Abstract
Background
Dihydropyrimidine dehydrogenase (DPD) deficiency is the main known cause of life-threatening fluoropyrimidine (FP)-induced toxicities. We conducted a meta-analysis on individual patient data to assess the contribution of deleterious DPYD variants *2A/D949V/*13/HapB3 (recommended by EMA) and clinical factors, for predicting G4-5 toxicity.
Methods
Study eligibility criteria included recruitment of Caucasian patients without DPD-based FP-dose adjustment. Main endpoint was 12-week haematological or digestive G4-5 toxicity. The value of DPYD variants *2A/p.D949V/*13 merged, HapB3, and MIR27A rs895819 was evaluated using multivariable logistic models (AUC).
Results
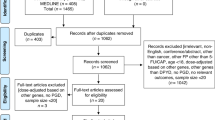
Among 25 eligible studies, complete clinical variables and primary endpoint were available in 15 studies (8733 patients). Twelve-week G4-5 toxicity prevalence was 7.3% (641 events). The clinical model included age, sex, body mass index, schedule of FP-administration, concomitant anticancer drugs. Adding *2A/p.D949V/*13 variants (at least one allele, prevalence 2.2%, OR 9.5 [95%CI 6.7–13.5]) significantly improved the model (p < 0.0001). The addition of HapB3 (prevalence 4.0%, 98.6% heterozygous), in spite of significant association with toxicity (OR 1.8 [95%CI 1.2–2.7]), did not improve the model. MIR27A rs895819 was not associated with toxicity, irrespective of DPYD variants.
Conclusions
FUSAFE meta-analysis highlights the major relevance of DPYD *2A/p.D949V/*13 combined with clinical variables to identify patients at risk of very severe FP-related toxicity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Individual patient data are not available for sharing. Agreement of the investigators from each study would be necessary to transfer such data. Some summary study-level data are available in this paper. Additional study-level summary data corresponding to the analyses mentioned in the paper may be provided on request.
References
Grothey A, Sobrero AF, Shields AF, Yoshino T, Paul J, Taieb J, et al. Duration of adjuvant chemotherapy for Stage III colon cancer. N. Engl J Med. 2018;378:1177–88.
Sharma BB, Rai K, Blunt H, Zhao W, Tosteson TD, Brooks GA. Pathogenic DPYD variants and treatment-related mortality in patients receiving fluoropyrimidine chemotherapy: a systematic review and meta-analysis. Oncologist. 2021;26:1008–16.
Barin-Le Guellec C, Lafay-Chebassier C, Ingrand I, Tournamille J-F, Boudet A, Lanoue M-C, et al. Toxicities associated with chemotherapy regimens containing a fluoropyrimidine: A real-life evaluation in France. Eur J Cancer. 2020;124:37–46.
van Kuilenburg ABP. Dihydropyrimidine dehydrogenase and the efficacy and toxicity of 5-fluorouracil. Eur J Cancer. 2004;40:939–50.
Pallet N, Hamdane S, Garinet S, Blons H, Zaanan A, Paillaud E, et al. A comprehensive population-based study comparing the phenotype and genotype in a pretherapeutic screen of dihydropyrimidine dehydrogenase deficiency. Br J Cancer. 2020;123:811–8.
Etienne MC, Lagrange JL, Dassonville O, Fleming R, Thyss A, Renée N, et al. Population study of dihydropyrimidine dehydrogenase in cancer patients. J Clin Oncol. 1994;12:2248–53.
Loriot M-A, Ciccolini J, Thomas F, Barin-Le-Guellec C, Royer B, Milano G. et al. [Dihydropyrimidine déhydrogenase (DPD) deficiency screening and securing of fluoropyrimidine-based chemotherapies: Update and recommendations of the French GPCO-Unicancer and RNPGx networks]. Bull Cancer. 2018;105:397–407.
Knikman JE, Gelderblom H, Beijnen JH, Cats A, Guchelaar H-J, Henricks LM. Individualized dosing of fluoropyrimidine-based chemotherapy to prevent severe fluoropyrimidine-related toxicity: what are the options? Clin Pharm Ther. 2021;109:591–604.
Amstutz U, Henricks LM, Offer SM, Barbarino J, Schellens JHM, Swen JJ, et al. Clinical pharmacogenetics implementation consortium (CPIC) guideline for dihydropyrimidine dehydrogenase genotype and fluoropyrimidine dosing: 2017 update. Clin Pharm Ther. 2018;103:210–6.
Offer SM, Fossum CC, Wegner NJ, Stuflesser AJ, Butterfield GL, Diasio RB. Comparative functional analysis of DPYD variants of potential clinical relevance to dihydropyrimidine dehydrogenase activity. Cancer Res. 2014;74:2545–54.
Kuilenburg ABPvan, Meijer J, Tanck MWT, Dobritzsch D, Zoetekouw L, Dekkers L-L, et al. Phenotypic and clinical implications of variants in the dihydropyrimidine dehydrogenase gene. Biochimica et Biophysica Acta (BBA) - Mol Basis Dis. 2016;1862:754–62.
fluorouracil-fluorouracil-related-substances-article-31-referral-ema-recommendations-dpd-testing_en.pdf [Internet]. [cité 20 déc 2021]. Available on: https://www.ema.europa.eu/en/documents/referral/fluorouracil-fluorouracil-related-substances-article-31-referral-ema-recommendations-dpd-testing_en.pdf.
Deenen MJ, Meulendijks D, Cats A, Sechterberger MK, Severens JL, Boot H, et al. Upfront Genotyping of DPYD*2A to Individualize Fluoropyrimidine Therapy: A Safety and Cost Analysis. J Clin Oncol. 2016;34:227–34.
Henricks LM, Lunenburg CATC, de Man FM, Meulendijks D, Frederix GWJ, Kienhuis E, et al. DPYD genotype-guided dose individualisation of fluoropyrimidine therapy in patients with cancer: a prospective safety analysis. Lancet Oncol. 2018;19:1459–67.
Tsiachristas A, Vallance G, Koleva-Kolarova R, Taylor H, Solomons L, Rizzo G, et al. Can upfront DPYD extended variant testing reduce toxicity and associated hospital costs of fluoropyrimidine chemotherapy? A propensity score matched analysis of 2022 UK patients. BMC Cancer. 2022;22:458.
Terrazzino S, Cargnin S, Del Re M, Danesi R, Canonico PL, Genazzani AA. DPYD IVS14+1G>A and 2846A>T genotyping for the prediction of severe fluoropyrimidine-related toxicity: a meta-analysis. Pharmacogenomics. 2013;14:1255–72.
Rosmarin D, Palles C, Church D, Domingo E, Jones A, Johnstone E, et al. Genetic markers of toxicity from capecitabine and other fluorouracil-based regimens: investigation in the QUASAR2 study, systematic review, and meta-analysis. J Clin Oncol. 2014;32:1031–9.
Meulendijks D, Henricks LM, Sonke GS, Deenen MJ, Froehlich TK, Amstutz U, et al. Clinical relevance of DPYD variants c.1679T>G, c.1236G>A/HapB3, and c.1601G>A as predictors of severe fluoropyrimidine-associated toxicity: a systematic review and meta-analysis of individual patient data. Lancet Oncol. 2015;16:1639–50.
Lee AM, Shi Q, Alberts SR, Sargent DJ, Sinicrope FA, Berenberg JL, et al. Association between DPYD c.1129-5923 C>G/hapB3 and severe toxicity to 5-fluorouracil-based chemotherapy in stage III colon cancer patients: NCCTG N0147 (Alliance). Pharmacogenet Genomics. 2016;26:133–7.
Meulendijks D, Rozeman EA, Cats A, Sikorska K, Joerger M, Deenen MJ, et al. Pharmacogenetic variants associated with outcome in patients with advanced gastric cancer treated with fluoropyrimidine and platinum-based triplet combinations: a pooled analysis of three prospective studies. Pharmacogenomics J. 2017;17:441–51.
Meulendijks D, Henricks LM, Amstutz U, Froehlich TK, Largiadèr CR, Beijnen JH, et al. Rs895819 in MIR27A improves the predictive value of DPYD variants to identify patients at risk of severe fluoropyrimidine-associated toxicity. Int J Cancer. 2016;138:2752–61.
Amstutz U, Offer SM, Sistonen J, Joerger M, Diasio RB, Largiadèr CR. Polymorphisms in MIR27A associated with early-onset toxicity in fluoropyrimidine-based chemotherapy. Clin Cancer Res. 2015;21:2038–44.
Etienne-Grimaldi M-C, Boyer J-C, Beroud C, Mbatchi L, van Kuilenburg A, Bobin-Dubigeon C, et al. New advances in DPYD genotype and risk of severe toxicity under capecitabine. PLoS One. 2017;12:e0175998.
Schwab M, Zanger UM, Marx C, Schaeffeler E, Klein K, Dippon J, et al. Role of genetic and nongenetic factors for fluorouracil treatment-related severe toxicity: a prospective clinical trial by the German 5-FU Toxicity Study Group. J Clin Oncol. 2008;26:2131–8.
Boige V, Vincent M, Alexandre P, Tejpar S, Landolfi S, Le Malicot K, et al. DPYD genotyping to predict adverse events following treatment with fluorouracil-based adjuvant chemotherapy in patients with stage iii colon cancer: a secondary analysis of the PETACC-8 randomized clinical trial. JAMA Oncol. 2016;2:655–62.
Deenen MJ, Tol J, Burylo AM, Doodeman VD, de Boer A, Vincent A, et al. Relationship between single nucleotide polymorphisms and haplotypes in DPYD and toxicity and efficacy of capecitabine in advanced colorectal cancer. Clin Cancer Res. 2011;17:3455–68.
Garg MB, Lincz LF, Adler K, Scorgie FE, Ackland SP, Sakoff JA. Predicting 5-fluorouracil toxicity in colorectal cancer patients from peripheral blood cell telomere length: a multivariate analysis. Br J Cancer. 2012;107:1525–33.
Boige V, Mendiboure J, Pignon J-P, Loriot M-A, Castaing M, Barrois M, et al. Pharmacogenetic assessment of toxicity and outcome in patients with metastatic colorectal cancer treated with LV5FU2, FOLFOX, and FOLFIRI: FFCD 2000-05. J Clin Oncol. 2010;28:2556–64.
Ducreux M, Adenis A, Pignon J-P, François E, Chauffert B, Ichanté JL, et al. Efficacy and safety of bevacizumab-based combination regimens in patients with previously untreated metastatic colorectal cancer: final results from a randomised phase II study of bevacizumab plus 5-fluorouracil, leucovorin plus irinotecan versus bevacizumab plus capecitabine plus irinotecan (FNCLCC ACCORD 13/0503 study). Eur J Cancer. 2013;49:1236–45.
Gross E, Busse B, Riemenschneider M, Neubauer S, Seck K, Klein H-G, et al. Strong association of a common dihydropyrimidine dehydrogenase gene polymorphism with fluoropyrimidine-related toxicity in cancer patients. PLoS One. 2008;3:e4003.
Loganayagam A, Arenas Hernandez M, Corrigan A, Fairbanks L, Lewis CM, Harper P, et al. Pharmacogenetic variants in the DPYD, TYMS, CDA and MTHFR genes are clinically significant predictors of fluoropyrimidine toxicity. Br J Cancer. 2013;108:2505–15.
Jennings BA, Loke YK, Skinner J, Keane M, Chu GS, Turner R, et al. Evaluating predictive pharmacogenetic signatures of adverse events in colorectal cancer patients treated with fluoropyrimidines. PLoS One. 2013;8:e78053.
Di Paolo A, Danesi R, Falcone A, Cionini L, Vannozzi F, Masi G, et al. Relationship between 5-fluorouracil disposition, toxicity and dihydropyrimidine dehydrogenase activity in cancer patients. Ann Oncol. 2001;12:1301–6.
Wettergren Y, Carlsson G, Odin E, Gustavsson B. Pretherapeutic uracil and dihydrouracil levels of colorectal cancer patients are associated with sex and toxic side effects during adjuvant 5-fluorouracil-based chemotherapy. Cancer. 2012;118:2935–43.
Budai B, Komlósi V, Adleff V, Pap É, Réti A, Nagy T, et al. Impact of SHMT1 polymorphism on the clinical outcome of patients with metastatic colorectal cancer treated with first-line FOLFIRI+bevacizumab. Pharmacogenet Genomics. 2012;22:69–72.
Wagner AD, Grothey A, Andre T, Dixon JG, Wolmark N, Haller DG, et al. Sex and Adverse Events of Adjuvant Chemotherapy in Colon Cancer: An Analysis of 34 640 Patients in the ACCENT Database. J Natl Cancer Inst. 2021;113:400–7.
Milano G, Etienne MC, Cassuto-Viguier E, Thyss A, Santini J, Frenay M, et al. Influence of sex and age on fluorouracil clearance. J Clin Oncol. 1992;10:1171–5.
Meulendijks D, van Hasselt JGC, Huitema ADR, van Tinteren H, Deenen MJ, Beijnen JH, et al. Renal function, body surface area, and age are associated with risk of early-onset fluoropyrimidine-associated toxicity in patients treated with capecitabine-based anticancer regimens in daily clinical care. Eur J Cancer. 2016;54:120–30.
Breton C, Aparicio T, Le Malicot K, Ducreux M, Lecomte T, Bachet J-B, et al. Predictive factors of severe early treatment-related toxicity in patients receiving first-line treatment for metastatic colorectal cancer: Pooled analysis of 2190 patients enrolled in Fédération Francophone de Cancérologie Digestive (FFCD) trials. Eur J Cancer. 2021;153:40–50.
Nie Q, Shrestha S, Tapper EE, Trogstad-Isaacson CS, Bouchonville KJ, Lee AM, et al. Quantitative Contribution of rs75017182 to Dihydropyrimidine Dehydrogenase mRNA Splicing and Enzyme Activity. Clin Pharm Ther. 2017;102:662–70.
Lunenburg CATC, van der Wouden CH, Nijenhuis M, Crommentuijn-van Rhenen MH, de Boer-Veger NJ, Buunk AM, et al. Dutch Pharmacogenetics Working Group (DPWG) guideline for the gene-drug interaction of DPYD and fluoropyrimidines. Eur J Hum Genet. 2020;28:508–17.
Knikman JE, Wilting TA, Lopez-Yurda M, Henricks LM, Lunenburg CATC, de Man FM, et al. Survival of Patients With Cancer With DPYD Variant Alleles and Dose-Individualized Fluoropyrimidine Therapy-A Matched-Pair Analysis. J Clin Oncol. 2023;41:5411–21.
Henricks LM, Lunenburg CATC, de Man FM, Meulendijks D, Frederix GWJ, Kienhuis E, et al. A cost analysis of upfront DPYD genotype-guided dose individualisation in fluoropyrimidine-based anticancer therapy. Eur J Cancer. 2019;107:60–7.
Baker SD, Bates SE, Brooks GA, Dahut WL, Diasio RB, El-Deiry WS, et al. DPYD Testing: Time to Put Patient Safety First. J Clin Oncol. 2023;41:2701–5.
Hertz DL. Assessment of the Clinical Utility of Pretreatment DPYD Testing for Patients Receiving Fluoropyrimidine Chemotherapy. J Clin Oncol. 2022;40:3882–92.
White C, Scott RJ, Paul C, Ziolkowski A, Mossman D, Ackland S. Ethnic Diversity of DPD Activity and the DPYD Gene: Review of the Literature. Pharmgenomics Pers Med. 2021;14:1603–17.
de With M, Knikman J, de Man FM, Lunenburg CATC, Henricks LM, van Kuilenburg ABP, et al. Dihydropyrimidine dehydrogenase phenotyping using pretreatment uracil: a note of caution based on a large prospective clinical study. Clin Pharm Ther. 2022;112:62–8.
Coenen MJH, Paulussen ADC, Breuer M, Lindhout M, Tserpelis DCJ, Steyls A, et al. Evolution of Dihydropyrimidine Dehydrogenase Diagnostic Testing in a Single Center during an 8-Year Period of Time. Curr Ther Res Clin Exp. 2019;90:1–7.
De Mattia E, Silvestri M, Polesel J, Ecca F, Mezzalira S, Scarabel L, et al. Rare genetic variant burden in DPYD predicts severe fluoropyrimidine-related toxicity risk. Biomed Pharmacother. 2022;154:113644.
Pellicer M, García-González X, García MI, Blanco C, García-Alfonso P, Robles L, et al. Use of exome sequencing to determine the full profile of genetic variants in the fluoropyrimidine pathway in colorectal cancer patients affected by severe toxicity. Pharmacogenomics. 2017;18:1215–23.
Hamzic S, Kummer D, Froehlich TK, Joerger M, Aebi S, Palles C, et al. Evaluating the role of ENOSF1 and TYMS variants as predictors in fluoropyrimidine-related toxicities: An IPD meta-analysis. Pharm Res. 2020;152:104594.
Traylor M, Walker JL, Corrigan AA, Hernandez MA, Newhouse SJ, Folarin AA, et al. Exome array analysis of adverse reactions to fluoropyrimidine-based therapy for gastrointestinal cancer. PLoS One. 2018;13:e0188911.
Acknowledgements
The authors would like to thank the Gustave Roussy library team (current head: Alexia Nerfié) for its support in the literature search. This study was supported by the French Ministry of Health (PHRC-K 14-193 FUSAFE) and the French « Ligue Nationale Contre le Cancer ». The funders of the study had no role in study design, data collection, data analysis, data interpretation, or writing of the report.
Author information
Authors and Affiliations
Consortia
Contributions
GLT, JPP, JCB, MCEG, NC and VB with the help of the steering committee members designed and supervised the study. MCEG, and JPP obtained funding. JPP, MCEG, JCB and the DPD working group (cf. appendix 2) searched and selected the studies. Steering committee members contributed to the identification and selection of the studies. JPP, NC, MCEG and VB collected and checked data with the help of the investigators who validated the re-analysis of their trials. GLT, JPP, and NC did the statistical analyses. GLT, JCB, JPP, MCEG, NC and VB wrote the draft, with revisions from the other investigators. All authors contributed to the interpretation of the results during the investigator meeting and the revision of the manuscript. All investigators listed in Web-Appendix 1 received the manuscript for revision.
Corresponding author
Ethics declarations
Competing interests
GM reports consulting fees from Servier. JHM Schellens reports patents on oral taxanes. MS reports payment and honoraria for lectures for CED Service GmbH. VB reports consulting fees from Merck, Serono, Bayer, Roche, Eisai and BMS. Q Shi reports grants from Janssen, BMS, Genetech, Novartis, Celgene and fees from Regeneron Pharmaceuticals and Chugai Pharmaceuticals. MC Etienne-Grimaldi reports honoraria for lectures for AMGEN. All other authors report no conflicts of interest.
Ethics approval
Included clinical studies were conducted in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Le Teuff, G., Cozic, N., Boyer, JC. et al. Dihydropyrimidine dehydrogenase gene variants for predicting grade 4-5 fluoropyrimidine-induced toxicity: FUSAFE individual patient data meta-analysis. Br J Cancer 130, 808–818 (2024). https://doi.org/10.1038/s41416-023-02517-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02517-2