Abstract
Neurons in the central nervous system (CNS) lose their intrinsic ability and fail to regenerate, but the underlying mechanisms are largely unknown. Polycomb group (PcG) proteins, which include PRC1 and PRC2 complexes function as gene repressors and are involved in many biological processes. Here we report that PRC1 components (polycomb chromobox (CBX) 2, 7, and 8) are novel regulators of axon growth and regeneration. Especially, knockdown of CBX7 in either embryonic cortical neurons or adult dorsal root ganglion (DRG) neurons enhances their axon growth ability. Two important transcription factors GATA4 and SOX11 are functional downstream targets of CBX7 in controlling axon regeneration. Moreover, knockdown of GATA4 or SOX11 in cultured DRG neurons inhibits axon regeneration response from CBX7 downregulation in DRG neurons. These findings suggest that targeting CBX signaling pathway may be a novel approach for promoting the intrinsic regenerative capacity of damaged CNS neurons.
Similar content being viewed by others
Introduction
Injured mature peripheral neurons successfully activate an array of regeneration-associated genes that enhance the intrinsic growth capacity to enable axon regeneration [1, 2]. In contrast, axons within the central nervous system (CNS) are not able to regenerate after injury largely because of the inhibitory extrinsic environment and their diminished intrinsic regenerative capability [3, 4]. The intrinsic growth capacity of neurons in CNS depends on the regulation of gene expression that supports axon growth, which gradually declines during neuronal maturation [4, 5]. Thus, defining gene expression that governs the intrinsic axon growth ability will provide critical perspectives for controlling axonal regeneration in the adult nervous system and develop novel therapeutic approaches to improve neuronal recovery following axon injury. However, our understanding of the important cellular and molecular mechanisms underlying intrinsic gene regulation within neurons that support axonal regrowth is very limited.
Epigenetic regulation, including DNA methylation, histone modification, and non-coding RNAs, is emerging to be a key cellular mechanism to control gene expression [6]. Several recent studies have explored the roles of epigenetic modifications such as histone acetylation and non-coding RNAs in axon regeneration. For example, the histone deacetylase 5 was identified as a novel injury-regulated tubulin deacetylase that has an essential role in growth cone dynamics and axon regeneration [7,8,9]. The other epigenetic mechanisms such as histone acetylation and non-coding RNAs also have pivotal roles in determining the regenerative capacity in neurons [1, 10].
Polycomb group (PcG) proteins function as gene repressors and are involved in the regulation of many processes in stem cell characteristics and differentiation into functional cells, which is first discovered as important developmental regulators in Drosophila [11]. In mammals, PcG proteins are found in several multiprotein complexes [12], the best characterized of which are Polycomb repressive complexes 1 and 2 (PRC1 and PRC2), which collaborate to repress gene transcription by catalyzing histone modifications [13]. PRC2 comprised three core components (EZH2, SUZ12, and EED) and PRC1 complex contains a single representative of the polycomb chromobox (CBX) family member, either CBX2, CBX4, CBX6, CBX7, or CBX8 [14]. The CBX family has important roles in the epigenetic regulation in many cell types [15,16,17]. CBX family members are involved in cancer progression and the Ink4a/ARF/Ink4b locus encoding three tumor suppressors is one of the earliest identified and the most well-known targets for CBX-containing PRC1 complex [18]. Consistently, knockdown of related CBX genes is generally associated with reduced cell proliferation [18]. Interestingly, several recent studies have shown that PcG also has important roles in the regulation of neuronal function [19, 20]. However, we still do not know which and how CBX subunits contributes to neuronal functions in postmitotic neurons, especially in regulation of axon growth and regeneration.
Here we examined whether CBX family members can respond to axon injury and have a role in axon regeneration. We found that CBX4, CBX6, and CBX7 gradually increase their expressions in the cortical tissues during development, and CBX family members respond differently after sciatic nerve injury in the peripheral nervous system. We provided evidences showing that CBX2, CBX7, and CBX8 are key regulators of axon regeneration. Especially, we showed that CBX7 is an intrinsic repressor of axon regeneration both in vitro and in vivo. Finally, we demonstrated that CBX7 modulates axon regeneration by repressing SOX11 and GATA4, and downregulation of SOX11 or GATA4 represses the axon regeneration ability of neurons with silenced CBX7.
Results
Distinct expression patterns of CBX family members that differentially affect CNS axon growth during cortical development
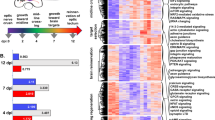
First, we assessed the expression level of all CBX family members in developing cortex by reverse-transcriptase PCR (RT-PCR) (Fig. 1a–e). The quantitative RT-PCR (qRT-PCR) revealed that expression levels of Cbx4, Cbx6, and Cbx7 gradually increased in the cortical tissues during development, reaching the highest level in adults (Fig. 1b–d). In contrast, Cbx2 gradually decreased during cortical development, whereas Cbx8 gradually increased to the highest level at P0 and then decreased to the same level as in E15 (Fig. 1a,e). Thus, CBX family members had distinct expression patterns during cortical development. To examine their expressions in neurons, we performed immunohistochemical staining of embryonic cortex tissue and immunochemical staining of cultured E15 cortical neurons. The results showed that CBX2, 4, 6, 7, and 8 were co-localized with neuronal markers TUJ1 and MAP2 (Fig. 1f,g).
CBX family genes are developmentally regulated during cortical development and differentially affect axon growth of embryonic cortical neurons. a–e Relative Cbx2, 4, 6, 7, and 8 expression levels in mouse cortical tissues during development from embryo to adult (n = 4, *p < 0.05, ** p < 0.01, ***p < 0.001). f Representative immunostaining images of CBX2, 4, 6, 7, and 8 with TUJ1 in E15 embryonic cortex tissues (TUJ1, green; CBXs, red). Scale bar, 100 μm. g Representative images of cultured E15 cortical neurons that electroporated with either NC-GFP plasmid or Cbx -shRNA-GFP vector at DIV 4. Neurons were immunostained with Tau-1 antibody, an axonal marker (red). Scale bar, 100 μm. h Cbx 2/6/7/8-shRNA-GFP vectors are efficient for knocking down corresponding CBX family member. RT-PCR data showed that the mRNA levels of CBX family member genes were dramatically decreased in E15 cortical neurons after 3 days of Cbx -shRNA vector electroporation (n = 3, *p < 0.05, **p < 0.01). i Representative images of E15 cortical neurons that electroporated with either NC-GFP plasmid or CBX-shRNA-GFP vector at DIV 4. Neurons were immunostained with Tau-1 (red). j Quantifications of axon length of E15 cortical neurons that were knocking down of CBX2, 6, 7, and 8 (n = 360 for control, n = 424 for Cbx2-shRNA, n = 322 for Cbx6-shRNA, n = 335 for Cbx7-shRNA, and n = 382 for CBX8-shRNA). Scale bar, 100 µm. (n = 3 repeats, *p < 0.05, ** p < 0.01). k. Cbx 6 mRNA level is increased after electroporation of Cbx6 overexpression plasmid (pCMV-Cbx6) into E15 cortical neurons (n = 3, *p < 0.05). l Cbx7 mRNA level is increased after electroporation of Cbx 7 overexpression plasmid (pCMV-CBX7) into E15 cortical neurons (n = 3, *p < 0.05). m Quantifications of axon length of overexpression Cbx6 and Cbx7 in E15 cortical neurons compared with control (n = 307 for control, n = 299 for Cbx6-OE and n = 329 for Cbx7-OE). (n = 3, *p < 0.05)
Next, to determine the intrinsic roles of CBX family members in the regulation of axon growth, we used mouse embryonic cortical neural cultures that are a well-established model system for studying axon growth. We constructed several short hairpin RNA (shRNA) vectors against each CBX family member and tested their efficiency and specificity for knocking down CBX in both E15 cortical neurons and adult dorsal root ganglion (DRG) neurons using electroporation. RT-PCR results showed that the constructed shRNA vectors significantly and specifically decrease the mRNA level of corresponding CBX family member in E15 cortical neurons (Fig. 1h and Supplemental Figures S1a-j). We then explored the functions of CBX family members in axon growth by electroporation of these Cbx-shRNA vectors into E15 cortical neurons. Quantitative analysis of axon length demonstrated that downregulation of Cbx4 has no effect on axon growth ability (Supplemental Figure S1k) but downregulation of Cbx2 or Cbx8 significantly impaired axon growth ability (Fig. 1i,j). Moreover, downregulation of Cbx6 or Cbx7 significantly promoted axon growth ability (Fig. 1i,j), suggesting that CBX6 and CBX7 may be intrinsic inhibitors of axon regeneration. To verify whether overexpression of Cbx6 and Cbx7 could inhibit axon growth in cortical neurons, we constructed pCMV-Cbx6 and pCMV-Cbx7 vectors that can significantly enhance expression levels of Cbx6 and Cbx7 after electroporation into E15 cortical neurons (Fig. 1k,l). As expected, the axon growth ability was dropped by 28.8% and 32.1%, respectively, after electroporation of pCMV-Cbx6 and pCMV-Cbx7 into cortical neurons compared with the control group (Fig. 1m, n = 3, *p < 0.05). These results indicated that CBX family members may be important intrinsic regulators for axon growth.
CBX family genes are dysregulated in adult sensory neurons after sciatic nerve injury
To determine whether CBX family members express in regenerating neurons, we did the immunochemical staining of cultured adult DRG neurons. CBX family proteins were mainly expressed in the nuclear and co-localized with TUJ1 in cultured DRG neurons (Fig. 2a). To test whether peripheral nerve injury is a stimulus for the in vivo activation of CBX family genes, we examined both mRNA and protein levels of CBX family members in injured DRGs, in comparison with the control naive group. Both qRT-PCR and western blotting results showed that CBX4, CBX6, and CBX7 were significantly downregulated in DRG neurons 1 week after the sciatic nerve injury (Fig. 2c,d,e). On the contrary, the level of CBX2 and CBX8 were drastically increased in adult DRGs 7 days after the sciatic nerve injury (Fig. 2b,f). Consistent with in vivo results, the expression levels of CBX family members were then verified in dissociated DRG neurons, which could mimic the in vivo injury after 3 days of culture (Supplemental Figures S2a-e). All these data suggested that peripheral nerve injury is a stimulus for the activation or deactivation of CBX family genes.
The expression levels of CBX family genes in adult DRG neurons are significantly responded to sciatic nerve injury in vivo. a Cultured adult DRG neurons were immunostained with CBX2/4/ 6/ 7/8, the neuronal marker Tuj1, and the nuclear marker DAPI. It is noteworthy that CBX proteins are mainly localized in the nucleus of DRG neurons (white arrowheads). b–f Comparison of CBX family members’ expression levels between naive and injured DRG tissues. b, Cbx2; c, Cbx4; d,Cbx 6; e,Cbx 7; f Cbx8. Left panel: quantification of CBX mRNA expression level (n = 5, *p < 0.05). Middle and right panels: western blotting and quantification of CBX protein levels (n = 6, *p < 0.05)
Downregulations of Cbx2 and Cbx8 impair axon regenerative ability of adult sensory neurons after sciatic nerve injury
We first studied the roles of CBX2 and CBX8 in the regulation of axon regeneration using adult sensory neurons from DRG, in which axon regenerate robustly after injury by reactivating their intrinsic regeneration capacity [21]. By using a recently developed electroporation model [1, 22], we first confirmed that Cbx2 and Cbx8-shRNA vectors could efficiently reduce Cbx2 and Cbx8 expression levels in cultured adult DRG neurons, respectively (Fig. 3a,b). As other CBX family members had no expression changes after Cbx2 and Cbx8 knockdown (Supplemental Figs. S3a and 3e), these shRNA vectors could specifically target CBX2 and CBX8 in DRG neurons. Functionally, we discovered that downregulation of Cbx2 or Cbx8 markedly suppressed regenerative axon growth of adult DRG neurons after 3 days of culture (Fig. 3c,d,e,f). Similar results were then obtained from embryonic cortical neurons (Fig. 1i,j), suggesting that CBX2 and CBX8 are intrinsic regulators of axon regeneration.
Downregulations of Cbx2 and 8 impair mammalian axon regeneration in DRG neurons. a, b The mRNA levels of Cbx2 and 8 after their knockdown by constructed shRNA-vectors. Cbx2-shRNA-1 and -2 in a, and Cbx8-shRNA-1 and -2 in b. The results showed that Cbx2 and Cbx8 genes dramatically decreased in DRG neurons after 4 days of the shRNA vectors electroporation (n = 3, *p < 0.05, **p < 0.01). c Representative images of adult DRG neurons expressing NC-GFP, CBX2-shRNA-1, and -2 vectors separately, as indicated, and fixed at DIV4 after electroporation. Neurons were immunostained with Tuj1-1 (red). Knocking down CBX2 with shRNAs in cultured adult DRG neurons significantly inhibited regenerative axon growth from adult DRG neurons. Scale bar, 100 µm. d Representative images of DRG neurons expressing NC-GFP, CBX8-shRNA-1, and 2 vectors separately, as indicated, and fixed at DIV4 after electroporation. Neurons were immunostained with neuronal marker Tuj1-1 antibody (red). Knocking down Cbx8 with shRNAs in cultured adult DRG neurons significantly inhibited regenerative axon growth ability. Scale bar, 100 µm. e The quantification of axon length after knocking down Cbx2 compared with control (n = 470 for control, n = 360 for Cbx2-shRNA-1 and n = 329 for CBX2-shRNA-2) (n = 3 repeats, *p < 0.05). f The quantification of axon lengths after knocking down Cbx8 compared with control (n = 470 for control, n = 318 for Cbx8-shRNA-1, and n = 302 for Cbx8-shRNA-2) (n = 3, *p < 0.05)
Downregulation of Cbx7 is necessary for regenerative axon growth of adult sensory neurons after sciatic nerve injury
To investigate the functions of CBX 4, 6, and 7 in the regulation of axon regeneration of adult DRG neurons, dissociated DRG neurons were electroporated with Cbx4-shRNA-1, -2, Cbx6-shRNA-1, -2, or Cbx7-shRNA-1 and -2 plasmids, which could significantly and specifically inhibit Cbx4, Cbx6, and Cbx7 expression levels in DRG neurons (Supplemental Figures S3b, S3c, S3d, S4a, and S4b). Quantitative analysis of axon growth demonstrated that downregulation of Cbx4 or Cbx6 had no effects on axon regeneration. However, downregulation of Cbx7 increased 43.9–53.6% of regenerative axon growth ability of adult DRG sensory neurons compared with the control group (Fig. 4a,b), which was very similar to that of embryonic cortical neurons (Fig. 1i,j). To further confirm the functional role of CBX7 in axon regeneration, we knocked down endogenous Cbx7 with a group of three different small interfering RNAs (siRNAs) that were designed to minimize the off-target effects (ON-TARGET plus, Shanghai GenePharma Co., Ltd) (Fig. 4c and Supplemental Figure S3f). We found that acute knockdown of Cbx7 also promoted regenerative axon growth of adult DRG neurons (Fig. 4d,e,f). Next, we constructed Cbx7 overexpression vector (pCMV-CBX7), which could highly express Cbx 7 in DRG neurons (Fig. 4g) and electroporated it into dissociated DRG neurons together with enhanced green fluorescent protein (EGFP) expression plasmid. We found that overexpression of Cbx7 significantly blocked axon growth of adult DRG neurons compared with the control (Fig. 4h,i). These findings further supported the conclusion that CBX7 may function as an intrinsic inhibitor of axon regeneration.
Downregulation of CBX7 promotes mammalian axon regeneration in DRG neurons in vitro. a Representative images of adult DRG neurons expressing NC-GFP, Cbx7-shRNA-1, and CBX7-shRNA-2 vectors separately, as indicated, and fixed at DIV4 after electroporation. Neurons were immunostained with neuronal marker Tuj1-1 antibody (red). Knocking down CBX7 with shRNAs in cultured adult DRG neurons significantly promoted regenerative axon growth ability from adult DRG neurons. Scale bar, 100 µm. b The quantification of axon lengths after knocking down CBX7 with CBX7-shRNA vectors (n = 310 for control, n = 331 for CBX7-shRNA-1 and n = 290 for CBX7-shRNA-2) (n = 3, *p < 0.05). Scale bar, 100 µm. c qRT-PCR data indicating Cbx7 mRNA level in adult DRGs in vivo 3 d after electroporation of the Cbx7 on-target siRNAs. (n = 3, *p < 0.05). d, e Knocking down Cbx7 with siRNAs resulted in significantly enhanced axon growth of adult DRG neurons that cultured on coverslips coated with poly-d-lysinse, which provided a less favorable condition for axon growth (n = 296 for control, n = 332 for CBX7-siRNA) (n = 5 repeats, *p < 0.05). f Cumulative distribution of the lengths of all individual axons measured (control, n = 296; si-CBX7, n = 332). (n = 5 repeats, *p < 0.05). g CBX7 high expression is shown after electroporation of Cbx7-OE plasmid into DRG neurons by western blotting. h Representative images of CBX7 overexpression in cultured adult DRG neurons with significantly inhibited axon regeneration ability. Scale bar, 200 µm. i Quantification of axon lengths of cultured adult DRG neurons with Cbx7 overexpression (Cbx7-OE) (n = 293 for control, n = 293 for Cbx6-OE, and n = 273 for Cbx7-OE). Scale bar, 200 µm. (n = 4, **p < 0.01)
Overexpression of CBX7 inhibits mammalian axon regeneration in DRG neurons in vivo
To investigate whether Cbx7 overexpression was able to prevent peripheral axotomy-induced axon regeneration in sensory neurons in vivo, we electroporated both Cbx7-OE and EGFP plasmids into L4 or L5 adult mouse DRG neurons [1, 22, 23]. The transfected DRGs were collected and detected CBX7 protein expression 3 days later. Western blotting assay showed that electroporation of Cbx7-OE plasmid into adult mouse DRGs could significantly increase CBX7 protein level in DRG tissues (Fig. 5b). With the same electroporation protocol, the mice were subjected to sciatic nerves crush procedure 2 days after in vivo electroporation, and axon regeneration were assessed 3 days later. We used whole-mount sciatic nerves to trace the full path of every EGFP-labeled axon from the crush site and found that sensory axons of CBX7-overexpressed neurons displayed significantly impaired axon regeneration compared with the control neurons that were expressing EGFP alone (Fig. 5a,c,d,e). Therefore, these results suggested that overexpression of CBX7 alone is sufficient to reduce axon regeneration ability in sensory neurons in vivo.
Overexpression of Cbx7 inhibits mammalian axon regeneration in DRG neurons in vivo. a Representative images of axon regeneration of sciatic nerves. After transfection of GFP and Cbx7-OE plasmid into the DRGs, mice were subjected to a sciatic nerve crush procedure. Using the whole-mount nerve segment, the lengths of all identifiable regenerating axons were measured from the crush site (marked by epineural suture, red line) to the distal axon tips. b CBX7 high expression is shown after electroporation of Cbx7-OE plasmid into adult DRG neurons by western blotting. c Quantification of axon regeneration length (GFP, n = 6; Cbx7-OE, n = 6) are shown (n = 284 for control; n = 268 for Cbx 7-OE). Scale bar, 1 mm. d In vivo regulation index of axon lengths of all individual axons measured (n = 284 for control; n = 268 for Cbx7-OE). e Cumulative distribution of the lengths of all individual axons measured (n = 284 for control; n = 268 for Cbx 7-OE)
CBX7 targets GATA4 and SOX11 for controlling axon regeneration in adult DRG neurons
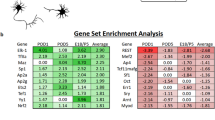
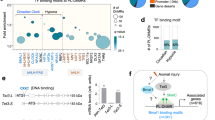
To investigate the molecular mechanism by which CBX7 regulates axon regeneration, we tested several potential targets of CBX7 in neurons by referencing its experimentally validated targets in other cell types and other published axon regeneration related genes (Fig. 6a). In embryonic stem cells (ESCs), CBX7 balances self-renewal and differentiation of ESCs by suppressing many important genes, such as GATA4 [16, 24, 25]. More recently, it has also been demonstrated that CBX7, as a suppressor, is able to differently modulate the expression of Wnt genes that are involved in cancer progression [26,27,28]. Our RT-PCR analysis showed that Gata4, Sox11, Wnt1, and Wnt10bb were dramatically increased in cultured adult DRG neurons with downregulation of Cbx7 expression (Fig. 6a). Meanwhile, mRNA levels of Wnt1, Wnt10b, Gata4, and Sox11 were also significantly upregulated in adult sensory neurons during peripheral nerve injury-induced axon regeneration (Fig. 6b,c,d,e). As CBX7 has been identified to bind the promoters for suppressing activity of its downstream target genes [29], we therefore tested whether CBX7 is able to bind to the promoters of the above tested downstream targets in adult DRG neurons. To do this, we performed chromatin ChiP using an anti-CBX7 antibody in DRGs tissues. Our qRT-PCR data showed that the binding of CBX7 within the promoter regions of GATA4 and SOX11 increased significantly in peripheral DRGs compared with IgG control (Fig. 6f). Next, we performed Chip assay on 3-day-cultured DRG neurons mimicking peripheral axotomized DRGs in which CBX7 expression was downregulated, and the results confirmed that the enrichment of CBX7 with the promoter regions of GATA4 and SOX11 was decreased significantly in 3-day-cultured DRG neurons compared with that in Day 0 DRGs (Fig. 6g). We then examined whether CBX7 is able to regulate GATA4 and SOX11 expression in adult DRG tissues. We found that expressions of endogenous GATA4 and SOX11 were drastically upregulated when CBX7 was knocking down in adult DRG tissues (Fig. 6h,i). Collectively, these findings clearly supported the idea that CBX7 directly repressed GATA4 and SOX11 expressions in adult DRG neurons.
CBX7 Targets SOX11 and GATA4 for controlling axon regeneration in adult DRG neurons. a Relative gene expression levels after CBX7 knockdown in cultured DRG sensory neurons (n = 4, *p < 0.05, **p < 0.01). b–e The mRNA levels of Wnt1, Wnt10b, Sox11, and Gata4 are significantly increased in sensory neurons in vivo 7 days after sciatic nerve injury. (b, n = 9; c, n = 6; d, n = 7; e, n = 4; f, n = 3; *p < 0.05, **p < 0.01, ***p < 0.0001). f The enrichment of CBX protein at genomic sequence upstream of the target gene locus in DRG tissues, as assessed by CBX7-specific chromatin immunoprecipitation (ChIP). ChIP with IgG antibody in DRG tissues was used as negative controls. Relative enrichment of CBX7 to each target gene in DRG tissues was calculated relative to IgG-only nonspecific control in the same group (n = 4, *p < 0.05, **p < 0.01). Quantities were calculated from an input DNA-generated standard curve. Two-way ANOVA, Bonferroni post test. g The enrichment of CBX protein at genomic sequence upstream of the target gene locus in regenerated cultured DRG neurons, as assessed by CBX7-specific ChIP. (n = 3, *p < 0.05, **p < 0.01). Quantities were calculated from an input DNA-generated standard curve. Two-way ANOVA, Bonferroni post test. h Gata4 mRNA (n = 3; ** P < 0.01) level was increased in adult DRGs 3 days after in vivo electroporation of Cbx 7 siRNA. i Sox11 mRNA (n = 3; ** P < 0.01) level was increased in adult DRGs 3 days after in vivo electroporation of Cbx7 siRNA
Downregulation of GATA4 and SOX11 blocks the axon regeneration ability of adult DRG neurons with CBX7 knockdown
Consistent with findings from previous in vivo sciatic nerve injury, we found that the expression levels of SOX11 and GATA4 were significantly upregulated in dissociated DRG neurons after 3 days of culture compared with freshly collected DRG neurons (Figs. 7a and 8a). As downregulation of CBX7 increased the expressions of SOX11 and GATA4, we first examined whether SOX11 and GATA4 had a role in axon regeneration in adult DRG neurons. With electroporation of a group of three different siRNAs that were designed to minimize the off-target effects (ON-TARGET plus, Shanghai GenePharma Co., Ltd), we found that acute downregulation of SOX11 resulted in impaired regenerative axon growth of adult DRG neurons (Fig. 7b–d), which was consistent with published findings [30, 31]. Similarly, acute knockdown of GATA4 also decreased regenerative axon growth of adult DRG neurons (Fig. 8c,d). To further validate SOX11 and GATA4 as the downstream targets of CBX7, we did electroporation of siRNAs of CBX7, SOX11, and GATA4 into DRG neurons, and found that downregulations of SOX11 (Fig. 7e,f) and GATA4 (Fig. 8e,f) blocked the axon regeneration ability of adult DRG neurons with CBX7 knockdown. Therefore, these data further supported that CBX7 acted as an intrinsic inhibitor of axon regeneration via downregulating the expressions of SOX11 and GATA4.
Downregulation of SOX11 restores the axon regeneration phenotype exhibited by CBX7 knockdown in adult DRG neurons. a Comparison of Sox11 expression level in cultured DRG sensory neurons between DIV0 and DIV3. Sox11 expression level is much higher at DIV3 than those at DIV0 (n = 6, **p < 0.01). b Sox11 mRNA level was tested after 3 days of siSOX11 electroporation into DRG sensory neurons (n = 3, *p < 0.05) and the mRNA level of Sox11 quantified using qRT-PCR and normalized to the expression of the GAPDH. c Adult DRG neurons were transfected with GFP plus siRNA ctrl and siRNAs against SOX11 (siSOX11) as indicated. At 3 days after transfection, neurons were fixed and immunostained with anti-GFP and anti-TuJ1 antibodies. Representative images of neurons transfected with siRNAs and GFP are shown. d Quantification of axon length of adult DRG neurons transfected with GFP or siSox11 (n = 148 for control, n = 206 for siSOX11) (n = 3, *P < 0.05). Scale bar, 100 µm. e Representative images of cultured adult sensory neurons showing that knocking down SOX11 (siSox11) restored regenerative axon growth phenotype by knocking down CBX7 (siCBX7). At Day 4 after transfection, neurons were fixed and immunostained with anti-GFP and anti-TuJ1 antibodies. Scale bar, 100 µm. f Quantification of axon length of adult DRG neurons transfected with GFP, siCBX7, siSOX11, and siCBX7 plus siSOX11 (n = 271 for control, n = 205 for siCBX7, n = 204 for siSOX11, and n = 177 for siCBX7 + siSOX11) (n = 5, *p < 0.05)
Downregulation of GATA4 restores the axon regeneration phenotype exhibited by Cbx7 knockdown in adult DRG neurons. a Comparison of GATA4 expression level in cultured DRG sensory neurons between DIV0 and DIV3 (n = 6, p < 0.01). b Gata4 mRNA level was tested after 3 days of GATA4 electroporation into DRG sensory neurons (n = 3, *p < 0.05). The Gata4 level was quantified using qRT-PCR and normalized to the expression of the GAPDH. c Adult DRG neurons were transfected with GFP siRNA ctrl and siRNAs against Gata4 (siGata4) as indicated. At 3 days after transfection, neurons were fixed and immunostained with anti-GFP and anti-TuJ1 antibodies. Representative images of neurons transfected with siRNAs and GFP are shown. d Quantification of axon length of adult DRG neurons transfected with GFP plus siRNA ctrl or siGata4 (n = 148 for control, n = 185 for siGATA4) (n = 3, *P < 0.05). Scale bar, 100 µm. e Downregulation of Gata4 could restore the increased axon length of adult DRG caused by downregulation of CBX7 as measured GFP and Tuj1 positive cells. Representative images are shown. Scale bar, 100 µm. f Quantification of axon length of adult DRG neurons transfected with GFP, siCbx7, siGata4, or siCbx7 plus siGata4 (n = 82 for control, n = 74 for siCbx7, n = 70 for siGata4, and n = 48 for siCbx7 + siGata4) (n = 4 repeats, *p < 0.05)
Discussion
CBX family proteins have emerged as key regulators of cancer progression [18], pluripotency, and differentiation of mouse ESCs [24]. This study demonstrates that CBX family members have differential expressions during the cortical development and response differently to sciatic nerve injury. CBX7 acts as a key intrinsic modulator of axon regeneration by directly suppressing the expressions of SOX11 and GATA4 in DRG neurons.
CBX family genes have been shown having different expression levels during the differentiation process of mouse ESCs [24], and distinct development-stage-specific expression patterns of CBX family members are also existed during hematopoietic differentiation [14]. In our study, both mRNA and protein levels of CBX2 gradually decreases during cortical development, and knocking down CBX2 in embryonic cortical neurons significantly inhibits their axon regeneration ability, suggesting that restoration of CBX2 expressions in adult neurons may be beneficial for axon regeneration. Remarkably, CBX family proteins in adult DRG neurons also show different responses to the sciatic nerve injury. Although CBX4, CBX6, and CBX7 are downregulated, CBX2 and CBX8 are upregulated in adult DRGs 1 week after the sciatic nerve injury. More functional analyses and possible interactions of CBX family members during cortical development and in response to sciatic nerve injury need to be elucidated in the future.
As a transcriptional suppressor, CBX7 regulates many downstream target genes that are responsible for controlling life span, stem cell pluripotency, or cancer progress [14, 18, 26]. In our study, we found that CBX7 is an intrinsic negative regulator of axon growth and regeneration in both embryonic cortical neurons and adult DRG neurons. At present, several specific pharmacological inhibitors of CBX7 have been identified and used in vivo [32, 33]. It will be of great interest to determine if systematic administration of these drugs would promote axon regeneration in the future. Furthermore, it will be important to determine if deleting CBX7, either genetically or pharmacologically, in cortical motor neurons would also promote corticospinal tract axon regeneration after spinal cord injury. Our findings of impaired axon regenerative ability of adult sensory neurons with silencing CBX2 or CBX8 suggest that targeting the signaling pathways involved by other CBX family members may also have therapeutic potentials for axon regeneration.
This study has identified SOX11 and GATA4 as the downstream targets of CBX7 in controlling axon regeneration in adult DRG neurons. SOX11 is a well-known regulator of axon regeneration; overexpression of SOX11 has been shown to be important for promoting axon regeneration [30, 31, 34]. Consistent with this, our study demonstrated that downregulation of SOX11 in adult DRG neurons impaired axon regeneration ability. GATA4 is a GATA zinc-finger transcription factor family member, which has been identified as a functional downstream target of CBX7 to regulate differentiation, growth, and survival of a wide range of cell types [35, 36]. Especially, GATA4 is an important transcription factor for heart development and repair after injury [37, 38]. Recent evidence have shown that GATA4 is regulated and modified by posttranscriptional protein modification such as acetylation, phosphorylation, and SUMOylation during heart development or ESC differentiation [39]. For the first time, our study identified GATA4 as a novel regulator of axon regeneration. Although downregulations of SOX11 and GATA4 have apparent rescue of normal neurite length in neurons with silenced CBX7, we cannot rule out the possibility that GATA4 and SOX11 generally affect neurite length after injury, regardless of the status of CBX7.
So far, several genes (e.g., KLF4 and PTEN) have been identified as important regulators of axon regeneration [40, 41]. Our finding suggests that manipulating multiple CBX genes along with GATA4 and SOX11 signaling pathways may also be a useful approach to promote the intrinsic regenerative capacity of mature CNS neurons damaged by injury or disease. Recent studies have indicated that CBX7 is involved in H3K27me3 modification [42, 43], a key epigenetic mechanism for maintaining the silencing of developmental regulators. Thus, future researches to identify more effector genes of CBX7 will be of great interest to elucidate how CBX7 regulates the axotomy-induced genetic program supporting axon regeneration.
Materials and methods
Animals
All animals involved in the experiments were performed in accordance with the animal protocol approved by the Institutional Animal Care and Use Committee of Institute of Zoology, Chinese Academy of Sciences. Eight- to 12-week-old adult female CD-1 mice (weighing 30–35 g) were purchased from Vital River Laboratory Animal Technology Co. Ltd and housed in the Institute Animal Facility.
Reagents and antibodies
The anti-β- III tubulin antibody (TuJ1) was bought from Covance (cat number Chantilly, VA). Antibodies AP17146C-EV20 CBX2 Antibody (Center) and AP2514A-EV20 PC2 (CBX4) Antibody (N-term) were purchased from Abgent. Antibody against CBX6 (Millipore 09–030) and CBX6 (sc-86355) was ordered from Millipore company and Santa-Cruz. Ab 21873 GR79485 Rabbit polyclonal to CBX7 was from Abcam. Antibody against CBX8 (AP2515b) was purchased from Abgent. The internal control actin antibody was from Sigma-Aldrich (St. Louis, MO). All fluorescence-tagged secondary antibodies such as goat anti-mouse Alexa Fluor 568 (A11031; Invitrogen), goat anti-rabbit Alexa Fluor 647 (A21245; Invitrogen), goat anti-rat Alexa Fluor 568 (A11077; Invitrogen), goat anti-rabbit Alexa Fluor 488 (A11008; Invitrogen), and goat anti-chicken Alexa Fluor 488 (A11039; Invitrogen) were from Life Technology Molecular Probes, Inc.
Primary neuron culture and in vitro transfection
Dissection and culture of mouse embryonic cortical and adult DRG neurons were performed as described previously [1, 23, 44]. DNA constructs or RNA oligos were transfected into dissociated neurons via electroporation using the 4D-Nucleofector system from Lonza (Cologne, Germany) as previously described [1, 44, 45]. Briefly, dissociated neurons were centrifuged to remove the supernatant and re-suspended in 80–100 μl of specified Amaxa electroporation buffer with plasmid DNA (3–10 μg and 2–3 μg for DRG and cortical neurons, respectively) or RNA oligos. Suspended cells were then transferred to a 100 µl single Nucleocuvette and electroporated with the Amaxa Nucleofector apparatus. After electroporation, cells were immediately mixed with the desired volume of pre-warmed culture medium and transferred to the culture dish. After neurons fully attached to the substrates (3–4 h), the medium was changed to remove the remnant transfection buffer.
Immunostaining, fluorescence microscopy, and image analysis
Neurons were fixed with 4% Paraformaldehyde (PFA) at room temperature for 10 min. Fixed neurons were washed with phosphate-buffered saline (PBS) and blocked in blocking solution (2% bovine serum albumin, 0.1% Triton X-100, and 0.1% sodium azide in PBS). Primary and secondary antibodies were diluted with the blocking buffer and PBS, respectively, then incubated for 1 h each at room temperature. After the immunostaining, the coverslips were extensively rinsed with PBS and mounted onto glass slides for observation. DRG neurons were viewed with Leica DMI3000 B inverted microscope. Images were captured with a charge-coupled device camera controlled by Leica Application Suite V4 software (Leica Microsystems, Ltd). For embryonic cortical neurons, the axons were manually traced and the axon lengths were recorded. For adult DRG neurons, the longest axon of each neuron was traced and measured. Axon length was measured with the “image J” software. For quantification of axon length, we restricted the analysis to neurons with processes longer than one cell body in diameter. The mean and SEM of neurite-bearing cells were calculated from at least three independent experiments.
In vivo electroporation of adult DRG neurons and quantification of axon regeneration
The in vivo electroporation of adult mouse DRGs was performed as described previously [46]. Briefly, under anesthesia induced by katamine (100 mg/kg) and xylazine (10 mg/kg), a small dorsolateral laminectomy was performed to expose the left L4–L5 DRGs. EGFP plasmid or EGFP plus CBX7 overexpression plasmid were injected into the DRGs using pulled glass capillaries (1.5 μl per ganglion). Immediately after injection, electroporation was performed by applying five pulses of current (35 V, 10 ms, 950 ms interval) using a custom-made tweezer-like electrode powered by the Electro Square Porator ECM830 (BTX Genetronics). The wound was then closed and the mice were allowed to recover. Two days after the electroporation, the sciatic nerves were crushed with fine forceps and the crushed sites were marked with nylon epineuria sutures. Three days later, the mice were perfused with 4% PFA in sodium phosphate buffer (pH 7.4). The whole nerve segment was then dissected out and further fixed in 4% PFA overnight at 4 °C.
For quantification of in vivo axon regeneration, the fluorescence images of whole mount nerves were first obtained. All identifiable EGFP-labeled axons in the sciatic nerve were then manually traced from the crush site to the distal growth cone to measure the length of axon regeneration.
Real-time PCR relative quantification of CBX family genes mRNA
Total RNA were extracted from tissues and cultured cells according to procedures using Trizol reagent(Invitrogen). Briefly, 1 μg of total RNA was reverse transcribed using either oligo(dT) primers or specific primers (TransScript One-Step gDNA Removal and cDNA synthesis Kit; TRANS) according to the manufacturer’s protocol. For real-time PCR analysis, according to the manufacturer’s instructions, using a SYBR Premix Ex Taq (Tli RNaseH Plus) (Takara), 25 ng of cDNA and 0.5 mM primers in a final volume of 20 μl. The PCR steps were performed 30 s pre-denaturation at 95 °C, followed by 45 cycles of 10 s denaturation at 94 °C, 30 s annealing at 60 °C, 30 s extension at 72 °C. The analysis of RT-PCR used the 2–ΔΔCT method. All values were normalized with respect to the Gapdh mRNA level in each sample. Primers for real-time RT-PCR are as follows: Gapdh forward: 5′-AGGTCGGTGTGAACGGATTTG-3′, Gapdh reverse: 5′-TGTAGACCATGTAGTTGAGGTCA-3′. The sequences of other primers used are listed in the Supplemental Table 2.
Western blot analysis
Protein samples from DRG tissues or dissociated DRG neurons were collected and lysed using the RIPA buffer. The extracted proteins were separated by 10% gradient SDS/PAGE gel electrophoresis, and transferred onto polyvinylidene fluoride membranes. Membranes were processed following the ECL Western blotting protocol (GE Healthcare). After blocking with 3% non-fat milk, membranes were incubated with primary antibodies (4 °C, overnight) followed by horseradish peroxidase-linked secondary antibodies (room temperature, 1 h). The densities of protein bands from three independent experiments were quantified using the Image J software. The artificial unit of each protein was calculated by normalizing the band of the protein of interest to the band of loading control, β-actin.
Chromatin immunoprecipitation
Chromatin immunoprecipitation was performed according to the published method [1, 6, 47]. Briefly, the DRG samples were cross-linked with 1% formaldehyde for 10 min at room temperature followed by quenched with glycine. Chromatin lysates were sonicated (6 pulses, 10 s each at a power output of 40%, with 1 min incubations on ice in between each pulse) to shear the genomic DNA into 200 and 1,000 base pair fragments. Immunoprecipitation was performed with anti-CBX7 (Abcam) and IgG was used as a negative control. Precipitated DNA was detected by qPCR with specific primers. The primers we used are as follows: Sox11 FW-5′-CCCTTTCCTCTGCCTAGCTT-3′, RV-5′-CTGGGACTGTAGGTGCCAAT-3′; GATA4: FW-5′-CGAGTGCCTTTCTTCAGTCC-3′, RV-5′- CAAAAACCGTGAGTGTGGTG-3′; Wnt1 FW-5′-TTGAGATCCCTGGTGAAAGG-3′, RV-5′-GCGCCTGCTATGAAAGAGAG-3′; Wnt10b: FW-5′- ACACAAAGAGAAGCGAGTTTCC-3′, RV-5′- TTTAACCCAACGGTACAAACC-3′.
Statistical analysis
Data are presented as mean ± SEM. The Student’s t-test was used to determine the statistical significance between different experimental groups, which was set at a value of p < 0.05.
References
Liu CM, Wang RY, Saijilafu, Jiao ZX, Zhang BY, Zhou FQ. MicroRNA-138 and SIRT1 form a mutual negative feedback loop to regulate mammalian axon regeneration. Genes Dev. 2013;27:1473–83.
Liu K, Tedeschi A, Park KK, He Z. Neuronal intrinsic mechanisms of axon regeneration. Annu Rev Neurosci. 2011;34:131–52.
Liu CM, Hur EM, Zhou FQ. Coordinating gene expression and axon assembly to control axon growth: potential role of GSK3 signaling. Front Mol Neurosci. 2012;5:3.
He Z, Jin Y. Intrinsic control of axon regeneration. Neuron. 2016;90:437–51.
Weng YL, Joseph J, An R, Song H, Ming GL. Epigenetic regulation of axonal regenerative capacity. Epigenomics. 2016;8:1429–42.
Liu C, Teng ZQ, Santistevan NJ, Szulwach KE, Guo W, Jin P, et al. Epigenetic regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell. 2010;6:433–44.
Cho Y, Sloutsky R, Naegle KM, Cavalli V. Injury-induced HDAC5 nuclear export is essential for axon regeneration. Cell. 2013;155:894–908.
Cho YC, Sloutsky R, Naegle KM, Cavalli V. Injury-induced HDAC5 nuclear export is essential for axon regeneration (vol 155, pg 894, 2013). Cell. 2015;161:691–691.
Cho Y, Cavalli V. HDAC5 is a novel injury-regulated tubulin deacetylase controlling axon regeneration. EMBO J. 2012;31:3063–78.
Puttagunta R, Tedeschi A, Soria MG, Hervera A, Lindner R, Rathore KI, et al. PCAF-dependent epigenetic changes promote axonal regeneration in the central nervous system. Nat Commun. 2014;5:3527.
Lewis EB. A gene complex controlling segmentation in Drosophila. Nature. 1978;276:565–70.
Simon JA, Kingston RE. Mechanisms of polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol. 2009;10:697–708.
Margueron R, Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469:343–9.
Klauke K, Radulovic V, Broekhuis M, Weersing E, Zwart E, Olthof S, et al. Polycomb Cbx family members mediate the balance between haematopoietic stem cell self-renewal and differentiation. Nat Cell Biol. 2013;15:353–62.
Sparmann A, van Lohuizen M. Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer. 2006;6:846–56.
O’Loghlen A, Munoz-Cabello AM, Gaspar-Maia A, Wu HA, Banito A, Kunowska N, et al. MicroRNA regulation of Cbx7 mediates a switch of Polycomb orthologs during ESC differentiation. Cell Stem Cell. 2012;10:33–46.
Landeira D, Sauer S, Poot R, Dvorkina M, Mazzarella L, Jorgensen HF, et al. Jarid2 is a PRC2 component in embryonic stem cells required for multi-lineage differentiation and recruitment of PRC1 and RNA Polymerase II to developmental regulators. Nat Cell Biol. 2010;12:618–U214.
Gil J, Bernard D, Martinez D, Beach D. Polycomb CBX7 has a unifying role in cellular lifespan. Nat Cell Biol. 2004;6:67–U19.
Henriquez B, Bustos FJ, Aguilar R, Becerra A, Simon F, Montecino M, et al. Ezh1 and Ezh2 differentially regulate PSD-95 gene transcription in developing hippocampal neurons. Mol Cell Neurosci. 2013;57:130–43.
Palomer E, Carretero J, Benvegnu S, Dotti CG, Martin MG. Neuronal activity controls Bdnf expression via Polycomb de-repression and CREB/CBP/JMJD3 activation in mature neurons. Nat Commun. 2016;7:11081.
Liu CM, Hur EM, Zhou FQ. Coordinating gene expression and axon assembly to control axon growth: potential role of GSK3 signaling. Front Mol Neurosci. 2012;5:3.
Saijilafu, Hur EM, Liu CM, Jiao Z, Xu WL, Zhou FQ. PI3K-GSK3 signalling regulates mammalian axon regeneration by inducing the expression of Smad1. Nat Commun. 2013;4:2690.
Jiang JJ, Liu CM, Zhang BY, Wang XW, Zhang M, Saijilafu, et al. MicroRNA-26a supports mammalian axon regeneration in vivo by suppressing GSK3beta expression. Cell Death Dis. 2015;6:e1865.
Morey L, Pascual G, Cozzuto L, Roma G, Wutz A, Benitah SA, et al. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell Stem Cell. 2012;10:47–62.
Morey L, Aloia L, Cozzuto L, Benitah SA. Di Croce L. RYBP and Cbx7 define specific biological functions of polycomb complexes in mouse embryonic stem cells. Cell Rep. 2013;3:60–69.
Forzati F, Federico A, Pallante P, Abbate A, Esposito F, Malapelle U, et al. CBX7 is a tumor suppressor in mice and humans. J Clin Invest. 2012;122:612–23.
Pallante P, Sepe R, Federico A, Forzati F, Bianco M, Fusco A. CBX7 modulates the expression of genes critical for cancer progression. PLoS ONE. 2014;9:e98295.
Kim HY, Park JH, Won HY, Lee JY, Kong G. CBX7 inhibits breast tumorigenicity through DKK-1-mediated suppression of the Wnt/beta-catenin pathway. FASEB J. 2015;29:300–13.
Pallante P, Forzati F, Federico A, Arra C, Fusco A. Polycomb protein family member CBX7 plays a critical role in cancer progression. Am J Cancer Res. 2015;5:1594–601.
Jing XT, Wang T, Huang SH, Glorioso JC, Albers KM. The transcription factor Sox11 promotes nerve regeneration through activation of the regeneration-associated gene Sprr1a. Exp Neurol. 2012;233:221–32.
Jankowski MP, McIlwrath SL, Jing XT, Cornuet PK, Salerno KM, Koerber HR, et al. Sox11 transcription factor modulates peripheral nerve regeneration in adult mice. Brain Res. 2009;1256:43–54.
Ren C, Morohashi K, Plotnikov AN, Jakoncic J, Smith SG, Li J, et al. Small-molecule modulators of methyl-lysine binding for the CBX7 chromodomain. Chem Biol. 2015;22:161–8.
Simhadri C, Daze KD, Douglas SF, Quon TTH, Dev A, Gignac MC, et al. Chromodomain antagonists that target the Polycomb-group methyllysine reader protein chromobox homolog 7 (CBX7). J Med Chem. 2014;57:2874–83.
Norsworthy MW, Bei FF, Kawaguchi R, Wang Q, Tran NM, Li Y, et al. Sox11 expression promotes regeneration of some retinal ganglion cell types but kills others. Neuron. 2017;94:1112–20.
Mohammad HP, Cai Y, McGarvey KM, Easwaran H, Van Neste L, Ohm JE, et al. Polycomb CBX7 promotes initiation of heritable repression of genes frequently silenced with cancer-specific DNA hypermethylation. Cancer Res. 2009;69:6322–30.
Xu M, Millard RW, Ashraf M. Role of GATA-4 in differentiation and survival of bone marrow mesenchymal stem cells. Prog Mol Biol Transl Sci. 2012;111:217–41.
Yu W, Huang X, Tian X, Zhang H, He L, Wang Y, et al. GATA4 regulates Fgf16 to promote heart repair after injury. Development. 2016;143:936–49.
Stefanovic S, Barnett P, van Duijvenboden K, Weber D, Gessler M, Christoffels VM. GATA-dependent regulatory switches establish atrioventricular canal specificity during heart development. Nat Commun. 2014;5:3680.
Kee HJ, Kook H. Roles and targets of class I and IIa histone deacetylases in cardiac hypertrophy. J Biomed Biotechnol. 2011;2011:928326.
Moore DL, Blackmore MG, Hu Y, Kaestner KH, Bixby JL, Lemmon VP, et al. KLF family members regulate intrinsic axon regeneration ability. Science. 2009;326:298–301.
Park KK, Liu K, Hu Y, Smith PD, Wang C, Cai B, et al. Promoting axon regeneration in the adult CNS by modulation of the PTEN/mTOR pathway. Science. 2008;322:963–6.
Zhen CY, Tatavosian R, Huynh TN, Duc HN, Das R, Kokotovic M, et al. Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin. Elife. 2016;5:e17667.
Escamilla-Del-Arenal M, da Rocha ST, Spruijt CG, Masui O, Renaud O, Smits AH, et al. Cdyl, a new partner of the inactive X chromosome and potential reader of H3K27me3 and H3K9me2. Mol Cell Biol. 2013;33:5005–20.
Hur EM, Saijilafu, Lee BD, Kim SJ, Xu WL, Zhou FQ. GSK3 controls axon growth via CLASP-mediated regulation of growth cone microtubules. Genes Dev. 2011;25:1968–81.
Hur EM, Yang IH, Kim DH, Byun J, Saijilafu XuWL, et al. Engineering neuronal growth cones to promote axon regeneration over inhibitory molecules. Proc Natl Acad Sci USA. 2011;108:5057–62.
Saijilafu HurEM, Zhou FQ. Genetic dissection of axon regeneration via in vivo electroporation of adult mouse sensory neurons. Nat Commun. 2011;2:543.
Liu C, Teng ZQ, McQuate AL, Jobe EM, Christ CC, von Hoyningen-Huene SJ, et al. An epigenetic feedback regulatory loop involving microRNA-195 and MBD1 governs neural stem cell differentiation. PLoS ONE. 2013;8:e51436.
Acknowledgements
We acknowledge Dr. Xingguo Li for critical manuscript revise by University of Rochester Medical Center. This work was supported by grants from the National Science Foundation of China (No. 91753140, No. 31571043, and No. 81771224 to C-ML; No. 81571212 to Z-QT), The National Key Research and Development Program of China Project (grant number 2016YFA0101402 to C-ML), and the Hundred Talents Program of Chinese Academy of Sciences to C-ML.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
:The authors declare that they have no conflict of interest.
Additional information
Edited by P Salomoni
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Duan, RS., Tang, GB., Du, HZ. et al. Polycomb protein family member CBX7 regulates intrinsic axon growth and regeneration. Cell Death Differ 25, 1598–1611 (2018). https://doi.org/10.1038/s41418-018-0064-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0064-0
This article is cited by
-
CBX7 silencing promoted liver regeneration by interacting with BMI1 and activating the Nrf2/ARE signaling pathway
Scientific Reports (2024)
-
Cerebellar granule neurons induce Cyclin D1 before the onset of motor symptoms in Huntington’s disease mice
Acta Neuropathologica Communications (2023)
-
Computational analysis of crosstalk between transcriptional regulators and RNA-binding proteins suggests mutual regulation of polycomb proteins and SRSF1 influencing adult hippocampal neurogenesis
Discover Mental Health (2023)
-
St18 specifies globus pallidus projection neuron identity in MGE lineage
Nature Communications (2022)
-
CBX7 binds the E-box to inhibit TWIST-1 function and inhibit tumorigenicity and metastatic potential
Oncogene (2020)