Abstract
Background
Ovarian cancer patients frequently develop chemotherapy resistance, limiting treatment options. We have previously shown that individuality in fibroblast growth factor 1 (FGF1) expression influences survival and chemotherapy response.
Methods
We used MTT assays to assess chemosensitivity to cisplatin and carboplatin following shRNA-mediated knockdown or heterologous over-expression of FGF1 (quantified by qRT-PCR and immunoblot analysis), and in combination with the FGFR inhibitors AZD4547 and SU5402, the ATM inhibitor KU55933 and DNA-PK inhibitor NU7026. Immunofluorescence microscopy was used to quantify the FGF1-dependent timecourse of replication protein A (RPA) and γH2AX foci formation.
Results
Pharmacological inhibition of FGF signalling reversed drug resistance in immortalised cell lines and in primary cell lines from drug-resistant ovarian cancer patients, while FGF1 over-expression induced resistance. Ataxia telangiectasia mutated (ATM) phosphorylation, but not DNA adduct formation was FGF1 dependent, following cisplatin or carboplatin challenge. Combining platinum drugs with the ATM inhibitor KU55933, but not with the DNA-PK inhibitor NU7026 re-sensitised resistant cells. FGF1 expression influenced the timecourse of damage-induced RPA and γH2AX nuclear foci formation.
Conclusion
Drug resistance arises from FGF1-mediated differential activation of high-fidelity homologous recombination DNA damage repair. FGFR and ATM inhibitors reverse platinum drug resistance, highlighting novel combination chemotherapy approaches for future clinical trial evaluation.
Similar content being viewed by others
Background
Ovarian cancer, the most deadly gynaecological malignancy, frequently presents when already advanced and adjuvant or neo-adjuvant chemotherapy combining cisplatin or, more frequently, carboplatin with paclitaxel is the most appropriate treatment [1]. Although many patients initially respond well, the development of treatment-limiting drug resistance is unfortunately inevitable in the majority of patients [2, 3]. There is therefore an urgent need to understand the molecular mechanisms underpinning drug resistance, and to use this information to develop clinical response biomarkers and propose novel combination chemotherapy regimens.
We have previously shown that individuality in fibroblast growth factor 1 (FGF1) expression significantly inversely influences both progression-free and overall survival in ovarian cancer patients [4]. FGF1 expression is significantly increased in cisplatin-resistant cell line models, where FGF1 knockdown in resistant A2780DPP cells re-sensitised to both cisplatin and carboplatin [4]. Consistent with our findings, others have confirmed that FGF1 is more highly expressed in platinum-resistant compared to drug-sensitive ovarian cancers [5]. FGF1 is one of 22 fibroblast growth factors, related by sequence conservation and structural similarity, the majority of which drive autocrine signalling by binding to FGF receptors (FGFR1-4) to induce receptor tyrosine kinase signalling cascades. These intracellular cascades are activated by phosphorylation of the adaptor protein FGFR Substrate 2α (FRS2α) following CRK-like proto-oncogene recruitment to FGFR, which drives signalling through the PLCγ, MEKK, ERK1/2 and AKT pathways [6,7,8,9,10,11,12,13]. Dysregulated fibroblast growth factor signalling is frequently observed in many cancers [14]; however, the molecular mechanisms rationalising a role for the FGF signalling pathway in promoting drug resistance remain poorly characterised.
Cisplatin and carboplatin are actively transported into the cell by the copper transporter CTR1 [15,16,17]. Intracellular translocation results in spontaneous drug aquation [18]—both drugs then promote DNA damage induced by the formation of DNA interstrand cross-links that exceed cellular repair capacity [19]. This in turn forces cells to introduce DNA double-strand breaks (DSBs) [20], producing genomic instability. As maintenance of genome stability is fundamental to eukaryotic cell survival [21], cells have evolved highly conserved networks of DNA damage repair mechanisms [22]. DSBs are most commonly repaired by low-fidelity non-homologous end joining (NHEJ) or high-fidelity homologous recombination (HR) [23, 24]. NHEJ promotes direct ligation of broken DNA ends, whereas HR involves the replacement of damaged DNA using template sister chromatid sequences and is an essential defence against genome instability [25, 26]. Alterations to DNA damage repair mechanisms commonly induce resistance to DNA damaging drugs, including cisplatin and carboplatin [27]. For example, delayed engagement of HR results in toxic chromatid fusions [28], while compromised HR, for example as a result of inherited BRCA1 or BRCA2 mutations, confers sensitivity to poly-ADP ribose polymerase inhibitors including Olaparib [29, 30], and increases sensitivity to cisplatin and carboplatin chemotherapy in ovarian cancer patients [31].
An early step in HR is the focal recruitment of replication protein A (RPA), which protects 3’ single-stranded DNA overhangs from degradation [32] and promotes recruitment of RAD51 recombinase that induces sister chromatid invasion to facilitate homology-directed error-free repair [33]. HR-specific proteins are activated at DSB sites following the phosphorylation of the central regulator of DSB repair, ataxia telangiectasia mutated (ATM) [34,35,36]. ATM is an important regulator of DSB repair that phosphorylates multiple substrates [34], including itself at Ser1981 to induce its activation [35], and H2AX at Ser139 [36] to signal the presence of DSBs and instigate focal recruitment of crucial DNA damage repair proteins involved in homology-directed DSB repair [37].
Confirmation of a mechanistic link between FGF signalling and the DNA damage response would not only rationalise the influence of individuality in FGF1 expression on disease progression and chemotherapy response in ovarian cancer patients, but may additionally identify candidate resistance biomarkers and novel combination chemotherapy/resistance pathway inhibitor approaches for future clinical trial evaluation.
Methods
Cell culture
The A2780 cell line (chemosensitive, derived from an untreated ovarian cancer patient [38]), cisplatin-resistant derivative A2780DPP (A2780Cis [39]), and the SK-OV-3 cell line (platinum drug-sensitive ovarian adenocarcinoma cell line [40]) were purchased from the European Collection of Authenticated Cell Cultures. The CaOV3 cell line (platinum drug-sensitive epithelial ovarian cancer cell line derived from tumour tissue of a patient with adenocarcinoma of the ovary [41]) was a gift from Dr Jozien Helleman, Erasmus University, Netherlands. Cells were maintained in RPMI1640 medium (Gibco, Renfrewshire, UK) supplemented with 10% v/v foetal bovine serum (FBS) (Gibco, Renfrewshire, UK) and 1% v/v penicillin/streptomycin (Sigma Aldrich, Dorset, UK), with the addition of 1 µM cisplatin (Sigma Aldrich, Dorset, UK) to A2780DPP cells on every third passage. Further information summarising the genetic background of our ovarian cancer cell line panel is summarised in Supplementary Fig. 1. Human embryonic kidney 293T cells were a gift from Dr Laureano de la Vega, University of Dundee, and were maintained in DMEM medium (Gibco, Renfrewshire, UK) supplemented with 10% v/v FBS. All cell lines were maintained in 37 °C incubators supplemented with 5% CO2. Cell lines were authenticated by short tandem repeat profiling (Centre for Life, Newcastle, UK) and routinely tested and found to be negative for mycoplasma contamination (Lonza Biologics, Slough, UK).
Cell culture of patient-derived cells
Patient ascites samples were collected following paracentesis and mixed 1:1 with ascites media (1:1 MCDB (Sigma Aldrich, Dorset, UK)/Media 199 (Gibco, Renfrewshire, UK), 10% v/v FBS, 1% v/v penicillin/streptomycin (Sigma Aldrich, Dorset, UK)) [42]. Cells were incubated for 3–5 days before differential trypsinisation (cells incubated in 1 mL trypsin for 30 s) to limit fibroblast contamination, then incubated for an additional 3 days before use in subsequent experiments.
Lentiviral shRNA-mediated FGF1 knockdown
shRNA constructs were prepared from bacterial glycerol stocks using midi-prep kits (Qiagen, Manchester, UK) according to the manufacturers’ instructions. 3 × 106 HEK293T cells were transfected in 10 cm dishes using 5 µL Lipofectamine 2000 (Invitrogen, Renfrewshire, UK) with 5 µg of shRNA construct, one of several constructs previously evaluated to target FGF1 (Sigma Aldrich, Dorset, UK; TRCN0000072524) [4], together with 3.2 µg psPAX2 packaging and 1.8 µg pMD2.G enveloping vectors (a gift from Dr Rita Moreno, University of Dundee) [43] in 3 mL Opti-MEM serum-free medium (Gibco) with 7 mL DMEM for 24 h before virus-containing supernatant was passed through a 0.45 µm filter (VWR, Leicestershire, UK). 2.5 × 105 A2780DPP target cells seeded 24 h previously in six-well plates were then cultured for 24 h in virus-containing medium supplemented with 200 µg/mL hexadimethrine bromide (Sigma Aldrich, Dorset, UK), and transduced cells selected by 24 h incubation in RPMI supplemented with 2 µg/mL puromycin to create the A2780DPP FGFKD cell line (Gibco, Renfrewshire, UK). A control cell line (A2780DPP EV) was similarly created, following transfection with the construct pLKO.1.
siRNA-mediated ATM knockdown
2.5 × 105 A2780DPP cells were seeded in six-well plates and incubated for 24 h before transient transfection using 5 µL Lipofectamine RNAiMAX (Thermo Fisher, Renfrewshire, UK) according to the manufacturers’ instructions with a final concentration of 5, 10 or 20 nM SMARTpool ON-TARGETplus ATM-targeted siRNA (L-003201-00-0005) or negative control non-targeted siRNA (D-001810-10-05, Horizon Discovery, Cambridge, UK) in Opti-MEM serum-free medium (Gibco, Renfrewshire, UK).
FGF1 over-expression by plasmid transfection
2.5 × 105 A2780 cells were seeded in six-well plates and pre-incubated for 24 h before transfection with either 1 µg pCMV3.1-EGFP-FGF1 plasmid (Sino Biological, Beijing, China) or 1 µg pCMV3.1-EGFP control plasmid (Sino Biological, Beijing, China) using 5 µL Lipofectamine 2000 (Invitrogen, Renfrewshire, UK) in 2 mL Opti-MEM reduced serum medium (Gibco, Renfrewshire, UK). Cells were incubated for 24 h in transfection mix and then selected with 5 µg/mL hygromycin (Gibco, Renfrewshire, UK) for 120 h.
RNA extraction
Total RNA was extracted from 2.5 × 105 cells using RNeasy Mini Kits (Qiagen, Manchester, UK), following the manufacturers’ guidelines, including an on-column DNase digestion (RNAs free DNase Kit, Qiagen, Manchester, UK). RNA yield and integrity were confirmed using a Nanodrop ND1000 spectrophotometer (Thermo Fisher, Renfrewshire, UK).
qRT-PCR analysis
RNA was reverse transcribed into cDNA (4 ng/µL final concentration of RNA) using TaqMan Reverse Transcription Reagents Kit (Thermo Fisher, Renfrewshire, UK) according to the manufacturer’s instructions, replacing oligo dT with random hexamers. FGF1 (Hs01092738_m1), ATM (Hs00175892_m1) and 18S ribosomal RNA expression (438839) was assessed in individual 20 µL reactions, combining 10 µL TaqMan universal master mix (Thermo Fisher, Renfrewshire, UK), 1 µL gene-specific probe, 1 µL cDNA and 8 µL nuclease-free water. Each reaction was performed in triplicate and run on the standard PCR programme (50 °C 2 min, 95 °C 10 min, and 40 cycles of 95 °C 15 s, 60 °C 1 min) on a QuantStudio5 qRT-PCR instrument (Thermo Fisher, Renfrewshire, UK). Baseline and threshold values were calculated automatically, and gene expression was quantified by cycle threshold (Ct) values, with relative gene expression comparing the Ct change in target gene and 18S rRNA control (ΔCt), as previously described [44]. Compound errors (s) were calculated using the formula s = ((standard deviation target gene)2 + (standard deviation 18S rRNA)2)½ (https://assets.thermofisher.com/TFS-Assets/LSG/manuals/cms_042380.pdf).
Immunoblot analysis
2.5 × 105 cells were seeded in six-well plates and incubated for 24 h prior to harvest or treatment with either an acute cisplatin challenge (3 µM) (Sigma Aldrich, prepared as a 2 mg/mL stock in sterile water), the FGFR inhibitors AZD4547 (ApexBio, Cambridge, UK, prepared as a 10 mM stock in DMSO) or SU5402 (Sigma Aldrich, Dorset, UK, prepared as a 10 mM stock in DMSO), or transfection as described above. Plates were placed on ice and cells were washed with ice-cold phosphate-buffered saline. Protein extracts were prepared in RIPA buffer (50 mM Tris-HCl (pH8), 150 mM NaCl, 0.1% SDS, 0.1% sodium deoxycholate, 1% NP-40, 2 mM EDTA, 2.1 g/L NaF), supplemented with EDTA-free protease inhibitor cocktail tablet (Sigma Aldrich, Dorset, UK) using a cell scraper. Lysates were collected following centrifugation at 2000 × g for 10 min at 4 °C, and protein concentrations were determined using the DC protein assay (BioRad, Hertfordshire, UK). Samples were diluted to 1 µg/µL with RIPA buffer, mixed with 10× sample reducing agent (500 mM dithiothreitol, Thermo Fisher, Renfrewshire, UK) and 4× BOLT sample buffer (Thermo Fisher, Renfrewshire, UK) and denatured at 100 °C for 10 min. Proteins were separated by electrophoresis in 12% lithium-dodecyl sulphate gels in Tris-glycine buffer (25 mM Tris pH8.3, 192 mM glycine, 0.1% SDS) with the addition of 500 µL NuPage antioxidant (Thermo Fisher, Renfrewshire, UK), then transferred to 0.45-µm-pore nitrocellulose membranes (GE Healthcare, Bucks, UK) in Tris-glycine-methanol buffer (25 mM Tris, 192 mM glycine, 20% methanol). Non-specific antibody binding was blocked by incubation in 5% non-fat dried milk in TBS-T (50 mM Tris (pH 7.9), 150 mM NaCl, 1% Tween-20), and membranes incubated overnight at 4 °C with either goat polyclonal anti-FGF1 (AF232, R&D Systems, Oxfordshire, UK, diluted 1:1000), rabbit polyclonal anti-GFP (G1544, Sigma Aldrich, diluted 1:2000) or anti-FRS2a (pY196) (PA5-64616 Thermo Fisher, diluted 1:1000), mouse monoclonal anti-FRS2α (MAB4-69, R&D Systems, diluted 1:1000), anti-β−Actin (sc-47778, Santa Cruz Biotechnology, Heidelberg, Germany, diluted 1:1000), or anti-γH2AX (05-636, Merck Life Science, Darmstadt, Germany), or rabbit monoclonal anti-ATM (S1981) (5883T, Cell Signaling Technology, Danvers, MA, USA, diluted 1:500), or anti-ATM (2873T, Cell Signaling Technology, diluted 1:1000). Membranes were washed in TBS-T and incubated for 1 h in HRP-conjugated rabbit anti-goat polyclonal secondary antibody (FGF1; 170-1034, BioRad, diluted 1:2000), HRP-conjugated goat anti-mouse polyclonal secondary antibody (FRS2α, β−Actin, γH2AX; 170-6516, BioRad, diluted 1:2000), or HRP-conjugated goat anti-rabbit polyclonal secondary antibody (GFP, FRS2α (Y196), ATM (S1981), ATM; 170-6515, BioRad, diluted 1:2000). Immunoblots were developed using Amersham enhanced chemiluminescence kit (GE Healthcare, Buckinghamshire, UK) using Amersham high-performance Hyperfilms (GE Healthcare, Buckinghamshire, UK).
Slot blot analysis
2 × 105 cells/well were seeded in six-well plates and incubated overnight. Cells were synchronised by treatment with 2 mM thymidine for 24 h before treatment with 3 µM cisplatin or 10 µM carboplatin for 3 h, harvested by trypsinisation and genomic DNA extracted using GeneJet Genomic DNA purification kit (Thermo Fisher, Renfrewshire, UK) according to the manufacturers’ instructions. DNA was diluted to 100 ng/150 µL in 0.4 M NaOH/10 mM EDTA and boiled at 100 °C for 10 min before immediate incubation on ice and the addition of ice-cold 150 µL 2 M NH4CH3CO2. DNA samples were bound to Protran nitrocellulose membranes (BioRad, Hertfordshire, UK) (pre-incubated in 0.9 M NaCl, 90 mM sodium citrate, pH 7.0) by vacuum manifold filtration and membranes washed with 0.3 M NaCl, 30 mM sodium citrate, air-dried for 4 h at room temperature then rehydrated in PBS-0.1% Tween, with non-specific antigen binding blocked with 5% BSA, 0.05% w/v NaN3. Membranes were incubated in anti-platinum-modified DNA Ab (1:1000; Abcam, Cambridge, UK) in 5% BSA, 0.05% w/v NaN3 at 4 °C overnight, washed in PBS-0.1% Tween, and incubated in anti-Rat IgG HRP, 1:2000, 5% BSA, 0.05% w/v NaN3, (BioRad, Herts, UK) for 1 h. Membranes were washed in PBS-0.1% Tween and slot blots were developed using Amersham enhanced chemiluminescence kit (GE Healthcare, Buckinghamshire, UK) using Amersham high-performance Hyperfilms (GE Healthcare, Buckinghamshire, UK).
In vitro chemosensitivity assays
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assays [45] were used to compare the chemosensitivity of cell lines. Cells were seeded in 96-well plates (5000 cells per well in 100 µL media). Cells were untreated or treated in triplicate with vehicle or with serial dilutions of cisplatin or carboplatin at concentrations relevant to typical ovarian cancer patient peak plasma levels (range 0–200%; cisplatin (0–25.33 µM), carboplatin (0–85.12 µM) [46], and in combination with FGFR, ATM or DNA-PK inhibitors (10 µM SU5402, AZD4547; 10 µM KU55933, 2 µM NU7026). Following 72 h (immortalised cell lines, as previously described [4, 45]) or 144 h (to allow for slower growth of primary patient-derived cells), media and drugs were removed and 100 µL 0.5 mg/mL MTT (Alfa Aesar, Lancashire, UK) solution (MTT in phenol-free RPMI, Gibco, Renfrewshire, UK) added and incubated for 3 h at 37 °C, 5% CO2. MTT solution was removed, and formazan crystals formed by viable cells were solubilised in 100 µL DMSO and quantified by absorbance at 570 nm. The percentage of viable cells remaining following drug treatment was calculated as a percentage of vehicle-treated control cells and associated IC50 values were calculated from log dose-response curves using Prism 9 software (GraphPad Software, Inc., La Jolla, CA, USA).
Immunofluorescence microscopy
Immunofluorescence microscopy was used to assess Pt-DNA adduct formation in ovarian cancer cells with different levels of FGF1 expression and HR-specific nuclear foci formation in response to cisplatin-induced DNA damage. 2 × 106 cells were seeded per well of a six-well plate containing a #1.5 glass coverslip ø22mm (VWR, Leicestershire, UK) and incubated overnight. Cells were synchronised with 2 mM thymidine (Alfa Aesar, Lancashire, UK) for 24 h and, following thymidine washout, treated for 3 h with 3 µM cisplatin, then fixed immediately to assess adduct formation or washed in pre-warmed PBS and incubated over a 24 h (RPA, RAD51) or 36 h (γH2AX) timecourse with cells fixed at appropriate timepoints. To fix cells, coverslips were washed in PBS, incubated for 10 min in 4% paraformaldehyde/NaOH (pH 7.2), then washed in PBS and permeabilised in KCM buffer (120 mM KCl, 20 mM NaCl, 10 mM Tris-HCl, pH 7.5, 0.1% v/v Triton-X100) for 10 min. Non-specific antigen binding was blocked by incubation in blocking buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 2% w/v BSA, 2% w/v fish gelatine, 0.1% Triton-X100) for 20 min at 37 °C, prior to incubation in primary antibody (cisplatin-modified DNA, rat monoclonal CP9/19, ab103261, Abcam (Cambridge, UK), diluted 1:1000; RPA, mouse monoclonal, RPA34-19, NA18 Calbiochem (Dorset, UK), diluted 1:1000; RAD51, rabbit monoclonal, EPR4030 (3) ab133534, Abcam, Cambridge, UK, diluted 1:1000; γH2AX, mouse monoclonal JBW301, 05-636, Merck (Dorset, UK), diluted 1:1000) in blocking buffer at 4 °C overnight in a damp chamber. Coverslips were washed in PBS-T (PBS, 0.1% Tween-20) and then incubated in secondary AlexaFluor-conjugated antibody (cisplatin-modified DNA, donkey anti-rat AlexaFluor488 A21208, Thermo Fisher; RPA and γH2AX goat anti-mouse AlexaFluor488 A11001, Thermo Fisher; RAD51 recombinase, donkey anti-rabbit AlexaFluor594 R37119, Thermo Fisher; all 1:2000) and DAPI (1:3000) in blocking buffer for 2 h in a damp chamber at RT. Coverslips were washed in PBS-T, air-dried for 3 h and mounted to SuperFrostPlus microscope slides (VWR, Leicestershire, UK) with the addition of 5 µL Prolong Gold anti-fade reagent (Thermo Fisher, Renfrewshire, UK). Confocal microscopy was performed on each coverslip using an LSM-710 microscope (Carl Zeiss, Oberkochen, Germany), using an inverted lens at ×40 magnification using oil immersion. Images were taken in Z-stacks where the maximum intensity projection was used for subsequent image analysis in OMERO insight (University of Dundee, UK), with a minimum of 40 cells counted in each experiment. Nuclear foci were quantified using Fiji Image J v2.0 open-source image processing package using Duke University Protocol (Durham, NC, USA) and data processed using Prism 9 software.
Statistical analysis
Statistical comparisons of data were performed using two-tailed Student’s t-tests for pairwise comparisons or one-way analysis of variance for multiple comparisons using GraphPad Prism 9, with p values <0.05 considered to represent statistical significance (*p < 0.05, **p < 0.01, ***p < 0.001).
Results
FGF1 influences sensitivity to cisplatin and carboplatin in ovarian cancer cells
To extend our previous analysis [4], we used lentivirus to create an A2780DPP derivative cell line with shRNA-mediated stable FGF1 knockdown (A2780DPP FGF1KD). FGF1 knockdown (274.4-fold change in expression, p = 0.004) was confirmed by qRT-PCR (Fig. 1a) and immunoblot (Fig. 1b) analysis and was stable for more than 20 passages (data not shown). Loss of FGF1 expression in A2780DPP cells again re-sensitised drug-resistant cells to cisplatin (3.84-fold change in IC50, p = 0.007, Fig. 1c) and carboplatin (6.9-fold change in IC50, p = 0.002, Fig. 1d), assessed by MTT assay. We further confirmed that FGF1 knockdown influenced colony formation following cisplatin and carboplatin challenge by clonogenicity assay (Supplementary Fig. 2).
A2780DPP cells were lentivirally transduced with an shRNA construct targeting FGF1 or an empty vector control and FGF1 expression assessed using a qRT-PCR analysis and b immunoblot analysis (EV empty vector). MTT assays were used to assess the effect of FGF1 knockdown on sensitivity to c cisplatin (0–25.33 µM) and d carboplatin (0–85.12 µM). FGF1 was overexpressed in A2780 cells using an FGF1-EGFP fusion plasmid and e immunoblot analysis and f green fluorescence imaging used to confirm FGF1 expression. MTT assays were used to assess the influence of FGF1 over-expression on sensitivity to g cisplatin (0–25.33 µM) and h carboplatin (0–85.12 µM). Results represent three independent experiments. Pairwise comparisons of mean IC50 values and relative gene expression were performed using Student’s t-tests. Scale bar = 100 µm. *p < 0.05, **p < 0.01, ***p < 0.001.
To complement these experiments, FGF1 expression was increased in drug-sensitive A2780 cells by transfection with a dual GFP/FGF1 expression plasmid. Increased expression of both FGF1 and GFP was confirmed by immunoblot analysis (Fig. 1e) and green fluorescence microscopy (Fig. 1f). As predicted, increased FGF1 expression induced cisplatin (3.54-fold change in IC50, p < 0.001, Fig. 1g) and carboplatin (3.29-fold change in IC50, p = 0.002, Fig. 1h) resistance in A2780 cells. Similarly, resistance to cisplatin and carboplatin was induced by heterologous FGF1 expression in chemonaive SK-OV-3 and CaOV3 ovarian cancer cells (Supplementary Fig. 3).
FGF receptor inhibition re-sensitises A2780DPP cells to cisplatin and carboplatin
To investigate whether pharmacological inhibition of the FGF signalling pathway in A2780DPP cells recapitulates FGF1 knockdown, the Type I inhibitors AZD4547 and SU5402 which inhibit all four FGF receptors were used. Treatment of A2780DPP cells with AZD4547 (Fig. 2a) and SU5402 (Fig. 2b) blocked FGFR signalling in a concentration-dependent manner, confirmed using immunoblot analysis to assess phosphorylation of FGF Receptor Substrate 2α (FRS2α, Y196) as a surrogate for pathway activity [13]. Co-treatment with AZD4547 re-sensitised A2780DPP cells to cisplatin (fold change IC50 = 2.32, p < 0.001, Fig. 2c) and carboplatin (fold change IC50 = 8.05, p = 0.004, Fig. 2d), as did co-treatment with SU5402 with cisplatin (fold change IC50 = 13.42, p = 0.004, Fig. 2e) and carboplatin (fold change IC50 = 3.19, p = 0.006, Fig. 2f), assessed by MTT assays. Neither inhibitor was independently toxic at the concentration used (data not shown).
Immunoblot analysis was used to assess blockade of the FGF signalling pathway in A2780DPP cells in response to a AZD4547 (0–10 µM) and b SU5402 (0–10 µM). MTT assays were used to investigate the viability of A2780DPP cells following co-treatment with 10 µM AZD4547 or 10 µM SU5402 and c, d cisplatin (0–25.33 µM) and e, f carboplatin (0–85.12 µM). Results represent three independent experiments. Pairwise comparisons of mean IC50 values were calculated by Student’s t-tests. *p < 0.05, **p < 0.01, ***p < 0.001. Ns not significant.
FGFR inhibitors reverse drug resistance in ascites-derived primary cell lines
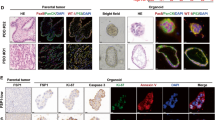
The Dundee Ovarian Cancer Study (DOCS) allows us to compare chemosensitivity and investigate associated resistance mechanisms in primary ascites-derived cell lines from chemotherapy naive drug-sensitive and matched drug-resistant ovarian cancer patients (Fig. 3a). To investigate whether FGFR inhibition also influenced cisplatin and carboplatin sensitivity in clinical samples in which the expression of both FGF1 and FGFR2 had been confirmed by qRT-PCR analysis (Fig. 3b), MTT assays were again used to assess chemosensitivity in two matched sample pairs before and after treatment with AZD4547, an ATP-competitive FGFR inhibitor, which was less toxic than SU5402 in previous clinical trials [47]. Both patients were initially clinically sensitive to cisplatin and carboplatin at disease presentation and relapsed with treatment-resistant disease. Although the primary ascites-derived cell line from Patient 1 showed higher expression of both FGF1 and FGFR2 than Patient 2, 10 µM AZD4547 in combination with either cisplatin (Fig. 3c, e) or carboplatin (Fig. 3d, f) again re-sensitised both drug-resistant cell lines—Patient 1 cisplatin fold change IC50 = 47.17, p = 0.006 and carboplatin fold change IC50 > 2, and Patient 2 cisplatin fold change IC50 = 14.13, p < 0.001 and carboplatin fold change IC50 > 2.17.
a The Dundee Ovarian Cancer Study; ‘A1’ sample = chemonaïve, ‘A2’ sample = disease relapse. b FGF1 and FGFR2 gene expression in chemoresistant ovarian cancer patient-derived cells was assessed by qRT-PCR analysis. MTT assays were used to compare viability of chemonaive cells, and cells from drug-resistant relapse samples from two patients in the presence or absence of combination treatments with 10 µM AZD4547 and c, e cisplatin (0–25.33 µM) or d, f carboplatin (0–85.12 µM). Pairwise comparisons of mean IC50 values and relative gene expression were calculated by Student’s t-tests. *p < 0.05, **p < 0.01, ***p < 0.001. Ns not significant.
Platinum-induced DNA adduct formation is independent of FGF1 expression
Following CTR1-mediated influx, cisplatin and carboplatin kill cells principally by crosslinking to and forming adducts between opposing DNA strands [15,16,17, 19]. CTR1 expression was not significantly different in A2780 and A2780DPP cells, and was not FGF1-dependent (data not shown). We then used immunofluorescence to quantify cis-[Pt(NH3)2]2+-DNA adduct formation in synchronised cells with different levels of FGF1 expression following 3 μM cisplatin or carboplatin challenge, using an antibody which detects platinum-modified DNA structures (Fig. 4a). FGF1 expression did not significantly influence cisplatin or carboplatin-induced DNA adduct formation, assessed both by immunofluorescence (Fig. 4b, c) and by slot blot analysis (Fig. 4d).
Cells were thymidine synchronised (2 mM, 24 h) prior to cisplatin challenge (3 µM cisplatin for 3 h). Immunofluorescence microscopy was used to detect platinum-DNA adducts using a CP9/19 cisplatin-modified DNA antibody. A representative maximum intensity projection of a stained cell is illustrated in a, where DAPI was used as a nuclear stain, and the yellow line represents the nuclear boundary. Platinum-DNA adduct formation was evaluated in response to 3 µM cisplatin and 10 µM carboplatin by immunofluorescence (b, c) and slot blot (d) analysis. Results are illustrative of three independent experiments. Foci formation was compared by one-way ANOVA. Scale bar = 100 µm. *p < 0.05, **p < 0.01, ***p < 0.001. Ns not significant, EV empty vector.
Double-strand break detection is influenced by FGF1 expression
As drug influx and Pt-DNA adduct formation were not FGF1-dependent, we next investigated whether FGF1 could influence the DNA damage repair response. Inhibition of the NHEJ pathway with 2 µM NU7026 DNA-PK inhibitor (DNA-PKi) did not influence cisplatin sensitivity in A2780DPP cells (Fig. 5a). Conversely, inhibition of ATM with 10 µM KU55933 (ATMi) completely re-sensitised A2780DPP cells to cisplatin (Fig. 5a), suggesting that the HR pathway may be more important to the drug-resistant phenotype of these cells. Again, neither inhibitor was independently toxic at the concentrations used (data not shown). To confirm that chemosensitivity was ATM-dependent, siRNA-mediated ATM depletion was optimised in A2780DPP cells (Fig. 5b). Following siRNA knockdown, A2780DPP cells were re-sensitised to cisplatin (fold change IC50 = 3.43, p = 0.004; Fig. 5c), assessed by MTT assay. We next used immunoblot analysis to investigate whether ATM activation (Ser1981 phosphorylation) in response to DNA damage (3 μM cisplatin challenge) was FGF1-dependent. A2780 cells induced maximal ATM phosphorylation 24 h following cisplatin challenge (Fig. 5d). In contrast, maximal ATM phosphorylation occurred 12 h earlier in A2780DPP cells (Fig. 5e), a phenotype reversible by FGF1 knockdown (Fig. 5f). Similar FGF1 dependency was observed for γH2AX expression. A2780 cells are sensitive to platinum drugs, and show basal γH2AX expression following cisplatin challenge (Fig. 5g). In contrast, resistant A2780DPP cells show a more transient γH2AX induction, 6 h following drug challenge (Fig. 5h), while re-sensitised A2780DPP FGF1KD cells delay γH2AX induction to 24 h (Fig. 5i), consistent with the FGF1-dependent p-ATM expression described above. Dependency of the DNA damage response was further confirmed by combining cisplatin challenge with the ATMi inhibitor KU55933 (Fig. 5j–l). KU55933 inhibited γH2AX phosphorylation 12 h following treatment in A2780 cells but not in A2780DPP cells, while inhibition was restored and phosphorylation inhibited 6 h following treatment in A2780DPP FGF1KD cells.
a MTT assays were used to compare viability of A2780, A2780DPP and A2780DPP cells treated with 10 µM KU55933 (ATMi) or 2 µM NU7026 (DNA-PKi) in response to cisplatin (0–25.33 µM). b qRT-PCR analysis was used to assess ATM expression relative to 18S rRNA in response to 5 nM, 10 nM and 20 nM ATM siRNA (±compound SD). c MTT assays were used to compare the viability of A2780, A2780DPP and A2780DPP cells transfected with 20 nM siRNA scrambled negative control or 20 nM siATM in response to cisplatin (0–25.33 µM). Representative immunoblot analysis comparing the phosphorylation of ATM (S1981) over indicated timepoints following 3 h treatment with 3 µM cisplatin in thymidine synchronised cells (2 mM, 24 h) in d A2780, e A2780DPP and f A2780DPP FGF1KD cells. A timecourse of FGF1-dependency H2AX (S139; γH2AX) phosphorylation was investigated before (g–i) and following treatment with 10 µM KU55933 (ATMi) (j–l) in A2780, A2780DPP and A2780DPP FGF1KD cells. Results are illustrative of four repeat experiments. Pairwise comparisons of mean IC50 values and relative gene expression were calculated by Student’s t-tests. Scale bar = 100 µm. *p < 0.05, **p < 0.01, ***p < 0.001. Ns not significant.
FGF1 promotes HR pathway activation in response to cisplatin
HR is initiated by focal recruitment of RPA and γH2AX, which recruits DSB repair proteins [37], inducing sister chromatid invasion to facilitate homology-directed error-free DSB repair [33] (Fig. 6a). Three micromolar cisplatin challenge was again used to induce DSB formation in synchronised cells, and immunofluorescence was used to assess the formation of RPA and γH2AX foci in A2780, A2780DPP and A2780DPP FGF1KD cells. In A2780 cells, RPA foci formed 12 h after cisplatin challenge. In contrast, foci were induced at 6 h in A2780DPP cells, with foci formation delayed to 12 h in A2780DPP FGF1KD cells (Fig. 6b). Similarly, γH2AX foci formation was observed 24 h following cisplatin challenge in A2780 cells, at 12 h in A2780DPP cells, with foci formation delayed to 24 h in A2780DPP FGF1KD cells (Fig. 6c), with a similar timecourse of FGF1-dependent RAD51 recombinase foci formation observed (Supplementary Fig. 4).
a Graphical abstract of HR-mediated DNA damage repair. ICL interstrand crosslink, DSB double-strand break, RPA replication protein A, ATM ataxia telangiectasia mutated, HR homologous recombination. Immunofluorescence and confocal microscopy were used to compare the formation and dissolution of b RPA foci and c γH2AX foci following 3 µM cisplatin challenge, following thymidine synchronisation (2 mM, 24 h). Typical representative maximum intensity projections are displayed. DAPI was used as a nuclear stain. n > 40 cells per treatment. Foci formation was compared by one-way ANOVA. ***p < 0.001. Scale bar = 20 µm.
Finally, to allow us to consider whether our observed FGF1-mediated chemosensitivity and DNA damage response changes were influenced by altered cell cycle parameters in A2780, A2780DPP and A2780DPP FGF1KD cells, we used flow cytometry to compare cell cycle profiles (a typical scatter plot, gating strategy and cell cycle distribution is illustrated in Supplementary Fig. 5a). While we saw a consistent modest increase in cells in S phase comparing the A2780 and A2780DPP cell lines (33.8% and 39.0%, respectively, p = 0.002), with a corresponding decrease in G1 cells (50.5% and 41.3%, respectively, p = 0.005), these changes were not FGF1-dependent, as FGF1 knockdown in A2780DPP cells did not have a significant influence on S and G1 phase cell numbers (Supplementary Fig. 5b–d, summarised in Panel e). In contrast, G2 cell numbers were FGF1 dependent, with a significant increase in A2780DPP cells compared to A2780 control cells (% increase 140.3, p = 0.003), which was significantly reduced following FGF1 knockdown (% reduction 82.3, p = 0.02, Supplementary Fig. 5f).
Taken together, our data therefore suggest that FGF1-induced cisplatin and carboplatin resistance is mediated by differential ATM phosphorylation and associated FGF1-dependent HR-mediated DSB repair.
Discussion
Response to first-line chemotherapy, an important determinant of clinical outcome, is frequently compromised by the development of drug-resistant disease in ovarian cancer patients [2, 3, 48]. We have previously shown that FGF1 expression modifies chemosensitivity to cisplatin and carboplatin in multiple ovarian cancer cell lines [4], consistent with data from Ryner et al. [5], describing increased expression of FGF pathway genes in the reactive stroma in chemoresistant ovarian cancer patients. We now confirm that introduction of FGF1 to drug-sensitive cells confers resistance, and that resistance is reversible by co-administration of pharmacological FGFR inhibitors in both immortalised cell lines and primary ascites-derived cell lines from drug-resistant ovarian cancer patients. Drug-resistant primary cell lines were re-sensitised by combination treatment with the FGFR inhibitor AZD4547—we prioritise extension of this clinical series as our DOCS study matures, and suggest that assessment of individuality in tumour and/or ascites FGF1 expression may have future clinical utility in patient selection for combination chemotherapy/FGFR inhibitor treatment. Importantly, AZD4547 is an orally bioavailable and well-tolerated Type I FGFR-specific inhibitor that has progressed to Phase II clinical trials for FGFR-positive squamous cell lung cancer (NCT02965378), breast, lung, and stomach cancer (NCT01795768) and solid tumours, lymphomas, and multiple myelomas (NCT02465060).
Platinum drug resistance has previously been reported to result from loss of drug-target interactions, due to either reduced intracellular drug accumulation (decreased drug influx or increased drug efflux) [49, 50], or from increased cytoplasmic drug sequestration [51]. In contrast, our data confirm that increased FGF1 expression confers drug resistance but does not directly influence Pt-DNA adduct formation, instead influencing the timecourse of HR-mediated DNA damage repair. Consistent with this hypothesis, previous studies have confirmed that platinum-DNA adducts are removed more rapidly in cisplatin-resistant ovarian cancer cells than in paired sensitive cell lines [52, 53]. We suggest that FGF1 influences DNA damage sensing and induces drug resistance by focal recruitment of the ssDNA protective protein RPA and associated phosphorylation of ATM, a kinase with a critical role in DSB sensing [36]. Differential FGF1-dependent ATM phosphorylation influences the kinetics of high-fidelity HR DNA damage repair activation, in preference to low-fidelity DSB repair by NHEJ, initiated by the binding of Ku70/80 heterodimers to blunt DNA ends [54]. ATM additionally promotes the removal of Ku heterodimers to inhibit NHEJ [55] and prevents the formation of NHEJ-induced chromatid fusions [28]. Our data therefore suggest that ATM is an important intermediate in the FGF1 signalling pathway, directly linking FGF signalling to HR-mediated DNA damage repair. As delayed engagement of HR results in the formation of toxic intermediates that promote apoptosis [28], we further describe an important role for FGF1 in accelerating HR pathway activation, which rationalises the resultant drug-resistant phenotype. It remains to be confirmed, however, whether DSBs formed by cisplatin-DNA adducts are induced, detected or repaired in an FGF1-dependent manner, and whether differential FGF1 expression influences the kinetics of adduct formation. FGF1-dependent RPA foci formation, however, suggests that FGF1 either promotes the formation or detection of DSBs. Consistent with our data, recent evidence in the gastrointestinal stromal tumour cell line T-1R shows that the alternative FGFR1-3 inhibitor BGJ398 (Infigratinib) or knockdown of FGFR2 attenuates DSB repair by HR and not by NHEJ [56]. We have not investigated the influence of FGF1 on other components of the HR pathway, for example, MRE11 and RAD51, in the current study - but highlight this as an interesting area for future focus, as differential expression of both genes has also previously been associated with platinum drug chemosensitivity in various cancers [57, 58], with RAD51 particularly related to platinum drug resistance in ovarian cancer following the restoration of genomic integrity through secondary reversion mutations [59]. It was particularly interesting to note that, although the effects are modest, we observed an FGF1-dependent increase in the G2 cell population in resistant A2780DPP cells. Although FGF1-dependent cell cycle regulation has not previously been associated with drug resistance, FGF1 has previously been shown to induce an ATM-dependent G2 arrest in RCS chondrocytes [60], and G2 arrest has previously been associated with cisplatin resistance in ovarian cell lines [61] and in other in cancer cell lines including lung adenocarcinoma [62] and gastric cancer [63]. We are therefore currently optimising experimental protocols to allow us to investigate the influence of cell cycle kinetics on the evolution of drug resistance in our DOCS study primary ovarian cancer cell models.
We have further shown that ATM inhibitors can re-sensitise immortalised platinum-resistant ovarian cancer cells, suggesting that ATM inhibition may be an additional novel candidate combination chemotherapy approach. We highlight the need to extend our analysis to clinical samples, and note that the orally bioavailable ATM inhibitor AZD0156 is currently in clinical trials in combination with other cytotoxic chemotherapies, including olaparib and irinotecan (NCT02588105). To further validate our hypothesis that the FGF1-mediated HR DNA damage response pathway is ATM-dependent, we additionally highlight the importance of future experiments to confirm that γH2AX foci formation is also reduced in the presence of FGFR inhibitors.
In summary, we therefore extend our previous data describing the promotion of drug resistance by a platinum chemotherapy-induced increase in FGF1 expression, and highlight that pharmacological inhibition of FGF signalling can re-sensitise both immortalised drug-resistant cells and primary cell lines from drug-resistant ovarian cancer patients. We describe a novel ATM kinase-mediated mechanism linking FGF and DNA damage response signalling, with cisplatin and carboplatin chemosensitivity influenced by FGF1-dependent regulation of the HR DNA damage response pathway, where FGF1 accelerates the kinetics of HR-mediated DNA damage repair.
Data availability
All data generated or analysed during this study are included in this published article [and associated Supplementary information files].
References
Scottish Intercollegiate Guidelines Network. Management of epithelial ovarian cancer. Healthcare Improvement Scotland. Edinburgh; 2018.
Clark TG, Stewart ME, Altman DG, Gabra H, Smyth JF. A prognostic model for ovarian cancer. Br J Cancer. 2001;85:944–52.
Vaughan S, Coward JI, Bast RCJ, Berchuck A, Balkwill FR. Rethinking ovarian cancer: recommendations for improving outcomes. Nat Rev Cancer. 2011;23:719–25.
Smith G, Ng MT, Shepherd L, Herrington CS, Gourley C, Ferguson MJ, et al. Individuality in FGF1 expression significantly influences platinum resistance and progression-free survival in ovarian cancer. Br J Cancer. 2012;107:1327–36.
Ryner L, Guan Y, Firestein R, Xiao Y, Choi Y, Rabe C, et al. Upregulation of periostin and reactive stroma is associated with primary chemoresistance and predicts clinical outcomes in epithelial ovarian cancer. Clin Cancer Res. 2015;21:2941–51.
Itoh N, Ornitz DM. Evolution of the Fgf and Fgfr gene families. Trends Genet. 2004;20:563–9.
Jackson A, Friedman S, han X, Engelka KA, Forough R, Maciag T. Heat shock induces the release of fibroblast growth factor 1 from NIH 3T3 cells. Proc Natl Acad Sci USA. 1992;89:10691–5.
Eswarakumar VP, Lax I, Schlessinger J. Cellular signaling by fibroblast growth factor receptors. Cytokine Growth Factor Rev. 2005;16:139–49.
Hoshi M, Nishida E, Sakai H. Activation of a Ca2’-inhibitable protein kinase that phosphorylates microtubule-associated protein 2 in vitro by growth factors, phorbol esters, andserum in quiescent cultured human fibroblasts. J Biol Chem. 1988;263:5398–401.
Kanazawa S, Fujiwara T, Matsuzaki S, Shingaki K, Taniguchi M, Miyata S, et al. bFGF regulates PI3-kinase-Rac1-JNK pathway and promotes fibroblast migration in wound healing. PLoS One. 2010;5:e12228.
Tan Y, Rouse J, Zhang A, Cariati S, Cohen P, Comb MJ. FGF and stress resulte CREB and ATF-1 via a pathway involving p38 MAP kinase and MAPKAP kinase-2. EMBO J. 1996;15:4629–42.
Seo JH, Suenaga A, Hatakeyama M, Taiji M, Imamoto A. Structural and functional basis of a role for CRKL in a fibroblast growth factor 8-induced feed-forward loop. Mol Cell Biol. 2009;29:3076–87.
Larsson H, Kilint P, Landgren E, Claesson-Welsh L. Fibroblast growth factor receptor-1-mediated endothelial cell proliferation is dependent on the Src homology (SH) 2/SH3 domain-containing adaptor protein Crk. J Biol Chem. 1999;274:25726–34.
Ornitz DM, Itoh N. The fibroblast growth factor signaling pathway. Wiley Interdiscip Rev Dev Biol. 2015;4:215–66.
Ishida S, Lee J, Thiele DJ, Herskowitz I. Uptake of the anticancer drug cisplatin mediated by the copper transporter Ctr1 in yeast and mammals. Proc Natl Acad Sci USA. 2002;99:14298–302.
Holzer AK, Katano K, Klomp LW, Howell SB. Cisplatin rapidly downregulates its own influx transporter hCTR1 in cultured human ovarian carcinoma cells. Clin Cancer Res. 2004;10:6744–9.
Holzer AK, Samimi G, Katano K, Naerdemann W, Lin XJ, Safaei R, et al. The copper influx transporter human copper transport protein 1 regulates the uptake of cisplatin in human ovarian carcinoma cells. Mol Pharm. 2004;66:817–23.
Reishus JW, Martin DSJ. cis-Dichlorodiammineplatinum(II). Acid hydrolysis and isotopic exchange of the chloride ligands. J Am Chem Soc. 1961;83:2457–62.
Onoa GB, Cervantes G, Moreno V, Prieto MJ. Study of the interaction of DNA with cisplatin and other Pd(II) and Pt(II) complexes by atomic force microscopy. Nucleic Acid Res. 1998;26:1473–80.
Hanada K, Budzowska M, Modesti M, Maas A, Wyman C, Essers J, et al. The structure-specific endonuclease Mus81-Eme1 promotes conversion of interstrand DNA crosslinks into double-strands breaks. EMBO J. 2006;25:4921–32.
Alabert C, Groth A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol. 2012;13:153–67.
McVey M. Strategies for DNA interstrand crosslink repair: insights from worms, flies, frogs and slime moulds. Environ Mol Mutagen. 2010;51:646–58.
Cousineau I, Abaji C, Belmaaza A. BRCA1 regulates RAD51 function in response to DNA damage and suppresses spontaneous sister chromatid replication slippage: implications for sister chromatid cohesion, genome stability, and carcinogenesis. Cancer Res. 2005;65:11384–91.
Wu PY, Frit P, Meesala S, Dauvillier S, Modesti M, Andres SN, et al. Structural and functional interaction between the human DNA repair proteins DNA ligase IV and XRCC4. Mol Cell Biol. 2009;29:3163–72.
Bhattacharyya A, Ear US, Koller BH, Weichselbaum RR, Bishop DK. The breast cancer susceptibility gene BRCA1 is required for subnuclear assembly of Rad51 and survival following treatment with the DNA cross-linking agent cisplatin. J Biol Chem. 2000;275:23899–903.
Kee Y, Huang M, Chang S, Moreau LA, Park E, Smith PG, et al. Inhibition of the Nedd8 system sensitizes cells to DNA interstrand cross-linking agents. Mol Cancer Res. 2012;10:369–77.
Galluzzi L, Senovilla L, Vitale I, Michels J, Martins I, Kepp O, et al. Molecular mechanisms of cisplatin resistance. Oncogene. 2012;31:1869–83.
Balmus G, Pilger D, Coates J, Demir M, Sczaniecka-Clift M, Barros AC, et al. ATM orchestrates the DNA-damage response to counter toxic non-homologous end-joining at broken replication forks. Nat Commun. 2019;10:87.
Bryant HE, Schultz N, Thomas HD, Parker AM, Flower D, Lopez E, et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature. 2005;434:913–7.
Farmer H, McCabe N, Lord CJ, Tutt ANJ, Johnson DA, Richardson TB, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature. 2005;434:917–21.
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast RC Jr, Beral V, et al. Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer. 2015;15:668–79.
Huertas P. DNA resection in eukaryotes: deciding how to fix the break. Nat Struct Mol Biol. 2010;17:11–6.
Nielsen FC, van Overeem Hansen T, Sorensen CS. Hereditary breast and ovarian cancer: new genes in confined pathways. Nat Rev Cancer. 2016;16:599–612.
Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER, Hurov KE, Luo JI, et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science. 2007;316:1160–6.
Bakkenist CJ, Kastan MB. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature. 2003;421:499–506.
Burma S, Chen BP, Murphy M, Kurimasa A, Chen DJ. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J Biol Chem. 2001;276:42462–7.
Stucki M, Clapperton JA, Mohammad D, Yaffe MB, Smerdon SJ, Jackson SP. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell. 2005;123:1213–26.
Hamilton TC, Young RC, Ozols RF. Experimental model systems of ovarian cancer: applications to the design and evaluation of new treatment approaches. Semin Oncol. 1984;11:285–98.
Masuda H, Ozols RF, Lai G-M, Fojo AT, Rothenberg M, Hamilton TC. et al. Repair as a mechanism of acquired resistance to cis-diamminedicloroplatinum (II) in human ovarian cancer cell lines. Cancer Res. 1988;48:5713–6.
Fogh J, Fogh JM, Orfeo T. One hundred and twenty seven cultured human tumor cell lines producing tumors in nude mice. J Nat Cancer Inst. 1977;59:221–6.
Buick RN, Pullano R, Trent JM. Comparative properties of five human ovarian adenocarcinoma cell lines. Cancer Res. 1985;45:3668–76.
Iwanicki MP, Davidowitz RA, Ng MR, Besser A, Muranen T, Merritt M, et al. Ovarian cancer spheroids use myosin-generated force to clear the mesothelium. Cancer Discov. 2011;1:144–57.
Renner F, Moreno R, Schmitz ML. SUMOylation-dependent localization of IKKepsilon in PML nuclear bodies is essential for protection against DNA-damage-triggered cell death. Mol Cell. 2010;37:503–15.
Vaidyanathan A, Sawers L, Gannon AL, Chakravarty P, Scott AL, Bray SE, et al. ABCB1 (MDR1) induction defines a common resistance mechanism in paclitaxel- and olaparib-resistant ovarian cancer cells. Br J Cancer. 2016;115:431–41.
Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods. 1983;65:55–63.
Konecny G, Crohns C, Pegram M, Felber M, Lude S, Kurbacher C, et al. Correlation of drug response with the ATP tumorchemosensitivity assay in primary FIGO stage III ovarian cancer. Gynecol Oncol. 2000;77:258–63.
Van Cutsem E, Bang YJ, Mansoor W, Petty RD, Chao Y, Cunningham D, et al. A randomized, open-label study of the efficacy and safety of AZD4547 monotherapy versus paclitaxel for the treatment of advanced gastric adenocarcinoma with FGFR2 polysomy or gene amplification. Ann Oncol. 2017;28:1316–24.
CRUK. Ovarian cancer survival by stage at diagnosis (England data). Cancer Research U.K. 2018. https://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/ovarian-cancer/survival#heading-Three.
Loh SY, Mistry P, Kelland LR, Abel G, Harrap KR. Reduced drug accumulation as a major mechanism of acquired resistance to cisplatin in a human ovarian carcinoma cell line: circumvention studies using novel platinum(I) and (IV) ammine/amine complexes. Br J Cancer. 1992;66:1109–15.
Katano K, Kondo A, Safaei R, Holzer A, Samimi G, Mishima M. et al. Acquisition of resistance to cisplatin is accompanied by changes in the cellular pharmacology of copper. Cancer Res. 2002;62:6559–65.
Lewis AD, Hayes JD, Wolf CR. Glutathione and glutathione-dependent enzymes in ovarian adenocarcinoma cell lines derived from a patient before and after the onset of drug resistance: intrinsic differences and cell cycle effects. Carcinogenesis. 1988;9:1283–7.
Sonego M, Pellizzari I, Dall’Acqua A, Pivetta E, Lorenzon I, Benevol S, et al. Common biological phenotypes characterize the acquisition of platinum-resistance in epithelial ovarian cancer cells. Sci Rep. 2017;7:7104.
Johnson SW, Swiggard PA, Handel LM, Brennan JM, Godwin AK, Ozols RF, et al. Relationship between platinum-DNA adduct formation and removal and cisplatin cytotoxicity in cisplatin-sensitive and -resistant human ovarian cancer cells. Cancer Res. 1994;54:5911–6.
Walker JR, Corpina RA, Goldberg J. Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair. Nature. 2001;412:607–14.
Chanut P, Britton S, Coates J, Jackson SP, Calsou P. Coordinated nuclease activities counteract Ku at single-ended DNA double-strand breaks. Nat Commun. 2016;7:12889.
Sergei B, Pavel D, Aigul G, Firyuza B, Ilmira N, Ilshat M, et al. Inhibition of FGFR2-signaling attenuates a homology-mediated DNA repair in GIST and sensitizes them to DNA-topoisomerase II inhibitors. Int J Mol Sci. 2020;21:352.
Murakami T, Shoji Y, Nishi T, Chang SC, Jachimowicz RD, Hoshimoto S, et al. Regulation of MRE11A by UBQLN4 leads to cisplatin resistance in patients with esophageal squamous cell carcinoma. Mol Oncol. 2021;15:1069–87.
Dev H, Chiang TW, Lescale C, de Krijger I, Martin AG, Pilger D, et al. Shieldin complex promotes DNA end-joining and counters homologous recombination in BRCA1-null cells. Nat Cell Biol. 2018;20:954–65.
Kondrashova O, Nguyen M, Shield-Artin K, Tinker AV, Teng NNH, Harrell MI, et al. Secondary somatic mutations restoring RAD51C and RAD51D associated with acquired resistance to the PARP inhibitor rucaparib in high-grade ovarian carcinoma. Cancer Discov. 2017;7:984–98.
Tran T, Kolupaeva V, Basilico C. FGF inhibits the activity of the cyclin B1/CDK1 kinase to induce a transient G(2)arrest in RCS chondrocytes. Cell Cycle. 2010;9:4379–86.
Tu Y, Kim E, Gao Y, Rankin GO, Li B, Chen YC. Theaflavin-3, 3’-digallate induces apoptosis and G2 cell cycle arrest through the Akt/MDM2/p53 pathway in cisplatin-resistant ovarian cancer A2780/CP70 cells. Int J Oncol. 2016;48:2657–65.
Gonzalez Rajal A, Marzec KA, McCloy RA, Nobis M, Chin V, Hastings JF, et al. A non-genetic, cell cycle-dependent mechanism of platinum resistance in lung adenocarcinoma. Elife. 2021;10:e65234.
Lu Y, Han D, Liu W, Huang R, Ou J, Chen X, et al. RNF138 confers cisplatin resistance in gastric cancer cells via activating Chk1 signaling pathway. Cancer Biol Ther. 2018;19:1128–38.
Acknowledgements
We acknowledge Dr Laureano de la Vega, Dr Adrian Saurin, Dr Rita Moreno and Dr Prasun Chakraborty for provision of plasmid vectors, inhibitors, and microscopy reagents, and for useful discussions. The authors thank the research nurses at the Stefani Unit, Ninewells Hospital, and Dr Wendy McMullen for their assistance with DOCS study patient recruitment. We additionally acknowledge Amber McWhirter and Caitlin MacDonald for their assistance with DOCS study data analysis.
Funding
This work was supported by a grant from Medical Research Scotland (PhD-1030-2016).
Author information
Authors and Affiliations
Contributions
Conception and study supervision: KJH, MJF and GS; data collection and analysis: HAN, LS and RGC; original draft writing: HAN and GS; and manuscript editing and final approval: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The Dundee Ovarian Cancer Study (DOCS) was approved by the East of Scotland Research Ethics Service REC1 Committee (13/ES/0035). Informed consent was obtained from all subjects, and the study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nicholson, H.A., Sawers, L., Clarke, R.G. et al. Fibroblast growth factor signalling influences homologous recombination-mediated DNA damage repair to promote drug resistance in ovarian cancer. Br J Cancer 127, 1340–1351 (2022). https://doi.org/10.1038/s41416-022-01899-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01899-z
This article is cited by
-
NPAS2 dampens chemo-sensitivity of lung adenocarcinoma cells by enhancing DNA damage repair
Cell Death & Disease (2024)
-
Radiation exposure lymphocyte damage assessed by γ-H2AX level using flow cytometry
Scientific Reports (2024)
-
Molecular panorama of therapy resistance in prostate cancer: a pre-clinical and bioinformatics analysis for clinical translation
Cancer and Metastasis Reviews (2024)
-
Fanconi anemia associated protein 20 (FAAP20) plays an essential role in homology-directed repair of DNA double-strand breaks
Communications Biology (2023)