Abstract
Amyotrophic lateral sclerosis (ALS) is one of the most common fatal neurodegenerative diseases in adults. ALS pathogenesis is associated with toxic SOD1 aggregates generated by mutant SOD1. Since autophagy is responsible for the clearance of toxic protein aggregates including SOD1 aggregates, autophagy induction has been considered as a potential strategy for treating ALS. Autophagic signaling is initiated by unc-51 like autophagy activating kinase 1 (ULK1) complex. We previously identified that BL-918 as a specific ULK1 activator, which exerted cytoprotective effect against Parkinson’s disease in vitro and in vivo. In this study we investigated whether BL-918 exerted a therapeutic effect against ALS, and characterized its pharmacokinetic profile in rats. In hSODG93A-NSC34 cells, treatment with BL-918 (5, 10 μM) dose-dependently induced ULK1-dependent autophagy, and eliminated toxic SOD1 aggregates. In SODG93A mice, administration of BL-918 (40, 80 mg/kg, b.i.d., i.g.) dose-dependently prolonged lifespan and improved the motor function, and enhanced the clearance of SOD1 aggregates in spinal cord and cerebral cortex through inducing autophagy. In the pharmacokinetic study conducted in rats, we found BL-918 and its 2 metabolites (M8 and M10) present in spinal cord and brain; after intragastric and intravenous administration, BL-918 reached the highest blood concentration compared to M8 and M10. Collectively, ULK1 activator BL-918 displays a therapeutic potential on ALS through inducing cytoprotective autophagy. This study provides a further clue for autophagic dysfunction in ALS pathogenesis.
Similar content being viewed by others
Introduction
Amyotrophic Lateral Sclerosis (ALS), one of the most common fatal neurodegenerative diseases in adults, is characterized by the selective loss of motor neurons in the ventral horn of spinal cord, cerebral cortex and the brain stem nucleus [1, 2]. Taking 10% of all ALS diseases, familial ALS is associated with genetic mutations such as C9orf72, SOD1, TARDBP [3,4,5]. Among these mutants, mutant of Cu/Zn superoxide dismutase 1 (SOD1) was the first to be discovered, accounting for 20%-25% familiar ALS [6]. Mutant SOD1 undergoes conformation changes easily, leading to abnormal protein aggregates [7,8,9]. Aggregates inclusions were found both in human familiar ALS and SODG93A mouse models which are used to investigate the pathogenesis of mutant SOD1 in ALS [10]. Moreover, ventral horn samples of ALS patients with SOD1 mutations displayed low autophagy capacity and more SOD1 aggregations [11]. Some research has reported that toxic SOD1 aggregates contribute to ALS through toxic endoplasmic reticulum (ER) stress induced by oxidative response and overload of unfolded protein [12, 13]. The aggregates seem to escape normal degradation process to exert toxic effect and impair autophagy [14]. Thus, induction of autophagy may be of considerable potential to treat ALS through eliminating toxic SOD1 aggregates [15].
Autophagy is a protein degradation pathway which involves formation of double-membrane autophagosomes and its fusion of lysosomes, and it deals with long-lived proteins and impaired organelles [16, 17]. Autophagic pathway is initiated by unc-51 like autophagy activating kinase 1 (ULK1) complex and involves the formation of a cytoplasmic double-membrane structure known as autophagosome in which cargos aggregate. ULK1, an important serine/threonine kinase in autophagy, is imperative in the recruitment of downstream autophagy-related proteins as well as in the formation of autophagosome. It can be activated by the direct phosphorylation of AMP-activated protein kinase (AMPK) at Ser317 and Ser777 and inhibited by mechanistic target of rapamycin (mTOR) at Ser757 phosphorylation [18]. Subsequently, ULK1, FIP200, ATG13 and ATG101 form a complex, called ULK1 complex, which regulates downstream pathways of autophagy [19,20,21]. Recently, accumulating evidence has illuminated that autophagy acts as an important way to degrade mutant SOD1 in ALS, reducing the toxicity of mutant SOD1 proteins [22,23,24,25]. Also, a study reported the involvement of mTOR pathway in human ALS SODG93A astrocytes, in which p-ULK1 Ser757 was highly expressed than control cells, indicating the impairment of autophagy in ALS SODG93A astrocytes [26]. Accordingly, attempts of using lithium, trehalose and other autophagy inducer to treat ALS displayed good outcome, including delayed disease onset and prolonged lifespan of SODG93A mice [27]. Thus, we hypothesized that targeting ULK1 to induce autophagy and ultimately eliminating toxic SOD aggregates may be a potential way to treat ALS.
In our previous study, BL-918 was identified as a specific ULK1 activator, presenting activity against Parkinson’s disease (PD) both in vitro and in vivo through autophagy induction by phosphorylating ULK1 [28]. Therefore, we assume that BL-918 may exhibit therapeutic potential on ALS by inducing autophagy thereby eradicating toxic SOD1 aggregates due to SOD mutations. Additionally, low molecular weight of BL-918 (Mw = 533.0808 Da) are considered suitable to penetrate the brain-blood barrier. Here, we investigated the pharmacokinetic profiles of BL-918, as well as the effect and possible mechanisms of this autophagy inducer on ALS using hSODG93A-NSC34 cells and SODG93A mice and provided a clue for autophagic dysfunction in ALS pathogenesis.
Materials and methods
Cell lines and cell culture
NSC34 cells were cultured in RPMI-1640 Medium containing 10% fetal bovine serum (FBS) (PAN-Biotech, Germany) and maintained at 37 °C in a humidified incubator in an atmosphere of 5% CO2. hSOD1WT-mCherry-NSC34 and hSOD1G93A-mCherry-NSC34 cells were generated by stable transduction of NSC34 cells (Jennio Biotech, Guangzhou, China) according to LipofectamineTM LTX and PlusTM Reagent (Invitrogen, Thermo Fisher, Waltham, MA, USA, 15338030) and cultivated in 1640 medium containing 250 μg/mL G418 (Sigma-Aldrich, Burlington, MA, USA) in bottles free of CO2, at 37 °C.
Antibodies and reagents
Antibodies used were as follows: superoxide dismutase 1 (SOD1) (Abcam, Cambridge, UK, ab16831), SQSTM1/p62 (Abcam, ab56416), LC3B (CST, Danvers, MA, USA, 2775 S), LC3B (Abcam, ab51520), ULK1 (CST, 8054 T), ULK1 (Abcam, ab167139), phospho-ULK1 (Ser757) (D7O6U) (CST, 14202 S), phospho-ULK1 (Ser555) (D1H4) (CST, 5869 S), phospho-ULK1 (Ser317) (CST, 37762 S), mTOR (Proteintech, Rosemont, IL, USA, 66888-1-IG), phospho-mTOR (Ser2448) (D9C2) XP® (CST, 5536 S), phospho-AMPKα (Thr172) (40H9) (CST, 2535 S), Beclin-1 (2A4) (CST, 4122 S), phospho-Beclin-1 (Ser15) (D4B7R) (CST, 84966 S), Atg101 (E1Z4W) (CST, 13492 S), phospho-ATG13 (S318) (Abnova, Walnut, CA, USA, PAB19948), FIP200 (D10D11) (CST, 12436 S), anti-choline acetyltransferase (ChAT) antibody (Merck-Millipore Biosciences, Darmstadt, Germany, AB144P), GAPDH antibody, β-actin antibody (Fude Biological Technology, FD0063, FD0060), donkey anti-goat IgG-HRP (Santa Cruz, Dallas, TX, USA, sc 2020), goat anti-rabbit-HRP, goat anti-mouse-HRP (Fude Biological Technology, FDR007, FDM007), BECN1 antibody (H-300) (Santa Cruz, sc-11427).
Chemicals
BL-918 (synthesized), Masson stain (Leagene, Beijing, China, DC0034), Nissl stain (Beyotime, Nantong, China, C0117), LipofectamineTM LTX PlusTM reagent, bafilomycin A1 (Sigma, Louis, MO, USA, B1793), rapamycin (Selleck, Houston, TX, USA, S1039), 3-MA (Sigma, M9281), TRIzol (Bio-rad, Hercules, California, USA), SYBR green Mix (Vazyme Biotech, Nanjing, China) were used.
Crystal violet staining
Transient transfected hSOD1WT-mCherry-NSC34 and hSOD1G93A-mCherry-NSC34 cells as above mentioned were incubated for 12 h transfection, and then were seeded in 96-well flat bottom microtiter plates. After 12 h, cells were treated with BL-918, M8, and M10, respectively (10 μM) and back to incubation for 36 h. The medium was removed and 100 μL of 10% methanol solution was added to fix the cells. Then methanol solution was removed and 100 μL crystal violet staining solution was added, the dying lasted for 20 min under 25 °C. The dyed cells were scaned under IN Cell 6000 HCA system, and the number of cells was quantified using ImageJ.
Autophagic flux assay
hSOD1G93A-mCherry-NSC34 cells and hSOD1WT-mCherry-NSC34 cells were transiently transfected with mRFP-GFP-LC3 for 24 h, and treated with 10 μM BL-918 as experimental group, starvation as positive control, treatment with Baf1 (80 nM) as negative control all for another 24 h. After that, cells were washed by PBS and fixed by 4% paraformaldehyde for 15 min, followed by PBS washing twice. Then the sampled cells were examined and analyzed by Confocal Laser Scanning Microscope and LC3-punctas were quantified by ImageJ.
Western blot analysis
The collected spinal cord and cerebral cortex were homogenized at 4 °C for 30 min, light-free, with 120 μL lysis buffer [29] containing protease inhibitor and 18.5 mg/mL iodoacetamide (IA) [30], a sulfhydryl-reactive alkylating reagent used to avoid air oxidation and disulfide-bond scrambling, which prevent the misfolded SOD1 from dissociating. Cell lysates were centrifuged at 12,000 rpm for 10 min at 4 °C and lysed in the same lysis buffer containing IA as these tissues, and then the supernatant lysates were collected and determined for the protein concentration with the BCA Protein Assay Kit. And equal amounts of protein (diluted with SDS-PAGE buffer without DL-Dithiothreitol (DTT) to avoid disulfide-bond reducting [30]) were separated by 10% SDS-PAGE, and then transferred onto the PDVF membranes. The membranes were blocked with TBST containing 5% skimmed milk for 1 h at room temperature, followed by incubation with indicated antibodies (dilution rate, 1:1000) overnight at 4 °C and then HRP-flagged secondary antibodies at room temperature for 2 h. Finally, the membranes were processed by ECL reagent and examined by electron imaging.
Pharmacokinetics experiments
Animal treatment and sample preparation
Male rats were separated into experimental group and blank control group with 9 rats in each group and treated with BL-918 (50 mg/kg) twice at an internal of 12 h. Plasma (0.5 mL) was collected at 0.5, 1, 2, 4, 6, 7, 8, 10, 12, 24, 36 h from external jugular vein and restored in heparinized plastic tubes to undergoing centrifugation at 12,000 r/min for 15 min. Urine and excrement were collected instantly after treatment for 24 h and centrifugated at 12,000 rpm for 10 min. Bile was collected after the second treatment, and spinal cord and cerebral cortex were also collected. All these samples underwent SPE.
UPLC-Q-TOF-MS analysis
The UPLC method employed a Waters ACQUITYTM BEH C18 column (2.1 mm × 50 mm, 1.7 μm) at the temperature of 15 °C. Peaks were separated using mobile phases of 0.1% formic acid-water and 0.1% formic acid-acetonitrile at a flow rate of 0.4 mL/min. MS method used electrospray ionization (ESI) and the data were acquired in centroid mode. The injection volume was 2 μL. Waters Masslynx 4.1 platform was used to acquire total ion chromatogram (TIC) in MSE mode. Metabolites were analyzed by MetabolynxTM.
Animals and treatment
SODG93A mice (170-day old) kept at room temperature 23 ± 2 °C, with 12 h light a day and sufficient food and water were randomly divided into four groups: (1) WT-NT group (treated with solvent within 0.3% CMC-Na and 0.4% tween 80); (2) TG-NT (treated the same with WT-NT group); (3) TG-BL-918-L (treated with 40 mg/kg BL-918 dissolved in same solvent); and (4)TG-BL-918-H (treated with 80 mg/kg BL-918 dissolved in same solvent).
Mice were intragastrically administered every two days with solvent solely or at different doses of BL-918 until the late stage of disease. Behavioral training began three days before administration and mice underwent behavior testing once a week simultaneous with the beginning of administration. Samples were collected after the last drug treatment.
Staining
After treatment, mice were narcotized with 10% chloral hydrate and dissected to expose the hearts, and then the needle was inserted from the apex of the left ventricle of the mouse heart, the right atrial appendage of the heart was cut, and the blood was rinsed in the mouse with normal saline. Spinal cord was taken out and L2-L4 parts of it were separated and fixed in 4% paraformaldehyde for 7 to 14 days. Then the spinal cord was dehydrated with 10%, 20%, 30% sucrose solution at 4 °C. Next, the spinal cord was embedded in OCT and then cut at the thickness of 10 μm under −20 °C. The sections of spinal cord were stained after heating and fixing by Nissl stain while the sections of gastrocnemius muscle were stained directly by Masson staining.
Behavioral experiment
Rotarod Test
Motor function and skeletal functional performance were assessed by using rotarod test (MK-610A, Muromachi Kikai, Tokyo, Japan). Mice were tested during the age of nineteen to forty-six weeks old. Mice were trained three days before drug treatment for at least three times. If the time which a mouse could remain on the rotating rod was under 5 min, we define the mouse as motor dysfunction. Mice were placed onto the apparatus at a constant speed of 20 rpm and the remaining time was recorded in three repeated attempts.
Hanging wire test
Mice were allowed to grip wire which is turned over (above the surface 30 cm) for 180 s. The time (in second) each mouse fall is recorded as described and each mouse undergoes three repeated attempts. Mice were trained three days before drug treatment for at least three times. Mice were assessed for signs of motor deficit and scored as previously described [31].
RNA-seq data analysis
hSOD1G93A-mCherry-NSC34 cells were treated with 10 μM BL-918 or control in triplicate for 24 h. RNA purity was checked by the NanoPhotomer® spectrophotometer (IMPLEN, CA, USA). RNA concentration was measured with Qubit® RNA Assay Kit in Qubit® 2.0 Flurometer (Life Technologies, CA, USA). RNA integrity was assessed by the RNA Nano 6000 Assay Kit of the Bioanalyzer 2100 system (Agilent Technologies, CA, USA). Sequencing libraries were generated using NEBNext® UltraTM RNA Library Prep Kit for Illumina® (NEB, USA) following manufacturer’s recommendations and index codes were added to attribute sequences to each sample. The clustering of the index-coded samples was performed on a cBot Cluster Generation System using TruSeq PE Cluster Kit v3-cBot-HS (Illumia) according to manufacturer’s instructions.
Fastq files were aligned to human reference genome hg19 using HiSat with default parameters. featureCounts v1.5.0-p3 was used to count the reads numbers mapped to each gene. And then FPKM of each gene was calculated based on the length of the gene and reads count mapped to this gene. The analysis of differentially expressed genes (DEGs) between the treatment and control groups was performed using the DESeq2 R package. P-values (padj) were adjusted using the Benjamini and Hochberg method [32]. A corrected P-value <0.05 and an absolute log2(fold change) >1 were set as the threshold for significantly differential expression. GO and KEGG pathway enrichment analyses of the DEGs were performed by using the DAVID and GSEA annotation. For the GO terms and KEGG pathway enrichment analyses, padj < 0.05 was used as a threshold for significant enrichment by DEGs.
qPCR
RNAs were extracted with Trizol according to the manufacturer’s instruction and fresh RNA underwent RT-PCR to gain corresponding cDNA. Next, a total of 10 μL liquid system containing SYBR green MIX, primers and ddH2O were used to run a qPCR protocol. The primers used are listed below:
Gene | Forward (5′-3′) | Reverse (5′-3′) |
|---|---|---|
ANAXA7 | CTGCTGGGGTCAGAATGTCA | TCCCATTGGAGGAAAGCCAC |
ATG2A | CATGGACACCTCCACCTTCC | AGACATAATCTCGCCGCAGG |
ATG16L2 | TGGGATGAGGGAGAGAAGGG | CACATCCACACAGTGGGACA |
HSPA8 | CCTACACCCCAGCAACCAT | TCAGTGTCCGTAAAGGCGAC |
GABARAPL1 | GACAGGGTCCCCGTGATTG | GTCCTCAGGTCTCAGGTGGA |
AMBRA1 | GCTTTGTCCCAAGGCGTTTC | GRRACTGGCCACAGCAGACT |
Statistical analysis
All the presented data and results were confirmed by at least three independent experiments. The data were expressed as means ± SEM and analyzed with GraphPad Prism 8.0 software. Statistical differences between two groups were determined using Student’s t test, while between multiple groups were determined using one-way analysis of variance. P < 0.05 was considered statistically significant.
Results
BL-918 induces autophagy
BL-918 was previously reported as an autophagy activator in Parkinson’s disease, and in the present study, we investigated the effect on hSODG93A-NSC34 cells. As the accumulation of autophagosomes is considered to reflect the level of autophagy, we observed the accumulation of autophagosomes in hSODG93A-NSC34 cells using transmission electron microscopy (TEM) [33,34,35]. Notably, hSODG93A-NSC34 cells and hSODWT-NSC34 cells treated with 10 μM BL-918 contained many autophagosomes, and few autophagosomes were observed in cells without BL-918 treatment (Fig. 1a and Supplementary Fig. S1). Additionally, compared to control group, more LC3 puncta were found in hSODG93A-NSC34 cells treated with BL-918 (Fig. 1b). As shown by Western blot, 5 μM and 10 μM BL-918 treatment increased the expression of LC3-II in PC12 cells (Fig. 1c), NSC34 cells (Fig. 1d), hSODWT-NSC34 cells (Fig. 1e), and hSODG93A-NSC34 cells (Fig. 1f), indicating the capacity of BL918 on inducing autophagy in different mouse neuron-like cells. Moreover, BL-918 treatment also increased autophagy in a time-dependent manner in hSODG93A-NSC34 cells (Fig. 1g). And the induction of autophagy caused by BL-918 could be reversed by 3-MA, an autophagy inhibitor (Fig. 1h). Moreover, we detected autophagy flux to confirm the effect of BL-918 in hSODG93A-NSC34 cells and hSODWT-NSC34 while using Bafilomycin A1 (a lysosome inhibitor). Compared to cells not treated with 10 μM BL-918, obvious accumulation of LC3 puncta was detected in the presence of BL-918 both in hSODG93A-NSC34 cells and hSODWT-NSC34 cells (Fig. 1i and Supplementary Fig. S2). Moreover, to exclude the effect of proteasome on obliterating SOD1 aggregates, we measured proteasome activity after BL-918 treatment both in vitro and in vivo, and found BL-918 could not eliminate SOD1 aggregates by promoting the activity of proteasome (Supplementary Fig. S3). Collectively, these results indicate that BL-918 induces autophagy in vitro.
a The image of hSODG93A-NSC34 cells under transmission electron microscopy (TEM). More autophagosomes were observed in BL-918 group (10 μM). Scale bar: 1 μm. b hSODG93A-NSC34 cells were transfected with adenovirus expressing mRFP-GFP-LC3, and then treated with BL-918 (10 μM) or rapamycin (50 μM) for 24 h, respectively. The LC3 puncta were detected by confocal microscopy. Scale bar: 15 μm. c–f PC12 cells, NSC34 cells, hSODWT-NSC34 cells and hSODG93A-NSC34 cells were treated with 5 μM and 10 μM BL-918 for 24 h. The expression levels of LC3-I, LC3-II, and p62 were analyzed by Western blot. The rate of LC3-II/LC3-I was quantitatively analyzed (n = 3). g hSODG93A-NSC34 cells were treated with BL-918 for 0 h, 6 h, 12 h, 24 h, and 36 h and the expression of LC3-I, LC3-II, and p62 were analyzed by Western blot. h hSODG93A-NSC34 cells were treated with 3-MA or BL-918 and 3-MA together, and the expression levels of LC3-I, LC3-II, and p62 were analyzed by Western blot (n = 3). i hSODG93A-NSC34 cells were transfected with adenovirus expressing mRFP-GFP-LC3, and then treated with BL-918 (10 μM), Baf1 (80 nM), or BL-918 and Baf1 together for 24 h. And the LC3 puncta were observed with confocal microscopy. Scale bar: 5 μm. ns not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. #P < 0.05, ##P < 0.01, ###P < 0.001.
BL-918 induces cytoprotective autophagy and eliminates toxic SOD1 aggregates
Mutations of SOD1 abolish both SOD1 protein structure and function, which result in abnormal SOD1 aggregates and thereby reducing the degradation of ROS, increasing oxidative activity and impairing the normal function of ER, collectively contributing to ALS [9, 36, 37]. Studies have indicated that wild-type SOD1 rarely contributed to the mutant-mediated toxicity on ALS [7]. Consistent with that, BL-918 treatment showed little effect on SOD1 monomer in hSODWT-NSC34 cells (Fig. 2a), while eliminating SOD1 monomer in hSODG93A-NSC34 cells (Fig. 2b, P < 0.01). Interestingly, we also observed a decrease of endogenous WT mouse SOD1 (mSOD1) in hSODG93A-NSC34 cells, which may be attributed to the clearance of misfolded WT-SOD1 aggregates induced by G93A mutation in hSODG93A-NSC34 cells [38]. As BL-918 was proved to induce autophagy in hSODG93A-NSC34 cells, we wonder if BL-918 could achieve its therapeutic effects through inducing cytoprotective autophagy and eliminating mutant SOD aggregates. NSC34 cells transfected with hSOD-G93A-mCherry showed more SOD1 aggregates than NSC34 cells (Supplementary Fig. S4), which can be reduced by 10 μM BL-918 treatment, accompanied with increased rate of LC3-II/LC3-I, indicating enhanced autophagy level. Moreover, results of MTT assay and crystal violet staining showed that BL-918 has no toxicity and promotes the proliferation of hSODG93A-NSC34 (Supplementary Figs. S5, S6). Additionally, rapamycin-induced autophagy may also decrease SOD1 aggregates (Fig. 2c). The immunofluorescence image further demonstrated that BL-918 treatment decreased SOD1 in hSODG93A-NSC34 cells, whose effect could be partially reversed by 3-MA (Fig. 2d). Additionally, the colocalization of LC3 puncta and SOD1 was enhanced in BL-918 group, with the pearson’s R value [39] of 0.7, which indicates high correlation of SOD1 elimination and LC3 accumulation (Fig. 2e). Taken together, BL-918 induces cytoprotective autophagy to eliminate toxic SOD1 aggregates in hSODG93A-NSC34 cells.
a, b Stably expressing cell lines of hSOD1WT-mCherry-NSC34 and hSOD1G93A -mCherry-NSC34 were treated with 5, 10 μM BL-918 or treated without serum for 24 h. The levels of hSOD1 monomer were examined by Western blot. c NSC34 cells transient transfected with hSOD1G93A-mCherry were treated with 10 μM BL-918 or 50 nM rapamycin for 24 h. The levels of SOD1 aggregates, SOD1 monomer, p62, LC3-I, and LC3-II were analyzed by Western blot. d Representative images showing that BL-918 significantly decrease the accumulation of SOD1, while the effect may be reversed by 3-MA. Scale bar: 5 μm. e Fluorescent microscope revealed that the aggregation of SOD1 in cytoplasm (red) colocalized with LC3 (green) and decreased after BL-918 treatment. Scale bar: 5 μm. ns not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. ##P < 0.05 vs BL-918+3-MA.
BL-918 induces ULK1-dependent autophagy in vitro
To explore how BL-918 induced autophagy, we conducted structure-based molecular docking of BL-918 and ULK1 (Fig. 3a, b). Importantly, ULK1 can be inhibited by mTOR through Ser757 phosphorylation [18, 40] and activated by AMPK through Ser317 and Ser555 phosphorylation [18, 41]. Thus, we detected the expressions of ULK1, p-ULK1 to determine whether BL-918 evokes autophagy through a ULK1-dependent way in hSODG93A-NSC34 cells. As demonstrated by Western blot, the expression of ULK1 shows no change whereas the expression of p-ULK1 (Ser757) decreased and p-ULK1 (Ser555) and p-ULK1 (Ser317) increased (Fig. 3c) in a time-dependent manner. Besides, we observed that the expression of p-ULK1 (Ser555) and p-ULK1 (Ser317) at 36 h BL-918 treatment was decreased than that at 24 h treatment, which may be attributed to that the phosphorylation of kinase is an instantaneous event. Next, we examined the expressions of its upstream proteins after BL-918 treatment to investigate whether BL-918 activates ULK1 through mTOR- or AMPK-dependent pathways. mTOR is phosphorylated and inhibited at Ser2448, relieving its suppression of autophagy [42,43,44,45]. The expression of mTOR shows no changes, while p-mTOR (Ser2448) decreased markedly, which is similar to the alterations of p-ULK1 (Ser757), indicating possible feedback modulation of ULK1 to mTOR. To further confirm the targeted effect of BL-918 is independent on mTOR, we knocked down mTOR and found BL-918 still evoked autophagy (Supplementary Fig. S7). Additionally, the expressions of AMPK and p-AMPK, display no changes, suggesting that BL-918 may activate ULK1 without affecting AMPK to induce autophagy in hSODG93A-NSC34 cells (Fig. 3d). Also, ULK1 forms ULK1 complex with mATG13, FIP200, and ATG101 to initiate autophagy. And beclin1 plays an important role in the formation of Beclin 1-Vps34-Vps15 core complexes, promoting autophagy [46,47,48]. Therefore, downstream proteins of ULK1, including beclin1, p-beclin1, FIP200, ATG101 and p-mATG13 were analyzed to confirm that BL-918 triggers ULK1-mediated autophagy. The level of p-beclin1, beclin1, p-mATG13 and ATG101 increased in a time-dependent manner after BL-918 treatment (Fig. 3e, P < 0.001). To further confirm that BL-918 induces autophagy through a ULK1-dependent way, we used shRNA to knockdown ULK1 in hSODG93A-NSC34 cells (Supplementary Fig. S8). BL-918 treatment failed to induce autophagy in ULK1-knockdown cells, while autophagy in cells transfected with blank vector was not affected (Fig. 3f). Consistent with that, the expressions of the downstream autophagic proteins of ULK1, such as beclin1, ATG101, mATG13 and LC3 were not increased by BL-918 treatment in ULK1-knockdown cells (Fig. 3g). These results indicate that BL-918 induces autophagy in a ULK1-dependent way. Moreover, SOD1 aggregates failed to be eliminated in ULK1-knockdown hSODG93A-NSC34 cells after BL-918 treatment (Fig. 3f), accompanied by decreased autophagy level, which were demonstrated by decreased expression of LC3-II and beclin1 (Fig. 3g). This suggests that BL-918 evokes ULK1-dependent autophagy to eliminate toxic SOD aggregates. Collectively, these results demonstrate that the cytoprotective autophagy induced by BL-918 in hSODG93A-NSC34 cells is dependent on ULK1.
a Structure of BL-918. b Binding mode of BL-918 and ULK1. c Western blot of ULK1 and related phosphorylated ULK1 proteins in hSODG93A-NSC34 cells treated with BL-918 for 0 h, 6 h, 12 h, 24 h, or 36 h. d Western blot of upstream proteins of ULK1 in hSODG93A-NSC34 cells. e Western blot of downstream proteins of ULK1 in hSODG93A-NSC34 cells treated with BL-918 for 0 h, 6 h, 12 h, 24 h, or 36 h. f Immunofluorescence images showing the elimination of SOD1 in cells transfected with si-NC or sh-ULK1. Scale bar: 15 μm. g Western blot of SOD1 and downstream proteins of ULK1 in hSODG93A-NSC34 cells transfected with si-NC or sh-ULK1. ns not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Analysis of the effect of BL918 in hSODG93A-NSC34 cells based on transcriptome sequencing
To further explore the mechanism of BL-918 on treating ALS, we conducted RNA-seq on hSODG93A-NSC34 cells after the treatment of 10 μM BL-918 to examine the altered gene expression. Based on transcriptome sequencing, 2765 genes of BL-918 samples displayed altered expression compared to NC samples in total. Among them, 1249 were upregulated and 1516 were downregulated (P < 0.05, FDR < 0.05 and FC > 2 or FC < 0.5; Fig. 4a). Notably, through gene ontology (GO) annotation analysis of the differentially expressed genes, 4 autophagy-related terms were enriched (Fig. 4b). Then, a heat map shown the list of genes enriched by the autophagy, and the all 33 genes were upregulated, indicating the induction of autophagy (Fig. 4c). Since we have demonstrated that BL-918 could induce ULK1-dependent autophagy in hSODG93A-NSC34 cells, we constructed protein-protein interaction (PPI) network of ULK1 from altered expression of gene to seek for potentially new mechanisms of BL-918 on treating ALS (Fig. 4d). As a result, 105 downregulated genes and 90 upregulated genes were interacted with ULK1, and all of autophagy genes were upregulated, indicating that ULK1 may regulate these genes to induce autophagy. Subsequently, we simply validate the results of RNA-seq with qPCR (Fig. 4e–j), and found except ANXA7 and AMBRA1, other autophagy-associated genes including ATG2A, ATG16L, HSPA8, GABARAPL1, which were also identified interacted with ULK1 in the PPI network, displayed consistently positive regulation after BL-918 treatment as the analysis indicated. These results suggest that BL-918 may induce ULK1 dependent autophagy through these downstream genes.
a Volcano plot of differentially expressed genes in hSODG93A-NSC34 cells treated with or without 10 μM BL-918. Red plots represent padj <0.05 and fold change >2; blue plots represent padj <0.05 and fold change <0.5. b Bubble plot of GO enriched by the differentially expressed genes. c Heatmap of autophagy-related genes with significant different expression in hSODG93A-NSC34 cells treated with or without 10 μM BL-918. d PPI network of ULK1. Upregulated genes are colored in red, in which 7 of them associated with autophagy are colored purple, while downregulated genes are colored blue. e–j The expression of AMBRA1, ANXA7, ATG2A, ATG16L2, GABARAPL1, and HSPA8 in hSODG93A-NSC34 cells treated with or without 10 μM BL-918 were measured by PCR
Pharmacokinetic (PK) profiles of BL-918
As BL-918 was identified as an activator of autophagy by promoting phosphorylation of ULK1, we investigated its pharmacokinetic profiles to evaluate its ability to penetrate blood-brain barrier. Rats are intragastrically administered 50 mg/kg BL-918 and samples were collected and analyzed by UPLC-Q-TOF-MS. MetabolynxTM data processing software is used to analyze the data from UPLC-Q-TOF-MS, while potential metabolites are screened by precise molecular weights and MS/MS secondary chromatography for fragmentation mode analysis. A total of 18 xenobiotics were tested in all the samples and only three metabolites were present in spinal cord and brain, indicating that BL-918 and its metabolites may penetrate blood-brain barrier and holds potential in treating neurodegenerative diseases such as ALS. Moreover, we demonstrated the three metabolites could promote the proliferation of hSODG93A-NSC34 cells, without significant toxicity even at high concentration (Supplementary Figs. S5, S6). Furthermore, based on the analysis of the metabolites of BL-918, we speculated that it undergoes oxidation, hydrolysis, dehydrogenation, glucuronidation, acetylation, sulfonation and other reactions in vivo (Fig. 5a). Next, we detected the blood concentration of BL-918 (M0), M8 and M10 and use non-compartmental model to calculate the pharmacokinetic parameters of these compound, including t1/2, Tmax, the area under the ROC curve (AUC0-t), mean residence time (MRT0-t), and absolute bioavailablity (F) (Fig. 5b-i). As a result, we found BL-918 reached the highest blood concentration after both intragastric and intravenous administration compared to M8 and M10, with t1/2 of 18.0 ± 1.6 h, Tmax at 10.7 ± 0.4 h, MRT0-t at 28.1 ± 0.3 h and F of 23.5% (Tables 1, 2), suggesting that the exposure of the prototype component (BL-918) in vivo is higher than its metabolites (M8 and M10).
a The potential proposed metabolic pathways of BL-918 (M0) in rat. b Total concentration-time profiles of M0, M8 and M10 when rats were administered intragastrically. c Concentration-time profiles of M0 when rats were administered intragastrically. d Total concentration-time profiles of M0, M8, and M10 when rats were injected intravenously. e Concentration-time profiles of M0 when rats were injected intravenously. f Concentration-time profiles of M8 when rats were administered intragastrically. g Concentration-time profiles of M10 when rats were administered intragastrically. h Concentration-time profiles of M8 when rats were injected intravenously. i Concentration-time profiles of M10 when rats were injected intravenously.
BL-918 displays a therapeutic potential on SODG93A mice
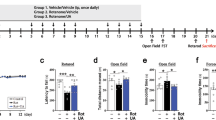
As BL-918 and its metabolites can penetrate brain-blood barrier, we speculated that BL-918 may display good efficacy on treating ALS in vivo. Thus, we investigated its therapeutic potential on ALS using SODG93A transgenic mice. Mice were randomly divided into four groups: (1) WT-NT (2) TG-NT (3) TG-BL-918-L (4) TG-BL-918-H and treated as methodology indicated and assessed with motor functions by behavioral experiments as well as recorded with life spans, weights and food intake [49]. As mice age, their motor functions degenerated, reflected by shorter time in rotarod test and hanging wire test and lower scores in neuro scoring. Notably, BL-918 treatment significantly delayed the loss of motor functions of TG-BL-918-L and TG-BL-918-H mice than that of TG-NT mice (Fig. 6a–c). Also, TG-BL-918-L and TG-BL-918-H mice showed increased life spans than TG-NT mice, with average of 288.5 ± 0.645 days of TG-BL-918-L mice and 293.9 ± 6.056 days of TG-BL-918-H mice (Fig. 6e–g). Additionally, food intake and weights of TG-BL-918-L and TG-BL-918-H mice increased markedly than TG-NT mice. These alterations indicated that BL-918 may exert therapeutic effects on SODG93A transgenic mice. Next, we wonder whether BL-918 could attenuate the loss of motor neurons and the muscular atrophy of SOD1G93A mice, which happened in the late stage of ALS [50, 51]. Therefore, we quantified the number of motor neurons in ventral horn and spinal cord of SOD1G93A mice at the end stage of disease and analyzed the formative changes by staining gastrocnemius muscles of WT and BL-918-treated mice with Masson’s trichrome. More motor neurons and less significant muscular atrophy were discovered in TG-BL-918-L and TG-BL-918L-H mice, while the TG-NT mice showed motor neurons degeneration and more severe muscular atrophy (Fig. 6d, j), consistent with the conclusion that BL-918 potentially ameliorates symptoms of SOD1G93A mice.
a BL-918 improves motor function assessed by rotarod test in SOD1G93A mice. The numbers of animals were as follows: WT-NT, n = 19; TG-NT, n = 10; TG-BL-918-L, n = 13; TG-BL-918-H, n = 14. Data are presented as mean ± SEM. The data were analyzed using two-way ANOVA with LSD post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001 vs. the TG-NT group. b BL-918 improves motor function assessed by hanging wire test in SOD1G93A mice. The numbers of animals were as follows: WT-NT, n = 19; TG-NT, n = 10; TG-BL-918-L, n = 13; TG-BL-918-H, n = 14. Data are presented as mean ± SEM. The data were analyzed using two-way ANOVA with LSD post hoc test. *P < 0.05 vs. the TG-NT group. c BL-918 improves motor function assessed by neurological score in SOD1G93A mice. The numbers of animals were as follows: WT-NT, n = 19; TG-NT, n = 10; TG-BL-918-L, n = 13; TG-BL-918-H, n = 14. Data are presented as mean ± SEM. The data were analyzed using two-way ANOVA with LSD post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001 vs. the TG-NT group. d A motor neuron survival was determined in the ventral horn of the lumbar spinal cord after Nissl staining from wild-type (WT), vehicle (NT), or BL-918 (40 mg/kg and 80 mg/kg) treated SOD1G93A mice. The yellow arrows indicate large (>25 μm) and intact mortor neurons in the ventral horn of spinal cord. The scale bar of upper picture is 500 μm. Higher magnification is shown in the below figure, scale bar = 200 μm. Motor neuron numbers showing that BL-918 treatment significantly attenuated neurodegeneration at end stage when compared with TG-NT group. Data were presented as mean ± SEM. The data were analyzed using independent-samples t test. **P < 0.01, ***P < 0.001 as compared with TG-NT group. The numbers of animals were as follows: WT-NT, n = 5; TG-NT, n = 4; TG-BL-918-L, n = 5; TG-BL-918-H, n = 3. e The results of Kaplan–Meier survival analysis showed the probability of survival in WT-NT, TG-NT, TG-BL-918-L and TG-BL-918-H mice. f, g The data of life span in TG-NT, TG-BL-918-L or TG-BL-918-H mice. There were 19 mice in the WT-NT group and 15 mice in each group of TG-NT, TG-BL-918-L and TG-BL-918-H group. The data were analyzed using two-way ANOVA with Dunnett’s post-hoc test. Data are presented as mean ± SEM. ***P < 0.001 as compared with TG-NT group. h Food intake in WT or SOD1G93A mice treated with BL-918 (40 mg/kg or 80 mg/kg) or vehicle. Data are presented as mean ± SEM. The data were analyzed using two-way ANOVA with LSD post hoc test. i The relative body weight of the mice of the indicated groups including WT-NT (n = 19), TG-NT (n = 10), TG-BL-918-L (n = 13), TG-BL-918-H (n = 14). *P < 0.05, **P < 0.01 vs the TG-NF group. j Transverse sections of gastrocnemius muscles from WT mice and vehicle or BL-918-treated transgenic mice were stained with Masson’s trichrome. Scale bar = 200 μm
BL-918 ameliorates ALS by inducing cytoprotective autophagy to eliminate SOD aggregates in vivo
To evaluate whether the amelioration of ALS symptoms in SODG93A mice and the clearance of SOD1 aggregates in vivo are associated with the induction of autophagy by BL-918, we examined the expression of autophagy protein in mice spinal cord and cerebral cortex (Fig. 7a, b). As was shown in the picture, the decreased expression of p62 and the increased rate of LC3-II/LC3-I were found in mice from spinal cord and cerebral cortex, demonstrating the induction of autophagy. Interestingly, the expression of ULK1 increased both in spinal cord and cerebral cortex, which may come down to the effects of endogenous substances. Consistent with the alterations in vitro, phosphorylation of ULK1 at Ser757 decreased while that at Ser555 and Ser317 increased in both spinal cord and cortex of SOD1G93A mice, demonstrating the activation of ULK (Fig. 7c, d). Notably, the accumulation of SOD1 in the spinal cord and cortex of SODG93A mice decreased after either 5 μM or 10 μM of BL-918 treatment (Fig. 7e, f), suggesting that toxic SOD1 aggregates were degraded after BL-918 treatment. Altogether, these results demonstrate that BL-918 exerts its protective effect on SODG93A mice by inducing cytoprotective autophagy.
a, b BL-918 induces the expression of autophagy-related proteins in the spinal cord and cortex at end stage of SOD1G93A mice. The protein level of ULK1, LC3, SQSTM1/p62 and ChAT of mice were analyzed by Western blot. GAPDH was used as loading control. Quantitative analysis of ULK1, SQSTM1/p62, ChAT to GAPDH and the ratio of LC3-II/LC3-I (n = 3, one-way ANOVA with LSD post-hoc test was used for statistical analysis). c, d The protein level of p-ULK1, p-mTOR and Beclin of mice were analyzed by Western blot. β-Actin was used as loading control. Quantitative analysis of p-ULK1, p-mTOR and Beclin to β-actin. e, f BL-918 enhances SOD1 aggregate clearance in the spinal cord and cortex at end stage of SOD1G93A mice. The protein level of SOD1 aggregate was analyzed by Western blot. GAPDH was used as loading control. Quantitative analysis of SOD1 aggregates to GAPDH (n = 3, one-way ANOVA with Bonferroni’s multiple comparisons test). Data were expressed as mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. ns not significant
Discussion
Autophagic defects are one of the pathological features of ALS, partially stemming from abundant ALS-linked gene mutations, including genes encoding for core autophagy proteins, for functional proteins. Moreover, some misfolded proteins of ALS-causing gene mutations, such as SOD1, aggravating autophagic stress [52]. Given the impaired autophagy in ALS, the induction of autophagy is emerging as a potential therapy for ALS.
Autophagy is regulated by both mTOR-dependent and mTOR-independent pathways. Previous study has reported aggravated neuronal loss and disease progression with the treatment of rapamycin, a mTOR-dependent autophagy activator, indicating the activation of mTOR-dependent autophagy may have some off-targets side effects on ALS, such as activated apoptosis [32]. mTOR prevents ULK1 activation by phosphorylating Ser757 under glucose sufficiency, thereby inhibiting autophagy [53]. Moreover, lithium was previously shown to prolong the mean survival time and delay disease duration of SODG93A mice and facilitate the clearance of SOD aggregates in motor neurons as a mTOR-independent autophagy enhancer [54]. However, the multiple-targeted lithium may exert additional effects, such as an increase of motor neurons adhesion and decrease of mitochondria, which is detrimental to the disease progression. Trehalose, another mTOR-independent small-molecule activator of autophagy, increases life span and delays disease onset of SOD1G86R mice possibly through promoting the transcription of autophagy-regulatory genes and the activation of FOXO1. Despite that there are no reports of detrimental effects of trehalose, the exact mechanism of its mTOR-independent induction of autophagy remains a mystery and may cause unexpected events. Moreover, carbamazepine was recently reported to delay the disease onset and extend the life span of SODG93A mice, as well as ameliorating the altered muscle morphology and reducing MNs loss. Autophagy induced by AMPK-ULK1 pathway was involved in eliminating mutant SOD aggregates, representing a therapeutic potential of autophagy enhancers in ALS [55].
Considering the adverse effects of inducing the mTOR-dependent autophagy, we assume that directly activating ULK1 and evoking autophagy may be of great advantage to treat ALS. Thus, in the present study, we investigated the therapeutic effect of BL-918, an autophagy activator targeting ULK1, on ALS. And BL-918 treatment eliminates toxic SOD1 aggregates through inducing ULK1-dependent cytoprotective autophagy in hSODG93A-NSC34 cells. In the present study, BL-918 increased Ser317 and Ser555 phosphorylation of ULK1, as well as reduced Ser757 phosphorylation of ULK1 and activated autophagy, which is reflected by augmented level of autophagic proteins LC3-II and autophagic flux and formation of autophagosomes. In addition, BL-918 failed to induce autophagy in ULK1-knockdown cells, suggesting the induced autophagy by BL-918 is ULK1-dependent. Moreover, the upstream regulators of ULK1, mTOR and AMPK and their phosphorylation forms have not changed after BL-918 treatment, which demonstrated that BL-918 could induce autophagy without affecting mTOR pathway or AMPK pathway, thus, it may avoid some side effects due to suppressed mTOR pathway and activated AMPK pathway.
One challenge in treating neurodegenerative disease with small molecules is that brain-blood barrier (BBB) prevents most drugs from entering brain tissues. Thus, we evaluated the Pharmacokinetic (PK) profiles of BL-918 and confirmed the protodrug and its metabolites M8 and M10 can penetrate BBB and exert their therapeutic effects, indicating good druggability of BL-918. Next, we confirmed the effect of BL-918 in vivo, and found that life span and quality of SODG93A mice have been improved after BL-918 treatment while toxic SOD aggregates were decreased through induction of autophagy, demonstrating by augmented autophagic flux and increased expression of autophagic proteins.
Importantly, G4C2 repeat expansion in the C9orf72 causes the most common forms of familial amyotrophic lateral sclerosis (ALS), and C9orf72 was reported to regulate autophagy via regulating ULK1 complex [56, 57], establishing associations between autophagy and C9-ALS [58]. However, no strategies targeting autophagy to treat C9-ALS have been reported. Furthermore, ULK1 was found interdependent with TBK1 in phosphorylating mitophagy receptor optineurin (OPTN), which may serve as a complementary way when ALS-linked TBK1 mutations disturb the clearance of damaged mitochondria by mitophagy [59]. Our study has confirmed the therapeutic effect of the autophagy enhancer, BL-918, on SOD mutant ALS models. In follow-up studies, attempts on treating C9-ALS or ALS harboring TBK1 mutations with ULK1 activator BL-918 representing a promising direction and warrant further investigation.
In summary, using hSODG93A-NSC34 cells and SODG93A mice, we investigate the therapeutic role and mechanism of ULK1 activator BL-918, an autophagy activator previously reported in Parkinson’s disease, in treating ALS. BL-918 treatment prolongs lifespan and improves the motor function of SODG93A mice, and enhances the clearance of SOD1 aggregates in spinal cord and cerebral cortex through inducing autophagy. Moreover, BL-918 treatment eliminates toxic SOD1 aggregates through inducing ULK1-dependent cytoprotective autophagy in hSODG93A-NSC34 cells. Collectively, ULK1 activator BL-918 displays a therapeutic potential on ALS through the induction of cytoprotective autophagy and provides a further clue for autophagic dysfunction in ALS pathogenesis.
References
Tandan R, Bradley WG. Amyotrophic lateral sclerosis: Part 1. Clinical features, pathology, and ethical issues in management. Ann Neurol. 1985;18:271–80.
van Es MA, Hardiman O, Chio A, Al-Chalabi A, Pasterkamp RJ, Veldink JH, et al. Amyotrophic lateral sclerosis. Lancet. 2017;390:2084–98.
Valdmanis PN, Rouleau GA. Genetics of familial amyotrophic lateral sclerosis. Neurology. 2008;70:144–52.
Al-Chalabi A, van den Berg LH, Veldink J. Gene discovery in amyotrophic lateral sclerosis: implications for clinical management. Nat Rev Neurol. 2017;13:96–104.
Tu LF, Zhang TZ, Zhou YF, Zhou QQ, Gong HB, Liang L, et al. GPX4 deficiency-dependent phospholipid peroxidation drives motor deficits of ALS. J Adv Res. 2022; https://doi.org/10.1016/j.jare.2022.02.016.
Renton AE, Chiò A, Traynor BJ. State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci. 2014;17:17–23.
Bruijn LI, Houseweart MK, Kato S, Anderson KL, Anderson SD, Ohama E, et al. Aggregation and motor neuron toxicity of an ALS-linked SOD1 mutant independent from wild-type SOD1. Science. 1998;281:1851–4.
Chattopadhyay M, Valentine JS. Aggregation of copper-zinc superoxide dismutase in familial and sporadic ALS. Antioxid Redox Signal. 2009;11:1603–14.
Kaur SJ, McKeown SR, Rashid S. Mutant SOD1 mediated pathogenesis of amyotrophic lateral sclerosis. Gene. 2016;577:109–18.
Dal Canto MC, Gurney ME. Neuropathological changes in two lines of mice carrying a transgene for mutant human Cu, Zn SOD, and in mice overexpressing wild type human SOD: a model of familial amyotrophic lateral sclerosis (FALS). Brain Res. 1995;676:25–40.
Tokuda E, Brännström T, Andersen PM, Marklund SL. Low autophagy capacity implicated in motor system vulnerability to mutant superoxide dismutase. Acta Neuropathol Commun. 2016;4:6.
Van Damme P, Robberecht W, Van Den Bosch L. Modelling amyotrophic lateral sclerosis: progress and possibilities. Dis Model Mech. 2017;10:537–49.
Bendotti C, Marino M, Cheroni C, Fontana E, Crippa V, Poletti A, et al. Dysfunction of constitutive and inducible ubiquitin-proteasome system in amyotrophic lateral sclerosis: implication for protein aggregation and immune response. Prog Neurobiol. 2012; 97:101–26.
Robberecht W, Philips T. The changing scene of amyotrophic lateral sclerosis. Nat Rev Neurosci. 2013;14:248–64.
Liu W, Wang G, Wang Z, Wang G, Huang J, Liu B. Repurposing small-molecule drugs for modulating toxic protein aggregates in neurodegenerative diseases. Drug Discov Today. 2022;27:1994–2007.
Nijholt DA, De Kimpe L, Elfrink HL, Hoozemans JJ, Scheper W. Removing protein aggregates: the role of proteolysis in neurodegeneration. Curr Med Chem. 2011;18:2459–76.
Ravikumar B, Sarkar S, Davies JE, Futter M, Garcia-Arencibia M, Green-Thompson ZW, et al. Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev. 2010;90:1383–435.
Kim J, Kundu M, Viollet B, Guan KL. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13:132–41.
Hara T, Takamura A, Kishi C, Iemura S, Natsume T, Guan JL, et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J Cell Biol. 2008;181:497–510.
Hosokawa N, Hara T, Kaizuka T, Kishi C, Takamura A, Miura Y, et al. Nutrient-dependent mTORC1 association with the ULK1-Atg13-FIP200 complex required for autophagy. Mol Biol Cell. 2009;20:1981–91.
Hosokawa N, Sasaki T, Iemura S, Natsume T, Hara T, Mizushima N. Atg101, a novel mammalian autophagy protein interacting with Atg13. Autophagy. 2009;5:973–9.
Nah J, Yuan J, Jung YK. Autophagy in neurodegenerative diseases: from mechanism to therapeutic approach. Mol Cells. 2015;38:381–9.
Nassif M, Hetz C. Targeting autophagy in ALS: a complex mission. Autophagy. 2011;7:450–3.
Djajadikerta A, Keshri S, Pavel M, Prestil R, Ryan L, Rubinsztein DC. Autophagy induction as a therapeutic strategy for neurodegenerative diseases. J Mol Biol. 2020;432:2799–821.
Sun J, Mu Y, Jiang Y, Song R, Yi J, Zhou J, et al. Inhibition of p70 S6 kinase activity by A77 1726 induces autophagy and enhances the degradation of superoxide dismutase 1 (SOD1) protein aggregates. Cell Death Dis. 2018;9:1–16.
Granatiero V, Sayles NM, Savino AM, Konrad C, Kharas MG, Kawamata H, et al. Modulation of the IGF1R-MTOR pathway attenuates motor neuron toxicity of human ALS SOD1G93A astrocytes. Autophagy. 2021;17:4029–42.
Galluzzi L, Bravo-San Pedro JM, Levine B, Green DR, Kroemer G. Pharmacological modulation of autophagy: therapeutic potential and persisting obstacles. Nat Rev Drug Discov. 2017;16:487–511.
Ouyang L, Zhang L, Zhang S, Yao D, Zhao Y, Wang G, et al. Small-molecule activator of UNC-51-like kinase 1 (ULK1) that induces cytoprotective autophagy for Parkinson’s disease treatment. J Med Chem. 2018;61:2776–92.
Peng J, Pan J, Mo J, Peng Y. MPO/HOCl facilitates apoptosis and ferroptosis in the SOD1G93A motor neuron of amyotrophic lateral sclerosis. Oxid Med Cell Longev. 2022;2022:8217663.
Karch CM, Prudencio M, Winkler DD, Hart PJ, Borchelt DR. Role of mutant SOD1 disulfide oxidation and aggregation in the pathogenesis of familial ALS. Proc Natl Acad Sci USA. 2009;106:7774–9.
Günther R, Suhr M, Koch JC, Bähr M, Lingor P, Tönges L. Clinical testing and spinal cord removal in a mouse model for amyotrophic lateral sclerosis (ALS). J Vis Exp. 2012;61:3936.
Zhang X, Li L, Chen S, Yang D, Wang Y, Zhang X, et al. Rapamycin treatment augments motor neuron degeneration in SOD1 G93A mouse model of amyotrophic lateral sclerosis. Autophagy. 2011;7:412–25.
Lucocq JM, Hacker C. Cutting a fine figure: On the use of thin sections in electron microscopy to quantify autophagy. Autophagy. 2013;9:1443–8.
Eskelinen EL, Reggiori F, Baba M, Kovács AL, Seglen PO. Seeing is believing: the impact of electron microscopy on autophagy research. Autophagy. 2011;7:935–56.
Klionsky DJ, Abdel-Aziz AK, Abdelfatah S, Abdellatif M, Abdoli A, Abel S, et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition)1. Autophagy. 2021;17:1–382.
Lovejoy DB, Guillemin GJ. The potential for transition metal-mediated neurodegeneration in amyotrophic lateral sclerosis. Front Aging Neurosci. 2014;6:173.
Pickles S, Semmler S, Broom HR, Destroismaisons L, Legroux L, Arbour N, et al. ALS-linked misfolded SOD1 species have divergent impacts on mitochondria. Acta Neuropathol Commun. 2016;4:43.
Woo TG, Yoon MH, Kang SM, Park S, Cho JH, Hwang YJ, et al. Novel chemical inhibitor against SOD1 misfolding and aggregation protects neuron-loss and ameliorates disease symptoms in ALS mouse model. Commun Biol. 2021;4:1397.
Lai CS, Tao Y, Xu F, Ng WWY, Jia Y, Yuan H, et al. A robust correlation analysis framework for imbalanced and dichotomous data with uncertainty. Inf Sci. 2019;470:58–77.
Shang L, Chen S, Du F, Li S, Zhao L, Wang X. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc Natl Acad Sci USA. 2011;108:4788–93.
Egan DF, Shackelford DB, Mihaylova MM, Gelino S, Kohnz RA, Mair W, et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science. 2011;331:456–61.
Chiang GG, Abraham RT. Phosphorylation of mammalian target of rapamycin (mTOR) at Ser-2448 is mediated by p70S6 kinase. J Biol Chem. 2005; 280: 25485–90.
Xu Z, Han X, Ou D, Liu T, Li Z, Jiang G, et al. Targeting PI3K/AKT/mTOR-mediated autophagy for tumor therapy. Appl Microbiol Biotechnol. 2020;104:575–87.
Querfurth H, Lee HK. Mammalian/mechanistic target of rapamycin (mTOR) complexes in neurodegeneration. Mol Neurodegener. 2021;16:44.
Figueiredo VC, Markworth JF, Cameron-Smith D. Considerations on mTOR regulation at serine 2448: implications for muscle metabolism studies. Cell Mol Life Sci. 2017;74:2537–45.
Zhang L, Ouyang L, Guo Y, Zhang J, Liu B. UNC-51-like kinase 1: from an autophagic initiator to multifunctional drug target. J Med Chem. 2018;61:6491–500.
Kang R, Zeh HJ, Lotze MT, Tang D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011;18:571–80.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402:672–6.
Weydt P, Hong SY, Kliot M, Möller T. Assessing disease onset and progression in the SOD1 mouse model of ALS. NeuroReport. 2003;14:1051.
Narasimhan K. Quantifying motor neuron loss in ALS. Nat Neurosci. 2006;9:304–304.
Gurney ME, Pu H, Chiu AY, Dal Canto MC, Polchow CY, Alexander DD, et al. Motor neuron degeneration in mice that express a human Cu, Zn superoxide dismutase mutation. Science. 1994;264:1772–5.
Amin A, Perera ND, Beart PM, Turner BJ, Shabanpoor F. Amyotrophic lateral sclerosis and autophagy: dysfunction and therapeutic targeting. Cells. 2020;9:2413.
Xiang H, Zhang J, Lin C, Zhang L, Liu B, Ouyang L. Targeting autophagy-related protein kinases for potential therapeutic purpose. Acta Pharm Sin B. 2020;10:569–81.
Fornai F, Longone P, Cafaro L, Kastsiuchenka O, Ferrucci M, Manca ML, et al. Lithium delays progression of amyotrophic lateral sclerosis. Proc Natl Acad Sci USA. 2008;105:2052–7.
Zhang JJ, Zhou QM, Chen S, Le WD. Repurposing carbamazepine for the treatment of amyotrophic lateral sclerosis in SOD1-G93A mouse model. CNS Neurosci Ther. 2018;24:1163–74.
Yang M, Liang C, Swaminathan K, Herrlinger S, Lai F, Shiekhattar R, et al. A C9ORF72/SMCR8-containing complex regulates ULK1 and plays a dual role in autophagy. Sci Adv. 2016;2:1–18.
Sullivan PM, Zhou X, Robins AM, Paushter DH, Kim D, Smolka MB, et al. The ALS/FTLD associated protein C9orf72 associates with SMCR8 and WDR41 to regulate the autophagy-lysosome pathway. Acta Neuropathol Commun. 2016;4:51.
Beckers J, Tharkeshwar AK, Van Damme P. C9orf72 ALS-FTD: recent evidence for dysregulation of the autophagy-lysosome pathway at multiple levels. Autophagy. 2021;17:3306–22.
Harding O, Evans CS, Ye J, Cheung J, Maniatis T, Holzbaur ELF. ALS- and FTD-associated missense mutations in TBK1 differentially disrupt mitophagy. Proc Natl Acad Sci USA. 2021;118:e2025053118.
Acknowledgements
We are grateful to Prof Sheng-yong Yang and Prof Can-hua Huang (Sichuan University) for their critical reviews on this manuscript. We thank Prof Ying Peng (Sun Yat-sen University) for providing us with NSC34 cells and hSODG93A-NSC34 cells. This work was supported by grants from National Natural Science Foundation of China (Grant No. 81873209, 82172649, 21314120 and 81922064), Key R&D Program of Sichuan Province (Grant No. 2021YFS0046), the Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program (grant number 2017BT01Y036) and GDUPS; Research Foundation for Administration of traditional Chinese medicine of Sichuan Province (Grant No. 2020HJZX002); CAMS Innovation Fund for Medical Science (Grant 2019-RC-HL-023); National Clinical Research Center for Geriatrics, West China Hospital, Sichuan University (Grant Z20201004); and Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (grant 2020-JKCS-014).
Author information
Authors and Affiliations
Contributions
WL, SOZ, YLG and LFT contributed equally to this work. LOY, BL and RRH designed the research. WL, SOZ,YLG, LFT, YQZ and RYZ performed all the experiments of this research. WL, SOZ and YLG contributed to the statistical analyses and interpretation of the results. WL, SOZ and LFT wrote the manuscript. HK, LOY, RRH and BL revised the paper. All authors edited and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, W., Zhu, So., Guo, Yl. et al. BL-918, a small-molecule activator of ULK1, induces cytoprotective autophagy for amyotrophic lateral sclerosis therapy. Acta Pharmacol Sin 44, 524–537 (2023). https://doi.org/10.1038/s41401-022-00972-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-022-00972-w
Keywords
This article is cited by
-
In Vitro Models of Amyotrophic Lateral Sclerosis
Cellular and Molecular Neurobiology (2023)