Abstract
Background
Breast fibroadenoma (FA) and phyllodes tumour (PT) often have variations of gene mediator complex subunit 12 (MED12) and mutations in the telomerase reverse transcriptase promoter region (TERTp). TERTp mutation is usually tested by Sanger sequencing. In this study, we compared Sanger sequencing and droplet-digital PCR (ddPCR) to measure TERTp mutations in FA and PT samples.
Methods
FA and PT samples were collected from 82 patients who underwent surgery at our institution from 2005 to 2016. MED12 mutations for all cases and TERTp mutations for 17 tumours were detected by Sanger sequencing. ddPCR was performed to analyse TERTp mutation in all cases.
Results
A total of 75 samples were eligible for analysis. Sanger sequencing detected MED12 mutations in 19/44 FA (42%) and 21/31 PT (68%). Among 17 Sanger sequencing-tested samples, 2/17 (12%) were TERTp mutation-positive. In ddPCR analyses, a significantly greater percentage of PT (19/31, 61%) was TERTp mutation-positive than was FA (13/44, 30%; P = 0.0046). The mutation positivity of TERTp and MED12 did not correlate, in either FA or PT.
Conclusions
ddPCR was more sensitive for detecting TERTp mutation than Sanger sequencing, being able to elucidate tumorigenesis in FA and PT.
Similar content being viewed by others
Background
Breast fibroepithelial tumours, which include fibroadenoma (FA) and phyllodes tumour (PT), are characterised by the biphasic proliferation of both epithelial and stromal components.1,2,3,4,5 FA is common benign tumour that is often observed in young women. It expresses oestrogen receptor (ER)-α in epithelium and ERβ in stromal components, and is known to be hormone dependent.6,7 Small FA (<3 cm) is usually followed up without resection, and 16–37% of FA cases reportedly regress or completely resolve spontaneously.8,9,10 PT is much less common than FA; it comprises only 2–3% of fibroepithelial breast tumours and accounts for <1% of all breast tumours. PT is histologically classified as benign, borderline or malignant type. Resection is generally recommended for PT because it often grows rapidly and can potentially become malignant. Occasionally, PT enlarges to huge sizes that require total mastectomy, and malignant PT has high risks of both local recurrences and distant metastasis compared with the other types of PT.1,2,3,10,11,12,13,14,15
FA and PT, especially when they remain relatively small, are morphologically so similar that differentiating them histologically is sometimes difficult. Some studies have suggested that FA could potentially progress to PT, especially FA with monoclonal stromal components; however, the mechanisms underlying their initiation and progression are unclear.10,11,16,17 Although FA and PT have been genomically analysed,18,19,20,21 little had been known about their genetic abnormalities. However, since next-generation sequencing (NGS) became widely used in research, several genetical alterations have been revealed in both tumours. The discovery of highly recurrent Mediator complex subunit 12 (MED12) somatic mutations in breast FA was surprising; almost nothing had previously been known about these mutations.22 As for PT, in addition to MED12,10,11,12,13,23,24,25,26,27 the telomerase reverse transcriptase (TERT) promoter has been shown to have repeated mutations in these tumours.28,29,30,31,32,33,34
In 2014, Lim et al. first found highly frequent MED12 exon 2 mutations in FA (58/98, 59%) using exome analysis.22 The MED12 gene, located on the X chromosome, encodes MED12 protein, a member of the multiprotein mediator complex that regulates transcription of all RNA polymerase II-dependent genes.35 Reportedly, up to 60% of FA, 80% of benign and borderline PT and 40% of malignant PT harbours somatic mutations in exon 2 of the MED12 gene,10,11,12,13,22,23,24,25,26,27 which suggests that FA and PT have much more in common in their origin or development than previously thought. However, the underlying mechanisms, and how this mutation generates or induces the progression of FA and PT, are unknown.
Since the discovery of frequent MED12 mutations in FA and PT, next-generation sequencing has been used to search for other gene mutations in these tumours. Many mutations found in PT, such as in RARA, EGFR, RB1 or TP53, are very uncommon compared with MED12 mutations,12,21,27,28,29,30 whereas mutations in the TERT promoter region (TERTp) are reportedly more frequent: 0–7% in FA, and 27–70.6% in PT.29,30,31,32,33,34 TERTp is considered to be a critical regulatory element for telomerase expression.35,36 Hotspots for mutations in this region in PT and FA are reported to be c.−146 C > T (C250T) and c.−124 C > T (C228T),29,31,32,33,34 which are concordant with those in other tumours, including central nervous system tumours, thyroid cancers, bladder cancers and skin melanoma.36,37 Previous studies found correlations between TERTp mutations (TERTpMut) and MED12 mutations (MED12Mut),28,29,30,31 which suggests that these mutations interact, especially in the development of PT.
Recently, McEvoy et al. showed that droplet-digital polymerase chain reaction (ddPCR) is very sensitive in detecting TERTpMut in melanoma compared with pyrosequencing or Sanger sequencing (SS).38,39 Digital PCR was developed to yield absolute measures for nucleic acid concentrations by a combination of limiting dilution, end-point PCR and Poisson statistics.40 ddPCR is a newer, more precise and less subjective assay to quantify DNA amplification, based on water–oil emulsion droplet technology. In ddPCR, a sample is fractionated into 20,000 droplets, and PCR amplification of the template molecules occurs in each individual droplet. ddPCR has also been shown to obtain high levels of partitioning at a low cost.41,42
TERTpMut was previously evaluated with ddPCR, but only in melanoma.38,39 We found no reports of studies that evaluated mutations in breast fibroepithelial lesions using ddPCR. In this study, we used ddPCR to measure TERTpMut in formalin-fixed, paraffin-embedded (FFPE) samples of resected FA or PT, and compared the results with those from conventional SS. We also analysed the relationships among TERTpMut and MED12Mut status and histopathological characteristics in FA and PT.
Methods
Tissue samples
We collected FFPE samples of 54 FA and 31 PT from 82 patients who underwent surgery at the University of Tokyo Hospital from 2005 to 2016. Three patients had two tumours excised at different times. All samples were diagnosed by two expert pathologists. FA was classified as intracanalicular type, pericanalicular type, mastopathic type, organoid type,43,44 complex fibroadenoma45 or juvenile fibroadenoma;46 PT was subclassified according to the WHO classification as benign, borderline or malignant lesions.2 All patients consented to the use of their stored tumour tissue. This study protocol was approved by the ethics committee at the University of Tokyo Hospital, Tokyo, Japan.
DNA extraction
For each specimen, two or three 10-µm FFPE sections were cut from a single representative block per case. Macrodissection was performed with a scalpel as needed to adjust the tumour content to be visually more than 20%. Microdissection was not performed. DNA was isolated using a GeneRead DNA FFPE Kit (Qiagen, Hilden, Germany), in accordance with the manufacturer’s instructions. Purified DNA was quantified using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA), with 0.5–14.0 µg of DNA recovered per section.
Sanger sequencing to detect MED12 mutation
FA and PT were analysed for mutations in exon 2 of MED12 by SS for all samples. Exon 2 was amplified with the following primers: (exon 2 forward) 5′-AACTAAACGCCGCTTTCCTG-3′, (exon 2 reverse) 5′-TTCCTTCAGCCTGGCAGAG-3′10,47 (Supplementary Table S1). The PCR products were purified using agarose gel electrophoresis, labelled with Big Dye Terminator (Applied Biosystems, Foster City, CA, USA) with bidirectional primers and subjected to 3130 × l Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) in accordance with standard protocols. The Catalogue of Somatic Mutations in Cancer (COSMIC) database48 was used to identify already-known somatic mutations and mutation types.
Sanger sequencing for detecting TERTp mutations
TERTp mutations (C228T and C250T) were analysed in FA and PT with the same extracted DNA as used for the MED12Mut analysis. We performed SS on 17 samples, which had been surgically resected in 2015 and 2016. TERTp was amplified with the following primers: (promoter forward) 5′-AGCGCTGCCTGAAACTCG-3′, (promoter reverse) 5′-CCTGCCCCTTCACCTTCCAG-3′31,34 (Supplementary Table S1). Q-Solution (Qiagen) was added to the reaction to amplify TERTp by PCR, in accordance with previous reports.38,39 The SS method was the same as that for the MED12Mut analysis described above.
ddPCR for detecting TERTp mutations
Analysis of TERTp mutations (C228T and C250T) with ddPCR was performed for all of the samples, using a previously described ddPCR method.38,39 Primers were the same as those used in SS (Supplementary Table S1). Two probes were used to detect the mutations: one (the mutant probe) was designated as “/56-FAM/CCC + C + T + T + CCGG/3IABkFQ/” to detect both C228T and C250T mutations (as both mutations result in the same sequencing string), and the other (the wild probe) was designated as “/5HEX/CCCC + C + T + CCGG/3IABkFQ/,” to recognise the C228 locus (Supplementary Fig. S1). The probes were custom-synthesised by Integrated DNA Technologies (Coralville, IA, USA). PCR reactions were performed in 20-mL reactions that contained 10 µL of Bio-Rad 2× ddPCR Supermix for Probes (no dUTP) (Bio-Rad, Hercules, CA, USA), 250 nmol/L probe, 900 nmol/L primers, 10–290 ng of DNA and water. Reaction mixtures were partitioned into emulsions of ~20,000 droplets in oil using a QX200 Droplet Generator (Bio-Rad). The droplets were then transferred to a 96-well PCR plate, heat-sealed and placed in a thermal cycler (Bio-Rad PX1). Droplets were generated and analysed using the QX200 Droplet Digital PCR System (Bio-Rad). The thermal cycling conditions were 1 cycle at 95 °C (2.5 °C/s ramp) for 10 min, 40 cycles at 94 °C (2.5 °C/s ramp) for 30 s and at 57 °C followed by 98 °C (2.5 °C/s ramp) for 10 min. Samples were held at 4 °C until further processing. After PCR, the PCR plates were loaded on a Bio-Rad QX200 droplet reader, and ddPCR absolute quantifications of mutant and wild-type alleles were estimated by modelling them as a Poisson distribution, using Bio-Rad QuantaSoft version 1.6.6 software. Thresholds were defined based on signals from empty droplets, wild-type DNA controls and mutant-positive controls, as described in the Droplet Digital Application Guide (Bio-Rad).
Statistical analysis
All analyses were performed using JMP Pro statistical software (ver. 14.1.0, SAS Institute, Tokyo, Japan). Fisher’s exact test or chi-squared test was used to analyse categorical data. Proportions were compared by two-sample tests. The t test, Mann–Whitney U test or Kruskal–Wallis test was used to analyse continuous variables. P < 0.05 was considered significant.
Results
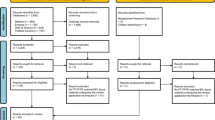
Among 85 FFPE tissue samples, 75 (44 FA and 31 PT) were eligible for analysis. Ten samples, including five FA samples with insufficient extracted DNA and five FA samples in which the MED12 amplicon could not be amplified to perform SS, were excluded. We successfully performed ddPCR to detect TERTp mutations in all PT samples and SS for TERTp regions in all but one PT sample.
Patient characteristics
Histological classifications of FA, histological grades of PT, ages and tumour sizes of each case are listed in Table 1. Among 44 FA, 18 samples were typed as intracanalicular, 6 as pericanalicular, 8 as mastopathic, 7 as organoid, 4 as complex and 1 as juvenile fibroadenoma. Among 31 PT samples, 17 were classified as benign, 9 as borderline and 5 as malignant. The FA ranged in size from 9 to 125 mm, and the PT ranged from 23 to 130 mm. The Mann–Whitney U test showed that patients with PT were significantly older (P = 0.006) and had larger tumours (P < 0.001) than those with FA.
MED12 mutations in FA and PT
A significantly higher percentage of PT (21/31, 68%) was MED12Mut+ than was FA (19/44, 42%, P = 0.035, two-sample test of proportions). Among FA, the intracanalicular type showed MED12 mutations more frequently than the other types (P = 0.046, chi-squared test). The frequency of MED12 mutations did not significantly differ among histological grades of PT (P = 0.81, chi-squared test; Supplementary Table S2).
Table 2 shows the types and locations of MED12 mutations. All of the missense mutations have already been reported in COSMIC,47 but many deletion and deletion/insertion patterns were newly discovered in this study (shown in red in Table 2).
TERTp mutations in FA and PT
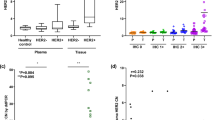
In this study, the only mutation site for TERTp was C228T; we saw no case of C250T. In 16 cases examined for TERTpMut+ by both SS and ddPCR, SS found that 12.5% (2/16) were TERTpMut+, whereas ddPCR found 37.5% (6/16; P = 0.037, two-sample test of proportions). Both cases determined as TERTpMut+ by SS were PT; no FA case carrying TERTp mutations was found (Fig. 1 and Supplementary Table S3).
SS was performed for 16 cases to detect TERTpMut+ and ddPCR was performed for all cases. SS found that 12.5% (2/16) were TERTpMut+, and both the cases were PT. ddPCR analyses of all tumours found that 42.7% (32/75) were TERTpMut+, a much higher rate than in SS detection. TERTpMut+ was significantly more frequent in PT (19/31, 61.3%) than in FA (13/44, 29.5%; P = 0.009, Fisher’s exact test).
ddPCR analyses of all tumours found that 42.7% (32/75) were TERTpMut+, which was significantly more frequent in PT (19/31; 61.3%) than in FA (13/44; 29.5%; P = 0.009, Fisher’s exact test, Fig. 1 and Table 3).
The two samples that were found to be TERTpMut+ by SS (16a and 16i) were strongly positive in ddPCR analysis, whereas those with discrepant results between SS and ddPCR (e.g. 16b and 16k) were weakly positive by ddPCR (Fig. 2).
Correlations between TERTp-mutation status and clinicopathological variables in FA and PT are shown in Table 3. Among FA, TERTpMut+ tumours were significantly larger than TERTpMut– ones (P = 0.015, Mann–Whitney U test), but this was not the case in PT (P = 1.0). None of the other variables, such as patients’ age, FA classification, PT grade and MED12-mutation status, differed significantly between TERTpMut+ and TERTpMut− tumours.
Fractional abundance of FAs and PTs with TERTp mutation
A total of 70 FA and PT samples with TERTp mutation were detected by ddPCR at a fractional abundance from 0.7% to 92% (Supplementary Table S4). PT had significantly higher fractional abundance of TERTp mutation than FA: the mean fractional abundance in FA was 20% (range: 1.0–89%), while that in PT was 40% (range: 0.7–92%) (P = 0.049, Mann–Whitney U test). Among the cases analysed by both SS and ddPCR, fractional abundance was relatively high in the two concordant cases (28.8% and 52.3%; Supplementary Table S5). Those cases that were both MED12-mutation-positive with SS and TERTp-mutation-positive with ddPCR did not show high fractional abundance compared with the others (P = 0.067, t test; Supplementary Table S4).
Concordance of MED12 and TERTp mutations between metachronous multiple tumours from the same individuals
In this study, three patients had metachronous multiple tumours. All three cases harboured the same MED12 mutation in the primary and secondary lesions (Table 4). However, only one case, of malignant PT, was TERTpMut+ in both primary and secondary tumours, whereas the other two cases carried this mutation only in the second tumour.
Discussion
In this study, we describe a method to detect TERT promoter mutations in FA and PT using ddPCR. TERT promoter mutation has been reported to be one of the most prevalent mutations in PT other than MED12 mutation. ddPCR not only succeeded in detecting TERT promoter mutations, but also revealed this mutation to be much more frequent among breast fibroepithelial tumours, especially in FA, than was previously known. The advantages of ddPCR are its high sensitivity, quantitative measurement and low running cost. As ddPCR can detect mutations that have few patterns of single-base substitution, it is entirely appropriate for TERTp mutations, because only two hotspots have been reported in this region.29,31,32,33,34 ddPCR can distinguish mutations with a very low frequency of allele variants that SS would not have detected. Here, we show the potential high sensitivity of ddPCR in detecting TERTpMut+ FA and PT, as in a previous report on melanoma by McEvoy et al.39
Around the mutation sites of C228T and C250T, TERTp region contains guanine and cytosine in high frequency. Such GC-rich regions affect the generation of secondary products during PCR, and as a consequence, inhibit DNA polymerases from copying these sites correctly. Besides, the two mutation hotspots, C228T and C250T, locate close to each other. These inconvenient factors reduce the sensitivity in sequencing analyses, including NGS. Such disadvantageous potentials remain challenging in ddPCR research; however, Corless BC et al.49 showed excellent sensitivity and specificity in detecting TERTp mutations by ddPCR. According to them, in ddPCR analysis of mutations in GC-rich regions, shorter amplicon length provides better detection sensitivity. They mentioned that DNA in FFPE samples is shortly fragmented due to the chemical reaction between formalin and nucleic acid, which helps primers bind to the target regions and generate short amplicons, leading to the efficient PCR reaction.49 Although FFPE samples are often considered to be unfavourable in sequencing analysis due to the low quality of nucleic acid conditions, they might be even advantageous in ddPCR analysis for TERTp-mutation detection.
There have been a few reports comparing the detection of genetic mutations with SS and digital PCR, including one from McEvoy et al.39 and another from Sho et al.50 McEvoy et al. examined mutations in BRAF, NRAS and TERTp in melanoma. According to them, the frequency of mutation detection in SS was 20% (8/40) in BRAF, 0% (0/40) in NRAS and 12.5% (5/40) in TERTp, whereas that with ddPCR was 55% (22/40) in BRAF, 10% (41/40) in NRAS and 37.5% (15/40) in TERTp.39 Sho et al. performed SS on resected specimens of the pancreas, and digital PCR on preoperative endoscopic ultrasound–fine-needle aspiration specimens. All 22 cases positive for mutation by SS were also determined to be mutation-positive by digital PCR, and digital PCR detected five more mutations. This report demonstrates the utility of digital PCR in cytology samples, which have a much lower tumour burden than surgical specimens.50 In these reports, digital PCR determined as mutation-positive for all cases in which SS detected the mutation, and this result is consistent with ours.
In this study, we used SS instead of NGS to detect TERTp mutations in some cases. NGS’s deep sequencing is well known to be more sensitive than Sanger sequencing, and it would be interesting to know what the results of NGS analysis would be in these samples. However, conducting NGS targeting only the TERT promoter region is too costly. In addition, because one of our future goals is to apply our method to clinical practice, we judged it impractical to perform additional NGS in this study from the beginning of the research.
With our TERTpMut analysis by ddPCR, the frequency of this mutation was shown with histological grade, from FA (29%) and benign PT (59%) to borderline PT (67%, Table 3). Nault et al. analysed TERTp mutations in cirrhotic dysplastic nodules, which are premalignant lesions of hepatocellular carcinoma (HCC), and in early and progressed HCC. They reported a strong relationship between TERTp mutations and hepatocarcinogenesis: the mutations were identified in 6% of low-grade dysplastic nodules, 19% of high-grade nodules, 61% of early HCC and 42% of small and progressed HCC.51 Our results in breast FA and PT were quite similar to their results in HCC, in that the frequency of TERTp mutations increased as tumours went from benign to malignant, which implies a sequential development by some FA into PT. Interestingly, our study showed a positive correlation between FA size and frequency of TERTp mutations (P = 0.015, Table 3). Some FA, especially large ones, may be genetically similar to benign PT, although morphologically distinguishable from them.
The relationship between MED12 and TERTp mutations varies in previous reports.29,30,31,33,34 Piscuoglio et al.,29 Pareja et al.30 and Liu et al.34 reported that just 50% and slightly over of PT were MED12Mut+ (13/25, 5/9 and 3/6), whereas Yoshida et al.31 and Garcia-Dios et al.33 reported that almost all of them were simultaneously MED12Mut+ (29/30 and 12/13, respectively). Only one study found TERTp mutations in FA:31 4 cases out of 58 (7%) and all of the TERTpMut+ FA also harboured MED12 mutations. Our study found no significant correlation between MED12 and TERTp mutations, in either FA or PT. Although MED12 mutations are most frequently observed in intracanalicular-type FA,10,11,21,23,24,28 TERTp mutations detected in our study had no statistical relationship with FA classification or PT grade (Table 3). These two mutations might independently affect the genesis or development of FA and PT.
Among the 72 patients in our study, three experienced recurrences (Table 4). When we compared the first resected tumour with later ones, two of them, an FA and a benign PT, showed TERTpMut status change from negative to positive. Considering that all cases harboured the same pattern of MED12 mutations within each case, the secondary resected tumours seemed to be genetically identical to the primary tumours, suggesting that they were truly recurrent tumours. Garcia-Dios et al. also reported evidence of recurrent PT that acquired TERTp mutations,33 but our result indicates that TERTp mutations can be acquired during tumour growth or recurrence in both FA and PT.
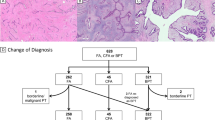
We detected TERTp mutations at a much higher rate in FA than were seen in previous studies,29,31,33 but some might doubt the accuracy of the FA diagnoses. In the current study, two expert pathologists diagnosed all of the tumours. The distinction between FA and PT is often challenging, and in fact, we had eight cases, all eventually diagnosed as benign PT, on which the pathologists did not initially agree whether they were intracanalicular-type FA or benign PT (data not shown). TERTp mutations were found in five of these controversial tumours (63%), which is similar to the mutation rate of PT as a whole (61%), but much higher than that of FA (30%, Table 3). We consider that these controversial cases were credibly distinguished from other FA, and would be similarly diagnosed by most pathologists.
Although we found ddPCR to be more sensitive than SS for TERTp mutations, technical challenges remain. Sufficient quality and quantity of DNA is essential for detecting low-frequency mutations; therefore, small tumours or hyalinised FA may be inadequate for mutation analysis with ddPCR. Another challenge to successful analysis is artefacts from formalin-fixed samples when using sensitive molecular assays causing false-positive calls.52,53 We set thresholds based on signals from empty droplets, wild-type DNA and mutant-positive controls, and determined samples as negative when they showed fewer than ten mutation calls, to decrease the possibility of error due to false positives. In other words, detecting very low-frequency mutations with fewer than ten calls is highly challenging. Additional studies are required to resolve these problems and improve accuracy.
To examine the influence of FFPE-derived artefacts of DNA in ddPCR analysis, it would be more robust to prepare an FFPE-derived negative and a positive control for both C228T and C250T mutation-positive controls. We set distilled water as a negative control, and used DNA collected from a cell line as a positive control, so the DNA of these controls did not derive from FFPE. However, in our positive control, only C228T positivity was confirmed; thus, it did not show what type of dot plot would be obtained by ddPCR when C250T had a mutation in our study, although this was depicted in previous reports. These matters can be considered limitations of this work.
Another limitation is the primers that we used in this study, which were the same as those reported by Yoshida et al.31 and Liu et al.34 The nucleotide sequences of these primers differ from those in the study by McEvoy et al.,39 albeit by only a few bases. As there is no previous report describing the performance of ddPCR analysis using our primers, it may have been necessary to verify the findings of the study by using the primers that McEvoy et al. used for validation.
Extracted tumour DNA in the present study contained mixtures of that from epithelium, stroma or somewhat normal mammary cells and lymphocytes surrounding the tumour. Earlier studies found mutations for TERTp and MED12 only in PT stromal components,29,31,32 so strictly speaking, DNA extraction from only the stromal components through microdissection would be necessary to identify TERTp mutations in FA and PT stroma, and to measure the allele frequency by ddPCR. Extracting DNA and measuring TERTp mutations from only the tumour’s stromal components may solve the problem of false positivity and -negativity more precisely, thus enhancing studies of relationships of TERTpMut allelic frequency with tumour growth rate and malignancy in FA and PT. Further study with targeted tumour cells is warranted.
In conclusion, we have presented the first assessment by ddPCR of TERTp mutations in FFPE breast FA and PT, and detected these mutations at a higher rate than previously reported. Our new findings reconfirm the genomic similarity of FA to PT and may help elucidate the biology of these tumours. We have shown ddPCR to be a robust method of detecting TERTp mutations, which suggest a wider clinical potential for this technology. A large-scale study is needed to determine whether TERTp mutation detection by ddPCR can be predictive, and has prognostic value for the surgical treatment of FA and PT.
References
Tavassoli, F. A. & Eusebi, V. Biphasic tumors. in AFIP Atlas of Tumor Pathology, Vol 10 (eds Tavassoli, F. A & Eusebi, V.) 315–340 (American Registry of Pathology, Washington, DC, 2009).
Tan, P. H., Tse, G., Lee, A., Simpson, J. F. & Hanby, A. M. Fibroepithelial tumours. in WHO Classification of Tumours of the Breast, 4th edn. (eds Lakhani, S. R., Ellis, I. O., Schnitt, S. J., Tan, P. H. & van de Vijver, M. J.) 142–147 (IARC Press, Lyon, France, 2012).
Yang, X., Kandil, D., Cosar, E. F. & Khan, A. Fibroepithelial tumors of the breast: pathologic and immunohistochemical features and molecular mechanisms. Arch. Pathol. Lab. Med. 138, 25–36 (2014).
Tan, P. H. & Ellis, I. O. Myoepithelial and epithelial-myoepithelial, mesenchymal and fibroepithelial breast lesions: updates from the WHO Classification of Tumours of the Breast. J. Clin. Pathol. 66, 465–470 (2012).
Brogi, E. Fibroepithelial neoplasms. in Rosen’s Breast Pathology, 4th edn. (eds Hoda, S. A., Brogi, E., Koerner, F. C. & Rosen, P. R.) 213–270 (Lippincott Williams and Willkins, Philadelphia, PA, 2009).
Sapino, A., Bosco, M., Cassoni, P., Castellano, I., Arisio, R., Cserni, G. et al. Estrogen receptor-beta is expressed in stromal cells of fibroadenoma and phyllodes tumors of the breast. Mod. Pathol. 19, 599–606 (2006).
Tan, W. J., Chan, J. Y., Thike, A. A., Lim, J. C., Md Nasir, N. D., Tan, J. S. et al. MED12 protein expression in breast fibroepithelial lesions: correlation with mutation status and oestrogen receptor expression. J. Clin. Pathol. 69, 858–865 (2016).
Carty, N. J., Carter, C., Rubin, C., Ravichandran, D., Royle, G. T. & Taylor, I. Management of fibroadenoma of the breast. Ann. R. Coll. Surg. Engl. 77, 127–130 (1995).
Dixon, J. M. Cystic disease and fibroadenoma of the breast: natural history and relation to breast cancer risk. Br. Med. Bull. 47, 258–271 (1991).
Mishima, C., Kagara, N., Tanei, T., Naoi, Y., Shimoda, A., Shimazu, K. et al. Mutational analysis of MED12 in fibroadenomas and phyllodes tumors of the breast by means of targeted next-generation sequencing. Breast Cancer Res. Treat. 152, 305–312 (2015).
Yoshida, M., Sekine, S., Ogawa, R., Yoshida, H., Maeshima, A., Kanai, Y. et al. Frequent MED12 mutations in phyllodes tumours of the breast. Br. J. Cancer 112, 1703–1708 (2015).
Cani, A. K., Hovelson, D. H., McDaniel, A. S., Sadis, S., Haller, M. J., Yadati, V. et al. Next-Gen sequencing exposes frequent MED12 mutations and actionable therapeutic targets in phyllodes tumors. Mol. Cancer Res. 13, 613–619 (2015).
Nagasawa, S., Maeda, I., Fukuda, T., Wu, W., Hayami, R., Kojima, Y. et al. MED12 exon 2 mutations in phyllodes tumors of the breast. Cancer Med. 4, 1117–1121 (2015).
Ben Hassouna, J., Damak, T., Gamoudi, A., Chargui, R., Khomsi, F., Mahjoub, S. et al. Phyllodes tumors of the breast: a case series of 106 patients. Am. J. Surg. 192, 141–147 (2016).
Guerrero, M. A., Ballard, B. R. & Grau, A. M. Malignant phyllodes tumor of the breast: review of the literature and case report of stromal overgrowth. Surg. Oncol. 12, 27–37 (2003).
Noguchi, S., Yokouchi, H., Aihara, T., Motomura, K., Inaji, H., Imaoka, S. et al. Progression of fibroadenoma to phyllodes tumor demonstrated by clonal analysis. Cancer 76, 1779–1785 (1995).
Noguchi, S., Aihara, T., Motomura, K., Inaji, H., Imaoka, S., Koyama, H. et al. Phyllodes tumor of the breast: pathology, genesis, diagnosis, and treatment. Breast Cancer 3, 79–92 (1996).
Millikan, R., Hulka, B., Thor, A., Zhang, Y., Edgerton, S., Zhang, X. et al. p53 mutations in benign breast tissue. J. Clin. Oncol. 13, 2293–2300 (1995).
Franco, N., Picard, S. F., Mege, F., Arnould, L. & Lizard-Nacol, S. Absence of genetic abnormalities in fibroadenomas of the breast determined at p53 gene mutations and microsatellite alterations. Cancer Res. 61, 7955–7958 (2001).
Vorkas, P. A., Poumpouridou, N., Agelaki, S., Kroupis, C., Georgoulias, V. & Lianidou, E. S. PIK3CA hotspot mutation scanning by a novel and highly sensitive high-resolution small amplicon melting analysis method. J. Mol. Diagn. 12, 697–704 (2010).
Loke, B. N., Md Nasir, N. D., Thike, A. A., Lee, J. Y. H., Lee, C. S., The, B. T. et al. Genetics and genomics of breast fibroadenomas. J. Clin. Pathol. 71, 381–387 (2018).
Lim, W. K., Ong, C. K., Tan, J., Thike, A. A., Ng, C. C., Rajasegaran, V. et al. Exome sequencing identifies highly recurrent MED12 somatic mutations in breast fibroadenoma. Nat. Genet. 46, 877–880 (2014).
Pfarr, N., Kriegsmann, M., Sinn, P., Klauchen, F., Endris, V., Herpel, E. et al. Distribution of MED12 mutations in fibroadenomas and phyllodes tumors of the breast–implications for tumor biology and pathological diagnosis. Genes Chromosomes Cancer 54, 444–452 (2015).
Piscuoglio, S., Murray, M., Fusco, N., Marchiò, C., Loo, F. L., Martelotto, L. G. et al. MED12 somatic mutations in fibroadenomas and phyllodes tumours of the breast. Histopathology 67, 719–729 (2015).
Lien, H. C., Huang, C. S., Yang, Y. W. & Jeng, Y. M. Mutational analysis of MED12 exon 2 in a spectrum of fibroepithelial tumours of the breast: implications for pathogenesis and histogenesis. Histopathology 68, 433–441 (2016).
Ng, C. C., Tan, J., Ong, C. K., Lim, W. K., Rajasegaran, V., Nasir, N. D. et al. MED12 is frequently mutated in breast phyllodes tumours: a study of 112 cases. J. Clin. Pathol. 68, 685–691 (2015).
Tan, J., Ong, C. K., Lim, W. K., Ng, C. C., Thike, A. A., Ng, L. M. et al. Genomic landscapes of breast fibroepithelial tumors. Nat. Genet. 47, 1341–1345 (2015).
Kim, J. Y., Yu, J. H., Nam, S. J., Kim, S. W., Lee, S. K., Park, W. Y. et al. Genetic and clinical characteristics of phyllodes tumors of the breast. Transl. Oncol. 11, 18–23 (2018).
Piscuoglio, S., Ng, C. K., Murray, M., Burke, K. A., Edelweiss, M., Geyer, F. C. et al. Massively parallel sequencing of phyllodes tumours of the breast reveals actionable mutations, and TERT promoter hotspot mutations and TERT gene amplification as likely drivers of progression. J. Pathol. 238, 508–518 (2016).
Pareja, F., Geyer, F. C., Kumar, R., Selenica, P., Piscuoglio, S., Ng, C. K. Y. et al. Phyllodes tumors with and without fibroadenoma-like areas display distinct genomic features and may evolve through distinct pathways. NPJ Breast Cancer 3, 40 (2017).
Yoshida, M., Ogawa, R., Yoshida, H., Maeshima, A., Kanai, Y., Kinoshita, T. et al. TERT promoter mutations are frequent and show association with MED12 mutations in phyllodes tumors of the breast. Br. J. Cancer 113, 1244–1248 (2015).
Tsang, J. Y. S., Hui, Y. K., Lee, M. A., Lacambra, M., Ni, Y. B., Cheung, S. Y. et al. Association of clinicopathological features and prognosis of TERT alterations in phyllodes tumor of breast. Sci. Rep. 8, 3881 (2018).
Garcia-Dios, D. A., Levi, D., Shah, V., Gillett, C., Simpson, M. A., Hanby, A. et al. MED12, TERT promoter and RBM15 mutations in primary and recurrent phyllodes tumours. Br. J. Cancer 118, 277–284 (2018).
Liu, S. Y., Joseph, N. M., Ravindranathan, A., Stohr, B. A., Greenland, N. Y., Vohra, P. et al. Genomic profiling of malignant phyllodes tumors reveals aberrations in FGFR1 and PI-3 kinase/RAS signaling pathways and provides insights into intratumoral heterogeneity. Mod. Pathol. 29, 1012–1027 (2016).
Clark, A. D., Oldenbroek, M. & Boyer, T. G. Mediator kinase module and human tumorigenesis. Crit. Rev. Biochem. Mol. Biol. 50, 393–426 (2015).
Yuan, P., Cao, J. L., Abuduwufuer, A., Wang, L. M., Yuan, X. S., Lv, W. et al. Clinical characteristics and prognostic significance of TERT promoter mutations in cancer: a cohort study and a meta-analysis. PLoS ONE 11, e0146803 (2016).
Vinagre, J., Almeida, A., Pópulo, H., Batista, R., Lyra, J., Pinto, V. et al. Frequency of TERT promoter mutations in human cancers. Nat. Commun. 4, 2185 (2013).
McEvoy, A. C., Calapre, L., Pereira, M. R., Giardina, T., Robinson, C., Khattak, M. A. et al. Sensitive droplet digital PCR method for detection of TERT promoter mutations in cell free DNA from patients with metastatic melanoma. Oncotarget 8, 78890–78900 (2017).
McEvoy, A. C., Wood, B. A., Ardakani, N. M., Pereira, M. R., Pearce, R., Cowell, L. et al. droplet digital PCR for mutation detection in formalin-fixed, paraffin-embedded melanoma tissues: a comparison with Sanger sequencing and pyrosequencing. J. Mol. Diagn. 20, 240–252 (2018).
Vogelstein, B. & Kinzler, K. W. Digital PCR. Proc. Natl Acad. Sci. USA 96, 9236–9241 (1999).
Hindson, B. J., Ness, K. D., Masquelier, D. A., Belgrader, P., Heredia, N. J., Makarewicz, A. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610 (2011).
Taylor, S. C., Carbonneau, J., Shelton, D. N. & Boivin, G. Optimization of droplet digital PCR from RNA and DNA extracts with direct comparison to RT-qPCR: clinical implications for quantification of oseltamivir-resistant subpopulations. J. Virol. Methods 224, 58–66 (2015).
Kuroda, H., Takeuchi, I., Ohnishi, K., Sakamoto, G., Akiyama, F., Toyozumi, Y. et al. Clinical and pathologic features of fibroadenoma of the mastopathic type. Surg. Today 36, 590–595 (2006).
Mori, I., Han, B., Wang, X., Taniguchi, E., Nakamura, M., Nakamura, Y. et al. Mastopathic fibroadenoma of the breast: a pitfall of aspiration cytology. Cytopathology 17, 233–238 (2006).
Sklair-Levy, M., Sella, T., Alweiss, T., Craciun, I., Libson, E. & Mally, B. Incidence and management of complex fibroadenomas. Am. J. Roentgenol. 190, 214–218 (2008).
Wechselberger, G., Schoeller, T. & Piza-Katzer, H. Juvenile fibroadenoma of the breast. Surgery 132, 106–107 (2002).
Je, E. M., Kim, M. R., Min, K. O., Yoo, N. J. & Lee, S. H. Mutational analysis of MED12 exon 2 in uterine leiomyoma and other common tumors. Int. J. Cancer 131, E1044–E1047 (2012).
Forbes, S. A., Beare, D., Gunasekaran, P., Leung, K., Bindal, N., Boutselakis, H. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–D811 (2015).
Corless, B. C., Chang, G. A., Cooper, S., Syeda, M. M., Shao, Y., Iman Osman, I. et al. Development of novel mutation-specific droplet digital PCR assays detecting TERT promoter mutations in tumor and plasma samples. J. Mol. Diagn. 21, 274–285 (2019).
Sho, S., Court, C. M., Kim, S., Braxton, D. R., Hou, S., Muthusamy, V. R. et al. Digital PCR improves mutation analysis in pancreas fine needle aspiration biopsy specimens. PLoS ONE 26, e0170897 (2017).
Nault, J. C., Calderaro, J., Di Tommaso, L., Balabaud, C., Zafrani, E. S., Bioulac-Sage, P. et al. Telomerase reverse transcriptase promoter mutation is an early somatic genetic alteration in the transformation of premalignant nodules in hepatocellular carcinoma on cirrhosis. Hepatology 60, 1983–1992 (2014).
Ye, X., Zhu, Z. Z., Zhong, L., Lu, Y., Sun, Y., Yin, X. et al. High T790M detection rate in TKI-naive NSCLC with EGFR sensitive mutation: truth or artifact? J. Thorac. Oncol. 8, 1118–1120 (2013).
Watanabe, M., Kawaguchi, T., Isa, S., Ando, M., Tamiya, A., Kubo, A. et al. Ultra-sensitive detection of the pretreatment EGFR T790M mutation in non-small cell lung cancer patients with an EGFR-activating mutation using droplet digital PCR. Clin. Cancer Res. 21, 3552–3560 (2015).
Acknowledgements
We thank Marla Brunker, from Edanz Group (www.edanzediting.com/ac), for editing a draft of this paper.
Author information
Authors and Affiliations
Contributions
K.O. and T.S. conceived the study. T.S., M.T. and Y.S. directed the study and supervised the research. K.O. collected tumour specimens. K.O. and T.S. confirmed the histopathological findings and interpreted the clinical data. K.O. performed Sanger sequencing and droplet-digital PCR analyses. K.O. wrote the paper, with assistance and final approval of all authors.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study protocol was approved by the ethics committee at the University of Tokyo Hospital, Tokyo, Japan. Written informed consent was obtained from all patients. The study was performed in accordance with the Declaration of Helsinki.
Consent to publish
Not applicable.
Data availability
All data supporting the study are available on request. No proprietary materials, except patient tissues, were used.
Competing interests
The authors declare no competing interests.
Funding information
This study was supported by “Project for the Advanced Genome-Based Medicine” from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Additional information
Note This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Otsuji, K., Sasaki, T., Tanabe, M. et al. Droplet-digital PCR reveals frequent mutations in TERT promoter region in breast fibroadenomas and phyllodes tumours, irrespective of the presence of MED12 mutations. Br J Cancer 124, 466–473 (2021). https://doi.org/10.1038/s41416-020-01109-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-020-01109-8