Abstract
Protocadherin-19 (PCDH19) developmental and epileptic encephalopathy causes an early-onset epilepsy syndrome with limbic seizures, typically occurring in clusters and variably associated with intellectual disability and a range of psychiatric disorders including hyperactive, obsessive-compulsive and autistic features. Previous quantitative neuroimaging studies revealed abnormal cortical areas in the limbic formation (parahippocampal and fusiform gyri) and underlying white-matter fibers. In this study, we adopted morphometric, network-based and multivariate statistical methods to examine the cortex and substructure of the hippocampus and amygdala in a cohort of 20 PCDH19-mutated patients and evaluated the relation between structural patterns and clinical variables at individual level. We also correlated morphometric alterations with known patterns of PCDH19 expression levels. We found patients to exhibit high-significant reductions of cortical surface area at a whole-brain level (left/right pvalue = 0.045/0.084), and particularly in the regions of the limbic network (left/right parahippocampal gyri pvalue = 0.230/0.016; left/right entorhinal gyri pvalue = 0.002/0.327), and bilateral atrophy of several subunits of the amygdala and hippocampus, particularly in the CA regions (head of the left CA3 pvalue = 0.002; body of the right CA3 pvalue = 0.004), and differences in the shape of hippocampal structures. More severe psychiatric comorbidities correlated with more significant altered patterns, with the entorhinal gyrus (pvalue = 0.013) and body of hippocampus (pvalue = 0.048) being more severely affected. Morphometric alterations correlated significantly with the known expression patterns of PCDH19 (rvalue = -0.26, pspin = 0.092). PCDH19 encephalopathy represents a model of genetically determined neural network based neuropsychiatric disease in which quantitative MRI-based findings correlate with the severity of clinical manifestations and had have a potential predictive value if analyzed early.
Similar content being viewed by others
Introduction
Protocadherin-19 (PCDH19) gene mutations cause an infantile-onset epilepsy syndrome with limbic seizures, typically occurring in clusters and variably associated with intellectual disability and a range of psychiatric disorders including autistic features [1]. Cognitive and affective dysfunctions in PCDH19 developmental and epileptic encephalopathy (DEE) impose a major medical and socio-economic burden and are increasingly recognized as core disease dimensions that can affect patients’ quality of life even more than the seizures themselves [2]. Kolc and colleagues [3] reviewed 271 PCDH19-variant individuals and found hyperactive, autistic and obsessive-compulsive features to be the most frequently observed neuropsychiatric manifestations, while no genotype-phenotype associations emerged in individuals with recurrent variants, or in the group overall. An earlier seizure onset was significantly associated with more severe intellectual disability, but no correlations were observed between the type or location of PCDH19 mutation, neuropsychiatric profile, and age at seizure onset.
PCDH19 is a calcium dependent cell-adhesion molecule involved in neuronal circuit formation. In individuals with PCDH19-gene mutations, gross human brain morphology appears usually normal on MRI [4], with mild, ill-defined malformations of the cortical mantle having been described in isolated patients [5]. However, quantitative neuroimaging reveals abnormal cortical areas in the limbic formation and underlying white-matter fibers [6]. In patients with earlier onset of seizures, worse autistic manifestations, and cognitive impairment, structural abnormalities in the limbic areas are more pronounced [6]. Using MRI, a correlation between abnormal mesio-temporal anatomy and autism has been reported in different etiological settings [7,8,9,10]. MRI has further disclosed altered patterns of brain expansion in children with autism [11], but a correlation between specific mutations and autism remains to be clarified in PCDH19-related phenotypes [3].
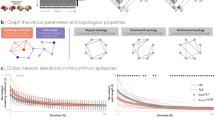
Developmental abnormalities involving the limbic structures, particularly the parahippocampal and entorhinal gyri, likely represent a measurable anatomic consequence of the altered expression of the PCDH19 protein on cortical folding and white matter organization in these anatomic areas [12], and are functionally reflected in the phenotypic features involving neuropsychiatric symptoms, cognitive and communicative skills and local epileptogenesis [3]. Recent studies on animal models have contributed to clarify how PCDH19 mutations alter brain network formation and maintenance [13], and have identified mechanisms by which focal mosaic mutations disrupt cortical networks [14] and impair neuronal activity and connectivity [15]. Exploring the structural organization of the limbic regions in PCDH19-related phenotypes could benefit from adopting network-based algorithms [16]. Such approaches, most often used to study atrophic patterns, may unveil broader inter-associations between brain regions, contributing to a deeper understanding of the underlying developmental relationships at a whole-brain level. Cortico-subcortical network alterations in epilepsy have been associated with genetic risk factors at large [17], but the links between neurobiological mechanisms and cortical features in single-gene developmental and epileptic encephalopathies remain poorly explored. Open databases focused on transcriptomic and proteomic provide a great resource for integrating macroscale neuroimaging observations with molecular-level spatial variances [18]. Syndrome-specific morphometric alterations may indeed reflect the spatial expression patters of PCDH19 and potentially give insight into the intricate interplay between neuroimaging data and molecular-scale information.
Neuroimaging research has moved on to disentangle the contributions of different hippocampal subregions and adjacent cortices, so as to bridge the gap between rodent and human data [19]. In previous quantitative MRI study of PCDH19-related encephalopathy [6], no volumetric alterations emerged in the hippocampus at a whole level, although surface patterns in the mesio-temporal cortex were abnormal. However, the hippocampus has a key role in limbic epilepsies [20], and PCDH19 has a prevalent expression in the hippocampal structures [21], particularly in the Cornu Ammonis (CA) [22]. Latest studies on murine models have highlighted that PCDH19 impairment contributes to develop structural and functional synaptic defects in the hippocampus [15].
In this study, we adopted morphometry and network-based models to detect abnormal structural patterns of brain regions in patients with PCDH19-related epilepsy, previously analyzed in the cortex and white matter, and comparatively studied patients’ subgroups stratified by different clinical severity at the individual level. We also adopted advanced neuroimaging methods to further investigate the substructure and shape of the hippocampus, CA and amygdala subunits, and evaluated the relation between structural patterns of abnormality and spatial expression of PCDH19. We found the limbic circuit to represent a structural epicenter of the anatomic alteration of the cortex and several hippocampal substructures to exhibit reduced volumes. We could also demonstrate that patients exhibiting significantly more severe atrophic patterns experienced more severe psychiatric comorbidities.
Patients and Methods
Participants
We studied a cohort of 20 patients seen at two tertiary pediatric neurology centers (Meyer Children’s Hospital IRCCS, 11 patients and 22 controls; IRCCS Bambino Gesù Pediatric Hospital, 9 patients and 18 controls). Patients (mean age 14.1 years, median 13.6, range 2.3-26.6) had been included in previous publications [1, 4, 6, 23, 24] and carried pathogenic PCDH19 gene mutations (canonical transcript NM_001184880.1) identified by next-generation sequencing, Sanger sequencing or multiplex ligation-dependent probe amplification. To mitigate confounding effects, we enrolled 40 controls that were matched with patients for acquisition center, sex and age ( ± 1year, mean age 14.2 years, median 13.4, range 3.4–27.0) and were healthy individuals who had received an MRI for uncomplicated headache.
Clinical and genetic data
For each patient, we collected clinical and genetic data using the REDCap (Research Electronic Data Capture) [25] tool hosted at Meyer Children’s Hospital IRCCS. Clinical data considered for the purpose of this study included age at seizure onset, seizure type, ictal and interictal EEG characteristics, neuropsychology testing and psychiatric manifestations (Table 1). At seizure onset (mean 10.8 months, median 8.0, range 4–28), 18 patients (90%) manifested focal seizures, two had focal to bilateral tonic and tonic-clonic seizures, while at follow-up 16 patients (80%) manifested more seizure types, i.e. focal, tonic, tonic-clonic and absences. The triggering factor for seizures was a febrile illness in 80% of patients. Ten patients presented focal seizures with prominent affective symptoms, suggesting an epileptogenic dysfunction involving the limbic system.
Formal cognitive assessment was performed according to age and verbal function, using one or more of the following neuropsychological tests: Brunet‐Lezine, Griffiths mental development scale, Bayley Scales of Infant and Toddler Development (Bayley‐III), Leiter International Performance Scale‐Revised (Leiter‐R), Raven’s Coloured Progressive Matrices, Stanford‐Binet test, Wechsler Preschool and Primary Intelligence test (WPPSI), Wechsler Intelligence Scale for Children (WISC), Wechsler Adult Intelligence Scale | Fourth Edition (WAIS‐IV), and Mini‐Mental State Examination (MMSE). Intellectual disability (ID) occurred in 14 patients (70%) (six with moderate level, five with mild, three with severe ID), while in six patients cognitive level was normal. Neurodevelopmental and psychiatric disorders were classified according to the Diagnostic and Statistical Manual of Mental disorders, 5th edition [26]. Most patients (17, 85%) developed psychiatric and behavioral disorders, including conduct disorder (n = 10), autism spectrum disorder (n = 9), oppositional defiant disorder (n = 5), obsessive-compulsive disorder (n = 4), intermittent explosive disorder (n = 3), attention deficit with or without hyperactivity disorder (n = 3), depressive and anxiety disorder (n = 1), social communication disorder (n = 1). The severity of psychiatric comorbidities was classified on an empirical basis as (0) “None”, (1) “Mild”, (2) “Moderate” and (3) “Severe”, considering a severity range spanning from the ‘mild’ category (1) where the psychiatric disorder would cause minimum functional impairment and the symptoms would be considered mild, to the ‘severe’ category (3) where there would be significant functional individual and social disruption and the symptoms would be considered severe or very severe.
Genetic testing revealed that all patients carried pathogenic PCDH19 mutations, 14 were missense substitutions, six were truncating (two stopgain, two frame-shift, one genomic deletion and one splicing). Eighteen patients were females, and two were males who carried mosaic PCDH19 mutations. In sixteen patients (80%) mutations occurred de novo, while in four they had been inherited (two from father, two from mother).
MRI data acquisition
We performed MRI acquisitions in participants adopting the same protocol on two 3 T systems (Achieva, Philips Healthcare, The Netherlands, at Meyer Hospital IRCCS; Magnetom Skyra, Siemens Healthineers, Germany, at IRCCS Bambino Gesù Hospital). MRI assessment included: 3D T1-weigthed fast-spoiled gradient echo (3D-T1), 3D T2-weighted Fluid Attenuation Inversion Recovery, T2-weighted Fast Spin-Echo, T2*-weighted Gradient-eco, and 2D T1-weigthed acquisitions. We acquired 3D-T1 whole-brain MRI by setting both 3 T scanners with the same sequence parameters (acquisition plane sagittal, TR/TE 8.0/3.7 ms, matrix 240 × 240, flip angle 8°, slice thickness 1 mm/no gap, FOV [240×240] mm2, number of slices 191, voxel resolution [1.0 × 1.0 × 1.0] mm3, acquisition time 6 min, 32 s). The study was approved by the Pediatric Ethics Committee of the Tuscany Region, Italy, in the context of the DESIRE project (FP7 EU call, grant agreement 602531) and its extension by the DECODE-EE (Tuscany Region Call for Health 2018) and the Human Brain Optical Mapping (Fondazione Cassa di Risparmio di Firenze 2021) projects. For all patients and controls, written informed consent for research use of clinical and genetic data was obtained from patients or their parents, or legal guardians in the case of minors or those with intellectual disability.
MRI data analysis
Visual analysis
Brain MRI images were reviewed by pediatric neuroradiologists looking for structural abnormalities and artifacts that could hamper computational analyses.
Morphometric analysis of cortical regions
We processed T1-3D MRI of each subject using the FreeSurfer processing pipeline (version 7.2.0, https://surfer.nmr.mgh.harvard.edu). The processing pipeline implies several stages, including segmentation of subcortical structures [27] segmentation of the cortical regions, and reconstruction of surfaces demarcating the boundaries between white and gray matter (white surface) and between gray matter and cerebrospinal fluid (CSF, pial surface) [28]. We performed the parcellation of the cortical mantle to divide the cortical surfaces into distinct gyral and sulcal regions [29] and generated detailed 3D high-resolution reconstructions for both white and pial surfaces for each hemisphere. To ensure accuracy, we carried out visual inspections and manual corrections on the subcortical segmentations and cortical surface reconstructions, when appropriate.
We conducted an analysis of the morphometric characteristics of the parceled regions of interest (ROIs). Our focus was to investigate cortical anatomy as it results from its developmental process and, to achieve this, we measured cortical thickness (CT), surface area (SA) and cortical volume (CV) for every vertex of the cortical surfaces. Cortical thickness, as quantified by measuring the distance between the white and pial surfaces [30], provides insights into neuronal migration and postmigrational organization [31], as well as on gray matter [32]. We determined the SA by summing the areas of the triangles surrounding a specific vertex. The SA is a trait with a high degree of heritability [33], and has been associated with cognitive abilities [34]. We determined cortical volume by multiplying the values of CT and SA. We averaged the values of these morphometric features (CT, SA and CV) for the parceled ROIs, as defined in the atlas provided by Desikan and colleagues (2006) and corrected for multiple comparison.
Network-based analysis of cortical and subcortical regions
We employed the processing pipeline introduced by Larivière et al. [16] to identify distinctive structural connectivity profiles exhibiting a significant correlation with morphometric abnormality maps. For this purpose, we used both CT and SA values to pinpoint regions in the cortex and subcortical structures. Regions were considered as epicenters due to their strong connections with regions showing high morphometric reductions and weaker connections with areas displaying lower levels of morphometric reductions.
The pipeline we used to identify these epicenters encompassed the following steps (detailed in https://enigma-toolbox.readthedocs.io): loading z-value maps for CT and SA, calculated as previously outlined; loading high-resolution structural (derived from diffusion-weighted tractography) connectivity matrix from a group of healthy adults in the Human Connectome Project, with parcellation based on Desikan et al. (2006); extracting seed-based structural connectivity matrices, utilizing each cortical and subcortical region as a seed.
Volumetric analysis of subcortical structures
We employed advanced computer-based modules for the subfield segmentations of subcortical structures. We adopted the method by Iglesias and colleagues [35] to provide an in-deep analysis of the hippocampal subfields. For the segmentation of amygdala nuclei, we adopted the method by Saygin et al. [36]. Such algorithms were optimized by these authors for subfield segmentations by adopting a computational atlas with a novel atlas-building algorithm based on Bayesian inference and working on T1-weighted images. To obtain more reliable segmentations, we included additional 3D T2-FLAIR MRIs in the processing pipeline. We finally inspected the subfield segmentations by visual analysis and manually corrected it where appropriate. For each subject, we calculated the volume of the segmented substructures of the hippocampus and amygdala.
Shape analysis of hippocampus and Cornu Ammonis
We used the segmentations obtained from the volumetric analysis to perform a shape analysis of the hippocampus and CA. We first considered the segmented masks and built the corresponding mesh of points using a MATLAB script (R2020b, MathWorks Inc., Natick, MA). We used the segmented volumes of the hippocampus and CA as inputs of SPHARM-MAT (https://www.nitrc.org/projects/spharm-mat), a tool based on a 3D Fourier surface representation method (SPHARM) that creates parametric surface models using the spherical harmonics [37]. The SPHARM processing pipeline included several steps. We pre-processed data to remove noises, therefore the 3D binary object presented a surface with a spherical topology. We repaired small vacancies and discontinuities between image voxels using the TopologyFix tool. We then adopted a parameterization algorithm to create a continuous and uniform mapping from the object surface to the surface of a unit sphere. We applied the CALD (control of area and length distortion) algorithm [38] to generate triangular meshes, consisting of vertices and faces. Then, with the SPHARM tool, we expanded the object surface into a complete set of spherical harmonic basis functions, which was a Fourier basis function defined on the sphere. By solving a linear system, we reconstructed the surface object using the Fourier coefficients up to a user-desired degree. Finally, with the SPHARM registration, we created a shape descriptor from a normalized set of SPHARM coefficients, which were comparable across objects for providing a group analysis. The registration used the first order ellipsoid for establishing surface correspondence and aligning objects. We applied surface-reconstruction steps for both the hippocampus and CA of both hemispheres.
To perform a comparison between patients and controls, we aligned models and carried a translation in the same center of reference using a high-resolution mesh (a six-level icosahendral mesh with 40962 vertices and 81920 faces). The same rotation matrix was used to register the CA in the same centroid with respect to the hippocampus and report the model in the center of the reference system.
Statistical analysis
We carried out statistical analyses on volumetric features using FreeSurfer [28] and FSL [39] tools and MATLAB R2020b (Statistics and Machine Learning Toolbox Version 11.6, MathWorks Inc., Natick, MA).
Morphometric and volumetric analysis of cortical and subcortical regions
To evaluate the effect of the diagnostic group (Group) on morphometric measurements, we performed a group analysis between PCDH19 patients and controls. To this purpose and mitigate confounding effects, we adopted a multivariate linear regression model using the center (Center) and age (Age) as covariates: \({Morph}={\beta }_{0}+{\beta }_{1}{Center}+{\beta }_{2}{Group}+{\beta }_{3}{Age}\). For volumetric measurements, we included in the model the estimated intra-cranial volume (β4eICV) as covariate.
Network-based analysis of cortical and subcortical regions
For each ROI, we evaluated the spatial correlation (rvalue) between the structural connectivity matrices and the z-value morphometry maps. We tested the statistical significance of each spatial correlation between seed-based cortico-cortical and subcortico-cortical connectivity and morphometric measurements using spin permutation tests (with 1,000 rotations and a significance threshold of pvalue < 0.1).
Relationship between morphometric analysis e gene-expression patterns
For each ROI, we extracted the spatial PCDH19-expression levels (εvalue) from the Allen Human Brain Atlas [18], a brain-wide gene expression atlas that includes measurement obtained through microarray technology from more than 20,000 genes, sampled across 3,702 spatially distinct tissue samples [40]. We calculated the spatial correlation (rvalue) between the PCDH19-expression levels (εvalue) and morphometry features representing the structural SA and CT alterations (βvalue) and assessed significance testing by spin permutation tests (with 1,000 rotations and a significance threshold of pspin < 0.1).
Relationship between morphometric and volumetric features and clinical findings
Maintaining the center and age as covariates, for each ROI, we estimated the effect of the morphometric and volumetric features on the clinical variables of patients using the following linear regression model: \({Clinics}={\beta }_{0}+{\beta }_{1}{Center}+{\beta }_{2}{Morph}+{\beta }_{3}{Age}\) (with Clinics as Age at seizure onset, Cognitive level, Psychiatric disorder, Habitual seizures, number of treatments).
Shape analysis of hippocampus and Cornu Ammonis
For the shape analysis, we assessed the difference between the mean shape of the two groups and calculated a t-test on the distribution of vertices of the volumetric meshes. We finally adopted a multivariate analysis of the covariance model (MANCOVA) to assess the covariance between the tridimensional meshes of the hippocampal shapes for each side.
Results
MRI visual analysis and clinical assessment
After visual inspection of diagnostic MRI sequences (Supplementary Fig. 1 and Table 1), one patient (no. 16) was excluded from the volumetric analysis due to movement artifacts, and another patient (no. 10) was excluded from the hippocampal analysis as an incomplete inversion of the left hippocampus was observed on the MRI scan (Supplementary Fig. 2). We assessed associations between clinical variables (Fig. 1) and found that an earlier age at seizure onset was significantly correlated with cognitive impairment (rvalue = -0.58, pvalue = 0.008). The severity of psychiatric comorbidities did not show a significant correlation with age at seizure onset (rvalue = -0.11, pvalue = 0.640).
Correlation patterns between age at seizure onset and cognitive level (A) and severity of psychiatric disorder (B) in the cohort of patients with PCDH19-gene-related syndrome. Cognitive level was classified as follow: (0) “normal” or (1) “mild ID” or (2) “moderate ID” or (3) “severe ID”. Clinical values are summarized in Table 1.
Morphometric analysis of cortical regions
In morphometric analysis, as shown in Supplementary Table 1 and depicted in Fig. 2, we observed a significant reduction of SA in cortical areas of the limbic circuit in PCDH19 patients compared to controls. ROIs with reduced SA were bilaterally the entorhinal (left/right pvalue = 0.002/0.327), parahippocampal (left/right pvalue = 0.230/0.016), fusiform (left/right pvalue = 0.004/0.013), inferior temporal (left/right pvalue = 0.002/0.062), superior temporal (left/right pvalue = 0.029/0.003), rostral-anterior cingulate (left/right pvalue = 0.018/0.069), medial orbito-frontal (left/right pvalue = 0.034/0.055) gyri, temporal pole (left/right pvalue = 0.001/0.004) and insula (left/right pvalue = 0.067/0.006). The reduction of SA contributed to a proportional reduction of CV within the same regions. A reduced CT was noted only in the parahippocampal gyri (left/right pvalue = 0.004/0.049).
Statistical ROI-based morphometric analysis of PCDH19-patients compared with matched controls. The results of the regression analysis on morphometric features, surface area (SA, A) and cortical thickness (CT, B), are represented by β2 maps (specified in Supplementary Table 1) superimposed on the pial surfaces of the left and right hemispheres. We observed statistically significant lower values of SA in left and right temporo-limbic cortices (blue ROIs, A), and normal SA in the surrounding regions (green ROIs, panel A). CT was significantly increased in right lateral occipital gyrus and parahippocampal regions bilaterally (yellow ROIs, B).
Network-based analysis of cortical and subcortical regions
Based on the results of structural morphometric analysis, we conducted a network-based analysis of SA and observed a statistically significant structural correlation in the limbic regions (as depicted in Fig. 3). Regions where emerged a significant negative correlation (SA reduction, for details see Supplementary Tables 2 and 3) were the fusiform (left/right pspin = 0.025/0.096), entorhinal (left/right pspin = 0.073/0.034), parahippocampal (left/right pspin = 0.230/0.016), inferior temporal (left/right pspin = 0.014/0.036) gyri, temporal pole (left/right pspin = 0.018/0.015), right insula (pspin = 0.074) and amygdala (left/right pspin = 0.003/0.033). We also noticed a positive correlation (SA increase) in paracentral (left/right pspin = 0.004/0.006), postcentral (left/right pspin = 0.030/0.009) and posterior cingulate (left/right pspin = 0.089/0.013) gyri.
Spatial correlations (rvalue, blue-red colormap) between SA patterns (β2) and seed-based cortico-cortical (A) and subcortico-cortical (B) connectivity profiles. In this example, we used as seed the left entorhinal gyrus. Correlation scores (rvalue) are detailed in Supplementary Tables 2 and 3. SA changes (β2) are indicated in Supplementary Table 1.
The network-based analysis of CT revealed no significant structural correlations (Supplementary Fig. 3 and Supplementary Tables 2 and 3).
Relationship between structural morphometric analysis and gene-expression
For each cortical and subcortical ROI, we extracted the spatial gene-expression coefficients (εvalue) from the Allen Human Brain Atlas [18] (Fig. 4). PCDH19 is significantly expressed in the cortical regions of the limbic circuit (Fig. 4A), in particular the fusiform (left/right εvalue = 0.717/0.701), entorhinal (left/right εvalue = 0.717/0.613), inferior temporal (left/right εvalue = 0.705/0.773), superior-temporal (left/right εvalue = 0.675/0.677), medial orbito-frontal (left/right εvalue = 0.741/0.788), and rostral-anterior cingulate (left/right εvalue = 0.696/0.792) gyri and temporal pole (left εvalue = 0.790). Among the limbic circuit, amygdala (left/right εvalue = 0.645/0.330) and hippocampus (left/right εvalue = 0.689/0.664) are the subcortical regions with the highest PCDH19 expression (Fig. 4B). We also noticed that cortical regions with positive correlations in network-based analysis (SA increase) had lower PCDH19 expression levels, i.e. paracentral (left/right εvalue = 0.434/0.494), postcentral (left/right εvalue = 0.433/0.521) and posterior cingulate (left/right εvalue = 0.520/0.570) gyri.
Spatial PCDH19 expression patterns (εvalue) in cortical (A) and subcortical (B) regions of interest and (C) correlation between levels of PCDH19 expression and surface area changes (β2 detailed in Supplementary Table 1) observed in cortical and subcortical regions. The statistical correlation highlighted that regions with higher level of PCDH19 expression exhibited significantly higher reductions in SA (rvalue = -0.262, pspin = 0.092).
We correlated morphometric alterations in the cortical and subcortical regions with the levels of PCDH19 expression (Fig. 4C) and observed that regions with highest PCDH19 levels had a statistically significant reduction of SA (rvalue = -0.262, pspin = 0.092). Subcortical regions exhibited no volumetric reductions, hence no correlations between volume reduction and expression level. We observed no statistically significant correlations between gene-expression levels and CT alterations (rvalue = -0.127, pspin = 0.185, Supplementary Fig. 4).
Relationship between morphometric analysis and severity of psychiatric comorbidities, habitual seizures and treatments
The effect of morphometric variables on psychiatric comorbidities (summarized in Table 1) was significant in several areas of the limbic cortex (Supplementary Table 4). Patients with more severe psychiatric symptoms exhibited lower values of SA in the left entorhinal (β2 = -0.0093, pvalue = 0.013), fusiform (β2 = -0.0018, pvalue = 0.046), inferior temporal (β2 = -0.0011, pvalue = 0.013), rostral anterior cingulate (β2 = -0.0033, pvalue = 0.025), rostral middle frontal (β2 = -0.0009, pvalue = 0.022) and right medial orbito-frontal (β2 = -0.0027, pvalue = 0.013) gyri (see Fig. 5). The effect on cortical thickness of habitual seizures was significant in two regions (Supplementary Table 5), the left caudal anterior (β2 = 1.1745, pvalue = 0.008) and posterior (β2 = 1.5761, pvalue = 0.008) cingulate. The pharmacological load (number of treatments, Supplementary Table 6) was significant for the cortical thickness of the entorhinal regions (left/right β2 = 1.4803/1.8152, pvalue = 0.014/0.010).
Significant regression patterns between the severity of psychiatric comorbidities and values of surface area (SA) of a representative subset of the cortical regions (left and right entorhinal, A and D; left and right fusiform, B and E; left and right inferiortemporal, C and F; left and right medialorbitofrontal, G and J; left and right rostralanteriorcingulate, H and K; left and right rostralmiddlefrontal, I and L, as reported in Supplementary Table 4) in PCDH19-mutated patients. The values, indicated by asterisks, are adjusted according to the linear regression model applied in the statistical analysis. The continuous line represents the fitting line, the dotted line indicated the 95% confidence bounds. We observed a negative trend in the fitting lines, that show a reduced SA in patients with more severe psychiatric disorder. Legend: lh, left hemisphere; rh, right hemisphere.
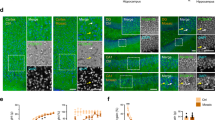
Analysis of subfields in subcortical structures
The group analysis of subcortical structures revealed volumetric reductions involving the hippocampal subfields bilaterally in the PCDH19 group compared to controls (Supplementary Table 7 and Supplementary Fig. 5). We observed a statistically significant volumetric reduction in the right hippocampal tail (pvalue = 0.048), hippocampal fissure (left/right pvalue = 0.003/0.014), right body of molecular layer (pvalue = 0.016), right GC-ML-DG region (pvalue = 0.007), head of the left CA3 region (pvalue = 0.002), body of the right CA3 (pvalue = 0.004), and of the right CA4 (pvalue = 0.011). The volume of other segments of CA were also reduced, although not reaching statistically significant values (body of CA1 left/right pvalue = 0.103/0.089; right CA3: pvalue = 0.084; body of the left CA3: pvalue = 0.078; head of the CA4: left/right pvalue = 0.115/0.119; body of the left CA4: pvalue = 0.083). We found that the volume of the body of the right hippocampus was significantly reduced (pvalue = 0.026), while the whole hippocampus was also reduced, but not significantly (left/right pvalue = 0.330/0.146), as we had previously measured [6].
In the left amygdala subunits (Supplementary Table 8), we found not significantly reduced values in central (pvalue = 0.100), medial (pvalue = 0.100) and cortical (pvalue = 0.082) nuclei, in patients with respect to controls.
Relationship between volume of subfields in subcortical structures and clinical findings
When we correlated the volumetric values with clinical variables (summarized in Table 1), we observed that patients with more severe psychiatric comorbidities exhibited lower volumetric values (Supplementary Table 9 and Supplementary Fig. 6), in particular in the heads of right CA3 (β2 = -0.0278, pvalue = 0.049) and left CA4 (β2 = -0.0302, pvalue = 0.038) and body of the right hippocampus (β2 = -0.0045, pvalue = 0.048). We observed no correlation between hippocampal volumetric assessments and age at seizure onset or cognitive level.
Shape analysis of hippocampus and Cornu Ammonis
We evaluated the mean difference between the shape of groups of PCDH19 patients and controls (Supplementary Fig. 7) by a t-test with a corresponding statistical significance (pvalue, Supplementary Fig. 8) and observed that the shape of the hippocampus was moderately different in several regions (Supplementary Fig. 8B and C). The main regions of significance in our comparative analyses were primarily located in the CA (Supplementary Fig. 9), which exhibited abnormal shapes in PCDH19 patients with respect to controls.
Discussion
In this cohort of patients with PCDH19 mutations we observed a significant correlation between age at seizure onset and ID, consistently with suggestions that a causal relationship exists in these genetic disorder between early onset seizures and adverse cognitive outcomes (Kolc et al. [3]).
In the morphometric analysis of the cortex, we observed PCDH19 patients to exhibit bilateral SA reductions in the cortical regions forming the limbic circuit, i.e. fusiform, entorhinal, parahippocampal, inferior and superior temporal, medial orbito-frontal and rostral-anterior cingulate gyri, temporal pole and insula. In our network-based analysis of SA the amygdala emerged as a structural epicenter in correlation with the abnormal patterns in cortical limbic regions. Morphometric and network-based analysis strengthen our previous surface-based findings [6], revealing clusters of reduced gyrification index in the limbic regions. Cortical gyrification represents the culmination of complex growth mechanisms influenced by genetic, molecular factors, and mechanical forces, initiating during fetal development and persisting throughout life [41, 42]. Cortical SA is proportional to the gyrification index [43], since the process of gyrification aids in augmenting the SA within finite spatial confines, and potentially enhances connectivity by reducing inter-regional distances [44]. SA is a highly heritable trait, yet little is known about genetic influences on regional cortical differentiation in humans [45, 46]. From the third trimester of pregnancy to the second year of age, the cortex expands dynamically in a regionally heterogeneous fashion [47]. SA is the result of such mechanisms of expansion [48]. We observed that reduced SA values significantly correlated with post-natal PCDH19-expression levels as reported in the atlas of the human brain transcriptome [40]. More precise correlations would be possible also considering prenatal PCDH19 expression levels, but this information is not fully available in public databases. We interrogated the Mouse Genome Informatics (MGI) database (https://www.informatics.jax.org/) and extracted PCDH19 expression levels (Supplementary Table 10), from embryonic day 0 (E0) to birth (postnatal day 0, P0). In mouse embryo, PCDH19 is expressed since early brain development (E8.5) and its expression has been detected at different stages in three regions participating to the limbic circuit, cerebral cortex (from E11.5 to E15.5), hippocampus (from E14.5 to E15.5), and thalamus (E18.5). In the mouse, E8.5 falls in the neurulation stage, the time windows from E11.5 to E15.5 and E18.5 are included in neurogenesis, neuronal proliferation and migration, and dendritic development and synaptogenesis [49]. Thus, developmental brain anomalies observed in patients carrying PCDH19 mutations might occur both during prenatal stages and postnatal maturation processes.
We quantitatively examined the volume and shape of the hippocampus, exploring all its components, and found patients to exhibit highly significant reductions in the volumes and shape differences in several subunits, particularly the CA. Studies exploring quantitative subfield alterations in TLE reported an overall effect of temporal lobe seizures manifesting as bilateral medial hippocampal atrophy, and a more selective effect of hippocampal seizures, leading to disease-proportional CA1 atrophy [50, 51]. The hippocampal formation acts as an epicenter of more intense structural changes even in epilepsy syndromes in which no clear indication of seizure genesis in the hippocampus exists, such as SCN1A-related Dravet syndrome [52] and idiopathic generalized epilepsy syndromes [20, 53]. In a previous study of PCDH19-related encephalopathy, we demonstrated altered gyrification in the parahippocampal and entorhinal regions, and diffusion abnormalities in the underlying white matter [6], but no alterations emerged in the hippocampus when it was examined as a whole. The novel findings we present in this study demonstrate that specific hippocampal subfields show significantly reduced volumes to an extent that did not emerge when analyzing the whole hippocampus.
We suppose that structural alterations in the hippocampus, amygdala and connected cortical regions of the limbic circuit might be the expression of specific PCDH19 defects underlying irregular growth dynamics. In-vitro studies and experimental animal models corroborate this hypothesis, since PCDH19 expression patterns play a key role in the formation of limbic regions [21, 54], more specifically the CA [22]. The PCDH19 protein is also abundant in the mossy fiber pathway [55], where its expression levels increase postnatally during synapse development [56]. The mossy fiber pathway terminates in synapses formed by dentate granule cells onto CA3 pyramidal neurons. Pcdh19 mRNA is expressed by both dentate granule cells and CA3 pyramidal neurons, indicating that PCDH19 regulates the development, stabilization, maintenance and function of both the pre- and post-synaptic neurons of mossy fiber synapses [13]. PCDH19 mutations might also impair columnar organization [57] and lead to deviations in developmental trajectories at a network level [58]. Animal models have highlighted how PCDH19 intervenes in regulating the formation and maintenance of brain networks [13], and how mosaic mutations disrupt local cortical networks [14]. Mouse models in which PCDH19 is expressed in mosaic fashion exhibit structural and functional synaptic abnormalities in the hippocampus, with impaired neuronal network activity and connectivity [15].
The limbic cortex and amygdala emerged as structural epicenters of SA alterations, i.e. regions strongly inter-connected with regions in which reduced SA emerged, and weakly inter-connected with regions with normal SA. Structural abnormalities within the limbic network may correspond to specific clinical manifestations in individuals with PCDH19 encephalopathy as suggested by the frequently observed pattern of focal seizures with affective symptoms [1], also confirmed in most patients within the current cohort. We also evaluated the effect on morphometry of pharmacological load and habitual seizures and found no significant effect at either the SA or cortical thickness globally, while the left caudal cingulate and entorhinal cortex revealed a weak correlation with active epilepsy.
PCDH19-related epilepsies are among the eight most common single-gene epilepsies, with an incidence of 1 per 20,600 live-born females, and a prevalence of 4.85/100,000 (95% CI 1.97–9.15) [59]. Conditions resulting from single-gene mutations provide an adequate paradigm for correlating specific imaging changes with behavioral and psychiatric phenotypes. Our observations indicate that patients exhibiting more severe psychiatric manifestations had more prominent pattern of anatomic aberration, notably involving the entorhinal and fusiform cortex, hippocampal body and CA3. Such findings align with our previous study demonstrating that in PCDH19 encephalopathy, more significant gyrification and diffusion abnormalities in the mesio-temporal regions occur in individuals with ASD [6]. A correlation between atypical mesio-temporal anatomy and autism has been demonstrated using MRI in various clinical contexts [7,8,9,10]. Post-mortem brain studies in individuals with autism have revealed abnormalities in the hippocampus, amygdala and connected limbic structures [60, 61]. Altered amygdala development has been demonstrated in children with autism [62, 63], but little is known about the simultaneous development of the brain regions directly connected with the amygdala [64]. Autism has been associated with genes related to the regulation of neuronal proliferation [65] and transcriptionally downregulated genes implicated in autism have been robustly associated with global changes in cortical thickness variability in children with autism [66]. Such findings corroborate our observations that the neurodevelopmental trajectory of brain maturation may be atypical in ASD [67], and that structural brain imaging offers clues about the effect of the genetic etiology of neuropsychiatric disorders [68].
Advanced neuroimaging holds great promise as a tool for categorizing patients with different neuropsychiatric manifestations [69] and may elucidate genotype-phenotype correlations at the patient level. Recent advances in machine learning, combined with quantitative neuroimaging methods, have demonstrated their remarkable capability to discern disparities in local morphological features across various brain subregions and elucidated variations between patients groups, a crucial step toward establishing a reliable psychiatric nosology, understanding disease progression and predicting individual disease trajectories [70]. MRI-based quantitative methods are a potential tool for early characterization and quantification of anatomical changes to predict the individual-level progression of clinical variables, such as the likelihood of experiencing cognitive disability and psychiatric comorbidities.
Although our methodological approaches have detected anatomic differences between patients with PCDH19 encephalopathy and a control population, several limitations must be acknowledged. We only included 20 patients, with a wide age range. However, PCDH19 encephalopathy remains rare, has its onset in the pediatric age and is associated with impaired cognitive function, which may pose challenges in systematically assessing psychiatric features. Since we analyzed images acquired from two different scanners, we adopted a multivariate statistical model to mitigate biases among centers. Furthermore, while analyzing the structural characteristics of hippocampal subfields presents challenges, the tools we applied have undergone extensive validation and have been adopted in numerous previous investigations [71]. Effects of genetic variant types (missense vs other) on morphometric features, expression patterns and clinical manifestations can be studied adopting suitable statistical methods for exploring data from larger cohorts [3].
Data availability
Data are available from corresponding author upon reasonable request.
References
Marini C, Darra F, Specchio N, Mei D, Terracciano A, Parmeggiani L, et al. Focal seizures with affective symptoms are a major feature of PCDH19 gene-related epilepsy. Epilepsia. 2012;53:2111–9.
Rodriguez-Cruces R, Royer J, Larivière S, Bassett DS, Caciagli L, Bernhardt BC. Multimodal connectome biomarkers of cognitive and affective dysfunction in the common epilepsies. Netw Neurosci. 2022;6:320-38.
Kolc KL, Sadleir LG, Scheffer IE, Ivancevic A, Roberts R, Pham DH, et al. A systematic review and meta-analysis of 271 PCDH19-variant individuals identifies psychiatric comorbidities, and association of seizure onset and disease severity. Mol Psychiatry. 2019;24:241–51.
Trivisano M, Pietrafusa N, Terracciano A, Marini C, Mei D, Darra F, et al. Defining the electroclinical phenotype and outcome of PCDH19-related epilepsy: A multicenter study. Epilepsia. 2018;59:2260–71.
Kurian M, Korff CM, Ranza E, Bernasconi A, Lübbig A, Nangia S, et al. Focal cortical malformations in children with early infantile epilepsy and PCDH19 mutations: case report. Dev Med Child Neurol. 2018;60:100–5.
Lenge M, Marini C, Canale E, Napolitano A, de Masi S, Trivisano M, et al. Quantitative MRI-based analysis identifies developmental limbic abnormalities in PCDH19 encephalopathy. Cereb Cortex. 2020;30:6039–50.
Travers BG, Adluru N, Ennis C, Tromp DPM, Destiche D, Doran S, et al. Diffusion Tensor Imaging in Autism Spectrum Disorder: A Review. Autism Res. 2012;5:289–313.
Wallace GL, Robustelli B, Dankner N, Kenworthy L, Giedd JN, Martin A. Increased gyrification, but comparable surface area in adolescents with autism spectrum disorders. Brain. 2013;136:1956–67.
Ecker C, Andrews D, Dell’Acqua F, Daly E, Murphy C, Catani M, et al. Relationship between cortical gyrification, white matter connectivity, and autism spectrum disorder. Cereb Cortex. 2016;26:3297–309.
Duret P, Samson F, Pinsard B, Barbeau EB, Boré A, Soulières I, et al. Gyrification changes are related to cognitive strengths in autism. NeuroImage Clin. 2018;20:415–23.
Hazlett HC, Gu H, Munsell BC, Kim SH, Styner M, Wolff JJ, et al. Early brain development in infants at high risk for autism spectrum disorder. Nature. 2017;542:348–51.
Gerosa L, Francolini M, Bassani S, Passafaro M. The Role of Protocadherin 19 (PCDH19) in Neurodevelopment and in the Pathophysiology of Early Infantile Epileptic Encephalopathy-9 (EIEE9). Dev Neurobiol. 2019;79:75–84.
Hoshina N, Johnson-Venkatesh EM, Hoshina M, Umemori H Female-specific synaptic dysfunction and cognitive impairment in a mouse model of PCDH19 disorder. Science (80-) 2021;372. https://doi.org/10.1126/science.aaz3893.
Lamers D, Landi S, Mezzena R, Baroncelli L, Pillai V, Cruciani F, et al. Perturbation of Cortical Excitability in a Conditional Model of PCDH19 Disorder. Cells. 2022;11:1–16.
Giansante G, Mazzoleni S, Zippo AG, Ponzoni L, Ghilardi A, Maiellano G et al. Neuronal network activity and connectivity are impaired in a conditional knockout mouse model with PCDH19 mosaic expression. Mol Psychiatry 2023. https://doi.org/10.1038/s41380-023-02022-1.
Larivière S, Rodríguez-Cruces R, Royer J, Caligiuri ME, Gambardella A, Concha L, et al. Network-based atrophy modeling in the common epilepsies: A worldwide ENIGMA study. Sci Adv. 2020;6:1–14.
Larivière S, Royer J, Rodríguez-Cruces R, Paquola C, Caligiuri ME, Gambardella A, et al. Structural network alterations in focal and generalized epilepsy assessed in a worldwide ENIGMA study follow axes of epilepsy risk gene expression. Nat Commun. 2022;13:1–16.
Arnatkevic̆iūtė A, Fulcher BD, Fornito A. A practical guide to linking brain-wide gene expression and neuroimaging data. Neuroimage. 2019;189:353–67.
Nees F, Pohlack ST. Functional MRI studies of the hippocampus. Hippocampus Clin Neurosci. 2014;34:85–94.
Whelan CD, Altmann A, Botía JA, Jahanshad N, Hibar DP, Absil J, et al. Structural brain abnormalities in the common epilepsies assessed in a worldwide ENIGMA study. Brain. 2018;141:391–408.
Kim SY, Mo JW, Han S, Choi SY, Han SB, Moon BH, et al. The expression of non-clustered protocadherins in adult rat hippocampal formation and the connecting brain regions. Neuroscience. 2010;170:189–99.
Kim SY, Chung HS, Sun W, Kim H. Spatiotemporal expression pattern of non-clustered protocadherin family members in the developing rat brain. Neuroscience. 2007;147:996–1021.
Specchio N, Marini C, Terracciano A, Mei D, Trivisano M, Sicca F, et al. Spectrum of phenotypes in female patients with epilepsy due to protocadherin 19 mutations. Epilepsia. 2011;52:1251–7.
Cappelletti S, Specchio N, Moavero R, Terracciano A, Trivisano M, Pontrelli G, et al. Cognitive development in females with PCDH19 gene-related epilepsy. Epilepsy Behav. 2015;42:36–40.
Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. Research electronic data capture (REDCap)-A metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inf. 2009;42:377–81.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders. American Psychiatric Association, 2013 https://doi.org/10.1176/appi.books.9780890425596.
Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C, et al. Whole brain segmentation: Automated labeling of neuroanatomical structures in the human brain. Neuron. 2002;33:341–55.
Fischl B. FreeSurfer. Neuroimage. 2012;62:774–81.
Fischl B, van der Kouwe A, Destrieux C, Halgren E, Ségonne F, Salat DH, et al. Automatically Parcellating the Human Cerebral Cortex. Cereb Cortex. 2004;14:11–22.
Fischl B, Dale AM. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc Natl Acad Sci USA. 2000;97:11050–5.
Rakic P. Specification of cerebral cortical areas. Science (80-). 1988;241:170–6.
Huttenlocher PR. Morphometric study of human cerebral cortex development. Neuropsychologia. 1990;28:517–27.
Panizzon MS, Fennema-Notestine C, Eyler LT, Jernigan TL, Prom-Wormley E, Neale M, et al. Distinct Genetic Influences on Cortical Surface Area and Cortical Thickness. Cereb Cortex. 2009;19:2728–35.
Vuoksimaa E, Panizzon MS, Chen C-H, Fiecas M, Eyler LT, Fennema-Notestine C, et al. The Genetic Association Between Neocortical Volume and General Cognitive Ability Is Driven by Global Surface Area Rather Than Thickness. Cereb Cortex. 2015;25:2127–37.
Iglesias JE, Augustinack JC, Nguyen K, Player CM, Player A, Wright M, et al. A computational atlas of the hippocampal formation using ex vivo, ultra-high resolution MRI: Application to adaptive segmentation of in vivo MRI. Neuroimage. 2015;115:117–37.
Saygin ZM, Kliemann D, Iglesias JE, van der Kouwe AJW, Boyd E, Reuter M, et al. High-resolution magnetic resonance imaging reveals nuclei of the human amygdala: manual segmentation to automatic atlas. Neuroimage. 2017;155:370–82.
Brechbühler C, Gerig G, Kübler O. Parametrization of closed surfaces for 3-D shape description. Comp Vis Graph Image Process. 1995;61:154–70.
Shen L, Makedon F. Spherical mapping for processing of 3D closed surfaces. Image Vis Comput. 2006;24:743–61.
Jenkinson M, Beckmann CF, Behrens TEJJ, Woolrich MW, Smith SM. Fsl. Neuroimage. 2012;62:782–90.
Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, Shen EH, Ng L, Miller JA, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 2012;489:391–9.
Llinares-Benadero C, Borrell V. Deconstructing cortical folding: genetic, cellular and mechanical determinants. Nat Rev Neurosci. 2019;20:161–76.
Hogstrom LJ, Westlye LT, Walhovd KB, Fjell AM. The structure of the cerebral cortex across adult life: Age-related patterns of surface area, thickness, and gyrification. Cereb Cortex. 2013;23:2521–30.
Schaer M, Cuadra MBB, Tamarit L, Lazeyras F, Eliez S, Thiran J. A Surface-Based Approach to Quantify Local Cortical Gyrification. IEEE Trans Med Imaging. 2008;27:161–70.
Docherty AR, Hagler DJ, Panizzon MS, Neale MC, Eyler LT, Fennema-Notestine C, et al. Does degree of gyrification underlie the phenotypic and genetic associations between cortical surface area and cognitive ability? Neuroimage. 2015;106:154–60.
Chen CH, Gutierrez ED, Thompson W, Panizzon MS, Jernigan TL, Eyler LT, et al. Hierarchical genetic organization of human cortical surface area. Science (80-). 2012;335:1634–6.
van der Meer D, Kaufmann T. Mapping the genetic architecture of cortical morphology through neuroimaging: progress and perspectives. Transl Psychiatry. 2022;12:1–11.
Huang Y, Wu Z, Wang F, Hu D, Li T, Guo L, et al. Mapping developmental regionalization and patterns of cortical surface area from 29 post-menstrual weeks to 2 years of age. Proc Natl Acad Sci. 2022;119:1–8.
Lyall AE, Shi F, Geng X, Woolson S, Li G, Wang L, et al. Dynamic Development of Regional Cortical Thickness and Surface Area in Early Childhood. Cereb Cortex. 2015;25:2204–12.
Chini M, Hanganu-Opatz IL. Prefrontal Cortex Development in Health and Disease: Lessons from Rodents and Humans. Trends Neurosci. 2021;44:227–40.
Maccotta L, Moseley ED, Benzinger TL, Hogan RE. Beyond the CA1 subfield: Local hippocampal shape changes in MRI-negative temporal lobe epilepsy. Epilepsia. 2015;56:780–8.
Hogan RE, Moseley ED, Maccotta L. Hippocampal Surface Deformation Accuracy in T1-Weighted Volumetric MRI Sequences in Subjects with Epilepsy. J Neuroimaging. 2015;25:452–9.
Lenge M, Balestrini S, Mei D, Macconi L, Caligiuri ME, Cuccarini V, et al. Morphometry and network-based atrophy patterns in SCN1A-related Dravet syndrome. Cereb Cortex. 2023;33:9532–41.
Bouyeure A, Patil S, Mauconduit F, Poiret C, Isai D, Noulhiane M. Hippocampal subfield volumes and memory discrimination in the developing brain. Hippocampus. 2021;31:1202–14.
Hirano S, Yan Q, Suzuki ST. Expression of a novel protocadherin, OL-protocadherin, in a subset of functional systems of the developing mouse brain. J Neurosci. 1999;19:995–1005.
Schaarschuch A, Hertel N. Expression profile of N-cadherin and protocadherin-19 in postnatal mouse limbic structures. J Comp Neurol. 2018;526:663–80.
Terauchi A, Johnson-Venkatesh EM, Toth AB, Javed D, Sutton MA, Umemori H. Distinct FGFs promote differentiation of excitatory and inhibitory synapses. Nature. 2010;465:783–7.
Cooper SR, Emond MR, Duy PQ, Liebau BG, Wolman MA, Jontes JD. Protocadherins control the modular assembly of neuronal columns in the zebrafish optic tectum. J Cell Biol. 2015;211:807–14.
Light SEW, Jontes JD. Multiplane calcium imaging reveals disrupted development of network topology in zebrafish pcdh19 mutants. eNeuro. 2019;6:1–15.
Symonds JD, Zuberi SM, Stewart K, Mclellan A, O’regan M, Macleod S, et al. Molecular epidemiology of monogenic epilepsies answers key clinical questions. Brain. 2019;142:2173–5.
Ben Shalom D. Memory in autism: Review and synthesis. Cortex. 2003;39:1129–38.
Bauman ML, Kemper TL. Neuroanatomic observations of the brain in autism: A review and future directions. Int J Dev Neurosci. 2005;23:183–7.
Amaral DG, Nordahl CW. Amygdala Involvement in Autism: Early Postnatal Changes, But What Are the Behavioral Consequences? Am J Psychiatry. 2022;179:522–4.
Shen MD, Swanson MR, Wolff JJ, Elison JT, Girault JB, Kim SH, et al. Subcortical Brain Development in Autism and Fragile X Syndrome: Evidence for Dynamic, Age- and Disorder-Specific Trajectories in Infancy. Am J Psychiatry. 2022;179:562–72.
Lee JK, Andrews DS, Ozturk A, Solomon M, Rogers S, Amaral DG, et al. Altered Development of Amygdala-Connected Brain Regions in Males and Females with Autism. J Neurosci. 2022;42:6145–55.
Stein JL, de la Torre-Ubieta L, Tian Y, Parikshak NN, Hernández IA, Marchetto MC, et al. A quantitative framework to evaluate modeling of cortical development by neural stem cells. Neuron. 2014;83:69–86.
Romero-Garcia R, Warrier V, Bullmore ET, Baron-Cohen S, Bethlehem RAI. Synaptic and transcriptionally downregulated genes are associated with cortical thickness differences in autism. Mol Psychiatry. 2019;24:1053–64.
Ecker C, Bookheimer SY, Murphy DGM. Neuroimaging in autism spectrum disorder: brain structure and function across the lifespan. Lancet Neurol. 2015;4422:1–14.
Radonjić NV, Hess JL, Rovira P, Andreassen O, Buitelaar JK, Ching CRK, et al. Structural brain imaging studies offer clues about the effects of the shared genetic etiology among neuropsychiatric disorders. Mol Psychiatry. 2021;26:2101–10.
Bahathiq RA, Banjar H, Bamaga AK, Jarraya SK Machine learning for autism spectrum disorder diagnosis using structural magnetic resonance imaging: Promising but challenging. Front Neuroinform 2022;16. https://doi.org/10.3389/fninf.2022.949926.
Yassin W, Nakatani H, Zhu Y, Kojima M, Owada K, Kuwabara H et al. Machine-learning classification using neuroimaging data in schizophrenia, autism, ultra-high risk and first-episode psychosis. Transl Psychiatry 2020;10. https://doi.org/10.1038/s41398-020-00965-5.
Sämann PG, Iglesias JE, Gutman B, Grotegerd D, Leenings R, Flint C et al. FreeSurfer-based segmentation of hippocampal subfields: A review of methods and applications, with a novel quality control procedure for ENIGMA studies and other collaborative efforts. Hum. Brain Mapp. 2020. https://doi.org/10.1002/hbm.25326.
Acknowledgements
This study was supported in part by funding from the ‘Current Research’ by the Italian Ministry of Health and by a grant from the Italian Ministry of Education, University and Research (grant code 20172C9HLW). This work was supported by grants to RG from the Tuscany Region Call for Health 2018 (project DECODE-EE) and Fondazione Cassa di Risparmio di Firenze (project Human Brain Optical Mapping). This work was also supported by #NEXTGENERATIONEU (NGEU) and funded by the Ministry of University and Research (MUR), National Recovery and Resilience Plan (NRRP), project MNESYS (PE0000006) – A Multiscale integrated approach to the study of the nervous system in health and disease (DN. 1553 11.10.2022). This research was also supported by the European Union – Next Generation EU – NRRP M6C2 – Investment 2.1 Enhancement and strengthening of biomedical research in the NHS and by a grant from the Italian Ministry of Health and Tuscany Region (grant code RF-2021-12372804, 2023-2026) Use of Transcranial Magnetic Stimulation (TMS) as a surrogate of Pathophysiology in genetic epilepsies – TMSpath.
Author information
Authors and Affiliations
Consortia
Contributions
ML: conception and design, acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; SB: analysis and interpretation of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; AN: acquisition of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; DM: acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; VC: analysis and interpretation of data, revising the article critically for important intellectual content, final approval of the version to be published; GB: acquisition of data, revising the article critically for important intellectual content, final approval of the version to be published; MT: acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; SP: acquisition of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; LM: analysis and interpretation of data, revising the article critically for important intellectual content, final approval of the version to be published; DL: acquisition of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; MCRE: acquisition of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; SC: acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content; final approval of the version to be published; LD’I: analysis and interpretation of data, drafting the article or revising it critically for important intellectual content; final approval of the version to be published; CB: acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content, final approval of the version to be published; NS: acquisition of data, analysis and interpretation of data, drafting the article or revising it critically for important intellectual content; final approval of the version to be published; and RG: conception and design, acquisition of data, analysis and interpretation of data; drafting the article or revising it critically for important intellectual content; final approval of the version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lenge, M., Balestrini, S., Napolitano, A. et al. Morphometric network-based abnormalities correlate with psychiatric comorbidities and gene expression in PCDH19-related developmental and epileptic encephalopathy. Transl Psychiatry 14, 35 (2024). https://doi.org/10.1038/s41398-024-02753-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-024-02753-x