Abstract
Background
Resistance to androgen receptor signalling inhibitors (ARSIs) represents a major clinical challenge in prostate cancer. We previously demonstrated that the ARSI enzalutamide inhibits only a subset of all AR-regulated genes, and hypothesise that the unaffected gene networks represent potential targets for therapeutic intervention. This study identified the hyaluronan-mediated motility receptor (HMMR) as a survival factor in prostate cancer and investigated its potential as a co-target for overcoming resistance to ARSIs.
Methods
RNA-seq, RT-qPCR and Western Blot were used to evaluate the regulation of HMMR by AR and ARSIs. HMMR inhibition was achieved via siRNA knockdown or pharmacological inhibition using 4-methylumbelliferone (4-MU) in prostate cancer cell lines, a mouse xenograft model and patient-derived explants (PDEs).
Results
HMMR was an AR-regulated factor that was unaffected by ARSIs. Genetic (siRNA) or pharmacological (4-MU) inhibition of HMMR significantly suppressed growth and induced apoptosis in hormone-sensitive and enzalutamide-resistant models of prostate cancer. Mechanistically, 4-MU inhibited AR nuclear translocation, AR protein expression and subsequent downstream AR signalling. 4-MU enhanced the growth-suppressive effects of 3 different ARSIs in vitro and, in combination with enzalutamide, restricted proliferation of prostate cancer cells in vivo and in PDEs.
Conclusion
Co-targeting HMMR and AR represents an effective strategy for improving response to ARSIs.

Similar content being viewed by others
Introduction
Prostate cancer (PCa) is the most frequently diagnosed cancer and the second leading cause of cancer-related death in men in the developed world [1]. A hallmark of PCa is its dependence on androgen signalling, hence the androgen receptor (AR) is the primary therapeutic target for advanced disease [2, 3]. While inhibition of AR signalling by androgen deprivation therapy (ADT) has been the mainstay of PCa treatment for almost 80 years, the disease invariably progresses to a lethal form known as castration-resistant prostate cancer (CRPC). A key feature of CRPC is the re-activation of AR signalling, which has underpinned the development of second-generation AR signalling inhibitors (ARSIs) [4]. There are four second-generation ARSIs currently approved by the Food and Drug Administration (FDA); the androgen biosynthesis inhibitor abiraterone acetate [5], and the AR antagonists enzalutamide [6, 7], apalutamide [8] and darolutamide [9]. While ARSI use improves survival, resistance to these agents has become one of the most important challenges in the clinical management of PCa. Intrinsic resistance to enzalutamide, for example, is observed in up to 50% of patients treated, and those who initially respond will generally acquire resistance within a period of months [10]. Mechanistically, resistance to ARSIs is largely attributed to aberrations in AR, such as mutations, amplification and splicing variants [11], highlighting the continued dependence of CRPC on AR signalling.
For a decade, combination approaches to target AR signalling in PCa have been postulated as essential to achieve long-term disease control [12], but combination strategies have still not been optimised. There are a multitude of PCa clinical trials investigating combinations of approved agents, including ADT, ARSIs, radiotherapy, chemotherapy, immunotherapy and small molecular inhibitors. Indeed, triplet therapy that combines ADT, chemotherapy and an ARSI may soon become the new gold standard for hormone-sensitive metastatic PCa based on recent trial results [13]. While this and other combination strategies have been successful in extending survival, they have largely been based on existing and available agents. The aim of our study was to rationally design a novel combination treatment strategy for PCa that more effectively inhibits AR signalling and PCa cell survival. Previously we reported that the AR antagonist enzalutamide does not effectively target the entire repertoire of genes regulated by the AR in PCa cells, and presented evidence that those remaining networks provide fuel for survival [14]. Herein, we investigate the function of one such gene, hyaluronan-mediated motility receptor (HMMR), and the therapeutic potential of targeting HMMR to sensitise tumour cells to enzalutamide.
Materials and Methods
Cells lines and reagents
VCaP, LNCaP, 22Rv1 and PC3 human prostate carcinoma cells were purchased from the American Type Culture Collection (ATCC, MD, USA). LNCaP-derived V16D (castration-resistant, enzalutamide-sensitive) and MR49F (castration-resistant, enzalutamide-resistant) human PCa cells were a kind gift from Prof. Amina Zoubeidi (Vancouver Prostate Centre, Vancouver, Canada) [15]. All cell lines were verified using short tandem repeat profiling in 2022 by ATCC or CellBank Australia. LNCaP, 22RV1, MR49F and V16D cells were maintained in RPMI-1640 supplemented with 10% FBS; the media for growth of MR49F cells was additionally supplemented with 10 µM enzalutamide. VCaP cells were maintained in Dulbecco’s Modified Eagle’s Medium containing 10% FBS, 1% sodium pyruvate, 1% MEM non–essential amino acids, and 0.1 nM 5α-dihydrotestosterone (DHT).
All chemicals, reagents and 4-methlumbelliferone were purchased from Sigma Aldrich (St Louis, MO, USA). Enzalutamide was obtained from Selleck Chemicals (Houston, TX, USA); apalutamide (ARN-509), darolutamide (ODM-201) from Sapphire Bioscience (Redfern, NSW, AUS) and Hyaluronic acid from Contipro (Dolní Dobrouč, Czech Republic). All drugs were dissolved in DMSO except for HA which was dissolved in RNA grade double distilled water.
Transient transfection
HMMR knockdown was carried out using Dharmacon ON-Target HMMR (3161) SMARTpool 5 nmol siRNA cat#: L-010409-00-0005 and control cells were transfected with Dharmacon ON-Target Plus siRNA cat#: D-001810-01-20 siRNA at a concentration of 12.5 nM for 6 well plates (5 × 105 cells), 5 nM for 24 and 96 well plates (3.5 × 104 cells and 3.5 × 103 cells respectively). The cells were reverse transfected using lipofectamine RNAiMAX (Life technologies, ThermoFisher Scientific, Scornsby, VIC, AUS) according to the manufacturer’s protocol. AR knockdown was achieved as previously described [14]. The following AR siRNAs were used: AR (Silencer Select #4390824/5; s1538 (siAR2), s1539 (siAR1) (Ambion; ThermoFisher Scientific), and Negative Control 2 #AM4637 (Ambion; ThermoFisher Scientific).
Cell viability and apoptosis assays
Cells were seeded in triplicate in 24 well plates at densities of 3.5 × 104 for LNCaP, 3 × 104 for MR49F and 2.5 × 104 for V16D. Cell viability was determined by manual counting using Trypan blue exclusion, as described previously [16]. Alternatively, cell viability was assessed by CyQuantTM Cell Proliferation Assay (ThermoFisher Scientific), according to the manufacturer’s instructions.
The effect of siRNA or drug treatment on apoptosis was carried out by seeding 5 × 104 cells/well in triplicate. Cells were collected three days post-treatment or knockdown and stained with Annexin V PE (BD PharmagenTM, CA, USA) and 1 mM 7-Aminoactiomycin D (Thermo Fisher Scientific) and analysed using a BD LSRFortessa X20 Flow Cytometer.
Quantification of combination index
The combination index (CI) was determined using Compusyn software (Compusyn Inc. Paramus, NJ, USA) based on the Chau-Talalay theorem [17]. The quantitative value of the CI is defined as follows: CI = 1: additive effect, CI > 1: antagonism, CI < 1: synergy.
Colony formation assay
Cells were seeded at 500 cells/well in 2 mL RPMI + 10% FBS and incubated at 37 °C for 2 weeks with the indicated treatments, fixed with 4% paraformaldehyde and stained with 1% crystal violet to identify colonies. Colony-forming efficiency was determined by counting colonies greater than 50 cells for each well.
Quantitative real-time polymerase chain reaction (RT-qPCR)
RNA (1 µg) was reverse transcribed using iScript cDNA synthesis kit (Bio-Rad, CA, USA), according to the manufacturer’s instructions. RT-qPCR was performed on 1:10 diluted cDNA using SYBR green and CFX384 Real-Time System (Bio-Rad) for 40 cycles. Gene expressions are presented relative to GUSB and L19 as determined by GeNorm [18]. All primers were purchased from Sigma Aldrich with primer sequences listed in Supplementary Table 1.
Immunoblotting
Whole-cell lysates were collected in RIPA lysis buffer supplemented with cOmplete ULTRA protease and phosphatase inhibitor (Cell Signalling Technology (CST), Danvers, MA, USA) and immunoblotting was performed as previously described [19]. Primary and secondary antibodies used in this study are enlisted in Supplementary Table 2.
Visualisation of pCMV-tagged AR protein
PC3 cells at a density of 4 × 104 cells/well were reverse transfected with 1 µg pCMV-AR in an 8-well Lab-Tek II chamber slide (ThermoFisher Scientific) using lipofectamine 2000 (ThermoFisher Scientific) according to the manufacturer’s protocol and allowed to adhere for 4 h. An overlay of drug 4-MU or docetaxel was added and incubated for 24 h. Cells were then treated with 1 nM DHT or vehicle for 4 h and fixed with 4% paraformaldehyde. Fixed cells were washed in PBS and incubated with an antibody against AR (AR-N20) overnight. AR was visualised using Alexa-Fluor 488 goat anti-rabbit IgG (ThermoFisher Scientific). Images were obtained using Leica TCS SP8X/MP confocal microscope (Leica Microsystem, Wetzler, Germany). Nucleoplasmic translocation was quantified using FIJI software (ImageJ: http://fiji.sc/Fiji; version 1.52 p) [20].
Subcellular fractionation of prostate cancer cells
LNCaP cells at 80% confluency were androgen starved for three days and treated with PRF-RPMI containing vehicle, 0.4 mM 4-MU for 24 h followed by 10 nM DHT or vehicle treatment for 4 h. Cells were trypsinised, 200 µL was collected as whole cell lysate and the remaining cell suspension centrifuged at 1500 rpm for 5 mins to pellet the cells. The cell pellet was washed in PBS and lysed in 500 µL lysis buffer 1 (10 mM Tris HCl (pH 7.9), 0.34 M sucrose, 3 mM CaCl2, 2 mM magnesium acetate, 0.1 mM EDTA, 1 mM DTT, 0.5% nonidet P-40 substitute) containing cOmplete ULTRA protease (CST, Danvers, MA, USA) and centrifuged at 3500 x g for 5 min. The supernatant was clarified by spinning at 20,000 x g for 15 mins and saved as the cytoplasmic fraction. The cell pellet was then washed with 300 µl lysis buffer 1 and pelleted by centrifugation at 3500 x g for 5 mins. The cell pellet was resuspended in 225 µl lysis buffer 2 (20 mM HEPES (pH 7.9), 3 mM EDTA, 10% glycerol, 150 mM potassium acetate, 1.5 mM MgCl2, 1 mM DTT, 0.1% Nonident P-40 substitute) containing cOmplete ULTRA protease and phosphatase inhibitor (CST, Danvers, MA, USA) and centrifuged at 15000 x g for 30 mins. The supernatant was saved as the nuclear fraction. The cell pellet was resuspended in 300 µl PBS and sonicated by BioRuptor sonication system (Diagenode, Belgium) at high intensity 4 × 30 s with 1 min rest in between. The sonicated solution was used as the chromatin fraction. Equivalent volumes of cytoplasmic and nuclear plus chromatin fractions were immunoblotted for AR(AR-N20). Cytoplasmic marker GAPDH and nuclear marker H3 were used as sub-fractionation markers.
In vivo studies
All animal experiments were approved by The University of Adelaide and Nagoya University (ethics approval numbers: M-2020-014 and 31460 respectively) ethics committees. Balb/cSlc (nu/nu) 6 weeks old male nude mice were subcutaneously injected with 1:1 PBS/matrigel suspension containing 2 × 106 V16D cells. Once tumours became palpable, mice were assigned randomly to one of the 4 groups (Vehicle, 225 mg/kg 4-MU, 10 mg/kg ENZ and 225 mg/kg 4-MU + 10 mg/kg ENZ). For 5 weeks, mice were fed with a powdered diet supplemented with vehicle or 4-MU, ENZ or 4-MU + ENZ. Tumour dimensions were measured every second-day using callipers and volumes were calculated using the formula (width2 x length x 0.5236). Mice were culled when tumour volume reached 1000 mm3 or at the end of 5 weeks of treatment. Tumours were resected and formalin-fixed for immunohistochemistry (IHC) analysis and plasma samples were collected at sacrifice.
Ex vivo studies
PCa specimens were obtained with informed written consent through St Andrew’s Hospital (Adelaide, Australia) or the Nagoya University Hospital (Nagoya, Japan) from men who underwent radical prostatectomy. Experiments were approved by The University of Adelaide (approval: H-2012-016) and Nagoya University (approval: 2020-01-17) human research ethics committees. Patient-derived explants (PDEs) were generated from 1–2 mm3 tissues cultured on gelatin sponges pre-soaked in media as described previously [21]. PDEs were treated with DMSO or 4-MU alone or in combination with enzalutamide, as indicated for each experiment for 48 h. Tissues were formalin-fixed and paraffin embedded for analysis by IHC. AR knockdown in PDEs was performed as previously described [22].
Immunohistochemistry (IHC)
Tissue sections (2 µm) on Superfrost plus slides were dewaxed and quenched for peroxidase activity using 3% H2O2 for 15 mins. Target antigens were stained using antibodies detailed in Supplementary Table 2. For animal studies, whole slide images of Ki67 stained sections were obtained using the visual slide system VS120 (Olympus, Tokyo, Japan) and images were quantified using semi-automated TissuemorphDPTM software (Visiopharm, Hørsholm, Denmark), according to the manufacturer’s instructions. For patient-derived explants, Ki67 stained slides were imaged and quantified in a blinded manner, as previously described [21].
Statistical analysis
Statistical significance was determined using GraphPad Prism version 9 with Student’s t-test, one-way or two-way ANOVA (with Tukey or Dunnett’s post hoc tests), as specified in the figure legends. A p-value ≤ 0.05 was considered statistically significant.
Results
Discovery of HMMR as a potential pathway to ARSI resistance
Using an unbiased transcriptomics approach previously reported by our team [14], HMMR was identified as one of the most differentially regulated genes by AR knockdown that remains unaffected by enzalutamide treatment (Fig. 1a). This differential regulation of HMMR was validated in LNCaP (Fig. 1b, c), VCaP (Supplementary Figure 1a, b) and V16D (Supplementary Figure 1c, d) PCa cells, where HMMR gene and protein expression were unaffected by enzalutamide compared to vehicle treatment, but significantly downregulated by two independent siRNA targeted to AR (siAR) compared to scrambled control siRNA (siCON). In contrast, the canonical AR-regulated gene KLK3, was significantly downregulated by both enzalutamide treatment and siAR treatment (Fig. 1b, Supplementary Figure 1a, Supplementary Figure 1c). In support of our findings, stable AR knockdown in PCa cells downregulated HMMR in an independent RNA-seq dataset (Supplementary Figure 1e) [23], and in patient-derived explants (PDEs), AR knockdown by siAR-loaded nanoparticles [22] reduced both AR and HMMR mRNA expression compared to control (Supplementary figure 1f). Similarly, the SU2C dataset confirmed our finding that pharmacological inhibition of AR using ARSIs does not inhibit HMMR gene expression, as HMMR remained unchanged by ARSI treatment in CRPC tumours compared to untreated CRPC (Supplementary Figure 1g) [24]. Given the increasing use of ARSIs in combination with ADT clinically [25], we evaluated HMMR expression in response to enzalutamide under androgen-deprived conditions. To simulate androgen deprivation in vitro, androgen-sensitive LNCaP and VCAP cells and androgen-independent V16D cells were androgen-starved for three days. Upon the addition of enzalutamide, HMMR expression remained unchanged compared to vehicle treatment, whereas KLK3 expression was significantly reduced in the presence of enzalutamide in all three cell lines (Supplementary Figure 1h–j).
a Volcano plot of differentially expressed genes affected by either siAR versus siCON (red; n = 599 genes), or Veh versus ENZ (black; n = 23 genes). b Validation of HMMR expression in response to siAR or 1 µM ENZ by RT-qPCR. Gene expression was normalised to GUSB and L19. Data is presented as mean ± SD of 3 biological replicates and are representative of 3 independent experiments. Controls (Veh and siCON) were set to 1 and data statistically evaluated using two-way ANOVA with Tukey’s multiple comparison test (treatment vs control; ****p < 0.0001). c AR and HMMR protein expression in response to siAR or 1 µM ENZ by Western Blot. GAPDH was used as a loading control.KLK3, KLK2, FKBP5 and TMPRSS2 expression in response to treatment with increasing dose of ARSIs apalutamide (d) or darolutamide (e) for 24 h, by RT-qPCR. Gene expression was normalised to GUSB and L19. Data are presented as mean ± SD of 3 biological replicates and are representative of three independent experiments. Control (Veh) was set to 1 and data was statistically evaluated using one-way ANOVA with Dunnett’s multiple comparison test. (*p < 0.05, **p < 0.01, ****p < 0.0001). AR, PSA and HMMR protein expression in response to treatment with ARSIs apalutamide (f) or darolutamide (g) for 24 h, by Western Blot. Numerals above each lane represent densitometric analysis of each protein relative to loading controls GAPDH (f) or β-Actin (g).
Having demonstrated that enzalutamide fails to modulate HMMR expression, in the presence or absence of androgens, we tested whether this was specific to this particular ARSI or a broader feature of second-generation AR antagonists. LNCaP cells were treated with increasing dose of clinical agents apalutamide or darolutamide and HMMR expression was measured by RT-qPCR and Western Blot. No significant change in HMMR gene (Fig. 1d, e) or protein (Fig. 1f, g) levels were observed upon either ARSI treatment compared to vehicle-treated cells. In contrast, the expression of androgen-regulated genes KLK3 (PSA), KLK2, FKBP5 and TMPRSS2, and PSA protein expression were potently and dose-dependently supressed by both agents (Fig. 1d–g). Collectively our results provide evidence that despite being inhibited by AR knockdown, HMMR evades pharmacological inhibition of AR with second-generation ARSIs.
HMMR expression is associated with advanced disease, treatment resistance and poor prognosis
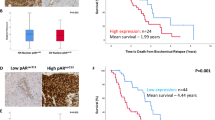
To determine whether HMMR has clinical relevance in PCa, its expression was interrogated in multiple transcriptomic and proteomic datasets. In clinical transcriptomic data from The Cancer Genome Atlas (TCGA) [26], HMMR expression was positively correlated with an established AR-regulated gene signature (Fig. 2a), and with acquired ADT resistance (Fig. 2b). This data supports that HMMR is AR-regulated in PCa [27], and that HMMR expression correlates with resistance to androgen deprivation and biochemical recurrence [28]. Increased HMMR mRNA expression has been reported in metastatic PCa [29] and CRPC [28] compared to localised disease. We demonstrate the association with metastatic disease is also observed at the protein level, with HMMR overexpressed in metastatic CRPC tumours compared to primary tumours in proteomic data obtained by mass spectrometry [30] (Fig. 2c). Importantly, survival analysis of three independent PCa patient cohorts revealed that high HMMR expression is associated with poorer progression-free (Fig. 2d), disease-free (Fig. 2e), and overall survival (Fig. 2f) [26, 31, 32]. In summary, HMMR expression is increased in advanced stages of PCa and is associated with treatment resistance and poorer prognosis.
Association between HMMR expression levels with an AR signalling signature (a) or acquired ADT resistance signature (b) from TCGA primary prostate cancer expression data. Scatter plot of the Spearman correlation analysis between HMMR expression and signature genes, where each symbol represents one patient. c Violin plot of HMMR protein expression in benign prostate tissues versus primary and metastatic castrate-resistant prostate cancer tissues. Plots show minimum and maximum values (lower and upper lines respectively). Data were analysed using one-way ANOVA with Tukey’s multiple comparison test (**p < 0.01). d–f HMMR mRNA expression is associated with shorter relapse-free survival in the TCGA (primary prostate cancer) and MSKCC (primary and metastatic CRPC) cohorts, and shorter overall survival rates in the SU2C (metastatic CRPC) cohort.
Targeting HMMR inhibits cell viability and induces cell death of hormone-sensitive and enzalutamide-resistant PCa models
Having established that HMMR is not modulated by ARSIs, we investigated its potential as a therapeutic target. Transient downregulation of HMMR using SMARTpool siRNA (siHMMR) markedly inhibited cell proliferation (Fig. 3a, b) and induced apoptosis (Fig. 3c) compared to control siRNA (siCON) in enzalutamide-sensitive LNCaP and enzalutamide-resistant MR49F cells. Additionally, siHMMR suppressed the colony-forming abilities of both LNCaP (Fig. 3d) and MR49F (Fig. 3e) cells, indicative of reduced survival ability. The efficiency of HMMR knockdown with the SMARTpool siRNA was confirmed at gene and protein levels by RT-qPCR and Western Blotting, respectively (Supplementary Figure 2a–c). Having demonstrated that HMMR knockdown regulates PCa cell growth and survival, we sought to inhibit HMMR using a pharmacological agent. While there are no direct inhibitors of HMMR, HA is a well-characterised ligand for HMMR [33], and the HA synthesis inhibitor, 4-methylumbeliferone (4-MU) has previously been reported to suppress HMMR expression [34]. We likewise found that 4-MU suppressed HMMR expression in a range of PCa cell lines (Supplementary Figure 2d–f). 4-MU dose-dependently reduced cell viability (Fig. 3f) and colony formation (Fig. 3g) compared to vehicle treatment in androgen-sensitive LNCaP cells. Importantly, 4-MU was also potent against models of CRPC, including 22Rv1 (Supplementary Figure 2g) and the enzalutamide-sensitive V16D and enzalutamide-resistant MR49F lines (Fig. 3h). In enzalutamide-resistant MR49F cells, 4-MU markedly induced apoptosis, as determined by flow cytometry using 7AAD/Annexin V-PE staining (Fig. 3i). To ensure that the growth inhibitory effects of 4-MU in this study were mediated through HA, we undertook a rescue experiment by co-treating PCa cells with 4-MU and low molecular weight HA (27 kDa). The addition of HA partially rescued the cell proliferative effects of 4-MU in V16D cells (Supplementary Figure 2h), supporting the on-target efficacy of the inhibitor. Encouraged by the therapeutic efficacy of 4-MU in PCa cell lines, we expanded our investigation into clinical prostate tumours using a PDE model [35]. Treatment of PCa PDE tissues with 4-MU for 48 h resulted in marked suppression of the proliferative marker Ki67, from a mean of 26.2% positively stained epithelial cells in control-treated PDE tissues to 5.4% and 1.5% in PDE tissues treated with 0.5 mM and 1 mM 4-MU, respectively (Fig. 3j). This finding was validated in an independent cohort of PCa PDEs, where 0.5 mM and 1 mM 4-MU again significantly reduced Ki67 positivity compared to the control (Supplementary Figure 2l). In summary, HMMR expression is critical for PCa cell growth and survival and represents a potential therapeutic target.
Knockdown of HMMR with sMARTpool siRNA (siHMMR) reduced proliferation of LNCaP (a) and MR49F (b) cells, as determined by CyQuant assay. Cell proliferation was evaluated by measure of fluorescence intensity on days 3-, 4-, 5- and 6 post-transfection. Data are presented as mean ± SD of 5 wells and represent three independent experiments. Data were statistically evaluated using two-way ANOVA with Tukey’s multiple comparison test (****p < 0.0001). c HMMR knockdown induced apoptosis of LNCaP and MR49F cells, as determined using flow cytometry-based 7-AAD/Annexin V assays at 3 days post-transfection. Data are presented as mean ± SD of 3 biological replicates and represent three independent experiments. Dead cell proportions upon HMMR knockdown were compared to the vehicle using ANOVA and Dunnett’s multiple comparison tests (ap<0.0001, bp < 0.001, cp < 0.01). HMMR knockdown with siHMMR inhibits colony formation of LNCaP (d) and MR49F (e) cells. Formalin-fixed cells were stained with 1% crystal violet then colonies containing >50 cells were manually counted. Data are presented as mean ± SD of 3 wells and represent three independent experiments. Data were statistically analysed using unpaired student’s T-test (**p < 0.01, ****p < 0.0001). (f) Pharmacological inhibition of HMMR with 4-MU inhibits LNCaP cell viability. Viable and dead cells were determined by Trypan blue exclusion assay after 3- and 6- days of treatment. Data are presented as mean ± SD of triplicate wells, are representative of two independent experiments, and were analysed using one-way ANOVA with Tukey’s multiple comparison test (*p < 0.05, ****p < 0.0001). g 4-MU dose-dependently suppresses colony formation in LNCaP cells. Data are presented as mean ± SD of 3 wells, are representative of two independent experiments, and were analysed using one-way ANOVA with Dunnett’s test. (**p < 0.01, ***p < 0.001, ***p < 0.0001). h 4-MU dose-dependently suppresses cell proliferation in V16D and MR49F PCa cells. Cells were counted using the Trypan blue dye exclusion method after 3- and 6- days of treatment. Data are presented as mean ± SD of triplicate wells and represent two independent experiments. Statistical analysis was performed using one-way ANOVA with Turkey’s multiple comparison test. (****p < 0.0001). i 4-MU treatment dose-dependently induces apoptosis in MR49F cells three days post-treatment. Data are presented as mean ± SD of 3 wells, represent two independent experiments and were analysed by two-way ANOVA with Turkey’s multiple comparison test (ap < 0.0001, cp < 0.01, dp < 0.05. j 4-MU inhibits proliferation in patient-derived prostate cancer explants (PDEs). PDEs (n = 7) were treated for 48 h with an increasing dose of 4-MU, then paraffin-embedded and formalin-fixed prior to immunohistochemistry (IHC) with proliferation marker Ki67. Digital images were manually counted. Data represents mean ± SEM and were statistically analysed using one-way ANOVA with Dunnett’s test (*p < 0.05, **p < 0.01). Quantification of Ki67 staining on the left and representative IHC images on the right.
Targeting HMMR inhibits AR nuclear localisation and transcriptional activity
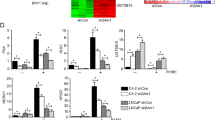
HMMR is a protein with dual functions: extracellularly, HMMR is a receptor for hyaluronic acid; intracellularly, HMMR associates with microtubules to regulate protein trafficking and spindle assembly prior to mitosis [36]. In PCa, the microtubule network is particularly significant as it directly facilitates AR nuclear translocation and transcriptional activation of downstream AR signalling [37, 38]. In fact, the reliance of AR on microtubules is attributed to the success of microtubule targeting agent docetaxel in PCa, which is the first-line treatment and standard-of-care for metastatic CRPC [39, 40]. To determine whether the microtubule aspect of HMMR underlies its role in PCa cell growth and survival, we investigated whether HMMR inhibition alters the cellular localisation of AR. Exogenous wild-type AR was transiently transfected into AR-negative PC3 PCa cells and treated with vehicle control or DHT in the presence or absence of 4-MU. As expected, AR was predominantly localised in the nucleus upon DHT treatment (83.19%) compared to control (51.74%) (Fig. 4a). Treatment with 4-MU alone (56.11%) did not induce AR localisation, and the presence of 4-MU prevented the nuclear import of AR upon DHT treatment (57.56%) (Fig. 4a), to a similar extent as the microtubule inhibitor docetaxel in the presence of DHT (66.63%) (Fig. 4a). We further showed by subcellular fractionation that co-treatment with 4-MU and DHT reduces nuclear localisation of AR in LNCaP cells compared to DHT treatment alone (Fig. 4b, Supplementary Figure 3a). We next evaluated PSA expression in LNCaP cells as a downstream readout of AR nuclear transcriptional activity, and found that 4-MU reduced DHT-mediated induction of PSA at both the gene and protein levels (Fig. 4c). Interestingly, we observed a decrease in AR steady-state protein levels with 4-MU treatment alone, which was stabilised by addition of DHT (Fig. 4c). In LNCaP, V16D and MR49F cells cultured in growth medium containing full serum, to represent physiological levels of androgens, 4-MU dose-dependently reduced expression of PSA at both the gene and protein levels (Fig. 4d–f). The effect of 4-MU on AR protein expression was again observed and most prominently in V16D Cells (Fig. 4d–f). To validate the effects of HMMR inhibition by 4-MU on AR signalling, we examined AR and PSA levels upon HMMR knockdown. Depletion of HMMR with siRNA reduced AR protein levels by up to 50% (Fig. 4g), and PSA gene and protein levels were subsequently decreased, in both LNCaP and MR49F cells (Fig. 4g–i). Collectively, these findings reveal that targeting HMMR results in the inhibition of AR signalling, mediated through decreased AR nuclear translocation and expression.
a Decreased nuclear localisation of pCMV-tagged wtAR (green) in transfected PC3 cells treated with 4-MU or docetaxel (DTX) ± dihydrotestosterone (DHT), captured by fluorescence microscopy. Nuclei were visualised with DAPI mounting media (blue). Data are presented as the mean ± SD of 2 wells and represent two independent experiments. Data were analysed using two-way ANOVA with Tukey’s multiple comparison test. (***p < 0.001, ****p < 0.0001). The scale bar represents 10 pixels/micron. b Western Blot of AR protein expression in LNCaP nuclear fraction upon treatment with vehicle, 10 nM DHT, 0.4 mM 4-MU or DHT + 4-MU. Acetylated Histone H3 was used as a nuclear marker. Numerals above each lane represent densitometric analysis of AR relative to H3. c LNCaP cells were androgen deprived for 3 days prior to treatment with DHT ± 4-MU. Cells were analysed by RT-qPCR (upper panel) or Western Blot (lower panel) for KLK3 expression. Genes were normalised to GUSB and L19. Loading control for Western Blot was GAPDH. PCa cells grown in the presence of androgens were treated with increasing doses of 4-MU. Cells were analysed by RT-qPCR (upper panel) or Western Blot (lower panel) for KLK3 expression in LNCaP (d), V16D (e) and MR49F (f) cells. Genes were normalised to GUSB and L19. Loading control for Western Blots was β-Actin. Numerals above each lane represent densitometric analysis of each protein relative to β-Actin. Data in c–f are presented as mean ± SD of 3 wells and analysed using one-way ANOVA with Dunnett’s test (***p < 0.001, ****p < 0.0001). g HMMR, AR and PSA protein expression after 72 h of siHMMR knockdown. Numerals above each lane represent densitometric analysis of each protein relative to the loading control β-Actin. Data represents two independent experiments. RT-qPCR of KLK3 expression in response to 72 h of siHMMR knockdown in LNCaP (g) and MR49F (h) cells. Genes were normalised to GUSB and L19. Data are presented as mean ± SD of three biological replicates and two independent experiments. siControl was set to one and data was analysed by unpaired student’s T-test (siHMMR vs siCON; **p < 0.01, ***p < 0.001).
Targeting HMMR enhances enzalutamide efficacy
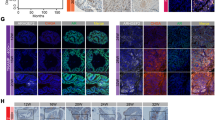
The observations that (1) HMMR is not modulated by enzalutamide, and (2) HMMR inhibition prevents AR transcriptional activity, provide a rationale for combining HMMR-targeted therapies with ARSIs as a potential new therapeutic strategy. Treatment of LNCaP and V16D PCa cells with enzalutamide in the presence or absence of 4-MU revealed that 4-MU enhances the efficacy of enzalutamide (Fig. 5a). Similarly, co-treatment with enzalutamide and siHMMR resulted in significant suppression of LNCaP viability compared to enzalutamide treatment or siHMMR treatment alone (Fig. 5b). Co-administration of 4-MU with darolutamide (Fig. 5c) or apalutamide (Fig. 5d) also significantly reduced cell viability compared to treatment with the individual agents. The Combination Index (CI) for each ARSI with 4-MU was calculated, which is a measure for determining whether a drug combination effect is synergistic (CI < 1), antagonistic (CI > 1) or additive (CI = 1) [17]. In both LNCaP and V16D cells, the CI for each combination was <1, indicative of synergy between 4-MU and ARSIs (Fig. 5e). To evaluate the combination of 4-MU and enzalutamide in vivo, we employed the V16D tumour xenograft model [41] according to the experimental plan depicted in Fig. 5f. Co-administration of 4-MU and enzalutamide markedly reduced tumour cell proliferation, as determined by immunostaining for the proliferative marker Ki67 (Fig. 5g). Tumour size was also significantly reduced by the combination of 4-MU and enzalutamide when compared to vehicle or enzalutamide treatment alone, but not compared to 4-MU single treatment (Supplementary Fig. 4a). An improvement in survival with the combination of 4-MU and enzalutamide was observed compared to the vehicle or single treatment groups (Supplementary Fig. 4b). Serum PSA levels remained unchanged across all treatments (Supplementary Figure 4c). Body weight measurements taken over the course of the study suggest no adverse effects of any treatment (Supplementary Figure 4d). Evaluation of the combination in PCa PDEs showed a significant reduction in epithelial cell proliferation from 29.42% Ki67 positively stained cells in control-treated PDE tissues to 11.01% upon co-treatment with 4-MU and enzalutamide (Fig. 5h). Compared to each agent when used individually, the combination was not significantly different to 4-MU treatment alone (24.51% Ki67 positively stained cells), but significantly suppressed proliferation compared to enzalutamide alone (21.91% ki67 positively stained cells) (Fig. 5h). Our results demonstrate in multiple models of PCa, including cell line and xenograft models of CRPC, and in clinically derived prostate tumours, that targeting HMMR enhances the anti-proliferative efficacy of enzalutamide.
a Combination treatment with 4-MU + ENZ significantly inhibits LNCaP and V16D PCa cell viability compared to Vehicle, 4-MU or ENZ treatment alone. Cells were counted on days 3 and 6 post-treatment using Trypan blue dye exclusion. Data are presented as mean ± SD of triplicate wells, represent two independent experiments, and were analysed using two-way ANOVA with Tukey’s multiple comparison test (****p < 0.0001). b Combination treatment with siHMMR+ENZ significantly inhibits LNCaP PCa cell viability compared to Vehicle, 4-MU or ENZ treatment alone, as assessed by Trypan blue dye exclusion. Data are presented as mean ± SD of triplicate wells, represent two independent experiments, and were analysed using two-way ANOVA with Tukey’s multiple comparison test (****p < 0.0001). c, d Combination treatment with 4-MU+darolutamide (DARO) or 4-MU+apalutamide (APA) significantly inhibits V16D PCa cell viability compared to Vehicle, 4-MU or ENZ treatment alone, as determined by the Trypan blue dye exclusion. Data are presented as mean ± SD of triplicate wells, represent two independent experiments, and were analysed using two-way ANOVA with Tukey’s multiple comparison test (****p < 0.0001). e The Chou-Talalay method [17] was used to determine the combination indices of the growth curves (A, B, D, E). f Schematic diagram of treatment regimen for in vivo subcutaneous xenograft model. g Combination treatment of 4-MU + ENZ significantly inhibits the proliferation of V16D PCa xenograft tumours compared to 4-MU or ENZ treatment alone. At the completion of the in vivo experiment, xenograft tumours were analysed for the proliferative marker Ki67 by immunohistochemistry (IHC). Quantification of Ki67 staining on the left and representative IHC images on the right. Data are presented as mean ± SD of 7 mice in each treatment group and analysed using one-way ANOVA with Turkey’s multiple comparison test (*p < 0.05, **p < 0.01). h Combination treatment with 4-MU + ENZ inhibits proliferation in patient-derived prostate cancer explants (PDEs). PDEs (n = 7) were treated as indicated for 48 h, then paraffin-embedded and formalin-fixed prior to immunohistochemistry (IHC) with proliferation marker Ki67. Digital images were manually counted. Quantification of Ki67 staining on the left and representative IHC images on the right. Data are presented as mean ± SD of 7 patients and analysed using one-way ANOVA with Dunnett’s multiple comparison test (*p < 0.05).
Discussion
ARSIs have revolutionised the treatment of advanced PCa but are unable to achieve durable responses, highlighting that combination approaches may be needed to overcome resistance. To rationally design an effective combination strategy, we used a data-driven approach wherein AR knockdown with siRNA was used as the benchmark of AR inhibition in PCa cells. HMMR was identified as an AR-regulated gene that is not suppressed by ARSIs enzalutamide, apalutamide or darolutamide. This study reveals HMMR as a survival factor in PCa and a promising co-target worth exploring to overcome resistance to ARSIs in PCa.
The potential role of HMMR in cancer development and progression is linked to two main functions, namely HA-induced cell migration and cell cycle progression [42,43,44]. HMMR overexpressing cells exhibit increased migratory and proliferative potential and the converse is true for HMMR downregulated cells [29, 45]. We observed a significant reduction in cell proliferation in a range of PCa cell lines following HMMR knockdown. Previous studies support our findings in enzalutamide-sensitive LNCaP cells [27, 46], whereas we provide the first-in-field evidence that HMMR is a potential target in CPRC, and importantly, in a setting of enzalutamide resistance. Given that there are no commercially available HMMR inhibitors, we employed a hyaluronic acid synthesis inhibitor 4-MU to suppress downstream HMMR activity [34, 47]. Hyaluronic acid is a glycosaminoglycan that forms an integral part of the extracellular matrix and has been implicated in the development and progression of PCa [47, 48]. Of note, high HA expression in the tumour stroma of PCa tissues is associated with increased proliferation, metastasis and disease recurrence post-radical prostatectomy [49,50,51,52]. The tumour-promoting roles of HA are linked to its interaction with two main receptors, CD44 and HMMR (RHAMM) [47]. Whilst both receptors have been detected in PCa cells, CD44 is reported to be highly expressed in androgen-independent and/or AR-negative cells whilst HMMR (RHAMM) is expressed in all PCa cell types [53,54,55]. Earlier studies also clearly show that CD44 expression does not correlate with poorer patient outcomes, PSA recurrence or metastasis in PCa patients [27, 49]. These observations suggest, however, that HMMR may be the predominant receptor-promoting HA-mediated oncogenic phenotypes in PCa. Indeed, we observed that treatment with 4-MU recapitulated mechanistic changes similar to HMMR knockdown cells.
Consistent with the microtubule-associated functions of HMMR [37], we saw a decrease in AR nuclear localisation with 4-MU treatment post DHT stimulation that resulted in decreased expression of well-known AR-regulated genes, and of AR itself. While it is possible that the localisation effects were due to the decreased AR steady-state levels observed upon 4-MU treatment, AR expression was not completely lost at the doses used in this study, and the AR that was expressed was clearly being held out of the nucleus in the presence of 4-MU. We propose that 4-MU instead works in a similar way to docetaxel, which is known to reduce AR expression by blocking AR nuclear localisation [56]. How this occurs mechanistically has not been defined for docetaxel, but studies into the regulation of AR by tumour suppressor PTEN may provide some insight. PTEN is known to suppress AR nuclear localisation, and the retention of AR in the cytoplasm promotes AR degradation via enzymatic factors [57].
In summary, as the AR continues to evolve in response to monotherapy, it is becoming more apparent that combination treatments may be critical for improving PCa survival. The work presented in this study highlights the importance of maximising the suppression of AR signalling using combinatorial approaches to achieve better control of cancer growth. We present strong evidence that persistent HMMR expression despite treatment with ARSIs provides a survival benefit to PCa cells and that co-targeting both HMMR and AR has promising therapeutic implications for the management of advanced PCa.
Data availability
1. Gillis, J. L., J. A. Hinneh, N. K. Ryan, S. Irani, M. Moldovan, L.-E. Quek, R. K. Shrestha, A. R. Hanson, J. Xie, A. J. Hoy, J. Holst, M. M. Centenera, I. G. Mills, D. J. Lynn, L. A. Selth and L. M. Butler (2021). NCBI Gene Expression Omnibus: GSE152254 “A feedback loop between the androgen receptor and 6-phosphogluoconate dehydrogenase (6PGD) drives prostate cancer growth.” eLife 10: e62592. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE152254
2. Gonit, M, Zhang, J, Salazar, MdA, Cui, H, Shatnawi, A, Trumbly, R & Ratnam, M 2011. NCBI Gene Expression Omnibus: GSE22483 ‘Hormone Depletion-Insensitivity of Prostate Cancer Cells Is Supported by the AR Without Binding to Classical Response Elements’, Molecular Endocrinology, vol. 25, no. 4, pp. 621-634. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE22483
3. Iglesias-Gato DThysell ETyanova SCrnalic SSantos ALima TSGeiger TCox JWidmark ABergh AMann MFlores-Morales AWikström P (2018). ProteomeXchange ID PXD009868. The Proteome of Prostate Cancer Bone Metastasis Reveals Heterogeneity with Prognostic Implications. https://clincancerres.aacrjournals.org/content/24
4. TCGA (2019) National Cancer Institute ID TCGA-PRAD. The Cancer Genome Atlas Prostate Adenocarcinoma (TCGA-PRAD). https://portal.gdc.cancer.gov/projects/TCGA-PRAD
5. Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E, Reva B, Antipin Y, Mitsiades N, Landers T, Dolgalev I, Major JE, Wilson M, Socci ND, Lash AE, Heguy A, Eastham JA, Scher HI, Reuter VE, Scardino PT, Sander CSawyers CL, Gerald WL (2010) NCBI Gene Expression Omnibus ID GSE21032.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA: Cancer J Clin. 2022;72:7–33.
Sandhu S, Moore CM, Chiong E, Beltran H, Bristow RG, Williams SG. Prostate cancer. Lancet. 2021;398:1075–90.
Miller DR, Ingersoll MA, Teply BA, Lin MF. Targeting treatment options for castration-resistant prostate cancer. Am J Clin Exp Urol. 2021;9:101–20. p
Rice, MA, Malhotra SV, and Stoyanova T, Second-generation antiandrogens: from discovery to standard of care in castration-resistant prostate cancer. Front Oncol, 2019. 9 https://doi.org/10.3389/fonc.2019.00801.
de Bono JS, Logothetis CJ, Molina A, Fizazi K, North S, Chu L. et al. Abiraterone and Increased Survival in Metastatic Prostate Cancer. N Engl J Med. 2011;364:1995–2005.
Scher HI, Fizazi K, Saad F, Taplin ME, Sternberg CN, Miller K. et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N Engl J Med. 2012;367:1187–97.
Beer TM, Armstrong AJ, Rathkopf DE, Loriot Y, Sternberg CN, Higano CS. et al. Enzalutamide in metastatic prostate cancer before chemotherapy. N Engl J Med. 2014;371:424–33.
Smith MR, Saad F, Chowdhury S, Oudard S, Hadaschik BA, Graff JN. et al. Apalutamide Treatment and Metastasis-free Survival in Prostate Cancer. New Engl J Med. 2018;378:1408–18.
Fizazi K, Shore N, Tammela TL, Ulys A, Vjaters E, Polyakov S. et al. Darolutamide in Nonmetastatic, Castration-Resistant Prostate Cancer. N Engl J Med. 2019;380:1235–46.
Schalken J, Fitzpatrick JM. Enzalutamide: targeting the androgen signalling pathway in metastatic castration-resistant prostate cancer. BJU Int. 2016;117:215–25.
Coutinho I, Day TK, Tilley WD, Selth LA. Androgen receptor signaling in castration-resistant prostate cancer: a lesson in persistence. Endocr Relat Cancer. 2016;23:T179–T197.
Al-Lazikani B, Banerji U, Workman P. Combinatorial drug therapy for cancer in the post-genomic era. Nat Biotechnol. 2012;30:679–92.
Thomas C, Baunacke M, Erb HHH, Füssel S, Erdmann K, Putz J, et al. Systemic triple therapy in metastatic hormone-sensitive prostate cancer (mHSPC): Ready for prime time or still to be explored? Cancers. 2022;14:8. p
Gillis JL, Hinneh JA, Ryan NK, Irani S, Moldovan M, Quek L-E. et al. A feedback loop between the androgen receptor and 6-phosphogluoconate dehydrogenase (6PGD) drives prostate cancer growth. eLife. 2021;10:e62592
Kuruma H, Matsumoto H, Shiota M, Bishop J, Lamoureux F, Thomas C. et al. A novel antiandrogen, Compound 30, suppresses castration-resistant and MDV3100-resistant prostate cancer growth in vitro and in vivo. Mol Cancer Ther. 2013;12:567–76.
Centenera MM, Carter SL, Gillis JL, Marrocco-Tallarigo DL, Grose RH, Tilley WD. et al. Co-targeting AR and HSP90 suppresses prostate cancer cell growth and prevents resistance mechanisms. Endocr Relat Cancer. 2015;22:805–18.
Chou TC, Talalay P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul. 1984;22:27–55.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3:research0034.1
Armstrong HK, Gillis JL, Johnson IRD, Nassar ZD, Moldovan M, Levrier C. et al. Dysregulated fibronectin trafficking by Hsp90 inhibition restricts prostate cancer cell invasion. Sci Rep. 2018;8:2090
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T. et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Centenera, MM, AD Vincent, M Moldovan, HM Lin, DJ Lynn, LG Horvath, et al., Harnessing the heterogeneity of prostate cancer for target discovery using patient-derived explants. Cancers, 2022. 14 https://doi.org/10.3390/cancers14071708.
Tieu T, Irani S, Bremert KL, Ryan NK, Wojnilowicz M, Helm M. et al. Patient-derived prostate cancer explants: a clinically relevant model to assess siRNA-based nanomedicines. Adv Healthc Mater. 2021;10:2001594
Gonit M, Zhang J, Salazar M, Cui H, Shatnawi A, Trumbly R. et al. Hormone depletion-insensitivity of prostate cancer cells is supported by the AR without binding to classical response elements. Mol Endocrinol. 2011;25:621–34.
Abida W, Cyrta J, Heller G, Prandi D, Armenia J, Coleman I. et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proce Natl Acad Sci. 2019;116:11428–36.
Fiorica F, Buttigliero C, Grigolato D, Muraro M, Turco F, Munoz F. et al. Addition of new androgen receptor pathway inhibitors to docetaxel and androgen deprivation therapy in metastatic hormone-sensitive prostate cancer: a systematic review and metanalysis. Curr Oncol. 2022;29:9511–24.
Network TCGAR. The molecular taxonomy of primary prostate cancer. Cell. 2015;163:1011–25.
Lin SL, Chang D, Chiang A, Ying SY. Androgen receptor regulates CD168 expression and signaling in prostate cancer. Carcinogenesis. 2008;29:282–90.
Korkes F, de Castro MG, de Cassio Zequi S, Nardi L, Del Giglio A, de Lima Pompeo AC. Hyaluronan-mediated motility receptor (RHAMM) immunohistochemical expression and androgen deprivation in normal peritumoral, hyperplasic and neoplastic prostate tissue. BJU Int. 2014;113:822–9.
Thangavel C, Boopathi E, Liu Y, Haber A, Ertel A, Bhardwaj A. et al. RB loss promotes prostate cancer metastasis. Cancer Res. 2017;77:982–95.
Iglesias-Gato D, Thysell E, Tyanova S, Crnalic S, Santos A, Lima TS. et al. The proteome of prostate cancer bone metastasis reveals heterogeneity with prognostic implications. Clin Cancer Res. 2018;24:5433–44.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM. et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Tolg, C, BJ Messam, JB McCarthy, AC Nelson, and EA Turley, Hyaluronan functions in wound repair that are captured to fuel breast cancer progression. Biomolecules, 2021. 11 https://doi.org/10.3390/biom11111551.
Lokeshwar VB, Lopez LE, Munoz D, Chi A, Shirodkar SP, Lokeshwar SD. et al. Antitumor activity of hyaluronic acid synthesis inhibitor 4-methylumbelliferone in prostate cancer cells. Cancer Res. 2010;70:2613–23.
Centenera MM, Hickey TE, Jindal S, Ryan NK, Ravindranathan P, Mohammed H. et al. A patient-derived explant (PDE) model of hormone-dependent cancer. Mol Oncol. 2018;12:1608–22.
Maxwell CA, Keats JJ, Crainie M, Sun X, Yen T, Shibuya E. et al. RHAMM is a centrosomal protein that interacts with dynein and maintains spindle pole stability. Mol Biol Cell. 2003;14:2262–76.
Darshan MS, Loftus MS, Thadani-Mulero M, Levy BP, Escuin D, Zhou XK. et al. Taxane-induced blockade to nuclear accumulation of the androgen receptor predicts clinical responses in metastatic prostate cancer. Cancer Res. 2011;71:6019–29.
Zhu ML, Horbinski CM, Garzotto M, Qian DZ, Beer TM, Kyprianou N. Tubulin-targeting chemotherapy impairs androgen receptor activity in prostate cancer. Cancer Res. 2010;70:7992–8002.
Tannock IF, de Wit R, Berry WR, Horti J, Pluzanska A, Chi KN. et al. Docetaxel plus Prednisone or Mitoxantrone plus Prednisone for advanced prostate cancer. N Engl J Med. 2004;351:1502–12.
Lowrance WT, Murad MH, Oh WK, Jarrard DF, Resnick MJ, Cookson MS. Castration-resistant prostate cancer: AUA Guideline Amendment 2018. J Urol. 2018;200:1264–72.
King CJ, Woodward J, Schwartzman J, Coleman DJ, Lisac R, Wang NJ, et al. Integrative molecular network analysis identifies emergent enzalutamide resistance mechanisms in prostate cancer. Oncotarget. 2017;8:111084–95. p
Turley EA, Austen L, Moore D, Hoare K. RAS transformed cells express both CD44 and RHAMM Hyaluran receptor: Only RHAMM is essensial for hyaluronan-promoted locomotion. Exp Cell Res. 1993;207:277–82.
Turley EA, Austen L, Vandeligt K, Clary C. Hyaluronan and a cell-associated hyaluronan binding protein regulate the locomotion of ras-transformed cells. J Cell Biol. 1991;112:1041–7.
Tolg C, Hamilton SR, Morningstar L, Zhang J, Zhang S, Esguerra KV. et al. RHAMM promotes interphase microtubule instability and mitotic spindle integrity through MEK1/ERK1/2 activity. J Biol Chem. 2010;285:26461–74.
Mele V, Soko L, Kölzer VH, Pfaff D, Muraro MG, Keller I, et al. The hyaluronan-mediated motility receptor RHAMM promotes growth, invasiveness and dissemination of colorectal cancer. Oncotarget. 2017;8:70617–29. p
Sun Y, Li Z, Song K. AR-mTOR-SRF axis regulates HMMR expression in human prostate cancer cells. Biomol Ther. 2021;29:667–77.
Tolg C, McCarthy JB, Yazdani A, Turley EA. Hyaluronan and RHAMM in wound repair and the “cancerization” of stromal tissues. Biomed Res Int. 2014;2014:10392
Liu M, Tolg C, Turley E. Dissecting the dual nature of hyaluronan in the tumor microenvironment. Front Immunol. 2019;10:947
Aaltomaa S, Lipponen P, Tammi R, Tammi M, Viitanen J, Kankkunen JP. et al. Strong stromal hyaluronan expression is associated with PSA recurrence in local prostate cancer. Urol Int. 2002;69:266–72.
Posey JT, Soloway MS, Ekici S, Sofer M, Civantos F, Duncan RC, et al. Evaluation of the prognostic potential of hyaluronic acid and hyaluronidase (HYAL1) for prostate cancer. Cancer Res. 2003;63:2638–44. p
Bharadwaj AG, Kovar JL, Loughman E, Elowsky C, Oakley GG, Simpson MA. Spontaneous metastasis of prostate cancer is promoted by excess hyaluronan synthesis and processing. Am J Pathol. 2009;174:1027–36.
Josefsson A, Adamo H, Hammarsten P, Granfors T, Stattin P, Egevad L. et al. Prostate cancer increases hyaluronan in surrounding nonmalignant stroma, and this response is associated with tumor growth and an unfavorable outcome. Am J Pathol. 2011;179:1961–8.
Tai S, Sun Y, Squires JM, Zhang H, Oh WK, Liang CZ. et al. PC3 is a cell line characteristic of prostatic small cell carcinoma. Prostate. 2011;71:1668–79.
SU C-Y, HUANG G-C, CHANG Y-C, CHEN Y-J, FANG H-W. Analyzing the expression of biomarkers in prostate cancer cell lines. In Vivo. 2021;35:1541–8.
Liu AY. Differential expression of cell surface molecules in prostate cancer cells. Cancer Res. 2000;60:3429–34. p
Kuroda K, Liu H, Kim S, Guo M, Navarro V, Bander NH. Docetaxel down-regulates the expression of androgen receptor and prostate-specific antigen but not prostate-specific membrane antigen in prostate cancer cell lines: implications for PSA surrogacy. Prostate. 2009;69:1579–85.
Lin HK, Hu YC, Lee DK, Chang C. Regulation of androgen receptor signaling by PTEN (phosphatase and tensin homolog deleted on chromosome 10) tumor suppressor through distinct mechanisms in prostate cancer cells. Mol Endocrinol. 2004;18:2409–23.
Acknowledgements
The authors thank Jianling Xie, Madison Helm, Samira Khabbazi, Elizabeth Collis, Niwa Sawako, Makoto Kamei and Randall Grose for expert assistance with data generation. We acknowledge the SAHMRI ACRF Cellular Imaging and Cytometry Core Facility which is generously supported by the Australian Cancer Research Foundation, Detmold Hoopman Group and the Australian Government through the Zero Childhood Cancer Programme and the SAHMRI Light Microscopy and Laser Capture Core Facility for the support and assistance in this work. Animal experiments were carried out at the Nagoya University Graduate School of Medicine animal facility, Tsurumai campus. LNCaP-V16D and LNCaP-MR49F cells were a kind gift from Dr. Amina Zoubeidi (Vancouver Prostate Centre, Vancouver, Canada). The authors are grateful to the study participants, as well as the urologists, nurses, and histopathologists who assisted in the recruitment and collection of patient information and pathology reports through the Australian Prostate Cancer BioResource and the Nagoya University Hospital.
Funding
This work was supported by grants from Cancer Australia (ID 1138766 to LMB, DJL and MMC; ID 2001432 to LAS and LMB; ID 2011672 to ZDN); the Movember Foundation and the Prostate Cancer Foundation of Australia through a Movember Revolutionary Team Award (MRTA-3 to LMB and LAS). JAH is supported by a Beacon of Enlightenment PhD Scholarship from the University of Adelaide. CYM is supported by an Early-Career Research Fellowship awarded by the Prostate Cancer Foundation of Australia. LAS and LMB are supported by Principal Cancer Research Fellowships (PRF2919 and PRF1117, respectively) awarded by the Cancer Council’s Beat Cancer project on behalf of its donors, the State Government through the Department of Health and the Australian Government through the Medical Research Future Fund. Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
JAH, Conceptualisation, Data curation, Investigation, Methodology, Project administration, Supervision, Validation, Visualisation, Writing—original draft, Writing—review and editing, approved the final manuscript; JLG, Conceptualisation, Data curation, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualisation, Writing—original draft, Writing—review and editing approved the final manuscript; CYM, Formal analysis, Data curation, Investigation, Validation, Visualisation, Writing—review and editing, approved the final manuscript; SI, Data curation, Investigation, Methodology, Writing—review and editing approved final manuscript; RKS, Data curation, Formal analysis, Investigation, Methodology, Validation, Visualisation, Writing – review and editing approved the final manuscript; NKR, Investigation, Data curation, Methodology, Validation, Writing—review and editing, approved final manuscript; EA, Investigation, Data curation, Formal analysis, Methodology, Supervision, Validation, Writing—review and editing approved final manuscript; ZN: Investigation, Data curation, Methodology, Formal analysis, Supervision, Validation, Writing—review and editing, approved the final manuscript; DJL, Conceptualisation, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Visualisation, Writing—review and editing, approved the final manuscript; LAS, Conceptualisation, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Supervision, Visualisation, Writing—review and editing, approved the final manuscript. MK: Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Visualisation, Writing—review and editing, approved final manuscript; MMC, Conceptualisation, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Supervision, Visualisation, Writing—original draft, Writing—review and editing, approved final manuscript; LMB, Conceptualisation, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Visualisation, Writing—original draft, Writing— review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Human participants: Prostate cancer tissues were obtained with written informed consent from men undergoing radical prostatectomy at St Andrew’s Hospital (Adelaide, Australia) through the Australian Prostate Cancer BioResource, and Nagoya University Hospital (Nagoya, Japan). Ethical approval for the use of human prostate tumours was obtained from the Human Research Ethics Committees of the University of Adelaide (approval number H-2012-016), St Andrew’s Hospital in Adelaide (approval number 80) and Nagoya University Graduate School of Medicine (approval number 2020-01-17). All experiments were performed in accordance with the guidelines of the National Health and Medical Research Council (Australia) and Ministry of Education, Culture, Sports, Science and Technology, and the Ministry of Health, Labour and Welfare (Japan). Studies were performed according to the Declaration of Helsinki. Animal Studies: Ethical approval for animal studies were obtained from the Animal Ethics Committees of the University of Adelaide (approval number M-2020-014) and Nagoya University Graduate School of Medicine (approval number 31460). Experiments were carried out in accordance with the recommendations of the National Health and Medical Research Council of Australia, the Science Council of Japan and the ARRIVE 2.0 guidelines.
Consent for publication
All authors consent to the publication of this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hinneh, J.A., Gillis, J.L., Mah, C.Y. et al. Targeting hyaluronan-mediated motility receptor (HMMR) enhances response to androgen receptor signalling inhibitors in prostate cancer. Br J Cancer 129, 1350–1361 (2023). https://doi.org/10.1038/s41416-023-02406-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02406-8