Abstract
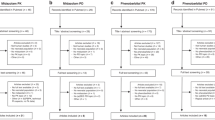
Variable responses to medications complicates perioperative care. As a potential solution, we evaluated and synthesized pharmacogenomic evidence that may inform anesthesia and pain prescribing to identify clinically actionable drug/gene pairs. Clinical decision-support (CDS) summaries were developed and were evaluated using Appraisal of Guidelines for Research and Evaluation (AGREE) II. We found that 93/180 (51%) of commonly-used perioperative medications had some published pharmacogenomic information, with 18 having actionable evidence: celecoxib/diclofenac/flurbiprofen/ibuprofen/piroxicam/CYP2C9, codeine/oxycodone/tramadol CYP2D6, desflurane/enflurane/halothane/isoflurane/sevoflurane/succinylcholine/RYR1/CACNA1S, diazepam/CYP2C19, phenytoin/CYP2C9, succinylcholine/mivacurium/BCHE, and morphine/OPRM1. Novel CDS summaries were developed for these 18 medications. AGREE II mean ± standard deviation scores were high for Scope and Purpose (95.0 ± 2.8), Rigor of Development (93.2 ± 2.8), Clarity of Presentation (87.3 ± 3.0), and Applicability (86.5 ± 3.7) (maximum score = 100). Overall mean guideline quality score was 6.7 ± 0.2 (maximum score = 7). All summaries were recommended for clinical implementation. A critical mass of pharmacogenomic evidence exists for select medications commonly used in the perioperative setting, warranting prospective examination for clinical utility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zheng SL, Sun J, Wiklund F, Gao Z, Stattin P, Purcell LD, et al. Genetic variants and family history predict prostate cancer similar to prostate-specific antigen. Clin Cancer Res. 2009;15:1105–11.
Nanji KC, Patel A, Shaikh S, Seger DL, Bates DW. Evaluation of perioperative medication errors and adverse drug events. Anesthesiology. 2016;124:25–34.
Weiss AJ, Elixhauser A, Bae J, Encinosa W. Origin of adverse drug events in U.S. Hospitals, 2011: Statistical Brief #158. Rockville (MD): Healthcare Cost and Utilization Project (HCUP) Statistical Briefs; 2006.
Wu CL, Raja SN. Treatment of acute postoperative pain. Lancet. 2011;377:2215–25.
Mavridou P, Dimitriou V, Manataki A, Arnaoutoglou E, Papadopoulos G. Patient’s anxiety and fear of anesthesia: effect of gender, age, education, and previous experience of anesthesia. A survey of 400 patients. J Anesth. 2013;27:104–8.
Denborough MA. Malignant hyperthermia. 1962. Anesthesiology. 2008;108:156–7.
Sangkuhl K, Dirksen RT, Alvarellos ML, Altman RB, Klein TE. PharmGKB summary: very important pharmacogene information for CACNA1S. Pharmacogenet Genom. 2020;30:34–44.
Relling MV, Evans WE. Pharmacogenomics in the clinic. Nature. 2015;526:343–50.
Kalow W. Pharmacogenetics and anesthesia. J Am Soc Anesthesiol. 1964;25:377–87.
Hopkins PM, Ruffert H, Snoeck MM, Girard T, Glahn KP, Ellis FR, et al. European malignant hyperthermia group guidelines for investigation of malignant hyperthermia susceptibility. Br J Anaesth. 2015;115:531–9.
Relling MV, Klein TE. CPIC: clinical pharmacogenetics implementation consortium of the pharmacogenomics research network. Clin Pharmacol Ther. 2011;89:464–7.
van der Wouden CH, Cambon-Thomsen A, Cecchin E, Cheung KC, Davila-Fajardo CL, Deneer VH, et al. Implementing pharmacogenomics in europe: design and implementation strategy of the ubiquitous pharmacogenomics consortium. Clin Pharmacol Ther. 2017;101:341–58.
Whirl-Carrillo M, McDonagh EM, Hebert JM, Gong L, Sangkuhl K, Thorn CF, et al. Pharmacogenomics knowledge for personalized medicine. Clin Pharmacol Ther. 2012;92:414–7.
Swen JJ, Wilting I, de Goede AL, Grandia L, Mulder H, Touw DJ, et al. Pharmacogenetics: from bench to byte. Clin Pharmacol Ther. 2008;83:781–7.
Borden BA, Galecki P, Wellmann R, Danahey K, Lee SM, Patrick-Miller L, et al. Assessment of provider-perceived barriers to clinical use of pharmacogenomics during participation in an institutional implementation study. Pharmacogenet Genom. 2019;29:31–8.
McKinnon RA, Ward MB, Sorich MJ. A critical analysis of barriers to the clinical implementation of pharmacogenomics. Ther Clin Risk Manag. 2007;3:751–9.
Wellmann R, Borden BA, Danahey K, Nanda R, Polite BN, Stadler WM, et al. Analyzing the clinical actionability of germline pharmacogenomic findings in oncology. Cancer. 2018;124:3052–65.
Kaufman AL, Spitz J, Jacobs M, Sorrentino M, Yuen S, Danahey K, et al. Evidence for clinical implementation of pharmacogenomics in cardiac drugs. Mayo Clin Proc. 2015;90:716–29.
Hussain S, Kenigsberg BB, Danahey K, Lee YM, Galecki PM, Ratain MJ, et al. Disease-drug database for pharmacogenomic-based prescribing. Clin Pharmacol Ther. 2016;100:179–90.
Danahey K, Borden BA, Furner B, Yukman P, Hussain S, Saner D, et al. Simplifying the use of pharmacogenomics in clinical practice: building the genomic prescribing system. J Biomed Inform. 2017;75:110–21.
Ratain MJ, Nakamura Y, Cox NJ. CYP2D6 genotype and tamoxifen activity: understanding interstudy variability in methodological quality. Clin Pharmacol Ther. 2013;94:185–7.
Thorn CF, Whirl-Carrillo M, Hachad H, Johnson JA, McDonagh EM, Ratain MJ, et al. Essential characteristics of pharmacogenomics study publications. Clin Pharmacol Ther. 2019;105:86–91.
Pratt VM, Del Tredici AL, Hachad H, Ji Y, Kalman LV, Scott SA, et al. Recommendations for clinical CYP2C19 genotyping allele selection: a report of the association for molecular pathology. J Mol Diagn. 2018;20:269–76.
O’Donnell PH, Bush A, Spitz J, Danahey K, Saner D, Das S, et al. The 1200 patients project: creating a new medical model system for clinical implementation of pharmacogenomics. Clin Pharmacol Ther. 2012;92:446–9.
O’Donnell PH, Danahey K, Jacobs M, Wadhwa NR, Yuen S, Bush A, et al. Adoption of a clinical pharmacogenomics implementation program during outpatient care-initial results of the University of Chicago “1,200 Patients Project”. Am J Med Genet C Semin Med Genet. 2014;166C:68–75.
O’Donnell PH, Wadhwa N, Danahey K, Borden BA, Lee SM, Hall JP, et al. Pharmacogenomics-based point-of-care clinical decision support significantly alters drug prescribing. Clin Pharmacol Ther. 2017;102:859–69.
Brouwers MC, Kho ME, Browman GP, Burgers JS, Cluzeau F, Feder G, et al. AGREE II: advancing guideline development, reporting and evaluation in health care. CMAJ. 2010;182:E839–42.
Collaboration A. Development and validation of an international appraisal instrument for assessing the quality of clinical practice guidelines: the AGREE project. Qual Saf Health Care. 2003;12:18–23.
Brouwers MC, Kho ME, Browman GP, Burgers JS, Cluzeau F, Feder G, et al. Development of the AGREE II, part 1: performance, usefulness and areas for improvement. CMAJ. 2010;182:1045–52.
Brouwers MC, Kho ME, Browman GP, Burgers JS, Cluzeau F, Feder G, et al. Development of the AGREE II, part 2: assessment of validity of items and tools to support application. CMAJ. 2010;182:E472–8.
Vlayen J, Aertgeerts B, Hannes K, Sermeus W, Ramaekers D. A systematic review of appraisal tools for clinical practice guidelines: multiple similarities and one common deficit. Int J Qual Health Care. 2005;17:235–42.
Truong TM, Apfelbaum J, Shahul S, Anitescu M, Danahey K, Knoebel RW, et al. The ImPreSS trial: implementation of point-of-care pharmacogenomic decision support in perioperative care. Clin Pharmacol Ther. 2019;106:1179–83.
Chidambaran V, Ngamprasertwong P, Vinks AA, Sadhasivam S. Pharmacogenetics and anesthetic drugs. Curr Clin Pharmacol. 2012;7:78–101.
Zhou S, Skaar DJ, Jacobson PA, Huang RS. Pharmacogenomics of medications commonly used in the intensive care unit. Front Pharmacol. 2018;9:1436.
Lam YW. Scientific challenges and implementation barriers to translation of pharmacogenomics in clinical practice. ISRN Pharmacol. 2013;2013:641089.
Johnson JA. Pharmacogenetics in clinical practice: how far have we come and where are we going? Pharmacogenomics 2013;14:835–43.
Criteria for Genetic TestingMalignant Hyperthermia Association of the United States. Available at: www.mhaus.org. Accessed November 30, 2019.
Henricks LM, Lunenburg C, de Man FM, Meulendijks D, Frederix GWJ, Kienhuis E, et al. DPYD genotype-guided dose individualisation of fluoropyrimidine therapy in patients with cancer: a prospective safety analysis. Lancet Oncol. 2018;19:1459–67.
Bank PCD, Swen JJ, Schaap RD, Klootwijk DB, Baak-Pablo R, Guchelaar HJ. A pilot study of the implementation of pharmacogenomic pharmacist initiated pre-emptive testing in primary care. Eur J Hum Genet. 2019;27:1532–41.
Greden JF, Parikh SV, Rothschild AJ, Thase ME, Dunlop BW, DeBattista C, et al. Impact of pharmacogenomics on clinical outcomes in major depressive disorder in the GUIDED trial: a large, patient- and rater-blinded, randomized, controlled study. J Psychiatr Res. 2019;111:59–67.
Smith DM, Weitzel KW, Elsey AR, Langaee T, Gong Y, Wake DT, et al. CYP2D6-guided opioid therapy improves pain control in CYP2D6 intermediate and poor metabolizers: a pragmatic clinical trial. Genet Med. 2019;21:1842–50.
Ignite: Implementing Genomics in Clinical Practice.Genomic Medicine Knowledge Base. www.gmkb.org. Accessed 30 Nov 2019.
Theken KN, Lee CR, Gong L, Caudle KE, Formea CM, Gaedigk A, et al. Clinical pharmacogenetics implementation consortium guideline (CPIC) for CYP2C9 and nonsteroidal anti-inflammatory drugs. Clin Pharmacol Ther. 2020;108:191–200.
Food and Drug Administration. Drug Label: Mivacurium. 2015. https://www.accessdata.fda.gov/drugsatfda_docs/label/2015/020098s018lbl.pdf.
Food and Drug Administration. Drug Label: Succinylcholine. 2010. https://www.accessdata.fda.gov/drugsatfda_docs/label/2010/008453s027lbl.pdf.
Clinical Pharmacogenetics Implementation Consortium. Genes-Drugs. 2020. https://cpicpgx.org/genes-drugs/.
Crews KR, Monte AA, Huddart R, Caudle KE, Kharasch ED, Gaedigk A, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) guideline for CYP2D6, OPRM1, and COMT genotype and select opioid therapy. Clin Pharmacol Ther. 2021. https://doi.org/10.1002/cpt.2149.
Altman RB. Pharmacogenomics: “noninferiority” is sufficient for initial implementation. Clin Pharmacol Ther. 2011;89:348–50.
Stamer UM, Zhang L, Book M, Lehmann LE, Stuber F, Musshoff F. CYP2D6 genotype dependent oxycodone metabolism in postoperative patients. PLoS ONE. 2013;8:e60239.
Samer CF, Daali Y, Wagner M, Hopfgartner G, Eap CB, Rebsamen MC, et al. Genetic polymorphisms and drug interactions modulating CYP2D6 and CYP3A activities have a major effect on oxycodone analgesic efficacy and safety. Br J Pharmacol. 2010;160:919–30.
Samer CF, Daali Y, Wagner M, Hopfgartner G, Eap CB, Rebsamen MC, et al. The effects of CYP2D6 and CYP3A activities on the pharmacokinetics of immediate release oxycodone. Br J Pharmacol. 2010;160:907–18.
Zwisler ST, Enggaard TP, Noehr-Jensen L, Pedersen RS, Mikkelsen S, Nielsen F, et al. The hypoalgesic effect of oxycodone in human experimental pain models in relation to the CYP2D6 oxidation polymorphism. Basic Clin Pharmacol Toxicol. 2009;104:335–44.
Zwisler ST, Enggaard TP, Mikkelsen S, Brosen K, Sindrup SH. Impact of the CYP2D6 genotype on post-operative intravenous oxycodone analgesia. Acta Anaesthesiol Scand. 2010;54:232–40.
Andreassen TN, Eftedal I, Klepstad P, Davies A, Bjordal K, Lundstrom S, et al. Do CYP2D6 genotypes reflect oxycodone requirements for cancer patients treated for cancer pain? A cross-sectional multicentre study. Eur J Clin Pharmacol. 2012;68:55–64.
Duncan LE, Ostacher M, Ballon J. How genome-wide association studies (GWAS) made traditional candidate gene studies obsolete. Neuropsychopharmacology. 2019;44:1518–23.
Hendricks-Sturrup RM, Linsky A, Lu CY, Vassy JL. Genomic testing is best integrated into clinical practice when it is actionable. Per Med. 2020;17:5–8.
Funding
This work was supported by National Institutes of Health (NIH)/NIGMS 5T32GM007019-41 (for EHJ and TMT as trainees), NIH/NHGRI 1R01HG009938-01A1 (PHO), by an Innovations Grant from the University of Chicago Medicine Office of Clinical Effectiveness (PHO), and the Benjamin McAllister Research Fellowship (TMT).
Author information
Authors and Affiliations
Contributions
BAB: data acquisition, data analysis, and drafting of paper. EHJ: study conception and design, data acquisition, data analysis, and drafting of paper. KD: data acquisition, data analysis. ES: data acquisition, data analysis. JLA: data analysis. MA: data analysis. RK: data analysis. SS: data analysis. TMT: data analysis. MJR: study conception and design. PHO: study conception and design, data acquisition, data analysis, and drafting of paper.
Corresponding author
Ethics declarations
Competing interests
EHJ is currently an employee of OneOme. MJR is a co-inventor holding patents related to pharmacogenetic diagnostics and receives royalties related to UGT1A1 genotyping outside of this work. All other authors declared no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Borden, B.A., Jhun, E.H., Danahey, K. et al. Appraisal and development of evidence-based clinical decision support to enable perioperative pharmacogenomic application. Pharmacogenomics J 21, 691–711 (2021). https://doi.org/10.1038/s41397-021-00248-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-021-00248-2