Abstract
Mental illnesses, such as depression, are highly prevalent and have been shown to impact an individual’s physical health. Recently, artificial intelligence (AI) methods have been introduced to assist mental health providers, including psychiatrists and psychologists, for decision-making based on patients’ historical data (e.g., medical records, behavioral data, social media usage, etc.). Deep learning (DL), as one of the most recent generation of AI technologies, has demonstrated superior performance in many real-world applications ranging from computer vision to healthcare. The goal of this study is to review existing research on applications of DL algorithms in mental health outcome research. Specifically, we first briefly overview the state-of-the-art DL techniques. Then we review the literature relevant to DL applications in mental health outcomes. According to the application scenarios, we categorize these relevant articles into four groups: diagnosis and prognosis based on clinical data, analysis of genetics and genomics data for understanding mental health conditions, vocal and visual expression data analysis for disease detection, and estimation of risk of mental illness using social media data. Finally, we discuss challenges in using DL algorithms to improve our understanding of mental health conditions and suggest several promising directions for their applications in improving mental health diagnosis and treatment.
Similar content being viewed by others
Introduction
Mental illness is a type of health condition that changes a person’s mind, emotions, or behavior (or all three), and has been shown to impact an individual’s physical health1,2. Mental health issues including depression, schizophrenia, attention-deficit hyperactivity disorder (ADHD), and autism spectrum disorder (ASD), etc., are highly prevalent today and it is estimated that around 450 million people worldwide suffer from such problems1. In addition to adults, children and adolescents under the age of 18 years also face the risk of mental health disorders. Moreover, mental health illnesses have also been one of the most serious and prevalent public health problems. For example, depression is a leading cause of disability and can lead to an increased risk for suicidal ideation and suicide attempts2.
To better understand the mental health conditions and provide better patient care, early detection of mental health problems is an essential step. Different from the diagnosis of other chronic conditions that rely on laboratory tests and measurements, mental illnesses are typically diagnosed based on an individual’s self-report to specific questionnaires designed for the detection of specific patterns of feelings or social interactions3. Due to the increasing availability of data pertaining to an individual’s mental health status, artificial intelligence (AI) and machine learning (ML) technologies are being applied to improve our understanding of mental health conditions and have been engaged to assist mental health providers for improved clinical decision-making4,5,6. As one of the latest advances in AI and ML, deep learning (DL), which transforms the data through layers of nonlinear computational processing units, provides a new paradigm to effectively gain knowledge from complex data7. In recent years, DL algorithms have demonstrated superior performance in many data-rich application scenarios, including healthcare8,9,10.
In a previous study, Shatte et al.11 explored the application of ML techniques in mental health. They reviewed literature by grouping them into four main application domains: diagnosis, prognosis, and treatment, public health, as well as research and clinical administration. In another study, Durstewitz et al.9 explored the emerging area of application of DL techniques in psychiatry. They focused on DL in the studies of brain dynamics and subjects’ behaviors, and presented the insights of embedding the interpretable computational models into statistical context. In contrast, this study aims to provide a scoping review of the existing research applying DL methodologies on the analysis of different types of data related to mental health conditions. The reviewed articles are organized into four main groups according to the type of the data analyzed, including the following: (1) clinical data, (2) genetic and genomics data, (3) vocal and visual expression data, and (4) social media data. Finally, the challenges the current studies faced with, as well as future research directions towards bridging the gap between the application of DL algorithms and patient care, are discussed.
Deep learning overview
ML aims at developing computational algorithms or statistical models that can automatically infer hidden patterns from data12,13. Recent years have witnessed an increasing number of ML models being developed to analyze healthcare data4. However, conventional ML approaches require a significant amount of feature engineering for optimal performance—a step that is necessary for most application scenarios to obtain good performance, which is usually resource- and time-consuming.
As the newest wave of ML and AI technologies, DL approaches aim at the development of an end-to-end mechanism that maps the input raw features directly into the outputs through a multi-layer network structure that is able to capture the hidden patterns within the data. In this section, we will review several popular DL model architectures, including deep feedforward neural network (DFNN), recurrent neural network (RNN)14, convolutional neural network (CNN)15, and autoencoder16. Figure 1 provides an overview of these architectures.
a Deep feedforward neural network (DFNN). It is the basic design of DL models. Commonly, a DFNN contains multiple hidden layers. b A recurrent neural network (RNN) is presented to process sequence data. To encode history information, each recurrent neuron receives the input element and the state vector of the predecessor neuron, and yields a hidden state fed to the successor neuron. For example, not only the individual information but also the dependence of the elements of the sequence x1 → x2 → x3 → x4 → x5 is encoded by the RNN architecture. c Convolutional neural network (CNN). Between input layer (e.g., input neuroimage) and output layer, a CNN commonly contains three types of layers: the convolutional layer that is to generate feature maps by sliding convolutional kernels in the previous layer; the pooling layer is used to reduce dimensionality of previous convolutional layer; and the fully connected layer is to make prediction. For the illustrative purpose, this example only has one layer of each type; yet, a real-world CNN would have multiple convolutional and pooling layers (usually in an interpolated manner) and one fully connected layer. d Autoencoder consists of two components: the encoder, which learns to compress the input data into a latent representation layer by layer, whereas the decoder, inverse to the encoder, learns to reconstruct the data at the output layer. The learned compressed representations can be fed to the downstream predictive model.
Deep feedforward neural network
Artificial neural network (ANN) is proposed with the intention of mimicking how human brain works, where the basic element is an artificial neuron depicted in Fig. 2a. Mathematically, an artificial neuron is a nonlinear transformation unit, which takes the weighted summation of all inputs and feeds the result to an activation function, such as sigmoid, rectifier (i.e., rectified linear unit [ReLU]), or hyperbolic tangent (Fig. 2b). An ANN is composed of multiple artificial neurons with different connection architectures. The simplest ANN architecture is the feedforward neural network (FNN), which stacks the neurons layer by layer in a feedforward manner (Fig. 1a), where the neurons across adjacent layers are fully connected to each other. The first layer of the FNN is the input layer that each unit receives one dimension of the data vector. The last layer is the output layer that outputs the probabilities that a subject belonging to different classes (in classification). The layers between the input and output layers are the hidden layers. A DFNN usually contains multiple hidden layers. As shown in Fig. 2a, there is a weight parameter associated with each edge in the DFNN, which needs to be optimized by minimizing some training loss measured on a specific training dataset (usually through backpropagation17). After the optimal set of parameters are learned, the DFNN can be used to predict the target value (e.g., class) of any testing data vectors. Therefore, a DFNN can be viewed as an end-to-end process that transforms a specific raw data vector to its target layer by layer. Compared with the traditional ML models, DFNN has shown superior performance in many data mining tasks and have been introduced to the analysis of clinical data and genetic data to predict mental health conditions. We will discuss the applications of these methods further in the Results section.
a An illustration of basic unit of neural networks, i.e., artificial neuron. Each input xi is associated with a weight wi. The weighted sum of all inputs Σwixi is fed to a nonlinear activation function f to generate the output yj of the j-th neuron, i.e., yj = f(Σwixi). b Illustrations of the widely used nonlinear activation function.
Recurrent neural network
RNNs were designed to analyze sequential data such as natural language, speech, and video. Given an input sequence, the RNN processes one element of the sequence at a time by feeding to a recurrent neuron. To encode the historical information along the sequence, each recurrent neuron receives the input element at the corresponding time point and the output of the neuron at previous time stamp, and the output will also be provided to the neuron at next time stamp (this is also where the term “recurrent” comes from). An example RNN architecture is shown in Fig. 1b where the input is a sequence of words (a sentence). The recurrence link (i.e., the edge linking different neurons) enables RNN to capture the latent semantic dependencies among words and the syntax of the sentence. In recent years, different variants of RNN, such as long short-term memory (LSTM)18 and gated recurrent unit19 have been proposed, and the main difference among these models is how the input is mapped to the output for the recurrent neuron. RNN models have demonstrated state-of-the-art performance in various applications, especially natural language processing (NLP; e.g., machine translation and text-based classification); hence, they hold great premise in processing clinical notes and social media posts to detect mental health conditions as discussed below.
Convolutional neural network
CNN is a specific type of deep neural network originally designed for image analysis15, where each pixel corresponds to a specific input dimension describing the image. Similar to a DFNN, CNN also maps these input image pixels to the corresponding target (e.g., image class) through layers of nonlinear transformations. Different from DFNN, where only fully connected layers are considered, there are typically three types of layers in a CNN: a convolution–activation layer, a pooling layer, and a fully connected layer (Fig. 1c). The convolution–activation layer first convolves the entire feature map obtained from previous layer with small two-dimensional convolution filters. The results from each convolution filter are activated through a nonlinear activation function in the same way as a DFNN. A pooling layer reduces the size of the feature map through sub-sampling. The fully connected layer is analogous to the hidden layer in a DFNN, where each neuron is connected to all neurons of the previous layer. The convolution–activation layer extracts locally invariant patterns from the feature maps. The pooling layer effectively reduces the feature dimensionality to avoid model overfitting. The fully connected layer explores the global feature interactions as in DFNNs. Different combinations of these three types of layers constitute different CNN architectures. Because of the various characteristics of images such as local self-similarity, compositionality, and translational and deformation invariance, CNN has demonstrated state-of-the-art performance in many computer vision tasks7. Hence, the CNN models are promising in processing clinical images and expression data (e.g., facial expression images) to detect mental health conditions. We will discuss the application of these methods in the Results section.
Autoencoder
Autoencoder is a special variant of the DFNN aimed at learning new (usually more compact) data representations that can optimally reconstruct the original data vectors16,20. An autoencoder typically consists of two components (Fig. 1d) as follows: (1) the encoder, which learns new representations (usually with reduced dimensionality) from the input data through a multi-layer FNN; and (2) the decoder, which is exactly the reverse of the encoder, reconstructs the data in their original space from the representations derived from the encoder. The parameters in the autoencoder are learned through minimizing the reconstruction loss. Autoencoder has demonstrated the capacity of extracting meaningful features from raw data without any supervision information. In the studies of mental health outcomes, the use of autoencoder has resulted in desirable improvement in analyzing clinical and expression image data, which will be detailed in the Results section.
Methods
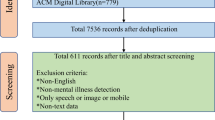
The processing and reporting of the results of this review were guided by the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines21. To thoroughly review the literature, a two-step method was used to retrieve all the studies on relevant topics. First, we conducted a search of the computerized bibliographic databases including PubMed and Web of Science. The search strategy is detailed in Supplementary Appendix 1. The literature search comprised articles published until April 2019. Next, a snowball technique was applied to identify additional studies. Furthermore, we manually searched other resources, including Google Scholar, and Institute of Electrical and Electronics Engineers (IEEE Xplore), to find additional relevant articles.
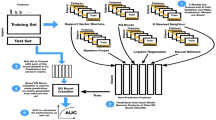
Figure 3 presents the study selection process. All articles were evaluated carefully and studies were excluded if: (1) the main outcome is not a mental health condition; (2) the model involved is not a DL algorithm; (3) full-text of the article is not accessible; and (4) the article is written not in English.
Results
A total of 57 articles met our eligibility criteria. Most of the reviewed articles were published between 2014 and 2019. To clearly summarize these articles, we grouped them into four categories according to the types of data analyzed, including (1) clinical data, (2) genetic and genomics data, (3) vocal and visual expression data, and (4) social media data. Table 1 summarizes the characteristics of these selected studies.
Clinical data
Neuroimages
Previous studies have shown that neuroimages can record evidence of neuropsychiatric disorders22,23. Two common types of neuroimage data analyzed in mental health studies are functional magnetic resonance imaging (fMRI) and structural MRI (sMRI) data. In fMRI data, the brain activity is measured by identification of the changes associated with blood flow, based on the fact that cerebral blood flow and neuronal activation are coupled24. In sMRI data, the neurological aspect of a brain is described based on the structural textures, which show some information in terms of the spatial arrangements of voxel intensities in 3D. Recently, DL technologies have been demonstrated in analyzing both fMRI and sMRI data.
One application of DL in fMRI and sMRI data is the identification of ADHD25,26,27,28,29,30,31. To learn meaningful information from the neuroimages, CNN and deep belief network (DBN) models were used. In particular, the CNN models were mainly used to identify local spatial patterns and DBN models were to obtain a deep hierarchical representation of the neuroimages. Different patterns were discovered between ADHDs and controls in the prefrontal cortex and cingulated cortex. Also, several studies analyzed sMRIs to investigate schizophrenia32,33,34,35,36, where DFNN, DBN, and autoencoder were utilized. These studies reported abnormal patterns of cortical regions and cortical–striatal–cerebellar circuit in the brain of schizophrenia patients, especially in the frontal, temporal, parietal, and insular cortices, and in some subcortical regions, including the corpus callosum, putamen, and cerebellum. Moreover, the use of DL in neuroimages also targeted at addressing other mental health disorders. Geng et al.37 proposed to use CNN and autoencoder to acquire meaningful features from the original time series of fMRI data for predicting depression. Two studies31,38 integrated the fMRI and sMRI data modalities to develop predictive models for ASDs. Significant relationships between fMRI and sMRI data were observed with regard to ASD prediction.
Challenges and opportunities
The aforementioned studies have demonstrated that the use of DL techniques in analyzing neuroimages can provide evidence in terms of mental health problems, which can be translated into clinical practice and facilitate the diagnosis of mental health illness. However, multiple challenges need to be addressed to achieve this objective. First, DL architectures generally require large data samples to train the models, which may pose a difficulty in neuroimaging analysis because of the lack of such data39. Second, typically the imaging data lie in a high-dimensional space, e.g., even a 64 × 64 2D neuroimage can result in 4096 features. This leads to the risk of overfitting by the DL models. To address this, most existing studies reported to utilize MRI data preprocessing tools such as Statistical Parametric Mapping (https://www.fil.ion.ucl.ac.uk/spm/), Data Processing Assistant for Resting-State fMRI40, and fMRI Preprocessing Pipeline41 to extract useful features before feeding to the DL models. Even though an intuitive attribute of DL is the capacity to learn meaningful features from raw data, feature engineering tools are needed especially in the case of small sample size and high-dimensionality, e.g., the neuroimage analysis. The use of such tools mitigates the overfitting risk of DL models. As reported in some selected studies28,31,35,37, the DL models can benefit from feature engineering techniques and have been shown to outperform the traditional ML models in the prediction of multiple conditions such as depression, schizophrenia, and ADHD. However, such tools extract features relying on prior knowledge; hence may omit some information that is meaningful for mental outcome research but unknown yet. An alternative way is to use CNN to automatically extract information from the raw data. As reported in the previous study10, CNNs perform well in processing raw neuroimage data. Among the studies reviewed in this study, three29,30,37 reported to involve CNN layers and achieved desirable performances.
Electroencephalogram data
As a low-cost, small-size, and high temporal resolution signal containing up to several hundred channels, analysis of electroencephalogram (EEG) data has gained significant attention to study brain disorders42. As the EEG signal is one kind of streaming data that presents a high density and continuous characteristics, it challenges traditional feature engineering-based methods to obtain sufficient information from the raw EEG data to make accurate predictions. To address this, recently the DL models have been employed to analyze raw EEG signal data.
Four articles reviewed proposed to use DL in understanding mental health conditions based on the analysis of EEG signals. Acharya et al.43 used CNN to extract features from the input EEG signals. They found that the EEG signals from the right hemisphere of the human brain are more distinctive in terms of the detection of depression than those from the left hemisphere. The findings provided shreds of evidence that depression is associated with a hyperactive right hemisphere. Mohan et al.44 modeled the raw EEG signals by DFNN to obtain information about the human brain waves. They found that the signals collected from the central (C3 and C4) regions are marginally higher compared with other brain regions, which can be used to distinguish the depressed and normal subjects from the brain wave signals. Zhang et al.45 proposed a concatenated structure of deep recurrent and 3D CNN to obtain EEG features across different tasks. They reported that the DL model can capture the spectral changes of EEG hemispheric asymmetry to distinguish different mental workload effectively. Li et al.46 presented a computer-aided detection system by extracting multiple types of information (e.g., spectral, spatial, and temporal information) to recognize mild depression based on CNN architecture. The authors found that both spectral and temporal information of EEG are crucial for prediction of depression.
Challenges and opportunities
EEG data are usually classified as streaming data that are continuous and are of high density. Despite the initial success in applying DL algorithms to analyze EEG data for studying multiple mental health conditions, there exist several challenges. One major challenge is that raw EEG data gathered from sensors have a certain degree of erroneous, noisy, and redundant information caused by discharged batteries, failures in sensor readings, and intermittent communication loss in wireless sensor networks47. This may challenge the model in extracting meaningful information from noise. Multiple preprocessing steps (e.g., data denoising, data interpolation, data transformation, and data segmentation) are necessary for dealing with the raw EEG signal before feeding to the DL models. Besides, due to the dense characteristics in the raw EEG data, analysis of the streaming data is computationally more expensive, which poses a challenge for the model architecture selection. A proper model should be designed relatively with less training parameters. This is one reason why the reviewed studies are mainly based on the CNN architecture.
Electronic health records
Electronic health records (EHRs) are systematic collections of longitudinal, patient-centered records. Patients’ EHRs consist of both structured and unstructured data: the structured data include information about a patient’s diagnosis, medications, and laboratory test results, and the unstructured data include information in clinical notes. Recently, DL models have been applied to analyze EHR data to study mental health disorders48.
The first and foremost issue for analyzing the structured EHR data is how to appropriately handle the longitudinal records. Traditional ML models address this by collapsing patients’ records within a certain time window into vectors, which comprised the summary of statistics of the features in different dimensions49. For instance, to estimate the probability of suicide deaths, Choi et al.50 leveraged a DFNN to model the baseline characteristics. One major limitation of these studies is the omittance of temporality among the clinical events within EHRs. To overcome this issue, RNNs are more commonly used for EHR data analysis as an RNN intuitively handles time-series data. DeepCare51, a long short-term memory network (LSTM)-based DL model, encodes patient’s long-term health state trajectories to predict the future outcomes of depressive episodes. As the LSTM architecture appropriately captures disease progression by modeling the illness history and the medical interventions, DeepCare achieved over 15% improvement in prediction, compared with the conventional ML methods. In addition, Lin et al.52 designed two DFNN models for the prediction of antidepressant treatment response and remission. The authors reported that the proposed DFNN can achieve an area under the receiver operating characteristic curve (AUC) of 0.823 in predicting antidepressant response.
Analyzing the unstructured clinical notes in EHRs refers to the long-standing topic of NLP. To extract meaningful knowledge from the text, conventional NLP approaches mostly define rules or regular expressions before the analysis. However, it is challenging to enumerate all possible rules or regular expressions. Due to the recent advance of DL in NLP tasks, DL models have been developed to mine clinical text data from EHRs to study mental health conditions. Geraci et al.53 utilized term frequency-inverse document frequency to represent the clinical documents by words and developed a DFNN model to identify individuals with depression. One major limitation of such an approach is that the semantics and syntax of sentences are lost. In this context, CNN54 and RNN55 have shown superiority in modeling syntax for text-based prediction. In particular, CNN has been used to mine the neuropsychiatric notes for predicting psychiatric symptom severity56,57. Tran and Kavuluru58 used an RNN to analyze the history of present illness in neuropsychiatric notes for predicting mental health conditions. The model engaged an attention mechanism55, which can specify the importance of the words in prediction, making the model more interpretable than their previous CNN model56.
Challenges and opportunities
Although DL has achieved promising results in EHR analysis, several challenges remain unsolved. On one hand, different from diagnosing physical health condition such as diabetes, the diagnosis of mental health conditions lacks direct quantitative tests, such as a blood chemistry test, a buccal swab, or urinalysis. Instead, the clinicians evaluate signs and symptoms through patient interviews and questionnaires during which they gather information based on patient’s self-report. Collection and deriving inferences from such data deeply relies on the experience and subjectivity of the clinician. This may account for signals buried in noise and affect the robustness of the DL model. To address this challenge, a potential way is to comprehensively integrate multimodal clinical information, including structured and unstructured EHR information, as well as neuroimaging and EEG data. Another way is to incorporate existing medical knowledge, which can guide model being trained in the right direction. For instance, the biomedical knowledge bases contain massive verified interactions between biomedical entities, e.g., diseases, genes, and drugs 59. Incorporating such information brings in meaningful medical constraints and may help to reduce the effects of noise on model training process. On the other hand, implementing a DL model trained from one EHR system into another system is challenging, because EHR data collection and representation is rarely standardized across hospitals and clinics. To address this issue, national/international collaborative efforts such as Observational Health Data Sciences and Informatics (https://ohdsi.org) have developed common data models, such as OMOP, to standardize EHR data representation for conducting observational data analysis60.
Genetic data
Multiple studies have found that mental disorders, e.g., depression, can be associated with genetic factors61,62. Conventional statistical studies in genetics and genomics, such as genome-wide association studies, have identified many common and rare genetic variants, such as single-nucleotide polymorphisms (SNPs), associated with mental health disorders63,64. Yet, the effect of the genetic factors is small and many more have not been discovered. With the recent developments in next-generation sequencing techniques, a massive volume of high-throughput genome or exome sequencing data are being generated, enabling researchers to study patients with mental health disorders by examining all types of genetic variations across an individual’s genome. In recent years, DL65,66 has been applied to identify genetic risk factors associated with mental illness, by borrowing the capacity of DL in identifying highly complex patterns in large datasets. Khan and Wang67 integrated genetic annotations, known brain expression quantitative trait locus, and enhancer/promoter peaks to generate feature vectors of variants, and developed a DFNN, named ncDeepBrain, to prioritized non-coding variants associated with mental disorders. To further prioritize susceptibility genes, they designed another deep model, iMEGES68, which integrates the ncDeepBrain score, general gene scores, and disease-specific scores for estimating gene risk. Wang et al.69 developed a novel deep architecture that combines deep Boltzmann machine architecture70 with conditional and lateral connections derived from the gene regulatory network. The model provided insights about intermediate phenotypes and their connections to high-level phenotypes (disease traits). Laksshman et al.71 used exome sequencing data to predict bipolar disorder outcomes of patients. They developed a CNN and used the convolution mechanism to capture correlations of the neighboring loci within the chromosome.
Challenges and opportunities
Although the use of genetic data in DL in studying mental health conditions shows promise, multiple challenges need to be addressed. For DL-based risk c/gene prioritization efforts, one major challenge is the limitation of labeled data. On one hand, the positive samples are limited, as known risk SNPs or genes associated with mental health conditions are limited. For example, there are about 108 risk loci that were genome-wide significant in ASD. On the other hand, the negative samples (i.e., SNPs, variants, or genes) may not be the “true” negative, as it is unclear whether they are associated with the mental illness yet. Moreover, it is also challenging to develop DL models for analyzing patient’s sequencing data for mental illness prediction, as the sequencing data are extremely high-dimensional (over five million SNPs in the human genome). More prior domain knowledge is needed to guide the DL model extracting patterns from the high-dimensional genomic space.
Vocal and visual expression data
The use of vocal (voice or speech) and visual (video or image of facial or body behaviors) expression data has gained the attention of many studies in mental health disorders. Modeling the evolution of people’s emotional states from these modalities has been used to identify mental health status. In essence, the voice data are continuous and dense signals, whereas the video data are sequences of frames, i.e., images. Conventional ML models for analyzing such types of data suffer from the sophisticated feature extraction process. Due to the recent success of applying DL in computer vision and sequence data modeling, such models have been introduced to analyze the vocal and/or visual expression data. In this work, most articles reviewed are to predict mental health disorders based on two public datasets: (i) the Chi-Mei corpus, collected by using six emotional videos to elicit facial expressions and speech responses of the subjects of bipolar disorder, unipolar depression, and healthy controls;72 and (ii) the International Audio/Visual Emotion Recognition Challenges (AVEC) depression dataset73,74,75, collected within human–computer interaction scenario. The proposed models include CNNs, RNNs, autoencoders, as well as hybrid models based on the above ones. In particular, CNNs were leveraged to encode the temporal and spectral features from the voice signals76,77,78,79,80 and static facial or physical expression features from the video frames79,81,82,83,84. Autoencoders were used to learn low-dimensional representations for people’s vocal85,86 and visual expression87,88, and RNNs were engaged to characterize the temporal evolution of emotion based on the CNN-learned features and/or other handcraft features76,81,84,85,86,87,88,89,90. Few studies focused on analyzing static images using a CNN architecture to predict mental health status. Prasetio et al.91 identified the stress types (e.g., neutral, low stress, and high stress) from facial frontal images. Their proposed CNN model outperforms the conventional ML models by 7% in terms of prediction accuracy. Jaiswal et al.92 investigated the relationship between facial expression/gestures and neurodevelopmental conditions. They reported accuracy over 0.93 in the diagnostic prediction of ADHD and ASD by using the CNN architecture. In addition, thermal images that track persons’ breathing patterns were also fed to a deep model to estimate psychological stress level (mental overload)93.
Challenges and opportunities
From the above summary, we can observe that analyzing vocal and visual expression data can capture the pattern of subjects’ emotion evolution to predict mental health conditions. Despite the promising initial results, there remain challenges for developing DL models in this field. One major challenge is to link vocal and visual expression data with the clinical data of patients, given the difficulties involved in collecting such expression data during clinical practice. Current studies analyzed vocal and visual expression over individual datasets. Without clinical guidance, the developed prediction models have limited clinical meanings. Linking patients’ expression information with clinical variables may help to improve both the interpretability and robustness of the model. For example, Gupta et al.94 designed a DFNN for affective prediction from audio and video modalities. The model incorporated depression severity as the parameter, linking the effects of depression on subjects’ affective expressions. Another challenge is the limitation of the samples. For example, the Chi-Mei dataset contains vocal–visual data from only 45 individuals (15 with bipolar disorder, 15 with unipolar disorder, and 15 healthy controls). Also, there is a lack of “emotion labels” for people’s vocal and visual expression. Apart from improving the datasets, an alternative way to solve this challenge is to use transfer learning, which transfers knowledge gained with one dataset (usually more general) to the target dataset. For example, some studies trained autoencoder in public emotion database such as eNTERFACE95 to generate emotion profiles (EPs). Other studies83,84 pre-trained CNN over general facial expression datasets96,97 for extracting face appearance features.
Social media data
With the widespread proliferation of social media platforms, such as Twitter and Reddit, individuals are increasingly and publicly sharing information about their mood, behavior, and any ailments one might be suffering. Such social media data have been used to identify users’ mental health state (e.g., psychological stress and suicidal ideation)6.
In this study, the articles that used DL to analyze social media data mainly focused on stress detection98,99,100,101, depression identification102,103,104,105,106, and estimation of suicide risk103,105,107,108,109. In general, the core concept across these work is to mine the textual, and where applicable graphical, content of users’ social media posts to discover cues for mental health disorders. In this context, the RNN and CNN were largely used by the researchers. Especially, RNN usually introduces an attention mechanism to specify the importance of the input elements in the classification process55. This provides some interpretability for the predictive results. For example, Ive et al.103 proposed a hierarchical RNN architecture with an attention mechanism to predict the classes of the posts (including depression, autism, suicidewatch, anxiety, etc.). The authors observed that, benefitting from the attention mechanism, the model can predict risk text efficiently and extract text elements crucial for making decisions. Coppersmith et al.107 used LSTM to discover quantifiable signals about suicide attempts based on social media posts. The proposed model can capture contextual information between words and obtain nuances of language related to suicide.
Apart from text, users also post images on social media. The properties of the images (e.g., color theme, saturation, and brightness) provide some cues reflecting users’ mental health status. In addition, millions of interactions and relationships among users can reflect the social environment of individuals that is also a kind of risk factors for mental illness. An increasing number of studies attempted to combine these two types of information with text content for predictive modeling. For example, Lin et al.99 leveraged the autoencoder to extract low-level and middle-level representations from texts, images, and comments based on psychological and art theories. They further extended their work with a hybrid model based on CNN by integrating post content and social interactions101. The results provided an implication that the social structure of the stressed users’ friends tended to be less connected than that of the users without stress.
Challenges and opportunities
The aforementioned studies have demonstrated that using social media data has the potential to detect users with mental health problems. However, there are multiple challenges towards the analysis of social media data. First, given that social media data are typically de-identified, there is no straightforward way to confirm the “true positives” and “true negatives” for a given mental health condition. Enabling the linkage of user’s social media data with their EHR data—with appropriate consent and privacy protection—is challenging to scale, but has been done in a few settings110. In addition, most of the previous studies mainly analyzed textual and image data from social media platforms, and did not consider analyzing the social network of users. In one study, Rosenquist et al.111 reported that the symptoms of depression are highly correlated inside the circle of friends, indicating that social network analysis is likely to be a potential way to study the prevalence of mental health problems. However, comprehensively modeling text information and network structure remains challenging. In this context, graph convolutional networks112 have been developed to address networked data mining. Moreover, although it is possible to discover online users with mental illness by social media analysis, translation of this innovation into practical applications and offer aid to users, such as providing real-time interventions, are largely needed113.
Discussion: findings, open issues, and future directions
Principle findings
The purpose of this study is to investigate the current state of applications of DL techniques in studying mental health outcomes. Out of 2261 articles identified based on our search terms, 57 studies met our inclusion criteria and were reviewed. Some studies that involved DL models but did not highlight the DL algorithms’ features on analysis were excluded. From the above results, we observed that there are a growing number of studies using DL models for studying mental health outcomes. Particularly, multiple studies have developed disease risk prediction models using both clinical and non-clinical data, and have achieved promising initial results.
Data bias
DL models “think to learn” like a human brain relying on their multiple layers of interconnected computing neurons. Therefore, to train a deep neural network, there are multiple parameters (i.e., weights associated links between neurons within the network) being required to learn. This is one reason why DL has achieved great success in the fields where a massive volume of data can be easily collected, such as computer vision and text mining. Yet, in the health domain, the availability of large-scale data is very limited. For most selected studies in this review, the sample sizes are under a scale of 104. Data availability is even more scarce in the fields of neuroimaging, EEG, and gene expression data, as such data reside in a very high-dimensional space. This then leads to the problem of “curse of dimensionality”114, which challenges the optimization of the model parameters.
One potential way to address this challenge is to reduce the dimensionality of the data by feature engineering before feeding information to the DL models. On one hand, feature extraction approaches can be used to obtain different types of features from the raw data. For example, several studies reported in this review have attempted to use preprocessing tools to extract features from neuroimaging data. On the other hand, feature selection that is commonly used in conventional ML models is also an option to reduce data dimensionality. However, the feature selection approaches are not often used in the DL application scenario, as one of the intuitive attributes of DL is the capacity to learn meaningful features from “all” available data. The alternative way to address the issue of data bias is to use transfer learning where the objective is to improve learning a new task through the transfer of knowledge from a related task that has already been learned115. The basic idea is that data representations learned in the earlier layers are more general, whereas those learned in the latter layers are more specific to the prediction task116. In particular, one can first pre-train a deep neural network in a large-scale “source” dataset, then stack fully connected layers on the top of the network and fine-tune it in the small “target” dataset in a standard backpropagation manner. Usually, samples in the “source” dataset are more general (e.g., general image data), whereas those in the “target” dataset are specific to the task (e.g., medical image data). A popular example of the success of transfer learning in the health domain is the dermatologist-level classification of skin cancer117. The authors introduced Google’s Inception v3 CNN architecture pre-trained over 1.28 million general images and fine-tuned in the clinical image dataset. The model achieved very high-performance results of skin cancer classification in epidermal (AUC = 0.96), melanocytic (AUC = 0.96), and melanocytic–dermoscopic images (AUC = 0.94). In facial expression-based depression prediction, Zhu et al.83 pre-trained CNN on the public face recognition dataset to model the static facial appearance, which overcomes the issue that there is no facial expression label information. Chao et al.84 also pre-trained CNN to encode facial expression information. The transfer scheme of both of the two studies has been demonstrated to be able to improve the prediction performance.
Diagnosis and prediction issues
Unlike the diagnosis of physical conditions that can be based on lab tests, diagnoses of the mental illness typically rely on mental health professionals’ judgment and patient self-report data. As a result, such a diagnostic system may not accurately capture the psychological deficits and symptom progression to provide appropriate therapeutic interventions118,119. This issue accordingly accounts for the limitation of the prediction models to assist clinicians to make decisions. Except for several studies using the unsupervised autoencoder for learning low-dimensional representations, most studies reviewed in this study reported using supervised DL models, which need the training set containing “true” (i.e., expert provided) labels to optimize the model parameters before the model being used to predict labels of new subjects. Inevitably, the quality of the expert-provided diagnostic labels used for training sets the upper-bound for the prediction performance of the model.
One intuitive route to address this issue is to use an unsupervised learning scheme that, instead of learning to predict clinical outcomes, aims at learning compacted yet informative representations of the raw data. A typical example is the autoencoder (as shown in Fig. 1d), which encodes the raw data into a low-dimensional space, from which the raw data can be reconstructed. Some studies reviewed have proposed to leverage autoencoder to improve our understanding of mental health outcomes. A constraint of the autoencoder is that the input data should be preprocessed to vectors, which may lead to information loss for image and sequence data. To address this, recently convolutional-autoencoder120 and LSTM-autoencoder121 have been developed, which integrate the convolution layers and recurrent layers with the autoencoder architecture and enable us to learn informative low-dimensional representations from the raw image data and sequence data, respectively. For instance, Baytas et al.122 developed a variation of LSTM-autoencoder on patient EHRs and grouped Parkinson’s disease patients into meaningful subtypes. Another potential way is to predict other clinical outcomes instead of the diagnostic labels. For example, several selected studies proposed to predict symptom severity scores56,57,77,82,84,87,89. In addition, Du et al.108 attempted to identify suicide-related psychiatric stressors from users’ posts on Twitter, which plays an important role in the early prevention of suicidal behaviors. Furthermore, training model to predict future outcomes such as treatment response, emotion assessments, and relapse time is also a promising future direction.
Multimodal modeling
The field of mental health is heterogeneous. On one hand, mental illness refers to a variety of disorders that affect people’s emotions and behaviors. On the other hand, though the exact causes of most mental illnesses are unknown to date, it is becoming increasingly clear that the risk factors for these diseases are multifactorial as multiple genetic, environmental, and social factors interact to influence an individual’s mental health123,124. As a result of domain heterogeneity, researchers have the chance to study the mental health problems from different perspectives, from molecular, genomic, clinical, medical imaging, physiological signal to facial, and body expressive and online behavioral. Integrative modeling of such multimodal data means comprehensively considering different aspects of the disease, thus likely obtaining deep insight into mental health. In this context, DL models have been developed for multimodal modeling. As shown in Fig. 4, the hierarchical structure of DL makes it easily compatible with multimodal integration. In particular, one can model each modality with a specific network and combine them by the final fully connected layers, such that parameters can be jointly learned by a typical backpropagation manner. In this review, we found an increasing number of studies have attempted to use multimodal modeling. For example, Zou et al.28 developed a multimodal model composed of two CNNs for modeling fMRI and sMRI modalities, respectively. The model achieved 69.15% accuracy in predicting ADHD, which outperformed the unimodal models (66.04% for fMRI modal-based and 65.86% for sMRI modal-based). Yang et al.79 proposed a multimodal model to combine vocal and visual expression for depression cognition. The model results in 39% lower prediction error than the unimodal models.
Model interpretability
Due to the end-to-end design, the DL models usually appear to be “black boxes”: they take raw data (e.g., MRI images, free-text of clinical notes, and EEG signals) as input, and yield output to reach a conclusion (e.g., the risk of a mental health disorder) without clear explanations of their inner working. Although this might not be an issue in other application domains such as identifying animals from images, in health not only the model’s prediction performance but also the clues for making the decision are important. For example in the neuroimage-based depression identification, despite estimation of the probability that a patient suffers from mental health deficits, the clinicians would focus more on recognizing abnormal regions or patterns of the brain associated with the disease. This is really important for convincing the clinical experts about the actions recommended from the predictive model, as well as for guiding appropriate interventions. In addition, as discussed above, the introduction of multimodal modeling leads to an increased challenge in making the models more interpretable. Attempts have been made to open the “black box” of DL59,125,126,127. Currently, there are two general directions for interpretable modeling: one is to involve the systematic modification of the input and the measure of any resulting changes in the output, as well as in the activation of the artificial neurons in the hidden layers. Such a strategy is usually used in CNN in identifying specific regions of an image being captured by a convolutional layer128. Another way is to derive tools to determine the contribution of one or more features of the input data to the output. In this case, the widely used tools include Shapley Additive Explanation129, LIME127, DeepLIFT130, etc., which are able to assign each feature an importance score for the specific prediction task.
Connection to therapeutic interventions
According to the studies reviewed, it is now possible to detect patients with mental illness based on different types of data. Compared with the traditional ML techniques, most of the reviewed DL models reported higher prediction accuracy. The findings suggested that the DL models are likely to assist clinicians in improved diagnosis of mental health conditions. However, to associate diagnosis of a condition with evidence-based interventions and treatment, including identification of appropriate medication131, prediction of treatment response52, and estimation of relapse risk132 still remains a challenge. Among the reviewed studies, only one52 proposed to target at addressing these issues. Thus, further efforts are needed to link the DL techniques with the therapeutic intervention of mental illness.
Domain knowledge
Another important direction is to incorporate domain knowledge. The existing biomedical knowledge bases are invaluable sources for solving healthcare problems133,134. Incorporating domain knowledge could address the limitation of data volume, problems of data quality, as well as model generalizability. For example, the unified medical language system135 can help to identify medical entities from the text and gene–gene interaction databases136 could help to identify meaningful patterns from genomic profiles.
Conclusion
Recent years have witnessed the increasing use of DL algorithms in healthcare and medicine. In this study, we reviewed existing studies on DL applications to study mental health outcomes. All the results available in the literature reviewed in this work illustrate the applicability and promise of DL in improving the diagnosis and treatment of patients with mental health conditions. Also, this review highlights multiple existing challenges in making DL algorithms clinically actionable for routine care, as well as promising future directions in this field.
References
World Health Organization. The World Health Report 2001: Mental Health: New Understanding, New Hope (World Health Organization, Switzerland, 2001).
Marcus, M., Yasamy, M. T., van Ommeren, M., Chisholm, D. & Saxena, S. Depression: A Global Public Health Concern (World Federation of Mental Health, World Health Organisation, Perth, 2012).
Hamilton, M. Development of a rating scale for primary depressive illness. Br. J. Soc. Clin. Psychol. 6, 278–296 (1967).
Dwyer, D. B., Falkai, P. & Koutsouleris, N. Machine learning approaches for clinical psychology and psychiatry. Annu. Rev. Clin. Psychol. 14, 91–118 (2018).
Lovejoy, C. A., Buch, V. & Maruthappu, M. Technology and mental health: the role of artificial intelligence. Eur. Psychiatry 55, 1–3 (2019).
Wongkoblap, A., Vadillo, M. A. & Curcin, V. Researching mental health disorders in the era of social media: systematic review. J. Med. Internet Res. 19, e228 (2017).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436 (2015).
Miotto, R., Wang, F., Wang, S., Jiang, X. & Dudley, J. T. Deep learning for healthcare: review, opportunities and challenges. Brief. Bioinformatics 19, 1236–1246 (2017).
Durstewitz, D., Koppe, G. & Meyer-Lindenberg, A. Deep neural networks in psychiatry. Mol. Psychiatry 24, 1583–1598 (2019).
Vieira, S., Pinaya, W. H. & Mechelli, A. Using deep learning to investigate the neuroimaging correlates of psychiatric and neurological disorders: methods and applications. Neurosci. Biobehav. Rev. 74, 58–75 (2017).
Shatte, A. B., Hutchinson, D. M. & Teague, S. J. Machine learning in mental health: a scoping review of methods and applications. Psychol. Med. 49, 1426–1448 (2019).
Murphy, K. P. Machine Learning: A Probabilistic Perspective (MIT Press, Cambridge, 2012).
Biship, C. M. Pattern Recognition and Machine Learning (Information Science and Statistics) (Springer-Verlag, Berlin, 2007).
Bengio, Y., Simard, P. & Frasconi, P. Learning long-term dependencies with gradient descent is difficult. IEEE Trans. Neural Netw. Learn. Syst. 5, 157–166 (1994).
LeCun, Y., Bottou, L., Bengio, Y. & Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 86, 2278–2324 (1998).
Vincent, P., Larochelle, H., Lajoie, I., Bengio, Y. & Manzagol, P. A. Stacked denoising autoencoders: learning useful representations in a deep network with a local denoising criterion. J. Mach. Learn. Res. 11, 3371–3408 (2010).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Cogn. modeling. 5, 1 (1988).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Cho, K., Van Merriënboer, B., Bahdanau, D. & Bengio, Y. On the properties of neural machine translation: encoder-decoder approaches. In Proc. SSST-8, Eighth Workshop on Syntax, Semantics and Structure in Statistical Translation 103–111 (Doha, Qatar, 2014).
Liou, C., Cheng, W., Liou, J. & Liou, D. Autoencoder for words. Neurocomputing 139, 84–96 (2014).
Moher, D., Liberati, A., Tetzlaff, J. & Altman, D. G. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Ann. Intern. Med. 151, 264–269 (2009).
Schnack, H. G. et al. Can structural MRI aid in clinical classification? A machine learning study in two independent samples of patients with schizophrenia, bipolar disorder and healthy subjects. Neuroimage 84, 299–306 (2014).
O’Toole, A. J. et al. Theoretical, statistical, and practical perspectives on pattern-based classification approaches to the analysis of functional neuroimaging data. J. Cogn. Neurosci. 19, 1735–1752 (2007).
Logothetis, N. K., Pauls, J., Augath, M., Trinath, T. & Oeltermann, A. Neurophysiological investigation of the basis of the fMRI signal. Nature 412, 150 (2001).
Kuang, D. & He, L. Classification on ADHD with deep learning. In Proc. Int. Conference on Cloud Computing and Big Data 27–32 (Wuhan, China, 2014).
Kuang, D., Guo, X., An, X., Zhao, Y. & He, L. Discrimination of ADHD based on fMRI data with deep belief network. In Proc. Int. Conference on Intelligent Computing 225–232 (Taiyuan, China, 2014).
Farzi, S., Kianian, S. & Rastkhadive, I. Diagnosis of attention deficit hyperactivity disorder using deep belief network based on greedy approach. In Proc. 5th Int. Symposium on Computational and Business Intelligence 96–99 (Dubai, United Arab Emirates, 2017).
Zou, L., Zheng, J. & McKeown, M. J. Deep learning based automatic diagnoses of attention deficit hyperactive disorder. In Proc. 2017 IEEE Global Conference on Signal and Information Processing (GlobalSIP) 962–966 (Montreal, Canada, 2017).
Riaz A. et al. Deep fMRI: an end-to-end deep network for classification of fMRI data. In Proc. 2018 IEEE 15th Int. Symposium on Biomedical Imaging. 1419–1422 (Washington, DC, USA, 2018).
Zou, L., Zheng, J., Miao, C., Mckeown, M. J. & Wang, Z. J. 3D CNN based automatic diagnosis of attention deficit hyperactivity disorder using functional and structural MRI. IEEE Access. 5, 23626–23636 (2017).
Sen, B., Borle, N. C., Greiner, R. & Brown, M. R. A general prediction model for the detection of ADHD and Autism using structural and functional MRI. PLoS ONE 13, e0194856 (2018).
Zeng, L. et al. Multi-site diagnostic classification of schizophrenia using discriminant deep learning with functional connectivity MRI. EBioMedicine 30, 74–85 (2018).
Pinaya, W. H. et al. Using deep belief network modelling to characterize differences in brain morphometry in schizophrenia. Sci. Rep. 6, 38897 (2016).
Pinaya, W. H., Mechelli, A. & Sato, J. R. Using deep autoencoders to identify abnormal brain structural patterns in neuropsychiatric disorders: a large-scale multi-sample study. Hum. Brain Mapp. 40, 944–954 (2019).
Ulloa, A., Plis, S., Erhardt, E. & Calhoun, V. Synthetic structural magnetic resonance image generator improves deep learning prediction of schizophrenia. In Proc. 25th IEEE Int. Workshop on Machine Learning for Signal Processing (MLSP) 1–6 (Boston, MA, USA, 2015).
Matsubara, T., Tashiro, T. & Uehara, K. Deep neural generative model of functional MRI images for psychiatric disorder diagnosis. IEEE Trans. Biomed. Eng. 99 (2019).
Geng, X. & Xu, J. Application of autoencoder in depression diagnosis. In 2017 3rd Int. Conference on Computer Science and Mechanical Automation (Wuhan, China, 2017).
Aghdam, M. A., Sharifi, A. & Pedram, M. M. Combination of rs-fMRI and sMRI data to discriminate autism spectrum disorders in young children using deep belief network. J. Digit. Imaging 31, 895–903 (2018).
Shen, D., Wu, G. & Suk, H. -I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 19, 221–248 (2017).
Yan, C. & Zang, Y. DPARSF: a MATLAB toolbox for “pipeline” data analysis of resting-state fMRI. Front. Syst. Neurosci. 4, 13 (2010).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Herrmann, C. & Demiralp, T. Human EEG gamma oscillations in neuropsychiatric disorders. Clin. Neurophysiol. 116, 2719–2733 (2005).
Acharya, U. R. et al. Automated EEG-based screening of depression using deep convolutional neural network. Comput. Meth. Prog. Biol. 161, 103–113 (2018).
Mohan, Y., Chee, S. S., Xin, D. K. P. & Foong, L. P. Artificial neural network for classification of depressive and normal. In EEG Proc. 2016 IEEE EMBS Conference on Biomedical Engineering and Sciences 286–290 (Kuala Lumpur, Malaysia, 2016).
Zhang, P., Wang, X., Zhang, W. & Chen, J. Learning spatial–spectral–temporal EEG features with recurrent 3D convolutional neural networks for cross-task mental workload assessment. IEEE Trans. Neural Syst. Rehabil. Eng. 27, 31–42 (2018).
Li, X. et al. EEG-based mild depression recognition using convolutional neural network. Med. Biol. Eng. Comput. 47, 1341–1352 (2019).
Patel, S., Park, H., Bonato, P., Chan, L. & Rodgers, M. A review of wearable sensors and systems with application in rehabilitation. J. Neuroeng. Rehabil. 9, 21 (2012).
Smoller, J. W. The use of electronic health records for psychiatric phenotyping and genomics. Am. J. Med. Genet. B Neuropsychiatr. Genet. 177, 601–612 (2018).
Wu, J., Roy, J. & Stewart, W. F. Prediction modeling using EHR data: challenges, strategies, and a comparison of machine learning approaches. Med. Care. 48, S106–S113 (2010).
Choi, S. B., Lee, W., Yoon, J. H., Won, J. U. & Kim, D. W. Ten-year prediction of suicide death using Cox regression and machine learning in a nationwide retrospective cohort study in South Korea. J. Affect. Disord. 231, 8–14 (2018).
Pham, T., Tran, T., Phung, D. & Venkatesh, S. Predicting healthcare trajectories from medical records: a deep learning approach. J. Biomed. Inform. 69, 218–229 (2017).
Lin, E. et al. A deep learning approach for predicting antidepressant response in major depression using clinical and genetic biomarkers. Front. Psychiatry 9, 290 (2018).
Geraci, J. et al. Applying deep neural networks to unstructured text notes in electronic medical records for phenotyping youth depression. Evid. Based Ment. Health 20, 83–87 (2017).
Kim, Y. Convolutional neural networks for sentence classification. arXiv Prepr. arXiv 1408, 5882 (2014).
Yang, Z. et al. Hierarchical attention networks for document classification. In Proc. 2016 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies 1480–1489 (San Diego, California, USA, 2016).
Rios, A. & Kavuluru, R. Ordinal convolutional neural networks for predicting RDoC positive valence psychiatric symptom severity scores. J. Biomed. Inform. 75, S85–S93 (2017).
Dai, H. & Jonnagaddala, J. Assessing the severity of positive valence symptoms in initial psychiatric evaluation records: Should we use convolutional neural networks? PLoS ONE 13, e0204493 (2018).
Tran, T. & Kavuluru, R. Predicting mental conditions based on “history of present illness” in psychiatric notes with deep neural networks. J. Biomed. Inform. 75, S138–S148 (2017).
Samek, W., Binder, A., Montavon, G., Lapuschkin, S. & Müller, K.-R. Evaluating the visualization of what a deep neural network has learned. IEEE Trans. Neural Netw. Learn. Syst. 28, 2660–2673 (2016).
Hripcsak, G. et al. Characterizing treatment pathways at scale using the OHDSI network. Proc. Natl. Acad. Sci. USA 113, 7329–7336 (2016).
McGuffin, P., Owen, M. J. & Gottesman, I. I. Psychiatric Genetics and Genomics (Oxford Univ. Press, New York, 2004).
Levinson, D. F. The genetics of depression: a review. Biol. Psychiatry 60, 84–92 (2006).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668 (2018).
Mullins, N. & Lewis, C. M. Genetics of depression: progress at last. Curr. Psychiatry Rep. 19, 43 (2017).
Zou, J. et al. A primer on deep learning in genomics. Nat. Genet. 51, 12–18 (2019).
Yue, T. & Wang, H. Deep learning for genomics: a concise overview. Preprint at arXiv:1802.00810 (2018).
Khan, A. & Wang, K. A deep learning based scoring system for prioritizing susceptibility variants for mental disorders. In Proc. 2017 IEEE Int. Conference on Bioinformatics and Biomedicine (BIBM) 1698–1705 (Kansas City, USA, 2017).
Khan, A., Liu, Q. & Wang, K. iMEGES: integrated mental-disorder genome score by deep neural network for prioritizing the susceptibility genes for mental disorders in personal genomes. BMC Bioinformatics 19, 501 (2018).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Salakhutdinov, R. & Hinton, G. Deep Boltzmann machines. In Proc. 12th Int. Conference on Artificial Intelligence and Statistics 448–455 (Clearwater, Florida, USA, 2009).
Laksshman, S., Bhat, R. R., Viswanath, V. & Li, X. DeepBipolar: Identifying genomic mutations for bipolar disorder via deep learning. Hum. Mutat. 38, 1217–1224 (2017).
Huang, K.-Y. et al. Data collection of elicited facial expressions and speech responses for mood disorder detection. In Proc. 2015 Int. Conference on Orange Technologies (ICOT) 42–45 (Hong Kong, China, 2015).
Valstar, M. et al. AVEC 2013: the continuous audio/visual emotion and depression recognition challenge. In Proc. 3rd ACM Int. Workshop on Audio/Visual Emotion Challenge 3–10 (Barcelona, Spain, 2013).
Valstar, M. et al. Avec 2014: 3d dimensional affect and depression recognition challenge. In Proc. 4th Int. Workshop on Audio/Visual Emotion Challenge 3–10 (Orlando, Florida, USA, 2014).
Valstar, M. et al. Avec 2016: depression, mood, and emotion recognition workshop and challenge. In Proc. 6th Int. Workshop on Audio/Visual Emotion Challenge 3–10 (Amsterdam, The Netherlands, 2016).
Ma, X., Yang, H., Chen, Q., Huang, D. & Wang, Y. Depaudionet: an efficient deep model for audio based depression classification. In Proc. 6th Int. Workshop on Audio/Visual Emotion Challenge 35–42 (Amsterdam, The Netherlands, 2016).
He, L. & Cao, C. Automated depression analysis using convolutional neural networks from speech. J. Biomed. Inform. 83, 103–111 (2018).
Li, J., Fu, X., Shao, Z. & Shang, Y. Improvement on speech depression recognition based on deep networks. In Proc. 2018 Chinese Automation Congress (CAC) 2705–2709 (Xi’an, China, 2018).
Yang, L., Jiang, D., Han, W. & Sahli, H. DCNN and DNN based multi-modal depression recognition. In Proc. 2017 7th Int. Conference on Affective Computing and Intelligent Interaction 484–489 (San Antonio, Texas, USA, 2017).
Huang, K. Y., Wu, C. H. & Su, M. H. Attention-based convolutional neural network and long short-term memory for short-term detection of mood disorders based on elicited speech responses. Pattern Recogn. 88, 668–678 (2019).
Dawood, A., Turner, S. & Perepa, P. Affective computational model to extract natural affective states of students with Asperger syndrome (AS) in computer-based learning environment. IEEE Access. 6, 67026–67034 (2018).
Song, S., Shen, L. & Valstar, M. Human behaviour-based automatic depression analysis using hand-crafted statistics and deep learned spectral features. In Proc. 13th IEEE Int. Conference on Automatic Face & Gesture Recognition 158–165 (Xi’an, China, 2018).
Zhu, Y., Shang, Y., Shao, Z. & Guo, G. Automated depression diagnosis based on deep networks to encode facial appearance and dynamics. IEEE Trans. Affect. Comput. 9, 578–584 (2018).
Chao, L., Tao, J., Yang, M. & Li, Y. Multi task sequence learning for depression scale prediction from video. In Proc. 2015 Int. Conference on Affective Computing and Intelligent Interaction (ACII) 526–531 (Xi’an, China, 2015).
Yang, T. H., Wu, C. H., Huang, K. Y. & Su, M. H. Detection of mood disorder using speech emotion profiles and LSTM. In Proc. 10th Int. Symposium on Chinese Spoken Language Processing (ISCSLP) 1–5 (Tianjin, China, 2016).
Huang, K. Y., Wu, C. H., Su, M. H. & Chou, C. H. Mood disorder identification using deep bottleneck features of elicited speech. In Proc. 2017 Asia-Pacific Signal and Information Processing Association Annual Summit and Conference (APSIPA ASC) 1648–1652 (Kuala Lumpur, Malaysia, 2017).
Jan, A., Meng, H., Gaus, Y. F. B. A. & Zhang, F. Artificial intelligent system for automatic depression level analysis through visual and vocal expressions. IEEE Trans. Cogn. Dev. Syst. 10, 668–680 (2017).
Su, M. H., Wu, C. H., Huang, K. Y. & Yang, T. H. Cell-coupled long short-term memory with l-skip fusion mechanism for mood disorder detection through elicited audiovisual features. IEEE Trans. Neural Netw. Learn. Syst. 31 (2019).
Harati, S., Crowell, A., Mayberg, H. & Nemati, S. Depression severity classification from speech emotion. In Proc. 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 5763–5766 (Honolulu, HI, USA, 2018).
Su, M. H., Wu, C. H., Huang, K. Y., Hong, Q. B. & Wang, H. M. Exploring microscopic fluctuation of facial expression for mood disorder classification. In Proc. 2017 Int. Conference on Orange Technologies (ICOT) 65–69 (Singapore, 2017).
Prasetio, B. H., Tamura, H. & Tanno, K. The facial stress recognition based on multi-histogram features and convolutional neural network. In Proc. 2018 IEEE Int. Conference on Systems, Man, and Cybernetics (SMC) 881–887 (Miyazaki, Japan, 2018).
Jaiswal, S., Valstar, M. F., Gillott, A. & Daley, D. Automatic detection of ADHD and ASD from expressive behaviour in RGBD data. In Proc. 12th IEEE Int. Conference on Automatic Face & Gesture Recognition 762–769 (Washington, DC, USA, 2017).
Cho, Y., Bianchi-Berthouze, N. & Julier, S. J. DeepBreath: deep learning of breathing patterns for automatic stress recognition using low-cost thermal imaging in unconstrained settings. In Proc. 2017 7th Int. Conference on Affective Computing and Intelligent Interaction (ACII) 456–463 (San Antonio, Texas, USA, 2017).
Gupta, R., Sahu, S., Espy-Wilson, C. Y. & Narayanan, S. S. An affect prediction approach through depression severity parameter incorporation in neural networks. In Proc. 2017 Int. Conference on INTERSPEECH 3122–3126 (Stockholm, Sweden, 2017).
Martin, O., Kotsia, I., Macq, B. & Pitas, I. The eNTERFACE'05 audio-visual emotion database. In Proc. 22nd Int. Conference on Data Engineering Workshops 8–8 (Atlanta, GA, USA, 2006).
Goodfellow, I. J. et al. Challenges in representation learning: A report on three machine learning contests. In Proc. Int. Conference on Neural Information Processing 117–124 (Daegu, Korea, 2013).
Yi, D., Lei, Z., Liao, S. & Li, S. Z.. Learning face representation from scratch. Preprint at arXiv 1411.7923 (2014).
Lin, H. et al. User-level psychological stress detection from social media using deep neural network. In Proc. 22nd ACM Int. Conference on Multimedia 507–516 (Orlando, Florida, USA, 2014).
Lin, H. et al. Psychological stress detection from cross-media microblog data using deep sparse neural network. In Proc. 2014 IEEE Int. Conference on Multimedia and Expo 1–6 (Chengdu, China, 2014).
Li, Q. et al. Correlating stressor events for social network based adolescent stress prediction. In Proc. Int. Conference on Database Systems for Advanced Applications 642–658 (Suzhou, China, 2017).
Lin, H. et al. Detecting stress based on social interactions in social networks. IEEE Trans. Knowl. Data En. 29, 1820–1833 (2017).
Cong, Q. et al. X-A-BiLSTM: a deep learning approach for depression detection in imbalanced data. In Proc. 2018 IEEE Int. Conference on Bioinformatics and Biomedicine (BIBM) 1624–1627 (Madrid, Spain, 2018).
Ive, J., Gkotsis, G., Dutta, R., Stewart, R. & Velupillai, S. Hierarchical neural model with attention mechanisms for the classification of social media text related to mental health. In Proc. Fifth Workshop on Computational Linguistics and Clinical Psychology: From Keyboard to Clinic 69–77 (New Orleans, Los Angeles, USA, 2018).
Sadeque, F., Xu, D. & Bethard, S. UArizona at the CLEF eRisk 2017 pilot task: linear and recurrent models for early depression detection. CEUR Workshop Proc. 1866 (2017).
Fraga, B. S., da Silva, A. P. C. & Murai, F. Online social networks in health care: a study of mental disorders on Reddit. In Proc. 2018 IEEE/WIC/ACM Int. Conference on Web Intelligence (WI) 568–573 (Santiago, Chile, 2018).
Gkotsis, G. et al. Characterisation of mental health conditions in social media using Informed Deep Learning. Sci. Rep. 7, 45141 (2017).
Coppersmith, G., Leary, R., Crutchley, P. & Fine, A. Natural language processing of social media as screening for suicide risk. Biomed. Inform. Insights 10, 1178222618792860 (2018).
Du, J. et al. Extracting psychiatric stressors for suicide from social media using deep learning. BMC Med. Inform. Decis. Mak. 18, 43 (2018).
Alambo, A. et al. Question answering for suicide risk assessment using Reddit. In Proc. IEEE 13th Int. Conference on Semantic Computing 468–473 (Newport Beach, California, USA, 2019).
Eichstaedt, J. C. et al. Facebook language predicts depression in medical records. Proc. Natl Acad. Sci. USA 115, 11203–11208 (2018).
Rosenquist, J. N., Fowler, J. H. & Christakis, N. A. Social network determinants of depression. Mol. Psychiatry 16, 273 (2011).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. In Proc. 2017 Int. Conference on Learning Representations (Toulon, France, 2017).
Rice, S. M. et al. Online and social networking interventions for the treatment of depression in young people: a systematic review. J. Med. Internet Res. 16, e206 (2014).
Hastie, T., Tibshirani, R. & Friedman, J. The elements of statistical learning: data mining, inference, and prediction. Springer Series in Statistics. Math. Intell. 27, 83–85 (2009).
Torrey, L. & Shavlik, J. in Handbook of Research on Machine Learning Applications and Trends: Algorithms, Methods, and Techniques 242–264 (IGI Global, 2010).
Yosinski, J., Clune, J., Bengio, Y. & Lipson, H. How transferable are features in deep neural networks? In Proc. Advances in Neural Information Processing Systems 3320–3328 (Montreal, Canada, 2014).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115 (2017).
Insel, T. et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am. Psychiatr. Assoc. 167, 748–751 (2010).
Nelson, B., McGorry, P. D., Wichers, M., Wigman, J. T. & Hartmann, J. A. Moving from static to dynamic models of the onset of mental disorder: a review. JAMA Psychiatry 74, 528–534 (2017).
Guo, X., Liu, X., Zhu, E. & Yin, J. Deep clustering with convolutional autoencoders. In Proc. Int. Conference on Neural Information Processing 373–382 (Guangzhou, China, 2017).
Srivastava, N., Mansimov, E. & Salakhudinov, R. Unsupervised learning of video representations using LSTMs. In Proc. Int. Conference on Machine Learning 843–852 (Lille, France, 2015).
Baytas, I. M. et al. Patient subtyping via time-aware LSTM networks. In Proc. 23rd ACM SIGKDD Int. Conference on Knowledge Discovery and Data Mining 65–74 (Halifax, Canada, 2017).
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-5®) (American Psychiatric Pub, Washington, DC, 2013).
Biological Sciences Curriculum Study. In: NIH Curriculum Supplement Series (Internet) (National Institutes of Health, USA, 2007).
Noh, H., Hong, S. & Han, B. Learning deconvolution network for semantic segmentation. In Proc. IEEE Int. Conference on Computer Vision 1520–1528 (Santiago, Chile, 2015).
Grün, F., Rupprecht, C., Navab, N. & Tombari, F. A taxonomy and library for visualizing learned features in convolutional neural networks. In Proc. 33rd Int. Conference on Machine Learning (ICML) Workshop on Visualization for Deep Learning (New York, USA, 2016).
Ribeiro, M. T., Singh, S. & Guestrin, C. Why should I trust you?: Explaining the predictions of any classifier. In Proc. 22nd ACM SIGKDD Int. Conference on Knowledge Discovery and Data Mining 1135–1144 (San Francisco, CA, 2016).
Zhang, Q. S. & Zhu, S. C. Visual interpretability for deep learning: a survey. Front. Inf. Technol. Electron. Eng. 19, 27–39 (2018).
Lundberg, S. M. & Lee, S. I. A unified approach to interpreting model predictions. In Proc. 31st Conference on Neural Information Processing Systems 4765–4774 (Long Beach, CA, 2017).
Shrikumar, A., Greenside, P., Shcherbina, A. & Kundaje, A. Not just a black box: learning important features through propagating activation differences. In Proc. 33rd Int. Conference on Machine Learning (New York, NY, 2016).
Gawehn, E., Hiss, J. A. & Schneider, G. Deep learning in drug discovery. Mol. Inform. 35, 3–14 (2016).
Jerez-Aragonés, J. M., Gómez-Ruiz, J. A., Ramos-Jiménez, G., Muñoz-Pérez, J. & Alba-Conejo, E. A combined neural network and decision trees model for prognosis of breast cancer relapse. Artif. Intell. Med. 27, 45–63 (2003).
Zhu, Y., Elemento, O., Pathak, J. & Wang, F. Drug knowledge bases and their applications in biomedical informatics research. Brief. Bioinformatics 20, 1308–1321 (2018).
Su, C., Tong, J., Zhu, Y., Cui, P. & Wang, F. Network embedding in biomedical data science. Brief. Bioinform. https://doi.org/10.1093/bib/bby117 (2018).
Bodenreider, O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32(suppl_1), D267–D270 (2004).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2014).
Acknowledgements
The work is supported by NSF 1750326, R01 MH112148, R01 MH105384, R01 MH119177, R01 MH121922, and P50 MH113838.
Author information
Authors and Affiliations
Contributions
C.S., Z.X. and F.W. planned and structured the whole paper. C.S. and Z.X. conducted the literature review and drafted the manuscript. J.P. and F.W. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Su, C., Xu, Z., Pathak, J. et al. Deep learning in mental health outcome research: a scoping review. Transl Psychiatry 10, 116 (2020). https://doi.org/10.1038/s41398-020-0780-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-0780-3
This article is cited by
-
Automated mood disorder symptoms monitoring from multivariate time-series sensory data: getting the full picture beyond a single number
Translational Psychiatry (2024)
-
Detecting your depression with your smartphone? – An ethical analysis of epistemic injustice in passive self-tracking apps
Ethics and Information Technology (2024)
-
Development of intelligent system based on synthesis of affective signals and deep neural networks to foster mental health of the Indian virtual community
Social Network Analysis and Mining (2024)
-
Prevalence and predictors of self-rated mental health among farm and non-farm adult rural residents of Saskatchewan
Current Psychology (2024)
-
Unraveling minds in the digital era: a review on mapping mental health disorders through machine learning techniques using online social media
Social Network Analysis and Mining (2024)