Abstract
The mammalian/mechanistic target of rapamycin (mTOR) is a serine/threonine protein kinase that integrates inputs from nutrients and growth factors to control many fundamental cellular processes through two distinct protein complexes mTORC1 and mTORC2. Recent mouse genetic studies have established that mTOR pathways play important roles in regulating multiple aspects of skeletal development and homeostasis. In addition, mTORC1 has emerged as a common effector mediating the bone anabolic effect of Igf1, Wnt and Bmp. Dysregulation of mTORC1 could contribute to various skeletal diseases including osteoarthritis and osteoporosis. Here we review the current understanding of mTOR signaling in skeletal development and bone homeostasis, as well as in the maintenance of articular cartilage. We speculate that targeting mTOR signaling may be a valuable approach for treating skeletal diseases.
Similar content being viewed by others
Introduction
The mechanistic (formerly “mammalian”) target of rapamycin, as indicated by its name, is highly sensitive to rapamycin, a drug clinically used for antifungal, immunosuppressive, and antitumor purposes. Rapamycin was initially isolated from bacteria in soil samples of Easter Island that can inhibit yeast proliferation1. Mechanistically, rapamycin was shown to exert its function by forming a complex with FKBP122. Subsequent studies identified the targets of FKBP12-rapamycin complex in yeasts and mammals, which were named as target of rapamycin (TOR) and mammalian target of rapamycin (mTOR), respectively3,4,5,6,7. Since its discovery, extensive research over the last twenty years has indicated that mTOR pathways play important roles in regulating development and homeostasis of mammalian tissues, and that their dysregulation is implicated in pathogenesis of many human diseases.
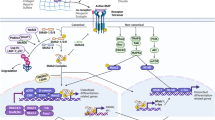
Biochemically, mTOR is an evolutionarily conserved serine/threonine protein kinase belonging to the phosphoinositide 3-kinase (PI3K)-related kinase family, and functions as a catalytic subunit in two distinct protein complexes: mTOR complex 1 (mTORC1) and complex 2 (mTORC2; Fig. 1). Initially, mTORC1 and mTORC2 were distinguished by virtue of their different sensitivities to rapamycin. Whereas mTORC1 is inhibited by acute rapamycin treatment, mTORC2 is resistant to such treatment. However, recent studies showed that prolonged rapamycin treatment also impaired mTORC2 signaling both in vitro and in vivo8,9. mTORC1 and mTORC2 differ in their components. While mTORC1 and mTORC2 do share two core components (mTOR, mLST8/GßL)10,11, they contain Raptor or Rictor as their respective unique core subunit. In addition, mTORC1 has two inhibitory subunits (PRAS40, DEPTOR)12,13,14,15, whereas mTORC2 contains an inhibitory subunit DEPTOR15 and two regulatory subunits (Protor1/2 and mSin1)16,17. Genetic studies revealed that ablation of mTOR blocked both mTORC1 and mTORC2 signaling whereas ablation of Raptor or Rictor only impaired mTORC1 or mTORC2 signaling, respectively10,11.
mTORC1 integrates a wide variety of intracellular and extracellular signals, including growth factors such as WNT and Insulin/IGF-1, the levels of oxygen, energy, stress, or amino acids, to regulate cell growth and metabolism through a number of downstream effectors18 (Fig. 1). One key upstream regulator of mTORC1 signaling is the Tsc1/Tsc2 complex, a GTPase-activating protein (GAP) for the small GTPase Rheb18. Rheb directly binds to mTORC1 and potently stimulates its activity, but Tsc1/Tsc2 negatively regulates mTORC1 by converting Rheb into its inactive GDP-bound form19,20. Whereas many upstream signals activate or inhibit mTORC1 activity by acting on Tsc1/Tsc218, regulation of mTORC1 activity by amino acid levels is independent of TSC1/2, and instead through Rag GTPases (RagA, RagB, RagC, and RagD) and their regulators21. Moreover, the presence of amino acids, in particular leucine and arginine, is required for other upstream signals to activate mTORC118. The lysosome has emerged as a key organelle mediating mTORC1 activation by both amino acids and growth factors. In a current model, functionally active heterodimers containing GTP-loaded RagA/B and GDP-loaded RagC/D accumulate on the cytoplasmic surface of the lysosome in response to amino acids that promote the formation of a supercomplex including the pentameric Regulator complex and the multi-subunit vacuolar ATPase complex. The active Rag heterodimer recruits mTORC1 to the lysosomal membrane where Rheb is also anchored, thus initiating mTORC1 activation. In support of the model, recent work has provided evidence that the solute carrier SLC38A9 likely functions as a sensor (“transceptor”) to arginine or glutamine concentration in the lysosome to initiate mTORC1 signaling through the Rag–Regulator complex22,23. Similarly, leucine stimulation of mTORC1 is dependent on the Rag GTPases but its potential transceptor in the lysosome is yet to be discovered24. Interestingly however, mTORC1 stimulation by glutamine appears to be independent of the Rag–Regulator complex, but requiring the small GTPase Arf124. Furthermore, mTORC1 may also be activated by amino acids on the Golgi membrane where another small GTPase Rab1A recruits mTORC1 to be activated by Rheb localized in the organell25. Thus, the mechanisms underlying amino acid regulation of mTORC1 are undoubtedly complex and likely function in an amino acid-specific manner.
One of the major functions of mTORC1 signaling is promoting anabolic processes, including protein and lipid synthesis. The stimulation of protein synthesis is mainly through phosphorylation of p70 S6 kinase (S6K1) and eukaryotic translation initiation factor 4E-binding protein 1 (4EBP1), whereas mTORC1 activates lipid synthesis through SREBP1/218. Besides its anabolic role, mTORC1 signaling inhibits catabolic processes, particularly autophagy by phosphorylating autophagy-initiating kinase Ulk1 and blocking its activation by AMPK26. In addition, mTORC1 has been shown to inhibit autophagy in part by inhibiting the nuclear translocation and activity of TFEB, a transcription factor important for the expression of autophagy and lysosomal genes27.
Similar to mTORC1 signaling, mTORC2 can be activated by various growth factors, including Wnt and Insulin/IGF128,29 (Fig. 1). In addition, mTORC2 is activated by mechanical stress and ribosomes in vitro, although the molecular mechanism is still unclear30,31. mTORC2 controls proliferation and survival through a distinct group of downstream targets, including members of the AGC family of kinases Akt, serum and glucocorticoid-induced protein kinase 1 (SGK1), and protein kinase C-α (PKC-α)10,11,32,33.
mTOR signaling in endochondral skeletal development
Mammalian bones are formed through two different mechanisms, endochondral versus intramembranous bone formation34. In contrast to intramembranous ossification where mesenchymal progenitors directly differentiate into osteoblasts, endochondral bone development begins with the condensation of mesenchymal progenitors due to increased cell–cell contact. Subsequently, the centrally-located cells within the mesenchymal condensations differentiate into chondrocytes, while cells at the periphery develop into the perichondrium. Following chondrogenesis, chondrocytes within the cartilage primordia initially proliferate rapidly, and then undergo a maturation process involving successive prehyertrophic, hypertrophic and terminal hypertrophic stages. Subsequently, blood vessels invade the hypertrophic cartilage and bring in progenitors for osteoclasts or osteblasts that are respectively responsible for resorbing the hypertrophic cartilage or depositing bone matrix.
Recent studies have implicated mTORC1 in regulating multiple aspects of cartilage development. Disruption of mTORC1 via deletion of Raptor in the early limb mesenchyme significantly reduced the size of limb bud cells and impaired chondrogenesis from the mesenchymal progenitors35. Similarly, rapamycin dramatically suppressed the formation of cartilage nodules from limb bud cells without affecting precartilaginous mesenchymal condensation35,36,37. In addition, rapamycin markedly reduced proteoglycan accumulation and the expression of chondrocyte markers in the chondrogenic ATDC5 cell line, perhaps through suppression of Sox9 expression35,36.
Studies of the growth plate cartilage in vivo have also revealed important roles for mTORC1 in chondrocytes. Immunofluorescence staining for phospho-S6, a common readout for mTORC1 signaling, revealed intense and nearly homogenous activity in prehypertrophic and early hypertrophic chondrocytes, but only sporadic signals in round chondrocytes36,38. In addition, mTORC1 was largely absent in much of the hypertrophic region except for the terminal hypertrophic chondrocytes36,38. Functionally, deletion of Raptor severely impaired skeletal growth through the reduction of chondrocyte size and matrix production, as well as the delay in chondrocyte hypertrophy and the eventual removal of the hypertrophic cartilage38. The decrease in chondrocyte size and matrix production is likely associated with the compromised protein synthesis rate in the Raptor-deficient chondrocytes. Surprisingly, ablation of Raptor did not have a major effect on chondrocyte proliferation or survival38. On the other hand, hyperactivation of mTORC1 signaling via Tsc1 deletion increased chondrocyte proliferation while impeding chondrocyte maturation39. This study further suggested that mTORC1 coordinated chondrocyte growth, proliferation, and differentiation through its downstream effector S6K1, which acts on Gli2 to stimulate transcription of parathyroid hormone-related peptide (PTHrP)39. However, the model is difficult to reconcile with the observation that mTORC1 activity is highest in the prehypertrophic and early hypertrophic chondrocytes but PTHrP is mainly expressed by the peri-articular chondrocytes. Moreover, global deletion of S6K1 in mice caused a milder skeletal phenotype than Raptor deletion did40,41. Thus, the mechanism underlying the importance of mTORC1 signaling in cartilage development remains to be fully elucidated.
In contrast to mTORC1 signaling, mTORC2 appears to play a minor role in endochondral skeletal development. Inactivation of mTORC2 signaling via ablation of Rictor only mildly affected limb growth42. This study further showed that deletion of Rictor did not affect chondrocyte proliferation, apoptosis, cell size, or matrix production, but instead caused a mild delay in chondrocyte hypertrophy in both embryos and postnatal mice42.
mTORC1 signaling in bone formation and resorption
Bone homeostasis is maintained through the balance of bone formation and bone resorption. Osteoblasts differentiated from mesenchymal stem/progenitor cells are the chief bone-forming cells, while HSC-derived osteoclasts are responsible for bone resorption. Inhibition of mTORC1 signaling by rapamycin was shown to impair both proliferation and osteogenic differentiation of mouse bone marrow stromal cells (BMSC) in vitro, and to cause trabecular bone loss in vivo43,44. Conversely, activation of mTORC1 by IGF-1, an abundant growth factor present in the bone matrix and released during bone resorption, activated mTORC1 signaling to stimulate osteoblast differentiation of BMSC44 (Fig. 2). Similarly, bone anabolic Wnt ligands such as Wnt3a and Wnt7b have been shown to activate mTORC1 through PI3K-AKT signaling45. Pharmacological inhibition of mTORC1 signaling prevented Wnt7b-induced osteoblast differentiation in ST2 cells45. More importantly, genetic deletion of Raptor in the osteoblastic lineage cells alleviated the Wnt7b-induced high bone mass phenotype in mice45, indicating that Wnt7b promotes bone formation in part through mTORC1 activation. Mechanistically, mTORC1 mediates the osteogenic effect of Wnt partly by promoting glutamine catabolism and integrated stress response (ISR), which in turn induces the expression of protein anabolism genes essential for osteoblast differentiation46,47. In addition, Bmp2 was recently reported to induce the osteogenic program partly through a mTORC1-dependent mechanism47. Furthermore, Bmp signaling through Bmpr1a stimulated osteoblast activity through mTORC1 signaling in mice48. Thus, mTORC1 appears to be a common effector downstream of multiple bone anabolic signals.
Recent studies have further demonstrated that mTORC1 is required for the transition of preosteoblasts to mature osteoblasts49,50 (Fig. 2). Genetic inactivation of mTORC1 in preosteoblasts by specifically deleting Raptor in preosteoblasts with Osx-Cre caused osteopenia in mice, mainly due to a defect in bone formation. Further analyses indicated that the raptor-deficient preosteoblasts were deficient in matrix synthesis and mineralization, exhibiting a transcriptional profile of immature osteoblasts, indicative of a failure to progress beyond the early stages of osteogenesis. Interestingly, these studies showed that deletion of Raptor impaired protein synthesis without overtly affecting autophagy. Together, these findings support that mTORC1 promotes the transition from preosteoblasts to mature osteoblasts through enhancing mRNA translation. However, others reported that inhibition of mTORC1 signaling with a low dose of rapamycin enhanced preosteoblast differentiation, but prevented their proliferation in cell cultures and in mice51. The conflicting results from these studies could be due to the different experimental approaches. Whereas genetic ablation of Raptor with Osx-Cre inactivates mTORC1 signaling mainly in the osteoblast lineage from the preosteoblast stage onward, systemic administration of rapamycin exerts broad inhibition both within the osteoblast lineage and beyond. In addition, preosteoblasts may respond differently to the different extent of mTORC1 inhibition caused by raptor deletion versus low-dose rapamycin.
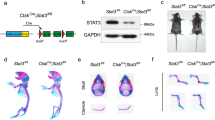
The importance of proper mTORC1 signaling in normal bone formation is further supported by the studies of the Tuberous Sclerosis (TSC) syndrome. TSC is an autosomal dominant disease with an estimated incidence of 1 in 5800 at birth and is caused by loss-of-function mutations of the TSC1 or TSC2 gene52,53,54. As heterodimeric TSC1 and TSC2 complex normally inhibits mTORC1 signaling by converting the active GTP-bound Rheb (a positive regulator of mTORC1) into the inactive GDP-bound form, the inactivating mutations of TSC1 or TSC2 cause hyperactive mTORC1 signaling in the TSC patients20. Although the main characteristics of TSC are benign tumors in skin, brain, kidney, and heart, 40–60% of the patients develop sclerotic bone lesions55,56. Recently, mice with TSC1 specifically deleted in neural crest cells were shown to exhibit sclerotic craniofacial bone lesions similar to those in TSC patients56. The study further revealed that deletion of TSC1 caused an expansion of osteoprogenitor cells at an early postnatal stage, leading to an increase in osteoblast number and consequently excessive bone formation. Remarkably, the sclerotic bone phenotype was completely reversed when rapamycin, a chemical inhibitor of mTORC1, was administered at an early postnatal stage, demonstrating that hyperactive mTORC1 signaling underlies the bone overgrowth caused by TSC1 deletion. In other studies, deletion of Tsc2 in mature osteoblasts or deletion of Tsc1 in preosteoblasts accelerated proliferation, but impaired osteoblast differentiation, probably through activating the STAT3/p63/Jagged/Notch pathway and suppressing Runx251,57. Thus, Tsc1/Tsc2 appears to function as an important modulator for proper mTORC1 signaling to ensure a balance of osteoblast proliferation and differentiation necessary for optimal bone formation.
The exact role of mTORC1 in regulating the osteoclast lineage is controversial at present. In one study, inactivation of mTORC1 by deletion of Raptor, or hyperactivation of mTORC1 by deleting Tsc1 in osteoclast precursors with LyzM-Cre either enhanced or impaired osteoclastogenesis, respectively58. The study further suggested that mTORC1 inhibits osteoclast differentiation through suppression of NF-kappaB and NFATc1, both critical transcription factors of osteoclastogenesis58. However, a recent study showed that inhibition of mTORC1 in bone marrow macrophages by either genetic deletion or rapamycin treatment suppressed osteoclast differentiation in vitro, which was rescued by over-expression of constitutively active S6K159. Moreover, mice with ablation of raptor in osteoclasts with Ctsk-Cre exhibited high bone mass phenotype due to decreased bone resorption59. Besides direct regulation, indirect inhibition of osteoclastogenesis and bone resorption has been reported for mTORC1 signaling in mesenchymal progenitors though not osteoblasts60. Therefore, mTORC1 signaling may exert stage-specific effects on the osteoclast lineage through both direct and indirect actions but a clear understanding about the roles and mechanisms warrants further investigation.
mTORC2 signaling in bone homeostasis and osteoporosis
Like mTORC1, mTORC2 is also implicated in regulating osteoblast differentiation and function. Bone marrow stromal cells (BMSC) lacking Rictor gene exhibited reduced osteogenic potential, but an increased capacity to undergo adipogenic differentiation in vitro30,42,61. Similarly, knockdown of rictor in primary cultures of preosteoblasts impaired their osteogenic differentiation62. Interestingly, expression of Rankl, but neither Opg nor M-CSF, was significantly downregulated in Rictor-deficient BMSC, which exhibited a diminished capability to supporting osteoclastogenesis in vitro42,63. Thus, besides a cell-autonomous role in stimulating osteoblast differentiation, mTORC2 signaling in the osteoblast precursors also indirectly promotes osteoclastogenesis by modulating the expression of Rankl. The stimulatory effect of mTORC2 on both osteoblasts and osteoclasts helps to explain the relatively normal trabecular bone mass when Rictor was deleted in the limb mesenchymal progenitors in the mouse even though the cortical bone mass was reduced42,63. The differential net effect on trabecular versus cortical bone mass in those mice may be due to the more active bone resorption normally occurring in the trabecular bone. Similarly, deletion of rictor in mature osteoblasts simultaneously decreased osteoblast activity and bone resorption in the mouse, leading to notably impaired cortical bone, along with some subtle changes in the trabecular bone mass62.
mTORC2 appears to be a common mediator for both mechanical and biochemical signals to stimulate osteoblast differentiation and bone formation (Fig. 3). Rictor-deficient bones exhibited a lesser anabolic response not only to mechanical loading, but also to the anti-sclerostin antibody therapy that enhances Wnt signaling in bone42,63. The later finding is consistent with biochemical studies demonstrating that bone anabolic Wnt ligands such as Wnt3a, Wnt7b or Wnt10b signal through Lrp5 to activate mTORC2 and to reprogram glucose metabolism28. In addition, mTORC2 was shown to participate in Hedgehog (Hh)-induced osteoblast differentiation, as Hh-Gli2 signaling induced Igf2 expression that activated the mTORC2-Akt-Gli2 cascade further stimulating Hh signaling and osteogenesis29.
Multiple lines of evidence have implicated mTORC2 in age-related osteoporosis. Studies have shown a decrease in the expression of rictor in osteoblastic lineage cells during aging and that decreased rictor expression could contribute to the age-related switch from osteoblast to adipocyte differentiation64,65. Deletion of rictor in osteoblasts accelerated age-related bone loss in the mouse64. Interestingly, several miRNA including miR-188 and miR-218 increase with aging in either BMSC or osteoblasts, and may be responsible for the age-dependent decrease in rictor expression64,65. Thus, mTORC2 may serve as a potential therapeutic target for treating age-related bone loss.
mTOR signaling and osteoarthritis
Osteoarthritis (OA) is a chronic degenerative joint disease characterized by gradual loss of articular cartilage, synovial inflammation, and subchondral bone remodeling. Recent studies have shown that mTOR was up-regulated in human OA cartilage and the articular cartilage of dogs and mice with injury-induced OA66,67. Moreover, activation of mTORC1 signaling via conditional ablation of Tsc1 in osteochondral progenitors with Col2a1-Cre caused spontaneous OA in mice, whereas inducible deletion of Tsc1 in chondrocytes in two-month-old mice promoted progression of aged-related and surgery-induced OA67. Mechanistically, activation of mTORC1 reduced expression of FGFR3 and PTH/PTHrP receptor in chondrocytes, probably through p73 and ERK1/267. Conversely, inhibition of mTORC1 signaling either pharmacologically or genetically attenuated OA pathology in animal models68,69,70,71. In particular, systemic administration of rapamycin significantly reduced cartilage degeneration and synovial inflammation in a murine model of OA. Similarly, local administration of rapamycin through intra-articular injection inhibited chondrocyte hypertrophy and the expression of angiogenic factor VEGF by the articular cartilage in a murine injury model, therefore attenuating OA progression. Likewise, intra-articular injection of Torin 1, a potent inhibitor of both mTORC1 and mTORC2, significantly alleviated articular cartilage degeneration in a rabbit model of collagenase-induced OA partly through suppression of MMP13 and VEGF71. Moreover, genetic ablation of mTOR in chondrocytes reduced chondrocyte apoptosis and the expression of MMP13 in a surgery-induced OA model, thus alleviating cartilage degradation66. The ablation of mTOR in chondrocytes also suppressed TGF-β/Smad3 signaling in synovial tissues, thus decreasing synovial fibrosis66. Thus, multiple lines of evidence support the notion that hyperactive mTOR signaling contributes to OA pathogenesis.
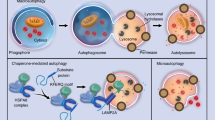
The mechanism underlying the contribution of aberrant mTORC1 activation to OA is not completely understood. Recent studies have implicated autophagy as an important downstream mediator of mTORC1 signaling in OA pathogenesis. Autophagy is an intracellular homeostatic mechanism responsible for degrading and recycling defective macromolecules and cytoplasmic organelles, and is critical for cell survival. A number of studies showed that the expression of major autophagy markers were suppressed in human OA cartilage as well as in animal models of OA66,72. Moreover, inhibition of autophagy caused chondrocyte apoptosis and OA-like pathogenesis in vitro and in vivo73,74. Consistent with the role of mTORC1 as a major suppressor of autophagy, chondrocyte-specific activation of mTORC1 reduced the expression of key autophagy genes in the articular cartilage and caused an OA phenotype in mice75. Conversely, inhibition of mTORC1 signaling by either Rapamycin or Torin or by genetic deletion of mTOR in chondrocytes increased autophagy and attenuated OA progression66,71. Strikingly, inhibition of autophagy negated the protective effects of rapamycin on OA phenotypes66. Thus, suppression of autophagy in response to hyperactive mTORC1 signaling appears to be an important contributor to OA progression.
Future directions
Despite the rapid progress in understanding the role of mTOR singaling in the skeleton, many challenges remain. For instance, the signal inputs to mTOR pathways and the corresponding mechanisms for activating mTORC1 versus mTORC2 are not fully understood76. Although multiple growth factors including Wnt, Igf, and Bmp can stimulate mTOR signaling in the skeleton, their relative contribution, likely dependent on the cellular context and the niche environment, is yet to be explored. Moreover, it is not clear how mTOR signaling is regulated by the nutrient status in chondrocytes, osteoblasts or osteoclasts. Acquiring such knowledge would require comprehensive biochemical studies in vitro, as well as skeleton-specific genetic studies in vivo.
It is important to identify specific downstream effector(s) mediating physiological or pathological functions of mTOR complexes in the skeleton. A recent report revealed that S6K1 only partially mediated the osteogenic effect of Wnt-mTORC1 signaling77. As previous work has implicated S6K1 in mediating mTORC1 signaling in aging, it would be of interest to determine whether S6K1 mediates the role of mTORC1 in the pathogenesis of OA, an age-related degenerative disease78. Such information could be of clinical value as specific S6K1 inhibitors have been developed and may be tested for therapeutic potentials in OA79.
A major challenge for targeting mTOR for therapeutic use lies with the very fact that mTOR signaling plays critical roles in many tissues and physiological processes. Although pharmacological inhibitors of mTORC1, such as rapamycin, may be adjusted to achieve partial inhibition of mTORC1 signaling, the long-term effect of mTORC1 inhibition is still uncertain. Moreover, truly specific inhibitors for mTORC1 versus mTORC2 are still lacking. Even though rapamycin is commonly considered as an mTORC1-specific inhibitor, prolonged rapamycin treatments also compromise mTORC2 signaling. Instead of directly suppressing mTOR, the future of drug development in this area may depend on tissue-specific mTOR modulators and/or process-specific downstream effectors. Identification of such modulators or effectors will also allow for development of agonists of the mTOR-dependent pathways that may be useful for stimulating bone growth in the case of osteoporosis and bone fractures.
References
Vezina, C., Kudelski, A. & Sehgal, S. N. Rapamycin (AY-22,989), a new antifungal antibiotic. I. Taxonomy of the producing streptomycete and isolation of the active principle. J. Antibiot. 28, 721–726 (1975).
Chung, J., Kuo, C. J., Crabtree, G. R. & Blenis, J. Rapamycin-FKBP specifically blocks growth-dependent activation of and signaling by the 70 kd S6 protein kinases. Cell 69, 1227–1236 (1992).
Heitman, J., Movva, N. R. & Hall, M. N. Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science 253, 905–909 (1991).
Cafferkey, R. et al. Dominant missense mutations in a novel yeast protein related to mammalian phosphatidylinositol 3-kinase and VPS34 abrogate rapamycin cytotoxicity. Mol. Cell. Biol. 13, 6012–6023 (1993).
Brown, E. J. et al. A mammalian protein targeted by G1-arresting rapamycin-receptor complex. Nature 369, 756–758 (1994).
Sabatini, D. M., Erdjument-Bromage, H., Lui, M., Tempst, P. & Snyder, S. H. RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs. Cell 78, 35–43 (1994).
Sabers, C. J. et al. Isolation of a protein target of the FKBP12-rapamycin complex in mammalian cells. J. Biol. Chem. 270, 815–822 (1995).
Sarbassov, D. D. et al. Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB. Mol. Cell 22, 159–168 (2006).
Lamming, D. W. et al. Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity. Science 335, 1638–1643 (2012).
Sarbassov, D. D. et al. Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr. Biol. 14, 1296–1302 (2004).
Jacinto, E. et al. Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat. Cell Biol. 6, 1122–1128 (2004).
Hara, K. et al. Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110, 177–189 (2002).
Kim, D. H. et al. GbetaL, a positive regulator of the rapamycin-sensitive pathway required for the nutrient-sensitive interaction between raptor and mTOR. Mol. Cell 11, 895–904 (2003).
Sancak, Y. et al. PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol. Cell 25, 903–915 (2007).
Peterson, T. R. et al. DEPTOR is an mTOR inhibitor frequently overexpressed in multiple myeloma cells and required for their survival. Cell 137, 873–886 (2009).
Jacinto, E. et al. SIN1/MIP1 maintains rictor-mTOR complex integrity and regulates Akt phosphorylation and substrate specificity. Cell 127, 125–137 (2006).
Pearce, L. R. et al. Identification of Protor as a novel Rictor-binding component of mTOR complex-2. Biochem. J. 405, 513–522 (2007).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Inoki, K., Li, Y., Xu, T. & Guan, K. L. Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev. 17, 1829–1834 (2003).
Tee, A. R., Manning, B. D., Roux, P. P., Cantley, L. C. & Blenis, J. Tuberous sclerosis complex gene products, Tuberin and Hamartin, control mTOR signaling by acting as a GTPase-activating protein complex toward Rheb. Curr. Biol. 13, 1259–1268 (2003).
Bar-Peled, L. & Sabatini, D. M. Regulation of mTORC1 by amino acids. Trends Cell Biol. 24, 400–406 (2014).
Wang, S. et al. Metabolism. Lysosomal amino acid transporter SLC38A9 signals arginine sufficiency to mTORC1. Science 347, 188–194 (2015).
Rebsamen, M. et al. SLC38A9 is a component of the lysosomal amino acid sensing machinery that controls mTORC1. Nature 519, 477–481 (2015).
Jewell, J. L. et al. Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 347, 194–198 (2015).
Thomas, J. D. et al. Rab1A is an mTORC1 activator and a colorectal oncogene. Cancer Cell 26, 754–769 (2014).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Martina, J. A., Chen, Y., Gucek, M. & Puertollano, R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8, 903–914 (2012).
Esen, E. et al. WNT-LRP5 signaling induces Warburg effect through mTORC2 activation during osteoblast differentiation. Cell Metab. 17, 745–755 (2013).
Shi, Y., Chen, J., Karner, C. M. & Long, F. Hedgehog signaling activates a positive feedback mechanism involving insulin-like growth factors to induce osteoblast differentiation. Proc. Natl Acad. Sci. USA 112, 4678–4683 (2015).
Sen, B. et al. mTORC2 regulates mechanically induced cytoskeletal reorganization and lineage selection in marrow-derived mesenchymal stem cells. J. Bone Miner. Res. 29, 78–89 (2014).
Zinzalla, V., Stracka, D., Oppliger, W. & Hall, M. N. Activation of mTORC2 by association with the ribosome. Cell 144, 757–768 (2011).
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005).
Garcia-Martinez, J. M. & Alessi, D. R. mTOR complex 2 (mTORC2) controls hydrophobic motif phosphorylation and activation of serum- and glucocorticoid-induced protein kinase 1 (SGK1). Biochem. J. 416, 375–385 (2008).
Long, F. & Ornitz, D. M. Development of the endochondral skeleton. Cold Spring Harb. Perspect. Biol. 5, a008334 (2013).
Jiang, M., Fu, X., Yang, H., Long, F. & Chen, J. mTORC1 Signaling Promotes Limb Bud Cell Growth and Chondrogenesis. J. Cell. Biochem. 118, 748–753 (2017).
Phornphutkul, C., Wu, K. Y., Auyeung, V., Chen, Q. & Gruppuso, P. A. mTOR signaling contributes to chondrocyte differentiation. Dev. Dyn. 237, 702–712 (2008).
Oh, C. D. et al. Immunosuppressant rapamycin inhibits protein kinase C alpha and p38 mitogen-activated protein kinase leading to the inhibition of chondrogenesis. Eur. J. Pharmacol. 427, 175–185 (2001).
Chen, J. & Long, F. mTORC1 signaling controls mammalian skeletal growth through stimulation of protein synthesis. Development 141, 2848–2854 (2014).
Yan, B. et al. mTORC1 regulates PTHrP to coordinate chondrocyte growth, proliferation and differentiation. Nat. Commun. 7, 11151 (2016).
Shima, H. et al. Disruption of thep70(s6k)/p85(s6k) gene reveals a small mouse phenotype and a new functional S6 kinase. EMBO J. 17, 6649–6659 (1998).
Selman, C. et al. Ribosomal protein S6 kinase 1 signaling regulates mammalian life span. Science 326, 140–144 (2009).
Chen, J., Holguin, N., Shi, Y., Silva, M. J. & Long, F. mTORC2 signaling promotes skeletal growth and bone formation in mice. J. Bone Mineral. Res. 30, 369–378 (2015).
Singha, U. K. et al. Rapamycin inhibits osteoblast proliferation and differentiation in MC3T3-E1 cells and primary mouse bone marrow stromal cells. J. Cell Biochem. 103, 434–446 (2008).
Xian, L. et al. Matrix IGF-1 maintains bone mass by activation of mTOR in mesenchymal stem cells. Nat. Med. 18, 1095–1101 (2012).
Chen, J. et al. WNT7B promotes bone formation in part through mTORC1. PLoS Genet. 10, e1004145 (2014).
Karner, C. M., Esen, E., Okunade, A. L., Patterson, B. W. & Long, F. Increased glutamine catabolism mediates bone anabolism in response to WNT signaling. J. Clin. Investig. 125, 551–562 (2015).
Karner, C. M., Lee, S. Y. & Long, F. Bmp Induces Osteoblast Differentiation through both Smad4 and mTORC1 Signaling. Mol. Cell. Biol. 37, pii: e00253-16 (2017).
Lim, J. et al. Dual function of Bmpr1a signaling in restricting preosteoblast proliferation and stimulating osteoblast activity in mouse. Development 143, 339–347 (2016).
Chen, J. & Long, F. mTORC1 signaling promotes osteoblast differentiation from preosteoblasts. PLoS ONE 10, e0130627 (2015).
Fitter, S. et al. mTORC1 Plays an important role in skeletal development by controlling preosteoblast differentiation. Mol. Cell. Biol. 37, pii: e00668-16 (2017).
Huang, B. et al. mTORC1 prevents preosteoblast differentiation through the notch signaling pathway. PLoS Genet. 11, e1005426 (2015).
Carbonara, C. et al. 9q34 loss of heterozygosity in a tuberous sclerosis astrocytoma suggests a growth suppressor-like activity also for the TSC1 gene. Hum. Mol. Genet. 3, 1829–1832 (1994).
Green, A. J., Smith, M. & Yates, J. R. Loss of heterozygosity on chromosome 16p13.3 in hamartomas from tuberous sclerosis patients. Nat. Genet. 6, 193–196 (1994).
Osborne, J. P., Fryer, A. & Webb, D. Epidemiology of tuberous sclerosis. Ann. NY Acad. Sci. 615, 125–127 (1991).
Fang, F., Wei, X., Hu, M. & Liu, F. A mouse model of craniofacial bone lesion of tuberous sclerosis complex. Musculoskelet Regen 1, pii: e814 (2015).
Fang, F. et al. Neural crest-specific TSC1 deletion in mice leads to sclerotic craniofacial bone lesion. J. Bone Mineral. Res. 30, 1195–1205 (2015).
Riddle, R. C. et al. Tsc2 is a molecular checkpoint controlling osteoblast development and glucose homeostasis. Mol. Cell. Biol. 34, 1850–1862 (2014).
Zhang, Y. et al. mTORC1 inhibits NF-kappaB/NFATc1 signaling and prevents osteoclast precursor differentiation, in vitro and in mice. J. Bone Miner. Res. 32, 1829–1840 (2017).
Dai, Q. et al. Inactivation of Regulatory-associated Protein of mTOR (Raptor)/Mammalian Target of Rapamycin Complex 1 (mTORC1) Signaling in Osteoclasts Increases Bone Mass by Inhibiting Osteoclast Differentiation in Mice. J. Biol. Chem. 292, 196–204 (2017).
Wu, H. et al. Bone size and quality regulation: concerted actions of mtor in mesenchymal stromal cells and osteoclasts. Stem Cell Reports. 8, 1600–1616 (2017).
Martin, S. K. et al. Brief report: the differential roles of mTORC1 and mTORC2 in mesenchymal stem cell differentiation. Stem Cells 33, 1359–1365 (2015).
Liu, D. M. et al. Rictor/mTORC2 loss in osteoblasts impairs bone mass and strength. Bone 90, 50–58 (2016).
Sun, W., Shi, Y., Lee, W. C., Lee, S. Y. & Long, F. Rictor is required for optimal bone accrual in response to anti-sclerostin therapy in the mouse. Bone 85, 1–8 (2016).
Lai, P. et al. Loss of Rictor with aging in osteoblasts promotes age-related bone loss. Cell Death Dis. 7, e2408 (2016).
Li, C. J. et al. MicroRNA-188 regulates age-related switch between osteoblast and adipocyte differentiation. J. Clin. Invest. 125, 1509–1522 (2015).
Zhang, Y. et al. Cartilage-specific deletion of mTOR upregulates autophagy and protects mice from osteoarthritis. Ann. Rheum. Dis. 74, 1432–1440 (2015).
Zhang, H. et al. mTORC1 activation downregulates FGFR3 and PTH/PTHrP receptor in articular chondrocytes to initiate osteoarthritis. Osteoarthritis Cartilage 25, 952–963 (2017).
Matsuzaki, T. et al. Intra-articular administration of gelatin hydrogels incorporating rapamycin-micelle reduces development of experimental osteoarthritis in a murine model. Biomaterials 21, 9904–9911 (2013).
Takayama, K. et al. Local intra-articular injection of rapamycin delays articular cartilage degeneration in a murine model of osteoarthritis. Arthritis Res. Ther. 16, 1–10 (2014).
Caramés, B. Autophagy activation by rapamycin reduces severity of experimental osteoarthritis. Ann. Rheum. Dis. 71, 575–581 (2012).
Cheng, N. T., Guo, A. & Cui, Y. P. Intra-articular injection of Torin 1 reduces degeneration of articular cartilage in a rabbit osteoarthritis model. Bone Joint Res. 5, 218–224 (2016).
Carames, B., Taniguchi, N., Otsuki, S., Blanco, F. J. & Lotz, M. Autophagy is a protective mechanism in normal cartilage, and its aging-related loss is linked with cell death and osteoarthritis. Arthritis Rheum. 62, 791–801 (2010).
Sasaki, H. et al. Autophagy modulates osteoarthritis-related gene expression in human chondrocytes. Arthritis Rheum. 64, 1920–1928 (2012).
Cheng, N. T. et al. Role of autophagy in the progression of osteoarthritis: The autophagy inhibitor, 3-methyladenine, aggravates the severity of experimental osteoarthritis. Int. J. Mol. Med. 39, 1224–1232 (2017).
Vasheghani, F. et al. PPARgamma deficiency results in severe, accelerated osteoarthritis associated with aberrant mTOR signaling in the articular cartilage. Ann. Rheum. Dis. 74, 569–578 (2015).
Saxton, R. A. & Sabatini, D. M. mTOR Signaling in Growth, Metabolism, and Disease. Cell 169, 361–371 (2017).
Gu, X. et al. Pharmacological inhibition of S6K1 impairs self-renewal and osteogenic differentiation of bone marrow stromal cells. J. Cell. Biochem. In press, https://doi.org/10.1002/jcb.26272 (2017).
Harrison, D. E. et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 460, 392–395 (2009).
Hollebecque, A. et al. A phase Ib trial of LY2584702 tosylate, a p70 S6 inhibitor, in combination with erlotinib or everolimus in patients with solid tumours. Eur. J. Cancer 50, 876–884 (2014).
Acknowledgements
This work is funded by the National Key R&D Program of China (2016YFC1100203) (J.C.), the Priority Academic Program Development of Jiangsu High Education Institutions (PAPD) (J.C.), the National Natural Science Foundation of China (81772294) (J.C.) and NIH R01 AR060456 and R01 AR055923 (F.L.).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Conflict of interest
:The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, J., Long, F. mTOR signaling in skeletal development and disease. Bone Res 6, 1 (2018). https://doi.org/10.1038/s41413-017-0004-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41413-017-0004-5
This article is cited by
-
p53 modulates kinase inhibitor resistance and lineage plasticity in NF1-related MPNSTs
Oncogene (2024)
-
Sericin promotes chondrogenic proliferation and differentiation via glycolysis and Smad2/3 TGF-β signaling inductions and alleviates inflammation in three-dimensional models
Scientific Reports (2024)
-
Inadequate linear catch-up growth in children born small for gestational age: Influencing factors and underlying mechanisms
Reviews in Endocrine and Metabolic Disorders (2024)
-
New insights into the mechanisms and therapeutic strategies of chondrocyte autophagy in osteoarthritis
Journal of Molecular Medicine (2024)
-
Screening for autophagy/hypoxia/ferroptosis/pyroptosis-related genes of tendon injury and repair in a rat model after celecoxib and lactoferrin treatment
Journal of Orthopaedic Surgery and Research (2023)