Abstract
Background and aims
CDH1 germline variants have been linked to heritability in diffuse gastric (DGC) and lobular breast cancer (LBC). Studies have not yet assessed whether CDH1 is a cancer-susceptibility gene in other cancers. Herein, we mapped the landscape of pathogenic and likely pathogenic (P/LP) germline variants in CDH1 across various cancers and ethnicities.
Methods
We evaluated CDH1 germline P/LP variants in 212,944 patients at one CLIA-certified laboratory (Invitae) and described their frequency in 7 cancer types. We screened for CDH1 variant enrichment in each cancer relative to a cancer-free population from The Genome Aggregation Database version 3 (gnomADv3).
Results
CDH1 P/LP variants were identified in 141 patients, most commonly in patients with DGC (27/408, 6.6%) followed by colorectal signet-ring cell cancer (CSRCC; 3/79, 3.8%), gastric cancer (56/2756, 2%), and LBC (22/6809, 0.3%). CDH1 P/LP variants were enriched in specific ethnic populations with breast cancer, gastric cancer, CRC, LBC, DGC, and CSRCC compared to matched ethnicities from gnomADv3.
Conclusion
We report for the first time the prevalence of P/LP CDH1 variants across several cancers and show significant enrichment in CDH1 P/LP variants for patients with CSRCC, DGC, and LBC across various ethnicities. Future prospective studies are warranted to validate these findings.
Similar content being viewed by others
Introduction
Germline variants in CDH1, which codes for the cell–cell adhesion protein E-cadherin, were first identified in families with hereditary diffuse gastric cancer (HDGC) [1]. Subsequent reports noted that individuals with germline CDH1 pathogenic variants were predisposed to both DGC and lobular breast cancer (LBC) [2]. Most recently, Massari et al. reported that 7% of all CDH1 mutations are present in non-gastric tumours with most being identified in patients with breast cancer [3]. Moreover, CDH1 mutations are more likely to be detected in areas with a low incidence of gastric cancer [4]. However, most reports have focused on highly penetrant HDGC families, in which the cumulative risk of HDGC for CDH1 mutation carriers is 70% by age 80 years for men and 56% for women, while the cumulative risk of LBC for women is estimated to be 42% by age 80 years [5]. A recent study found that among patients not exclusively ascertained based on strict HDGC criteria, the cumulative incidence of gastric cancer for individuals with pathogenic CDH1 variants is significantly lower (42% at age 80 years) than what has been previously reported [6, 7].
Although there is no evidence that the risk of other cancer types in individuals with a CDH1 variant is significantly increased [8], multiple case reports have noted the occurrence of CRC and appendiceal Signet-Ring Cell Carcinomas in CDH1 variant carriers [5]. In one study, colon cancer was reported in 3 of 238 (1%) CDH1 pathogenic variant carriers, with 1 case of SRCC. Most recently, Benesch et al. postulated through clinical observation and data from SEER that SRCC might be enriched among CDH1 variant carriers with signet ring cell gastric cancer [9]. Notably, there was no increased risk of colon cancer in CDH1 carriers compared to that of the SEER population [6].
The prevalence of CDH1 pathogenic variants in patients with gastric cancer and other cancer types is unknown. Here we examine the prevalence of CDH1 variants among 212,944 patients referred for genetic testing. We also examine germline cancer-risk variants by ethnicity across several tumour subtypes and identify, for the first time, CDH1 variant enrichment in individuals with CRC and colorectal signet-ring cell cancer (CSRCC).
Methods
Patient cohort
Personal history information for 212,944 independent probands with cancer was obtained from submitted requisition forms and medical records. The cohort included patients with breast, CRC, gastric, head and neck, ovarian, pancreatic, and prostate cancer types. All patients completed clinical germline genetic testing at Invitae (San Francisco, CA) between 09/2014 and 06/2020. Patient data were de-identified before analysis and the Western Institutional Review Board provided study oversight and approval. Western Institutional Review Board protocol number 1167406 waived the requirement to obtain written patient informed consent.
Germline genetic testing
The genes selected for sequencing for each patient were chosen by the ordering clinician; all the patients reported here had been chosen for CDH1 analysis. Genomic DNA was extracted from whole blood using a QiaSymphony (Qiagen, Hilden, Germany). Targeted genes including CDH1 were captured using Agilent (Santa Clara, CA) SureSelect probes or Integrated DNA Technologies (Coral, IL) xGen Lockdown probes at positions where SureSelect yield was inadequate. Clinically important regions of CDH1 including all the coding exons and 10 to 20 base pairs of adjacent intronic sequences on either side of the coding exons were covered. Next-generation sequencing [10] was performed on the Illumina (San Diego, CA) MiSeq or HiSeq 2500 to at least 350× average coverage of 2 × 150 reads, with a minimum of 50× required at every targeted position. Stringent process controls were used to minimise read-depth variability, and up to eight anonymous blood samples were used as control specimens in each run to measure remaining coverage variability [11].
Germline variant calling and assessment
Small indels and single-nucleotide variants were analysed using the Genome Analysis Toolkit [12]. Copy-number variant calls were performed using CNVitae [11]. Large structural variants were detected using split-read analysis. Candidate CDH1 variants were classified as pathogenic or likely pathogenic (P/LP) if: they affected the structure of CDH1; conferred a truncating, initiation codon or splice donor/acceptor effect; if functional data showed an impact on protein function; or if pathogenicity was otherwise reported in the published literature [13]. Orthogonal technology was used to validate P/LP variants via Sanger sequencing or Multiplex Ligation-Dependent Probe Amplification [14]. Confirmed variants were interrogated using a refined American College of Medical Genetics and Genomics criteria (Sherloc) [15]. For each of the examined tumour subtypes, the frequency of pathogenic germline variants in CDH1 relative to the number of patients sequenced was calculated.
Ethnicity and enrichment analysis
Ethnicity was provided by all patients at the time of test ordering and was grouped based on categories reported in population databases. The following ethnicities were considered in the analysis: Ashkenazi Jewish, Asian, Black/African American, White/Caucasian, and Hispanic. For each ethnicity, we calculated the frequency of pathogenic germline variants relative to the number of patients in which the gene was tested.
For every cancer subtype, we compared the frequency of pathogenic variants in CDH1 to the frequency of gene variants in an independent population derived from The Genome Aggregation Database version 3 (gnomAD v3) [16]. Two independent methods were followed: (1) All CDH1 variants in gnomAD were reviewed in ClinVar [17]. Variants in gnomAD deemed P/LP variants in ClinVar were retained. (2) Variants reported in gnomAD at a frequency >0.01% were excluded. Moreover, missense and synonymous variants in CDH1 from both the Invitae and gnomAD cohorts were excluded and only loss-of-function (LOF) variants were retained. LOF variants included frameshift, splice site, and nonsense variants, variants in the initiator codon, and exonic deletions.
For both analyses, the frequency of germline variants in each of the ethnic populations (Ashkenazi Jewish, Asian, Black/African American, Hispanic, and White/Caucasian) in the Invitae cancer cohort were compared to the frequency of these gene variants across various ancestries derived from gnomAD v3. CDH1 variants were considered enriched in a cancer subtype if they met both criteria: (1) they were significantly more likely to occur in a specific ethnic population with cancer when compared to the same ancestry in gnomAD and (2) the p-value was significant in both “LOF” and “ClinVar” analyses.
Statistical analysis
Two-sided Fisher’s exact test was used to calculate the odds ratios, 95% confidence intervals (CIs), and p-values of all enrichment analyses. For the enrichment analysis, we applied Benjamini–Hochberg correction for the number of independent tests conducted (significant q-value cutoff of <0.05).
Results
Germline landscape of CDH1 variants in the Invitae cohort
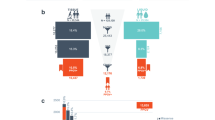
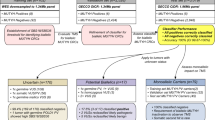
Of the 212,944 patients with cancer and available CDH1 sequencing data, 151,465 had breast cancer (71.1%), 27,915 had CRC (13.1%), 15,225 had ovarian cancer (7.1%), and 18,339 (8.6%) had other cancer types (Fig. 1a and Table 1). Detailed clinical history was available on all 141 patients with P/LP variants in CDH1 (Table S1.1). The most common cancer types in patients with P/LP CDH1 variants were breast cancer (77 of 141, 54.6%), gastric cancer (56 of 141, 39.7%), and CRC (14 of 141, 9.9%). Among patients with breast cancer of known histology (n = 30), 8 (27%) had ductal and 22 (73%) had LBC. Notably, five probands with CDH1 P/LP variants had concomitant gastric cancer and CRC. Variant type and location are shown in Fig. 1b and did not vary according to cancer type. The median age of onset of breast, colorectal, and gastric cancers among patients with CDH1 P/LP variants was lower than that of the general population from the SEER cohort (Table S1.2) [18].
a Frequency of P/LP germline variants in CDH1 in each of the seven cancer types. The numbers above each bar indicate the total number of patients tested for each cancer type; the number within each box indicates the number of P/LP variants. Y axis is frequency. b Landscape of P/LP germline variants in CDH1 in 212,944 patients with cancer. The variants are labeled with carrier counts and coloured by their respective carriers’ ancestry (Caucasian: blue, African American: red, Asian: green, Ashkenazi Jewish: black, non-white Hispanic: orange) for breast cancer, gastric cancer, and colorectal cancer. c Frequency of P/LP germline variants in CDH1 in each of the five ethnicities in the gnomAD v3 cohort. The numbers above each bar indicate the total number of subjects assessed by sequencing; the number within each box indicates the number of P/LP variants.
Among all major cancer types, gastric cancer had the highest frequency of P/LP variants (56 of 2756, 2%, 95% CI = 1.5–2.6%, Fig. 1a and Table S1.3) followed by head and neck (1 of 159, 0.6%, 95% CI = 0–3.5%) and breast cancer (77 of 151,465, 0.05%, 95% CI = 0.04–0.06%). Acknowledging that there were relatively small numbers of subjects in some categories (e.g. Ashkenazi Jewish with gastric cancer), none of the five ethnicities showed significantly higher CDH1 P/LP variant frequency for any cancer type (p-values of pairwise comparisons in Table S1.4). Various population groups and their cohort sizes are shown in Supplementary Table S1.5.
Enrichment analysis of major cancer subtypes
We then conducted enrichment analysis (see methods) and found that the odds of P/LP germline variants in CDH1 among African Americans, Asians, Caucasians, and Hispanics with gastric cancer in our cohort was significantly higher (126-fold to infinity) in comparison to the odds in the corresponding gnomAD ancestry cohorts (Figs. 1c and 2a, b and Table S1.5). Caucasians with breast cancer and CRC were enriched for CDH1 germline variants compared to Caucasian controls from gnomAD. The odds ratio was also increased for CDH1 variants in several other ethnicities for each of breast and colorectal cancer, though in general to a smaller degree, and false discovery rate-corrected q-values were not significant.
Enrichment of P/LP CDH1 variants in patients with various cancer types using two independent methods (see Supplementary Methods). Fisher’s exact test was used to calculate the odds ratios (ORs) and 95% confidence intervals (CIs). A two-sided binominal test was used to compute the p-values. a Enrichment analysis including variants in gnomAD deemed P/LP in ClinVar. b Enrichment analysis including only loss-of-function variants in gnomAD and Invitae found at a frequency ≤0.01. Applying a false discovery rate of <0.05 (horizontal dotted line), germline variants in CDH1 were enriched in Caucasians with CRC; African Americans/Blacks, Asians, Caucasians, and Hispanics with gastric cancer; and Caucasians with breast cancer when compared to corresponding ancestries from gnomAD v3. BRCA breast cancer, CRC colorectal cancer, GC gastric cancer.
Enrichment of CDH1 variants in DGC and LBC
We then analysed histology-specific associations and found germline P/LP variants in CDH1 in 6.6% (27 of 409) of patients with DGC and 0.3% (22 of 6955) of patients with LBC. The median age at diagnosis of patients with DGC and LBC harbouring CDH1 P/LP variants was 42 (range 19–75) and 48 years (range 41–66), respectively. Interestingly, among patients with DGC in our cohort, Caucasians harboured significantly more CDH1 P/LP germline variants compared to Asians and Hispanics (20/192, 10.4% vs 1/63, 1.6%, p = 0.032 and 3/94, 3.2% vs 0/13670, 0%, p = 0.038 respectively, Fig. 3a and Table S1.6). However, in LBC, none of the five ethnicities showed significantly higher CDH1 P/LP variant frequency (Fig. 3b and Table S1.6).
Enrichment analysis with gnomAD showed that African Americans and Caucasians with LBC and DGC were enriched for P/LP CDH1 germline variants. Ashkenazi Jews, Asians, and Hispanics harboured significantly more CDH1 P/LP variants compared to controls from gnomAD in LBC and DGC, respectively (Table S1.7).
Prevalence of CDH1 variants in CSRCC and enrichment compared to gnomAD
Prior case reports suggested that an association between CSRCC and CDH1 germline carriers exists [5, 19]. We found that 3.8% of patients with CSRCC (3 of 79) harboured CDH1 P/LP variants. Age at diagnosis for two of the three patients was known (35 and 41 years). Compared to corresponding controls from gnomAD, African Americans and Caucasians with CSRCC were enriched for P/LP CDH1 germline variants (African American LOF analysis: q = 0.0017, OR = 1365, 95% CI = 24–16,384; African American ClinVar Analysis: q = 0.001, OR = 1365, 95% CI = 24–16,384; Caucasian LOF analysis: q = 0.0001, OR = 226, 95% CI = 24–996; Caucasian ClinVar Analysis: q < 0.0001, OR = 969, 95% CI = 80–8192; Fig. 2a, b and Table S1.7, see “Methods”). However, we consider these findings tentative given the small number of patients within each ethnic group in our study (Fig. 3c and Table S1.6).
Discussion
The prevalence of CDH1 germline variants in patients with various cancer types is still not well-described. Herein, we found CDH1 germline carriers in about 7% of patients with DGC and 3.8% of patients with CSRCC. To date, there is conflicting evidence regarding the prevalence of CDH1 P/LP germline variants in LBC. In our study, among 6809 patients with LBC, approximately 0.3% harboured germline variants in CDH1. This frequency is lower than what is reported in prior studies (1–8%), which focused mainly on patients with early-onset or bilateral disease [20, 21]. In a comprehensive review of hereditary LBC, Corso et al. emphasised the importance of surveillance in CDH1 P/LP variant carriers [22]. With increasing knowledge about LBC risk factors, CDH1 germline genetic testing in high-risk families remains paramount.
Prior studies have led to conflicting results regarding the prevalence of CDH1 germline variants in Asians vs non-Asians. Despite the high incidence of gastric cancer in East Asian countries, previous work has suggested that low detection rates for germline CDH1 variants are identified in Asians compared to countries with a lower incidence of gastric cancer [23]. More recently, a study [24] of 105 Japanese patients with DGC found that germline variants in CDH1 occured in 14 patients (13.3%) and showed that Japanese patients with gastric cancer were four times more likely to harbour CDH1 variants compared to TCGA (The Cancer Genome Atlas) non-Asian populations with gastric cancer. In our study, we found that the prevalence of CDH1 P/LP variants is significantly higher in Caucasians with DGC (10.4%) compared to Asians (1.6%, p = 0.032) and Hispanics (3.2%, p = 0.038). One possible explanation for this difference is that non-Japanese Asians, which likely represent a sizable portion of our Asian population, may have lower levels of CDH1 variants compared to Japanese Asians. Another observation is that all 14 variants identified by Suzuki et al. [24] are classified as benign, likely benign, or of uncertain significance according to Invitae guidelines and ClinVar reports, and hence would not be P/LP variants by our criteria. Whether those 14 variants are truly non-pathogenic or are unrecognised but significant variants in CDH1 remains to be determined.
Histology-specific enrichment analysis validated prior associations between CDH1 germline variants in LBC and DGC and identified a novel association with CSRCC [9]. CSRCC is an aggressive adenocarcinoma subtype with a poor prognosis overall and thus determining cancer-risk susceptibility genes presents an unmet need. Guidelines for genetic testing factor in the underlying likelihood of identifying a germline variant [25]. Thus, accurate estimates of germline prevalence, identified herein, may alter recommendations for different patient populations. Consideration should also be given to colonoscopy surveillance in CDH1 carriers. Notably, prior work [6] did not identify an increased risk of developing CRC in CDH1 carriers compared to patients from the SEER cohort but was underpowered to assess for associations in patients with CSRCC. In our cohort, CDH1 variant enrichment was observed among Caucasian patients with CRC.
When comparing the prevalence of CDH1 P/LP variants across the different ethnicities within each cancer type or subtype, none of the five ethnic populations showed significantly higher CDH1 P/LP variant frequency, except for the aforementioned observation in DGC. Thus, the enrichment seen when comparing to gnomAD cohorts is independent of ethnicity but was not seen in some comparisons likely due to the lack of statistical power.
CDH1 is a tumour suppressor, and the vast majority of variants detected here were LOF (128 of 141, 90.8%, 95% CI = 84.9–94.5%), and distributed throughout the coding sequence. A prior study reported significant enrichment of CDH1 germline variants located in the PRE-PRO region (amino acid 1–115) in HDGC families affected by CRC. In addition, patients harbouring CDH1 variants in the linker region (regions shown in white, Fig. 1B) were significantly less likely to develop breast cancer [26]. In this dataset, there was no association between the location of CDH1 variant and the development of individual cancers.
Our study has several limitations. First, the selection of patients for genetic testing was influenced by clinical judgement and was likely skewed towards individuals with a significant suspicion for heritable pathogenic variants. Second, personal and family history data were obtained from genetic testing requisition forms and were not confirmed by direct review of the medical records or other data sources. Ethnicity information was provided by the subjects with no confirmation. Similarly, there was no central pathological confirmation of either the cancer type or subtype. We caution against overinterpreting ethnicity-specific CDH1 associations as only a handful of carriers may drive enrichment in a small cancer cohort.
In the largest study to date evaluating CDH1 germline variants, we found significant enrichment of P/LP variants in patients with CSRCC, CRC, breast, and gastric cancer. Importantly, we found that the frequency of P/LP CDH1 variants in multiple cancer types did not vary according to ethnicity for the most part. This is the first report on the prevalence of CDH1 variants in African American and Hispanic populations and indicates that these populations have the same frequency of P/LP variants as Caucasians and should be subject to the same germline testing and screening considerations.
Data availability
Data will be available in Supplementary Table S1 and Supplementary Materials.
References
Guilford P, Hopkins J, Harraway J, McLeod M, McLeod N, Harawira P, et al. E-cadherin germline mutations in familial gastric cancer. Nature. 1998;392:402–5.
Xie ZM, Li LS, Laquet C, Penault-Llorca F, Uhrhammer N, Xie XM, et al. Germline mutations of the E-cadherin gene in families with inherited invasive lobular breast carcinoma but no diffuse gastric cancer. Cancer. 2011;117:3112–7.
Massari G, Magnoni F, Favia G, Peradze N, Veronesi P, La Vecchia C, et al. Frequency of CDH1 germline mutations in non-gastric cancers. Cancers. 2021;13:2321.
Corso G, Corso F, Bellerba F, Carneiro P, Seixas S, Cioffi A, et al. Geographical distribution of E-cadherin germline mutations in the context of diffuse gastric cancer: a systematic review. Cancers. 2021;13:1269.
van der Post RS, Vogelaar IP, Carneiro F, Guilford P, Huntsman D, Hoogerbrugge N, et al. Hereditary diffuse gastric cancer: updated clinical guidelines with an emphasis on germline CDH1 mutation carriers. J Med Genet. 2015;52:361–74.
Roberts ME, Ranola JMO, Marshall ML, Susswein LR, Graceffo S, Bohnert K, et al. Comparison of CDH1 penetrance estimates in clinically ascertained families vs families ascertained for multiple gastric cancers. JAMA Oncol. 2019;5:1325–31.
Blair VR, McLeod M, Carneiro F, Coit DG, D’Addario JL, van Dieren JM, et al. Hereditary diffuse gastric cancer: updated clinical practice guidelines. Lancet Oncol. 2020;21:e386–97.
Hansford S, Kaurah P, Li-Chang H, Woo M, Senz J, Pinheiro H, et al. Hereditary diffuse gastric cancer syndrome: CDH1 mutations and beyond. JAMA Oncol. 2015;1:23–32.
Benesch MGK, Bursey SR, O’Connell AC, Ryan MG, Howard CL, Stockley CC, et al. CDH1 gene mutation hereditary diffuse gastric cancer outcomes: analysis of a large cohort, systematic review of endoscopic surveillance, and secondary cancer risk postulation. Cancers. 2021;13:2622.
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG, et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 2008;456:53–9.
Lincoln SE, Kobayashi Y, Anderson MJ, Yang S, Desmond AJ, Mills MA, et al. A systematic comparison of traditional and multigene panel testing for hereditary breast and ovarian cancer genes in more than 1000 patients. J Mol Diagn. 2015;17:533–44.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Kurian AW, Hare EE, Mills MA, Kingham KE, McPherson L, Whittemore AS, et al. Clinical evaluation of a multiple-gene sequencing panel for hereditary cancer risk assessment. J Clin Oncol. 2014;32:2001–9.
Hogervorst FB, Nederlof PM, Gille JJ, McElgunn CJ, Grippeling M, Pruntel R, et al. Large genomic deletions and duplications in the BRCA1 gene identified by a novel quantitative method. Cancer Res. 2003;63:1449–53.
Nykamp K, Anderson M, Powers M, Garcia J, Herrera B, Ho YY, et al. Sherloc: a comprehensive refinement of the ACMG-AMP variant classification criteria. Genet Med. 2017;19:1105–17.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018;46:D1062–7.
Howlader NNA, Krapcho M, Miller D, Brest A, Yu M, Ruhl J, Tatalovich Z, Mariotto A, Lewis DR, Chen HS, Feuer EJ, Cronin KA, editors. SEER Cancer Statistics Review, 1975–2018, National Cancer Institute. 2020. https://seer.cancer.gov/csr/1975_2018/. Accessed Apr 2021.
Pharoah PD, Guilford P, Caldas C, International Gastric Cancer Linkage Consortium. Incidence of gastric cancer and breast cancer in CDH1 (E-cadherin) mutation carriers from hereditary diffuse gastric cancer families. Gastroenterology. 2001;121:1348–53.
Petridis C, Shinomiya I, Kohut K, Gorman P, Caneppele M, Shah V, et al. Germline CDH1 mutations in bilateral lobular carcinoma in situ. Br J Cancer. 2014;110:1053–7.
Schrader KA, Masciari S, Boyd N, Salamanca C, Senz J, Saunders DN, et al. Germline mutations in CDH1 are infrequent in women with early-onset or familial lobular breast cancers. J Med Genet. 2011;48:64–8.
Corso G, Veronesi P, Sacchini V, Galimberti V. Prognosis and outcome in CDH1-mutant lobular breast cancer. Eur J Cancer Prev. 2018;27:237–8.
Sugimoto S, Komatsu H, Morohoshi Y, Kanai T. Recognition of and recent issues in hereditary diffuse gastric cancer. J Gastroenterol. 2015;50:831–43.
Suzuki A, Katoh H, Komura D, Kakiuchi M, Tagashira A, Yamamoto S, et al. Defined lifestyle and germline factors predispose Asian populations to gastric cancer. Sci Adv. 2020;6:eaav9778.
Corso G, Figueiredo J, La Vecchia C, Veronesi P, Pravettoni G, Macis D, et al. Hereditary lobular breast cancer with an emphasis on E-cadherin genetic defect. J Med Genet. 2018;55:431–41.
Lo W, Zhu B, Sabesan A, Wu HH, Powers A, Sorber RA, et al. Associations of CDH1 germline variant location and cancer phenotype in families with hereditary diffuse gastric cancer (HDGC). J Med Genet. 2019;56:370–9.
Acknowledgements
We thank all individuals who participated in this study.
Funding
The authors received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
EA, TZ, AHN, EWA, DJK, and GS designed this study. EA, TZ, AHN, EWA, SAA, THM, KH, SMN, EDE, HQR, TKC, DJK, GS conceived this study. EA, TZ, AHN, and EWA analyzed the results and generated the figures. EA, TZ, AHN, DJK, and GS drafted the manuscript. All authors reviewed the manuscript, approved the final version and take responsibility for all aspects of this work.
Corresponding authors
Ethics declarations
Competing interests
DJK reports serving as a consultant for Novartis and AADi. TKC reported receiving institutional and personal funds from Analysis Group, AstraZeneca, Alexion, Bayer, Bristol Myers-Squibb/ER Squibb and sons LLC, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Ipsen, Tracon, Genentech, Roche, Roche Products Limited, F. Hoffmann-La Roche, GlaxoSmithKline, Lilly, Merck, Novartis, Peloton, Pfizer, Prometheus Labs, Corvus, Calithera, Sanofi/Aventis, Takeda; reported receiving honoraria from the Analysis Group, AstraZeneca, Alexion, Sanofi/Aventis, Bayer, Bristol Myers-Squibb/ER Squibb and sons LLC, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Genentech, Roche, Roche Products Limited, F. Hoffmann-La Roche, GlaxoSmithKline, Heron Therapeutics, Lilly, Merck, Novartis, Peloton, Pfizer, EMD Serono, Prometheus Labs, Corvus, Ipsen, Up-to-Date, NCCN, Analysis Group, NCCN, Michael J. Hennessy (MJH) Associates, Inc. (Healthcare Communications Company with several brands, such as OnClive, PeerView, and PER), L-path, Kidney Cancer Journal, Clinical Care Options, Platform Q, Navinata Healthcare, Harborside Press, American Society of Medical Oncology, NEJM, Lancet Oncology; having a consulting or advisory role at Analysis Group, AstraZeneca, Alexion, Sanofi/Aventis, Bayer, Bristol Myers-Squibb/ER Squibb and sons LLC, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Genentech, Heron Therapeutics, Lilly, Roche, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, EMD Serono, Prometheus Labs, Corvus, Ipsen, Up-to-Date, NCCN. No speaker’s bureau. No leadership or employment in for-profit companies. Other present or past leadership roles: Director of GU Oncology Division at Dana-Farber and past President of Medical Staff at Dana-Farber), member of NCCN Kidney panel and the GU Steering Committee, past chairman of the Kidney Cancer Association Medical and Scientific Steering Committee). Patents, royalties, or other intellectual properties: International Patent Application No. PCT/US2018/12209, entitled “PBRM1 Biomarkers Predictive of Anti-Immune Checkpoint Response”, filed January 3, 2018, claiming priority to U.S. Provisional Patent Application No. 62/445,094, filed January 11, 2017; International Patent Application No. PCT/US2018/058430, entitled “Biomarkers of Clinical Response and Benefit to Immune Checkpoint Inhibitor Therapy,” filed October 31, 2018, claiming priority to U.S. Provisional Patent Application No. 62/581,175, filed November 3, 2017. Travel, accommodations, expenses, in relation to consulting, advisory roles, or honoraria. Medical writing and editorial assistance support may have been funded by Communications companies funded by pharmaceutical companies (ClinicalThinking, Envision Pharma Group, Fishawack Group of Companies, Health Interactions, and Parexel, others). The institution (Dana-Farber Cancer Institute) may have received additional independent funding of drug companies or/and royalties potentially involved in research around the subject matter. CV provided upon request for the scope of clinical practice and research. EDE, KH, and SN are employees of Invitae Corporation, a testing laboratory that furnished the diagnostic test results used in this study. EDE and KH reported being a shareholder of Invitae. GS: Advisory board: Pfizer, BMS, Genentech, EMD Serono, Novartis, Merck, Sanofi, Seattle Genetics/Astellas, Astrazeneca, Exelixis, Janssen, Amgen, Eisai, Bicycle Therapeutics; research support to institution: Boehringer-Ingelheim, Bayer, Pfizer, Merck, Sanofi, Astrazeneca; Travel costs: BMS, Astrazeneca; speaking fees: Physicians Education Resource (PER), Onclive, Research to Practice, Clinical Care Options; writing fees: Uptodate; steering committee of trials: BMS, Bavarian Nordic, Seattle Genetics, QED (all unpaid), and Astrazeneca and Debiopharm (both paid).
Ethics approval and consent to participate
Western Institutional Review Board protocol number 1167406 waived the requirement to obtain written patient informed consent. The study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Adib, E., El Zarif, T., Nassar, A.H. et al. CDH1 germline variants are enriched in patients with colorectal cancer, gastric cancer, and breast cancer. Br J Cancer 126, 797–803 (2022). https://doi.org/10.1038/s41416-021-01673-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01673-7
This article is cited by
-
circ0005027 Acting as a ceRNA Affects the Malignant Biological Behavior of Hypopharyngeal Squamous Cell Carcinoma by Modulating miR-548c-3p/CDH1 Axis
Biochemical Genetics (2024)
-
Extensive review on breast cancer its etiology, progression, prognostic markers, and treatment
Medical Oncology (2023)
-
Second primary malignancies in non-Hodgkin lymphoma: epidemiology and risk factors
Annals of Hematology (2023)