Abstract
The acute promyelocytic leukemia (APL) driver ZBTB16/RARα is generated by the t(11;17) (q23;q21) chromosomal translocation, which is resistant to combined treatment of all-trans retinoic acid (ATRA) and arsenic trioxide (ATO) or conventional chemotherapy, resulting in extremely low survival rates. In the current study, we investigated the effects of hyperthermia on the oncogenic fusion ZBTB16/RARα protein to explore a potential therapeutic approach for this variant APL. We showed that Z/R fusion protein expressed in HeLa cells was resistant to ATO, ATRA, and conventional chemotherapeutic agents. However, mild hyperthermia (42 °C) rapidly destabilized the ZBTB16/RARα fusion protein expressed in HeLa, 293T, and OCI-AML3 cells, followed by robust ubiquitination and proteasomal degradation. In contrast, hyperthermia did not affect the normal (i.e., unfused) ZBTB16 and RARα proteins, suggesting a specific thermal sensitivity of the ZBTB16/RARα fusion protein. Importantly, we found that the destabilization of ZBTB16/RARα was the initial step for oncogenic fusion protein degradation by hyperthermia, which could be blocked by deletion of nuclear receptor corepressor (NCoR) binding sites or knockdown of NCoRs. Furthermore, SIAH2 was identified as the E3 ligase participating in hyperthermia-induced ubiquitination of ZBTB16/RARα. In short, these results demonstrate that hyperthermia could effectively destabilize and subsequently degrade the ZBTB16/RARα fusion protein in an NCoR-dependent manner, suggesting a thermal-based therapeutic strategy that may improve the outcome in refractory ZBTB16/RARα-driven APL patients in the clinic.
Similar content being viewed by others
Introduction
Hematological malignancies are mainly due to perturbed functions of transcription factors, which arise from chromosomal translocations, loss- or gain-of-function mutations, or deregulated expression [1,2,3]. According to the French–American–British classification system, acute promyelocytic leukemia (APL) belongs to the M3 subtype of acute myeloid leukemia (AML) [4]. Of note, APL has become a highly curable disease due to the introduction of all-trans retinoic acid (ATRA) and arsenic trioxide (ATO) into frontline treatment, with the five-year survival rate reaching up to ~95% [4, 5]. Generally, APL is caused by the t(15;17) (q24; q21) chromosomal translocation, which results in the expression of the PML/RARα fusion gene and protein, eventually leading to the overproduction of immature promyelocytes in the bone marrow [5, 6]. However, ~2% of APL cases are attributed to the occurrence of the ZBTB16 (zinc finger and BTB domain containing 16, previously named PLZF)-RARα fusion protein, which is generated from the t(11;17)(q23;q21) translocation [7]. ZBTB16/RARα fusion proteins, as dominant-negative forms of either RARα or ZBTB16, display leukemogenic activity by disrupting the normal functions of ZBTB16 and RARα, such as RAR/RXR-mediated signaling, DNA binding activity, hematopoiesis, and p53 protein destabilization [8,9,10,11].
ATO has been adopted as the first-line drug for the treatment of classical APL, which can directly target the PML part of the PML/RARα fusion protein, induce protein destabilization (i.e., transfer from RIPA soluble supernatant into the insoluble pellet fraction), and enhance SUMOylation and ubiquitination of the fusion protein, ultimately resulting in degradation of the fusion protein through the proteasome pathway [12]. However, ATRA is able to dissociate nuclear receptor corepressors (NCoR1/SMRT) from the PML/RARα and histone deacetylase complexes to reactivate the transcription of repressed genes, leading to terminal differentiation of immature leukemic promyelocytes into normal mature granulocytes [13]. However, ZBTB16/RARα-driven APL patients showed intrinsic resistance to standard treatment of ATRA with ATO or chemotherapy [9, 11, 14]. Mechanistically, ATO is unable to induce ZBTB16/RARα fusion protein destabilization, degradation, or APL cell differentiation in vitro, which might be due to the absence of PML [12, 13, 15]. Notably, the NCoR (including NCoR1 and SMRT) complex binds to only the ligand binding domain (LBD) in the RARα part of the PML/RARα fusion protein, while it can bind to both the POZ and LBD domains in the ZBTB16 and RARα parts of the ZBTB16/RARα fusion protein, respectively [16]. Although ATRA can dissociate NCoR1/SMRT corepressors from the RARα part through competitive binding with the LBD domain, it is ineffective in dissociating the corepressors from the POZ domain within the ZBTB16 part [9, 17], which could explain why ZBTB16/RARα-associated APL is resistant to ATRA. In addition, ZBTB16/RARα-driven APL showed a limited response to chemotherapeutic drugs for AML, such as cytarabine and anthracycline or idarubicin [18]. A higher incidence of refractory/relapsed cases (30%) and relapse rate of 45%–80% were reported in ZBTB16/RARα patients [19, 20]. Therefore, the development of new treatment approaches or drugs for ZBTB16/RARα-driven APL patients is urgently needed.
Oncogenic fusion proteins are well-defined drivers in many leukemias, thus, targeting and destroying oncogenic fusion proteins are considered to be a promising strategy for hematological malignancies [9, 16, 21, 22]. For instance, ATO cures APL mainly by targeting and destabilizing the PML/RARα fusion protein. Since there are also emerging cases of ATO resistance, novel therapeutic strategies are in progress for these cases [23, 24]. Recently, we reported hyperthermia as a novel strategy that exerts potential therapeutic effects against PML/RARα (including ATO-resistant mutants)-driven APL [25]. Hyperthermia could selectively destabilize wild-type and drug-resistant mutants of the PML/RARα fusion protein, in which NCoRs are essential for PML/RARα aggregation, nuclear matrix targeting, and subsequent degradation [25, 26]. Moreover, clinical data further proved that hyperthermia has superior effects on refractory central nervous system-relapsed APL patients [25]. Similar to the PML/RARα fusion protein, ZBTB16/RARα also has a strong binding affinity to NCoRs; therefore, we assumed that hyperthermia may influence the stability of the ZBTB16/RARα protein and eventually induce its degradation.
To prove the above notion, we attempted to determine the effects of hyperthermia on oncogenic ZBTB16/RARα fusion protein stability. We found that the ZBTB16/RARα oncogenic fusion protein can be destabilized by hyperthermia at 42 °C, and this process is irreversible. Subsequently, ZBTB16/RARα was robustly ubiquitinated and further degraded by the proteasome. However, no appreciable effects were observed on either unfused ZBTB16 or RARα protein following hyperthermia. Moreover, we found that NCoRs play an important role in the destabilization of the ZBTB16/RARα fusion protein by hyperthermia. Taken together, our results provide a prospective hyperthermia-based therapeutic strategy for refractory ZBTB16/RARα-driven APL.
Materials and methods
Reagents
All reagents used were of chemical grade. Sodium arsenite (iAsIII), ATRA, MG132 and cycloheximide (CHX) were purchased from Sigma, St. Louis, MO, USA. Stock solutions of arsenic (10 mmol/L), ATRA (1 mmol/L), CHX (20 mmol/L), and MG132 (10 mmol/L) were prepared and stored at −20 °C. Dilutions were made according to standard protocols on a daily basis prior to use.
Cell culture
HeLa and 293T cells were purchased from the Cell Bank of the Chinese Academy of Sciences. OCI-AML3 cell lines were purchased from Zhejiang Meisen Cell Technology Co., Ltd. All cell lines were authenticated using DNA fingerprinting to assess cross-contamination during the study and tested for Mycoplasma contamination once a year. HeLa and 293T cells were cultured and maintained in DMEM (Gibco, 12800-082), while OCI-AML3 cell lines were cultured and maintained in RPMI-1640 medium (Gibco, 12800-017). All the culture media were supplemented with 10% fetal bovine serum (Gibco, 10270-106), 100 U/mL penicillin, and 100 µg/mL streptomycin. Cells were kept at 37 °C in a 5% CO2 atmosphere.
Plasmid construction and protein expression
The ZBTB16 protein and ZBTB16/RARα fusion gene were synthesized and inserted into the PCMV-Tag2B expression vector by Tsingke Biotechnology Co., Ltd., China. PCMV-Tag2B-PML, PCMV-Tag2B-PML/RARα, and PCMV-HA-Ub were constructed previously [25]. PCMV-Tag2B-ZBTB16/RARα-based functional domain deletion mutants, including ΔDBD, ΔLBD, ΔPOZ, Δ(POZ + LBD) and all point mutations, were generated by homologous recombination using a ClonExpress II One Step Cloning Kit (C112-02, Vanzyme, China) according to the manufacturer’s instructions. The mutants were confirmed by sequencing prior to transfection experiments. The primer sequences for Z/R mutants are listed in Supplementary Table S1. Transfection was carried out using Lipofectamine 3000 (L3000-008, Invitrogen, Carlsbad, USA) according to the manufacturer’s instructions. After transfection for 24–36 h, the cells were treated as indicated and then subjected to various analyses.
Gene silencing
Gene silencing was conducted by transfection of siRNAs using Lipofectamine 3000 (L3000-008, Invitrogen) according to the manufacturer’s instructions. Thirty-six hours after transfection, the cells were treated as indicated and subjected to various analyses. The target sequences for SMRT, NCoR1 and SIAH2 siRNAs are as follows: NCoR1#1, 5'-CTATCAGCCAGAGGTTGTTAA-3'; NCoR1#2, 5'-GCCATCAAACACAATGTCAAA-3'; SMRT#1, 5'-GCAGCGCATCAAGTTCATCAA-3'; SMRT#2, 5'-GCAGTGTAAGAACTTCTACTT-3'; SIAH2#1, 5'-CCATGATGTGACTTTCGTAAA-3'; SIAH2#2, 5'-ACACAGCCATAGCACATCTTT-3'.
Antibodies
The following primary antibodies were used for immunoblotting, immunoprecipitation, and immunofluorescence: anti-Flag (F1804, Sigma), anti-PRAP (sc-98819, Santa Cruz Biotechnology, CA), anti-NCoR1 (5948, Cell Signaling Technology [CST], Danvers, MA, USA), anti-SMRT (62370, CST), anti-SIAH2 (A17423, Abclonal, Wuhan, China), anti-HSF1 (AF1570, Beyotime, Shanghai, China), anti-HA (AF0039, Beyotime), anti-Ub (AF1705, Beyotime), anti-β-actin (AC026, Abclonal), and anti-GAPDH (60004, Proteintech, Chicago, IL). The HRP-tagged secondary antibodies used for immunoblotting were as follows: anti-mouse (A0216, Beyotime) and anti-rabbit (A0208, Beyotime). DyLight 488 and the anti-mouse secondary antibody used for immunofluorescence were purchased from Abbkine, Wuhan, China (A23210).
Protein extraction and Western blot analysis
Protein extractions for different research purposes were carried out according to the methods described previously [25]. (1) For protein stability analysis, the cells were lysed in RIPA lysis buffer (50 mmol/L Tris, 150 mmol/L NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, pH 7.5, 0.2 mmol/L PMSF, and a complete mini protease inhibitor tablet). Samples were incubated on ice for 15 min with vortexing at 5-min intervals and centrifuged for 30 min at 4 °C and 13,000 rpm to obtain the soluble fraction (supernatant, S) and insoluble fraction (pellet, P). The insoluble fraction (pellet, P) was further washed twice with PBS and lysed in SDS lysis buffer (1×TBS, 10% glycerol, 0.015% EDTA, 50 mmol/L DTT, and 2% SDS). Both S and P fractions were prepared for Western blot analysis. (2) For total protein extraction, cells were lysed in RIPA buffer with 8 M urea to solubilize detergent-insoluble pellets and incubated on ice for 15 min with vortexing at 5 min intervals. Cell lysate was centrifuged at 4 °C and 13,000 rpm for 30 min, and then, the supernatant was obtained for Western blot analysis. (3) For immunoprecipitation (IP), cells were suspended in IP buffer (50 mmol/L Tris-HCl pH 7.5, 10% glycerol, 150 mmol/L NaCl, 2 mmol/L EDTA, 0.5% NP-40, and 1 mmol/L PMSF plus protease inhibitors). Sonication was used to break cells apart at 15% amplitude for 70 pulses, where each pulse consisted of 2 s of sonication followed by a 3-s rest period. After centrifugation, the supernatant was used for IP analysis. Protein concentrations were measured using the BCA Protein Quantification Kit (20201ES76, Yeasen Biotech, Shanghai, China). Each protein sample (25 μg) was boiled at 95 °C for 10 min, resolved by SDS‒PAGE, and blotted onto PVDF membranes. The membranes were blocked with fat-free milk and incubated overnight with different primary antibodies at 4 °C, followed by incubation with HRP-labeled secondary antibodies for 1 h at room temperature. Then, the protein bands were visualized by enhanced chemiluminescence (E412-01, Vanzyme).
Immunoprecipitation assay
Immunoprecipitation was performed according to an established protocol as previously published [25]. Cells were seeded in 10-cm culture dishes and transfected with 12 μg of each indicated plasmid. Twenty-four to thirty-six hours after transfection, the cells were treated as indicated, scraped down, and collected in ice-cold PBS. Then, the cells were lysed on ice by sonication in IP buffer (50 mmol/L Tris-HCl pH 7.5, 10% glycerol, 150 mmol/L NaCl, 2 mmol/L EDTA, 0.5% NP-40, and 1 mmol/L PMSF plus protease inhibitors). Immunoprecipitation experiments were performed using primary antibodies and protein A/G PLUS-Agarose (sc-2003, Santa Cruz Biotechnology, CA) according to the manufacturer’s instructions.
Immunofluorescence microscopy
HeLa cells were grown in culture plates or wells containing glass disks following transfection and recommended treatments. The glass disks were transferred onto slides and washed twice with PBS. After fixation with 4% paraformaldehyde, the cells were permeabilized with 0.1% Triton X-100. Slides were blocked with 2% BSA in PBS, followed by incubation with primary antibodies overnight at 4 °C. The next day, slides were washed thrice with PBS and incubated with fluorescent-labeled secondary antibodies at room temperature for 4 h and washed thrice with PBS. Slides were mounted using DAPI Fluoromount-G (0100-20, SouthernBiotech) and stored in the dark at 4 °C. The fluorescent signals were visualized under a Zeiss (Göttingen, Germany) 510 confocal microscope.
Fluorescence recovery after photobleaching (FRAP)
HeLa cells overexpressing GFP-ZBTB16/RARα were subjected to hyperthermia at 42 °C for 1 h before the assay. FRAP assays were carried out on a Nikon A1R confocal microscope in a live-cell imaging chamber. Droplets were bleached with a 488-nm laser pulse (60% intensity, 1 s). Recovery was recorded every 5 s for a total of 180 s after bleaching. Analysis of the recovery curves was carried out with FIJI/ImageJ software.
Results
The normal ZBTB16 protein contains three major functional domains, while the RARα protein consists of six modular domains, as shown in Fig. 1a. Owing to chromosomal translocation, the N-terminus of ZBTB16 (amino acids 1-455) is fused with the C-terminus of RARα (amino acids 62–462) to generate the ZBTB16/RARα (Z/R) fusion protein (Fig. 1a). We transiently transfected HeLa cells with normal ZBTB16 and Z/R fusion plasmids and found that ZBTB16 and Z/R fusion proteins were completely diffused in the nucleus (Fig. S1a, b), which were different from the morphology of PML and PML/RARα proteins (Fig. S1c, d).
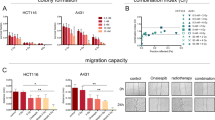
Schematic representation of the ZBTB16, RARα, and oncogenic ZBTB16/RARα (Z/R) fusion proteins with functional domains and breakpoints (a). PML/RARα- and Z/R-transfected HeLa cells were exposed to ATO (0.5, 1 µM) for 6 and 12 h (b and c) or ATRA (1, 2 µM) for 6 and 12 h (d and e). Protein expression in soluble (S) and pellet (P) fractions of RIPA lysate was analyzed by Western blotting with anti-Flag and anti-β-actin monoclonal antibodies.
ATO directly binds to cysteine residues in the zinc finger motifs of the PML portion, resulting in destabilization of PML/RARα proteins (e.g., a soluble protein shift to an insoluble protein) (Fig. 1b) and subsequent degradation by the proteasome [12]. Given that the Z/R fusion protein also contains cysteine-rich zinc finger motifs (Fig. 1a), we investigated whether ATO could destabilize the Z/R fusion protein. The results showed that ATO had no effect on the stability and degradation of the Z/R protein (Fig. 1c), which is consistent with previous reports [13]. Likewise, ATRA effectively degraded the PML/RARα protein (Fig. 1d), but it had a moderate effect on the Z/R protein (Fig. 1e), confirming that ATO and ATRA are less likely to degrade the Z/R fusion protein, which might be the main reason for clinical failure. Furthermore, chemotherapeutic agents such as cytarabine, azacitidine, venetoclax, and vincristine failed to affect Z/R protein stability (Fig. S2a–d). These findings demonstrate, at least in part, the resistance of Z/R-driven APL to chemotherapy.
Recently, we reported that hyperthermia could effectively induce destabilization and degradation of the PML/RARα oncogenic fusion protein as well as ATO- and ATRA-resistant mutants in an NCoR-dependent manner [25]. Since Z/R is also an NCoR-interacting fusion protein, we hypothesize that hyperthermia might have a comparable effect on the Z/R fusion protein. To test this hypothesis, we subjected HeLa, 293T and OCI-AML3 cells that exogenously expressed the Z/R fusion protein to hyperthermia. The results showed that the Z/R fusion protein was robustly destabilized (i.e., shifted from the RIPA soluble supernatant to the insoluble pellet) by mild hyperthermia in a time- and/or temperature-dependent manner (Fig. 2a–c and Fig. S3a–c). As a positive control for heat stress, phosphorylated heat shock transcription factor 1 (pHSF1) was also significantly increased under the same conditions (Fig. 2a–c and Fig. S3-c). Notably, hyperthermia could induce the transfer of Z/R fusion protein from RIPA soluble into insoluble fraction in the short term, while the total Z/R fusion protein level remained unchanged at the same time, indicating that hyperthermia rapidly destabilizes but does not degrade Z/R protein (Fig. S3d, e). Interestingly, hyperthermia had a negligible influence on normal ZBTB16 and RARα proteins (Fig. S4a, b), suggesting that hyperthermia selectively targets oncogenic proteins rather than normal proteins.
Exogenous Z/R fusion protein-expressing HeLa cells were exposed to hyperthermia in a temperature-dependent (a) and time-dependent manner (b and c). The changes in the Z/R fusion protein in the supernatant (S) and pellet fractions (P) of RIPA lysate were determined by Western blotting. Phosphorylated HSF1 (pHSF1) protein was used as a positive control for the heat shock response. Morphological changes in the Z/R oncoprotein with or without hyperthermia were determined by confocal microscopy (d). The scale bar is 5 µm. Dynamic changes in the Z/R protein by hyperthermia were determined by FRAP (e). The normal and hyperthermia-treated cells were exposed to 10 µg/mL CHX for the indicated time intervals. Then, the Z/R protein levels were analyzed by Western blotting (left panel) and quantified by ImageJ (right panel) (f). The apoptosis-associated proteins PRAP and Caspase-3 were measured by Western blotting (g). HT indicates hyperthermia.
However, hyperthermia did not affect the morphological features of the Z/R oncoprotein, but it significantly reduced its protein mobility (Fig. 2d, e), which is probably associated with Z/R protein solubility changes. In addition, hyperthermia was able to significantly shorten the half-life of the Z/R protein (Fig. 2f), implying that hyperthermia accelerates the degradation of Z/R oncogenic proteins. However, cleavage of PRAP-1 and caspase-3 was not observed under these treatment conditions (Fig. 2g), indicating that hyperthermia-induced Z/R protein stability changes and degradation are not caused by apoptosis.
Of note, nuclear receptor corepressors (NCoRs) could bind to the Z/R fusion protein through the POZ domain in the ZBTB16 part and the LBD domain in the RARα part. To validate whether NcoRs participate in hyperthermia-induced Z/R destabilization, we constructed several functional domain deletion truncation mutants (Fig. 3a). Deletion of the POZ domain or LBD domain alone (but not the DBD domain) resulted in partial resistance to hyperthermia (Fig. 3b, c), while dual deletion of POZ and LBD led to complete resistance (Fig. 3d). We further observed that hyperthermia promoted the interaction between NCoR1/SMRT and Z/R through the POZ and LBD domains, which verified the important role of NCoR1/SMRT in this process (Fig. 3e).
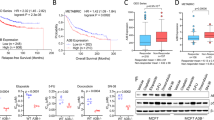
HeLa cells were transiently transfected with four different truncated Z/R plasmids, including single deletion of POZ, LBD, DBD, or double deletion of POZ and LBD (a), and then subjected to hyperthermia. Stability changes of Z/R and its mutants were determined by Western blots (b, c, d). The interaction of NCoR1/SMRT with Z/R and its mutants was analyzed by an immunoprecipitation (IP) assay using an anti-Flag antibody (e). Wild-type (f) and POZ domain-truncated (g) Z/R-overexpressing HeLa cells were pretreated with ATRA (2 µM) for 4 h and then subjected to hyperthermia at 42 °C for 1 h. Z/R proteins in soluble (S) and pellet (P) fractions of RIPA lysate were determined by Western blots. Effect of the corepressor NCoR1 and SMRT knockdown by siRNA on hyperthermia-induced stability changes of Z/R (h). si-NT, nontargeting siRNA; si-N/S, si-NCoR1, and si-SMRT. HT indicates hyperthermia.
It has been reported that ATRA can disassociate NCoRs from the LBD domain but not the POZ domain of the Z/R fusion protein. When cells were pretreated with ATRA, destabilization of the Z/R fusion protein by hyperthermia was partially impeded (Fig. 3f), and the mutant Z/R protein lacking the POZ domain showed complete resistance to hyperthermia (Fig. 3g). Furthermore, we simultaneously overexpressed Z/R and knocked down NCoR1/SMRT in HeLa cells and found that depletion of NCoR1/SMRT significantly alleviated hyperthermia-mediated Z/R destabilization (Fig. 3h and Fig. S5a). Taken together, these data suggest that the interaction with NCoR1/SMRT is essential for hyperthermia-induced Z/R protein destabilization.
For determination of whether destabilization of the Z/R fusion protein by hyperthermia is reversible, heat-treated cells were recovered at 37 °C. We found that the insoluble Z/R fusion protein was significantly decreased, accompanied by reappearance of Z/R in the soluble fraction following recovery from heat (Fig. 4a, compare lane 3 with 2). Then, the protein synthesis inhibitor CHX was used to exclude the influence of the newly synthesized Z/R fusion protein. Following treatment with CHX combined with hyperthermia, the Z/R fusion protein was unable to reappear in the soluble fraction (S) after recovery at 37 °C and gradually degraded in the pellet (Fig. 4a, compare lane 5 with 4; Fig. 4b), suggesting that destabilization of the Z/R protein is irreversible.
Z/R-overexpressing HeLa cells were pretreated with or without 10 µg/ml cycloheximide (CHX) for 1 h, followed by exposure to hyperthermia at 42 °C for 1 h. Afterward, the cells were allowed to recover at 37 °C for 12 h in the presence or absence of CHX (a). Likewise, cells were also recovered at 37 °C in the presence of CHX in a time-dependent manner (b). The abovementioned cells were recovered at 37 °C in the presence of MG132 (10 µM) or CQ (20 µM) to determine the pathway of hyperthermia-induced insoluble Z/R protein (i.e., in pellet) degradation (c). Z/R protein changes in the supernatants (S) and pellet fractions (P) were assessed by Western blot with an anti-Flag antibody. Additionally, ubiquitination of the Z/R protein by hyperthermia was assessed. HeLa cells cotransfected with HA-Ub and Flag-Z/R were exposed to hyperthermia. Ubiquitination of Z/R was analyzed by immunoprecipitation (IP) with anti-Flag (d) and anti-HA (e) antibodies. Wild-type (f), three lysine residue mutants (e.g., ubiquitin-binding sites; K242R, K387R, K396R) (g), POZ and LBD domain-deleted Flag-Z/R (h) were transfected into HeLa cells. NCoR1/SMRT (i) and SIAH2 (j) were silenced in the Z/R-overexpressing HeLa cells by siRNAs. These transfected HeLa cells were subjected to hyperthermia at 42 °C for 1 h, followed by immunoprecipitation with an anti-Flag antibody. The ubiquitination changes were analyzed by Western blots. si-NT, nontargeting siRNA; HT indicates hyperthermia.
The ubiquitin–proteasome system and the autophagy-lysosome pathway are the two major routes for protein clearance in eukaryotic cells. To determine which route is involved in hyperthermia-induced insoluble Z/R protein degradation, we used the proteasome inhibitor MG132 and lysosome inhibitor chloroquine (CQ). Interestingly, degradation of the Z/R protein mediated by hyperthermia was completely inhibited by MG132 but barely affected by CQ (Fig. 4c; compare lanes 5 and 4 with 3), implying that the Z/R fusion protein is predominantly degraded through the proteasome pathway.
Since the conjugation of polyubiquitin (or SUMO) chains is a major signal for proteasome-mediated degradation, we further assessed hyperthermia-induced modifications of the Z/R fusion protein. Hyperthermia markedly enhanced both exogenous (Fig. 4d, e) and endogenous (Fig. 4f) ubiquitination of the Z/R fusion protein, while no SUMOylation was found (Fig. S5b). A previous study indicated that three lysine residues, K242, K387, and K396, in ZBTB16 are the major ubiquitination sites [27, 28]. We found that single or double mutations slightly attenuated Z/R ubiquitination (Fig. S5c, d), while mutation of three lysine residues (e.g., K242R, K387R, and K396R) completely blocked Z/R ubiquitination by hyperthermia (Fig. 4g), indicating that hyperthermia mainly ubiquitinated the Z/R protein at these three lysine residues. However, the three lysine residue-mutated Z/R could still be destabilized by hyperthermia (Fig. S5e). Additionally, the dual POZ and LBD deletion or NCoR1/SMRT knockdown that impeded the destabilization of Z/R prevented ubiquitination after hyperthermia (Fig. 4h, i). These results suggest that destabilization is a prerequisite for ubiquitination and proteasomal degradation of Z/R by hyperthermia.
SIAH2, known as an E3 ubiquitin ligase for NCoR1, plays a crucial role in hyperthermia-mediated ubiquitination of the PML/RARα oncoprotein [25]. We reasoned that SIAH2 might also be involved in hyperthermia-mediated Z/R protein ubiquitination. Indeed, knocking down SIAH2 significantly decreased the ubiquitination of the Z/R protein (Fig. 4j and Fig. S5f). In addition, we investigated the involvement of Hsp70 and Hsp90 in this process. Inhibition of either Hsp70 or Hsp90 activity by their corresponding specific inhibitors failed to prevent both destabilization and ubiquitination of the Z/R fusion protein by hyperthermia (Fig. S6a–c), suggesting that SIAH2 but not Hsp70 and Hsp90 was important for hyperthermia-induced ubiquitination of the Z/R fusion protein.
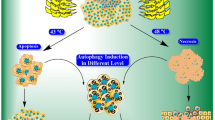
Thus, we illustrated the mechanism of Z/R protein degradation by hyperthermia through the following steps: hyperthermia induces Z/R fusion protein destabilization in an NCoR1/SMRT-dependent manner and then promotes the ubiquitination of destabilized fusion proteins through the E3 ligase SIAH2, finally leading to fusion protein degradation via the proteasome pathway (Fig. 5).
Discussion
Generally, oncogenic fusion proteins are well-defined drivers in many leukemias [16, 21, 22, 29,30,31]. Among them, more than 95% of APL cases are driven by chromosomal translocation between chromosomes 15 and 17 t(15;17)(q24;q21), which results in the generation of PML/RARα oncogenic fusion proteins [4, 32]. The development of ATRA and ATO dramatically improves the survival of patients with APL, making the deadly disease curable (complete remission rate of nearly 95%) [4, 12, 33]. Of note, subgroups of acute leukemia caused by RARα-related variant translocations have also been found clinically, and the morphologic, biologic, and clinical features of these variants are similar to those of classic APL [9, 16, 34]. For instance, these rearrangement partners of RARα include ZBTB16, NUMA1, NPM1, and STAT5B [34,35,36,37]. Among these variant fusions, ZBTB16/RARα (Z/R) is the most common (~2% of APL cases) and displays extreme resistance to the ATRA and ATO combination regimen [9, 38]. Although chemotherapy has shown a positive effect on certain Z/R-driven APL, it could not significantly improve the prognosis of patients. In addition, although modern drugs, such as tyrosine kinase inhibitors, chimeric antigen receptor T-cell therapy, and monoclonal antibodies, have dramatically changed the treatment landscape and overall survival in patients with AML [25, 39,40,41], there are no such drugs available for Z/R-driven APL. Thus, the discovery of new drugs or therapeutic strategies is urgently needed to improve outcomes in patients with Z/R-driven APL.
Here, we confirmed that the Z/R fusion protein was completely resistant to ATO and slightly responsive to ATRA, quite different from PML/RARα that can be effectively targeted and degraded by both drugs. We previously demonstrated that the arsenic derivative phenylarsine oxide is able to degrade the Z/R protein, which was found to be dependent on apoptosis [14].
Hyperthermia for cancer treatment was first introduced by William Coley by using bacterial infection to induce fever, but it was not widely adopted due to its unrevealed mechanism. However, fever has inspired the development of a strategy involving the use of heat in cancer treatment to some extent [42]. Currently, hyperthermia is either administered independently or, more often, in combination with radiotherapy or chemotherapy for management of solid tumor or other conditions and has shown satisfactory outcomes [25, 43, 44].
Very recently, we revealed for the first time that hyperthermia has superior therapeutic effects on relapsed and refractory APL patients by selectively destabilizing PML/RARα fusion proteins (including all drug-resistant mutations) in an NCoRs-dependent manner [25]. Given that Z/R is comparable to PML/RARα with respect to strong binding to NCoR1/SMRT corepressors, we speculated that Z/R can be destabilized by hyperthermia as well. Strikingly, hyperthermia robustly destabilized the Z/R protein within a very short time (0.5–1 h) without compromising cell viability, suggesting that the Z/R fusion protein is indeed susceptible to hyperthermia. In contrast to PML/RARα, hyperthermia could not induce apparent aggregation in the Z/R fusion protein, implying that destabilization of the Z/R protein is not tightly associated with aggregation.
The Z/R fusion protein could interact with NCoR1/SMRT corepressors through the POZ and LBD domains in the ZBTB16 and RARα parts, respectively. We further found that deletion of either the POZ or LBD partially inhibited fusion protein destabilization by hyperthermia, while protein destabilization was completely blocked by simultaneous deletion of these two domains. Likewise, hyperthermia promoted the NCoR1/SMRT and Z/R interaction, and knocking down endogenous NCoR1/SMRT completely blocked Z/R fusion protein destabilization. These results confirm that hyperthermia-mediated Z/R destabilization is dependent on the interaction of NCoR1/SMRT with the POZ and LBD domains of Z/R. In addition, protein destabilization by hyperthermia led to a decrease in Z/R mobility, and this process was found to be irreversible. Interestingly, knockdown of NCoRs that impeded hyperthermia-induced Z/R destabilization and prevented further ubiquitination, while mutation of three lysine residues that completely blocked ubiquitination could still be destabilized by hyperthermia, suggesting that the process of destabilization occurred prior to ubiquitination. Moreover, SIAH2 was identified as an E3 ligase for hyperthermia-mediated ubiquitination. Heat shock proteins, including HSF1, Hsp70, and Hsp90, can be rapidly upregulated by heat stress stimulation, which assists in the refolding of misfolded proteins or aids in the elimination of damaged proteins [45, 46]. However, neither Hsp70 nor Hsp90 protein participated in heat-induced Z/R destabilization and ubiquitination.
Based on our results, the effects of hyperthermia on the Z/R fusion protein are carried out in several steps: the 1st step is rapid destabilization (protein solubility shift), the 2nd step is Ub modification, and the final step is proteasomal degradation. Although the current study is based on in vitro observations, these findings strongly imply that Z/R’s thermal instability represents a biophysical vulnerability and that by exploiting this, a hyperthermia-based therapeutic method could be developed to cope with Z/R-driven APL.
References
Bradner JE, Hnisz D, Young RA. Transcriptional Addiction in cancer. Cell. 2017;168:629–43.
Zhuang JJ, Liu Q, Wu DL, Tie L. Current strategies and progress for targeting the “undruggable” transcription factors. Acta Pharmacol Sin. 2022. https://doi.org/10.1038/s41401-021-00852-9.
Hasserjian RP, Nardi V. Bedside to bench and back: identifying a new clinically relevant driver in pediatric acute myeloid leukemia. Blood Cancer Discov. 2022;3:173–5.
Wang ZY, Chen Z. Acute promyelocytic leukemia: from highly fatal to highly curable. Blood. 2008;111:2505–15.
Jiao B, Ren ZH, Liu P, Chen LJ, Shi JY, Dong Y, et al. 8-CPT-cAMP/all-trans retinoic acid targets t(11;17) acute promyelocytic leukemia through enhanced cell differentiation and PLZF/RARα degradation. Proc Natl Acad Sci USA. 2013;110:3495–500.
De Thé H, Chen Z. Acute promyelocytic leukaemia: novel insights into the mechanisms of cure. Nat Rev Cancer. 2010;10:775–83.
Girard N, Tremblay M, Humbert M, Grondin B, Haman A, Labrecque J, et al. RARα-PLZF oncogene inhibits C/EBPα function in myeloid cells. Proc Natl Acad Sci USA. 2013;110:13522–7.
Sanz MA, Fenaux P, Tallman MS, Estey EH, Löwenberg B, Naoe T, et al. Management of acute promyelocytic leukemia: updated recommendations from an expert panel of the European LeukemiaNet. Blood. 2019;133:1630–43.
Hussain L, Maimaitiyiming Y, Islam K, Naranmandura H. Acute promyelocytic leukemia and variant fusion proteins: PLZF-RARα fusion protein at a glance. Semin Oncol. 2019;46:133–44.
George B, Kantarjian H, Baran N, Krocker JD, Rios A. Tp53 in acute myeloid leukemia: molecular aspects and patterns of mutation. Int J Mol Sci. 2021;22:10782.
Mengeling BJ, Phan TQ, Goodson ML, Privalsky ML. Aberrant corepressor interactions implicated in PML-RARα and PLZF-RARα leukemogenesis reflect an altered recruitment and release of specific NCoR and SMRT splice variants. J Biol Chem. 2011;286:4236–47.
Zhang XW, Yan XJ, Zhou ZR, Yang FF, Wu ZY, Sun HB, et al. Arsenic trioxide controls the fate of the PML-RARα oncoprotein by directly binding PML. Science. 2010;328:240–3.
Koken MHM, Daniel MT, Gianni M, Zelent A, Licht J, Buzyn A. et al. Retinoic acid, but not arsenic trioxide, degrades the PLZF/RARα fusion protein, without inducing terminal differentiation or apoptosis, in a RA-therapy resistant t(11;17)q23;q21) APL patient. Oncogene. 1999;18:1113–8.
Hussain L, Maimaitiyiming Y, Su L, Wang QQ, Naranmandura H. Phenylarsine oxide can induce degradation of PLZF-RARα variant fusion protein of acute promyelocytic leukemia. Chem Res Toxicol. 2019;32:548–50.
Kitamura K, Hoshi S, Koike M, Kiyoi H, Saito H, Naoe T. Histone deacetylase inhibitor but not arsenic trioxide differentiates acute promyelocytic leukaemia cells with t(11;17) in combination with all- trans-retinoic acid. Br J Haematol. 2000;108:696–702.
Grignani F, De Matteis S, Nervi C, Tomassoni L, Gelmetti V, Cioce M, et al. Fusion proteins of the retinoic acid receptor-α recruit histone deacetylase in promyelocytic leukaemia. Nature. 1998;391:815–8.
Wong CW, Privalsky ML. Components of the SMRT corepressor complex exhibit distinctive interactions with the POZ domain oncoproteins PLZF, PLZF-RARalpha, and BCL-6. J Biol Chem. 1998;273:27695–702.
Morimoto S, Kondo T, Taya T, Matsuo H, Teramoto Y, Mizumoto C, et al. Successful allogeneic bone marrow transplantation in a case of variant acute promyelocytic leukemia with ZBTB16-RARA. Ann Hematol. 2022;101:1129–32.
Strehl S, König M, Boztug H, Cooper BW, Suzukawa K, Zhang SJ, et al. All-trans retinoic acid and arsenic trioxide resistance of acute promyelocytic leukemia with the variant STAT5B-RARA fusion gene. Leukemia. 2013;27:1606–10.
Cicconi L, Testi AM, Montesinos P, Rego E, Zhu HH, Takahashi H, et al. Characteristics and outcome of acute myeloid leukemia with uncommon retinoic acid receptor-alpha (RARA) fusion variants. Blood Cancer J. 2021;11:167. https://doi.org/10.1038/s41408-021-00561-w.
Geoffroy MC, de Thé H. Classic and variants APLs, as viewed from a therapy response. Cancers. 2020;12:967.
Renneville A, Gasser JA, Grinshpun DE, Jean Beltran PM, Udeshi ND, Matyskiela ME, et al. Avadomide induces degradation of ZMYM2 fusion oncoproteins in hematologic malignancies. Blood Cancer Discov. 2021;2:250–65.
Wang QQ, Wang HF, Zhao JZ, Naranmandura H, Jin J, Zhu HH. Venetoclax for arsenic-resistant acute promyelocytic leukaemia. Br J Haematol. 2022;197:e58–e60.
Jiang YH, Chen YJ, Wang C, Lan YF, Yang C, Wang QQ, et al. Phenylarsine oxide can induce the arsenite-resistance mutant PML protein solubility changes. Int J Mol Sci. 2017;18:247.
Maimaitiyiming Y, Wang QQ, Yang C, Ogra Y, Lou Y, Smith CA, et al. Hyperthermia selectively destabilizes oncogenic fusion proteins. Blood Cancer Discov. 2021;2:388–401.
Wu HC, Rérolle D, de Thé H. PML/RARα destabilization by hyperthermia: a new model for oncogenic fusion protein degradation? Blood Cancer Discov. 2021;2:300–1.
Chao TT, Chang CC, Shih HM. SUMO modification modulates the transrepression activity of PLZF. Biochem Biophys Res Commun. 2007;358:475–82.
Kang SI, Choi HW, Kim IY. Redox-mediated modification of PLZF by SUMO-1 and ubiquitin. Biochem Biophys Res Commun. 2008;369:1209–14.
Melnick A, Licht JD. Deconstructing a disease: RARalpha, its fusion partners, and their roles in the pathogenesis of acute promyelocytic leukemia. Blood. 1999;93:3167–215.
Mistry AR, Pedersen EW, Solomon E, Grimwade D. The molecular pathogenesis of acute promyelocytic leukaemia: Implications for the clinical management of the disease. Blood Rev. 2003;17:71–97.
Yun S, Vincelette ND, Yu X, Watson GW, Fernandez MR, Yang C, et al. TFEB Links MYC signaling to epigenetic control of myeloid differentiation and acute myeloid leukemia. Blood Cancer Discov. 2021;2:162–85.
Liquori A, Ibañez M, Sargas C, Sanz MÁ, Barragán E, Cervera J. Acute promyelocytic leukemia: a constellation of molecular events around a single PML-RARA fusion gene. Cancers. 2020;12:624.
Lagunas-Rangel FA, Chávez-Valencia V, Gómez-Guijosa MÁ, Cortes-Penagos C. Acute myeloid leukemia—genetic alterations and their clinical prognosis. Int J Hematol Stem Cell Res. 2017;11:328–39.
Zhou GB, Li G, Chen SJ, Chen Z. From dissection of disease pathogenesis to elucidation of mechanisms of targeted therapies: leukemia research in the genomic era. Acta Pharmacol Sin. 2007;28:1434–49.
Gurnari C, Voso MT, Girardi K, Mastronuzzi A, Strocchio L. Acute promyelocytic leukemia in children: a model of precision medicine and chemotherapy-free therapy. Int J Mol Sci. 2021;22:642.
Song Y, Hou J, Wan L, Liu K, Zhou C, Wei S, et al. A short report of novel RARG-HNRNPM fusion gene in resembling acute promyelocytic leukemia. Hematology. 2022;27:518–22.
Chen Y, Li S, Zhou C, Li C, Ru K, Rao Q, et al. TBLR1 fuses to retinoid acid receptor α in a variant t(3;17)(q26;q21) translocation of acute promyelocytic leukemia. Blood. 2014;124:936–45.
Zhang X, Sun J, Yu W, Jin J. Current views on the genetic landscape and management of variant acute promyelocytic leukemia. Biomark Res. 2021;9:33.
Stoddart A, Wang J, Fernald AA, Davis EM, Johnson CR, Hu C, et al. Cytotoxic therapy–induced effects on both hematopoietic and marrow stromal cells promotes therapy-related myeloid neoplasms. Blood Cancer Discov. 2020;1:32–47.
Alotaibi AS, Yilmaz M, Kanagal-Shamanna R, Loghavi S, Kadia TM, DiNardo CD, et al. Patterns of resistance differ in patients with acute myeloid leukemia treated with type I versus type II FLT3 inhibitors. Blood Cancer Discov. 2021;2:125–34.
McCarter AC, Della Gatta G, Melnick A, Kim E, Sha C, Wang Q, et al. Combinatorial ETS1-dependent control of oncogenic NOTCH1 enhancers in T-cell leukemia. Blood Cancer Discov. 2020;1:178–97.
Hobohm U. Fever and cancer in perspective. Cancer Immunol Immunother. 2001;50:391–6.
Ahmed K, Zaidi SF, Mati-ur-Rehman, Rehman R, Kondo T. Hyperthermia and protein homeostasis: cytoprotection and cell death. J Therm Biol. 2020;91:102615.
Maimaitiyiming Y, Yang T, Wang QQ, Feng Y, Chen Z, Björklund M, et al. Heat treatment promotes ubiquitin-mediated proteolysis of SARS-CoV-2 RNA polymerase and decreases viral load. Research. 2022;2022:9802969.
Petrucelli L, Dickson D, Kehoe K, Taylor J, Snyder H, Grover A, et al. CHIP and Hsp70 regulate tau ubiquitination, degradation and aggregation. Hum Mol Genet. 2004;13:703–14.
Taipale M, Jarosz DF, Lindquist S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat Rev Mol Cell Biol. 2010;11:515–28.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (No. 81872942, 82003875, 82000155, 82000145, 82200160), the China Postdoctoral Science Foundation (No. 2021M702875, 2021M702877), and the Leading Innovative and Entrepreneur Team Introduction Program of Zhejiang, Grant/Award Number: 2020R01006. We thank the Core Facilities of Zhejiang University School of Medicine for technical support.
Author information
Authors and Affiliations
Contributions
The study was conceived by YFM and HN; the experiments were performed by QQW, LH, and YFM; PHY, CYZ, CY, YYK, SCW, TY, and WJY helped with data processing and presentation. QQW and LH wrote the draft. YFM and HN approved and revised the manuscript. PHY, CYZ, YFM, and WJY assisted with the revision and polishing of the manuscript; all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Qq., Hussain, L., Yu, Ph. et al. Hyperthermia promotes degradation of the acute promyelocytic leukemia driver oncoprotein ZBTB16/RARα. Acta Pharmacol Sin 44, 822–831 (2023). https://doi.org/10.1038/s41401-022-01001-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-022-01001-6