Cells Mol Dis. 2008;41:95–9.
Phan RT, Dalla-Favera R. The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. Nature. 2004;432:635–39.
Phan RT, Saito M, Basso K, Niu H, Dalla-Favera R. BCL6 interacts with the transcription factor Miz-1 to suppress the cyclin-dependent kinase inhibitor p21 and cell cycle arrest in germinal center B cells. Nat Immunol. 2005;6:1054–60.
Lecona E, Fernandez-Capetillo O. Targeting ATR in cancer. Nat Rev Cancer. 2018;18:586–95.
Enders A, Bouillet P, Puthalakath H, Xu Y, Tarlinton DM, Strasser A. Loss of the pro-apoptotic BH3-only Bcl-2 family member Bim inhibits BCR stimulation-induced apoptosis and deletion of autoreactive B cells. J Exp Med. 2003;198:1119–26.
Labi V, Woess C, Tuzlak S, Erlacher M, Bouillet P, Strasser A, et al. Deregulated cell death and lymphocyte homeostasis cause premature lethality in mice lacking the BH3-only proteins Bim and Bmf. Blood. 2014;123:2652–62.
Woess C, Tuzlak S, Labi V, Drach M, Bertele D, Schneider P, et al. Combined loss of the BH3-only proteins Bim and Bmf restores B-cell development and function in TACI-Ig transgenic mice. Cell Death Differ. 2015;22:1477–88.
Delbridge ARD, Pang SHM, Vandenberg CJ, Grabow S, Aubrey BJ, Tai L, et al. RAG-induced DNA lesions activate proapoptotic BIM to suppress lymphomagenesis in p53-deficient mice. J Exp Med. 2016;213:2039–48.
Erlacher M, Michalak EM, Kelly PN, Labi V, Niederegger H, Coultas L, et al. BH3-only proteins Puma and Bim are rate-limiting for gamma-radiation- and glucocorticoid-induced apoptosis of lymphoid cells in vivo. Blood. 2005;106:4131–38.
Erlacher M, Labi V, Manzl C, Böck G, Tzankov A, Häcker G, et al. Puma cooperates with Bim, the rate-limiting BH3-only protein in cell death during lymphocyte development, in apoptosis induction. J Exp Med. 2006;203:2939–51.
Villunger A, Labi V, Bouillet P, Adams J, Strasser A. Can the analysis of BH3-only protein knockout mice clarify the issue of ‘direct versus indirect’ activation of Bax and Bak? Cell Death Differ. 2011;18:1545–46.
Gutierrez-Martinez P, Hogdal L, Nagai M, Kruta M, Singh R, Sarosiek K, et al. Diminished apoptotic priming and ATM signalling confer a survival advantage onto aged haematopoietic stem cells in response to DNA damage. Nat Cell Biol. 2018;20:413–21.
Hodgkin PD, Lee JH, Lyons AB. B cell differentiation and isotype switching is related to division cycle number. J Exp Med. 1996;184:277–81.
Wiedemann E-M, Peycheva M, Pavri R. DNA replication origins in immunoglobulin switch regions regulate class switch recombination in an R-loop-dependent manner. Cell Rep. 2016;17:2927–42.
Duffy KR, Wellard CJ, Markham JF, Zhou JHS, Holmberg R, Hawkins ED, et al. Activation-induced B cell fates are selected by intracellular stochastic competition. Science. 2012;335:338–41.
Zhang Y, Hunter T. Roles of Chk1 in cell biology and cancer therapy. Int J Cancer. 2014;134:1013–23.
Casola S, Cattoretti G, Uyttersprot N, Koralov SB, Seagal J, Segal J, et al. Tracking germinal center B cells expressing germ-line immunoglobulin gamma1 transcripts by conditional gene targeting. Proc Natl Acad Sci USA. 2006;103:7396–401.
Schuler F, Weiss JG, Lindner SE, Lohmüller M, Herzog S, Spiegl SF, et al. Checkpoint kinase 1 is essential for normal B cell development and lymphomagenesis. Nat Commun. 2017;8:1697.
Sutherland DB, Suzuki K, Fagarasan S. Fostering of advanced mutualism with gut microbiota by Immunoglobulin A. Immunol Rev. 2016;270:20–31.
Klein F, Diskin R, Scheid JF, Gaebler C, Mouquet H, Georgiev IS, et al. Somatic mutations of the immunoglobulin framework are generally required for broad and potent HIV-1 neutralization. Cell. 2013;153:126–38.
Pappas L, Foglierini M, Piccoli L, Kallewaard NL, Turrini F, Silacci C, et al. Rapid development of broadly influenza neutralizing antibodies through redundant mutations. Nature. 2014;516:418–22.
Dogan I, Bertocci B, Vilmont V, Delbos F, Mégret J, Storck S, et al. Multiple layers of B cell memory with different effector functions. Nat Immunol. 2009;10:1292–99.
McHeyzer-Williams LJ, Milpied PJ, Okitsu SL, McHeyzer-Williams MG. Class-switched memory B cells remodel BCRs within secondary germinal centers. Nat Immunol. 2015;16:296–305.
Frankenberger S, Davari K, Fischer-Burkart S, Böttcher K, Tomi N-S, Zimber-Strobl U, et al. Checkpoint kinase 1 negatively regulates somatic hypermutation. Nucleic Acids Res. 2014;42:3666–74.
Wang Q, Kieffer-Kwon K-R, Oliveira TY, Mayer CT, Yao K, Pai J, et al. The cell cycle restricts activation-induced cytidine deaminase activity to early G1. J Exp Med. 2017;214:49–58.
Khalil AM, Cambier JC, Shlomchik MJ. B cell receptor signal transduction in the GC is short-circuited by high phosphatase activity. Science. 2012;336:1178–81.
Enomoto M, Goto H, Tomono Y, Kasahara K, Tsujimura K, Kiyono T, et al. Novel positive feedback loop between Cdk1 and Chk1 in the nucleus during G2/M transition. J Biol Chem. 2009;284:34223–30.
Mouhamad S, Besnault L, Auffredou MT, Leprince C, Bourgeade MF, Leca G, et al. B cell receptor-mediated apoptosis of human lymphocytes is associated with a new regulatory pathway of Bim isoform expression. J Immunol. 2004;172:2084–91.
Takada E, Furuhata M, Nakae S, Ichijo H, Sudo K, Mizuguchi J. Requirement of apoptosis-inducing kinase 1 for the induction of bronchial asthma following stimulation with ovalbumin. Int Arch Allergy Immunol. 2013;162:104–14.
Fischer SF, Bouillet P, O’Donnell K, Light A, Tarlinton DM, Strasser A. Proapoptotic BH3-only protein Bim is essential for developmentally programmed death of germinal center-derived memory B cells and antibody-forming cells. Blood. 2007;110:3978–84.
Greenow KR, Clarke AR, Jones RH. Chk1 deficiency in the mouse small intestine results in p53-independent crypt death and subsequent intestinal compensation. Oncogene. 2009;28:1443–53.
Boles NC, Peddibhotla S, Chen AJ, Goodell MA, Rosen JM. Chk1 haploinsufficiency results in anemia and defective erythropoiesis. PLoS ONE. 2010;5:e8581.
Höglund A, Nilsson LM, Muralidharan SV, Hasvold LA, Merta P, Rudelius M, et al. Therapeutic implications for the induced levels of Chk1 in Myc-expressing cancer cells. Clin Cancer Res. 2011;17:7067–79.
Murga M, Campaner S, Lopez-Contreras AJ, Toledo LI, Soria R, Montaña MF, et al. Exploiting oncogene-induced replicative stress for the selective killing of Myc-driven tumors. Nat Struct Mol Biol. 2011;18:1331–35.
Ferrao PT, Bukczynska EP, Johnstone RW, McArthur GA. Efficacy of CHK inhibitors as single agents in MYC-driven lymphoma cells. Oncogene. 2012;31:1661–72.
Myklebust JH, Brody J, Kohrt HE, Kolstad A, Czerwinski DK, Wälchli S, et al. Distinct patterns of B-cell receptor signaling in non-Hodgkin lymphomas identified by single-cell profiling. Blood. 2017;129:759–70.
Lam MH, Liu Q, Elledge SJ, Rosen JM. Chk1 is haploinsufficient for multiple functions critical to tumor suppression. Cancer Cell. 2004;6:45–59.
Bouillet P, Metcalf D, Huang DC, Tarlinton DM, Kay TW, Köntgen F, et al. Proapoptotic Bcl-2 relative Bim required for certain apoptotic responses, leukocyte homeostasis, and to preclude autoimmunity. Science. 1999;286:1735–38.
Lang MJ, Brennan MS, O’Reilly LA, Ottina E, Czabotar PE, Whitlock E, et al. Characterisation of a novel A1-specific monoclonal antibody. Cell Death Dis. 2014;5:e1553–e1553.
Vikstrom I, Carotta S, Lüthje K, Peperzak V, Jost PJ, Glaser S, et al. Mcl-1 is essential for germinal center formation and B cell memory. Science. 2010;330:1095–99.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Acknowledgements
We thank I. Gaggl and K. Rossi for expert technical assistance; M. Saurwein, N. Heinrich, and N. Schöpf for animal care; I. Lengenfelder and C. Rahm for administrative assistance; the present and former V.L. and A.V. lab members and E. Derudder for critical comments and suggestions; K. Rajewsky for Cg1-cre mice; A. Strasser for Bim− mice, T.W. Mak for Chk1F mice, M. Herold for providing an antibody against BCL2a1 (clone 6D6), and B. Weinberger for technical help with ELISPOT analysis. This work was supported by grants from the Austrian Science Fund (FWF) to AV (P 26856; I1298) and to NH-K (P 28694-B30), the Tyrolean Science Fund to VL (UNI-0404/1696) and the Austrian Cancer Aid to VL (KH15017). KS was supported by the FWF-funded Doctoral College “Molecular Cell Biology and Oncology” (W1101), and received a DOC PhD fellowship from the Austrian Academy of Sciences (ÖAW).
Author information
Authors and Affiliations
Contributions
Conceptualization, methodology, project administration, writing—original draft, and writing—review & editing: VL; investigation: KS, BJ, JH, CS, AA and VL; resources and funding acquisition: VL, AV, NH-K; visualization: VL, KS and BJ.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by C. Borner
Supplementary information
Rights and permissions
About this article
Cite this article
Schoeler, K., Jakic, B., Heppke, J. et al. CHK1 dosage in germinal center B cells controls humoral immunity. Cell Death Differ 26, 2551–2567 (2019). https://doi.org/10.1038/s41418-019-0318-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0318-5