Abstract
Plk1-interacting checkpoint helicase (PICH) is a DNA translocase involved in resolving ultrafine anaphase DNA bridges and, therefore, is important to safeguard chromosome segregation and stability. PICH is overexpressed in various human cancers, particularly in lymphomas such as Burkitt lymphoma, which is caused by MYC translocations. To investigate the relevance of PICH in cancer development and progression, we have combined novel PICH-deficient mouse models with the Eμ-Myc transgenic mouse model, which recapitulates B-cell lymphoma development. We have observed that PICH deficiency delays the onset of MYC-induced lymphomas in Pich heterozygous females. Moreover, using a Pich conditional knockout mouse model, we have found that Pich deletion in adult mice improves the survival of Eμ-Myc transgenic mice. Notably, we show that Pich deletion in healthy adult mice is well tolerated, supporting PICH as a suitable target for anticancer therapies. Finally, we have corroborated these findings in two human Burkitt lymphoma cell lines and we have found that the death of cancer cells was accompanied by chromosomal instability. Based on these findings, we propose PICH as a potential therapeutic target for Burkitt lymphoma and for other cancers where PICH is overexpressed.
Similar content being viewed by others
Introduction
Burkitt lymphoma (BL) is the most frequent non-Hodgkin’s malignant lymphoma (NHML) to affect children in Europe, and also accounts for 2% of adult NHML [1]. It is an aggressive tumor that arises in various parts of the lymphatic system and is one of the fastest-growing tumors [2]. Although most patients respond well to standard chemotherapy, some patients, especially adults, with relapsed disease have a poor prognosis [3]. Thus, it is still necessary to identify mechanisms and molecular targets that may provide early prognostic markers and/or critical targets for the development of new drugs that block BL initiation and progression. In this regard, targeting pathways involved in the maintenance of chromosomal stability has emerged in the last few years as a promising cancer research area [4].
Faithful chromosomal segregation is critical for the maintenance of genomic stability in every cell division. Polo-like kinase 1 (Plk1)-interacting checkpoint helicase (PICH), encoded by the gene ERCC6L, is required for the resolution of anaphase Ultrafine DNA Bridges (UFBs) and hence for proper chromosome segregation. PICH localizes in the cytosol during interphase but associates with the nuclear DNA after the nuclear envelope breaks down during mitosis [5, 6]. PICH is a DNA translocase with high affinity for DNA under tension [7] and characteristically localizes to UFBs and contributes to their resolution [5, 8]. UFBs are catenated DNA structures derived from unresolved DNA replication intermediates that link the sister chromatids of mitotic chromosomes in anaphase [9, 10]. UFBs can arise from different chromosomal loci which are difficult to replicate: telomeres, centromeres, common fragile sites, and ribosomal DNA [9,10,11]. Failure in the resolution of UFBs at the end of anaphase may lead to chromosomal instability in the daughter cells [8, 12]. UFBs may also play beneficial roles in the maintenance of chromosome structure in rapidly proliferating cells [9]. Since UFBs are dechromatinized and cannot be stained with DNA dyes like DAPI, they can only be visualized via proteins that are associated with them, such as PICH or the Bloom’s syndrome helicase (BLM) [8, 10]. PICH recruits other UFB-associated factors such as BLM or RIF1 [6, 8] and stimulates the catalytic activity of the TOP2α topoisomerase in order to resolve UFBs [12, 13].
We have recently generated a Pich knock-out (KO) mouse model that contributed to understanding PICH function in vivo [14]. PICH is essential for embryonic development since the loss of Pich generates chromosomal instability, DNA damage, p53 activation, and apoptosis in Pich KO embryos that are incompatible with sustained viability [14]. More recently, it has been proposed that PICH supports embryonic hematopoiesis by suppressing a cGAS-STING-mediated type I interferon production [15].
The contribution of PICH deficiency-induced chromosomal instability to tumorigenesis in vivo is largely unknown. In fact, whether chromosomal instability (CIN) causes tumor development is still a matter of debate [16, 17]. In contrast, our previous findings indicate that PICH is of critical importance for supporting intense bouts of cell proliferation such as those that occur during embryonic development and the proliferation of transformed cells [14]. Recently, several papers support the notion that PICH activity is relevant to sustain the proliferation of certain types of cancerous cells [18,19,20,21,22,23,24,25]. In addition, PICH is overexpressed in several types of solid human cancers [18, 19, 21,22,23,24,25,26,27,28]. High expression of PICH has been linked to worse outcomes and decreased overall survival in cancer patients [14, 18, 19, 21, 23, 24, 27,28,29]. However, the role of PICH in hematopoietic malignancies and, in particular, in BL has not been explored yet.
In this study, we have investigated the consequences of PICH depletion in adult mice; and the impact of PICH depletion in BL, using mouse models and human BL cells. Our results indicate that PICH is highly expressed in BL. We also show that Pich deficiency delays the development of Myc-induced B-cell lymphoma. Furthermore, inducible systemic Pich deletion in adult Eµ-Myc mice increases their survival without causing toxic effects. PICH-depleted tumors derived from these mice and human BL cells have a significantly elevated frequency of apoptosis, persistent DNA bridges, bi- and poly nucleation, and micronuclei. Thus, we demonstrate that PICH is required for the growth of Myc-induced B-cell lymphoma in vivo and for human BL cells in vitro. In view of these results, we propose PICH as a potential novel target for the treatment of human BL.
Methods
Analysis of PICH expression from human databases
PICH mRNA expression in human cell lines and tissues was obtained from Cancer Cell Line Encyclopedia (CCLE) and the TCGA, respectively. RNA-seq expression data from TCGA (http://cancergenome.nih.gov/) were obtained through GEPIA (http://gepia.cancer-pku.cn/) to compare PICH expression levels between tumors and paired normal tissues. PICH mRNA expression in human samples from different subtypes of mature B-cell malignancies were obtained from an MD Anderson Cancer Center [30] study via cBioportal. DepMap portal (https://depmap.org/portal/) was used to analyze PICH expression levels across various human cancer cell lines from the Cancer Cell Lines Encyclopedia.
Mice
The Pich conditional KO (PICH cKO) and constitutive Pich KO mouse models were generated in our laboratory and have been described previously [14]. The Pich cKO strain has a mixed 129S2; C57BL/6 N background. The UBC-Cre-ERT2 (Jax strain 007001) and Eµ-Myc (Jax strain 002728) [31] strains have been described previously. Activation of the inducible UBC-Cre-ERT2 recombinase was achieved by injecting the Pich cKO with Tamoxifen (TAM, T5648, Sigma-Aldrich) into the peritoneum once every 24 h for 5 consecutive days followed by one injection per week in the subsequent 2 weeks. All mice were housed under standard conditions at the animal facility of the Department of Experimental Medicine at the University of Copenhagen and the research was monitored by the Institutional Animal Care and Use Committee. All animal experiments were approved by the Danish Animal Ethics Committee (licenses 2016-15-0201-011111 and 2019-15-0201-00344) and performed in compliance with Danish and European regulations.
Mouse genotyping
Mice were genotyped for Pich gene using three primers as described in [14] Pich_F1 (5′-CTATGCCTGATCCTCCCCAG-3′), Pich_R1 (5′-GCTAACAGACAAAATGGCCCT-3′) and Pich_F2 (5′-AAAGCCCAACTACAGTGTGG-3′). A 390-bp WT fragment and a 495-bp PichLox fragment were amplified using primers Pich _F1 and Pich _R1. The Pich KO allele was amplified by the Pich _R1 and Pich _F2 primer and generated a 220-bp fragment. Other primers used for genotyping were gCre5 (5′-TGGTTTCCCGCAGAACCTGAAG-3′ and gCre3: 5′-GAGCCTGTTTTGCACGTTCACC-3′) for Cre gene amplification, and OIMR0375 (5′-CAGCTGGCGTAATAGCGAAGAG-3′), OIMR0376 (5′- CTGTGACTGGTGAGTACTCAACC-3′), Tshb_Fw (5′- CTA CAT GAG CAG GCA GAC TGG A-3′) and Tshb_Rv (5′-AAT GGA CGG GTG CTT CTA TC-3′) for amplification of the Myc gene.
Immunohistochemistry
Tissues were fixed in 4% neutral buffered formalin (Sigma-Aldrich), embedded in paraffin and processed according to standard procedures. Immunohistochemistry (IHC) of mouse samples was carried out as described previously [14]. The primary antibodies used for mouse samples were PICH (Cell Signaling Technology, 8886, D4G8; 1:50;), γH2AX (MilliporeSigma, 05-636, JBV301; 1:25,000;) and cleaved caspase-3 (Cell Signaling Technology, 9661; 1:750).
The human lymphoma tissue microarray (TMA), was acquired from US Biomax (LM482c). Paraffin-embedded sections were deparaffinized in xylene and rehydrated through graded concentrations of ethanol in water. Following antigen retrieval with citrate buffer (pH 6.0), endogenous peroxidase was blocked using 3% hydrogen peroxide and sections were incubated overnight at 4 °C with anti-PICH (MyBiosource, MBS2519675, 1:100). Sections were then washed and incubated with secondary antibody for 30 min. IHC reaction was developed using 3,30-diaminobenzidine tetrahydrochloride (DAB) (Chromomap DAB, Ventana, Roche; DAB+ Chromogen System, Dako) and nuclei were counterstained with hematoxylin. Images were acquired using a slide scanner (AxioScan Z1, Zeiss).
Cell culture
Raji and Ramos BL cells were grown in RPMI (Roswell Park Memorial Institute) medium supplemented with 10% of fetal bovine serum (FBS) (Life Technologies) and 1% of penicillin/streptomycin (p/s) (Life Technologies). HEK293T cells were grown in DMEM supplemented with 10% of FBS, 1% of p/s and 1% of l-glutamin. All cells were incubated at 5% CO2 and 37 °C in a ThermoQuest incubator (Forma Scientific, Marietta, OH, USA).
Western blotting
Cells were lysed in RIPA buffer (Tris-HCl 50 mM, pH 7.4, NP-40 1%, Na-deoxycholate 0.25%, NaCl: 150 mM, EDTA 1 mM) supplemented with complete protease inhibitor cocktail tablet (Roche), 5 mM β-glycerophosphate (Sigma), 5 mM sodium fluoride (Sigma) and 1 mM sodium orthovanadate (Sigma). The total amount of protein was determined by DC Protein Assay kit (Bio-Rad) according to supplier’s instructions. Cell lysates containing 30 µg protein were incubated in NuPAGE™ LDS Sample Buffer 4x (Novex) with 10 mM DTT for 15 min at 70 °C. Samples were resolved by SDS-PAGE and Western blotting was carried out as described previously [32]. Antibodies against PICH (Cell Signaling Technology, 8886; 1:1000) and β-actin (Sigma, A5441; 1:1000) were used. Goat anti-rabbit immunoglobulin G (IgG) HRP (Sigma, A6667; 1:10000) and goat anti-mouse IgG HRP (Sigma-Aldrich, A4416; 1:10000) were used as secondary antibodies. Images were acquired on an Amersham™ Imager 600 (GE Healthcare Life Sciences). Measurements for the protein of interest were normalized to the loading control.
Lentivirus synthesis
HEK293T cells were used for the synthesis of third-generation lentiviruses containing short hairpin RNA (shRNA) targeting human PICH (shPICH) or the shRNA control pLKO.1 (shCtrl) (#8453, Addgene) vector. Briefly, 6 × 106 of cells were reverse transfected using lipofectamine 2000 with 10 μg of the plasmid of interest and the plasmids coding for the lentivirus packaging components (6.5 µg pMMDLRRE, 2.5 µg PRSVREV and 3.5 µg PMDGVSVG) (#12251, #12253 and #12259, respectively, Addgene). Forty-eight hours post-transfection, the viruses were filtered (0.25 µm) and collected. shPICH were acquired from Sigma (MISSION® shRNA Bacterial Glycerol Stock, SHCLNG-NM_017669) and their sequences were as follows: shPICH-1 5’-TATTCTGAGCACTAGCTTAAT-3’; shRNA-2 5’- ACAAGATCTCTCCAGTATAAA -3’; shRNA-3 5’- GATCATGCCAACCAATCTTAT -3’.
Cell Infection
A total of 2 × 105 Raji or Ramos cells were infected with lentivirus containing the vector of interest and 10 µg/ml of polybrene to facilitate the infection and were incubated for 3 h. Then, the cells and lentivirus mix were then seeded into a 12-well plate. Two days after transduction, cells were selected with 2 µg/mL of puromycin for three days. The selected cells were cultured and used as indicated.
Cell proliferation assay
Cells were seeded in triplicates in 12-well plates at a density of 5 × 105 cells per well. Cells were mixed with trypan blue (1:1 ratio) (Sigma) for staining and manually counted using a hemocytometer at the time points indicated in the figures over 7 days after selection with puromycin. Non-stained live cell numbers were plotted using GraphPad Prism software (GraphPad Software, San Diego, CA, USA, www.graphpad.com). Doubling times (DT) were calculated by non-linear regression using an exponential growth equation.
Viability assays
Three independent methods were used to evaluate the viability of PICH knockdown (KD) cells. For all methods, cells were seeded six days after lentiviral infection and viability was assessed four days later. Three technical replicates were seeded for each condition. For manual counting, cells were seeded and stained with trypan blue as described above. For the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazoliumbromide (MTT) assay, cells were seeded in 96-well plates at a density of 5 × 104 cells per well. Three days later, cells were incubated with MTT solution (Sigma-Aldrich, 1 mg/ml) for four hours. Formazan was solubilized in DMSO and absorbances at 570 nm and at 690 nm (background) were measured using a Varioskan plate reader (Thermo Fisher Scientific). Normalized absorbance values relative to shCtrl cells were plotted using GraphPad Prism. Finally, Hoechst 33342 and To-Pro-3 staining assay was performed to evaluate the number of total and dead cells, respectively. PICH KD cells were seeded in µCLEAR 96-well plates (Grenier Bio-One) at a density of 1.5 × 104 cells per well. Next, cells were incubated with Hoechst 33342 and To-Pro-3 at a final concentration of 5 µg/ml and 1 µM, respectively, for 30 min in a 5% CO2 incubator at 37 °C. Images were acquired automatically by a High-Content Screening System (MetaXpress) and analyzed with the MetaXpress High-Content Image Analysis Software (Molecular Devices). The number of living cells was calculated by subtracting the number of dead cells, given by the To-Pro-3 stain, from the total cell number, given by the Hoechst stain, and plotted using GraphPad Prism.
Immunofluorescence and microscopy
Fifty thousand cells per well were seeded in triplicates in µCLEAR 96-well plates (Grenier Bio-One) and left steady for 30 min. Cells were fixed with 4% formaldehyde (VWR Chemicals, Radnor, PA, USA) for 15 min at room temperature (RT) and permeabilized with 0.5% Triton X-100 (v/v) in PBS for 10 min. Then, cells were washed twice with PBS supplemented with 0.05% Tween-20, blocked with 3% BSA (Sigma) for 30 min and then labeled with α-tubulin primary antibody (Sigma, St. Louis, MO, USA, T9026, 1:500). Next, fluorescence-tagged secondary antibody (Alexa Fluor™ Goat Anti-mouse IgG 488 (Invitrogen) was added for 2 h at RT in the dark. DAPI was used for nuclear staining. Images were automatically acquired using a High-Content Screening System (MetaExpress) and analyzed with the MetaXpress software (Molecular Devices). Single plane images corresponding to Z positions of the maximal DAPI signal were acquired. At least nine images were acquired per well.
Annexin V-FITC kit
Apoptosis was evaluated by flow cytometry analysis using an Annexin V-FITC apoptosis detection kit from Miltenyi Biotec (Bergisch Gladbach, Germany) following the manufacturer’s instructions. Briefly, 1 × 106 cells were resuspended in binding buffer, stained with Annexin V-FITC antibody for 15 min in the dark and washed with binding buffer. Next, cells were stained with propidium iodide (PI) immediately prior to analysis in a FACSCalibur cytometer from BD Biosciences (San Jose, CA, USA). Data were analyzed using BD CellQuest Pro Software.
Cell cycle analysis
A total of 1 × 106 cells were fixed in ice-cold 70% ethanol in PBS, then pelleted and resuspended in PBS containing 100 µg/mL RNase A and 20 µg/mL PI(Sigma P4170) for 30 min. Cells were captured by a FACSCalibur cytometer from BD Biosciences (San Jose, CA, USA) and analyzed using ModFit Software (Verity Software House).
EdU incorporation
The DNA replication rate was determined by EdU incorporation using Click-iT technology following the manufacturer’s protocol (Life Technologies). Briefly, 5 × 105 cells were seeded in a T25 flask the day before the staining. EdU (Life Technologies A10044) was added to the culture medium at a final concentration of 10 μM for 60 min prior to fixing the cells with 4% formaldehyde (VWR Chemicals, Radnor, PA, USA) and permeabilizing with 0.1% Triton X-100 (v/v) in PBS. Next, a dilution of 1 mM ascorbic acid was prepared fresh, and a Click-it reaction mix was performed by mixing PBS, 2 mM CuSO4, 2.5 µg/mL Azide 488, and 1 mM ascorbic acid in the given order, which was then added to each cell condition for 60 min (protected from light). Cells were stained with 20 µg/mL PI and treated with 100 µg/mL RNAse for 30 min at room temperature and analyzed by FACSCalibur cytometer. Data were plotted and analyzed using ModFit Software (Verity Software House).
Statistical analysis
Statistical analyses were performed using GraphPad Prism 8 (GraphPad Software, San Diego California, USA, www.graphpad.com). The significance was determined by either an unpaired t-test or one-way ANOVA with Tukey post-test. The log-rank test was used to determine P values for all Kaplan–Meier survival curve analyses. P-values are indicated in each graph and figure legends. In vitro experiments were performed with at least three biological replicates.
Results
PICH is overexpressed in BL
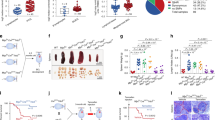
To investigate the clinical relevance of PICH in human cancer, we evaluated PICH mRNA expression in different human cancer tissues and cell lines. We compared PICH mRNA expression in the different cancer types from The Cancer Genome Atlas PanCancer Study and their paired normal tissues. PICH mRNA expression was elevated in multiple malignancies, including myeloid and lymphoid neoplasms, compared with normal tissues (Fig. 1A). PICH showed significantly higher expression in BL among different lymphoma subtypes (Fig. 1B). Accordingly, when analyzing RNA expression data from the Cancer Cell Line Encyclopedia (CCLE) database which consisted of 1,379 cell lines from a diversity of human tissues, PICH mRNA levels were higher in cancer cells from tumors of hematopoietic and lymphoid tissues and, particularly, the highest PICH expression was found in BL (Fig. 1C). BL cell lines also exhibited higher PICH protein expression levels than the non-cancerous and non-transformed human retinal pigment epithelial-1 (RPE-1) cell line (Fig. 1D). Moreover, we performed PICH immunohistochemistry (IHC) analysis of clinical samples from lymphoma and normal lymph nodes, and from mouse spleen and tumors derived from the Eμ-Myc mouse model (Jax strain 2728). Again, human cancer samples showed higher PICH expression compared to normal lymph nodes (Fig. 1E). Mouse spleen tumors derived from the Eμ-Myc mouse model, which develops Myc-induced B-cell lymphoma, also exhibited high PICH expression, in contrast to its lower expression in healthy spleen (Fig. 1F) and in most mouse adult tissues (Fig. 2D). PICH is located in the cytosol of cells in interphase, and its presence on chromatin is restricted to mitosis. As expected, PICH IHC revealed a predominantly cytosolic pattern of PICH localization in interphase cells. Based on these data, we hypothesized that PICH may have a relevant role in BL and that Eμ-Myc mice are a relevant model to investigate this in vivo.
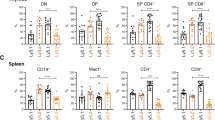
A PICH mRNA expression (TPM, Transcripts per million) in the different cancer types from The Cancer Genome Atlas PanCancer Study and paired normal tissue (Adapted from GEPIA). Fold-change in PICH expression in tumors compared to normal tissue is shown in the right y-axis. Studies are ranked by fold-change. The study acronyms are LUAD: Lung adenocarcinoma, DLBC Lymphoid Neoplasm Diffuse Large B-cell Lymphoma, THYM Thymoma, CHOL Cholangiocarcinoma, GBM Glioblastoma multiforme, PAAD Pancreatic adenocarcinoma, READ Rectum adenocarcinoma, LUSC Lung squamous cell carcinoma, COAD Colon adenocarcinoma, BRCA Breast invasive carcinoma, STAD Stomach adenocarcinoma, SARC Sarcoma, BLCA Bladder Urothelial Carcinoma, SKCM Skin Cutaneous Melanoma, LIHC Liver hepatocellular carcinoma, CESC Cervical squamous cell carcinoma and endocervical adenocarcinoma, TGCT Testicular Germ Cell Tumors, UCEC Uterine Corpus Endometrial Carcinoma, OV Ovarian serous cystadenocarcinoma, UCS Uterine Carcinosarcoma, LGG Brain Lower Grade Glioma, KIRP Kidney renal papillary cell carcinoma, ACC Adrenocortical carcinoma, PCPG Pheochromocytoma and Paraganglioma, ESCA Esophageal carcinoma, HNSC Head and Neck squamous cell carcinoma, KICH Kidney Chromophobe, PRAD Prostate adenocarcinoma, THCA Thyroid carcinoma, LAML Acute Myeloid Leukemia. B Box plot of PICH mRNA expression in different lymphoma subtypes in the Mature B-cell malignancies study from MD Anderson Cancer Center. Boxplot shows 10–90 percentile. n = 60 (Burkitt Lymphoma), n = 95 (Diffuse Large B-cell Lymphoma), n = 65 (Follicular Lymphoma), n = 43 (Mantle Cell Lymphoma), and n = 23 (Marginal Zone Lymphoma). Mean and SEMs are indicated. The levels of PICH expression are significantly higher in BL than in the other four lymphoma subtypes (Diffuse Large B-Cell L., Follicular L., Mantle Cell L., and Marginal Zone L.). Significance was assessed by Tukey’s multiple comparisons test. ****p ≤ 0.0001. The levels are also significantly higher in Diffuse Large B-Cell L. than in Follicular L., Mantle Cell L., and Marginal Zone L. with a P-value ****p ≤ 0.0001 with Tukey’s multiple comparisons test. C PICH expression in human cancer cell lines from the Cancer Cell Line Encyclopedia, adapted from DepMap. Cell lines are grouped by lineage and subtypes are shown for the lymphoid lineage. Boxplot shows 10–90 percentile. D Representative immunoblots of PICH protein levels in BL cell lines and the primary non-tumor cell line RPE-1. β-Actin was used as a control for protein load. E Representative pictures (left) and quantification (right) of PICH expression in BL and normal lymph node tissue array assessed by IHC staining. Scale bars are indicated, 100 μm. Quantification of the percentage of PICH-positive cells is shown on the right. Data shown correspond to six fields per tissue, counting at least 1000 cells per field. Four normal lymph nodes and three BL tumors were assessed. Means and SEMs are indicated. Significance was assessed by unpaired t-test ****p ≤ 0.0001. F Representative images of spleens collected from Myc wild-type (Myc+/+) and Emu-Myc transgenic (Myc+/tg) mice (left) and immunohistochemical staining against Pich on paraffin sections of spleens from mice with the indicated genotypes (right). Scale bar, 50 μm.
A Schematic representation of the alleles of the constitutive Pich-deficient mouse model. Two LoxP sites were inserted flanking the endogenous exon 2 of the Pich locus to generate Pich conditional KO (PichLox) mESC. This PichLox allele was later combined with the UBC-CMV-Cre allele expressing the Cre recombinase to generate mice harboring the Pich KO allele. Coding exons are indicated by filled boxes, and UTR are indicated by hatched boxes. LoxP sites are depicted as red triangles. B Kaplan–Meier survival curves for Pich+/+ (n = 18; blue; median survival 102.5 weeks) and Pich+/- (n = 15; red; median survival 99 weeks) mice. P-value = 0.6704 using the log-rank (Mantel–Cox) test. C Incidence of spontaneous tumors in Pich+/- mice compared to Pich+/+ littermates observed for 150 weeks. The differences observed in tumor incidence were not statistically significant according to the χ2 test. D PICH IHC staining of the indicated tissue sections collected from 50-week-old Pich+/+ and Pich+/- mice. PICH is present in cells of the intestine, spleen, thymus and ovary of both Pich+/+ and Pich+/- mice, but absent in the liver. Scale bar, 50 μm. E Kaplan–Meier survival curves for Pich+/+; Eµ-Myc+/tg (n = 28; blue; median survival 15 weeks), and Pich+/-; Eµ-Myc+/tg (n = 27; red; median survival 21 weeks) mice. P-value = 0.0003 using the log-rank (Mantel–Cox) test; ***p ≤ 0.001. F Immunohistochemical staining against Pich on spleen or tumoral spleen paraffin sections from Pich+/+; Eµ-Myc+/tg, Pich+/+ and Pich+/-; Eµ-Myc+/tg mice (from left to right). Scale bar, 20 μm.
Pich deficiency in heterozygosity does not induce cancer
To investigate the impact of PICH deficiency in cancer, we used a Pich-deficient mouse model previously generated in our lab [14]. This model consists of a Pich conditional KO mouse model with two loxP sites flanking the exon 2 of the murine Pich gene. Pich+/Lox females were combined with UBC-CMV-Cre mice to generate mice harboring a Pich KO allele (Fig. 2A). Whereas complete Pich depletion is embryonic lethal by 13.5 days of embryonic development, Pich heterozygous mice (Pich+/-) are viable and do not show any obvious abnormalities [14]. Given that Pich is located in the X chromosome, only heterozygous females could be investigated, thus, only female mice were included in the cohorts. To test the long-term effects of Pich deficiency, we evaluated the tumor-free survival and spontaneous tumor formation of Pich+/+ and Pich+/- mice for up to 150 weeks. Mice were euthanized when they had noticeable tumors or were visibly ill, as observed by rapid weight loss, hunched posture, rough hair coat, labored breathing, lethargy, impaired mobility or abdominal swelling. Females from both cohorts survived at least 74 weeks and did not show significant differences in their median tumor-free survival rates (Fig. 2B). All Pich+/- mice tested remained healthy and did not show weight loss or behavioral changes during this time. In addition, the percentage of mice that developed spontaneous tumors at the end of the experiment was similar in both Pich+/+ and Pich+/- mice (Fig. 2C). We did not find differences in the number of spontaneous tumors per mouse nor in the tumor types between both cohorts (Supplementary Fig. 1). Histological examination of tissues collected from 50-week-old Pich+/- mice (not included in the survival curve) did not reveal structural abnormalities (Fig. 2D). Analysis of adult tissues from Pich+/+ mice confirmed high PICH expression in tissues with an elevated percentage of proliferating cells, including those of the intestines, spleen, thymus, and ovaries in contrast to the liver that do not express PICH at detectable levels. Interestingly, Pich+/- mice exhibit similar PICH expression patterns to Pich+/+ females during adulthood (Fig. 2D). Overall, these results indicate that Pich heterozygosity does not promote tumorigenesis in vivo nor have negative effects on the physiology of healthy mice.
Pich deficiency in heterozygosity delays the onset of tumors in Eμ-Myc mice
To investigate the effect of Pich depletion in Myc-induced lymphoma development in vivo, Pich heterozygous females were crossed with the transgenic Eμ-Myc cancer mouse model. In this tumor model, c-Myc gene expression is driven by the IgH enhancer and is expressed in B-lymphoid cells, which mimics the translocated Myc genes in B-cell lymphoma. These mice spontaneously develop B-cell lymphoma with 100% penetrance and a median latency of 16–20 weeks [33]. We evaluated the survival of Pich+/+ and Pich+/- females in the Eµ-myc transgenic background. Pich deficiency, in heterozygosity, significantly delayed the onset of Myc-induced lymphoma (Fig. 2E). While the control cohort of Pich+/+; Eµ-Myc+/tg mice had a median survival rate of 15 weeks, Pich+/-; Eµ-Myc+/tg mice had a median survival rate of 21 weeks. The median survival was increased by 50%, with some Pich+/- mice surviving up to 90 weeks of age (Fig. 2E).
PICH expression was analyzed by IHC in tumors derived from Pich+/-; Eµ-Myc+/tg mice. Remarkably, we observed a homogeneous PICH expression in contrast to the patchy pattern present in tissues from Pich+/- mice and similar to the one found in Pich+/+; Eµ-Myc+/tg BL tumors (Fig. 2F). Note that Pich is located on the X chromosome and, therefore, heterozygous females are expected to exhibit mosaic PICH expression. Based on these results, X-chromosome inactivation (XCI) seems to be biased towards the allele containing the Pich KO allele in Pich+/-; Eµ-Myc+/tg cancer cells, suggesting that PICH is necessary for the development of these tumors.
Pich deletion limits Myc-induced lymphoma progression
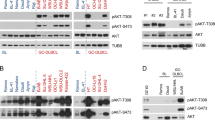
We next investigated the effect of PICH depletion in Myc-induced B-cell lymphoma progression. To this end, we generated Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T mice. UBC-Cre-ERT2 mice (Jax strain 007001) ubiquitously express an inducible Cre-ERT2 recombinase, selectively activated only in the presence of 4-hydroxytamoxifen, a metabolite of the prodrug tamoxifen [34]. In combination with the Lox-conditional allele of Pich, the UBC-Cre-ERT2 system allows efficient Pich deletion in all tissues of adult mice upon tamoxifen treatment. Pich deletion was induced at 11 weeks of age, when formation of tumors is expected to be underway. Both females and males with PichLox/Lox and PichLox genotypes, respectively, were included in the cohorts. Eleven-week-old Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T mice, along with Myc+/tg; Pich+/Lox; UBC-Cre-ERT2+/T and control animals (Myc+/tg; Pich+; UBC-Cre-ERT2+/T), were injected with tamoxifen into the peritoneum once every 24 h for a total of 5 consecutive days, plus 2 repeated injections the next 2 weeks (Fig. 3A). Activation of the UBC-Cre-ERT2 by tamoxifen treatment led to efficient recombination of the Pich locus and reduced PICH protein expression in non-tumoral tissues from Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T (Fig. 3B–D (left panel)). Survival of tamoxifen-treated Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T mice was increased compared to control mice (median survival from 21 to 35.5 weeks) (Fig. 3E). In addition, Myc+/tg; Pich+/Lox; UBC-Cre-ERT2+/T mice treated with tamoxifen also displayed intermediate increased survival. Thus, PICH seems to be required to sustain Myc-induced lymphoma, and the proliferation capacity of these tumors is diminished upon Pich deletion, which impacts the survival of the mice.
A Experimental overview to test the effect of deleting Pich in lymphoma progression. Myc+/tg; Pich+; UBC-Cre-ERT2+/T, Myc+/tg; Pich+/Lox; UBC-Cre-ERT2+/T, and Myc+/tg; PichLox; UBC-Cre-ERT2+/T or Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T mice were generated. Eleven-week-old mice were injected with tamoxifen daily for a total of 5 consecutive days, plus 2 repeated injections for the next 2 weeks to induce Pich deletion and monitored for survival. B Representation of the deletion of the Pich allele in the Pich conditional KO mouse model. Recombination of the floxed conditional allele of Pich after UBC-Cre-ERT2 activation upon tamoxifen treatment. Coding exons are indicated by filled boxes, and UTR regions are indicated by hatched boxes. LoxP sites are depicted as red triangles. C Genomic PCR analysis of PichLox allele recombination after treatment with tamoxifen. Genotyping PCR analysis from genomic DNA isolated from clip tails of tamoxifen-treated Myc+/tg; Pich+; UBC-Cre-ERT2+/T, and Myc+/tg; PichLox; UBC-Cre-ERT2+/T mice collected at humane endpoint. Migration of the expected unrecombined PichLox allele (495 bp), the wild-type Pich+ allele (440 bp) and the ablated Pich- allele (220 bp) are indicated. D PICH IHC staining of tissue sections collected from the indicated tamoxifen-treated mice. Left panel: non-tumoral spleen from healthy Myc+/tg; PichLox; UBC-Cre-ERT2+/T mice 20 weeks after tamoxifen treatment. Central panel: tumoral spleen collected from tamoxifen-treated Myc+/tg; PichLox; UBC-Cre-ERT2+/T mice at humane endpoint. Right panel: tumoral spleen collected from tamoxifen-treated Myc+/tg; Pich+; UBC-Cre-ERT2+/T mice at humane endpoint. E Kaplan–Meier survival curves for Myc+/tg; Pich+; UBC-Cre-ERT2+/T (n = 6; blue; median survival 21 weeks), Myc+/tg; Pich+/Lox; UBC-Cre-ERT2+/T (n = 13; hatched green; median survival 26.5 weeks) and Myc+/tg; PichLox; UBC-Cre-ERT2+/T or Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T (n = 20; pink; median survival 35.5 weeks) mice treated with tamoxifen at 11 weeks of age. Black dots indicate mice alive at the end of the experiment. The log-rank (Mantel–Cox) test was performed to assess differences in survival of PichLox or PichLox/Lox (p = 0.0173) and Pich+/Lox (p = 0.3475) mice compared to Pich+/+ mice, respectively. F Pich genotyping PCR analysis from genomic DNA isolated from individual tumors of Myc +/tg; Pich+; UBC-Cre-ERT2+/T, Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T mice at the humane endpoint. Each lane corresponds to a tumor from an individual mouse. The gender of each mouse is indicated in the figure. Migration of the expected unrecombined PichLox allele (495 bp), the wild-type Pich+ allele (440 bp) and the ablated Pich- allele (220 bp) are indicated.
We also analyzed PICH expression in tumors derived from the mice sacrificed at the designated humane endpoint (determined by rapid weight loss, hunched posture, rough hair coat, labored breathing, lethargy, impaired mobility or abdominal swelling) and compared it to non-tumoral spleens. Most of the Pich expression was lost in non-tumoral spleens expressing the Eµ-Myc allele that presented a normal histology upon the induction of Pich deletion with tamoxifen (Fig. 3D, left panel). In contrast, Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/T tumors exhibited a high percentage of cells with positive Pich staining (Fig. 3D, central panel). Moreover, analysis of genomic DNA isolated from these tumors revealed the presence of unrecombined PichLox alleles (Fig. 3F). Thus, Pich-positive tumor cells detected in the tumorigenic spleen by Pich staining result from the partial recombination of the PichLox allele, indicating that tumors have a preference for PICH expression to develop and hence there is a tumoral selective pressure for retaining PICH expression. Nevertheless, the recombination of the PichLox allele achieved in the mice of our study was sufficient to extend the survival of the mice. These observations reinforce the hypothesis that Pich is important for B-cell lymphoma progression. Interestingly, we identified one lymphoma with an efficient recombination of the Pich locus, suggesting that PICH is not completely essential and other factors may compensate for its absence in some cases.
Next, we investigated the mechanisms that contributed to the reduced Myc-induced tumor progression observed in Pich-depleted mice. We first evaluated DNA damage and apoptosis in tissues from tamoxifen-treated tumors at the designated humane endpoint. Of note, tumors with total or partial recombination of the Pich Lox allele were used for these analyses. A high number of apoptotic cells in tumors from Pich-depleted mice compared to tumors for mice with normal Pich expression was observed (Fig. 4A), as determined by IHC staining for cleaved caspase-3. However, the number of tumor cells positive for ϒ-H2AX, a marker of DNA damage was similar in both mice (data not shown). It has been shown that PICH deficiency results in abnormal chromosome segregation and cytokinesis, generating polyploid and binucleated cells [12]. This is supported by the observation that these cells exhibit a larger nuclear area compared to euploid cells, which has been shown to correlate with increased ploidy [35, 36]. Similarly, Pich depletion increased the percentage of tumoral cells with higher nuclear area in our mouse model, suggesting an increase of polyploid cells (Fig. 4B). Additionally, we examined the presence of DNA bridges in Pich-deficient tumors in the hematoxylin/eosin stainings. A higher percentage of cells with apparent DNA bridges was observed in tumors from tamoxifen-treated Myc+/tg; PichLox; UBC-Cre-ERT2+/T and Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/- mice compared to tumors from Pich wild-type counterparts (Fig. 4C).
A Representative immunohistochemical staining using anti-cleaved Caspase-3 antibody of paraffin-fixed tumor sections obtained at humane end point of Myc +/tg; Pich+; UBC-Cre-ERT2+/T (left) and Myc +/tg; PichLox; UBC-Cre-ERT2+/T (right) mice treated with tamoxifen. Scale bar, 50 μm. Quantification of the percentage of cleaved caspase-3 positive cells is shown on the right. Points represent the percentage of cleaved caspase-3 positive cells on each field. Data shown correspond to four to six fields per tumor, counting at least 800 cells per field. Tumors from four Pich wild-type and three Pich KO mice were assessed. Numbers indicate the mean percentage of cleaved caspase-3 positive cells. Means and SEMs are indicated. Significance was assessed by unpaired t-test ****p ≤ 0.0001. B Representative immunohistochemical staining using anti-PICH antibody of paraffin-fixed tumor sections obtained at humane endpoint of Pich+; Eµ-Myc+/tg; UBC-Cre-ERT2+/T (left) and PichLox; Eµ-Myc+/tg; UBC-Cre-ERT2+/T (right) treated with tamoxifen. Scale bar, 20 μm. Quantification of the nuclear area is shown on the right. Points represent individual nuclei. Data shown correspond to four to six fields per tumor, counting at least 400 cells per field. Tumors from three Pich wild-type and three Pich KO mice were assessed. Numbers indicate the percentage of cells with a nuclear area larger than 8 arbitrary units (a.u.). Cells with a nuclear area at least double that of the mean of cells from Myc +/tg; Pich+; UBC-Cre-ERT2+/T animals (3.7). Means and SEMs are indicated. Significance was assessed by unpaired t-test. *p ≤ 0.05. C B-cell lymphoma tumors from Pich-deficient tamoxifen-treated Myc-induced mice exhibit increased percentage of cells with DNA bridges. Quantification of the percentage of cells with DNA bridges is shown on the left. Data shown correspond to four to six fields per tumor, counting at least 400 cells per field. Tumors from three Myc +/tg; Pich+; UBC-Cre-ERT2+/T and three Myc +/tg; PichLox; UBC-Cre-ERT2+/T tamoxifen-treated mice obtained at humane endpoint were assessed. Means and SEMs are indicated. Significance was assessed by unpaired t-test. *p ≤ 0.05. Representative images of DNA bridges from Pich-deficient tumors are shown on the right. Scale bar, 5 μm.
Taken together, these results demonstrate that Pich ablation in Myc-induced mice gives rise to defects in chromosomal segregation and chromosomal instability that result in increased apoptosis and a reduction in proliferative capacity, which ultimately limits tumor growth.
Systemic depletion of PICH in adult mice is well tolerated
Tumor-selective cancer treatments are urgently needed to avoid the deleterious consequences on healthy tissues, which cause toxic side effects in patients. One of the strategies to achieve tumor selectivity is the design of drugs that target pathways essential for the survival of cancer cells, but not normal cells. In this line, we examined whether Pich was required for normal homeostasis in adult mice. In previous experiments, Pich deletion in Myc+/tg; PichLox/Lox; UBC-Cre-ERT2+/- mice was induced in all tissues and did not cause apparent systemic alterations. However, since these mice develop BL and die at a young age (median 35.5 weeks, Fig. 3E), we wanted to investigate the potential longer-term consequences of PICH deletion in adult tissues. At the age of 11 weeks, Pich+/Lox; UBC-Cre-ERT2+/- or PichLox/Lox; UBC-Cre-ERT2+/- mice were treated with tamoxifen as described above and monitored until they had noticeable tumors, reached the designated humane endpoint or up to 135 weeks. Survival of PichLox/Lox; UBC-Cre-ERT2+/- and control mice was similar (Fig. 5A). With regard to spontaneous tumor formation, none of the tamoxifen-treated mice developed spontaneous tumors before 47 weeks of age (Fig. 5B), and the percentage of mice that developed tumors was similar in both cohorts (Fig. 5B). In addition, we observed no noticeable differences in appearance among the mice with different genotypes, as reflected by their weight at 1 year of age (Fig. 5C).
A Kaplan–Meier survival curves for tamoxifen-treated Pich+/Lox; UBC-Cre-ERT2+/T (n = 8; green; median survival 98 weeks) and PichLox/Lox; UBC-Cre-ERT2+/T (n = 8; purple; median survival 78 weeks) mice. P-value = 0.3496 using the log-rank (Mantel–Cox) test. B Incidence of spontaneous tumors in Pich+/Lox; UBC-Cre-ERT2+/T mice compared to PichLox/Lox; UBC-Cre-ERT2+/T littermates treated with tamoxifen at eleven weeks of age. The differences observed in tumor incidence were not statistically significant according to the log-rank (Mantel–Cox) test. C Total body weights of adult mice at one year of age. Means and SEMs are indicated. Significance was assessed by unpaired t-test. P-value = 0.3746. D Analysis of PichLox allele recombination in tamoxifen-treated Pich conditional KO mice. PICH IHC staining of spleen tissue sections collected from 1-year-old Pich+/Lox; UBC-Cre-ERT2+/T and PichLox/Lox; UBC-Cre-ERT2+/T mice. Scale bar, 20 μm. Tissues from six mice were extracted for anatomical examination. E Pich genotyping by PCR of genomic DNA isolated from individual tissues of Pich+/Lox; UBC-Cre-ERT2+/T and PichLox/Lox; UBC-Cre-ERT2+/T mice at humane endpoint. Migration of the expected unrecombined PichLox allele (495 bp), the wild-type Pich+ allele (440 bp) and the ablated Pich- allele (220 bp) is indicated. F Representative immunohistochemical staining using anti-cleaved Caspase-3 antibody of fixed spleen sections obtained from Pich+/Lox; UBC-Cre-ERT2+/T (left) and PichLox/Lox; UBC-Cre-ERT2+/T littermates 33 weeks after tamoxifen treatment. Scale bar, 50 μm. Quantification of the percentage of cleaved caspase-3 positive cells is shown on the right. Points represent the percentage of cleaved caspase-3 positive cells on each field. Data shown correspond to four to six fields per spleen, counting at least 800 cells per field. Normal proliferative spleen for four Pich+/Lox and four PichLox/Lox healthy mice were assessed. Numbers indicate the mean percentage of cleaved caspase-3 positive cells. Means and SEMs are indicated. The differences observed were not statistically significant according to the unpaired t-test (p ≤ 0.05). G Representative immunohistochemical staining using anti-PICH antibody of fixed spleen sections obtained from Pich+/Lox; UBC-Cre-ERT2+/T and PichLox/Lox; UBC-Cre-ERT2+/T littermates 33 weeks after tamoxifen treatment. Scale bar, 20 μm. Quantification of the nuclear area is shown on the right. Points represent individual nuclei. Data shown correspond to four to six fields per spleen, counting at least 500 cells per field. Normal proliferative spleen from four Pich+/Lox and four PichLox/Lox healthy mice were assessed. Numbers indicate the percentage of cells with a nuclear area larger than 5 arbitrary units (a.u.). Cells with a nuclear area at least double that of the mean of cells from Pich+/Lox; UBC-Cre-ERT2+/T animals (2.5). Means and SEMs are indicated. The differences observed were not statistically significant according to the unpaired t-test (p ≤ 0.05).
Tissues from tamoxifen-treated mice were extracted for histological examination and to ascertain recombination efficiency of the PichLox allele, at 1, 6, and 52 weeks after tamoxifen treatment. PichLox/Lox; UBC-Cre-ERT2+/- tissues displayed efficient recombination of the PichLox allele, as assessed by IHC (Fig. 5D and Supplementary Fig. 2A). The presence of the recombined Pich KO allele in the genomic DNA was also confirmed by PCR in all tissues tested (Fig. 5E and Supplementary Fig. 2B). The histological analysis did not reveal histological differences between the organs of one-year-old tamoxifen-treated PichLox/Lox; UBC-Cre-ERT2+/- mice and those of control animals, with all exhibiting normal architecture (Fig. 5D and Supplementary Fig. 2A). In addition, Pich deletion in spleen and intestines did not lead to apoptosis, determined by IHC staining for cleaved caspase-3 (Fig. 5F and Supplementary Fig. 2C), nor changes in the number of polyploid cells (Fig. 5G and Supplementary Fig. 2D).
Overall, these findings demonstrate that systemic PICH deletion in adult mice is well tolerated and does not cause major toxic effects. Hence, PICH seems to be essential for tumor progression but not for cells of healthy adult organs, which places PICH as a promising therapeutic target.
PICH knockdown inhibits proliferation and induces cell death in human BL cells
To determine the relevance of PICH in human BL, we analyzed the consequences of losing PICH in human BL tumoral cells. Three shRNAs were transduced into Ramos and Raji BL cells to deplete PICH expression and the knockdown efficiency was assessed by Western blotting (Fig. 6A and Supplementary Fig. 3A). The effect of PICH knockdown on cell proliferation was measured over 14 days after lentiviral infection. PICH silencing strongly decreased the proliferative capacity of human BL cells (Fig. 6B). This reduction in the proliferation rates was not accompanied by major defects in DNA replication, as assessed by EdU incorporation assay (Supplementary Fig. 3B). Next, we evaluated the impact of PICH deficiency on cell viability by three different techniques, namely the MTT assay (Fig. 6C, D), trypan blue staining (Fig. 6E), and by Hoechst and To-Pro DNA fluorescence staining (Fig. 6F). Targeting PICH also led to a strong depletion of BL cell viability. Cell viability was notably reduced in the shPICH cells compared to the control cells (Fig. 6C-F). Next, apoptosis was determined using Annexin V/PI staining eight days after lentivirus infection and an increased number of apoptotic PICH-KD cells compared to cells expressing normal PICH levels were found (Fig. 6G). Thus, the effect of PICH depletion inhibiting BL cell viability was, at least partially, accounted for by the induction of apoptosis.
A Representative immunoblots of PICH protein levels in Ramos (left) and Raji (right) BL cells seven days after transduction with lentiviruses expressing the indicated shPICH or shControl (shCTRL). β-Actin was used as loading control. B Cell proliferation curve of Ramos (left) and Raji BL cells (right) infected with shCTRL or the indicated shPICH. PICH KD cells proliferated at a lower rate than shCTRL cells and stopped proliferating after four passages. Data shown correspond to a representative experiment (with technical triplicates) out of three biological replicates. The corresponding doubling times (DT) were calculated by non-linear regression and are shown in each graph. SEMs from each data point are indicated and the p-value for each dataset is shown. **p < 0.01; ****p < 0.0001. C Representative images of Ramos (top) and Raji (bottom) BL cells 10 days after infection with either shCTRL or shPICH. The images were captured at 4x magnification with a light optical microscope. Scale bar, 50 μm. Images were acquired immediately prior to the addition of MTT into the wells. D Relative cell viability of Ramos (left) and Raji (right) BL cells upon depletion of PICH compared to transfection with control shRNA (shCTRL) assessed by MTT assay 10 days after infection. Data are represented as relative absorbance at 570 nm and correspond to four independent experiments with six technical replicates for each experiment. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. ****p ≤ 0.0001. E Relative cell viability of Ramos (left) and Raji (right) BL cells upon depletion of PICH compared to transfection with control shRNA (shCTRL) determined by cell counting after labeling cells with trypan blue 10 days after infection. Histograms show the mean of the number of non-trypan blue-stained live cells of PICH depleted cells relative to shCTRL cells and correspond to three independent experiments with three technical replicates for each experiment. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. ***p ≤ 0.001; ****p ≤ 0.0001. F Cell viability of Ramos (left) and Raji (right) BL cells 10 days after transduction with lentiviruses expressing the indicated shPICH or shCTRL. Growing cells were incubated with Hoechst 33342 and To-Pro-3 dyes for 30 min. After labeling, images were acquired with a High-Content Screening System (MetaXpress) at 10× magnification. Quantification of Hoechst 33342-positive and To-Pro-3 positive cells were performed with the MetaXpress High-Content Image Analysis Software. Live cells were calculated by subtracting the number of To-Pro-3 positive cells from the Hoechst 33342-positive total cell number. Data correspond to three independent experiments with three technical replicates for each experiment. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. ****p ≤ 0.0001. G Effect of PICH KD on the apoptosis of Ramos and Raji BL cancer cells measured by Annexin V-FITC/Propidium iodide and flow cytometry 7 days after infection with either shCTRL or shPICH. The percentage of apoptotic (Annexin V-positive) cells is indicated. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. *p ≤ 0.05; **p ≤ 0.01.
PICH depletion causes chromosomal instability in human BL cells
To characterize the cause of cell lethality in PICH-depleted BL cells, we analyzed defects in cell division. Abnormal chromosome segregation usually leads to cytokinesis problems and abnormal cell division that is manifested as G1-associated aberrations, such us micronuclei and bi- or poly-nucleation. PICH knockdown in human cells has been shown to increase the occurrence of chromosomal abnormalities [12]. We examined whether silencing PICH in BL cells showed an alteration in the frequency of chromosome abnormalities. Of note, proliferative BL Ramos and Raji cell lines PICH positive persistent bridges (Supplementary Fig. 3C). In agreement with the results observed in PICH-deficient Eµ-Myc tumors, PICH-depleted human BL cells exhibited an increased frequency of persistent DNA bridges revealed by Hoechst 33342 staining of nuclei (Fig. 7A). PICH-deficient BL cells also had a significantly elevated frequency of binucleation (cells with two decondensed daughter nuclei in the same plasma membrane), polynucleation (Fig. 7B), and micronucleus formation, revealed by DAPI staining of nuclei and immunofluorescence staining for α-tubulin (Fig. 7C). Altogether, these data suggest that PICH deficiency results in defective cell division in BL cells, thus suppressing cell proliferation and inducing cell death.
A Frequency of DNA bridges in asynchronously Ramos and Raji BL growing cells was determined by Hoechst 33342 nuclear staining (white) 7 days after PICH depletion with the indicated shRNAs. After staining, samples were examined with High-Content Screening System MetaXpress at ×40 magnification. Data shown correspond to biological triplicates of at least 300 cells per condition. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. *p ≤ 0.05; **p ≤ 0.01. Representative images are shown. Scale bar, 5 μm. B Frequency of bi- and multi-nucleation in asynchronously growing Ramos BL cells was determined by immunofluorescence staining of α-tubulin (green) combined with DAPI nuclear staining (white) 7 days upon PICH depletion with shRNAs. After immunostaining, samples were examined with High-Content Screening System MetaXpress at 40× magnification. Data shown correspond to biological triplicates of at least 500 cells per condition. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. ***p ≤ 0.001 ****p ≤ 0.0001. Representative images are shown. Scale bar, 5 μm. C Frequency of micronucleus formation in cells stained with DAPI and obtained as described in Fig. 7B. Mean and SEMs are indicated. Significance was assessed by unpaired t-test. *p ≤ 0.05. An example of a scored micronucleus is shown. Red arrow indicates a micronucleus in the representative image. Scale bar, 5 μm.
Discussion
PICH is a DNA translocase that plays a role in the resolution of UFBs and is necessary for the maintenance of chromosomal stability in proliferating cells. Previously, we found that PICH is essential for embryonic development and several groups showed that PICH is highly expressed in several cancerous cells and is relevant for their proliferation in vitro [18,19,20,21,22,23,24,25]. Moreover, a recent study has shown that PICH deficiency limits the development of mammary tumors in mice [37]. Given that PICH expression is highest in BL among all cancer types, we have used a mouse model of Myc-induced B-cell lymphoma (Eμ-Myc), in combination with the Pich conditional KO model. We have found that PICH depletion limits the progression of MYC-induced lymphomas and, critically, does not have major toxic effects in non-tumoral tissues.
It is important to stress the contrast between the consequences of PICH depletion during embryonic development, which is incompatible with life, and PICH depletion in adult mice, which is very well tolerated. This is particularly relevant when considering PICH as a potential therapeutic target. Many proteins are required for the proliferation of cancerous cells, but few of them are highly specific for cancerous compared to normal tissues. Thus, we believe that PICH inhibitors may be useful for cancer therapy, and we are currently working on the identification and development of small molecules with PICH inhibitory capacity.
We have also investigated the long-term effect of PICH depletion in adult mice without additional alterations because PICH depletion could potentially lead to chromosomal instability (CIN) and increased cancer risk. However, we did not observe an increased incidence of any cancer type. Indeed, whether CIN itself may lead to cancer or not is still a matter of debate, and may vary between different cellular and genetic contexts [38].
Previous molecular analyses performed by us and other groups [7, 11, 14], and our observations of polyploid cells and chromosomal abnormalities in PICH-deficient cancer cells, suggest that the molecular mechanism by which PICH favors the proliferation of cancer cells is the resolution of UFBs and maintenance of chromosomal stability. However, other studies have reported that PICH also plays an additional role stabilizing the replication fork [39] and regulates the PI3K/AKT and NK- κB pathways [22] that may also contribute to cancer progression. Of note, increased levels of replication stress, such as those caused by MYC [40] overexpression, could lead to the formation of UFBs. It is well characterized that PICH binds to UFBs, where it recruits and modulates the activity of topoisomerase 2 and the Topoisomerase 3a–Rmi1–Rmi2 complex for the resolution of UFBs [12, 41]. Moreover, it has been shown that PICH ATPase activity is necessary for its DNA translocase activity and for UFB resolution [12]. Therefore, compounds blocking PICH ATPase activity are predicted to have antitumoral properties.
In the current study, we have specifically studied the role of PICH in BL, an aggressive non-Hodgkin B-cell lymphoma caused by translocations that lead to C-MYC overexpression. There are sporadic cases and endemic cases associated with HIV, Epstein-Barr Virus, and malaria. While most children respond well to standard chemotherapy (including cyclophosphamide, vincristine, prednisolone, and doxorubicin), many adults undergo a relapse of the disease that is associated with a poor prognosis. Moreover, the side effects associated with chemotherapy pose an important risk of infections and secondary pathologies, especially in developing countries. Thus, BL patients could benefit from a more specific and less toxic therapy. Although we have used BL as a cancer model, we believe that a therapy based on PICH inhibition could be useful for other cancer types with high PICH expression. However, higher PICH expression could just reflect changes in the cell cycle or the proliferation rate of some cancer types. Thus, the role of PICH should be investigated with experimental studies in each case, as was recently done for breast cancer [37]. Data from the DepMap portal indicate that PICH is essential in only 164 out of 1095 cancer cell lines (DepMap Public 23Q2+Score, Chronos). Contrary to our in vivo and in vitro data, BL cell lines are not significantly affected by PICH depletion according to DepMap portal data. One possible explanation for this discrepancy is that PICH was not efficiently knocked out in these screens since this was not verified by western blotting or other means.
It would be relevant to investigate mutations or other conditions that increase sensitivity to PICH deficiency, as well as mutations that could confer resistance to PICH inhibitors. In this regard, MYC amplification and MYC overexpression do not drive ERCC6L dependence in a pan-cancer analysis of data from the DepMap portal (DepMap Public 23Q2+Score, Chronos). These data reinforce the idea that PICH inhibition could be useful for other non-MYC-driven cancer types. Interestingly, a recent study has shown that PICH becomes more important for cell viability in cells deficient for DNA maintenance genes, such as those related to DNA repair, DNA condensation, centromere stability, and regulators of the replication fork stability such as FIRRM [42]. In addition, a recent study has found that PICH deficiency is synthetic lethal with SUMOylation inhibitors [43]. In this line, treatments combining PICH inhibitors and compounds targetting these DNA repair pathways could potentially offer higher efficacy.
In summary, the cumulative evidence in the literature of the relevance of PICH for the proliferation of different cancer cells, together with our results presented here using a Pich conditional KO model, indicate that PICH is a promising therapeutic target to treat BL and possibly other cancers with high PICH expression.
Data availability
The data that support the findings of this study are available on request to the corresponding author. andres.lopez@cabimer.es.
References
IARC. International Agency for Research on Cancer [Internet]. Globocan. 2018. Available at: https://www.iccp-portal.org/news/globocan-2018.
Molyneux EM, Rochford R, Griffin B, Newton R, Jackson G, Menon G, et al. Burkitt’s lymphoma. Lancet. 2012;379:1234–44.
Jacobson C, La Casce A. How I treat Burkitt lymphoma in adults. Blood. 2014;124:2913–20.
Mcclelland SE, Mcclelland SE. Role of chromosomal instability in cancer progression. Endocr Relat Cancer. 2017;24:T23–31.
Baumann C, Körner R, Hofmann K, Nigg EA. PICH, a centromere-associated SNF2 family ATPase, is regulated by plk1 and required for the spindle checkpoint. Cell. 2007;128:101–14.
Chan KL, North PS, Hickson ID. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. EMBO J. 2007;26:3397–409.
Biebricher A, Hirano S, Enzlin JH, Wiechens N, Streicher WW, Huttner D, et al. PICH: a DNA translocase specially adapted for processing anaphase bridge DNA. Mol Cell. 2013;51:691–701.
Hengeveld RCCCC, de Boer HR, Schoonen PMM, de Vries EGE, Lens SMAMA, van Vugt MATM, et al. Rif1 Is Required for Resolution of Ultrafine DNA Bridges in Anaphase to Ensure Genomic Stability. Dev Cell. 2015;34:466–74.
Bizard AH, Hickson ID. Anaphase: a fortune-teller of genomic instability. Curr Opin Cell Biol. 2018;52:112–9.
Liu Y, Nielsen CF, Yao Q, Hickson ID. The origins and processing of ultra fine anaphase DNA bridges. Curr Opin Genet Dev. 2014;26:1–5.
Nielsen CF, Hickson ID. PICH promotes mitotic chromosome segregation: identification of a novel role in rDNA disjunction. Cell Cycle. 2016;15:2704.
Nielsen CF, Huttner D, Bizard AH, Hirano S, Li T-N, Palmai-Pallag T, et al. PICH promotes sister chromatid disjunction and co-operates with topoisomerase II in mitosis. Nat Commun. 2015;6:8962.
Rouzeau S, Cordelières FP, Buhagiar-Labarchède G, Hurbain I, Onclercq-Delic R, Gemble S, et al. Bloom’s syndrome and PICH helicases cooperate with topoisomerase IIα in centromere disjunction before anaphase. PLoS ONE. 2012;7:e33905.
Albers E, Sbroggiò M, Pladevall-Morera D, Bizard AH, Avram A, Gonzalez P, et al. Loss of PICH results in chromosomal instability, p53 activation, and embryonic lethality. Cell Rep. 2018;24:3274–84.
Geng X, Zhang C, Li M, Wang J, Ji F, Feng H. et al. PICH supports embryonic hematopoiesis by suppressing a cGAS-STING-mediated interferon response. Adv Sci. 2022;9:e2103837.
Giam M, Rancati G. Aneuploidy and chromosomal instability in cancer: a jackpot to chaos. Cell Div. 2015;10:1–12.
Bach DH, Zhang W, Sood AK. Chromosomal instability in tumor initiation and development. Cancer Res. 2019;79:3995–4002.
Pu S-Y, Yu Q, Wu H, Jiang J-J, Chen X-Q, He Y-H, et al. ERCC6L, a DNA helicase, is involved in cell proliferation and associated with survival and progress in breast and kidney cancers. Oncotarget. 2017;8:42116–24.
Huang Y, Li W, Yan W, Wu J, Chen L, Yao X, et al. Loss of PICH promotes chromosome instability and cell death in triple-negative breast cancer. Cell Death Dis. 2019;10:428.
Xie Y, Yu J, Wang F, Li M, Qiu X, Liu Y, et al. ERCC6L promotes cell growth and invasion in human colorectal cancer. Oncol Lett. 2019;18:237–46.
Zhang G, Yu Z, Fu S, Lv C, Dong Q, Fu C, et al. ERCC6L that is up-regulated in high grade of renal cell carcinoma enhances cell viability in vitro and promotes tumor growth in vivo potentially through modulating MAPK signalling pathway. Cancer Gene Ther. 2019;26:323–33.
Chen H, Wang H, Yu X, Zhou S, Zhang Y, Wang Z, et al. ERCC6L promotes the progression of hepatocellular carcinoma through activating PI3K/AKT and NF-κB signaling pathway. BMC Cancer. 2020;20:853.
Chen D, Liu Q, Cao G. ERCC6L promotes cell growth and metastasis in gastric cancer through activating NF-κB signaling. Aging. 2021;13:20218–28.
Hou G, Lu Z, Bi Y, Deng J, Yang X. ERCC6L is a biomarker and therapeutic target for non-small cell lung adenocarcinoma. Med Oncol. 2022;39:51.
Zhang G, Ma J, Xiong J, Huang X, Han X, Yu X, et al. Upregulation of excision repair cross-complementation group 6-like (ERCC6L) promotes tumor growth in hepatocellular carcinoma. Dig Dis Sci. 2021;66:1097–109.
Zhou H, Yang Z, Yue J, Chen Y, Chen T, Mu T, et al. Identification of potential hub genes via bioinformatics analysis combined with experimental verification in colorectal cancer. Mol Carcinog. 2020;59:425–38.
Yu B, Liang H, Ye Q, Wang Y. Upregulation of ERCC6L is associated with tumor progression and unfavorable prognosis in hepatocellular carcinoma. J Gastrointest Oncol. 2020;11:1009–23.
Xu N, Chen SH, Lin TT, Cai H, Ke ZB, Dong RN, et al. Development and validation of hub genes for lymph node metastasis in patients with prostate cancer. J Cell Mol Med. 2020;24:4402–14.
Yin X, Wang J, Zhang J. Identification of biomarkers of chromophobe renal cell carcinoma by weighted gene co-expression network analysis. Cancer Cell Int. 2018;18:206.
Ma MCJ, Tadros S, Bouska A, Heavican T, Yang H, Deng Q, et al. Subtype-specific and co-occurring genetic alterations in B-cell non-Hodgkin lymphoma. Haematologica. 2022;107:690–701.
Adams JM, Harris AW, Pinkert CA, Corcoran LM, Alexander WS, Cory S, et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature. 1985;318:533–8.
Pladevall-Morera D, Castejón-Griñán M, Aguilera P, Gaardahl K, Ingham A, Brosnan-Cashman JA, et al. ATRX-deficient high-grade glioma cells exhibit increased sensitivity to RTK and PDGFR inhibitors. Cancers. 2022;14:1790.
Harris AW, Pinkert CA, Crawford M, Langdon WY, Brinster RL, Adams JM, et al. The E mu-myc transgenic mouse. A model for high-incidence spontaneous lymphoma and leukemia of early B cells. J Exp Med. 1988;167:353–71.
Ruzankina Y, Pinzon-Guzman C, Asare A, Ong T, Pontano L, Cotsarelis G, et al. Deletion of the developmentally essential gene ATR in adult mice leads to age-related phenotypes and stem cell loss. Cell Stem Cell. 2007;1:113–26.
de Cárcer G, Venkateswaran SV, Salgueiro L, El Bakkali A, Somogyi K, Rowald K, et al. Plk1 overexpression induces chromosomal instability and suppresses tumor development. Nat Commun. 2018;9:1–14. 91. 2018
Matsuura T, Ueda Y, Harada Y, Hayashi K, Horisaka K, Yano Y, et al. Histological diagnosis of polyploidy discriminates an aggressive subset of hepatocellular carcinomas with poor prognosis. Br J Cancer. 2023;15:1–10.
Yang H, Zhen X, Yang Y, Zhang Y, Zhang S, Hao Y, et al. ERCC6L facilitates the onset of mammary neoplasia and promotes the high malignance of breast cancer by accelerating the cell cycle. J Exp Clin Cancer Res. 2023;42:227.
Bakhoum SF, Cantley LC. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell. 2018;174:1347–60.
Tian T, Bu M, Chen X, Ding L, Yang Y, Han J, et al. The ZATT-TOP2A-PICH axis drives extensive replication fork reversal to promote genome stability. Mol Cell. 2021;81:198–211.e6.
Curti L, Campaner S. MYC-induced replicative stress: a double-edged sword for cancer development and treatment. Int J Mol Sci. 2021;22:6168.
Bizard AH, Allemand JF, Hassenkam T, Paramasivam M, Sarlós K, Singh MI, et al. PICH and TOP3A cooperate to induce positive DNA supercoiling. Nat Struct Mol Biol. 2019;26:267–74.
Stok C, Tsaridou S, van den Tempel N, Everts M, Wierenga E, Bakker FJ, et al. FIRRM/C1orf112 is synthetic lethal with PICH and mediates RAD51 dynamics. Cell Rep. 2023;42:112668.
Hertz EPT, Vega IA, Kruse T, Wang Y, Hendriks IA, Bizard AH, et al. The SUMO-NIP45 pathway processes toxic DNA catenanes to prevent mitotic failure. Nat Struct Mol Biol. 2023;30:1303–13.
Acknowledgements
This work was supported by grants from Danish National Research Foundation (DNRF115), European Research Council (ERC-2015-STG-679068), the Spanish Ministry of Science and Innovation-Agencia Estatal de Investigacion (PID2020-119329RB-I00), the Novo Nordisk Foundation (NNF19OC0055203), and the Junta de Andalucía (P20_00755, PAIDI2020). M.C.-G. was supported with a postdoctoral fellowship from the Junta de Andalucía and the European Social Fund (202099908313734) and with a postdoctoral “Juan de la Cierva” fellowship from the MICINN (FJC2020-045915-I). D.P.-M. was supported with a PhD scholarship from the Lundbeck Foundation (R218-2016-415). L.S.-C. was supported with a Postdoctoral Research Contract from the AECC Scientific Foundation (Grant Number POSTD211274SIMÓ). P.A. was supported with a postdoctoral fellowship from the Junta de Andalucía and the European Social Fund (202099908313486). A.G. is funded with a PhD fellowship from the Junta de Andalucía. E.P. is supported by the HORIZON-MSCA ITN RepliFate (101072903). We thank Prof. Fernandez-Capetillo for kindly providing human lymphoma cell lines. We thank Patricia González from the CNIO Histopathology Core Unit, Irene Delgado, and Alexandra Avram for technical support.
Author information
Authors and Affiliations
Contributions
AJL-C conceived, designed, and supervised the study. MC-G designed, performed, and analyzed most of the experiments. EA performed Pich heterozygous mouse-related work. DP-M helped with the tamoxifen injections. EL, AI, LS-C, PA, AG-B, and EP provided technical help for the mouse-related experiments. LS-C, IDH, and ML provided critical input and advice during the development of the project. AJL-C and MC-G wrote the manuscript. All of the authors read, edited, and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Castejón-Griñán, M., Albers, E., Simón-Carrasco, L. et al. PICH deficiency limits the progression of MYC-induced B-cell lymphoma. Blood Cancer J. 14, 16 (2024). https://doi.org/10.1038/s41408-024-00979-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41408-024-00979-y