Abstract
Schizophrenia (SCZ) is a complex disorder that typically arises in late adolescence or early adulthood. Age at onset (AAO) of SCZ is associated with long-term outcomes of the disease. We explored the genetic architecture of AAO with a genome-wide association study (GWAS), heritability, polygenic risk score (PRS), and copy number variant (CNV) analyses in 4 740 subjects of European ancestry. Although no genome-wide significant locus was identified, SNP-based heritability of AAO was estimated to be between 17 and 21%, indicating a moderate contribution of common variants. We also performed cross-trait PRS analyses with a set of mental disorders and identified a negative association between AAO and common variants for SCZ, childhood maltreatment and attention-deficit/hyperactivity disorder. We also investigated the role of copy number variants (CNVs) in AAO and found an association with the length and number of deletions (P-value = 0.03), whereas the presence of CNVs previously reported in SCZ was not associated with earlier onset. To our knowledge, this is the largest GWAS of AAO of SCZ to date in individuals from European ancestry, and the first study to determine the involvement of common variants in the heritability of AAO. Finally, we evidenced the role played by higher SCZ load in determining AAO but discarded the role of pathogenic CNVs. Altogether, these results shed light on the genetic architecture of AAO, which needs to be confirmed with larger studies.
Similar content being viewed by others
Introduction
Schizophrenia (SCZ) is a complex disorder influenced by an intricate interplay of genetic and environmental factors. SCZ patients show substantial heterogeneity in clinical characteristics such as symptomatology, cognitive ability, course, overall functioning, and age at onset (AAO). AAO has been consistently included among the most important determinants of disease outcome and is widely accepted as a significant clinical and prognostic factor [1, 2]. For instance, an earlier AAO is associated with a higher likelihood of having relatives with SCZ [3, 4], and has been correlated with an increased number of hospitalizations and illness episodes, more frequent negative symptoms, and poorer cognition, overall functioning, and global outcome [5,6,7]. In general, men have an earlier AAO, usually between 20 and 24 years of age, while in women, the onset occurs between 25 and 35 years of age [8,9,10]. In addition, women appear to have a secondary peak around menopause, between 50 and 54 years old [7, 11].
The genetic architecture of SCZ is complex. Heritability estimates from twin and population-based studies range between 64 and 81% [12, 13], in which common genetic variants account for a large proportion (24.4%) [14]. There is also evidence that both rare single-nucleotide variants and rare copy number variants (CNVs) contribute to the risk of developing SCZ [15,16,17,18,19]. In fact, individuals with a pathogenic CNV represent more than 2% of the confirmed cases [20]. CNVs are highly penetrant and may cause early-onset forms of developmental delay or autism spectrum disorders. Thus, similarly, it has been suggested that the presence of CNVs may play an important role in the onset of SCZ, although their contribution is still unclear [20].
The heritability of the AAO has been estimated in sibling pairs at 33% [21], indicating a moderate genetic basis. However, in contrast with the vast amount of information on the genetics of SCZ obtained from genome-wide association studies (GWAS) [14, 22, 23], the genetic determinants underlying AAO remain largely unknown. To date, only three GWAS have been performed in relatively small cohorts (<3000 individuals) and none of them identified any genomic loci associated with AAO at genome-wide significance [24,25,26]. On the other hand, recent GWAS carried out based on the age at onset of both Bipolar Disorder (BD) and Major Depression Disorder (MDD) with larger sample sizes have determined a significant SNP-based heritability and shared genetic risk with other psychiatric disorders [27, 28]. Thus, further studies with larger sample sizes are required to estimate the contribution of common genetic variants to AAO and the heritability they may explain.
Ultimately, identifying and researching the genetic factors that influence the AAO of SCZ may improve our understanding of the development and progression of this disease, provide new targets for therapy, and facilitate the development of personalized therapeutic interventions and preventive measures. In this study, we aimed to explore the genetic architecture of AAO by performing (1) a GWAS meta-analysis of nearly 5000 subjects of European ancestry, (2) heritability estimates based on common genetic variants, (3) polygenic risk score (PRS) analyses with a set of mental traits, and finally 4) an assessment of the influence of known CNVs on AAO.
Material and methods
Sample
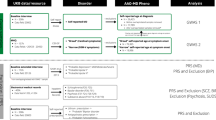
Four different datasets, two from Europe (CIBERSAM, PsyCourse) and two from USA (GAIN [29], and nonGAIN), were obtained and combined to perform a GWAS meta-analysis on AAO comprising 4740 patients of European ancestry (Table 1). In all datasets, subjects met the variat for SCZ, schizoaffective disorder, schizophreniform disorder, delusional disorder, brief psychotic disorder, or psychotic disorder not otherwise specified, in the Diagnostic and Statistical Manual of Mental Disorders version IV (DSM-IV).
The CIBERSAM and PsyCourse datasets were collected by the authors, while GAIN and nonGAIN datasets were obtained from public resources. Participants in the CIBERSAM (Biomedical Research Network in Mental Health) dataset were recruited from psychiatric in-patient units at seven different hospitals in Spain [30]. Participation was approved by the ethical committees at the hospitals involved in the recruitment. Finally, samples were genotyped using the Illumina Infinium PsychArray at the Broad Institute as part of the wave 3 meta-analysis GWAS of SCZ of the Psychiatric Genomics Consortium (PGC-SCZ wave 3) [31]. The PsyCourse samples were part of a multi-site German/Austrian longitudinal study (www.psycourse.de) that was conducted between 1 January 2012 and 31 December 2019. The study collected deep phenotypic, neuropsychologic, and omics data from patients with brief psychotic disorder, major depressive disorder (MDD), bipolar disorder (BD), SCZ, schizoaffective disorder, and healthy individuals. Adult participants were referred by the clinical staff or identified by querying patient registries. Study protocols were reviewed and approved by the ethics committees of the Medical Centers and Faculties involved in the recruitment, in accordance with the Declaration of Helsinki. All participants provided written informed consent. The phenotype information was gathered using the v4.1 version of the PsyCourse data release. These samples were genotyped using the Illumina Infinium PsychArray [32]. Finally, GAIN and nonGAIN datasets were both obtained from the dbGaP repository (accession numbers phs000021.v3.p2 and phs000167.v1.p1, respectively), and genotyped with the Affymetrix Genome-Wide Human SNP Array 6.0 as described elsewhere [33].
Age at onset
In the CIBERSAM dataset, AAO was defined as the onset of the first psychotic symptoms. The patient (and/or family members) and the psychiatrist defined when the first psychological symptoms appeared. In cases where this information was not available, we used the date of the first psychiatric visit due to a psychotic episode. In the PsyCourse dataset, AAO was collected as both the age at first outpatient and inpatient treatment. Subjects with information available for either of these data were included, and when both data were available, the earliest age was used. For the GAIN and nonGAIN datasets, AAO was defined as the most likely AAO of psychotic symptoms consistent with the onset of SCZ. A consensus diagnostician (PI or senior research clinician delegate) reviewed the diagnostic ratings made independently by two research diagnosticians (one of which could be the consensus diagnostician as well) and assigned a final diagnosis and AAO if the ratings were in agreement. The Kolmogorov-Smirnov test was used to determine pair-wise differences in AAO between datasets.
GWAS and functional analyses
Quality control (QC) was conducted for each dataset separately (CIBERSAM, PsyCourse, GAIN and nonGAIN) using PLINK 1.9 [34], according to standard procedures for GWAS [35]. Briefly, genetic variants with missingness rate >2%, minor allele frequency (MAF) < 5%, Hardy-Weinberg equilibrium (HWE) P-value < 1e−06, and those belonging to non-autosomal chromosomes were excluded from downstream analyses. Ambiguous and multiallelic variants were also removed. Subjects with a missingness rate > 2%, increased or decreased heterozygosity rates (defined as ±3 standard deviations away from the sample mean), and relatedness >12.5% (PI_HAT > 0.125) were excluded. Sex was imputed based on X chromosome heterozygosity/homozygosity rates, before removing sex chromosomes. Principal component analyses (PCA) were conducted using SMARTPCA from EIGENSOFT 6.1.4 [36]. To keep only subjects with European ancestry, we did not use those individuals who were beyond ±3 standard deviations from the mean of the first two principal components (PCs) of the European cluster of the 1000 Genomes Project [37] Phase I. We also removed four individuals who clustered in the Finnish subgroup of the European cluster. Before genotype imputation, a PCA was performed again on the remaining subjects and the top 10 PCs were kept for further analysis.
Genotype imputation was conducted for each resulting dataset independently using the TOPMed reference panel [38]. Imputed datasets were filtered according to an imputation quality score (r-squared) <0.9 and converted to binary files using PLINK’s --vcf flag. Then, a post-imputation QC was conducted for each dataset separately, using PLINK. Only single-nucleotide polymorphisms (SNPs) were kept for further analyses. In addition, ambiguous and multiallelic SNPs, as well as SNPs with MAF < 1%, and an HWE P-value < 1e−06 were excluded. The resulting datasets were lifted over to genome build 19 using the UCSC liftOverPlink tool [39]. The CIBERSAM dataset included: 1704 subjects and 4,962,031 SNPs; PsyCourse: 499 subjects and 5,338,835 SNPs; GAIN: 1278 subjects and 6,042,664 SNPs; and nonGAIN: 1259 subjects and 6,074,765 SNPs. A GWAS was performed by linear regression for each dataset in PLINK, using normalized AAO as outcome and sex and the top 10 PCs as covariates. Since AAO was different between data sets a meta-analysis was then conducted using the tool METAL [40] applying an inverse variance strategy. As a result, we obtained information on 4740 subjects and 6,540,522 SNPs. As an alternative method, individual-level imputed genotype data for each of the four separate datasets was merged and a GWAS was conducted in parallel for comparison purposes. From here on, this approach is called the merged approach (Supplementary Note).
Genomic loci showing suggestive associations with AAO (P-value < 1e-05) were identified and explored using the FUMA software [41]. Each genomic risk locus was represented by the top lead SNP which had the minimum P-value in the locus. Lead SNPs were defined as associated SNPs (P-value < 1e−05) that were independent of each other at LD r2 < 0.1. Independent significant SNPs were defined as SNPs with a P-value < 1e−05, and independent of each other at a linkage disequilibrium (LD) threshold r2 < 0.6. The genomic risk loci were mapped to protein-coding genes by positional mapping based on ANNOVAR [42] and eQTL mapping with GTEx v8 and BRAINEAC databases. Finally, pathway enrichment analyses of the mapped genes were conducted using KOBAS-i [43]. In all the analyses, a 5% false discovery rate (FDR) was considered for multiple testing correction.
SNP-based heritability
The proportion of phenotypic variance explained by SNPs was estimated using two different methods. First, SNP-based heritability was estimated with the Linkage Disequilibrium Score Regression (LDSC) method [44]. To reduce the standard error given our relatively small sample size, the intercept was constrained to 1 after testing that it was not significantly higher than 1 [45]. The LDSC intercept has been widely employed to distinguish between inflation due to confounding factors (such as population stratification and cryptic relatedness) and inflation due to polygenicity. Deviation of the intercept from 1 is indicative of residual confounding, thus observing an intercept not significantly higher than 1 can be interpreted as there being minimal confounding bias [44]. Second, we also estimated heritability using individual-level genotypes with the Genome-based Restricted Maximum Likelihood (GREML) approach implemented in the Genome-wide Complex Trait Analysis (GCTA) tool [46], adjusting for sex, the dataset and the top 10 PCs.
Cross-trait polygenic risk score
PRS analyses were conducted using the PRS-cs software [47] between ten mental phenotypes and AAO. Specifically, we obtained summary statistics data on seven psychiatric disorders downloaded from the Psychiatric Genomics Consortium (https://www.med.unc.edu/pgc/) [48], including SCZ, BD, MDD, attention-deficit/hyperactivity disorder (ADHD), autism spectrum disorder (ASD), obsessive-compulsive disorder (OCD), and cannabis use disorder (CUD). In addition, we obtained GWAS data on three conditions previously associated with AAO: neuroticism [49], educational attainment (EA) [11], and childhood maltreatment [50] (Table S1). The downloaded summary statistics were filtered to remove ambiguous, multiallelic and duplicated SNPs, and SNPs with an imputation score <0.9 (if the information was provided). The summary statistics of the mental phenotypes were used as base data, and the individual-level genotype dataset was used as target data. We calculated the PRS for each mental phenotype using the “auto” mode (the shrinkage parameter phi was determined from the data with a Bayesian approach). Then, the scores obtained for each individual in our dataset were regressed out against sex, age, and batch to obtain new adjusted-PRS scores. Linear regressions were performed with the normalized AAO values as outcome and the adjusted-PRSs as independent variables to evaluate the association of each PRS with AAO. Finally, p-values were corrected for multiple correction using FDR.
Copy number variation analysis
CNV analyses were conducted using signal intensity data from those individuals in the four cohorts from whom we obtained the intensity files (N = 4630). The raw CNVs were obtained using PennCNV [51]. Briefly, quality control of CNV calls was based on a sample-level criterion that examined the relationship between the standard deviation of the logarithm R Ratio (LRR_SD) and the number of CNV calls (NumCNV). At the end of the process, adjacent calls were merged together into one single call. Thresholds were carefully chosen to include as many subjects as possible but reduce false positives. Thus, subjects with LRR_SD > 0.35, BAF > 0.01, WF > 0.05, and NumCNV > 150 were not included. With all the identified CNVs we performed linear regression analyses to test whether the AAO was associated with either the number or the total length of CNVs, deletions or duplications. Sex, batch and the 10 first PCs were used as covariables. In parallel, 12 CNVs previously described as significantly associated with SCZ (SCZ-CNV) were obtained from the literature [52]. The BEDTools intersect [53] was used to look for overlaps between the described SCZ-CNVs and our data. Briefly, we selected as SCZ-CNV carriers only those individuals for whom at least 90% of the SCZ-CNV overlapped with a detected CNV (-f 0.9) and/or with at least 80% of reciprocal overlap (-r -f 0.8). To determine whether the presence of SCZ-CNVs could be influencing AAO, Wilcoxon and Kruskal–Wallis tests were performed in R4.1.2 [54], using non-normalized AAO values.
Results
Age at onset
Mean AAO was 23.35 (SD = 8.26). Mean AAO varied across datasets, ranging from 21.11 to 26.26 (Table 1). In the whole sample 1525 subjects (32.4%) were females (mean AAO = 25.09), and 3182 subjects (67.6%) were males (mean AAO = 22.52). Significant differences were detected between the AAO of CIBERSAM and GAIN (P-value < 2.2e−16), CIBERSAM and nonGAIN (P-value < 2.2e−16), PsyCourse and GAIN (P-value = 2.2e−16), and PsyCourse and nonGAIN (P-value = 4.44e−16). However, differences were not detected between CIBERSAM and PsyCourse (P-value = 0.11), and neither between GAIN and nonGAIN (P-value = 0.33, Fig. S1). Since the distribution of AAO was right-skewed (Fig. S1), it was normalized using a rank-based inverse-normal transformation and used in all subsequent analyses.
GWAS and functional analyses
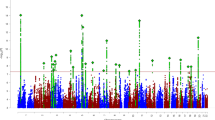
A total of 4740 subjects of European ancestry and 6,540,522 SNPs were included in the GWAS meta-analysis. Although none of the analyzed SNPs reached the genome-wide significant threshold (P-value < 5e−08, Fig. 1), 25 lead SNPs were identified, corresponding to 22 genomic risk loci that were mapped to 183 genes (Table 2). Using the Linkage Disequilibrium Score Regression (LDSC) method [44], an intercept of 1.03 (SE = 0.0083) was obtained. Mapped genes were enriched in categories such as transport of small molecules (FDR-adj P-value = 1.7e−02), vesicle-mediated transport (FDR-adj P-value = 1.8e−02), metabolism (FDR-adj P-value = 2.55e−02), MHC class II antigen presentation (FDR-adj P-value = 2.6e−02), and Asparagine N-linked glycosylation (FDR-adj P-value = 3.3e−02), among others (Table S2).
SNP-based heritability
The SNP-based heritability (h2SNP) was estimated using two methods that showed consistent results with moderate and significant SNP-based heritability. First, with LDSC, we obtained h2SNP = 0.21 (SE = 0.07). SNP-based heritability was also estimated using individual-level genotypes with GCTA-GREML, adjusting for sex, dataset and the top 10 PCs, resulting in an estimate of h2SNP = 0.17 (SE = 0.06; P-value = 3.33e−03). These results were consistent with the heritability estimates obtained from the GWAS merged approach (h2SNP = 0.13, Supplementary Note).
Cross-trait polygenic risk score
Ten mental phenotypes were examined through a cross-trait PRS analysis. Only the PRSs calculated based on ADHD, SCZ and childhood maltreatment sumstats were associated with AAO in our dataset (FDR < 0.05). All adjusted PRS beta coefficients were negative, corresponding to the higher burden of disease/condition risk variants being associated with earlier onset of SCZ (Fig. 2 and Table S3). The variance explained by the adjusted-PRS of these three phenotypes was low but significant (1.2e−03; 1.3e−03 and 1.1e−03, respectively for ADHD, SCZ, and childhood maltreatment). Among the other mental phenotypes, only BD-PRS was nominally associated with AAO (P = 0.02).
Beta estimates (x-axis) for the adjusted-PRS of the linear model are shown. Non-significant values are colored in grey (FDR > 0.05) and significant results (FDR < 0.05) in red. Adjusted R2 values of the linear models are shown for each phenotype on the right side of the panel. SCZ Schizophrenia, OCD obsessive-compulsive disorder, Neu Neuroticism, MDD major depressive disorder, EA educational attainment, CUD Cannabis use disorder, CM childhood maltreatment, BD bipolar disorder, ASD Autism spectrum disorder, ADHD attention-deficit/hyperactivity disorder.
CNV analysis
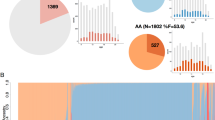
After quality control, a total of 3965 individuals remained for CNV downstream analyses and a total of 107,668 deletions (mean number = 27.18 ± 17.12; mean length = 1434 kb ± 1617 kb) and 59,336 duplications (mean number = 15.05 ± 10.09; mean length = 2094 kb ± 6817 kb) were identified. 3962 individuals (99.92%) carried at least one deletion and 3942 (99.41%) individuals carried at least one duplication. AAO was significantly associated with the total number of CNVs (beta = −0.015; P-value = 0.025), and also with the number of deletions (beta = −0.02; P-value = 0.03) and the length of the deletions (beta = −1.8e−7; P-value = 0.03, Table 3). Out of 3965, 117 individuals (3%) carried SCZ-CNVs previously associated with SCZ, and 4 individuals carried two SCZ-CNVs. Among the SCZ-CNV-carriers, we detected 36 (29.8%) carrying 15q11.2del, 20 (16.53%) 22q11.2del, 13 (10.74%) 16p12.1del, 12 (9.92%) 16p11.2dup, 11 (9.09%) 16p13.11dup, 10 (8.26%) 15q13.3del, 6 (4.96%) 3q29del, 4 (3.31%) carrying 1q21.1dup, 4 (3.31%) 15q11-q13dup, 3 (2.48%) 1q21.1del, and 2 (1.65%) 7q11.23dup individuals. The deletion corresponding to 2p16.3 was not present in our sample. In our dataset, the presence of SCZ-CNVs was not associated with an earlier AAO (Wilcoxon test; P-value = 0.73, Fig. S2A). In addition, no differences in the AAO were found across SCZ-CNVs (Kruskal–Wallis test; P-value = 0.49, Fig. S2B).
Discussion
This study explored the genetic architecture of AAO of SCZ. To this end, we performed a case-only GWAS of European ancestry in the largest sample collected to date. Although no genome-wide significant signals were detected, we successfully estimated for the first time the SNP-based heritability of AAO using two different approaches that showed consistent results of moderate heritability, ranging from 17 to 21%. The fact that we did not identify any specific genetic variants associated with AAO suggests that the heritability of the trait is likely to be complex, involving multiple genetic and environmental factors. We also provided evidence of negative genetic associations of cross-trait PRS derived from ADHD, SCZ, and childhood maltreatment with AAO. Finally, we determined that the burden of deletions was associated with AAO in our dataset.
In our study, the strongest association signal was found in a genomic locus at chromosome 4 (lead SNP rs111289733 P-value = 1.66e−07), which harbored the long non-coding RNA RP11-180C1.1. The mapped genes belonging to the suggestive associations were enriched in transport of small molecules, vesicle-mediated transport, Asparagine N-linked glycosylation, and MHC class II antigen presentation, among others. Both the transport of molecules and, specifically, the vesicular transport mechanism might participate in the pathogenesis of SCZ by triggering dysfunctional neuroexocytosis [55, 56]. In fact, previous studies reported abnormal reductions in synaptic vesicle proteins being associated with SCZ [57,58,59]. Also, MHC has been strongly associated with the risk of SCZ [60]. In addition, glycosylation involves processes critical for normal brain development and has been suggested to contribute to the abnormal neuronal signaling and connectivity observed in SCZ [61]. Altogether, these categories are promising candidates for further studies on pathways associated with AAO.
Over the years, many studies have evaluated the role of genetics in AAO of SCZ [62, 63], estimating a heritability of AAO ranging from approximately 20 to 58% [7]. In our study, based on two different methodologies we estimated the SNP-based heritability of AAO to be between 17 and 21%. In fact, our estimates show consistent results suggesting a moderate but significant contribution of common variants to AAO. Interestingly, SNP-based heritability is slightly higher than that of AAO in BD and MDD, which has recently been estimated at 5 and 6% respectively, using larger sample sizes [27]. Further studies with larger sample sizes are needed to obtain more accurate estimates in SCZ AAO.
Cross-trait PRSs constructed with ADHD, SCZ, and childhood maltreatment were significantly associated with AAO, showing that a higher risk of developing these conditions is associated with an earlier AAO in SCZ. However, they explained a very small fraction of AAO variation [64]. Previous studies reported that ADHD was among the commonest comorbidities in children and adolescents with SCZ [65], and it has been argued that the genetic architecture of ADHD has a large link with SCZ [66]. In this line, our results suggest that an earlier AAO may be related to a more severe neurodevelopmental impairment. AAO has been suggested as a potential endophenotype of SCZ, reflecting the underlying genetic architecture of the disorder. Our finding that the PRS of SCZ can predict AAO supports this notion and suggests that AAO could be a useful predictor of disease severity. It has also been suggested that individuals with higher genetic loadings for SCZ are at a higher risk of early onset [67], and similarly for MDD [28]. Moreover, patients with histories of being abused as children show an earlier onset of symptoms [68]. Here, we also report a negative association between AAO and the PRS of both SCZ and childhood maltreatment. However, it is still unknown how the genetic architectures of these traits are linked.
A recent study reported that the prevalence of recurrent CNVs was higher in early onset psychosis than in the general population, as well as CNV pathogenicity [67]. In addition, some of these CNVs cause earlier-onset disorders such as developmental delay or ASD, but not SCZ [20]. Here, we observed that the burden of CNVs, especially deletions (either the length or the total number), but not duplications, were associated with an earlier onset of SCZ, suggesting a combined role of common variation and CNVs in determining AAO. However, in our study, neither the presence of pathogenic SCZ-CNV, nor any specific SCZ-CNVs were associated with AAO. Nevertheless, we detected a 1.9% prevalence of the pathogenic SCZ-CNVs, which is close to the previously reported prevalence of 2.6% [20].
Our study has some strengths and limitations that deserve discussion. Despite the lack of genome-wide significant associations at the SNP-level, which could be expected given the relatively small sample size of our study, we were able to estimate the SNP-based heritability of AAO and identify relevant associations with the PRS of three mental phenotypes for the first time. This finding represents an important step toward elucidating the genetic architecture of AAO. It is worth noting that studying AAO is both technically and conceptually challenging, as there is often considerable variability in how AAO is defined and measured between studies. Although these measures have been shown to be correlated and to occur within a relatively short time frame [6, 69], it is possible that our results may be confounded by different definitions of AAO between datasets. In our study, we acknowledge that AAO differed between the Europe and US cohorts in our study and therefore performed a GWAS meta-analysis to minimize any potential bias due to phenotype heterogeneity. However, the fact that AAO did not differ within the European (CIBERSAM and PsyCourse) and the US (GAIN and nonGAIN) cohorts suggests that the phenotypes were obtained in a homogeneous manner. In the future, more reliable estimates of AAO may lead to the identification of significant signals and an increase in the heritability explained by SNPs. Also, the rank-based transformation applied to the AAO values may have affected our GWAS and subsequent analyses; however, it is considered as one of the best approaches to use. Some studies have reported that for small sample sizes or genetic effects, there is an improvement in sensitivity for rank-based transformations that outweighs a slight increase in the false-positive rate [70]. In addition, a recent study has demonstrated that these transformation tests outperform the standard untransformed association test, both in terms of power and type I error rate control [71]. Moreover, we were not able to control for putative differences between the different recruitment sites. All the datasets included in the study might be multicenter; thus, heterogeneity within datasets could be considerable. Such phenotypic heterogeneity has been reported to affect genetic analyses [27, 72], which indicates that phenotype harmonization is as important as a larger sample size for improving the power to detect significant associations and avoiding a biased view of genetic architectures.
Conclusions
In conclusion, we report on the largest GWAS of AAO in SCZ to date, providing the first SNP-based heritability estimate of AAO in individuals of European ancestry. Although no genome-wide significant SNP was detected, we provide evidence of a genetic background for AAO and a negative association with the PRS of ADHD, SCZ and childhood maltreatment. In addition, we demonstrate that the burden of deletions is associated with the AAO of the disease. Larger sample sizes are needed to fully determine the genetic architecture of AAO, which could help us understand further the pathogenesis of SCZ and contribute to the development of better strategies for the early detection of SCZ. Nonetheless, our study provides an important step forward in understanding the genetic architecture of AAO.
Data availability
Given the collaborative nature of the CIBERSAM cohort and the involvement of multiple institutions and research groups, requests for access to the genotype data should be addressed to the corresponding author [muntaneg@peremata.com] to initiate the data access request. Acces to Psycourse dataset can be requested thorugh a “Secondary Analysis Proposal” at http://www.psycourse.de/openscience-en.html.
Change history
30 November 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41398-023-02651-8
References
Delisi LE. The significance of age of onset for schizophrenia. Schizophr Bull. 1992;18:209–15.
Öngür D, Lin L, Cohen BM. Clinical characteristics influencing age at onset in psychotic disorders. Compr Psychiatry. 2009;50:13–9.
Kendler KS, MacLean CJ. Estimating familial effects on age at onset and liability to schizophrenia. I. Results of a large sample family study. Genet Epidemiol. 1990;7:409–17.
Sham PC, Jones P, Russell A, Gilvarry K, Bebbington P, Lewis S, et al. Age at onset, sex, and familial psychiatric morbidity in schizophrenia. Camberwell collaborative psychosis study. Br J Psychiatry. 1994;165:466–73.
Rajji TK, Ismail Z, Mulsant BH. Age at onset and cognition in schizophrenia: meta-analysis. Br J Psychiatry. 2009;195:286–93.
Immonen J, Jääskeläinen E, Korpela H, Miettunen J. Age at onset and the outcomes of schizophrenia: a systematic review and meta-analysis: age at onset and the outcomes of schizophrenia. 2017. https://doi.org/10.1111/eip.12412.
Musket CW, Kuo SS, Rupert PE, Almasy L, Gur RC, Prasad K, et al. Why does age of onset predict clinical severity in schizophrenia? A multiplex extended pedigree study. Am J Med Genet, Part B: Neuropsychiatric Genet. 2020;183:403–11.
Aleman A, Kahn RS, Selten JP. Sex differences in the risk of schizophrenia: evidence from meta-analysis. Archives General Psychiatry. 2003;60:565–71.
Leung M. Sex differences in schizophrenia, a review of the literature. Acta Psychiatr Scand Suppl. 2000;401:3–38.
Neill E, Tan EJ, Lin Toh W, Selvendra A, Morgan VA, Rossell SL, et al. Examining which factors influence age of onset in males and females with schizophrenia. 2020. https://doi.org/10.1016/j.schres.2020.08.011.
Ochoa S, Usall J, Cobo J, Labad X, Kulkarni J. Gender differences in schizophrenia and first-episode psychosis: a comprehensive literature review. Schizophr Res. Treatment. 2012;2012:1–9.
Sullivan PF, Kendler KS, Neale MC. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Archives General Psychiatry. 2003;60:1187–92.
Lichtenstein P, Yip BH, Björk C, Pawitan Y, Cannon TD, Sullivan PF, et al. Common genetic influences for schizophrenia and bipolar disorder: a population-based study of 2 million nuclear families. Lancet. 2009;373:1–14.
Consortium TSWG of the PG, Ripke S, Walters JT, O’Donovan MC. Mapping genomic loci prioritises genes and implicates synaptic biology in schizophrenia. medRxiv 2020; 2020.09.12.20192922.
Levinson DF, Duan J, Oh S, Wang K, Sanders AR, Shi J, et al. Copy number variants in schizophrenia: Confirmation of five previous finding sand new evidence for 3q29 microdeletions and VIPR2 duplications. Am J Psychiatry. 2011;168:302–16.
Rees E, Walters JTR, Georgieva L, Isles AR, Chambert KD, Richards AL, et al. Analysis of copy number variations at 15 schizophrenia-associated loci. Br J Psychiatry. 2014;204:108–14.
Rees E, Kendall K, Pardiñas AF, Legge SE, Pocklington A, Escott-Price V, et al. Analysis of intellectual disability copy number variants for association with schizophrenia. JAMA Psychiatry. 2016;73:963–9.
Marshall CR, Howrigan DP, Merico D, Thiruvahindrapuram B, Wu W, Greer DS, et al. Contribution of copy number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat Genet. 2016;49:27–35.
Halvorsen M, Huh R, Oskolkov N, Wen J, Netotea S, Giusti-Rodriguez P, et al. Increased burden of ultra-rare structural variants localizing to boundaries of topologically associated domains in schizophrenia. Nat Commun. 2020;11:1–13.
Kirov G, Rees E, Walters JTR, Escott-Price V, Georgieva L, Richards AL, et al. The penetrance of copy number variations for schizophrenia and developmental delay. Biol Psychiatry. 2014;75:378–85.
Hare E, Glahn DC, Dassori A, Raventos H, Nicolini H, Ontiveros A, et al. Heritability of age of onset of psychosis in schizophrenia. Am J Med Genet, Part B: Neuropsychiatric Genet. 2010;153:298–302.
Ripke S, O’Dushlaine C, Chambert K, Moran JL, Kähler AK, Akterin S, et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet. 2013;45:1150–9.
Working Group of the Psychiatric Genomics Consortium S. Biological insights from 108 schizophrenia-associated genetic loci. 2014. https://doi.org/10.1038/nature13595.
Wang K-S, Liu X, Zhang Q, Aragam N, Pan Y. Genome-wide association analysis of age at onset in schizophrenia in a European-American sample. Am J Med Genet Part B: Neuropsychiatric Genet. 2011;156:671–80.
Bergen SE, O’Dushlaine CT, Lee PH, Fanous AH, Ruderfer DM, Ripke S, et al. Genetic modifiers and subtypes in schizophrenia: Investigations of age at onset, severity, sex and family history. Schizophr Res. 2014;154:48–53.
Woolston AL, Hsiao P-CC, Kuo P-HH, Wang S-HH, Lien Y-JJ, Liu C-MM, et al. Genetic loci associated with an earlier age at onset in multiplex schizophrenia. Sci Rep. 2017;7:6486.
Kalman JL, Olde Loohuis LM, Vreeker A, McQuillin A, Stahl EA, Ruderfer D, et al. Characterisation of age and polarity at onset in bipolar disorder. Br J Psychiatry. 2021;219:659–69.
Harder A, Nguyen T-D, Pasman JA, Mosing MA, Hägg S, Lu Y. Genetics of age-at-onset in major depression. Transl Psychiatry. 2022;12:1–7.
The GAIN Collaborative Research Group. New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nat Genet. 2007;39:1045–51.
Salagre E, Arango C, Artigas F, Ayuso-Mateos JL, Bernardo M, Castro-Fornieles J, et al. CIBERSAM: Ten years of collaborative translational research in mental disorders. Revista de Psiquiatría y Salud Mental (English Edition). 2019;12:1–8.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Budde M, Anderson-Schmidt H, Gade K, Reich-Erkelenz D, Adorjan K, Kalman JL, et al. A longitudinal approach to biological psychiatric research: The PsyCourse study. Am J Med Genet, Part B: Neuropsychiatric Genet. 2019;180:89–102.
Shi J, Levinson DF, Duan J, Sanders AR, Zheng Y, Péer I, et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature. 2009;460:753–7.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Marees AT, de Kluiver H, Stringer S, Vorspan F, Curis E, Marie-Claire C, et al. A tutorial on conducting genome-wide association studies: Quality control and statistical analysis. Int J Methods Psychiatr Res. 2018;27:e1608.
Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS Genet. 2006;2:e190.
1000 Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium, Taliun D, Harris DN, Kessler MD, Carlson J, Szpiech ZA, et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature. 2021;590:290–9.
Navarro Gonzalez J, Zweig AS, Speir ML, Schmelter D, Rosenbloom KR, Raney BJ, et al. The UCSC Genome Browser database: 2021 update. Nucleic Acids Res. 2021;49:D1046–57.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Watanabe K, Taskesen E, Van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1–11.
Wang K, Li M, Hakonarson H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:1–7.
Bu D, Luo H, Huo P, Wang Z, Zhang S, He Z, et al. KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 2021;49. https://doi.org/10.1093/nar/gkab447.
Bulik-Sullivan B, Loh PR, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Nievergelt CM. International meta-analysis of PTSD genome-wide association studies identifies sex-and ancestry-specific genetic risk loci. https://doi.org/10.1038/s41467-019-12576-w.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: A tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Ge T, Chen C-Y, Ni Y, Feng Y-CA, Smoller JW. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat Commun. 2019;10:1776.
Sullivan PF, Agrawal A, Bulik CM, Andreassen OA, Børglum AD, Breen G, et al. Psychiatric genomics: an update and an agenda. https://doi.org/10.1176/appi.ajp.2017.17030283.
Goodwin RD, Fergusson DM, Horwood LJ. Neuroticism in adolescence and psychotic symptoms in adulthood. Psychol Med. 2003;33:1089–97.
Varese F, Smeets F, Drukker M, Lieverse R, Lataster T, Viechtbauer W, et al. Childhood adversities increase the risk of psychosis: A meta-analysis of patient-control, prospective-and cross-sectional cohort studies. Schizophr Bull. 2012;38:661–71.
Wang K, Li M, Hadley D, Liu R, Glessner J, Grant SFA, et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. https://doi.org/10.1101/gr.6861907.
Warland A, Kendall KM, Rees E, Kirov G, Caseras X. Schizophrenia-associated genomic copy number variants and subcortical brain volumes in the UK Biobank. Mol Psychiatry. 2020;25:854–62.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Team RC. R: A language and environment for statistical computing. 2021.
Egbujo CN, Sinclair D, Hahn C-G. Dysregulations of synaptic vesicle trafficking in schizophrenia. Curr Psychiatry Rep. 2016;18:77.
Schubert KO, Föcking M, Prehn JHM, Cotter DR. Hypothesis review: are clathrin-mediated endocytosis and clathrin-dependent membrane and protein trafficking core pathophysiological processes in schizophrenia and bipolar disorder? Mol Psychiatry. 2012;17:669–81.
Nakato M, Shiranaga N, Tomioka M, Watanabe H, Kurisu J, Kengaku M, et al. ABCA13 dysfunction associated with psychiatric disorders causes impaired cholesterol trafficking. J Biol Chem. 2021;296:100166.
Osimo EF, Beck K, Reis Marques T, Howes OD. Synaptic loss in schizophrenia: a meta-analysis and systematic review of synaptic protein and mRNA measures. Mol Psychiatry. 2019;24:549–61.
Onwordi EC, Halff EF, Whitehurst T, Mansur A, Cotel M-C, Wells L, et al. Synaptic density marker SV2A is reduced in schizophrenia patients and unaffected by antipsychotics in rats. Nat Commun. 2020;11:246.
Mokhtari R, Lachman HM. The Major Histocompatibility Complex (MHC) in Schizophrenia: A Review. J Clin Cell Immunol. 2016;07. https://doi.org/10.4172/2155-9899.1000479.
Williams SE, Mealer RG, Scolnick EM, Smoller JW, Cummings RD. Aberrant glycosylation in schizophrenia: a review of 25 years of post-mortem brain studies. Mol Psychiatry. 2020;25:3198–207.
Vares M, Saetre P, Deng H, Cai G, Liu X, Hansen T, et al. Association between methylenetetrahydrofolate reductase (MTHFR) C677T polymorphism and age of onset in schizophrenia. Am J Med Genet. 2010;153B:610–8.
Guo S, Liu J, Li W, Yang Y, Lv L, Xiao X, et al. Genome wide association study identifies four loci for early onset schizophrenia. Transl Psychiatry. 2021;11:248.
Anttila V, Bulik-Sullivan B, Finucane HK, Walters RK, Bras J, Duncan L, et al. Analysis of shared heritability in common disorders of the brain. Science. 2018;360. https://doi.org/10.1126/science.aap8757.
Ross RG, Heinlein S, Tregellas H. High rates of comorbidity are found in childhood-onset schizophrenia. Schizophr Res. 2006;88:90–5.
Hamshere ML, Stergiakouli E, Langley K, Martin J, Holmans P, Kent L, et al. Shared polygenic contribution between childhood attention-deficit hyperactivity disorder and adult schizophrenia. Br J Psychiatry. 2013;203:107–11.
Bearden CC, Glahn D, Brownstein C, Douard E, Mollon J, Cabral K, et al. Prevalence of rate of deleterious copy number variants similar in early onset psychosis and autism spectrum disorders: implications for clinical practice. Biol Psychiatry. 2022;91:S56–7.
Kaufman J, Torbey S. Child maltreatment and psychosis. Neurobiol Dis. 2019;131:104378.
Jones PB. Adult mental health disorders and their age at onset. Br J Psychiatry. 2013;202:s5–s10.
Goh L, Yap VB. Effects of normalization on quantitative traits in association test. BMC Bioinformatics. 2009;10. https://doi.org/10.1186/1471-2105-10-415.
McCaw ZR, Lane JM, Saxena R, Redline S, Lin X. Operating characteristics of the rank-based inverse normal transformation for quantitative trait analysis in genome-wide association studies. Biometrics. 2020;76:1262–72.
Cai N, Revez JA, Adams MJ, Andlauer TFM, Breen G, Byrne EM, et al. Minimal phenotyping yields genome-wide association signals of low specificity for major depression. Nat Genet. 2020;52:437–47.
Acknowledgements
This work was supported by Instituto de Salud Carlos III through the projects PI18/00514 and PI21/00612 and co-funded by the European Union, by the Catalan Agency of Research and Universities (AGAUR, 2017SGR-00444 and 2021SGR01065). The PsyCourse study was supported by DFG (SCHU 1603/4-1, 5-1, 7-1, FA241/16-1).
Author information
Authors and Affiliations
Contributions
ES-F carried out the analyses with the help of SA. GM conceived and designed the study. GM and EV, with the help of SP, JLK and TGS, participated in the scientific discussion and supervision of the project. The other authors contributed to data collection. All authors critically revised and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sada-Fuente, E., Aranda, S., Papiol, S. et al. Common genetic variants contribute to heritability of age at onset of schizophrenia. Transl Psychiatry 13, 201 (2023). https://doi.org/10.1038/s41398-023-02508-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02508-0