Abstract
Background
Adoptive immunotherapy with tumour-infiltrating lymphocytes (TIL) may benefit from the use of selective markers, such as PD-1, for tumour-specific T-cell enrichment, and the identification of predictive factors that help identify those patients capable of rendering tumour-reactive TILs. We have investigated this in ovarian cancer (OC) patients as candidates for TIL therapy implementation.
Methods
PD-1− and PD-1+ CD8 TILs were isolated from ovarian tumours and expanded cells were tested against autologous tumour cells. Baseline tumour samples were examined using flow cytometry, multiplexed immunofluorescence and Nanostring technology, for gene expression analyses, as well as a next-generation sequencing gene panel, for tumour mutational burden (TMB) calculation.
Results
Tumour-reactive TILs were detected in half of patients and were exclusively present in cells derived from the PD-1+ fraction. Importantly, a high TIL density in the fresh tumour, the presence of CD137+ cells within the PD-1+CD8+ TIL subset and their location in the tumour epithelium, together with a baseline T-cell-inflamed genetic signature and/or a high TMB, are features that identify patients rendering tumour-reactive TIL products.
Conclusion
We have demonstrated that PD-1 identifies ovarian tumour-specific CD8 TILs and has uncovered predictive factors that identify OC patients who are likely to render tumour-specific cells from PD-1+ TILs.
Similar content being viewed by others
Background
The adoptive transfer of tumour-infiltrating T lymphocytes (TILs) has shown clinical efficacy in patients with metastatic melanoma1 and cervical cancer.2 TIL therapy consists of the ex vivo expansion of T cells from tumour material and transfers back into the same patient after a lymphodepleting preparative regimen. Accumulating evidence supports ovarian cancer (OC) as a candidate for TIL therapy.3,4 However, attempts to treat OC patients with TILs have been generally disappointing.5,6,7
An important factor that determines the success of this therapy is the antitumour reactivity of the infused cellular product.2,8 During the TIL expansion process, there is an inter-clonal competition with different T-cell clones increasing or decreasing in frequency. Therefore, the more tumour-specific clones there are in the starting culture, the greater the chance of maintaining tumour-specific clones at an appreciable frequency in the final product. This may explain why the TIL therapy has yielded objective responses in cancers with high numbers of tumour-specific TILs, such as melanoma9,10 and virus-associated tumours,2 but not in cancers with lower numbers of tumour-specific TILs, such as OC.5,6,7
Surface markers, such as a programmed cell death protein 1 (PD-1) and CD137, have been proven to be able to identify melanoma-specific CD8 TILs.11,12 The CD8+CD137+ population is minority and largely confined to the CD8+PD-1+ TIL subset. For this reason, PD-1 has been proposed as the marker that more comprehensively identifies the repertoire of tumour-specific TILs.11 Recently, we have shown that the enrichment and separate amplification of PD-1+ CD8 TILs improves the antitumour efficacy of TIL therapy in mouse models of solid tumours.13 Our results were further confirmed by Jing et al. in haematological tumours.14
Overall, these findings have important implications since PD-1 may enable the isolation of rare tumour-specific lymphocytes and allow TIL therapy to be extended to less immunogenic tumours. Several pieces of evidence suggest that PD-1 may also demarcate tumour-specific CD8 T cells in ovarian tumours. Thus, the expression of PD-1 in intraepithelial CD8 TILs has been associated with a favourable prognosis in high-grade serous ovarian carcinomas.15 Prior studies have shown that NY-ESO-1-specific CD8+ TILs from OC patients expresses PD-1.16 In addition, ovarian tumours with deficient homologous recombination (HR) have a higher predicted neoantigen (NeoAg) load and infiltrating CD8 T cells in these tumours have increased PD-1 expression.17
Recently, investigation of potential biomarkers that may predict sensitivity to immunotherapy has become an area of active research. In the case of TIL therapy, a number of promising biomarkers have been discovered in the infused cell product that could predict clinical response.18,19 However, little is known about the factors in the original tumour that may help identify patients able to render tumour-reactive TIL products and, therefore, that might be eligible for this therapy.
In this study, we have investigated the role of PD-1 as a selective marker for the pre-enrichment, before expansion, of tumour-specific CD8 TILs in OC. Using multiple molecular and cellular analyses, we have also examined tumours at baseline to identify parameters that help us predict from which patients we can expect to obtain tumour-specific TILs.
Methods
Patients and tumour processing
We evaluated 10 chemotherapy-naive patients with different epithelial OC subtypes, including 7 high-grade serous carcinomas, 2 endometrioid carcinomas and 1 mucinous cystadenocarcinoma, at different cancer stages as described in Supplementary Table S1. The study was approved by the Institutional Review Boards of the Clinica Universidad de Navarra. Tumour fragments (n = 2–3, 0.7–1 cm3) were mechanically and enzymatically disrupted and the single-cell suspension was used to obtain tumour and non-tumour cell-enriched fractions (Supplementary Fig. S1), as detailed in Supplementary Methods. Both fractions were separately cryopreserved until further use.
Flow cytometry (FC) analysis
Fresh tumour single-cell suspensions were incubated with Zombie NIR dye (Biolegend) and, subsequently, were stained with fluorochrome-conjugated monoclonal antibodies (mAbs) against EPCAM, CD45, CD3, CD4, CD8, PD-1 and CD137. Information on mAb clones and fluorochromes is found in Supplementary Methods. Cells were acquired in a FACSCanto-II cytometer (BD Biosciences, Franklin Lakes, NJ, USA) and analysed using FlowJo software (BD Biosciences).
TIL isolation and expansion
Non-tumour cell-enriched fractions were thawed and rested overnight in T-cell media [1:1 mix of RPMI-1640-glutamax (Gibco) and AIMV (Gibco), supplemented with 5% heat-inactivated human serum (SIGMA), 12.5 mM HEPES, 100 U/ml penicillin, 100 μg/ml streptomycin and 10 μg/ml gentamicin (Gibco)]. The next day, cells were stained with fluorochrome-conjugated mAbs against CD8 and PD-1, and 7-amino-actinomycin D (dead cell marker), and were sorted into PD-1-negative (PD-1−) and PD-1 high (PD-1hi) CD8+ T cells using a FACSAria cell sorter (BD Biosciences). Isolated cells were separately expanded by Rapid expansion protocol (REP) in T-cell media containing soluble anti-CD3 mAb (OKT3) (30 ng/ml, Biolegend), human IL-2 (3000 IU/ml, Proleukin) and 3 × 107 irradiated peripheral blood leucocytes pooled from 3 different donors. Detailed information is found in Supplementary Methods and Supplementary Table S2. After 12–15 days of expansion, T cells were cryopreserved until further analysis.
TIL reactivity assessment
Expanded CD8 TILs and autologous tumour-enriched cells were thawed and separately rested overnight in T-cell media without IL-2. The next day, TILs (5 × 104 cells/well) were co-cultured either alone or with target cells [autologous tumour-enriched cells or the H929 plasmacytoma cell line (ATCC), as unrelated tumour cells] (105 cells/well) in ELIIP plates (Millipore) coated with purified anti-human IFN-γ mAb (Mabtech), in the presence or absence of HLA-I blocking mAb (W6/32, Bio-x-cell). Thirty-six hours later, plates were developed as described in Supplementary Methods. The results were expressed as the number of IFN-γ+ spots. When autologous tumour-enriched cells were available, TIL reactivity was confirmed by assessment of CD137. Briefly, TILs (1 × 105 cells/well) were cultured, either alone or with target cells (0.5–1 × 105 cells/well), in T-cell media, and 24 h later, cells were analysed for CD137 expression by FC.
Multiplex immunofluorescence staining and analysis
Multiplex immunofluorescence (IF) staining, validation and analysis are detailed in Supplementary Methods and Supplementary Figs. S2 and S3. Briefly, 4-μm sections of formalin-fixed paraffin-embedded (FFPE) tissue samples were deparaffinised and antigen retrieval was performed using DAKO PT-Link heat-induced antigen retrieval with a low- (pH 6) or high-pH (pH 9) solution (DAKO). Samples were stained with mAbs targeting cytokeratin (CK), CD4, CD8, FOXP3, PD-1 and CD137 followed by TSA visualisation with fluorophores Opal 520, Opal 540, Opal 570, Opal 620, Opal 650 and Opal 690 (Akoya Biosciences), as described earlier.20 Each tissue section was put through several sequential rounds of antibody staining. In the seventh round, nuclei were counterstained with spectral DAPI (Akoya Biosciences) and sections mounted with Faramount Aqueous Mounting Medium (Dako). Multiplexed immunofluorescence slides were scanned on a Vectra-Polaris Automated Quantitative Pathology Imaging System (Akoya Biosciences). Tissue imaging, spectral unmixing and phenotyping were performed using inForm software (version 2.4.8, Akoya Biosciences), as described previously.20
Somatic mutation estimation
The mutational load was estimated using the Trusight Tumour 170 panel (TST170) from Illumina. FFPE-tumour DNA extraction, library preparation and sequencing (NextSeq 500 using high-output cartridge and v2 chemistry) were executed by Macrogen (Korea) according to Illumina’s instructions. Sequence alignment and variant calling were performed using the TST170 BaseSpace application. Variant annotation was performed using Annovar (dbSNP150, 1000G, gnomAD and COSMIC). Positions with depth <100, variant call quality <40, NC (fraction of bases that were uncalled or with base call quality below the minimum threshold) ≥0.03 (for SNVs), strandbias score > −80 and variant allele frequency (VAF) <0.05 were ignored. Germline polymorphism filtering was performed by filtering variant allele frequency (VAF) between 0.4–0.6 and > 0.8. Polymorphisms were excluded if their minor allele frequencies (MAF) in European non-finnish (ENF) population (the ethnic group of our patients) according to gnomAD were > 0. Mutations with MAF(ENF) = 0 or not found in gnomad were also not found in 1000G database. All putative somatic SNVs and INDELs were carefully checked by manual inspection of the sequenced reads using the Integrative Genome Browser (IGV). About 100% of SNVs were “truly SNVs”, whereas 60% of identified somatic indels were “False positive”. Due to the high rate of false-positive Indels and the lower proportion of these variants in OC (~3% of the number of indels + SNVs),21 only SNVs were considered for the mutational load estimation. Tumour mutational burden (TMB) was calculated as the number of somatic SNVs per 0.524 megabase pairs (the targeted genomic regions of TST170). For more detailed information, see Supplementary Methods and Supplementary Tables S3–S11.
NanoString-based gene expression profiling
FFPE-tumour RNA was extracted using the RNeasy FFPE kit (QIAGEN). nCounter PanCancerImmune Profiling panel analysis (Nanostring) was performed by IMIBIC (Spain) following the manufacturer’s instructions. Analyses of expression were performed using nSolver Analysis Software (v 4.0), with normalisation utilising positive and negative control probes, as well as the most stable housekeeping (HK) genes across samples. p values < 0.05 were considered to identify differentially expressed genes (DEG). Volcano plots and unbiased clustering of DEG were generated using R (v3.5.3). For analysis of immune signatures, after HK normalisation, a log10 transformation was applied, and the signature score was calculated by averaging the expression level of those genes included in the IFN-γ signature, Expanded immune signature and T-cell inflamed signature.22 For more information, see Supplementary Methods.
Statistical analysis
The statistical tests used are detailed in each figure legend. For detailed information, see Supplementary Methods.
Results
TILs in fresh ovarian tumours display variable expression of PD-1 and CD137
Single-cell suspensions of fresh human ovarian tumours comprised both CD45+ cells and EpCAM+ cancer cells (Fig. 1 and Supplementary Fig. S4). The percentages of CD4+ and CD8+ cells within the CD45+ population (Supplementary Fig. S5A) ranged from 3.6 to 36.1% and from 5.9 to 31.6%, respectively. CD4+ and CD8+ TILs expressed PD-1 at variable levels (range of 1.73–72.7% and of 0.1–88.6% for CD4+ and CD8+ cells, respectively) (Fig. 1 and Supplementary Fig. S5B). Expression of CD137 was much lower than that of PD-1 and confined to the PD-1+ subsets. Interestingly, CD137 in CD8+ TILs was almost exclusively expressed on PD-1hi cells. The level of CD137 expression within the PD-1+CD4+ and the PD-1+CD8+ subset varied among patients (Supplementary Fig. S5C).
Tumour single-cell suspensions were analysed by FC as detailed in ‘Methods’. Gating strategy is described in Supplementary Fig. S4. The figure shows three representative patients (P05, P06 and P07). Names at the top indicate the parental population. Numbers indicate the percentage of gated cells with respect to the parental population.
CD8 TIL reactivity against autologous tumour was confined to the PD-1high compartment
To determine if PD-1 may enrich tumour-specific T cells in OC, we isolated CD8 TILs with extreme expression of PD-1, namely PD-1− and PD-1hi cells, from 10 resected ovarian tumours and expanded them separately. The number of isolated PD-1− and PD-1hi CD8 TILs varied among patients. Both subsets expanded efficiently (Supplementary Table S2). Next, we tested the ability of the expanded cells (also referred to as TIL products) to recognise the autologous tumour using the enriched tumour cell fraction obtained from enzyme-digested tumours. TIL products were cultured alone or together with autologous tumour cells or unrelated tumour cells (H929) in the presence or absence of HLA-I blockade. Figure 2a, b shows data from patient P05. Notably, cells derived from the PD-1hi CD8 TIL subset, but not from the negative counterparts, were tumour-reactive (TR) cells, as determined by IFN-γ secretion (Fig. 2a) and CD137 upregulation (Fig. 2b). We found TR TILs in 5 out of 10 patients (Fig. 2c and Supplementary Fig. S6). Remarkably, the antitumour reactivity was harboured by cells derived from the PD-1hi compartment, as deduced by IFN-γ ELISPOT. Recognition of autologous tumour by PD-1hi-derived cells was HLA-I-restricted (Fig. 2 and Supplementary Fig. S6). Only PD-1−- derived cells from patient P06 were able to recognise autologous tumour, but this recognition was not HLA-I-restricted (Fig. 2c and Supplementary Fig. S6). Recognition was tumour specific since TILs did not respond to unrelated H929 tumour cells (Fig. 2 and Supplementary Fig. S6). Our data indicate that, although only PD-1+-derived T cells were able to recognise autologous tumour cells, the ability of PD-1-selected cells to render TR TIL products varied among patients. Accordingly, patients were divided into two groups: patients with TR TILs (P01, P02, P04, P05 and P06) and patients with non-tumour-reactive (NTR) TILs (P03, P07, P08, P09 and P10).
Cells expanded from the PD-1− or the PD-1hi CD8 TIL subset isolated from different patients were co-cultured with the respective autologous tumour cells, or with unrelated tumour cells (H929), in the presence or absence of HLA-I blocking antibody (W6/32), and tumour recognition was assessed by measuring IFN-γ release by ELISPOT (a, c) and the frequency of CD137+ cells by FC (b). a, b Antitumour reactivity of cells derived from PD-1− and PD-1hi CD8+ TILs from patient P05. a Left: picture of the ELISPOT plate. Plate-bound anti-CD3 mAb was used as a positive control. Right: the graph shows the number of spots per 5 × 104 cells (mean ± SD). b Dot plots display the frequency of CD137+ cells in PD-1− (up) and PD-1hi (down) derived CD8 TILs 24 h after co-culture with target cells. Cells are gated on blastic CD45+CD3+CD8+cells. c Antitumour reactivity of cells derived from PD-1− and PD-1hi CD8+ TILs from each patient. The number of target-specific IFN-γ spots was determined by calculating the difference between the number of spots generated in the presence of target cells (autologous tumour or nonrelated tumour) and twice the number of spots generated in the absence of targets cells (medium). Reactivities >30 specific IFN-γ spots were considered positive (horizontal dotted line). Each line matches the PD-1− and PD-1hi TIL products derived from the same patient. Mean ± SEM. *P ≤ 0.05. Wilcoxon matched-pair signed-rank test, two-tailed, 95% confidence level.
Detection of CD137+ cells within the PD-1hiCD8+ TIL subset in the fresh tumour correlates with the antitumour reactivity of the final TIL product
We retrospectively analysed FC data from tumour infiltrates to find any feature that distinguished patients from the TR and NTR groups. No statistically significant differences were found in the percentage of CD45+ cells between both groups (Fig. 3a). Notably, those patients delivering TR TILs presented a significantly higher frequency of CD8+ and CD4+ lymphocytes in the fresh tumour (Fig. 3a, b). These patients also showed a higher proportion of PD-1+CD8+ and PD-1+CD4+ TILs (Fig. 3a). Interestingly, while no difference was found in the percentage of CD137+PD-1+CD4+ cells, those patients who rendered TR TILs showed a higher proportion of CD137+PD-1+CD8+ TILs (Fig. 3a). These patients also presented a higher frequency of CD137+ cells within the PD-1hiCD8+ population (Fig. 3b), the cell subset sorted for further expansion. No differences were found either in the percentage of PD-1+ cells within the CD8+ and CD4+ TIL compartment, or in the percentage of CD137+ cells within the PD-1hiCD4+ population (Fig. 3b).
The phenotypic traits of CD4 and CD8 TILs in the fresh tumour (Supplementary Fig. S5) from patients who rendered TR TILs were compared with those from patients with NTR TIL products (as defined in Fig. 3). a Frequency of the indicated subsets in living cells (Zombi NIR−). b Frequency of CD8+, CD4+, PD-1hi and CD137+ cells in the respective parental population. Each dot represents 1 patient’s sample. Mean ± SEM. *P ≤ 0.05; **P ≤ 0.01, Mann–Whitney test, two-tailed, 95% confidence level.
Patients with TR TIL products exhibited a higher percentage of CD137+PD-1+CD8+ TILs in the tumour epithelium
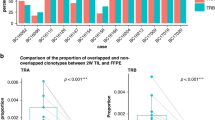
We subsequently analysed tumours by multiplexed IF (Fig. 4). Representative fluorescence images from patients P05 (TR group) and P07 (NTR group) in Fig. 4a show the presence of CD4+ and CD8+ T cells in the tumour infiltrate. CD4+ and CD8+ TILs were more prominent in the tumour stroma than in the tumour epithelium (Fig. 4b, c). PD-1, CD137 and forkhead box P3 (Foxp3) expression was also detected in the tumour infiltrate, with PD-1 being the most notable marker (Fig. 4a).
Tumours were analysed by multiplexed IF using mAbs targeting CK (cyan), CD4 (yellow), CD8 (red), PD-1 (green), CD137 (orange) and Foxp3 (magenta) molecules. a Representative fluorescence images corresponding to patient P05 (TR group) and patient 07 (NTR group). Immunofluorescent multiplexing images were scanned with the Akoya Biosciences’ multispectral image platform. Figures at the top compile the entire staining (merge). b, c Total CD8+ and CD4+ cells, PD-1+CD8+ and PD-1+CD4+ cells (regardless of whether or not they express CD137) and CD137+PD-1+CD8+ and CD137+PD-1+CD4+ cells were quantified using multiplexed quantitative analysis. In order to evaluate TIL subpopulations, first the percentage of each cell population was calculated from the total number of cells in 20 random fields (representing in total 6.95 mm2 of tumour area for each case of ovarian cancer). Then, the average of the percentage of all fields was calculated. This evaluation was done in both the stroma (b) and epithelium (c) compartments. The CD137+PD-1+CD8+ cell population represented a very small subpopulation inside the tumour (nearly null for few patients). Patients who rendered TR TILs were compared with those with NTR TILs. No significant differences were found in the percentage of Foxp3+ TILs, neither in the epithelium nor in the stroma, between both groups (data not shown). Each dot represents 1 patient. Mean ± SEM. Mann–Whitney test, one-tailed, 90% confidence level, Monte Carlo exact p-value estimation.
Since FC analyses showed that patients who rendered TR TILs had a higher percentage of CD137+PD-1+CD8+ cells in the tumour, we also analysed this phenotype by multiplexed quantitative IF. Total CD4+ and CD8+ TILs, PD-1+CD4+ and PD-1+CD8+ TILs (regardless of whether or not they expressed CD137) and CD137+PD-1+CD4+ and CD137+PD-1+CD8+ TIL subsets were quantified (Fig. 4b, c). Multiplexed quantitative IF showed that the percentage of CD137+PD-1+CD8+ cells among the total number of cells was very small. A relatively large tumour area was evaluated (6.95 mm2 of tumour area); thus, CD137+PD-1+CD8+ cells represented a very small cell population in the tumour. Patients with TR TIL products exhibited a slightly higher, but statistically significant, percentage of CD137+PD-1+CD8+ TILs in the tumour epithelium (Fig. 4c).
Antitumour reactivity of the final TIL products was associated with higher mutational load
Somatic mutations may potentially generate neoAgs capable of eliciting a highly potent and tumour-specific immune response. In order to determine if the number of mutations was related to the antitumour reactivity of the final TIL products, we estimated the mutational load of our patients using the TST170 panel as detailed in Supplementary Methods. Of the 2068 variants identified (Supplementary Table S3), 126 were SNVs and only 29 met the criteria for being somatic (Supplementary Table S4 and S5). Mutations were distributed along 26 genes (Fig. 5a). The median of SNVs/Mbs was 3.8, ranging from 0 to 15.3 (Supplementary Table S5). Remarkably, those patients with the highest mutational load (P02 and P06) belonged to the group with TR TILs (Supplementary Table S5). Interestingly, the SNV load of the TR group was borderline significantly higher than that of the NTR group (p = 0.0556) (Fig. 5b).
a Oncoprint displaying non-synonymous somatic SNVs in 26 cancer genes across studied patients (Supplementary Table S4). The digits on the left indicate the percentage of patients who presented mutation in the indicated gene. The bars on the right show the total number of mutations in each gene. b Estimation of SNV load with TST170 panel. Patients who rendered TR TILs were compared with those with NTR TILs. Each dot represents 1 patient. Mann–Whitney test, two-tailed, 95% confidence level.
We also analysed mutations in genes from the DNA damage repair (DDR) system. Defects in these pathways may accumulate an enormous number of mutations due to genome instability during the carcinogenic process and may account for the different mutational load found in our patients. Twenty-eight genes out of about 120 genes involved in direct DDR23 were contained in the TST170 panel (Supplementary Table S9–S11). A total of 53 variants were detected (5.3 ± 1.9) affecting 18 genes (Supplementary Table S9). To identify disturbing mutations affecting these genes, the degree of certainty of pathogenicity was assessed in different databases, as detailed in Supplementary Methods. Only 3 variants met the criteria to be putatively clinically relevant mutations (Supplementary Table S9). Notably, these variants were solely found in those patients with the highest mutational load. Indeed, P02 exhibited clinically relevant mutations in the ATM and FBXW7 genes, whereas P06 presented a likely pathogenic mutation in ERCC2.
Immune activation gene signature at the tumour site identified patients rendering TR TILs
Whereas high values of TMB seemed to be related to the antitumour reactivity of the TIL products, the fact that patients with relatively low TMB values also gave TR TILs suggests that other mechanisms independent of neoantigenicity might also account for the capability of rendering TR TILs. Gene signatures of activated T cells have been shown to predict response to anti-PD-1 therapy.22 In order to determine if the gene expression profile (GEP) may also predict patients capable of eliciting TR TIL products, we analysed the expression of 770 immune-related genes in 8/10 patients with FFPE-tumour RNA available. Four ovaries with non-malignant disease were included as a control. Transcriptome analysis revealed a set of 36 DEG between the TR and NTR groups (Fig. 6a). Unsupervised hierarchical clustering clearly separated patients from TR and NTR groups, as well as the control ovaries (Fig. 6b).
The expression of 770 immune-related genes was profiled using the PanCancerImmune Profiling Panel. Only FFPE-tumour RNA samples that met the quality control requirements [percentage of 300 nucleotide-long RNA fragments >50% and high purity] were included in the study. They included RNAs from 8 patients (P02, P04, P05 and P06, from TR group, and P03, P07, P08 and P10, from NTR group) and 4 ovaries with non-malignant disease (C11–C14). a Volcano plot for DEG between the TR and NTR groups. The coloured circles indicate significant DEG (p < 0.05) with fold change >2 (green) or <0.5 (red). b Heatmap showing the assortment of patients and healthy ovaries (based on the top 36 DEG between TR and NTR groups). Rows represent genes and columns patients and controls. Gene expression was standardised by the mean of the samples. The rows and columns have been grouped using unsupervised hierarchical clustering. c Gene expression signatures associated with TR TIL products. GEP score was calculated by averaging the expression level of the genes included in the IFN-γ signature, Expanded immune signature and T-cell inflamed signature. d GEP score was plotted against the TMB value in each patient. *P ≤ 0.05; **P ≤ 0.01. Mean ± SEM. Mann–Whitney test, two-tailed, 95% confidence level.
Gene Ontology analysis identified biological processes and pathways relevant to the antitumour response that were significantly overrepresented in tumours from the TR group (Supplementary Fig. S7A and B). Interestingly, high baseline expression of genes involved in antigen processing/presentation [PSMB9, TAP1, TAP2, HLA-I (A, B and C) and NLRC5], T-cell activation [CD247 (CD3z), CD3E, CD3D, LCK, CD7, CD38 and ICAM3], proliferation/differentiation of cytotoxic T cells (IL2RB and IL2RG), cytotoxic activity (PRF1 and GZMB), IFN‑γ signalling pathway (STAT1) and chemoattraction (CXCL9, CXCL13, CXCR3 and CCR5), as well as inhibitory mechanisms (HLA-G, FOXP3 and TIGIT), was associated with TR TIL products (Fig. 6b). In addition, patients with TR TIL products exhibited a higher expression of the TNFRSF9 (CD137) transcript that was close to statistical significance (Supplementary Fig. S7C). In contrast, higher expression of genes involved in wound healing (COL3A1, FN1, THBD, THBS1 and TNFRSF1A), mesenchymal transition and cell adhesion (COL3A1 and FN1), angiogenesis and blood vessel development (COL3A1 and THBS1), invasiveness (DOCK9 and THBS1) as well as NF-Κb [IKBKB and TNFRSF1A (TNFR1)] and Ras-MAPK (LRRN3) pathways, was associated with NTR TIL products.
Recently, three immune-related gene expression signatures that predict clinical response to PD-1 blockade across multiple cancer types have been identified: the IFN-γ signature, the Expanded immune gene signature and the T-cell inflamed signature.22 Interestingly, patients rendering TR TIL products exhibited significantly higher scores for the three mentioned signatures, in particular for the T-cell inflamed GEP (Fig. 6c).
Although the correlation between TMB and GEP was low, both parameters stratified TR and NTR patients (Fig. 6d). Therefore, patients with high values of either TMB (P02, P05 and P06) or GEP (P04) rendered TR TIL products, whereas those with low values of both biomarkers (P03, P07, P08 and P10) did not.
Finally, we also analysed the expression of cancer testis antigens (CTA) included in the panel. No differentially expressed CTA genes were detected between the TR and the NTR groups (Supplementary Fig. S7C). Curiously, patient P05, who rendered highly TR TILs (Fig. 2), stood out for expressing a large number of CTA genes.
Discussion
Adoptive immunotherapy using TILs in patients with melanoma1 and cervical2 cancer frequently results in durable complete responses. However, in OC patients, responses to TIL therapy have been less dramatic5,6,7 due in part to technical difficulties in identifying and expanding the TR TIL subset. In an effort to improve TIL therapy in OC, in this study, we have investigated the ability of PD-1 marker for ovarian tumour-specific T-cell enrichment. We have found that autologous tumour-specific T cells were exclusively present in cells derived from PD-1+CD8+ TILs, but not from their PD-1− counterparts. These findings provide a method to preselect ovarian tumour-specific T cells directly from tumour infiltrates, without the need for complex immunological screening technologies that require antigen prediction/identification to evaluate T cells. Recently, it has been shown that PD-1 also allows the identification of tumour-specific T cells in mismatch repair-proficient gastrointestinal cancer patients.24 This finding together with our results indicates that the utility of PD-1 to select for tumour-specific T cells is not restricted to highly immune-reactive tumours.11,13,14,25
However, only in 5 out of 10 patients studied, the PD-1+-derived cells were able to recognise autologous tumour. Interestingly, those patients with TR TIL products exhibited a higher frequency of CD137+ cells within the sorted PD-1+CD8+ population. Our findings reopen the debate on which marker, PD-1 or CD137, best identifies the repertoire of tumour-specific T lymphocytes.11,12 In the only patient studied, Ye et al. found that PD-1+CD137+ cells, but not the PD-1+CD137− subset, harboured tumour-specific T cells in OC.12 These results appear to contradict Rosenberg group’s findings in melanoma,11 which showed that both populations comprised tumour-specific CD8 TILs. CD137 expression is an indication of ongoing or recent activation of T cells,26 in contrast to PD-1 expression, which is maintained long after antigen recognition. On the other hand, the PD-1+ TIL population is heterogeneous and houses bystander and tumour-specific T cells,27 some of which may be recently activated cells (PD-1+CD137+), whereas the large majority are cells activated long ago (PD-1+CD137−). The ratio between bystander and tumour-specific TILs varies between tumours,28 and this could explain why in OC, with smaller numbers of tumour-specific lymphocytes in comparison to melanoma,11 only CD137+ cells12 or PD-1+ cells containing a high proportion of CD137+ cells (our study) were able to render TR TILs. A more exhausted state of PD-1+ tumour-specific TILs in OC may also account for the difficulty of obtaining TR TIL products from this population. Patient-related factors may also be implicated. Thus, similar to what we have found in OC, the ability of PD-1 to enrich for tumour-specific TILs in melanoma patients has also been shown to be inconsistent.25
Therefore, the use of PD-1 as a single marker to isolate tumour-specific cells appears to be controversial. On the other hand, the expression of CD137 is transitory. Consequently, a large proportion of tumour-specific T cells may be missed in the CD137− fraction. A recent study in colorectal and lung cancer has shown that CD39 accurately distinguishes between tumour-specific (PD-1+CD39+) and bystander (PD-1+CD39−) TILs.27 In another study, co-expression of PD-1 with markers of tissue-resident memory cells, such as CD39 and CD103, seemed to identify tumour-specific TILs.29 Presumably, combinations of PD-1 with other markers would be necessary to accurately identify the full repertoire of tumour-specific TILs, especially in tumours in which these cells are rare.
The finding that those patients rendering TR TILs had a higher frequency of CD137+PD-1+CD8+ TILs located in the tumour epithelium, suggests that their tumours may have special features that have enabled the activation of T cells, e.g., the presence of relevant antigens or/and an immune-permissive tumour microenvironment (TME). TMB provides an indirect assessment of tumour antigenicity because a high level of mutations will increase the likelihood of neoAg generation. The TMB values in our patients, as estimated using the TST170 panel, ranged from 0 to 15.3 SNVs/mpb. These values were within the range of those found for OC using whole-exome sequencing (WES), although the median values (3.8 SNVs/Mbp) were higher than those estimated with this method (~1.5 mutations/Mb).30,31 The use of a gene panel enriched in known cancer genes, which are more likely to be mutated in tumours, often leads to TMB overestimation.32 The precise estimation of the mutational load may also have been affected by the low genomic coverage of the TST170 panel (0.53 Mbp), a problem that most seriously affects tumours with low/moderate TMB levels, as is the case of OC.33,34 However, previous studies have shown that the numbers of mutations detected with the TST170 panel strongly correlate (Pearson correlation 0.97) with those observed with the gold standard WES.34 In addition, exploratory analyses demonstrated that the TST170 panel-based TMB discriminates between immunotherapy responders and non-responders.35,36 Therefore, although the panel used cannot provide an accurate estimation of the mutational load, it can still be valid to distinguish between patients. Remarkably, those patients with the highest SNV numbers (P02 and P06) belonged to the TR group. Although our data need to be confirmed in a larger cohort of patients using a higher panel size, they suggest that, in some patients at least, the existence of possible neoAgs might account for the capability of rendering TR TILs.
Several intrinsic mechanisms, including deficiencies in DDR genes, have been implicated in contributing to neoAg load.37 Interestingly, patient P02 showed deleterious mutations in ATM and FBXW7 genes, involved in HR and non-homologous end-joining pathways, respectively, whereas patient P06 presented a likely pathogenic mutation in ERCC2, entailed in the nucleotide excision repair pathway. ATM gene defects strongly correlate with spontaneous TIL infiltration in breast cancer and OC.38 Recently, it has been shown that bladder cancer patients with mutated ATM had better benefits from immune checkpoint inhibitor (ICI) therapy.39 Inactivation of FBXW7 causes genomic instability.40 Importantly, mutations in ATM and FBXW7 genes correlate with higher TMB levels in several cancer types, including OC.41,42,43,44,45,46,47 Mutated ERCC2 has also been associated with high TMB values.41 Although our findings require further investigation, they suggest that deleterious alterations in DDR genes may influence the competence to render TR TILs by leading to an increased TMB, and subsequently increased number of neoAgs.
The finding that some patients from the TR group showed relatively low mutational load suggests that mechanisms independent of neoantigenicity might also account for the capability of rendering TR TILs. The expression profile of immune-related genes reflects the nature of the TME, which is also an important feature of cancer immunobiology. Interestingly, tumours from patients with TR TIL products showed a high expression of genes involved in antigen processing/presentation, T-cell activation, proliferation/differentiation of T cells, cytotoxic activity, IFNγ-signalling, chemoattraction as well as mechanisms to regulate T-cell response. This GEP is characteristic of immune-reactive tumours with prolonged survival and response to ICIs.48,49 A similar GEP has also been associated with high tumour T-cell infiltration in OC patients.50,51 In line with the transcriptomic data, FC analysis of tumour infiltrates revealed a significantly higher frequency of CD8 and CD4 T cells in those patients who rendered TR TILs. Likewise, a high expression of genes involved in wound healing, mesenchymal transition, extracellular matrix remodelling, angiogenesis and invasiveness, was associated with the lack of antitumour reactivity of the final TIL product. The concurrent overexpression of genes involved in these processes has been associated with the presence of immunosuppressive TME,52 T-cell dysfunctionality53 and resistance to anti-PD-1 therapy.48
Three multigene expression signatures (IFN-γ, expanded immune and T-cell-inflamed signature) have been associated with clinical benefit from anti-PD-1 treatment in several cancer types.22 Importantly, patients rendering TR TIL products exhibited significantly higher scores for the three mentioned signatures, in particular for the T-cell inflamed GEP. These findings suggest that GEP may represent a useful tool to identify patients who are likely to render TR TILs and, ultimately, who may benefit from TIL therapy.
Interestingly, while the TMB values did not correlate with the GEP score, both parameters stratified TR and NTR groups. Thereby, patients with high values of either TMB and/or GEP rendered TR TIL products, whereas those with low values of both TMB and GEP did not. Our data show some similarities with the predictive value of TMB and GEP to anti-PD-1 therapy across different tumour types.54,55 The ability of these two parameters to reflect overlapping but not always coincident aspects of tumour immunology may explain their lack of correlation.
The fact that patients with relatively low TMB rendered TR TILs suggests that the quality of neoAgs, but not the quantity, or the presence of relevant shared tumour-associated antigens, may have determined the antitumour reactivity of the final TIL product. In the case of OC, several groups have identified TILs specific for private56 and hot-spot mutations,57 as well as for CTA.4,16 Interestingly, the patient who in our study stood out for rendering highly TR TILs (P05), exhibited a high expression of very potent CTA, such as NY-ESO-1, MAGEC2 and TPTE. In the future, it will be interesting to determine what type of antigens account for the antitumour reactivity of TIL products in OC.
The present study has notable limitations, namely the low number of patients and the inclusion of distinct histological subtypes of epithelial OC. Although our findings warrant additional investigation and need to be validated in larger-scale studies, they provide a rationale for further exploring the utility of biomarkers to (i) identify and selectively expand tumour-specific TILs, and (ii) serve as guides for selecting patients eligible for TIL therapy. In order to increase the tumour reactivity of the final TIL product, other selective markers may be considered together with PD-1 to distinguish tumour-specific from bystander T cells. The combination of several parameters, such as TIL density and the presence of intraepithelial CD137+ cells in the fresh tumour together with the GEP and the TMB, could guide future steps to defining an extensive predictive biomarker panel to identify patients eligible for TIL therapy. The use of such a predictive biomarker panel together with newer ways to select tumour-specific TILs, will help to extend this cell therapy to less immune-reactive tumours, such as OC.
References
Goff, S. L., Dudley, M. E., Citrin, D. E., Somerville, R. P., Wunderlich, J. R., Danforth, D. N. et al. Randomized, prospective evaluation comparing intensity of lymphodepletion before adoptive transfer of tumor-infiltrating lymphocytes for patients with metastatic melanoma. J. Clin. Oncol. 34, 2389–2397 (2016).
Stevanović, S., Draper, L. M., Langhan, M. M., Campbell, T. E., Kwong, M. L., Wunderlich, J. R. et al. Complete regression of metastatic cervical cancer after treatment with human papillomavirus-targeted tumor-infiltrating T cells. J. Clin. Oncol. 33, 1543–1550 (2015).
Sato, E., Olson, S. H., Ahn, J., Bundy, B., Nishikawa, H., Qian, F. et al. Intraepithelial CD8+ tumor-infiltrating lymphocytes and a high CD8+/regulatory T cell ratio are associated with favorable prognosis in ovarian cancer. Proc. Natl Acad. Sci. USA 102, 18538–18543 (2005).
Westergaard, M. C. W., Andersen, R., Chong, C., Kjeldsen, J. W., Pedersen, M., Friese, C. et al. Tumour-reactive T cell subsets in the microenvironment of ovarian cancer. Br. J. Cancer 120, 424–434 (2019).
Aoki, Y., Takakuwa, K., Kodama, S., Tanaka, K., Takahashi, M., Tokunaga, A. et al. Use of adoptive transfer of tumor-infiltrating lymphocytes alone or in combination with cisplatin-containing chemotherapy in patients with epithelial ovarian cancer. Cancer Res. 51, 1934–1939 (1991).
Freedman, R. S., Edwards, C. L., Kavanagh, J. J., Kudelka, A. P., Katz, R. L., Carrasco, C. H. et al. Intraperitoneal adoptive immunotherapy of ovarian carcinoma with tumor-infiltrating lymphocytes and low-dose recombinant interleukin-2: A pilot trial. J. Immunother. 16, 198–210 (1994).
Pedersen, M., Westergaard, M. C. W., Milne, K., Nielsen, M., Borch, T. H., Poulsen, L. G. et al. Adoptive cell therapy with tumor-infiltrating lymphocytes in patients with metastatic ovarian cancer: a pilot study. Oncoimmunology 7, e1502905 (2018).
Rothermel, L. D., Sabesan, A. C., Stephens, D. J., Chandran, S. S., Paria, B. C., Srivastava, A. K. et al. Identification of an immunogenic subset of metastatic uveal melanoma. Clin. Cancer Res. 22, 2237–2249 (2016).
Besser, M. J., Shapira-Frommer, R., Treves, A. J., Zippel, D., Itzhaki, O., Schallmach, E. et al. Minimally cultured or selected autologous tumor-infiltrating lymphocytes after a lympho-depleting chemotherapy regimen in metastatic melanoma patients. J. Immunother. 32, 415–423 (2009).
Dudley, M. E., Gross, C. A., Langhan, M. M., Garcia, M. R., Sherry, R. M., Yang, J. C. et al. CD8+ enriched ‘Young’ tumor infiltrating lymphocytes can mediate regression of metastatic melanoma. Clin. Cancer Res. 16, 6122–6131 (2010).
Gros, A., Robbins, P. F., Yao, X., Li, Y. F., Turcotte, S., Tran, E. et al. PD-1 identifies the patient-specific CD8(+) tumor-reactive repertoire infiltrating human tumors. J. Clin. Invest 124, 2246–2259 (2014).
Ye, Q., Song, D. G., Poussin, M., Yamamoto, T., Best, A., Li, C. et al. CD137 accurately identifies and enriches for naturally occurring tumor-reactive T cells in tumor. Clin. Cancer Res. 20, 44–55 (2014).
Fernandez-Poma, S. M., Salas-Benito, D., Lozano, T., Casares, N., Riezu-Boj, J.-I., Mancheño, U. et al. Expansion of Tumor-Infiltrating CD8 + T cells Expressing PD-1 Improves the Efficacy of Adoptive T-cell Therapy. Cancer Res. 77, 3672–3684 (2017).
Jing, W., Gershan, J. A., Blitzer, G. C., Palen, K., Weber, J., McOlash, L. et al. Adoptive cell therapy using PD-1+ myeloma-reactive T cells eliminates established myeloma in mice. J. Immunother. Cancer 5, 51 (2017).
Webb, J. R., Milne, K. & Nelson, B. H. PD-1 and CD103 are widely coexpressed on prognostically favorable intraepithelial CD8 T cells in human ovarian cancer. Cancer Immunol. Res. 3, 926–935 (2015).
Matsuzaki, J., Gnjatic, S., Mhawech-Fauceglia, P., Beck, A., Miller, A., Tsuji, T. et al. Tumor-infiltrating NY-ESO-1-specific CD8+ T cells are negatively regulated by LAG-3 and PD-1 in human ovarian cancer. Proc. Natl Acad. Sci. USA 107, 7875–7880 (2010).
Strickland, K. C., Howitt, B. E., Shukla, S. A., Rodig, S., Ritterhouse, L. L., Liu, J. F. et al. Association and prognostic significance of BRCA1/2-mutation status with neoantigen load, number of tumor-infiltrating lymphocytes and expression of PD-1/PD-L1 in high grade serous ovarian cancer. Oncotarget 7, 13587–13598 (2016).
Zhou, J., Shen, X., Huang, J., Hodes, R. J., Rosenberg, S. A. & Robbins, P. F. Telomere length of transferred lymphocytes correlates with in vivo persistence and tumor regression in melanoma patients receiving cell transfer therapy. J. Immunol. 175, 7046–7052 (2005).
Rosenberg, S. A., Yang, J. C., Sherry, R. M., Kammula, U. S., Hughes, M. S., Phan, G. Q. et al. Durable complete responses in heavily pretreated patients with metastatic melanoma using T-cell transfer immunotherapy. Clin. Cancer Res. 17, 4550–4557 (2011).
Abengozar-Muela, M., Esparza, M. V., Garcia-Ros, D., Vásquez, C. E., Echeveste, J. I., Idoate, M. A. et al. Diverse immune environments in human lung tuberculosis granulomas assessed by quantitative multiplexed immunofluorescence. Mod. Pathol. https://doi.org/10.1038/s41379-020-0600-6 (2020).
Turajlic, S., Litchfield, K., Xu, H., Rosenthal, R., McGranahan, N., Reading, J. L. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Ayers, M., Lunceford, J., Nebozhyn, M., Murphy, E., Loboda, A., Kaufman, D. R. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Chae, Y. K., Anker, J. F., Carneiro, B. A., Chandra, S., Kaplan, J., Kalyan, A. et al. Genomic landscape of DNA repair genes in cancer. Oncotarget 7, 23312–23321 (2016).
Gros, A., Tran, E., Parkhurst, M. R., Ilyas, S., Pasetto, A., Groh, E. M. et al. Recognition of human gastrointestinal cancer neoantigens by circulating PD-1+ lymphocytes. J. Clin. Invest. 129, 4992–5004 (2019).
Inozume, T., Hanada, K., Wang, Q. J., Ahmadzadeh, M., Wunderlich, J. R., Rosenberg, S. A. et al. Selection of CD8+PD-1+ lymphocytes in fresh human melanomas enriches for tumor-reactive T cells. J. Immunother. 33, 956–964 (2010).
Wolfl, M., Kuball, J., Ho, W. Y., Nguyen, H., Manley, T. J., Bleakley, M. et al. Activation-induced expression of CD137 permits detection, isolation, and expansion of the full repertoire of CD8+ T cells responding to antigen without requiring knowledge of epitope specificities. Blood 110, 201–210 (2007).
Simoni, Y., Becht, E., Fehlings, M., Loh, C. Y., Koo, S. L., Teng, K. W. W. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Scheper, W., Kelderman, S., Fanchi, L. F., Linnemann, C., Bendle, G., de Rooij, M. A. J. et al. Low and variable tumor reactivity of the intratumoral TCR repertoire in human cancers. Nat. Med 25, 89–94 (2019).
Duhen, T., Duhen, R., Montler, R., Moses, J., Moudgil, T., De Miranda, N. F. et al. Co-expression of CD39 and CD103 identifies tumor-reactive CD8 T cells in human solid tumors. Nat. Commun. 9, 2724 (2018).
Tian, W., Shan, B., Zhang, Y., Ren, Y., Liang, S., Zhao, J. et al. Association between DNA damage repair gene somatic mutations and immune-related gene expression in ovarian cancer. Cancer Med. https://doi.org/10.1002/cam4.2849 (2020).
Alexandrov, L. B., Nik-Zainal, S., Wedge, D. C., Aparicio, S. A. J. R., Behjati, S., Biankin, A. V. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Fancello, L., Gandini, S., Pelicci, P. G. & Mazzarella, L. Tumor mutational burden quantification from targeted gene panels: Major advancements and challenges. J. Immunother. Cancer https://doi.org/10.1186/s40425-019-0647-4 (2019).
Buchhalter, I., Rempel, E., Endris, V., Allgäuer, M., Neumann, O., Volckmar, A. L. et al. Size matters: dissecting key parameters for panel-based tumor mutational burden analysis. Int J. Cancer 144, 848–858 (2019).
Budczies, J., Allgäuer, M., Litchfield, K., Rempel, E., Christopoulos, P., Kazdal, D. et al. Optimizing panel-based tumor mutational burden (TMB) measurement. Ann. Oncol. 30, 1496–1506 (2019).
Zhang, S., So, A. S., Kaplan, S., Kruglyak, K. M. Comprehensive evaluation of Illumina’s TruSight® Tumor 170 Panel to estimate tumor mutational burden. Abstract 5358. Cancer Res. 2017, 5358–5358 (2017)
Endris, V., Buchhalter, I., Allgäuer, M., Rempel, E., Lier, A., Volckmar, A. L. et al. Measurement of tumor mutational burden (TMB) in routine molecular diagnostics: in silico and real-life analysis of three larger gene panels. Int J. Cancer 144, 2303–2312 (2019).
Ma, J., Setton, J., Lee, N. Y., Riaz, N. & Powell, S. N. The therapeutic significance of mutational signatures from DNA repair deficiency in cancer. Nat. Commun. 9, 1–12 (2018).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Yi, R., Lin, A., Cao, M., Xu, A., Luo, P. & Zhang, J. ATM Mutations benefit bladder cancer patients treated with immune checkpoint inhibitors by acting on the tumor immune microenvironment. Front Genet. 11, 933 (2020).
Zhang, Q., Karnak, D., Tan, M., Lawrence, T. S., Morgan, M. A. & Sun, Y. FBXW7 Facilitates nonhomologous end-joining via K63-linked polyubiquitylation of XRCC4. Mol. Cell 61, 419–433 (2016).
Mamdani, H., Chen, J., Kim, S., Ibrahim, Y., Asad, M. F. B., Nieva, J. J. et al. DNA damage response and repair (DDR) gene mutations and correlation with tumor mutation burden (TMB) in non-small cell lung cancer (NSCLC). J. Clin. Oncol. 37, 9100–9100 (2019).
Ricciuti, B., Cheng, M. L., Recondo, G., Nishino, M., Umeton, R., Sholl, L. M. et al. DNA damage response gene alterations are associated with high tumor mutational burden and clinical benefit from programmed death 1 axis inhibition in non-small cell lung cancer. J. Clin. Oncol. 37, 9077–9077 (2019).
Tian, W., Shan, B., Zhang, Y., Ren, Y., Liang, S., Zhao, J. et al. Association of high tumor mutation (TMB) with DNA damage repair (DDR) alterations and better prognosis in ovarian cancer. J. Clin. Oncol. 36, 5512–5512 (2018).
Sun, L., Wang, R. C., Zhang, Q. & Guo, L. L. ATM mutations as an independent prognostic factor and potential biomarker for immune checkpoint therapy in endometrial cancer. Pathol. Res. Pract. 216, 153032 (2020).
Wang, M., Fan, W., Ye, M., Tian, C., Zhao, L., Wang, J. et al. Molecular profiles and tumor mutational burden analysis in Chinese patients with gynecologic cancers. Sci. Rep. 8, 1–9 (2018).
Ali, S. M., Yao, M., Yao, J., Wang, J., Cheng, Y., Schrock, A. B. et al. Comprehensive genomic profiling of different subtypes of nasopharyngeal carcinoma reveals similarities and differences to guide targeted therapy. Cancer 123, 3628–3637 (2017).
Eder, T., Hess, A. K., Konschak, R., Stromberger, C., Jöhrens, K., Fleischer, V. et al. Interference of tumour mutational burden with outcome of patients with head and neck cancer treated with definitive chemoradiation: a multicentre retrospective study of the German Cancer Consortium Radiation Oncology Group. Eur. J. Cancer 116, 67–76 (2019).
Hugo, W., Zaretsky, J. M., Sun, L., Song, C., Moreno, B. H., Hu-Lieskovan, S. et al. Genomic and transcriptomic features of response to Anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Chen, P. L., Roh, W., Reuben, A., Cooper, Z. A., Spencer, C. N., Prieto, P. A. et al. Analysis of immune signatures in longitudinal tumor samples yields insight into biomarkers of response and mechanisms of resistance to immune checkpoint blockade. Cancer Disco. 6, 827–837 (2016).
Leffers, N., Fehrmann, R. S. N., Gooden, M. J. M., Schulze, U. R. J., Ten Hoor, K. A., Hollema, H. et al. Identification of genes and pathways associated with cytotoxic T lymphocyte infiltration of serous ovarian cancer. Br. J. Cancer 103, 685–692 (2010).
Dangaj, D., Bruand, M., Grimm, A. J., Ronet, C., Barras, D., Duttagupta, P. A. et al. Cooperation between Constitutive and Inducible Chemokines Enables T Cell engraftment and immune attack in solid tumors. Cancer Cell 35, 885–900.e10 (2019).
Sangaletti, S., Chiodoni, C., Tripodo, C. & Colombo, M. P. Common extracellular matrix regulation of myeloid cell activity in the bone marrow and tumor microenvironments. Cancer Immunol. Immunother. 66, 1059–1067 (2017).
Chen, L., Gibbons, D. L., Goswami, S., Cortez, M. A., Ahn, Y. H., Byers, L. A. et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat. Commun. 5, 5241 (2014).
Cristescu, R., Mogg, R., Ayers, M., Albright, A., Murphy, E., Yearley, J. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science https://doi.org/10.1126/science.aar3593 (2018).
Ott, P. A., Bang, Y. J., Piha-Paul, S. A., Abdul Razak, A. R., Bennouna, J., Soria, J. C. et al. T-cell–inflamed gene-expression profile, programmed death ligand 1 expression, and tumor mutational burden predict efficacy in patients treated with pembrolizumab across 20 cancers: KEYNOTE-028. J. Clin. Oncol. 37, 318–327 (2019).
Bobisse, S., Genolet, R., Roberti, A., Tanyi, J. L., Racle, J., Stevenson, B. J. et al. Sensitive and frequent identification of high avidity neo-epitope specific CD8 + T cells in immunotherapy-naive ovarian cancer. Nat. Commun. 9, 1092 (2018).
Deniger, D. C., Pasetto, A., Robbins, P. F., Gartner, J. J., Prickett, T. D., Paria, B. C. et al. T-cell responses to TP53 ‘Hotspot’ Mutations and unique neoantigens expressed by human ovarian cancers. Clin. Cancer Res. 24, 5562–5573 (2018).
Salas-Benito, D., Conde, E., Tamayo-Uria, I., Mancheño, U., Elizalde, E., Garcia-Ros, D. et al. The mutational load and a T-cell inflamed tumor phenotype identify ovarian cancer patients rendering tumor-reactive T cells from PD-1+ tumor-infiltrating lymphocytes. Research Square. https://doi.org/10.21203/rs.3.rs-25670/v1 (2020)
Acknowledgements
We particularly acknowledge all patients for their participation and V. Villar (Biobank, University of Navarra), Dr. D. Alignani and I. Rodriguez (Flow Cytometry Unit, CIMA), Dr. A. Patiño (Genomic Medicine Unit, Clinica Universidad de Navarra) and Dr. A. Jimenez-Arranz (Unidad de Genómica, IMIBIC, Cordoba, Spain), for their collaboration in this study. Members of Biobank of University of Navarra and Hospital Costa del Sol, and Genomics Unit of CIMA are also acknowledged. We are grateful to Dr. P.W. Miller for English editing. We also acknowledge Research Square platform for the free preprint posting of our original submission.58
Author information
Authors and Affiliations
Contributions
D.S.B. and M.P.S. were involved in study conceptualisation, acquired ethical approvals for the study and curated patient data. D.S.B. also analysed FC, multiplex IF and genetic data, and wrote the original paper draft. E.C. and E.E. performed DNA and RNA isolation from FFPE tissues. E.C. and I.T.U. performed bioinformatics and statistical analysis and analysed publicly available datasets. U.M. and E.E. performed tissue processing, cell isolation and expansion, FC staining and TIL reactivity assays. D.G.R. performed multiplex IF staining. J.M.A., J.A., J.A.M. and J.A.T. were involved in tissue procurement. F.G.G. was involved in statistical analysis. J.C.M., A.G.M., P.S., J.J.L. and C.E.A. were involved in study conceptualisation. C.E.A. also developed and validated multiplex quantitative IF staining methodology, analysed IF data and verified the adequacy of tumour in FFPE samples. S.H.S. conceived the study, performed the experimental design, oversaw the project, wrote and edited the final paper. J.J.L., C.E.A. and S.H.S. acquired funding to perform this research. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The study was performed in accordance with the Declaration of Helsinki and approved by the Ethics and Scientific Committee of the Clinica Universidad de Navarra and Hospital Costa de Sol (2016.145). Written informed consent was obtained from all patients. Samples and data from patients included in the study were provided by the Biobank of the University of Navarra and Hospital Costa del Sol and were processed following standard operating procedures approved by the Ethics and Scientific Committees.
Consent to publish
Not applicable.
Data availability
All data generated or analysed during this study are included in this published article and its Supplementary files.
Competing interests
The authors declare no competing interests.
Funding information
S.H.S. is founded by ISCIII/FEDER, UE (PI15/02027, PI18/00556), FEDER/MICIU-AEI/RTC-2017-6745-1, Fundación AECC (IDEAS18121HERV) and Gobierno de Navarra grants [Ayudas a Centros Tecnológicos (PT039 EstraTeC) and Departamento de Salud (045-2017)] co-financed (50%) with FEDER funds (UE, FEDER 2014–2020 “Una manera de hacer Europa”). E.C., J.J.L. and S.H.S. are funded by Fundación Ramón Areces. C.E.A. and D.G.R. are funded by the Instituto de Salud Carlos III (AC14/00034) y co-financed by Fondos FEDER (Spain).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salas-Benito, D., Conde, E., Tamayo-Uria, I. et al. The mutational load and a T-cell inflamed tumour phenotype identify ovarian cancer patients rendering tumour-reactive T cells from PD-1+ tumour-infiltrating lymphocytes. Br J Cancer 124, 1138–1149 (2021). https://doi.org/10.1038/s41416-020-01218-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-020-01218-4
This article is cited by
-
Immunomic longitudinal profiling of the NeoPembrOv trial identifies drivers of immunoresistance in high-grade ovarian carcinoma
Nature Communications (2024)
-
Improving the cytotoxic response of tumor-infiltrating lymphocytes towards advanced stage ovarian cancer with an oncolytic adenovirus expressing a human vIL-2 cytokine
Cancer Gene Therapy (2023)
-
Tumor-infiltrating lymphocyte treatment for anti-PD-1-resistant metastatic lung cancer: a phase 1 trial
Nature Medicine (2021)