Abstract
Background
Identifying biomarkers to predict immune checkpoint inhibitor (ICI) efficacy is warranted. Considering that somatic mutation-derived neoantigens induce strong immune responses, patients with a high tumour mutational burden reportedly tend to respond to ICIs. However, there are several conflicting data. Therefore, we focused on the original function of neoantigenic mutations and their impact on the tumour microenvironment (TME).
Methods
We evaluated 88 high-frequency microsatellite instability (MSI-H) colorectal cancers and analysed the function of the identified neoantigenic mutations and their influence on programmed cell death 1 (PD-1) blockade efficacy. The results were validated using The Cancer Genome Atlas (TCGA) datasets.
Results
We identified frameshift mutations in RNF43 as a common neoantigenic gene mutation in MSI-H tumours. However, loss-of-function RNF43 mutations induced noninflamed TME by activating the WNT/β-catenin signalling pathway. In addition, loss of RNF43 function induced resistance to PD-1 blockade even in neoantigen-rich tumours. TCGA dataset analyses demonstrated that passenger rather than driver gene mutations were related to the inflamed TME in diverse cancer types.
Conclusions
We propose a novel concept of “paradoxical neoantigenic mutations” that can induce noninflamed TME through their original gene functions, despite deriving neoantigens, suggesting the significance of qualities as well as quantities in neoantigenic mutations.

Similar content being viewed by others
Background
Cancer develops through the acquisition of immune escape mechanisms [1], including the use of suppressive immune checkpoint molecules such as programmed cell death-1 (PD-1) and PD-1 ligand 1 (PD-L1). Antitumor immune responses can be restored by blocking these immunosuppressive pathways [2]. Cancer immunotherapy with immune checkpoint inhibitors (ICIs), including monoclonal antibodies (mAbs) against cytotoxic T lymphocyte-associated protein 4 (CTLA-4) and PD-1/PD-L1, rescues dysfunctional cytotoxic effector CD8+ T cells, leading to tumour regression [3, 4]. ICIs have been demonstrated to be effective against diverse cancer types, including gastrointestinal cancers, leading to a paradigm shift in cancer treatment [5,6,7]. However, more than half of patients do not respond to ICIs, and identifying biomarkers to predict clinical benefits is imperative.
Somatic mutation-derived neoantigens, which can be recognised as non-self-antigens, reportedly induce strong immune responses similar to those of foreign antigens [8, 9]. Thus, neoantigens are presumed to induce an inflamed tumour microenvironment (TME), which is substantial for the ICI response, and the number of neoantigens is reportedly correlated with the inflamed TME [9,10,11]. Therefore, tumour mutational burden (TMB) is one of the predictive biomarker candidates for ICIs [10, 12, 13], and ICIs have been approved for treatment of high-frequency microsatellite instability (MSI-H) cancers, which generally have an extremely high TMB [14,15,16]. However, there are several conflicting data [17,18,19], and the TMB does not seem to be a sufficient predictor of ICI response. Even in MSI-H cancers, the response rates of PD-1 blockade therapies are only ~50% [20, 21].
Tumours also evade antitumor immunity through “immune editing”, which involves the elimination of highly immunogenic tumour cells or mutation of their own human leukocyte antigen (HLA) molecules to decrease antigen presentation [2, 22,23,24]. Furthermore, in the process of accumulating somatic mutations, certain cancer signalling pathways promote immune evasion [25,26,27,28,29,30]. Therefore, quantitative assessment of somatic mutations seems to be insufficient for predicting ICI response, and it can be essential to elucidate the effects of mutated gene functions on the TME. In this study, we focused on the functions of neoantigenic driver gene mutations in MSI-H colorectal cancers. In our MSI-H cohort [31], neoantigens derived from mutations in RNF43 were commonly observed, as previously reported [32, 33]. However, tumours with RNF43 mutations did not always have an inflamed TME. We demonstrated that some forms of mutated RNF43 could suppress antitumor immunity and induce resistance to PD-1 blockade, even when neoantigens were present, by activating the WNT/β-catenin signalling pathway [29, 30, 34, 35]. These results were validated using The Cancer Genome Atlas (TCGA) datasets. We propose a novel concept of “paradoxical neoantigenic mutations” that can induce the noninflamed TME due to their original gene functions, despite deriving neoantigens. When evaluating neoantigens, it is essential to consider the functions of the neoantigenic gene mutations in addition to the number of neoantigens, as the functions of mutated molecules, such as mutated RNF43, can affect antitumor immunity.
Methods
Study design
Eighty-eight patients with MSI-H colorectal cancer who underwent surgery were enrolled in this study (Table S1) [31]. Written informed consent was obtained prior to participation. The clinical information of patients was obtained from their medical records.
Whole-exon sequencing
Exome sequencing data were obtained from a publicly available dataset [31]. Genomic DNA was isolated from each sample and subjected to exonic fragment enrichment using a SureSelect Human All Exon Kit v5 (Agilent Technologies, Santa Clara, CA). Massively parallel sequencing of isolated fragments was conducted with a HiSeq2500 (Illumina, San Diego, CA) using the paired-end option. Paired-end whole-exon sequencing (WES) reads were independently aligned to a human reference genome (hg38) using BWA [36], Bowtie2 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml), and NovoAlign (http://www.novocraft.com/products/novoalign/). Somatic mutations were called using MuTect (http://www.broadinstitute.org/cancer/cga/mutect), SomaticIndelDetector (http://www.broadinstitute.org/cancer/cga/node/87), and VarScan (http://varscan.sourceforge.net). Mutations were discarded if (i) the read depth was <20 or the variant allele frequency was <0.1, (ii) they were supported by only one genome strand, or (iii) they were present in normal human genomes in either the 1000 Genomes Project dataset (http://www.internationalgenome.org/) or our in-house database. Gene mutations were annotated with SnpEff (http://snpeff.sourceforge.net).
RNA-sequencing
Transcriptome sequencing data were obtained through a previous study; these data have already been reported and are publicly available [31]. Total RNA was extracted using RNA Bee reagent (Tel-Test Inc., Friendswood, TX) and treated with DNase I (Qiagen, Venlo, NL) using a Rneasy Mini Kit (Qiagen). RNA integrity was evaluated using either a Bioanalyzer (Agilent Technologies) or TapeStation (Agilent Technologies). RNA with a high RNA integrity number underwent RNA-seq using an NEBNext Ultra Directional RNA Library Prep Kit (New England BioLabs, Ipswich, MA), with complementary DNA (cDNA) prepared from polyA-selected RNA. RNA with a low RNA integrity number was subjected to RNA-seq using a TruSeq RNA Access Library Prep Kit (Illumina), with cDNA generated using random primers. Prepared RNA-seq libraries underwent next-generation sequencing of 120 bp from both ends (paired-end reads). The expression level of each gene was computed using DESeq2 (http://bioconductor.org/packages/release/bioc/html/DESeq2.html) with VST transformation, and gene fusions were detected using a deFuse pipeline (https://bitbucket.org/dranew/defuse) and STAR (https://github.com/alexdobin/STAR).
Long-read sequencing
We determined the precise sequences of ~3500 bp regions between the 5′ and 3′ untranslated region (UTR) of HLA-ABC gene using single-molecule real-time sequencing conducted on a Sequel platform (Pacific Biosciences, Menlo Park, CA). The genotypes of HLA-A, HLA-B, and HLA-C genes were determined by comparing the obtained sequences with reference sequences and were subsequently validated using NGSengine software.
Neoantigen prediction
Neoantigens were predicted as reported in a previous study [24]. We assumed that the genomic DNA sequences carrying somatic nonsynonymous mutations that were also detected in transcriptome sequencing reads produced abnormal peptides. The affinities of abnormal peptides (nine amino acids long) for MHC class I molecules in individual tumours were predicted using NetMHCpan (Ver. 4.0) [37]. Abnormal peptides that were presumed to result from frameshift (fs) mutations were also included in our cohort analysis. The peptides in the top two percentiles for affinity were defined as neoantigens.
Immunohistochemistry
Formalin-fixed paraffin-embedded (FFPE) specimens from human clinical samples were cut into 4 µm sections and mounted on glass slides. Antigen retrieval was performed in citrate buffer (pH 6) at 121 °C for 20 min. CD8+ cell infiltration within the tumour gland was observed and blindly scored by an experienced pathologist according to the following criteria: 0 (0–5% CD8 staining); 1 (5–10% CD8 staining); 2 (10–15% CD8 staining); and 3 (>15% CD8 staining). Simultaneously, CD8+ cells were quantified using an automated imaging system (Vectra 3.0; Perkin-Elmer, Waltham, MA, USA) equipped with the analytical software InForm 2.2 (Perkin-Elmer).
Frozen tissue specimens from mice were cut into 6 µm sections and mounted on glass slides. Individual slides were incubated overnight at 4 °C with anti-CD11c mAb (D1V9Y; Cell Signalling Technology, Danvers, MA) and anti-CD8 mAb (D4W2Z; Cell Signaling Technology). The slides were then incubated in SignalStain® Boost IHC Detection Reagent (Cell Signaling Technology), and the colour was developed using a Signalstain® DAB Substrate Kit (Cell Signalling Technology). CD11c- and CD8-positive cells were counted; three fields (×400) comprising tumour cells were randomly selected and counted for each slide. The mean of the three area counts for each tumour was used for statistical analysis.
Gene set enrichment analysis
Enriched pathways were determined using the GSEA tool available from the Broad Institute website (http://software.broadinstitute.org/gsea/index.jsp) [38].
Cell lines and reagents
The HEK293T (human embryonic kidney cell), CT26 (murine colon cancer), RENCA (murine renal cell carcinoma), and A20 (murine lymphoma) cell lines were purchased from ATCC (Manassas, VA). The HEK293T and CT26 cell lines were maintained in DMEM (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) supplemented with 10% foetal calf serum (FCS; Cytiva, Tokyo, Japan). The RENCA and A20 cell lines were maintained in RPMI medium (FUJIFILM Wako Pure Chemical Corporation) supplemented with 10% FCS. All tumour cells were confirmed to be Mycoplasma (−) using a PCR Mycoplasma Detection Kit (TaKaRa, Shiga, Japan) according to the manufacturer’s instructions before use. Recombinant human Wnt3a protein was obtained from R&D (Minneapolis, MN). Recombinant murine Wnt3a protein was purchased from PeproTech (Cranbury, NJ). Rat anti-mouse PD-1 mAb (RMP1-14) and control rat IgG2a mAb (RTK2758) were obtained from BioLegend (San Diego, CA).
Constructs, virus production, and transfection
The pGreenFire 2.0 TCF/LEF Reporter lentivirus was purchased from System Biosciences (Palo Alto, CA). Human RNF43 WT, 117 fs, and 659 fs cDNA subcloned into separate pMSCV expression retroviral vectors were purchased from Vectorbuilder (Chicago, IL). Human RNF43 I48T was subcloned from human RNF43 WT using a KOD-Plus-Mutagenesis Kit (TOYOBO, Osaka, Japan). A CTAG1B-subcloned pMSCV expression retroviral vector was purchased from Vectorbuilder. A lentiviral vector carrying a short hairpin RNA (shRNA) sequence to knock down murine Rnf43 expression (sh-Rnf43) was purchased from Vectorbuilder. Viral vectors were transfected into packaging cells using the Lipofectamine 3000 Reagent (Thermo Fisher Scientific, Waltham, MA). After 48 h, the supernatant was concentrated and transfected into each cell line.
Reverse transcription quantitative PCR
Total RNA was reverse transcribed into cDNA using PrimeScript RT Master Mix (TaKaRa), and reverse transcription quantitative PCR (RT-qPCR) was performed using TB Green Premix Ex Taq II (TaKaRa) according to the manufacturer’s instructions. Human GAPDH or murine Gapdh was used as an internal control. Atf3 and Ccl4 gene expression levels were measured after cells were treated with 10 ng/mL murine Wnt3a protein for 24 h. The experiments were performed in triplicate. The primers used are listed in Table S2.
Luciferase assay
Each human RNF43 mutation was introduced into HEK293T cells with the pGreenFire 2.0 TCF/LEF Reporter. After treatment with a human Wnt3a protein at 500 ng/mL for 24 h, luciferase activities were measured using the ONE-StepTM Luciferase Assay System (System Biosciences). The luciferase activities of mock, human RNF43 WT, human RNF43 117 fs, human RNF43 659 fs, and human RNF43 I48T-overexpressing HEK293T cells were compared. All experiments were repeated independently in triplicate.
Western blotting
After treatment with 10 ng/mL murine Wnt3a for 24 h, subconfluent cells were washed with phosphate-buffered saline buffer and harvested using 1% sodium dodecyl sulphate (SDS). Whole-cell lysates were separated using SDS-PAGE and blotted onto a polyvinylidene fluoride membrane. Post-blocking, the membrane was probed with a primary antibody. After rinsing twice with tris-buffered saline buffer, the membrane was incubated with a horseradish peroxidase-conjugated secondary antibody and washed, followed by visualisation and quantification using an ECL detection system and a ChemiDoc imaging system (Bio-Rad, Hercules, CA). Antibody specific for nonphosphorylated (active) β-catenin (D13A1) was purchased from Cell Signaling Technology. Antibody against β-actin (A5441) was purchased from Sigma–Aldrich (St. Louis, MO).
In vivo animal models
Female Balb/c mice (6–8 weeks old) were purchased from SLC Japan (Shizuoka, Japan). C57BL/6J-Prkdc<scid>/Rbrc mice (SCID; RBRC01346) were provided by RIKEN BRC (Tsukuba, Japan) through the National BioResource Project of the MEXT/AMED, Japan. CT26 (1 × 106), RENCA (2 × 106), or A20 cells (4 × 106) were inoculated subcutaneously, and tumour volume was monitored every 3 days. The mean of the long and short diameters was used to generate tumour growth curves. The mice were grouped when the tumour volume reached ~100 mm3 (Day 0), and an anti-PD-1 mAb (200 μg/mouse) or a control mAb was administered intraperitoneally every 3 days thereafter (three times total). The tumours were harvested 14 days post-tumour cell inoculation and evaluated using immunohistochemistry (IHC). All in vivo experiments were performed at least twice (n = 4–6 per group). All mice were maintained under specific pathogen-free conditions in the Institute of Biophysics animal facility.
TCGA dataset analysis
WES and RNA-seq data for each cancer type were acquired from the TCGA database published in cBioPortal (https://www.cbioportal.org/) [39, 40]. The TMB was defined as the sum of the total number of nonsynonymous single-nucleotide variants (SNVs) and fs mutations. Peptide sequence information was obtained from UniProt (https://www.uniprot.org/) [41]. HLA typing data for each patient was available at TCIA (https://tcia.at/home) [42]. Cancer types with no HLA data and haematological malignancies were excluded from the analyses. The affinity of abnormal peptides for HLA class I molecules in individual tumours was predicted using NetMHCpan 4.0, and the peptides in the top two percentiles for affinity were defined as neoantigens. We defined the genes listed in the Cancer Gene Census (https://cancer.sanger.ac.uk/census) of COSMIC (https://cancer.sanger.ac.uk/cosmic) as “driver genes” and the others as “passenger genes” (Table S3) [43]. The average expression of CD8A, GZMA, and PRF1 was computed from the RNA-seq data of the same patients to derive an “immune activity score”. The correlations between the TMB, driver/passenger mutational burden, total neoantigen load or neoantigen load derived from driver/passenger mutations, and the immune activity score were compared by computing Pearson’s correlation coefficient (R). The analysis was conducted on cases for which the TMB and predicted neoantigen loads were within the 95% confidence interval.
Statistics
GraphPad Prism 9 (GraphPad Software, San Diego, CA) was utilised for statistical analyses. Frequencies were compared between groups using Fisher’s exact test. The relationships of continuous variables between or among groups were compared using t-test or one-way analysis of variance (ANOVA), respectively. The relationships among tumour volume curves were compared using two-way ANOVA. Overall survival was defined as the time from the date of surgery to that of death from any cause. The log-rank test was used to compare Kaplan–Meier curves. The correlations between two variables were evaluated by computing Pearson’s correlation coefficient (R). For multiple comparisons testing, Bonferroni corrections were employed. All tests were two-tailed, and P values < 0.05 were considered statistically significant.
Results
Neoantigenic RNF43 mutations that are frequently observed in MSI-H colorectal cancer do not consistently induce an inflamed TME
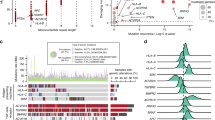
In this study, we evaluated 88 MSI-H colorectal cancer patients using WES, RNA-sequencing, and HLA long-read sequencing data (Table S1) [24]. We predicted neoantigens using NetMHCpan 4.0 and identified that peptides derived from driver gene RNF43 fs mutations frequently became neoantigens (47 patients; Arg117fs, 7; Gly659fs, 37) (Fig. 1a, b) [37]. Although we reported that patients with HLA abnormalities had noninflamed TME and that the abnormalities were related to neoantigens, there were a considerable number of patients with intact HLA [24]. Focusing on this HLA-intact population (n = 58), tumours with RNF43 fs mutations had significantly higher CD8+ T cell infiltration than those without it (Fig. S1A), which seems to be consistent with previous studies demonstrating that neoantigens induce an inflamed TME [9,10,11]. Alternatively, when RNF43 fs mutations were divided, patients with the RNF43 659 fs mutation were associated with more CD8+ T cell infiltration than those without RNF43 fs mutations; however, patients with the other RNF43 fs mutations, such as the 117 fs mutation, were not (Fig. 1c). With regard to TMB, there was no significant correlation with CD8+ T cell infiltration or the RNF43 fs mutations (Figs. 1d and S1B). These results suggest that not all RNF43 fs mutations result in an inflamed TME despite being common neoantigens.
a Predicted neoantigens in our high-frequency microsatellite instability (MSI-H) cohort. Eighty-eight patients with MSI-H colorectal cancer who underwent surgical resection were enrolled in this study. Whole-exon sequencing (WES), RNA-sequencing, and long-read HLA sequencing were performed with the resected tumour samples. The NetMHCpan algorithm (version 4.0) was used to predict possible neoantigens from WES, RNA-sequencing, and long-read HLA sequencing data. b Distributions of RNF43 frameshift (fs) mutations. All RNF43 fs mutations derived from 88 MSI-H colorectal cancer samples are depicted with arrows. c Comparison of CD8+ T cell infiltration according to the RNF43 status in patients with intact HLA. Summary of CD8+ T cell infiltration in patients with the RNF43 659 fs mutation, with the other fs mutations, and without any fs mutations is depicted. d Comparison of tumour mutational burden (TMB) according to the RNF43 status in patients with intact HLA. The summary of TMB in patients with the RNF43 659 fs mutation, with the other fs mutations, and without any fs mutations is depicted. One-way ANOVA with the Bonferroni correction is used in (c) and (d) for statistical analyses. The means and SDs are depicted. *P < 0.05; ns not significant.
RNF43 fs loss-of-function mutations activate the WNT/β-catenin signalling pathway
Considering the diverse TMEs resulting from RNF43 fs mutations in our cohort, we analysed the functions of RNF43 fs mutations. RNF43 is a single-chain, transmembrane E3 ubiquitin ligase comprising 783 amino acids (Fig. 1b). The binding and ubiquitination of the WNT receptor, involving the RING domain, are mapped to the N-terminus, followed by the Disheveled-2 (DVL2) binding domain. No role has been assigned to the C-terminal extension. RNF43 negatively regulates the WNT/β-catenin signalling pathway and activates the signalling pathway in the presence of loss-of-function mutations [44]. We generated HEK293T cell lines with a TCF/LEF luciferase reporter (TCF/LEF Reporter-HEK293T cells) overexpressing human RNF43 wild-type (WT); I48T, a representative WNT/β-catenin hyperactivating mutation [44]; 117 fs; or 659 fs (Fig. S2A). These cell lines were used to evaluate the activity of the WNT/β-catenin signalling pathway. We observed that RNF43 WT and RNF43 659 fs suppressed the WNT/β-catenin signalling pathways, whereas RNF43 117 fs did not (Fig. 2a), suggesting that RNF43 117 fs is a loss-of-function mutation, as previously reported [34, 45, 46]. Furthermore, WNT/β-catenin signalling pathway activation was observed in patients with the RNF43 fs mutations other than 659 fs by gene set enrichment analysis (GSEA) of the RNA-sequencing data for the HLA-intact population in our MSI-H cohort (Fig. S3A). In addition, patients with RNF43 fs mutations, except 659 fs, had significantly shorter overall survival (OS) compared with patients with the other mutations (Fig. S3B and Table S4).
a Luciferase reporter assay. The luciferase activity in each human RNF43-overexpressing TCF/LEF Reporter-HEK293T cell line treated with human Wnt3a was analysed. The fold changes determined by comparison to mock-overexpressing control cells are presented. b Western blotting. Human RNF43-overexpressing CT26 cells treated with murine Wnt3a were analysed using western blotting. Each band was quantified, and the fold change was computed by comparison to mock-overexpressing control cells. Representative blots (left) and the corresponding summary (right) are depicted. β-actin was used as an internal control. c Atf3 and Ccl4 gene expression in CT26 cells. Gene expression post-treatment with murine Wnt3a was analysed using RT-qPCR. Murine Gapdh was used as an internal control. The fold changes determined by comparison to mock-overexpressing control cells are presented. d Western blotting. Rnf43-knockdown (sh-Rnf43) RENCA cells treated with murine Wnt3a were analysed using western blotting. Each band was quantified, and the fold change was computed by comparison to sh-Control cells. Representative blots (left) and the corresponding summary (right) are depicted. β-actin was used as an internal control. e Atf3 and Ccl4 gene expression in RENCA cells. Gene expression post-treatment with murine Wnt3a was analysed using RT-qPCR. The fold changes determined by comparison to sh-Control cells are presented. All in vitro experiments were performed in triplicate. One-way ANOVA with the Bonferroni correction is used in (a–c), and t-test is used in (d) and (e) for statistical analyses. The means and SEMs are depicted. *P < 0.05; **P < 0.01; ***P < 0.001; **** < 0.0001; ns not significant.
RNF43 fs loss-of-function mutation reduces CCL4 levels by activating the WNT/β-catenin signalling pathway
A previous report demonstrated that WNT/β-catenin signalling pathway activation induces a noninflamed TME via ATF3/CCL4 [29]. The transcriptional repressor ATF3 suppresses CCL4 and prevents effector CD8+ T cell infiltration by reducing dendritic cell (DC) infiltration [29]. Therefore, to evaluate the impact of RNF43 fs mutations on TME, we examined WNT/β-catenin signalling pathway activation and Atf3 and Ccl4 expression using murine cell lines. We used the human RNF43 WT- or 117fs-overexpressing CT26 (murine colorectal cancer) cell line, as this cell line had low mouse Rnf43 expression (Fig. S2B, C). Western blotting analyses demonstrated that active β-catenin was suppressed in human RNF43 WT-overexpressing CT26 cells, but not in human RNF43 117fs-overexpressing cells (Fig. 2b). Furthermore, the human RNF43 117 fs loss-of-function mutation led to higher Atf3 and lower Ccl4 expression than human RNF43 WT (Fig. 2c). In addition, we performed mouse Rnf43 knockdown using shRNA in RENCA and A20 cell lines because these cell lines had high Rnf43 expression (Fig. S2B, D). Rnf43 knockdown increased active β-catenin expression in both cell lines, resulting in higher Atf3 and lower Ccl4 expression (Figs. 2d, e, S2E, F). Collectively, these results indicate that RNF43 fs loss-of-function mutations activate the WNT/β-catenin signalling pathway, leading to a reduction in the level of CCL4, a crucial chemokine for the inflamed TME.
RNF43 loss-of-function mutations induce a noninflamed TME and resistance to PD-1 blockade even if they produce neoantigens
We explored the impact of RNF43 loss-of-function mutations on the therapeutic effects of ICIs and TMEs using mouse models. We used the human RNF43 WT or 117fs-overexpressing CT26 cell lines, and compared with control tumours, human RNF43 WT-overexpressing tumours demonstrated reduced growth and improved response to an anti-PD-1 mAb (Fig. 3a). Conversely, human RNF43 117fs-overexpressing tumours grew similarly to control tumours, and PD-1 blockade was less effective against these tumours than against human RNF43 WT-overexpressing tumours (Fig. 3a). These effects were not observed in immunodeficient mice (Fig. S4A). IHC revealed that human RNF43 117fs-overexpressing tumours had a lesser DC and CD8+ T cell infiltration than human RNF43 WT-overexpressing tumours (Fig. 3b, c), which is consistent with a previous study and the results of our in vitro analysis [29]. Similarly, Rnf43-knockdown tumours (RENCA), generated using shRNA, were resistant to PD-1 blockade (Fig. 4a). Previous studies have demonstrated that NY-ESO-1 is a neoantigen in Balb/c mice [47, 48]; therefore, we generated CTAG1B (encoding NY-EOS-1)-overexpressing cell lines (RENCA and A20 cell lines) to mimic neoantigen-rich models. Accordingly, CTAG1B-overexpressing tumours showed improved response to PD-1 blockade compared with control tumours (Figs. 4a, b, and S4B). However, Rnf43 knockdown abolished the efficacy against CTAG1B-overexpressing tumours (Figs. 4b and S4B). Efficacy against CTAG1B-overexpressing tumours was not observed in immunodeficient mice (Fig. S4C). IHC demonstrated that Rnf43 knockdown impaired DC and CD8+ T cell infiltration even in CTAG1B-overexpressing tumours (Fig. 4c, d). Overall, RNF43 loss-of-function mutations could induce a noninflamed TME and resistance to PD-1 blockade even if they produced neoantigens.
a Tumour growth of human RNF43-overexpressing tumours. CT26 cells (1 × 106) were inoculated subcutaneously, and tumour volume was monitored every 3 days. The means of the long and short diameters were used to generate tumour growth curves. The mice were grouped when the tumour volume reached ~100 mm3 (Day 0), and an anti-PD-1 monoclonal antibody (mAb) or a control mAb was administered intraperitoneally every 3 days thereafter for three times total (n = 6 per group). b, c Immunohistochemistry (IHC) for CD11c (b) and CD8 (c). Tumours were harvested 14 days post-tumour cell inoculation for IHC. The average count of three fields (400×) for each tumour is used for statistical analysis. Representative staining figures (left) and the corresponding summaries (right) are depicted. All in vivo experiments were performed in duplicate, and similar results were obtained. Two-way ANOVA with the Bonferroni correction is used in (a), and one-way ANOVA with the Bonferroni correction is used in (b) and (c) for statistical analyses. The means and SEMs are depicted. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; ns not significant.
a, b Efficacy of PD-1 blockade against RENCA (a) or CTAG1B-overexpressing RENCA (b) tumours. RENCA cells (2 × 106) were inoculated subcutaneously. Subsequent steps were performed as described in Fig. 3 (n = 5 per group). c, d IHC for CD11c (c) and CD8 (d). Tumours were harvested 14 days post-tumour cell inoculation for IHC. The average count of three fields (400×) for each tumour was used for statistical analysis. Representative staining figures (left) and the corresponding summaries (right) are depicted. All in vivo experiments were performed in duplicate, and similar results were obtained. Two-way ANOVA is used in (a) and (b), and one-way ANOVA with the Bonferroni correction is used in (c) and (d) for statistical analyses. The means and SEMs are depicted. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; ns not significant.
Neoantigens derived from passenger rather than driver gene mutations are related to the inflamed TME
Considering that several driver gene mutations, in addition to RNF43 mutations, have been reported to have immunosuppressive functions [25,26,27,28,29,30], we compared the effects of neoantigens derived from driver and passenger gene mutations on immune responses. We defined “driver genes” as genes listed in the Cancer Gene Census of COSMIC and the other genes as “passenger genes” (Table S3). We computed the average expression of CD8A and cytotoxicity-related genes (GZMA and PRF1) to produce an “immune activity score” based on a previous report [11] and evaluated the correlations between the TMB or predicted neoantigen loads and this score using TCGA datasets. As previously reported, there were certain positive correlations between the TMB or predicted neoantigen loads and immune activity scores in several cancer types (Table S5) [11]. When the driver and passenger mutations were evaluated separately, the latter had more positive and significant correlations with immune activity scores (Fig. 5, Tables S6 and 7).
a The heatmap for Pearson’s correlation coefficients (R) between the driver/passenger mutational burden and immune activity scores (left) and the difference in R (ΔR) for each cancer type (right) are depicted. b The heatmaps for R between the neoantigen loads derived from driver/passenger mutations and immune activity scores (left) and ΔR for each cancer type (right) are depicted. Cancer types with P values < 0.05 for either driver or passenger mutations are presented. Bold R numbers indicate P values < 0.05. UCS uterine carcinosarcoma, BRCA breast invasive carcinoma, CA colorectal adenocarcinoma, CESC cervical squamous cell carcinoma, STAD stomach adenocarcinoma, UCEC uterine corpus endometrial carcinoma, ACC adrenocortical carcinoma, PRAD prostate adenocarcinoma, HNSC head and neck squamous cell carcinoma.
Discussion
The need to identify biomarkers to predict the response to ICIs is imperative in clinical settings. Even though several basic and clinical studies on biomarkers have been conducted, we still cannot predict the response accurately. The TMB can be a predictive biomarker as it reflects the number of neoantigens [10, 12, 13]. However, the neoantigen theory, which states that neoantigens induce an inflamed TME, is not always accurate as the correlation between neoantigens and the TME immune status is not always substantial, and the effect of ICI therapy is not well predicted [17, 18]. In this study, we demonstrated that the functions of genetic variants could paradoxically induce a noninflamed TME even if the variants served as neoantigens (“paradoxical neoantigenic mutations”). Considering that several driver gene mutations reportedly play substantial roles in antitumor immunity evasion as well as in cell proliferation and cell death [25,26,27,28,29,30], we should evaluate the qualities in addition to quantities of neoantigenic mutations.
We evaluated 88 MSI-H colorectal cancer patients and identified that fs mutations in the driver gene RNF43 were shared neoantigens among patients, as previously reported [32]. Tumours with neoantigens derived from these RNF43 fs mutations tended to have an inflamed TME, which is consistent with the neoantigen theory. However, the TME demonstrated differences among the fs mutations in our study. RNF43 is a tumour suppressor gene that suppresses the WNT/β-catenin signalling pathway, and its RING domain plays a substantial role in its function [49]. Although fs mutations generally result in a loss of function, some previous studies have demonstrated that functions differ among RNF43 fs mutations [34, 35, 45, 46]. We demonstrated that RNF43 117fs, a common fs mutation in the N-terminal side of the RING domain, is a loss-of-function mutation that activates the WNT/β-catenin signalling pathway [33,34,35, 45]. Conversely, RNF43 659fs, which is in the C-terminal side of the RING domain, was comparable to RNF43 WT in suppressing the WNT/β-catenin signalling pathway [34, 35, 45, 46]. Previous reports have demonstrated that WNT/β-catenin signalling pathway suppresses immunity [26, 29]. Accordingly, the WNT/β-catenin signalling pathway was not activated via the 659fs mutation, resulting in an inflamed TME owing to neoantigens. Alternatively, the WNT/β-catenin signalling pathway activation resulting from loss-of-function fs mutations led to a noninflamed TME even in the presence of neoantigens and resistance to PD-1 blockade. To our knowledge, this is the first study demonstrating that neoantigenic RNF43 mutations (i.e. 117 fs) can induce a noninflamed TME by WNT/β-catenin signalling pathway activation because of the loss of function.
We previously reported that even when somatic mutations increased, tumour cells could escape antitumor immunity by increasing the activity of immunosuppressive signalling pathways such as the WNT/β-catenin pathway [28, 30]. We also reported that tumour cells evade antitumor immunity by mutating HLA genes and that driver gene mutations play significant roles in not only proliferation and survival but also antitumor immunity evasion [24, 27, 28]. Moreover, the present study demonstrates that some of the neoantigenic mutations, particularly driver mutations, can paradoxically act as antitumor immunity suppressors through the original gene functions. These observations could explain the weak correlation with the immune status of the TME and the inadequacy in predicting the effects of ICIs [24, 27, 28, 30].
Several reports have demonstrated the presence of RNF43 fs mutations in MSI-H colorectal cancer [32,33,34,35]. In addition, the relationship between RNF43 mutations and the WNT/β-catenin signalling pathway has been reported [34, 35, 45, 46]. However, these studies did not focus on antitumor immunity and PD-1 blockade-mediated efficacy based on the neoantigenic RNF43 mutations. Previous studies have reported that WNT/β-catenin signalling pathway activation induces a noninflamed TME [29, 30]; however, this study, to our knowledge, has for the first time demonstrated that neoantigenic RNF43 mutations (i.e. 117 fs) can induce the noninflamed TME by WNT/β-catenin signalling pathway activation because of the loss of function. These detailed analyses are limited to MSI-H cancer as there are few shared driver genes and more individualised mutations reported as candidate genes for neoantigens [8, 50]. In addition, there are few reports of common neoantigens in other cancer types [32], making such analyses difficult. Conversely, using TCGA dataset analyses, we demonstrated that passenger rather than driver gene mutations were related to the inflamed TME. Thus, even if such functional driver gene mutations become neoantigens, patients with these neoantigens could have a noninflamed TME because of gene functions. These findings suggest the need to evaluate the qualities as well as quantities of neoantigenic mutations. To validate these results, further research is warranted.
In summary, we identified “paradoxical neoantigenic mutations” that could induce a noninflamed TME owing to their original gene function, despite deriving neoantigens. We propose the need to assess the qualities as well as the quantities of neoantigenic gene mutations, particularly driver gene mutations, as predictive biomarkers for ICI response. Further studies using large cohorts are warranted to determine the clinical applicability of our results.
Data availability
The data that support the results of this study are available from the corresponding author, YT, upon reasonable request. Raw sequencing data were deposited in the Japanese Genotype-Phenotype Archive (http://trace.ddbj.nig.ac.jp/jga), which is hosted by the DNA Data Bank (https://doi.org/10.1093/nar/gku1120) of Japan, under accession number JGAS00000000113 (NBDC number: hum0094).
References
Dunn GP, Old LJ, Schreiber RD. The immunobiology of cancer immunosurveillance and immunoediting. Immunity 2004;21:137–48.
Schreiber RD, Old LJ, Smyth MJ. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 2011;331:1565–70.
Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJ, Robert L, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 2014;515:568–71.
Zou W, Wolchok JD, Chen L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci Transl Med. 2016;8:328rv4–rv4.
Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366:2455–65.
Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012;366:2443–54.
Kang Y-K, Boku N, Satoh T, Ryu M-H, Chao Y, Kato K, et al. Nivolumab in patients with advanced gastric or gastro-oesophageal junction cancer refractory to, or intolerant of, at least two previous chemotherapy regimens (ONO-4538-12, ATTRACTION-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;390:2461–71.
Schumacher TN, Schreiber RD. Neoantigens in cancer immunotherapy. Science 2015;348:69–74.
Matsushita H, Vesely MD, Koboldt DC, Rickert CG, Uppaluri R, Magrini VJ, et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 2012;482:400–4.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015;348:124–8.
Rooney Michael S, Shukla Sachet A, Wu Catherine J, Getz G, Hacohen N. Molecular and Genetic Properties of Tumors Associated with Local Immune Cytolytic Activity. Cell 2015;160:48–61.
Cristescu R, Mogg R, Ayers M, Albright A, Murphy E, Yearley J, et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science. 2018;362:eaar3593.
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, et al. Genetic Basis for Clinical Response to CTLA-4 Blockade in Melanoma. N Engl J Med. 2014;371:2189–99.
Mandal R, Samstein RM, Lee KW, Havel JJ, Wang H, Krishna C, et al. Genetic diversity of tumors with mismatch repair deficiency influences anti-PD-1 immunotherapy response. Science 2019;364:485–91.
Willis JA, Reyes-Uribe L, Chang K, Lipkin SM, Vilar E. Immune Activation in Mismatch Repair–Deficient Carcinogenesis: More Than Just Mutational Rate. Clin Cancer Res. 2020;26:11–7.
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N Engl J Med. 2015;372:2509–20.
McGrail DJ, Pilié PG, Rashid NU, Voorwerk L, Slagter M, Kok M, et al. High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann Oncol. 2021;32:661–72.
Voorwerk L, Slagter M, Horlings HM, Sikorska K, van de Vijver KK, de Maaker M, et al. Immune induction strategies in metastatic triple-negative breast cancer to enhance the sensitivity to PD-1 blockade: the TONIC trial. Nat Med. 2019;25:920–8.
Rizvi H, Sanchez-Vega F, La K, Chatila W, Jonsson P, Halpenny D, et al. Molecular Determinants of Response to Anti-Programmed Cell Death (PD)-1 and Anti-Programmed Death-Ligand 1 (PD-L1) Blockade in Patients With Non-Small-Cell Lung Cancer Profiled With Targeted Next-Generation Sequencing. J Clin Oncol. 2018;36:633–41.
Marabelle A, Le DT, Ascierto PA, Di Giacomo AM, De Jesus-Acosta A, Delord JP, et al. Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair-Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. J Clin Oncol. 2020;38:1–10.
Overman MJ, Lonardi S, Wong KYM, Lenz HJ, Gelsomino F, Aglietta M, et al. Durable Clinical Benefit With Nivolumab Plus Ipilimumab in DNA Mismatch Repair-Deficient/Microsatellite Instability-High Metastatic Colorectal Cancer. J Clin Oncol. 2018;36:773–9.
Milo I, Bedora-Faure M, Garcia Z, Thibaut R, Périé L, Shakhar G, et al. The immune system profoundly restricts intratumor genetic heterogeneity. Sci Immunol. 2018;3:eaat1435.
Montesion M, Murugesan K, Jin DX, Sharaf R, Sanchez N, Guria A, et al. Somatic HLA Class I Loss Is a Widespread Mechanism of Immune Evasion Which Refines the Use of Tumor Mutational Burden as a Biomarker of Checkpoint Inhibitor Response. Cancer Disco. 2021;11:282–92.
Kawazu M, Ueno T, Saeki K, Sax N, Togashi Y, Kaneseki T, et al. HLA Class I analysis provides insight into the genetic and epigenetic background of immune evasion in colorectal cancer with high microsatellite instability. Gastroenterology 2022;162:799–812.
Keenan TE, Burke KP, Van Allen EM. Genomic correlates of response to immune checkpoint blockade. Nat Med. 2019;25:389–402.
Spranger S, Gajewski TF. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat Rev Cancer. 2018;18:139–47.
Sugiyama E, Togashi Y, Takeuchi Y, Shinya S, Tada Y, Kataoka K, et al. Blockade of EGFR improves responsiveness to PD-1 blockade in EGFR-mutated non-small cell lung cancer. Sci Immunol. 2020;5:eaav3937.
Kumagai S, Togashi Y, Sakai C, Kawazoe A, Kawazu M, Ueno T, et al. An Oncogenic Alteration Creates a Microenvironment that Promotes Tumor Progression by Conferring a Metabolic Advantage to Regulatory T Cells. Immunity 2020;53:187–203.e8.
Spranger S, Bao R, Gajewski TF. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 2015;523:231–5.
Takeuchi Y, Tanegashima T, Sato E, Irie T, Sai A, Itahashi K, et al. Highly immunogenic cancer cells require activation of the WNT pathway for immunological escape. Sci Immunol. 2021;6:eabc6424.
Sato K, Kawazu M, Yamamoto Y, Ueno T, Kojima S, Nagae G, et al. Fusion Kinases Identified by Genomic Analyses of Sporadic Microsatellite Instability–High Colorectal Cancers. Clin Cancer Res. 2019;25:378–89.
Roudko V, Bozkus CC, Orfanelli T, McClain CB, Carr C, O’Donnell T, et al. Shared Immunogenic Poly-Epitope Frameshift Mutations in Microsatellite Unstable Tumors. Cell 2020;183:1634–49.e17.
Seeber A, Battaglin F, Zimmer K, Kocher F, Baca Y, Xiu J, et al. Comprehensive Analysis of R-Spondin Fusions and RNF43 Mutations Implicate Novel Therapeutic Options in Colorectal Cancer. Clin Cancer Res. 2022;28:1863–70.
Li S, Lavrijsen M, Bakker A, Magierowski M, Magierowska K, Liu P, et al. Commonly observed RNF43 mutations retain functionality in attenuating Wnt/β-catenin signaling and unlikely confer Wnt-dependency onto colorectal cancers. Oncogene 2020;39:3458–72.
Yamamoto D, Oshima H, Wang D, Takeda H, Kita K, Lei X, et al. Characterization of RNF43 frameshift mutations that drive Wnt ligand- and R-spondin-dependent colon cancer. J Pathol. 2022;257:39–52.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–60.
Jurtz V, Paul S, Andreatta M, Marcatili P, Peters B, Nielsen M. NetMHCpan-4.0: Improved Peptide-MHC Class I Interaction Predictions Integrating Eluted Ligand and Peptide Binding Affinity Data. J Immunol. 2017;199:3360–8.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Consortium TU. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 2020;49:D480–D9.
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, et al. Pan-cancer Immunogenomic Analyses Reveal Genotype-Immunophenotype Relationships and Predictors of Response to Checkpoint Blockade. Cell Rep. 2017;18:248–62.
Tate JG, Bamford S, Jubb HC, Sondka Z, Beare DM, Bindal N, et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 2018;47:D941–D7.
Tsukiyama T, Fukui A, Terai S, Fujioka Y, Shinada K, Takahashi H, et al. Molecular Role of RNF43 in Canonical and Noncanonical Wnt Signaling. Mol Cell Biol. 2015;35:2007–23.
Thornton AM, Fang L, Lo A, McSharry M, Haan D, O’Brien C, et al. eVIP2: Expression-based variant impact phenotyping to predict the function of gene variants. PLoS Comput Biol. 2021;17:e1009132.
Tu J, Park S, Yu W, Zhang S, Wu L, Carmon K, et al. The most common RNF43 mutant G659Vfs*41 is fully functional in inhibiting Wnt signaling and unlikely to play a role in tumorigenesis. Sci Rep. 2019;9:18557.
Muramatsu T, Noguchi T, Sugiyama D, Kanada Y, Fujimaki K, Ito S, et al. Newly emerged immunogenic neoantigens in established tumors enable hosts to regain immunosurveillance in a T-cell-dependent manner. Int Immunol. 2021;33:39–48.
Nishikawa H, Sato E, Briones G, Chen LM, Matsuo M, Nagata Y, et al. In vivo antigen delivery by a Salmonella typhimurium type III secretion system for therapeutic cancer vaccines. J Clin Investig. 2006;116:1946–54.
Zheng N, Wang P, Jeffrey PD, Pavletich NP. Structure of a c-Cbl-UbcH7 complex: RING domain function in ubiquitin-protein ligases. Cell 2000;102:533–9.
Miao D, Margolis CA, Vokes NI, Liu D, Taylor-Weiner A, Wankowicz SM, et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat Genet. 2018;50:1271–81.
Acknowledgements
We wish to thank Yoshitaka Hippo for useful discussions. We would like to thank Noriko Sakurai, Joji Nagasaki and Takao Morinaga for their technical assistance. The results shown here are in part based upon data generated by the TCGA Research Network (https://www.cancer.gov/tcga). We created our graphical abstract with BioRender.com (Toronto, Canada). We would like to thank Editage (www.editage.com) for English language editing.
Funding
This study was supported by Grants-in-Aid for Scientific Research [B grant no. 21H02772 (MK) and 20H03694 (YT)], Challenging Exploratory Research [no. 22K1945904 (YT)], and Research Activity Start-up no. 21K20859 (TI)] from the Japan Society for the Promotion of Science (JSPS); the Project for Cancer Research and Therapeutic Evolution [no. 21cm0106502 (MK) and no. 21cm0106383 (YT)]; Practical Research for Innovative Cancer Control [no. 19ck0106521h0001 (YT) and 22ck0106723h0001 (MK and YT)] from the Japan Agency for Medical Research and Development (AMED); the Fusion Oriented Research for disruptive Science and Technology [FOREST, no. 21-211033868 (YT)] from Japan Science and Technology Agency (JST); the Chiba Prefecture Research Grant (MK and YT); the Takeda Science Foundation (YT); the Mochida Memorial Foundation (YT); the Japanese Foundation for Multidisciplinary Treatment of Cancer Foundation (YT); the KANAE Foundation for the Promotion of Medical Science (YT); the Yasuda Memorial Foundation for Medicine (YT); the MSD Life Science Foundation (YT); the Kowa Life Science Foundation (YT); the Senri Life Science Foundation (YT); the Uehara Memorial Foundation (YT); the GSK Japan foundation (YT); the Ono Medical Research Foundation (YT); the Inamori Foundation (YT); the Astellas Foundation for Research on Metabolic Disorders (YT); the Japan Respiratory Foundation (YT); the Kato Memorial Bioscience Foundation (YT); the Naito Foundation (YT); Research Grant of the Princess Takamatsu Cancer Research Fund [no. 21-25329 (YT)]; the Ube Industries Foundation (YT); The Wesco Foundation (YT); the Pharmacology Research Foundation (YT). Open access funding provided by Okayama University.
Author information
Authors and Affiliations
Contributions
Conception and design: YT. Development of the methodology: TI, SK, ET, MK and YT Collection of clinical samples and data: SI, TU and MK. Acquisition of experimental data: TI, SK, ET, YU and MK. Writing, review, and/or revision of the paper: TI, MK and YT. Study supervision: SO, HM and NK. All authors read and approved the final paper. YT and MK act as guarantor for this study.
Corresponding authors
Ethics declarations
Competing interests
SO received honoraria from Bayer, Eisai, Eli Lilly and Chugai Pharmaceutical, and research grants from Bayer, Eisai, Eli Lilly, Chugai Pharmaceutical and AstraZeneca. NK received honoraria from MSD, Bristol-Myers Squibb Company, Chugai Pharmaceutical, and AstraZeneca, and a research grant from Chugai Pharmaceutical. YT has received honoraria from Ono Pharmaceutical, Bristol-Myers Squibb, Chugai Pharmaceutical, and MSD and research grants from Ono Pharmaceutical, Bristol-Myers Squibb, Daiichi-Sankyo, KOTAI Biotechnologies, and KORTUC outside of this study. All other authors declare that they have no competing financial interests.
Ethics approval and consent to participate
The clinical protocol for this study was approved by the appropriate institutional review boards and ethics committees at the National Cancer Center (#2015-202) and the University of Tokyo (G10063 and G10094). The study was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained prior to participation. Mouse experiments were approved by the Animal Committee for Animal Experimentation of the Chiba Cancer Center (21–3) and Okayama University (629). All experiments met the U.S. Public Health Service Policy on Humane Care and Use of Laboratory Animals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ishino, T., Kawashima, S., Tanji, E. et al. Somatic mutations can induce a noninflamed tumour microenvironment via their original gene functions, despite deriving neoantigens. Br J Cancer 128, 1166–1175 (2023). https://doi.org/10.1038/s41416-023-02165-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02165-6
This article is cited by
-
“Paradoxical neoantigenic mutations”: not simply a driver vs passenger dualism
British Journal of Cancer (2023)